Abstract
The insulin‐like growth factor‐1 receptor (IGF‐1R) is a receptor tyrosine kinase (RTK) that plays critical roles in cancer. Microarray, computational, thermodynamic, and cellular imaging studies reveal that activation of IGF‐1R by its cognate ligand IGF1 is inhibited by shorter, soluble heparan sulfate (HS) sequences (e.g., HS06), whereas longer polymeric chains do not inhibit the RTK, a phenomenon directly opposed to the traditional relationship known for GAG‐protein systems. The inhibition arises from smaller oligosaccharides binding in a unique pocket in the IGF‐1R ectodomain, which competes with the natural cognate ligand IGF1. This work presents a highly interesting observation on preferential and competing inhibition of IGF‐1R by smaller sequences, whereas polysaccharides are devoid of this function. These insights will be of major value to glycobiologists and anti‐cancer drug discoverers.
Keywords: Drug Discovery, Glycosaminoglycans, Growth Factors, Heparan Sulfate, Receptor Tyrosine Kinases
The work provides a new insight into insulin‐like growth factor‐1 receptor (IGF‐1R), which is preferentially inhibited by smaller heparan sulfate chains, e.g., HS06, in contrast to longer polysaccharidic chains, e.g., HS36, which are devoid of the inhibition function.

Introduction
The insulin‐like growth factor‐1 receptor (IGF‐1R) drives IGF1‐initiated physiologic processes such as growth and development.[ 1 , 2 ] Recent studies on multiple stem cells, including cancer stem‐like calls (CSCs), indicate that IGF‐1R is critical for stem cell self‐renewal and pluripotency.[ 3 , 4 ] In fact, IGF‐1R plays key roles in aberrant cellular growth, tumor invasion, and metastasis, and contributes to the development of breast, colorectal, lung and liver cancers.[ 5 , 6 , 7 , 8 , 9 ]
IGF‐1R belongs to the family of cell surface proteins called receptor tyrosine kinases (RTKs). Yet, unlike other RTKs that are monomers, IGF‐1R is a constitutive dimer, [10] which introduces differences in the mechanism of receptor activation. Whereas, some RTKs, e.g., ErbB and PDGFR[ 11 , 12 ] rely on their ligands and/or co‐receptors to dimerize, and others, e.g., FGFR, VEGFR and c‐MET use surface‐bound proteoglycans in addition to their cognate ligands, for receptor dimerization,[ 13 , 14 , 15 , 16 , 17 ] IGF‐1R does not need dimerization because it is a constitutive dimer. In fact, the inter‐domain interactions between monomers in the native site maintain IGF‐1R in an auto‐inhibitory dimeric state (Figure 1A). [10] Upon IGF1 binding, a major conformational change is initiated in the extracellular domain that brings the two transmembrane domains together (Figure 1B), thereby unleashing cross‐phosphorylation of the intra‐cellular tyrosine kinase domains. [18]
Figure 1.
A) Structural model of IGF‐1R. Each monomeric ectodomain consists of L1, L2, CR, Fn1, Fn2, and Fn3 domains that assemble into a unique Π‐shape formed by two inverted “J‐shaped” monomer chains running opposite to each other. B) Binding of one IGF1 molecule initiates a conformational change that brings the two TM domains in close proximity resulting in activation. C) Structures of 24 HS oligosaccharides printed on the microarray. See underneath for notation of residues. D) Plot showing quantitative fluorescence intensities for each HS oligosaccharide of the microarray. Error bars are ±1 SE. (E and F) Spectrofluorimetry‐based studies of IGF‐1R binding to HS06 (E) and HS12 (F) in the presence of varying concentrations of IGF1 (10–80 nM). The fluorescence signal from AlexFluorTM labeled IGF‐1R was used to monitor the interactions. G) Shows the variation in K D,obs as a function of IGF1 concentration for IGF‐1R complexes with HS06 (circles) and HS12 (squares). Solid lines represent linear fits. H) describes the equilibria based on these studies. A IGF‐1R‐HS06‐IGF1 ternary complex is unlikely to form because HS06 and IGF1 compete with each other for IGF‐1R, while HS12 does not compete with IGF1 for binding to IGF‐1R.
A number of RTKs are modulated by cell surface or extracellular “niche” proteoglycans (PGs).[ 13 , 19 ] Whereas cell surface PGs may serve as co‐receptors in modulating receptor activation, niche PGs including their glycosaminoglycan (GAG) chains, may serve as reservoirs for release of cognate growth factors that induce receptor activation. [20] Considering that IGF‐1R biology is strongly dependent on the state of cell surface and extracellular environment,[ 21 , 22 , 23 , 24 ] we reasoned that GAGs may play a role in IGF‐1R function. Yet, GAG modulation of IGF‐1R is unknown.
We hypothesized that GAGs may bind to IGF‐1R and modulate its activation. In this work, we find that the IGF‐1R‐IGF1 system is exquisitely regulated by heparan sulfate (HS) oligosaccharides. More specifically, we find that smaller oligosaccharides, e.g., HS06, bind potently and preferentially to the extracellular domain (ECD) of IGF‐1R, whereas longer polysaccharides, e.g., HS36, bind weakly. This phenomenon defies the traditional well‐established relationships for GAG‐protein systems wherein affinity increases monotonously with the length of the GAG chain. [25] Further, smaller oligosaccharides, e.g., HS06, compete with the cognate ligand IGF1 for binding to IGF‐1R. This is a fundamentally different role of a GAG oligosaccharide as compared to that known for other RTK‐cognate ligand systems wherein the GAG role is cooperative, and not competitive. At a functional level, HS06 inhibits the IGF1‐dependent activation of IGF‐1R. Finally, our mutagenesis studies indicate that HS06 binds in a unique pocket at the interface of the L1 and CR domains of IGF‐1R. These unique structural and functional insights bring forth a new phenomenon of glycan regulation of IGF‐1R for the first time.
