Abstract
Aims:
Evidence suggests alterations of thyroid hormone levels can disrupt normal bone development. Most data suggest the major targets of thyroid hormones to be the Htra1/Igf1 pathway. Recent discovery by our group suggests involvement of targets WNT pathway, specifically overexpression of antagonist Sfrp4 in the presence of exogenous thyroid hormone.
Main methods:
Here we aimed to model these interactions in vitro using primary and isotype cell lines to determine if thyroid hormone drives increased Sfrp4 expression in cells relevant to craniofacial development. Transcriptional profiling, bioinformatics interrogation, protein and function analyses were used.
Key findings:
Affymetrix transcriptional profiling found Sfrp4 overexpression in primary cranial suture derived cells stimulated with thyroxine in vitro. Interrogation of the SFRP4 promoter identified multiple putative binding sites for thyroid hormone receptors. Experimentation with several cell lines demonstrated that thyroxine treatment induced Sfrp4 expression, demonstrating that Sfrp4 mRNA and protein levels are not tightly coupled. Transcriptional and protein analyses demonstrate thyroid hormone receptor binding to the proximal promoter of the target gene Sfrp4 in murine calvarial pre-osteoblasts. Functional analysis after thyroxine hormone stimulation for alkaline phosphatase activity shows that pre-osteoblasts increase alkaline phosphatase activity compared to other cell types, suggesting cell type susceptibility. Finally, we added recombinant SFRP4 to pre-osteoblasts in combination with thyroxine treatment and observed a significant decrease in alkaline phosphatase positivity.
Significance:
Taken together, these results suggest SFRP4 may be a key regulatory molecule that prevents thyroxine driven osteogenesis. These data corroborate clinical findings indicating a potential for SFRP4 as a diagnostic or therapeutic target for hyperostotic craniofacial disorders.
Keywords: Sfrp4, Thyroxine, Osteogenesis, mRNA, EMSA, Protein
1. Introduction
The effects of thyroid hormone on somatic growth is well established [1–4]. Homeostasis is key in this paradigm where hyper-, hypo-, and other thyroid associated diseases and replacement medications have been shown to have profound effects on development. The known primary effects of thyroid hormones on bone involve local and circulating IGF (insulin-like growth factor) levels and its associated serine proteinase, HTRA1 [3,5–9]. Thyroid hormone is generally thought to be pro-anabolic within this system with its primary effect being increased activity of the osteoblasts [10,11,11–16]. However, as with any hormone, balance plays a key role and alterations to osteoclast activity have also been observed with excess or deficiency in thyroid hormones [10,17–20].
Our research team has worked in the area of environmental effects on growth and development of the craniofacial skeleton and gene environment interactions involving thyroid hormones [21–26]. To date, our work has suggested that, at least in vitro, thyroxine (T4) exposure, commonly prescribes clinically as leveothyroxine, can drive increases in osteoblast activity of calvarial derived pre-osteoblast cells and precipitate marked changes in IGF pathway-associated genes (Htra1, Igf1) [21]. Interestingly, in our own interrogation of the perisutural cranial areas after in utero thyroid hormone exposures, we did not observe marked changes to these same IGF-associated markers. WNT pathway targets, specifically antagonists including Dkk and Sfrp genes were, however, differentially expressed [25]. As canonical WNT signaling is associated with increased osteoblastogenesis [27], interrogation of these relationships is warranted.
The association of many WNT-related genes has been linked to bone growth disorders including those of the craniofacial skeleton such as craniosynostosis [28]. Craniosynostosis involves the premature fusion of the cranial suture prior to the completion of brain expansion. This disorder has a reported incidence of 1 in 1800–2500 live births in the United States and often presents with concomitant co-morbidities including eye related symptomologies, increased intracranial pressure, and altered cranial volumes [29–31]. Although craniosynostosis is described as a genetic disorder, only a minority of the cases are classified into the classic FGF, TWIST, or TGFβ gene mutation related disorders. The majority of craniosynostosis cases are non-syndromic with unknown etiology [29,32–35]. Human craniosynostosis cohort data as well as our pre-clinical modeling of excess thyroid hormone gene-environment interaction in skull growth suggest involvement of WNT and WNT antagonist related genes[4,21,23–26]. We now have data that point to potential involvement of Sfrp4, a WNT antagonist, which may be a novel target in the identification and diagnosis of thyroid hormone driven growth disorders. Thus, we set out to confirm that Sfrp4 would be differentially regulated in mesenchymal derived cells including those of the craniofacial skeleton.
2. Material and Methods
2.1. Isolation and Culture of Primary Suture Derived Cells
Animal use protocols were approved by the Medical University of South Carolina Institutional Animal Care and Use Committee (AR#3341) and all procedures and the reporting thereof are in compliance with the Animal Research: Reporting in Vivo Experiments (ARRIVE) guidelines[36]. Murine calvarial primary suture derived cells were harvested from postnatal day 15 C57Bl6J pups per published protocol [24] and grown to confluence. Cells were then seeded at a density of 65,000 cells per well and treated with standard Dulbecco’s Modified Eagle Medium (DMEM, ThermoFisher Scientific, Pittsburgh PA) or DMEM supplemented with 10−6 mol/liter thyroxine for 7 days [21].
2.2. Affymetrix Analysis of in vitro Thyroxine Treated Primary Suture Derived Cells
Total cellular RNA was extracted using the Qiagen RNeasy mini kit (Qiagen, Valencia, CA, USA). GeneChip® Mouse Gene 2.0 ST Array (Affymetrix, Santa Clara, CA) was utilized to determine dysregulated gene products after 7 days of thyroxine treatment. Total RNA was screened for quality and quantity by bioanalyzer analysis (Agilent 2100, Santa Clara, CA), then 250 ng of RNA was used to prepare biotin labeled product for each sample. All results have been deposited at NCBI GEO under accession GSE66149.
2.3. RNA Isolation and qrtPCR Confirmation on Multiple Cell Types
Murine calvarial primary suture derived cells (as above), murine bone marrow stem cells (BMSC) [37], as well as isotype cell lines myoblast C2C12 and pre-osteoblasts MC3T3-E1 (ATCC, Manassas, VA) were cultured and RNA was isolated at 6 hours, 1 day, 3 days, or 7 days after the onset of thyroxine treatment. cDNA synthesis was performed with a qScript cDNA Synthesis Kit (Beverly, MA) following manufacturer’s instructions and subjected to real time PCR using Taqman assay for Sfrp4 (Mm00840104_m1) and 18s rRNA (Mm03928990_g1) as a housekeeping gene. The qRT-PCR was performed using a Taqman Fast Advanced Master Mix (ThermoFisher) and ABI StepOne Plus. Relative gene expression was quantified between control and thyroxine treatment at each time point.
2.4. Bioinformatic Interrogation of SFRP4 Promoter Region
Using Genomatix software, analysis was performed on the mouse Sfrp4 annotated promoter sequence (GXP_12095), which extended from −602 to +317 relative to the mouse Sfrp4 transcription start site (NCBI NM_016687.3). Orthologous segments of human and rat Sfrp4 genes were identified using the UCSC Genome BLAST search tool and then accessed through NCBI Nucleotide resources. Canonical thyroid hormone receptor (THR) binding sites were identified in the mouse Sfrp4 promoter using Genomatix MatInspector. Conservation of canonical THR binding sites in human and rat Sfrp4 promoters were evaluated with the Genomatix Common TFs tool. Multiple sequence alignment was done with Clustal Omega web service software [38].
2.5. Protein Isolation and Studies
Primary suture derived cells, BMSCs, C2C12s, and MC3T3-E1s were seeded at 300,000 cells per 9.6 cm2 well. Control media and thyroxine supplemented media were used to treat the cells for 24 hours. At isolation, supernatant media was aspirated, collected, spun down to remove debris and stored until use in protein studies. Adhered cells were washed with PBS and lysed with radioimmunoprecipitation assay buffer (RIPA, Thermofisher) and incubated under agitation at 4°C for 30 minutes. Isolation was completed using a cell scraper and centrifugation. Standard Bradford Assay determined total protein concentration in processed supernatants and lysates.
Cell supernatants and lysates were subject to ELISA analysis for SFRP4 (MBS907810, MyBioSource, San Diego, CA), following manufacturer’s instructions. Briefly, samples were diluted in provided sample diluent based on protein concentration (supernatants diluted 1:3; BMSC lysates diluted 1:3, all other lysate were not diluted). Standards were prepared per instruction. Samples and standards were incubated for 2 hours in provided plate at 37°C. After, biotin antibody was added and incubated for one hour followed by washes before the addition of HRP-avidin and a one-hour incubation at 37°C. After incubation, substrate was added and incubated for 15 minutes followed by addition of stop solution. Optical density was read at 450nm with corrections at 540nm and 570nm and recorded. Concentration of SFRP4 was determined based on standard curve.
2.6. Sfrp4 Promoter Region Luciferase Assay
A 217-bp fragment of the Sfrp4 gene extending from −63 to +154 relative to the transcriptional start site was isolated by PCR from MC3T3-E1 genomic DNA (Fw:GATCGGTACCTTGCTCTCGCCGCTGCCAAG; Rev: GGCGCAGCCACAGGCATAAC) and subcloned into the Kpn I and Sac I sites of pGL4.23, a firefly luciferase reporter vector containing a minimal promoter (Promega, Madison, WI). The identity of the fragment was confirmed by DNA sequencing. The CMV-Renilla luciferase vector pGL4.75 (Promega) was used as a control for transfection efficiency.