Results and Discussion
Human IGF‐1R Surface Electrostatics Supports GAG‐Binding Property
We first analyzed hIGF‐1R's primary sequence for the presence of a heparin‐binding Cardin‐Weintraub sequence and found three plausible regions (Figure S1). [26] Three‐dimensional surface electrostatics, calculated as described in our earlier works,[ 27 , 28 ] found an extended area of contiguous electropositivity lining the L1 and CR domains (Figure S2). Such extended basic regions are fairly common in GAG‐binding RTKs, e.g., FGFR, VEGFR, and c‐MET (Figure S3), which implied a high probability of hIGF‐1R binding favorably with GAGs. Yet, IGF‐1R ECD displayed a different electropositive landscape as compared to other RTKs. Specifically, hIGF‐1R ECD presented several electropositive regions scattered in its L1, CR and L2 domains that may potentially engage GAGs.
hIGF‐1R Binds to HS Oligosaccharides
To quickly assess the possibility of GAG binding, we utilized a commercial microarray of 24 HS sequences. The 24 sequences ranged from tetra‐ to nona‐ mer in length; carried as many as nine sulfate groups; and were primarily based on glucosamine (GlcN)‐glucuronic acid (GlcA) building blocks with few iduronic acid residues (IdoA) (Figure 1C). Exposing the array to His‐tagged IGF‐1R followed by detection using an antibody against the HIS‐tag led to a characteristic recognition profile (Figure 1D). Of the 24 sequences, IGF‐1R preferentially recognized sequences XX–XXIV, which are sulfated hexa‐ and octa‐ saccharides. These microarray results presented the first evidence that this dimeric receptor binds directly to some HS oligosaccharides.
hIGF‐1R Preferentially Binds to Smaller Oligosaccharides, e.g., HS06 (hexasaccharide) with High Affinity
To better elucidate the nature of HS recognition by IGF‐1R, we performed thermodynamic fluorimetry‐based affinity studies. Titrations of the C‐terminal fluorophore‐labeled ECD against full‐length hIGF1 (see Methods) at pH 7.4 displayed a saturable response, which yielded an affinity of 90±20 nM (Figure S4). This was only 2‐fold lower than that measured using isothermal titration calorimetry for an IGF‐1R construct devoid of nearly 20 amino acid residues, [10] thereby validating the commercial ECD of hIGF‐1R.
We then screened a library of HS oligosaccharides, including di‐, tetra‐, and hexa‐ saccharides against hIGF‐1R. Whereas the di‐ and tetra‐ saccharides bound the receptor rather poorly (K D 20–1000 μM), the hexasaccharides, i.e., HS06 (Figure S5), bound with an affinity of 190 nM (Figure 1E), which was similar to that of the IGF‐1R‐IGF1 complex. To ascertain that the fluorescence labeling of IGF‐1R does not impact HS06 binding, we used intrinsic fluorescence of unlabeled IGF‐1R and found the affinity to be 113 nM (Figure S6). This indicated that the IGF‐1R‐HS06 interaction is a high affinity interaction.
To evaluate the selectivity of recognition, we measured affinities of CS06 and DS06, two sequences of identical chain length from the chondroitin sulfate (CS) and dermatan sulfate (DS) class of GAGs (Figure S5), respectively. CS06 and DS06 displayed a loss of 5–6‐fold in affinity (Figure S7, Table S1), which implied that IGF‐1R preferentially recognized HS06.
To assess the chain length dependence of HS interaction, we studied HS12, which is nearly twice as long as HS06. In contrast to our expectations based on first principles, HS12 did not display higher affinity. In fact, the IGF‐1R‐HS12 interaction displayed a significantly lower affinity of 830 nM (Figure 1, Table S1). Measurements using intrinsic fluorescence of unlabeled IGF‐1R yielded a K D of 565 nM (Figure S6).
We then studied HS36 and unfractionated heparin (UFH), which are >6‐fold longer than HS06, and found the affinities to be more than 3‐fold lower (Table S1). Whereas UFH is highly sulfated, our preparation of HS36 did not contain 3‐O‐ and 2‐O‐ sulfate groups. Irrespective of these changes, a polymeric ligand carrying multiple sites of binding, e.g., HS12, HS36, or UFH v/s HS06, is expected to display higher affinity because of a large statistical advantage. [25] This has been a fundamental expectation of nearly all GAG‐protein systems, especially for GAG‐growth factor interactions.[ 29 , 30 ] The result that smaller oligosaccharides, e.g., HS06, bind better than longer polysaccharides was against known rules established for GAG—protein systems.
Smaller Oligosaccharides, e.g., HS06, Compete with IGF1 for IGF‐1R
To assess whether HS can modulate the interaction of IGF‐1R with its cognate ligand IGF1, we first evaluated whether HS oligosaccharides can bind to hIGF1. Surface electrostatics predicted that hIGF1 will bind to sulfated oligosaccharides (Figure S8). In fact, fluorophore‐labeled HS06 and HS12 yielded affinities of 1.5 and 1.1 μM (Figure S9, Table S1), respectively, which are much weaker in comparison that for IGF‐1R. Hence, we studied their competition with IGF1 for binding to IGF‐1R. We chose IGF1 levels such that IGF1‐HS complex interference would be minimal. Figures 1E and F shows profiles for HS06 and HS12, respectively, in the presence of IGF1. Figure 1G, compiling the change in measured affinities, shows that the two oligosaccharides displayed strikingly different profiles. Whereas the HS06 affinity increased monotonously with increasing IGF1, that for HS12 remained rather invariant. This suggests that the presence of IGF1 impedes HS06 from binding to IGF‐1R, whereas HS12‐IGF‐1R interaction remains unaffected by the cognate ligand. These spectrofluorimetric studies lead to a rather unique structure‐function relationship (Figure 1H). The studies indicate that a ternary complex with the smaller oligosaccharide, i.e., IGF‐1R‐HS06‐IGF1, is unlikely to form because of competition between HS06 and IGF1 for IGF‐1R. In striking contrast, the longer oligosaccharide HS12 does not compete with IGF1 for IGF‐1R.