The MC3T3-E1 cell line was used in transfection studies. Briefly, cells were seeded in a 96 well white flat bottom plate at a density of 10,000 cells per well (0.32 cm2). After 24 hours, media was replaced with serum free media supplemented with retinoic acid (Sigma Aldrich, St. Louis, MO) at a concentration of 1.52 μg/ml as retinoic receptors participate in a heterodimeric relationship to mediate thyroid receptor activities. Firefly luciferase plasmids with or without Sfrp4 sequences were mixed at a ratio of approximately 1:9 to the CMV-Renilla luciferase plasmid to a total concentration of 100 ng of DNA with 0.2 μl of P3000 reagent and 0.15 μl of Lipofectamine 3000 reagent following manufacturer’s protocol for Lipofectamine 3000 (Invitrogen, Waltham, MA). Cells were incubated for an additional 24 hours after treatment. Six hours prior to assay, thyroxine at a concentration of 10−6 mol/liter was added to experimental wells. The Dual-Glo Luciferase System (Promega) was used to assess luciferase reporter activity following the manufacturer’s protocol. Firefly and Renilla luciferase reads occurred at 20 minutes post-exposure to the reagents. Replicates were averaged and compared for change in luminescence intensity due to treatment with thyroxine when normalized to empty firefly plasmid pGL4.23 (background luminescence) and Renilla plasmid pGL4.75 (cellular density).
2.7. Sfrp4 Thyroid Response Element Electrophoretic Mobility Gel Shift Assay (EMSA)
All reactions and detections were performed with a LightShift Chemiluminescent EMSA kit (Thermo Scientific, Rockford IL), using the manufacturer’s suggested conditions. Unlabeled probes and probes with 5’-biotinylation were designed from the mouse Sfrp4 DNA sequence and obtained from Integrated DNA Technologies. The upstream probe encompassed the putative thyroid hormone receptor binding site at −30 to −57 relative to the transcriptional start site and was of the sequence 5’-CTGCCAAGGCTGCTGAGCCCACGTCAGGGGACGCGTCTGGATAAA-3’ (coding strand). The downstream probe encompassed two adjacent putative thyroid hormone receptor binding sites at +104 to +151 and was of the sequence 5’-CGCCCAGGGGTGATCGACGGCCAACCCCCCAGAACTCCTCATCAGTGCAGACTGGAGCT-3’ (coding strand). Nuclear extracts were isolated from MC3T3-E1 cells using NE-PER Nuclear and Cytoplasmic Extraction Reagents (Thermo Scientific). Cells were plated in T175 (75 cm2) flasks and allowed to reach a confluency of 80%. Six hours prior to isolation, half the flasks were treated with standard media (growth media replaced) or media supplemented with thyroxine at a concentration of 10−6 mol/liter. Isolations followed the manufacturer’s protocol and were frozen at −80°C until use in EMSA studies. Briefly, 10 μg of nuclear extracts were mixed with 20 fmol of biotinylated probe in LightShift standard binding buffer plus 1 μg/μl poly (dI·dC) and incubated for 20 min at room temperature, prior to addition of loading buffer and electrophoresis in a 5% non-denaturing (TBE) polyacrylamide gel (Bio-Rad, Hercules, CA). For competitive inhibition, 4 pmol unlabeled probe was added to the reaction 10 min prior to addition of the labeled probe. For supershifts, 1 or 2 μl of anti-thyroid hormone receptor beta antiserum (ChIP grade; Abcam ab5622), or 5 μg purified IgG (Sigma) was added to the reaction 30 min prior to addition of the labeled probe. Following electrophoresis, complexes were transferred to nylon membrane (Hybond N+, GE Healthcare) and UV-crosslinked (Stratalinker 1800; Stratagene), followed by detection using LightShift kit reagents and X-ray film.
2.8. Detection of Alkaline Phosphatase by Enzyme Immunoassays
At the time of confluence, primary suture derived cells, BMSCs, C2C12s, and MC3T3-E1s were seeded at a density of 4,000 cells per 0.32 cm2 well for alkaline phosphatase (ALP) assays (Sigma-Aldrich, USA). Comparison was made between control and thyroxine treated cells by kinetics of absorbance read at 405 nm on a Gen5 plate reader (BioTek, Winooski, VT, USA). ALP assay was repeated for MC3T3-E1 pre-osteoblasts with media as above supplemented with 0, 250, or 1000ng/ml of rhSFRP4 (R&D Systems, Minneapolis, MN). All assays were conducted after 7 days in culture.
2.9. Statistics
Statistical analysis of Affymetrix data was performed via t-test to determine those gene targets driven to increased or decreased expression after thyroxine exposure, with p<0.05 and expression change of 1.5 fold considered significant. For mRNA analysis, statistical differences were determined based on calculated delta-delta Ct at each post culture time point by cell type [39]. A three-way Analysis of Variance was conducted for ELISA analysis modeled as time in culture by treatment (control or thyroxine treated) by sample type (lysate or supernatant). Raw data were screened for normality and homogeneity of variance and violations were corrected by rank order transformation. Significant interaction effects are described in results as well as main effects and post-hoc Bonferroni analyses. Luciferase reporter assay conditions (control vs. thyroxine treated) were compared by t test. For quantitative alkaline phosphatase analysis, t tests were used to compare differences of control and thyroxine treated cells. All experiments were repeated at least three independent times and results are displayed as mean ± standard error of the mean.
3. Results
3.1. Genome Wide Array Reveals Sfrp4 Overexpression as Novel Target of Thyroxine
Genome Wide Array results can be found deposited in GeoAccession (GSE66149). Table 1 summarizes statistically significant results screened for false discovery. Several genes related to the IGF pathway (Htra1, Ehd1, Il6) were identified as being differentially regulated in murine primary suture derived cells after in vitro thyroxine exposure. A competitive antagonist of the canonical WNT pathway, Sfrp4, was upregulated and identified as a novel gene of interest in thyroxine treated primary suture derived cells.
Table 1:
Affymetrix Results
| UPREGULATED GENES | |||||
| ID | Gene | Mean Difference Base 2 | Fold Increase | t-value | p-value |
| 17363626 |
Pgm5
Hsp90aa |
0.79 | 1.73 | 6.43 | 0.008 |
| 17278998 | 1 | 0.75 | 1.68 | 4.19 | 0.025 |
| 17285546 | Sfrp4 | 0.67 | 1.59 | 4.98 | 0.016 |
| 17458960 | Vmn1r20 | 0.67 | 1.59 | 5.98 | 0.009 |
| 17448735 | Slain2 | 0.64 | 1.56 | 3.88 | 0.030 |
| 17477832 | Ftl1 | 0.64 | 1.56 | 4.07 | 0.027 |
| 17242839 | Atp5d | 0.63 | 1.55 | 3.92 | 0.030 |
| 17465824 | Fam180a | 0.61 | 1.53 | 3.93 | 0.029 |
| 17396024 | Stmn2 | 0.61 | 1.53 | 3.94 | 0.029 |
| 17365883 | Afap1l2 | 0.61 | 1.53 | 7.05 | 0.006 |
| 17547608 | Actg1 | 0.6 | 1.52 | 3.97 | 0.029 |
| 17318794 | Apol7c | 0.6 | 1.52 | 4.73 | 0.018 |
| 17477695 | Rpl13a | 0.59 | 1.51 | 3.9 | 0.030 |
| 17414738 | Orm3 | 0.59 | 1.51 | 4.77 | 0.018 |
| 17339392 | Myl12a | 0.58 | 1.49 | 3.66 | 0.035 |
| 17548153 | Cat | 0.57 | 1.48 | 4.07 | 0.027 |
| 17483912 | Htra1 | 0.57 | 1.48 | 5.36 | 0.013 |
| 17358214 | Klf9 | 0.56 | 1.47 | 7.13 | 0.006 |
| 17305401 | Gm3633 | 0.55 | 1.46 | 5.01 | 0.015 |
| 17548123 | Ehd1 | 0.53 | 1.44 | 3.3 | 0.046 |
| DOWNREGULATED GENES | |||||
| 17305327 | Gm3543 | −0.57 | −1.48 | −4.4 | 0.022 |
| 17435725 | Il6 | −0.57 | −1.48 | −5.27 | 0.013 |
| 17417858 | Mir1957 | −0.6 | −1.52 | −4.65 | 0.019 |
| 17357688 | Ms4a6b | −0.57 | −1.48 | −3.38 | 0.043 |
| 17337580 | Olfr111 | −0.54 | −1.45 | −3.89 | 0.030 |
| 17362555 | Stxbp3b | −0.77 | −1.71 | −6.5 | 0.007 |
3.2. Thyroxine Treatment Increases Sfpr4 Expression in Relevant Cells
Confirmation of Sfrp4 mRNA upregulation after thyroxine exposure was interrogated using murine suture derived primary cells, murine bone marrow-derived stem cells (BMSCs), myoblasts (C2C12s) and murine calvarial pre-osteoblasts (MC3T3-E1s). These cells are likely relevant to mesenchymal cells in general and the dynamic mammalian perisutural tissue area comprising several cell types including fibroblasts, stem cells, and pre-osteoblasts. Data suggest upregulation of Sfrp4 mRNA in all cell types at 6 hours, 1 day, 3 days, and 7 days in culture with thyroxine treatment. Greatest change in relative expression was observed for the pre-osteoblast cells (MC3T3-E1), peaking at an almost 10-fold increase at the 6-hour time point and an 8-fold increase at 7-day time point (Figure 1).
Figure 1:
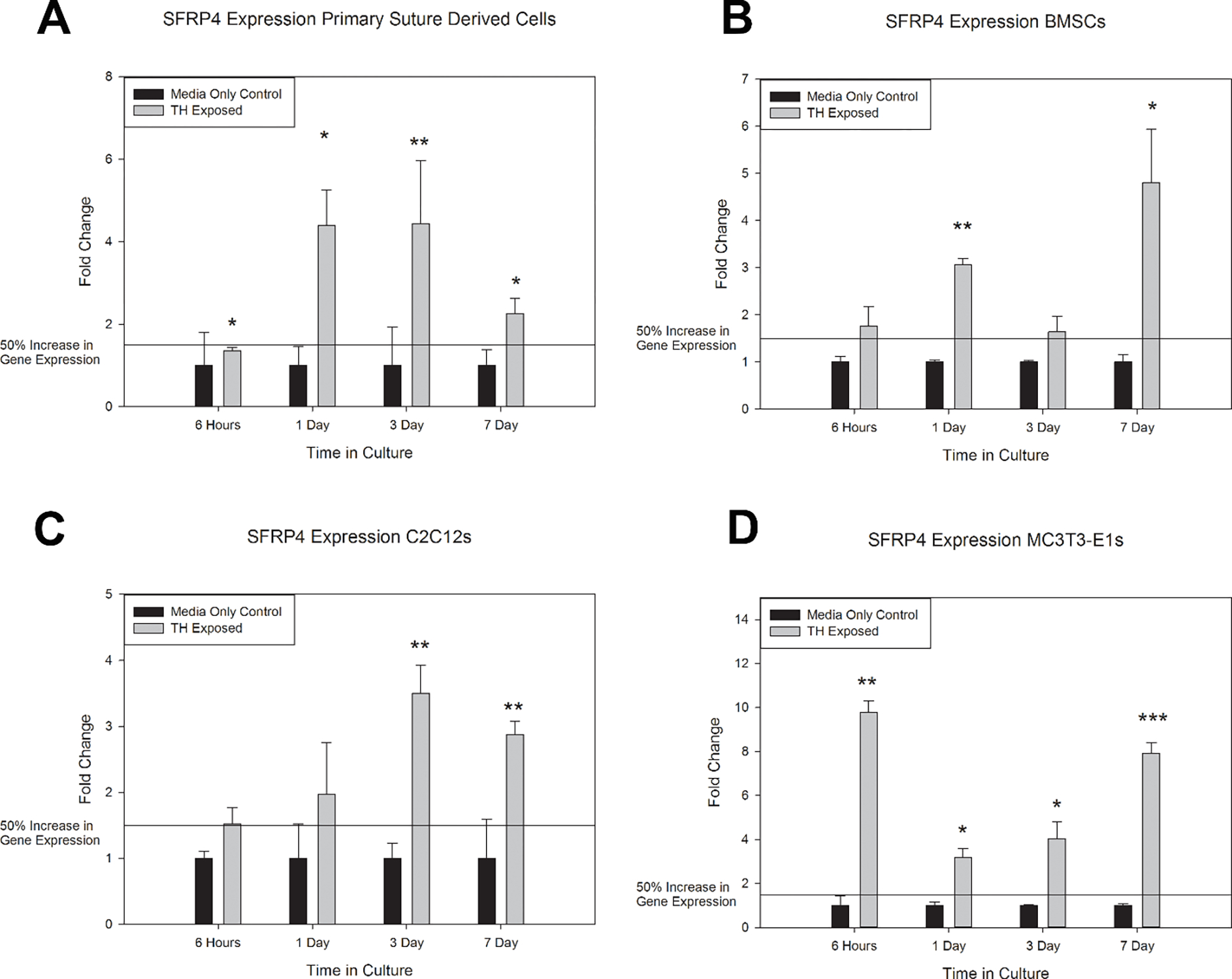
In Vitro mRNA Responses to Thyroxine Treatment. Results are represented by fold change and 50% increase of gene expression noted with reference line. A) Primary suture derived cells responded to thyroxine (TH) exposure by statistically significant increases in Sfrp4 mRNA expression at all time points. B) Bone marrow mesenchymal stem cells (BMSCs) responded to thyroxine exposure by a greater than 50% increase in Sfrp4 mRNA expression at all time points and statistically significant increase at 1 and 7 days. C) Murine myofibroblast cell line C2C12 responded to thyroxine exposure by a greater than 50% increase in Sfrp4 mRNA expression at all time points and statistically significant increase at 3 and 7 days. D) Murine pre-osteoblast cell line MC3T3-E1 responded to thyroxine exposure by statistically significant increases in Sfrp4 mRNA expression at all timepoints. * p<0.05, ** p<0.01, *** p<0.001. Represented as mean +/− SE. n=3 per comparison.
3.3. Sfrp4 Protein Concentrations Differs by Cell Type
Protein levels were similarly interrogated in murine cells including the primary suture derived cells, BMSCs, C2C12, and MC3T3-E1. ELISA was used to quantify the amounts of Sfrp4 in the lysates and supernatant after 24 hours of thyroid hormone treatment to capture protein levels when Sfrp4 mRNA expression was significantly increased in relevant cell types due to treatment with Thyroxine. Suture-derived primary cells harbored very high Sfrp4 content in the lysate samples. BMSCs also expressed relatively high amounts of Sfrp4, whereas C2C12 and MC3T3-E1 had less Sfrp4 in both lysates and supernatants. The MC3T3-E1s were shown to have a much smaller concentration of Sfrp4 in both the lysate and supernatant. Statistical analysis demonstrated significant differences by cell type (p<0.001) where both primary suture derived cells and BMSCs had greater concentrations of Sfrp4 than C2C12 and MC3T3-E1s (p<0.001 for all comparisons). There were no significant differences for primary suture derived cell to BMSC, or C2C12 to MC3T3-E1 comparisons. There were two significant statistical interactions, cell by sample type (p<0.001) and sample type by treatment. Data suggest BMSCs and C2C12 have no discernable difference in protein concentration between supernatants and lysates, where MC3T3-E1s appear to have elevated concentration in the supernatant, and primary suture derived cells have greater concentrations in the lysates. Interestingly, thyroid hormone treatment was observed to decrease the concentration of Sfrp4 in the lysates and remained similar in the supernatant where all other comparisons are collapsed (Figure 2, Supplementary Figure 1, Supplementary Figure 2).
Figure 2:
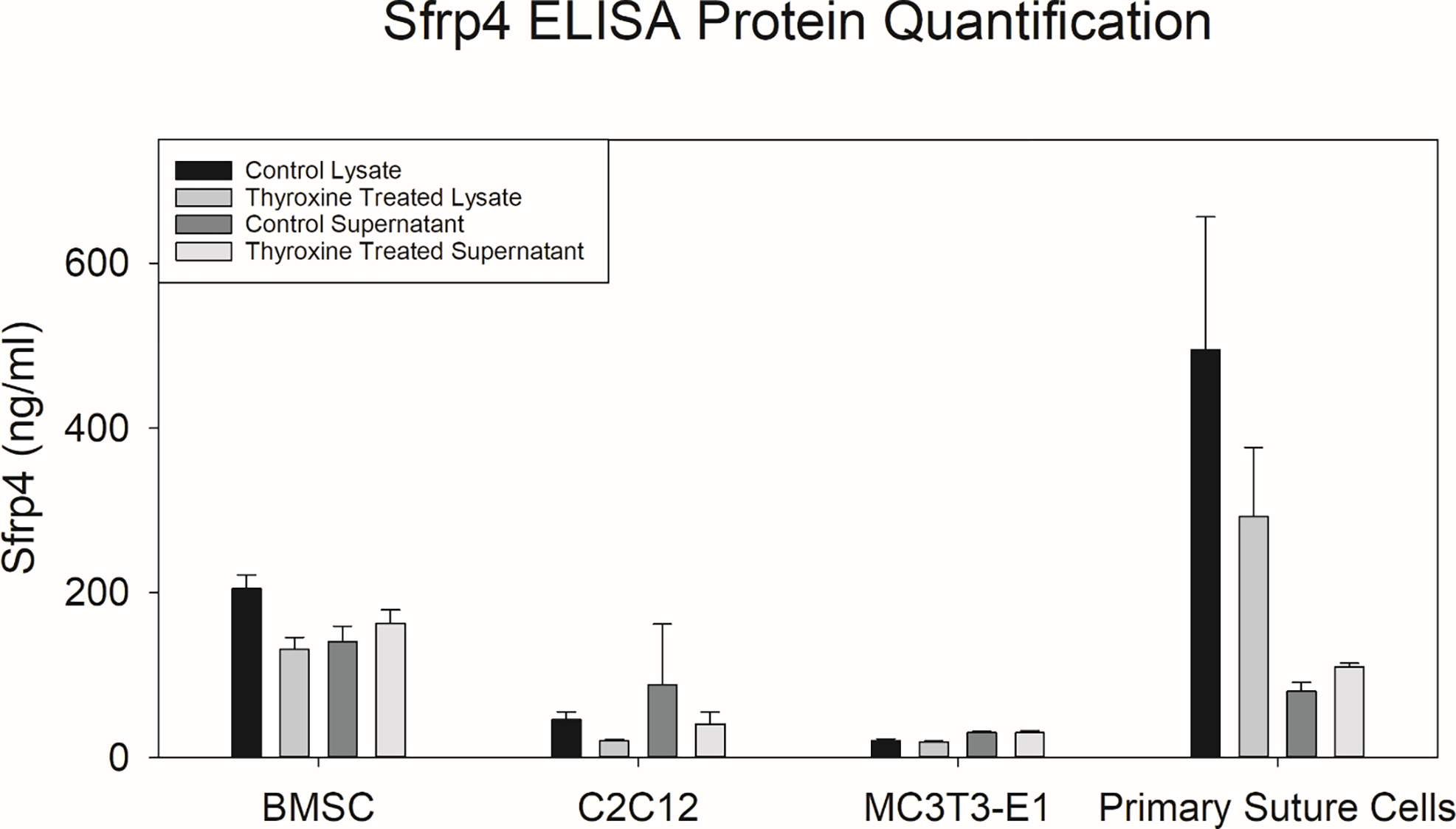
In Vitro Protein Responses to Thyroxine Treatment. For each cell type ELISA was performed 24 hours after thyroxine treatment. Statistical analysis was conducted using a three-way ANOVA for sample type (Lysate or Supernatant), by cell type (BMSC, C2C12, MC3T3-E1 and primary derived suture cells), and by treatment (Control or Thyroxine Treated) after rank-order transformation. There was a significant difference in Sfrp4 concentration by cell type (p<0.001), with post-hoc analysis suggesting segregating of BMSCs and primary suture cells having greater Sfrp4 concentrations than both C2C12s, and MC3T3-E1s (p<0.001 for all comparisons respectively). Two significant interactions were also observed cell type by sample type (p<0.001) and sample type by treatment (p=0.049). Note that the interaction effect for cell type by sample type is driven by no difference observed between lysate and supernatant concentrations for BMSCs and primary suture cells, MC3T3-E1s showing greater Sfrp4 concentration in the supernatant, and C2C12 showing greater concentration in the lysate. Note that the interaction effect for sample type by treatment is driven by decreases in thyroxine treated lysates where no change is observed in the supernatant. (For Sub analysis of Interactions see Supplementary Figure 1).