To more directly assess whether HS06 and HS12 bind to different sites on IGF‐1R, we prepared rhodamine‐labeled HS06 (HS06*) and HS12 (HS12*) and studied cross‐competition with unlabeled HS12 and HS06, respectively. Whereas IGF‐1R‐HS06* co‐complex showed an increase in fluorescence in the presence of unlabeled HS12, practically no change in fluorescence was observed for IGF‐1R‐HS12* co‐complex upon introduction of unlabeled HS06 (Figure S11). Alternatively, unlabeled HS12 impacts the fluorescence of HS06*, while unlabeled HS06 does not impact the fluorescence of HS12*. Such a phenomenon is only possible because their sites of binding on IGF‐1R are different. To the best of our knowledge, such a phenomenon is unique among GAG‐protein systems.
Smaller Oligosaccharides (HS06), but Not Longer Polysaccharides (HS36), Bind Preferentially to Cell Surface IGF‐1R and Compete with IGF1
Cell surface IGF‐1R, typically upregulated on cancer spheroids,[ 3 , 31 ] may be influenced by its interaction with soluble ligands, e.g., GAGs. Earlier, we have shown that IGF‐1R is overexpressed in HT‐29 cells grown as spheroids and also regulates CSC self‐renewal. [4] To understand whether HS modulates cell surface IGF‐1R, we chose HS06 and HS36 as representative smaller oligosaccharide and longer polysaccharide, respectively.
Both HS06 and HS36 were covalently tagged at the non‐reducing end with an AlexaFluor (AF) dye for visualization and imaging (see Methods). Imaging of HT‐29 spheroids treated with the two HS species showed a significant dose‐dependent binding compared to the dye control (Figure S12). Further, both preparations bound HT‐29 spheroids fast with signals reaching optimal intensity within 20 minutes (Figure S13). Studies with an alternative AF dye gave similar results (Figure S14). Likewise, studies using HCT‐116, another colon cancer cell line, grown as spheroids also yielded results similar to HT‐29 suggesting the generality of HS binding to colorectal spheroids (Figure S15).
To assess the selectivity of binding, we generated an HT‐29 cell line in which IGF‐1R expression was knocked down (KD) >70 % using shRNA technology, i.e., IGF‐1R KD, as assessed by QPCR (Figure S16), also described in our prior work. [4] An equivalent cell line generated with scrambled shRNA served as the control. As expected, fluorophore‐labeled HS06 bound to scrambled HT‐29 spheroids; however, spheroids with IGF‐1R KD showed minimal HS06 binding (Figure 2A). On the contrary, the binding of the polysaccharides (HS36) was independent of the expression of IGF‐1R levels (Figure 2B). To further validate the observed selectivity, we studied whether three distinct anti‐IGF‐1R antibodies (i.e., Ab1, Ab2 and Ab3) can antagonize HS06 and HS36 binding. Interestingly, all three Abs competed effectively with the smaller oligosaccharide (HS06); yet, Ab2 targeting the region spanning L1 and CR (i.e., residues 170–275), was particularly effective (Figures 2C; see Figures S17&S18 for Ab1 and Ab3). In stark contrast, none of the three Abs reversed HS36 binding to HT‐29 spheroids. Finally, we studied the competitive influence of IGF1 on HS06 and HS36 recognition of cell surface IGF‐1R. Pre‐treatment with IGF1 induced a dose‐dependent decrease in the binding of HS06, but not HS36 (Figures 2E and 2F). Together, these results imply that the smaller oligosaccharide HS06 preferentially binds to cell surface hIGF‐1R, whereas the longer polysaccharide (HS36) engages other target(s), while also possibly binding to IGF‐1R.
Figure 2.

Smaller oligosaccharides, e.g., HS06, but not polysaccharide (HS36), bind preferentially to cell surface IGF‐1R and compete with IGF1. A) and B) Images of fluorescence stained HT‐29 spheroids prepared from either IGF‐1R knockdown (KD) or scrambled cell lines and incubated with 20 μM HS06‐AF594 (red, Panel A) or 20 μM HS36‐AF594 (red, Panel B) for 20 mins. C) and D) Representative images of fluorescence stained HT‐29 spheroids pre‐treated with IGF‐1R N‐terminal antibodies Ab2184–283 (0 to 10 ng mL−1) for 2 hrs and then incubated with 20 μM HS06‐AF594 (red; C) or 20 μM HS36‐AF594 (red; D) for 20 mins. (E and F). Representative images of fluorescence stained HT‐29 spheroids pre‐treated with 0–20 ng mL−1 IGF1 followed by treatment with 20 μM HS06‐AF594 (red, Panel E) or 20 μM HS36‐AF594 (red, Panel F) for 20 min. For all panels, nuclei were counterstained with DAPI (blue). Average relative fluorescence units (RFUs) at the single cell level (i.e., total intensity/number of cells) are shown in bar graphs. The scale bar in each image represents 30 μm. Data represent mean±S.E. *p<0.005 vs. scrambled and vehicle controls (A&B; C&D). *p<0.01 vs. vehicle controls (E&F).