3.4. The Promoter Region of SFRP4 Harbors Several Binding Sites for Thyroid Receptors
Bioinformatic interrogation of the promoter region of Sfrp4 revealed three areas of relatively conservative consensus sequence (murine and human) that represent putative binding sites for thyroid receptors (Figure 3). These data suggest a potential direct interaction between thyroid hormone and Sfrp4 transcription.
Figure 3:
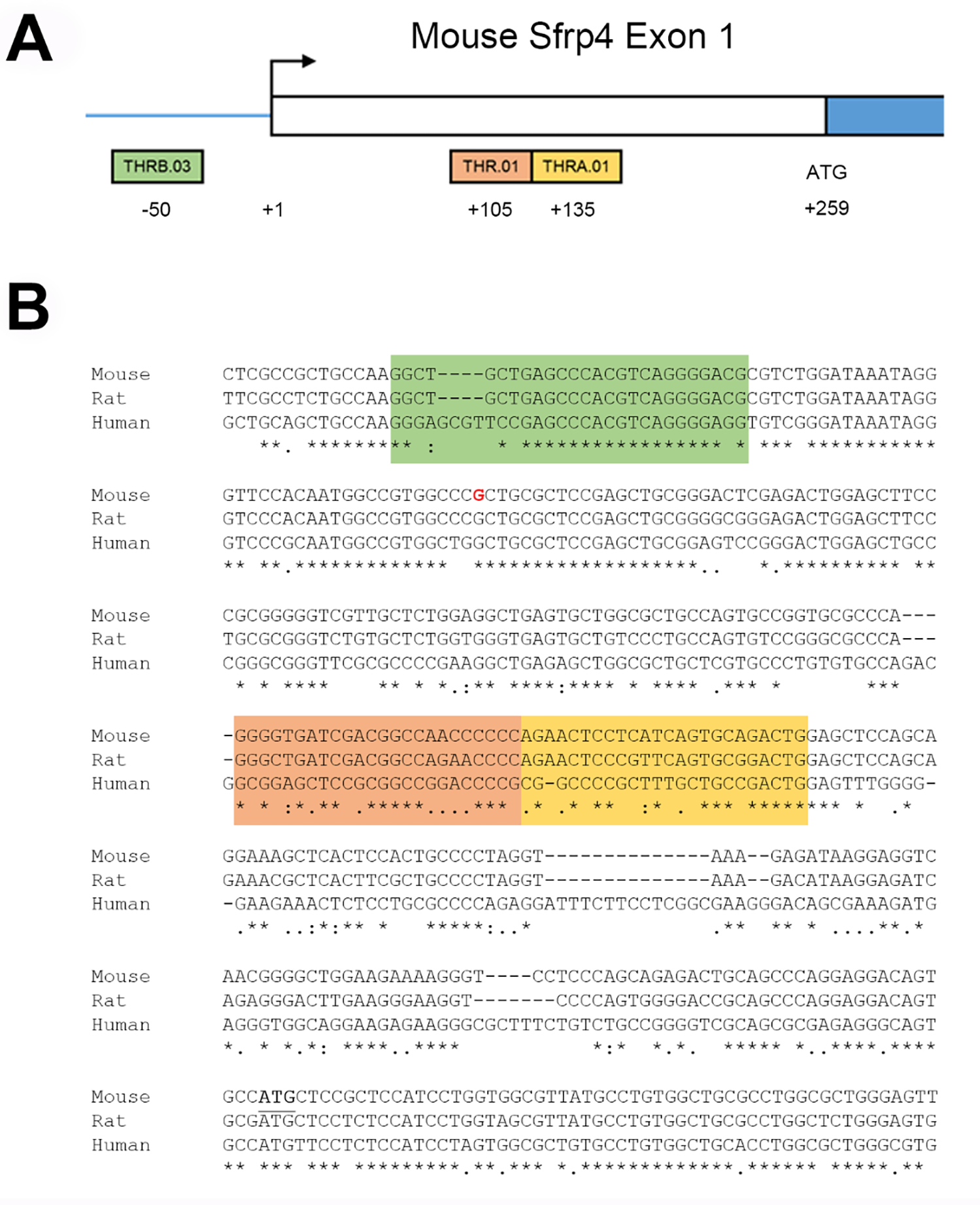
Thyroid Hormone Response Elements in Sfrp4 Promoter. Sfrp4 exon 1 structure (A) and sequence (B) with the transcription start site (base shown in red) are adapted from RefSeq mRNA sequence NM_016687 mapped onto mouse genome build mm39. Sequences consistent with thyroid hormone response elements were identified in the murine Sfrp4 promoter region. There are three canonical thyroid hormone receptor (THR) binding sites (family V$RXRF, matrix V$THR), with one site upstream of the transcription start site (THRB.03, −50 (green)) and two sites downstream of the start site (THR.01, +105 (orange); THRA.01, +135(yellow)). Analysis of orthologous sequences in rat and human Sfrp4 genes showed the upstream site was most highly conserved, having 100% identity with rat and 84.6% identity with human. The transcription start codon is underlined for mouse Sfrp4.
3.5. Reporter Assay and EMSA Suggest Direct Effects of Thyroid Hormone on Sfrp4
Luciferase reporter assays demonstrated increased activity of the Sfrp4 promoter in MC3T3-E1 cells treated with thyroid hormone for 6 hours in vitro (p=0.031). These data suggest an affinity for direct transcriptional effects on the Sfrp4 promoter region due to exogenous thyroid hormone treatments (Figure 4A). To confirm binding of thyroid hormone receptors to their putative sites in the Sfrp4 promoter, EMSA was performed using nuclear extracts from untreated and thyroxine treated cells. Figure 4B show that extracts from MC3T3-E1 cells produced a specific shift of the upstream Sfrp4 probe that is supershifted with addition of an antibody to the thyroid hormone receptor THRβ. Similar results were obtained from MC3T3-E1 extracts and the downstream Sfrp4 probe (Figure 4C). Notably, extracts from thyroxine-treated MC3T3-E1s engendered a more intense shift of the upstream probe than extracts from untreated cells, and this was seen in replicates from three independently isolated sets of extracts (not shown). This consistent increase in binding was not produced by any other extract/probe combination.
Figure 4:
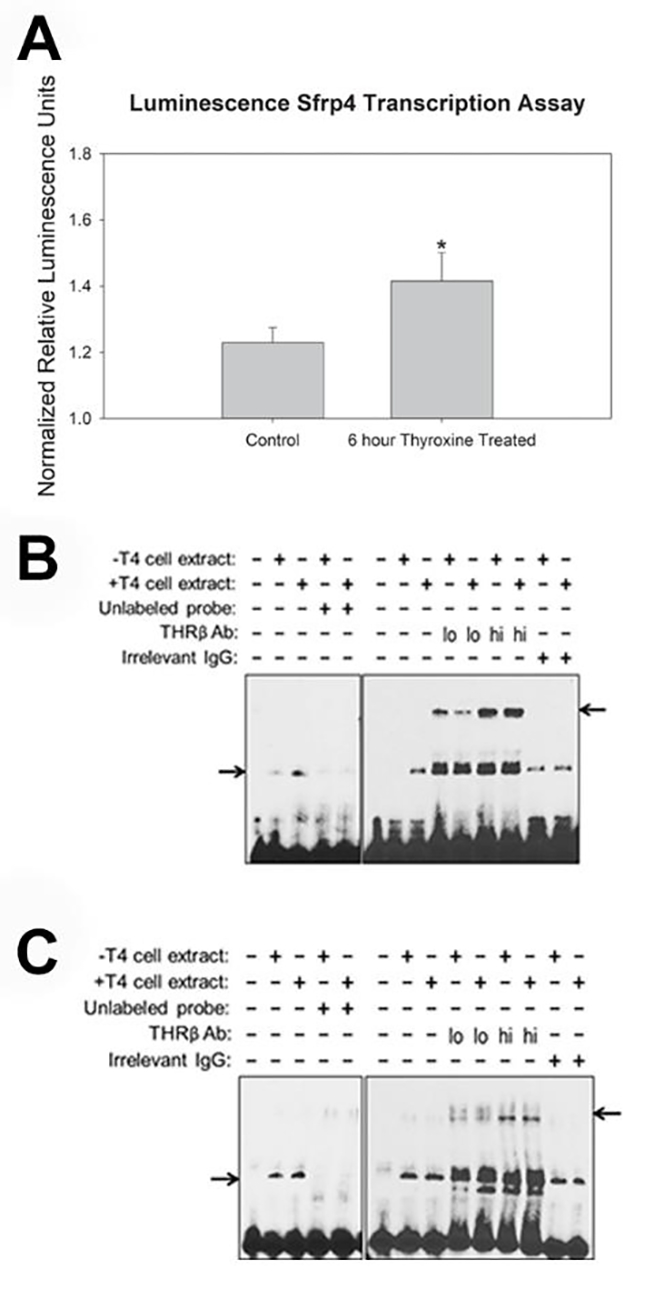
Increased SFRP4 Transcription and Binding of Thyroid Hormone Receptor to Sfrp4 Promoter Region as a Result of Thyroid Hormone Exposure. A) Note significantly increased transcription evidenced by the luciferase assay for MC3T3-E1 cells after thyroid hormone treatment for 6 hours. * p<0.031 Represented as mean +/− SE, one tailed student t-test, n=4 (triplicates x triplicated per plate, 4 plates). B, C) For all panels, the five left lanes demonstrate a specific shift caused by addition of nuclear extracts from cells treated with standard growth media or media supplemented with thyroxine (T4) at a concentration of 10−6 mol/liter to putative THR binding sites in Sfrp4. Specificity is shown by absence of signal upon addition of unlabeled probe to the reaction. The remaining lanes demonstrate a supershift upon addition of low or high amounts of antibody against THRβ, but a lack of supershift upon addition of irrelevant IgG. Panels show the following combinations of nuclear extracts and probes: (B) MC3T3-E1 extracts + upstream probe; (C) MC3T3-E1 extracts + downstream probe. Lower and upper arrows indicate the probe shifts and supershifts, respectively.
3.6. Alkaline Phosphatase Increases after Thyroxine Treatment
Alkaline phosphatase is an early marker of osteoblastogenesis. Figure 5A confirms that only pre-osteoblasts (MC3T3-E1) increase ALP activity after thyroxine treatment. Further, when physiological levels of recombinant SFRP4 protein are added to the thyroxine-laden media, alkaline phosphatase levels are partially inhibited, suggesting that SFRP4 can inhibit thyroxine-driven osteoblastogenesis in differentiating bone cells (Figure 5B).
Figure 5:
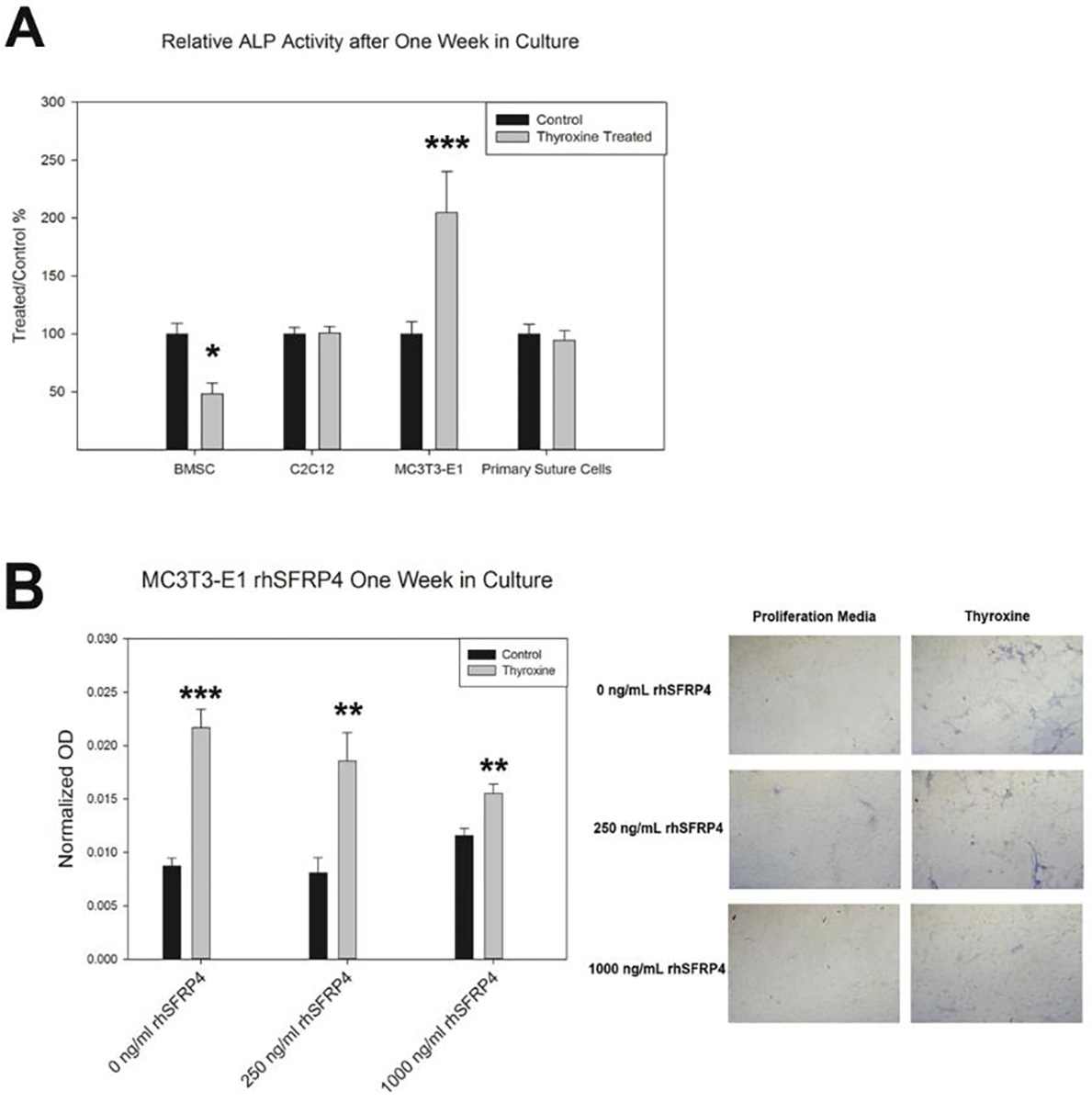
ALP Response to Thyroxine Treatment. A) Only MC3T3-E1s are stimulated to increased ALP activity by thyroxine treatment in vitro (p<0.001). Note decreases in basal ALP activity for BMSCs (p=0.0153). B) MC3T3-E1s responded to rhSFRP4 treatment with decreased ALP activity (Quantified, right, Qualitatively less blue staining, left); * p<0.05, * p<0.01, ** p<0.001. Data compared control to treated samples by t-test. n=3 per comparison. Represented as mean +/− SE.
4. Discussion
Here we sought to determine if treatment with exogenous thyroid hormone could alter Sfrp4 expression in cells that are relevant to growth and development of the craniofacial skeleton in order to confirm Sfrp4 a novel gene of interest for thyroid hormone driven growth disorders. We further sought to determine if SFRP4 could act in regulation of osteoblast activity. Our data have confirmed that thyroid hormone exposure upregulates mRNA expression of Sfrp4 in all cell types tested. This is supported by molecular and protein data suggesting the presence of thyroid receptor binding sites within the promoter region of this gene. Further, we have now demonstrated that increased Sfrp4 transcription should be expected as a result of thyroid hormone exposure evidenced by transfection study and specificity of binding of thyroid hormone via response elements evidenced by EMSA.
Interestingly, despite similarity in increases in mRNA transcription of Sfrp4 after thyroid hormone treatment, there is wide variability between overall SFPR4 protein levels in the cells studied here. Our heterogeneous primary suture derived cells exhibited substantial Sfrp4 in the lysate, BMSC showed a marked amount, C2C12 myoblasts had less, and pre-osteoblast MC3T3-E1s had the least protein concentrations. Only when comparison groups were collapsed did we see alterations in Sfrp4 in only the lysates after thyroid hormone treatment. These data suggest that, in the case of this gene product, transcriptional upregulation does not necessarily equate to greater steady-state protein levels. Such uncoupling of gene and protein expression can be the result of the stability of both RNA transcripts and proteins as well as interference by μRNA. The data concerning alkaline phosphatase suggest thyroxine only increases osteoblastogenesis in pre-osteoblasts from the calvarium and are met with little change or even decreases in ALP activity in stem cells, myoblasts, and primary cells. This correlates with our previously published data indicating no change in proliferation in primary cells and an increase in proliferation followed by an increase in differentiation in MC3T3-E1 cells [21,24]. Further this validates bone lineage cells as particularly susceptible to changes in Sfrp4 expression even when expression changes are driven by exogenous thyroid hormone[40–42]. Given the short course of treatment and the lack of treatment with an osteogenic differentiation media for cells not primed for osteogenesis a lack of differentiation marked by ALP activity is expected. However, the higher levels of SFRP4 protein especially in the stem cell and primary suture derived cells used here may suggest and important role for WNT antagonists in thyroxine driven osteoblastogenesis[40–42]. We attempted to address this paradigm and find consistent data as the calvarial pre-osteoblasts express absolutely lower levels of SFRP4, and adding additional recombinant protein at least partially blocked or reduce alkaline phosphatase activity.