Computational Studies Identify the Site of Binding of Smaller Oligosaccharides, e.g., HS06, on IGF‐1R
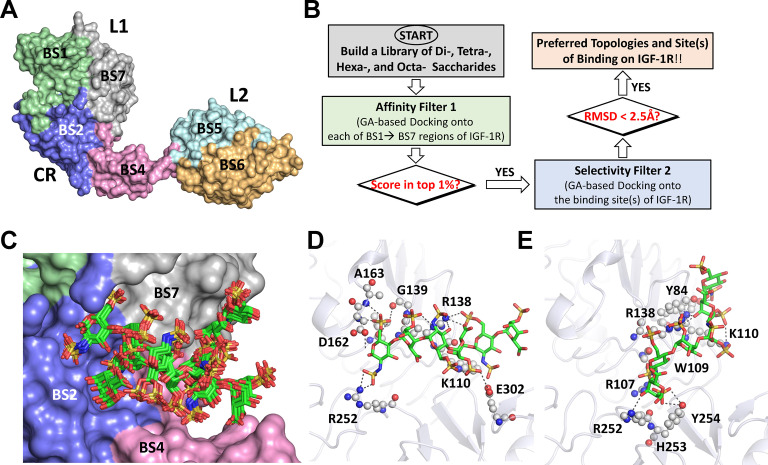
To decipher the site of binding, we resorted to the computational, dual‐filter combinatorial virtual library screening (CVLS) algorithm that we had developed earlier for GAGs. [32] This algorithm elucidates the selectivity of GAG binding to proteins and has now been validated on a large number of proteins, especially with regard to chain length and sulfation.[ 27 , 28 , 33 , 34 , 35 ] Although several structures of IGF‐1R are reported in the free[ 10 , 36 ] and ligand‐bound forms,[ 10 , 37 , 38 ] our studies indicated HS06 competing with IGF1, which implied that the structure of free IGF‐1R (PDB ID: 1IGR) [36] would be most appropriate for CVLS studies. The ECD of IGF‐1R was divided into seven overlapping potential sites of binding (Figures 3A and 3B) for unbiased screening of a library of HS di‐, tetra‐, hexa‐ and octa‐ saccharides using CVLS.
Figure 3.
Identification of the putative HS binding site on human IGF‐1R using CVLS algorithm. A) The surface of extracellular domain of hIGF‐1R was divided into 7 overlapping regions (BS1–BS7; Note: BS3 is on the back side) in an unbiased manner as potentially engaging HS chain(s). B) The dual‐filter CVLS algorithm used to study di‐ (HS02) to octa‐ (HS08) sequences binding to IGF‐1R. GA=genetic algorithm; RMSD=root‐mean‐square‐deviation. See review Curr. Opin. Struct. Biol. 2018, 50, 91–100 for more details. C) Overlay of the preferred poses of HS06 identified by CVLS onto IGF‐1R in the interface between the L1 and CR domains. CVLS predicted two preferred binding poses. HS06 is shown as sticks (green & red). Note: Results for HS02‐HS08 sequences are shown in Figure S16. D) and E) Recognition of the two preferred poses of HS06 by amino acid residues in the interface of L1 and CR domain. Multiple directional H‐bonds with R107, K110, R138 and R252 of IGF‐1R were predicted by CVLS. The interacting residues are shown as white/grey spheres.
For di‐ and tetra‐ saccharides, CVLS identified several plausible sites of binding on the IGF‐1R (Figures S19 and S20). The predicted in silico interaction scores were modest indicating rather weak binding. In contrast, HS06 sequences bound in the interfacial region between the L1 and CR domains with a high in silico interaction score. This site was also the only putative site of binding on IGF‐1R that met the criteria of high affinity and high consistency of binding (Figures 3C, Table S2). This consistency of binding arose from the engagement of R107, K110, R138 and R252 of IGF‐1R forming key interactions with sulfate groups of the HS06 (Figures 3D and E). Backbone atoms of W109, A163, and G139, and −OH groups of Y84 and Y254 were also found to contribute to recognition of the HS06 sequences.
We then compared the CVLS modeled HS06‐IGF‐1R complex with the literature reported IGF1‐IGF‐1R crystal structure (Figure S21).[ 37 , 38 ] The visualization revealed that the extended loop of IGF1 overlapped with the site of binding of HS06, which meant that the two species are likely to compete for IGF‐1R.
When HS08 was studied using CVLS, it was predicted to engage an extended region on the L1 domain with a moderate interaction score (not as high as HS06, Figure S22 & Table S2). The reason for HS08 to not bind in the HS06 site with high affinity appears to be the ridge lining the site (Figure S23). In fact, replacing I160 of the ridge with Ala does enable better binding of the HS08 chain. Thus, the site for binding of smaller oligosaccharides, e.g., HS06, at the interface of L1 and CR domain appears to be rather well‐defined and capable of preferentially hosting smaller GAGs. Alternatively, this site is not favored by longer polysaccharides, e.g., HS36.
Smaller Oligosaccharides (e.g., HS06) but Not Longer Polysaccharides (e.g., HS36), Inhibit Cell Surface IGF‐1R, which is Reversed by IGF1
To determine whether HS06 binding to IGF‐1R has consequences on its activation, we studied receptor autophosphorylation. In these experiments, we used higher concentrations (20 μM) of HS06 and HS36 in comparison to exogenously added IGF1 (≈2.5 μM) anticipating the presence of high endogenous levels of IGF1 in IGF‐1R knockdown cells. A dose‐dependent effect of HS06 pre‐treatment on IGF‐1R activation (pIGF‐1R levels) in HT‐29 spheroids was observed when probed with anti‐pIGF‐1R antibodies (Figure 4A). Inhibition of IGF‐1R was observed early (<10 min) and persisted for ≈30 min suggesting a direct effect of smaller oligosaccharides binding to IGF‐1 (Figure S24). In contrast, longer polysaccharides (HS36) were not effective at reducing IGF‐1R activation (Figure 4B). In fact, the longer polysaccharides appeared to activate IGF‐1R. This is a novel observation. Yet, at this point it is not clear whether this is because of HS36 alone or HS36 in combination with high levels of IGF1. Further studies are need to clarify its mechanism of action.
Figure 4.