The SFPR4 gene has been implicated in several disease processes including Pyle’s disease [43–45], various cancers [46–50], and potentiation of adipogenesis [51–57]. The identification of overexpression of a WNT antagonist (SFRP4), in human cases of craniosynostosis [28] is puzzling as WNT antagonism, in theory, would reduce or at least attenuate any hyper-osteogenic processes that lead to disease[58,59]. Our confirmation of upregulation of this target in sutural tissue from mice treated in utero with levothyroxine, which concomitantly disrupts and alters calvarial growth, was suggestive of a potential interaction between thyroid hormone and craniofacial development, with the gene SFRP4 playing a potential role [25]. Importantly, the interaction of Sfrp4 and bone is complex and bone type specific. For example, a deficiency of Sfrp4 expression as seen in Pyle’s disease results in increased trabecular bone and decreased cortical bone demonstrating that the same level of Sfrp4 can have differing effects on the type and location of bone lineage cells[40–42,45].
Our data suggest upregulation of the Sfrp4 similar to that seen in the human patients and mouse models[40–42,45,59], but not a translative effect in total protein. Only those cells primed for differentiation to osteoblast phenotype respond to thyroxine by increasing alkaline phosphatase activity, a critical marker of osteoblastogenesis. Thus, what may be at work here are several mechanisms that will require further studies. Firstly, the incongruence between gene expression and protein resulting in phenotypic alterations in growth could be explained by several intervening mechanisms including RNA and/or protein stability and post-transcription regulation such as potential micro-RNAs (preliminary interrogated candidates include 124-3p, 125b-3p, 135a-5p, 511-5p). Secondly, a key component in maintaining normal developmental processes after thyroid hormone insult may be the differential level of this target between the participating cells.
Based on available data, it is likely that cross talk between different cell types (stem and progenitor cells, pre-osteoblasts, osteoblasts, fibroblasts) exists within and around the calvarial sutures [22,60–64]. As Sfrp4 is a secreted factor, it is possible that low-Sfrp4 producing and containing cells (e.g. pre-osteoblasts) are affected by exposure to the higher amount of Sfrp4 produced by stem cells and fibroblasts in a growth niche such as the cranial suture. Interestingly, our primary cells showed the highest amount of SFRP4, which could be explained by the mixed cell population which is still only naively characterized (i.e. suture mesenchymal niche) [22]. Thus, lack of thyroid homeostasis may not be sufficient to cause a disorder like craniosynostosis but would more likely drive alterations by gene-environment interactions supported here by data that suggest SFRP4 can at least partially alter osteoblastogenesis. Gene alterations that prime or allow for greater numbers of osteoblasts within the sutures (i.e. Twist, Fgfr) might be highly susceptible to interactive or additive effects of exogenous thyroid hormone as replacement or in disease.
Overall, these data contribute to future directions related to the defining the increased risk of craniosynostosis concurrent with maternal and endogenous thyroid hormone disorders [65–71]. Our group and others have identified the presence of thyroid receptors within the calvaria, allowing for interpretation of a direct effect [23,25,72]. However, our data previously published on ex vivo expression after in utero exposures [25] and available human patient expression data [28] lead to the question of why targets within the WNT pathway, particularly antagonists, would materialize within the craniosynostosis spectrum. Here we confirm that Sfrp4 is a transcriptional target of thyroid hormone due to the presence of binding sites for the receptor in the promoter region. However, the lack of alterations in translation assessed by protein concentrations begs the question of its role in hyperostosis disorders. The susceptible cell type here appears to be the pre-osteoblast, with its sensitivity to thyroid hormone resulting in increased alkaline phosphatase activity. The ability for SFRP4 to decrease osteoblast activity does allow for the design of downstream future experimentation aimed at understanding gene/environment interactions and potential therapeutic interventions.
5. Conclusion
Here we confirmed that Sfrp4 mRNA is upregulated by exposure to exogenous thyroid hormone. Further alterations to protein production were not overtly observed but concentration differences by cell type were informative. However, our data do suggest a direct effect of thyroid hormone on Sfrp4 expression and specificity of binding to the promoter region of this gene. In the case of the pre-osteoblast, when exposed to exogenous excess SFRP4, thyroid hormone driven alkaline phosphatase activity is decreased. This target, identified as overexpressed in patients with craniosynostosis and now modeled in preclinical in vivo and in vitro experiments, warrants further study.
Supplementary Material
6. Acknowledgments
The authors would like to thank Xing Ming Shi for isolation of the bone marrow mesenchymal cells. The following funding sources were used for this study: Cleft Palate Foundation Grant, NIH/NIDCR R03DE023350A, 5T32DE017551, and F31DE026684, NIH/NCATS UL1 TR000062, NIH/NIGM P30GM103331, GM103342 and GM103499, and MUSC’s Office of the Vice President for Research.
Footnotes
Conflict of Interest Statement
The authors declare that there are no conflicts of interest.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
10. Data Availability Statement
The datasets generated for this study can be found in the NCBI GEO under accession GSE66149.
7 References
- [1].Wojcicka A, Bassett JHD, Williams GR, Mechanisms of action of thyroid hormones in the skeleton, Biochim Biophys Acta. 1830 (2013) 3979–3986. 10.1016/j.bbagen.2012.05.005. [DOI] [PubMed] [Google Scholar]
- [2].Bassett JHD, Williams GR, Role of Thyroid Hormones in Skeletal Development and Bone Maintenance, Endocrine Reviews. 37 (2016) 135–187. 10.1210/er.2015-1106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Gouveia CHA, Miranda-Rodrigues M, Martins GM, Neofiti-Papi B, Thyroid Hormone and Skeletal Development, Vitam Horm. 106 (2018) 383–472. 10.1016/bs.vh.2017.06.002. [DOI] [PubMed] [Google Scholar]
- [4].Williams GR, Bassett JHD, Thyroid diseases and bone health, Endocrinol Invest. 41 (2018) 99–109. 10.1007/s40618-017-0753-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Bassett JHD, Boyde A, Zikmund T, Evans H, Croucher PI, Zhu X, Park JW, Cheng S, Williams GR, Thyroid hormone receptor α mutation causes a severe and thyroxine-resistant skeletal dysplasia in female mice, Endocrinology. 155 (2014) 3699–3712. 10.1210/en.2013-2156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].Pitetti J-L, Calvel P, Zimmermann C, Conne B, Papaioannou MD, Aubry F, Cederroth CR, Urner F, Fumel B, Crausaz M, Docquier M, Herrera PL, Pralong F, Germond M, Guillou F, Jégou B, Nef S, An Essential Role for Insulin and IGF1 Receptors in Regulating Sertoli Cell Proliferation, Testis Size, and FSH Action in Mice, Mol Endocrinol. 27 (2013) 814–827. 10.1210/me.2012-1258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [7].Robson H, Siebler T, Shalet SM, Williams GR, Interactions between GH, IGF-I, Glucocorticoids, and Thyroid Hormones during Skeletal Growth, Pediatr Res. 52 (2002) 137–147. 10.1203/00006450-200208000-00003. [DOI] [PubMed] [Google Scholar]
- [8].Salani B, Ravera S, Amaro A, Salis A, Passalacqua M, Millo E, Damonte G, Marini C, Pfeffer U, Sambuceti G, Cordera R, Maggi D, IGF1 regulates PKM2 function through Akt phosphorylation, Cell Cycle. 14 (2015) 1559–1567. 10.1080/15384101.2015.1026490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [9].Wang Y, Zhang S, Expression and regulation by thyroid hormone (TH) of zebrafish IGF-I gene and amphioxus IGFl gene with implication of the origin of TH/IGF signaling pathway, Comp Biochem Physiol A Mol Integr Physiol. 160 (2011) 474–479. 10.1016/j.cbpa.2011.08.005. [DOI] [PubMed] [Google Scholar]
- [10].Bassett JHD, Williams GR, The skeletal phenotypes of TRalpha and TRbeta mutant mice, J Mol Endocrinol. 42 (2009) 269–282. 10.1677/JME-08-0142. [DOI] [PubMed] [Google Scholar]