HS06, but not HS36, preferentially inhibits cell surface IGF‐1R, which is competitively reversed by IGF1. (Panels A and B): Representative images of immunofluorescence stained HT‐29 spheroids with pIGF‐1R (green; AlexaFluor488). The spheroids were pre‐incubated with either 0–50 μM HS06 (A) or HS36 (B) for 20 mins. (Panels C and D): Representative images of immunofluorescence stained HT‐29 spheroids pre‐treated with 0–20 ng mL−1 IGF1 followed by treatment with either 20 μM HS06 (C) or 20 μM HS36 (D) for 20 min. Nuclei were counterstained with DAPI (blue). Average relative fluorescence units (RFUs) at the single cell level (A&B) or precent change in relative fluorescence (ΔF/F 0) normalized to respective vehicle controls (C&D) corresponding to the experiments are shown in bar graphs The scale bar in each image represents 30 μm. Data represent mean±S.E. *p<0.05 vs. respective vehicle controls. Although several biological replicates were studied, the averages presented are from two biological replicates, each of which involve a minimum count of 150–200 cells.
We also studied downstream effects of HS06‐dependent reduction in IGF‐1R activation. Significant inhibition of insulin receptor substrate (IRS)‐1, a downstream adapter protein phosphorylated by IGF‐1R, was observed (Figure S25). Likewise, phosphorylation of AKT, another downstream protein belonging to the IGF‐1R signaling system, was also inhibited. Thus, smaller oligosaccharides (HS06) can be expected to induce significant IGF‐1R‐dependent modulation intracellular signals.
We then studied whether HS06 competes with IGF1 in activation of IGF‐1R. In line with the above results, HS06‐mediated reduction of pIGF‐1R levels on HT‐29 spheroids was reversed in a concentration‐dependent manner by pre‐treatment with IGF1 (Figure 4C). In contrast, the presence of HS36 did not reduce activation of IGF‐1R on spheroids at low levels of IGF1, whereas IGF‐1R activation appeared to increase at higher levels of IGF1 (Figure 4D). Whether this increase is due to the longer polysaccharide alone or its combination with IGF1 will require additional work; however, prior work has shown that HS36 does not inhibit colorectal spheroid growth. [39] These results imply that smaller oligosaccharides, e.g., HS06, and longer polysaccharides, e.g., HS36, have considerably different impacts on IGF‐1R signaling.
Validation of the Predicted Site of Binding of SmallerOligosaccharides, e.g., HS06
We generated HT‐29 cells that expressed mutant hIGF‐1R with replacement in two of the four basic residues hypothesized to be important for the interaction, i.e., R107 and R138. HT‐29 cells, in which wild‐type hIGF‐1R was significantly reduced (described above), were used to introduce either R107H, R138A, or R107H+R138A mutations‐containing constructs on pBABE‐bleo, a mammalian expression vector, to generate the corresponding mutant cell lines. Simultaneously, HT‐29 cells were also transfected with a wild‐type IGF‐1R pBABE‐bleo vector to generate the WT control. R017H mutation was selected because it has been observed in urothelial and lymphoid cancers. Ala replacement at R138 position was chosen to evaluate the role of electrostatics.
First, the level of IGF‐1R expression in the respective cell lines was assessed using AF‐labeled antibodies against IGF‐IR and found to be nearly identical (Figure 5A). Next, binding of HS06 to WT and mutant cells was evaluated using AF594‐labeled HS06. In comparison to the WT HT‐29 cell line, R107H, R138A, and R107H+R138A mutant cell lines displayed 88±8, 69±13 and 88±15 % losses, respectively, in fluorophore intensities (Figure 5B). This indicated a significant defect in the binding of smaller oligosaccharides owing to replacement of R107 and R138 residues.
Figure 5.

Identification of the site of HS06 binding on hIGF‐1R. (Panels A and B): Representative immunofluorescence images of HT‐29 spheroids from expressing wild‐type or mutant IGF‐1R. The generation of HS06 binding site mutant cell lines is described in the Methods section. A) Immunofluorescence staining with IGF‐1R (red; AlexaFluor568); B) Immunofluorescence staining of spheroids pre‐incubated with 20 μM HS06 for 20 mins. ΔF/F 0 (%) represents relative fluorescence change compared to WT controls. Data represent mean±S.E. *p<0.05. (Panels C–F) Representative images of fluorescence‐stained HS06 binding site mutant HT‐29 spheroids for WT (C), R107 (D), R138A (E) and R107+R138A (F) with either vehicle (water) or 20 μM HS06‐AlexaFluor594 for 20 min. Nuclei were counterstained with DAPI (blue). ΔF/F 0 (%) corresponding to the experiments are shown underneath the images from the average relative fluorescence units (RFUs) at the single cell level (i.e., total intensity/number of cells). The change in fluorescence was calculated from the observed fluorescence minus background (AF594). The scale bar in each image represents 30 μm. Data represent mean ± S.E. *p<0.005 vs. vehicle control. Although several biological replicates were studied, the averages presented are from two biological replicates, each of which involve a minimum count of 150–200 cells.
Following this, we studied whether receptor activation is impacted by the introduction of the three mutations. As described in Figure 4, activation was studied using AF568‐labeled antibodies against pIGF‐1R. In the absence of HS06, autophosphorylation of the receptor on HT‐29 spheroids generated from the wild‐type and mutant cell lines was essentially identical (Figure S26). In contrast, the presence of HS06 led to significant differences in autophosphorylation. Figures 5C through F show the results for WT, R107H, R138A and R107H+R138A HT‐29 spheroids and Figure 5G quantifies the change in activation originating from the exposure of HS oligosaccharides relative to the vehicle control. HS06 significantly reduces WT receptor activation (−78±16 %, p<0.005), as one would predict from prior results. In contrast, none of the three mutants, including the double mutant spheroids, displayed a significant change (p<0.01) in receptor activation as compared to the corresponding vehicle‐treated HT‐29 spheroids. This implies that IGF‐1R activation is significantly impaired if either or both HS06‐binding, basic residues are mutated. Thus, smaller oligosaccharides, e.g., HS06, recognize and inhibit IGF‐1R through specific residues, e.g., R107 and R138, which line the interface between the L1 and CR domains.
Conclusion
This work reveals several new insights. With regard to RTK biology, it has been known HSPGs contribute to the cellular processes of different receptors, e.g., FGFRs, PDGFR, Tie1, and VEGFRs.[ 13 , 19 , 40 , 41 ] IGF‐1R also contributes to many cellular processes such as growth, differentiation, invasion/metastasis, angiogenesis and anti‐apoptosis.[ 2 , 3 , 42 ] More pertinently, HS/HSPGs have been implicated in IGF‐1R regulation;[ 43 , 44 , 45 ] however, GAG oligosaccharide‐based modulation of IGF‐1R had escaped researchers until now.
We find that smaller oligosaccharides of HS bind to and modulate IGF‐1R activation. More specifically, hexasaccharide sequences of HS, e.g., HS06, potently and preferentially inhibits IGF1‐induced activation of IGF‐1R. To be more precise, HS06 also binds to IGF1, albeit with much weaker affinity. Alternatively, smaller oligosaccharides prefer IGF‐1R over IGF1.
HSPGs generally have been found to contribute to activation of RTKs by either directly binding to the RTK or by facilitating the interaction of the cognate ligands, e.g., FGFs, PDGF and VEGF, with the receptor. [13] Interestingly for some RTKs, e.g., FGFRs, HS oligosaccharides have been shown to either inhibit or activate the receptor depending on their sulfation patterns. [46] Our work presents a novel phenomenon that smaller oligosaccharides, but not longer polysaccharides, directly bind and inhibit IGF‐1R activation. Among the smaller oligosaccharides, the preparation of HS06 stands out because of its ≈200 nM affinity for IGF‐1R. In comparison, slightly longer chains, e.g., HS12, bind ≈1.5‐fold weaker, whereas polysaccharides, e.g., HS36, are much weaker by ≈9‐fold. This discovery has important implications because smaller oligosaccharides of different lengths can be produced in vivo through the actions of heparanase, an enzyme present in the tumor microenvironment and known to degrade HSPGs.[ 13 , 47 , 48 , 49 ]
This work also presents a novel insight on GAG binding to proteins. Traditionally, proteins prefer longer GAG chains because their affinity is higher. This has been shown for several RTKs, and especially for FGFRs.[ 19 , 29 , 30 ] Further, the longer and shorter GAG chains generally bind to the same site on the RTK. This confers a major advantage because natural GAGs are comprised of both shorter and longer chains. In contrast, the smaller oligosaccharides, e.g., HS06, prefer to bind to a defined site on IGF‐1R, which is not engaged by polysaccharides, e.g., HS36. This should alert glycobiologists that longer and shorter HS chains may not bind in the same site on proteins.
A key question arises whether naturally present smaller oligosaccharides, e.g., HS06, modulate cell surface IGF‐1R. The role of endogenous HS06‐like species remains unknown. In fact, the levels of endogenous smaller oligosaccharides, especially in the context of cancer, remains unknown. It has been shown in 1995 that circulating HS is a mixture of shorter and longer oligosaccharides. [50] Likewise, smaller oligosaccharides, such as hexasaccharides, have been found in the plasma of mucopolysaccharidosis patients. [51] Recently, circulating HS fragments, including hexasaccharides, were detected 23‐fold higher under inflammatory conditions. [52] This suggests that smaller oligosaccharides are likely to be present in the vicinity of CSCs owing to the actions of tumor niche heparanase.[ 47 , 48 , 49 , 53 ] In contrast, polymeric HS has been known to stimulate the growth of most cells including normal stem cells. [54] Thus, it is possible that the engagement of the interfacial HS06‐binding site on IGF‐1R, which induces receptor downregulation, may provide a biologic advantage. Although the consequences of longer polysaccharides, e.g., HS36, were not fully studied here, it is possible to speculate that nature may be advantageously using different functions (e.g., inhibition, neutral and activation) in a context‐dependent manner.
IGF‐1R has been touted as anti‐cancer drug target for a long time.[ 55 , 56 ] IGF‐1R activation on CSCs helps propagate self‐renewal and pluripotency, which stimulates cancerous growth.[ 3 , 31 ] Therefore, smaller oligosaccharide‐based inhibition of IGF‐1R should lead to reduction in tumor. In fact, prior work with smaller oligosaccharides, especially HS06, has already been shown to inhibit CSCs, [39] which express high levels of IGF‐1R. Likewise, a synthetic mimetic of HS06 has also been shown to selectively inhibit proliferation of CSCs, while exhibiting potent anti‐tumor properties in vivo.[ 57 , 58 ] Hence, we predict HS‐based smaller oligosaccharides or mimetics could yield novel anti‐cancer agents.
Overall, this work presents unique structural and functional insights on the modulation of hIGF‐1R by smaller oligosaccharides, e.g., HS06. Yet, interesting questions arise in terms of IGF‐1R regulation by longer polysaccharides, e.g., HS36. Where do these longer sulfated polysaccharides bind? Do these co‐operatively or independently work with IGF1? Do these activate the receptor? If so, what intracellular signals are transduced? Answering such questions will help develop novel anti‐cancer agents, while also presenting new light on RTK biology.
Experimental Procedures/Data
Details on experimental methods are provided in Supporting Information.
Conflict of interest
The authors declare no conflict of interest.
1.
Supporting information
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Acknowledgements
This work was supported in part by VA Merit Award 5I01BX000837 and 1I01BX004584, and Commonwealth Health Research Board grant 811‐04‐19 to BBP; NIH grants HL141954, CA241951 and HL151333 to URD. Computing resources from the National Center for Research Resources (S10RR027411) to VCU are also highly appreciated.