- [11].Fonseca TL, Teixeira MBCG, Rodrigues-Miranda M, Silva MV, Martins GM, Costa CC, D.Y. [UNIFESP Arita, J.D. [UNIFESP Perez, D.E. [UNIFESP Casarini, P.C. Brum, C.H.A. Gouveia, Thyroid hormone interacts with the sympathetic nervous system to modulate bone mass and structure in young adult mice, (2014). 10.1152/ajpendo.00643.2013. [DOI] [PubMed] [Google Scholar]
- [12].Gu WX, Stern PH, Madison LD, Du GG, Mutual up-regulation of thyroid hormone and parathyroid hormone receptors in rat osteoblastic osteosarcoma 17/2.8 cells, Endocrinology. 142 (2001) 157–164. 10.1210/endo.142.1.7905. [DOI] [PubMed] [Google Scholar]
- [13].Huang BK, Golden LA, Tarjan G, Madison LD, Stern PH, Insulin-like growth factor I production is essential for anabolic effects of thyroid hormone in osteoblasts, J Bone Miner Res. 15 (2000) 188–197. 10.1359/jbmr.2000.15.2.188. [DOI] [PubMed] [Google Scholar]
- [14].Lakatos P, Foldes J, Nagy Z, Takacs I, Speer G, Horvath C, Mohan S, Baylink DJ, Stern PH, Serum insulin-like growth factor-I, insulin-like growth factor binding proteins, and bone mineral content in hyperthyroidism, Thyroid. 10 (2000) 417–423. 10.1089/thy.2000.10.417. [DOI] [PubMed] [Google Scholar]
- [15].Ramajayam G, Vignesh RC, Karthikeyan S, Senthil Kumar K, Karthikeyan GD, Veni S, Sridhar M, Arunakaran J, Michael Aruldhas M, Srinivasan N, Regulation of insulin-like growth factors and their binding proteins by thyroid stimulating hormone in human osteoblast-like (SaOS2) cells, Mol Cell Biochem. 368 (2012) 77–88. 10.1007/s11010-012-1345-4. [DOI] [PubMed] [Google Scholar]
- [16].Sampath TK, Simic P, Sendak R, Draca N, Bowe AE, O’Brien S, Schiavi SC, McPherson JM, Vukicevic S, Thyroid-stimulating hormone restores bone volume, microarchitecture, and strength in aged ovariectomized rats, J Bone Miner Res. 22 (2007) 849–859. 10.1359/jbmr.070302. [DOI] [PubMed] [Google Scholar]
- [17].Bassett JHD, Williams GR, The molecular actions of thyroid hormone in bone, Trends Endocrinol Metab. 14 (2003) 356–364. 10.1016/s1043-2760(03)00144-9. [DOI] [PubMed] [Google Scholar]
- [18].Britto JM, Fenton AJ, Holloway WR, Nicholson GC, Osteoblasts mediate thyroid hormone stimulation of osteoclastic bone resorption, Endocrinology. 134 (1994) 169–176. 10.1210/endo.134.1.8275930. [DOI] [PubMed] [Google Scholar]
- [19].Cordero DR, Brugmann S, Chu Y, Bajpai R, Jame M, Helms JA, Cranial neural crest cells on the move: their roles in craniofacial development, Am J Med Genet A. 155A (2011) 270–279. 10.1002/ajmg.a.33702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Williams GR, Thyroid Hormone Actions in Cartilage and Bone, Eur Thyroid J. 2 (2013) 3–13. 10.1159/000345548. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Cray JJ, Khaksarfard K, Weinberg SM, Elsalanty M, Yu JC, Effects of Thyroxine Exposure on Osteogenesis in Mouse Calvarial Pre-Osteoblasts, PLOS ONE. 8 (2013) e69067. 10.1371/journal.pone.0069067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Durham E, Howie RN, Larson N, LaRue A, Cray J, Pharmacological exposures may precipitate craniosynostosis through targeted stem cell depletion, Stem Cell Res. 40 (2019) 101528. 10.1016/j.scr.2019.101528. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Durham E, Howie RN, Parsons T, Bennfors G, Black L, Weinberg SM, Elsalanty M, Yu JC, Cray JJ, Thyroxine exposure effects on the cranial base, Calcif Tissue Int. 101 (2017) 300–311. 10.1007/s00223-017-0278-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Durham EL, Howie RN, Black L, Bennfors G, Parsons TE, Elsalanty M, Yu JC, Weinberg SM, Cray JJ , Effects of thyroxine exposure on the Twist 1 +/− phenotype: A test of gene-environment interaction modeling for craniosynostosis, Birth Defects Res A Clin Mol Teratol. 106 (2016) 803–813. 10.1002/bdra.23543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Howie RN, Durham EL, Black L, Bennfors G, Parsons TE, Elsalanty ME, Yu JC, Weinberg SM, Cray JJ, Effects of In Utero Thyroxine Exposure on Murine Cranial Suture Growth, PLoS One. 11 (2016) e0167805. 10.1371/journal.pone.0167805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Kesterke MJ, Judd MA, Mooney MP, Siegel MI, Elsalanty M, Howie RN, Weinberg SM, Cray JJ, Maternal environment and craniofacial growth: geometric morphometric analysis of mandibular shape changes with in utero thyroxine overexposure in mice, Journal of Anatomy. 233 (2018) 46–54. 10.1111/joa.12810. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Monroe DG, McGee-Lawrence ME, Oursler MJ, Westendorf JJ, Update on Wnt signaling in bone cell biology and bone disease, Gene. 492 (2012) 1–18. 10.1016/j.gene.2011.10.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [28].Stamper BD, Park SS, Beyer RP, Bammler TK, Farin FM, Mecham B, Cunningham ML, Differential Expression of Extracellular Matrix-Mediated Pathways in Single-Suture Craniosynostosis, PLOS ONE. 6 (2011) e26557. 10.1371/journal.pone.0026557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Durham EL, Howie RN, Cray JJ, Gene/environment interactions in craniosynostosis: A brief review, Orthod Craniofac Res. 20 Suppl 1 (2017) 8–11. 10.1111/ocr.12153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [30].Delahaye S, Bernard JP, Rénier D, Ville Y, Prenatal ultrasound diagnosis of fetal craniosynostosis, Ultrasound Obstet Gynecol. 21 (2003) 347–353. 10.1002/uog.91. [DOI] [PubMed] [Google Scholar]
- [31].Robin NH, Molecular genetic advances in understanding craniosynostosis, Plast Reconstr Surg. 103 (1999) 1060–1070. [PubMed] [Google Scholar]
- [32].Maruyama T, Mirando AJ, Deng C-X, Hsu W, The balance of WNT and FGF signaling influences mesenchymal stem cell fate during skeletal development, Sci Signal. 3 (2010) ra40. 10.1126/scisignal.2000727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Wilkie AOM, Johnson D, Wall SA, Clinical genetics of craniosynostosis, Curr Opin Pediatr. 29 (2017) 622–628. 10.1097/MOP.0000000000000542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [34].Twigg SRF, Wilkie AOM, A genetic-pathophysiological framework for craniosynostosis, The American Journal of Human Genetics. 97 (2015) 359–377. 10.1016/j.ajhg.2015.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [35].Johnson D, Wilkie AOM, Craniosynostosis, Eur J Hum Genet. 19 (2011) 369–376. 10.1038/ejhg.2010.235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Kilkenny C, Browne WJ, Cuthill IC, Emerson M, Altman DG, Improving bioscience research reporting: The ARRIVE guidelines for reporting animal research, J Pharmacol Pharmacother. 1 (2010) 94–99. 10.4103/0976-500X.72351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [37].Zhang W, Ou G, Hamrick M, Hill W, Borke J, Wenger K, Chutkan N, Yu J, Mi Q-S, Isales CM, Shi X-M, Age-related changes in the osteogenic differentiation potential of mouse bone marrow stromal cells, J Bone Miner Res. 23 (2008) 1118–1128. 10.1359/jbmr.080304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [38].Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG, Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega, Mol Syst Biol. 7 (2011) 539. 10.1038/msb.2011.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Yuan JS, Reed A, Chen F, Stewart CN, Statistical analysis of real-time PCR data, BMC Bioinformatics. 7 (2006) 85. 10.1186/1471-2105-7-85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Haraguchi R, Kitazawa R, Mori K, Tachibana R, Kiyonari H, Imai Y, Abe T, Kitazawa S, sFRP4-dependent Wnt signal modulation is critical for bone remodeling during postnatal development and age-related bone loss, Sci Rep. 6 (2016) 25198. 10.1038/srep25198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [41].Nakanishi R, Shimizu M, Mori M, Akiyama H, Okudaira S, Otsuki B, Hashimoto M, Higuchi K, Hosokawa M, Tsuboyama T, Nakamura T, Secreted Frizzled-Related Protein 4 Is a Negative Regulator of Peak BMD in SAMP6 Mice, Journal of Bone and Mineral Research. 21 (2006) 1713–1721. 10.1359/jbmr.060719. [DOI] [PubMed] [Google Scholar]
- [42].Nakanishi R, Akiyama H, Kimura H, Otsuki B, Shimizu M, Tsuboyama T, Nakamura T, Osteoblast-Targeted Expression of Sfrp4 in Mice Results in Low Bone Mass, Journal of Bone and Mineral Research. 23 (2008) 271–277. 10.1359/jbmr.071007. [DOI] [PubMed] [Google Scholar]
- [43].Arboleya L, Queiro R, Alperi M, Lorenzo JA, Ballina J, Pyle’s Disease: A human model of differentiated cortical and trabecular homeostasis, Reumatol Clin (Engl Ed). 16 (2020) 56–58. 10.1016/j.reuma.2018.01.002. [DOI] [PubMed] [Google Scholar]