R. S. Boothello, N. V. Sankaranarayanan, J. C. Sistla, B. Nagarajan, C. Sharon, J. E. Chittum, R. Y. Niyaz, S. Roy, A. Nandi, C. P. O'Hara, R. N. Gangji, D. K. Afosah, R. Ongolu, B. B. Patel, U. R. Desai, Angew. Chem. Int. Ed. 2022, 61, e202211320; Angew. Chem. 2022, 134, e202211320.
Contributor Information
Prof. Dr. Bhaumik B. Patel, Email: Bhaumik.Patel@va.gov.
Prof. Dr. Umesh R. Desai, Email: urdesai@vcu.edu.
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
- 1. Adams T. E., Epa V. C., Garrett T. P., Ward C. W., Cell. Mol. Life Sci. 2000, 57, 1050–1093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Yuan J., Yin Z., Tao K., Wang G., Gao J., Oncol. Lett. 2018, 15, 41–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Teng C. F., Jeng L. B., Shyu W. C., Cell. Transplant. 2018, 27, 1313–1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Sharon C., Baranwal S., Patel N. J., Rodriguez-Agudo D., Pandak W. M., Majumdar A. P., Krystal G., Patel B. B., Oncotarget 2015, 6, 15332–15347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Yin M., Guan X., Liao Z., Wei Q., Am. J. Transl. Res. 2009, 1, 101–114. [PMC free article] [PubMed] [Google Scholar]
- 6. Gross J. M., Yee D., Cancer Metastasis Rev. 2003, 22, 327–336. [DOI] [PubMed] [Google Scholar]
- 7. Mitsiades C. S., Mitsiades N. S., McMullan C. J., Poulaki V., Shringarpure R., Akiyama M., Hideshima T., Chauhan D., Joseph M., Libermann T. A., García-Echeverría C., Pearson M. A., Hofmann F., Anderson K. C., Kung A. L., Cancer Cell 2004, 5, 221–230. [DOI] [PubMed] [Google Scholar]
- 8. Ouban A., Muraca P., Yeatman Y., Coppola D., Hum. Pathol. 2003, 34, 803–808. [DOI] [PubMed] [Google Scholar]
- 9. Gallagher E. J., LeRoith D., Endocrinology 2011, 152, 2546–2551. [DOI] [PubMed] [Google Scholar]
- 10. Xu Y., Kong G. K., Menting J. G., Margetts M. B., Delaine C. A., Jenkin L. M., Kiselyov V. V., De Meyts P., Forbes B. E., Lawrence M. C., Nat. Commun. 2018, 9, 821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Papadopoulos N., Lennartsson J., Mol. Aspects Med. 2018, 62, 75–88. [DOI] [PubMed] [Google Scholar]
- 12. Yang Y., Yuzawa S., Schlessinger J., Proc. Natl. Acad. Sci. USA 2008, 105, 7681–7686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Lanzi C., Cassinelli G., Biochem. Pharmacol. 2020, 178, 114084. [DOI] [PubMed] [Google Scholar]
- 14. Grünewald F. S., Prota A. E., Giese A., Ballmer-Hofer K., Biochim. Biophys. Acta Proteins Proteomics 2010, 1804, 567–80. [DOI] [PubMed] [Google Scholar]
- 15. Xie M., Li J. P., Cell. Signalling 2019, 54, 115–121. [DOI] [PubMed] [Google Scholar]
- 16. Schlessinger J., Plotnikov A. N., Ibrahimi O. A., Eliseenkova A. V., Yeh B. K., Yayon A., Linhardt R. J., Mol. Cell 2000, 6, 743–750. [DOI] [PubMed] [Google Scholar]
- 17. Pellegrini L., Burke D. F., von Delft F., Mulloy B., Blundell T. L., Nature 2000, 407, 1029–1034. [DOI] [PubMed] [Google Scholar]
- 18. Kavran J. M., McCabe J. M., Byrne P. O., Connacher M. K., Wang Z., Ramek A., Sarabipour S., Shan Y., Shaw D. E., Hristova K., Cole P. A., Leahy D. J., eLife 2014, 3, e03772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Pomin V. H., Biochimie 2016, 127, 214–226. [DOI] [PubMed] [Google Scholar]
- 20. Iozzo R. V., Sanderson R. D., J. Cell. Mol. Med. 2011, 15, 1013–1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Chen P. C., Kuo Y. C., Chuong C. M., Huang Y. H., Front. Cell. Dev. Biol. 2021, 8, 625943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Tzanakakis G. N., Giatagana E. M., Berdiaki A., Spyridaki I., Hida K., Neagu M., Tsatsakis A. M., Nikitovic D., Cancers (Basel) 2021, 13, 2478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Chitnis M. M., Yuen J. S., Protheroe A. S., Pollak M., Macaulay V. M., Clin. Cancer Res. 2008, 14, 6364–6370. [DOI] [PubMed] [Google Scholar]
- 24. Pollak M., Nat. Rev. Cancer 2012, 12, 159–169. [DOI] [PubMed] [Google Scholar]
- 25. Olson S. T., Halvorson H. R., Björk I., J. Biol. Chem. 1991, 266, 6342–6352. [PubMed] [Google Scholar]
- 26. Cardin A. D., Weintraub H. J., Arteriosclerosis 1989, 9, 21–32. [DOI] [PubMed] [Google Scholar]
- 27. Sankaranarayanan NV., Nagarajan B., Desai U. R., Curr. Opin. Struct. Biol. 2018, 50, 91–100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Sankaranarayanan NV., Desai U. R., Glycobiology 2014, 24, 1323–1333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Xu R., Ori A., Rudd T. R., Uniewicz K. A., Ahmed Y. A., Guimond S. E., Skidmore M. A., Siligardi G., Yates E. A., Fernig D. G., J. Biol. Chem. 2012, 287, 40061–40073. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Feyzi E., Lustig F., Fager G., Spillmann D., Lindahl U., Salmivirta M., J. Biol. Chem. 1997, 272, 5518–5524. [DOI] [PubMed] [Google Scholar]
- 31. Chang W. W., Lin R. J., Yu J., Chang W. Y., Fu C. H., Lai A., Yu J. C., Yu A. L., Breast Cancer Res. 2013, 15, R39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Raghuraman A., Mosier P. D., Desai U. R., J. Med. Chem. 2006, 49, 3553–3562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Sankaranarayanan NV., Strebel T. R., Boothello R. S., Sheerin K., Raghuraman A., Sallas F., Mosier P. D., Watermeyer N. D., Oscarson S., Desai U. R., Angew. Chem. Int. Ed. 2017, 56, 2312–2317; [DOI] [PMC free article] [PubMed] [Google Scholar]; Angew. Chem. 2017, 129, 2352–2357. [Google Scholar]
- 34. Sankaranarayanan NV., Nagarajan B., Desai U. R., Int. J. Mol. Sci. 2021, 22, 7542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Sankaranarayanan NV., Bi Y., Kuberan B., Desai U. R., Comput. Struct. Biotechnol. J. 2020, 18, 933–941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Garrett T. P., McKern N. M., Lou M., Frenkel M. J., Bentley J. D., Lovrecz G. O., Elleman T. C., Cosgrove L. J., Ward C. W., Nature 1998, 394, 395–399. [DOI] [PubMed] [Google Scholar]
- 37. Li J., Choi E., Yu H., Bai X. C., Nat. Commun. 2019, 10, 4567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Xu Y., Kirk N. S., Venugopal H., Margetts M. B., Croll T. I., Sandow J. J., Webb A. I., Delaine C. A., Forbes B. E., Lawrence M. C., Structure 2020, 28, 786–798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Patel N. J., Sharon C., Baranwal S., Boothello R. S., Desai U. R., Patel B. B., Oncotarget 2016, 7, 84608–84622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Ohkawa Y., Wade A., Lindberg O. R., Chen K. Y., Tran V. M., Brown S. J., Kumar, Kalita M., James C. D., Phillips J. J., Mol. Cancer Res. 2021, 19, 150–161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Griffin M. E., Sorum A. W., Miller G. M., W. A. Goddard III , Hsieh-Wilson L. C., Nat. Chem. Biol. 2021, 17, 178–186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Baserga R., Oncogene 2000, 19, 5574–5581. [DOI] [PubMed] [Google Scholar]
- 43. Qiu H., Jiang J. J., Liu M., Huang X., Ding S. J., Wang L., Mol. Cell. Proteomics 2013, 12, 2160–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Mytilinaiou M., Nikitovic D., Berdiaki A., Papoutsidakis A., Papachristou D. J., Tsatsakis A., Tzanakakis G. N., Exp. Cell Res. 2017, 361, 9–18. [DOI] [PubMed] [Google Scholar]
- 45. Rapraeger A. C., Ell B. J., Roy M., Li X., Morrison O. R., Thomas G. M., Beauvais D. M., FEBS J. 2013, 280, 2194–206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Guimond S. E., Turnbull J. E., Curr. Biol. 1999, 9, 1343–1346. [DOI] [PubMed] [Google Scholar]
- 47. Lanzi C., Zaffaroni N., Cassinelli G., Curr. Med. Chem. 2017, 24, 2860–2886. [DOI] [PubMed] [Google Scholar]
- 48. Masola V., Zaza G., Gambaro G., Franchi M., Onisto M., Semin. Cancer Biol. 2020, 62, 86–98. [DOI] [PubMed] [Google Scholar]
- 49. Vlodavsky I., Gross-Cohen M., Weissmann M., Ilan N., Sanderson R. D., Trends Biochem. Sci. 2018, 43, 18–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Volpi N., Cusmano M., Venturelli T., Biochim. Biophys. Acta Gen. Subj. 1995, 1243, 49–58. [DOI] [PubMed] [Google Scholar]
- 51. Byers S., Rozaklis T., Brumfield L. K., Ranieri E., Hopwood J. J., Mol. Genet. Metab. 1998, 65, 282–290. [DOI] [PubMed] [Google Scholar]
- 52. Schmidt E. P., Li G., Li L., Fu L., Yang Y., Overdier K. H., Douglas I. S., Linhardt R. J., J. Biol. Chem. 2014, 289, 8194–8202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Doviner V., Maly B., Kaplan V., Gingis-Velitski S., Ilan N., Vlodavsky I., Sherman Y., Mod. Pathol. 2006, 19, 878–888. [DOI] [PubMed] [Google Scholar]
- 54. Kraushaar D. C., Dalton S., Wang L., Biol. Chem. 2013, 394, 741–751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Singh P., Alex J. M., Bast F., Med. Oncol. 2014, 31, 805. [DOI] [PubMed] [Google Scholar]
- 56. Buck E., Mulvihill M., Expert Opin. Invest. Drugs 2011, 20, 605–621. [DOI] [PubMed] [Google Scholar]
- 57. Boothello R. S., Patel N. J., Sharon C., Abdelfadiel E. I., Morla S., Brophy D. F., Lippman H. R., Desai U. R., Patel B. B., Mol. Cancer Ther. 2019, 18, 51–61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Patel N. J., Karuturi R., Al-Horani R. A., Baranwal S., Patel J., Desai U. R., Patel B. B., ACS Chem. Biol. 2014, 9, 1826–33. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
As a service to our authors and readers, this journal provides supporting information supplied by the authors. Such materials are peer reviewed and may be re‐organized for online delivery, but are not copy‐edited or typeset. Technical support issues arising from supporting information (other than missing files) should be addressed to the authors.
Supporting Information
Data Availability Statement
The data that support the findings of this study are available from the corresponding author upon reasonable request.