- [44].Galada C, Shah H, Shukla A, Girisha KM, A novel sequence variant in SFRP4 causing Pyle disease, J Hum Genet. 62 (2017) 575–576. 10.1038/jhg.2016.166. [DOI] [PubMed] [Google Scholar]
- [45].Kiper POS, Saito H, Gori F, Unger S, Hesse E, Yamana K, Kiviranta R, Solban N, Liu J, Brommage R, Boduroglu K, Bonafé L, Campos-Xavier B, Dikoglu E, Eastell R, Gossiel F, Harshman K, Nishimura G, Girisha KM, Stevenson BJ, Takita H, Rivolta C, Superti-Furga A, Baron R, Cortical-Bone Fragility — Insights from sFRP4 Deficiency in Pyle’s Disease, New England Journal of Medicine. 374 (2016) 2553–2562. 10.1056/NEJMoa1509342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [46].Deshmukh A, Arfuso F, Newsholme P, Dharmarajan A, Regulation of Cancer Stem Cell Metabolism by Secreted Frizzled-Related Protein 4 (sFRP4), Cancers (Basel). 10 (2018) E40. 10.3390/cancers10020040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [47].Deshmukh A, Kumar S, Arfuso F, Newsholme P, Dharmarajan A, Secreted Frizzled-related protein 4 (sFRP4) chemo-sensitizes cancer stem cells derived from human breast, prostate, and ovary tumor cell lines, Sci Rep. 7 (2017) 2256. 10.1038/s41598-017-02256-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [48].Ghoshal A, Ghosh SS, Antagonizing canonical Wnt signaling pathway by recombinant human sFRP4 purified from E. coli and its implications in cancer therapy, Mol Cell Biochem. 418 (2016) 119–135. 10.1007/s11010-016-2738-6. [DOI] [PubMed] [Google Scholar]
- [49].Han X, Saiyin H, Zhao J, Fang Y, Rong Y, Shi C, Lou W, Kuang T, Overexpression of miR-135b-5p promotes unfavorable clinical characteristics and poor prognosis via the repression of SFRP4 in pancreatic cancer, Oncotarget. 8 (2017) 62195–62207. 10.18632/oncotarget.19150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [50].Sandsmark E, Andersen MK, Bofin AM, Bertilsson H, Drabløs F, Bathen TF, Rye MB, Tessem M-B, SFRP4 gene expression is increased in aggressive prostate cancer, Sci Rep. 7 (2017) 14276. 10.1038/s41598-017-14622-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [51].Almario RU, Karakas SE, Roles of circulating WNT-signaling proteins and WNT-inhibitors in human adiposity, insulin resistance, insulin secretion, and inflammation, Horm Metab Res. 47 (2015) 152–157. 10.1055/s-0034-1384521. [DOI] [PubMed] [Google Scholar]
- [52].Batsali AK, Pontikoglou C, Koutroulakis D, Pavlaki KI, Damianaki A, Mavroudi I, Alpantaki K, Kouvidi E, Kontakis G, Papadaki HA, Differential expression of cell cycle and WNT pathway-related genes accounts for differences in the growth and differentiation potential of Wharton’s jelly and bone marrow-derived mesenchymal stem cells, Stem Cell Res Ther. 8 (2017) 102. 10.1186/s13287-017-0555-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [53].Ehrlund A, Mejhert N, Lorente-Cebrián S, Aström G, Dahlman I, Laurencikiene J, Rydén M, Characterization of the Wnt inhibitors secreted frizzled-related proteins (SFRPs) in human adipose tissue, J Clin Endocrinol Metab. 98 (2013) E503–508. 10.1210/jc.2012-3416. [DOI] [PubMed] [Google Scholar]
- [54].Guan H, Zhang Y, Gao S, Bai L, Zhao S, Cheng XW, Fan J, Liu E, Differential Patterns of Secreted Frizzled-Related Protein 4 (SFRP4) in Adipocyte Differentiation: Adipose Depot Specificity, Cell Physiol Biochem. 46 (2018) 2149–2164. 10.1159/000489545. [DOI] [PubMed] [Google Scholar]
- [55].Jeong JY, Kim JS, Nguyen TH, Lee H-J, Baik M, Wnt/β-catenin signaling and adipogenic genes are associated with intramuscular fat content in the longissimus dorsi muscle of Korean cattle, Anim Genet. 44 (2013) 627–635. 10.1111/age.12061. [DOI] [PubMed] [Google Scholar]
- [56].Park J-R, Jung J-W, Lee Y-S, Kang K-S, The roles of Wnt antagonists Dkk1 and sFRP4 during adipogenesis of human adipose tissue-derived mesenchymal stem cells, Cell Prolif. 41 (2008) 859–874. 10.1111/j.1365-2184.2008.00565.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [57].Visweswaran M, Schiefer L, Arfuso F, Dilley RJ, Newsholme P, Dharmarajan A, Wnt Antagonist Secreted Frizzled-Related Protein 4 Upregulates Adipogenic Differentiation in Human Adipose Tissue-Derived Mesenchymal Stem Cells, PLOS ONE. 10 (2015) e0118005. 10.1371/journal.pone.0118005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [58].Estrada K, Styrkarsdottir U, Evangelou E, Hsu Y-H, Duncan EL, Ntzani EE, Oei L, Albagha OME, Amin N, Kemp JP, Koller DL, Li G, Liu C-T, Minster RL, Moayyeri A, Vandenput L, Willner D, Xiao S-M, Yerges-Armstrong LM, Zheng H-F, Alonso N, Eriksson J, Kammerer CM, Kaptoge SK, Leo PJ, Thorleifsson G, Wilson SG, Wilson JF, Aalto V, Alen M, Aragaki AK, Aspelund T, Center JR, Dailiana Z, Duggan DJ, Garcia M, Garcia-Giralt N, Giroux S, Hallmans G, Hocking LJ, Husted LB, Jameson KA, Khusainova R, Kim GS, Kooperberg C, Koromila T, Kruk M, Laaksonen M, Lacroix AZ, Lee SH, Leung PC, Lewis JR, Masi L, Mencej-Bedrac S, Nguyen TV, Nogues X, Patel MS, Prezelj J, Rose LM, Scollen S, Siggeirsdottir K, Smith AV, Svensson O, Trompet S, Trummer O, van Schoor NM, Woo J, Zhu K, Balcells S, Brandi ML, Buckley BM, Cheng S, Christiansen C, Cooper C, Dedoussis G, Ford I, Frost M, Goltzman D, González-Macías J, Kähönen M, Karlsson M, Khusnutdinova E, Koh J-M, Kollia P, Langdahl BL, Leslie WD, Lips P, Ljunggren Ö, Lorenc RS, Marc J, Mellström D, Obermayer-Pietsch B, Olmos JM, Pettersson-Kymmer U, Reid DM, Riancho JA, Ridker PM, Rousseau F, Slagboom PE, Tang NLS, Urreizti R, Van Hul W, Viikari J, Zarrabeitia MT, Aulchenko YS, Castano-Betancourt M, Grundberg E, Herrera L, Ingvarsson T, Johannsdottir H, Kwan T, Li R, Luben R, Medina-Gómez C, Palsson ST, Reppe S, Rotter JI, Sigurdsson G, van Meurs JBJ, Verlaan D, Williams FMK, Wood AR, Zhou Y, Gautvik KM, Pastinen T, Raychaudhuri S, Cauley JA, Chasman DI, Clark GR, Cummings SR, Danoy P, Dennison EM, Eastell R, Eisman JA, Gudnason V, Hofman A, Jackson RD, Jones G, Jukema JW, Khaw K-T, Lehtimäki T, Liu Y, Lorentzon M, McCloskey E, Mitchell BD, Nandakumar K, Nicholson GC, Oostra BA, Peacock M, Pols HAP, Prince RL, Raitakari O, Reid IR, Robbins J, Sambrook PN, Sham PC, Shuldiner AR, Tylavsky FA, van Duijn CM, Wareham NJ, Cupples LA, Econs MJ, Evans DM, Harris TB, Kung AWC, Psaty BM, Reeve J, Spector TD, Streeten EA, Zillikens MC, Thorsteinsdottir U, Ohlsson C, Karasik D, Richards JB, Brown MA, Stefansson K, Uitterlinden AG, Ralston SH, Ioannidis JPA, Kiel DP, Rivadeneira F, Genome-wide meta-analysis identifies 56 bone mineral density loci and reveals 14 loci associated with risk of fracture, Nat Genet. 44 (2012) 491–501. 10.1038/ng.2249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [59].Maeda K, Kobayashi Y, Koide M, Uehara S, Okamoto M, Ishihara A, Kayama T, Saito M, Marumo K, The Regulation of Bone Metabolism and Disorders by Wnt Signaling, Int J Mol Sci. 20 (2019) 5525. 10.3390/ijms20225525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [60].Ikegame M, Tabuchi Y, Furusawa Y, Kawai M, Hattori A, Kondo T, Yamamoto T, Tensile stress stimulates the expression of osteogenic cytokines/growth factors and matricellular proteins in the mouse cranial suture at the site of osteoblast differentiation, Biomed Res. 37 (2016) 117–126. 10.2220/biomedres.37.117. [DOI] [PubMed] [Google Scholar]
- [61].Ikegame M, Ishibashi O, Yoshizawa T, Shimomura J, Komori T, Ozawa H, Kawashima H, Tensile stress induces bone morphogenetic protein 4 in preosteoblastic and fibroblastic cells, which later differentiate into osteoblasts leading to osteogenesis in the mouse calvariae in organ culture, J Bone Miner Res. 16 (2001) 24–32. 10.1359/jbmr.2001.16.1.24. [DOI] [PubMed] [Google Scholar]
- [62].Merrill AE, Bochukova EG, Brugger SM, Ishii M, Pilz DT, Wall SA, Lyons KM, Wilkie AOM, Maxson RE, Cell mixing at a neural crest-mesoderm boundary and deficient ephrin-Eph signaling in the pathogenesis of craniosynostosis, Hum Mol Genet. 15 (2006) 1319–1328. 10.1093/hmg/ddl052. [DOI] [PubMed] [Google Scholar]
- [63].Park S, Zhao H, Urata M, Chai Y, Sutures Possess Strong Regenerative Capacity for Calvarial Bone Injury, Stem Cells Dev. 25 (2016) 1801–1807. 10.1089/scd.2016.0211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [64].Yen EH, Pollit DJ, Whyte WA, Suga DM, Continuous stressing of mouse interparietal suture fibroblasts in vitro, J Dent Res. 69 (1990) 26–30. 10.1177/00220345900690010301. [DOI] [PubMed] [Google Scholar]
- [65].Ardalan M, Rafati A, Nejat F, Farazmand B, Majed M, El Khashab M, Risk factors associated with craniosynostosis: a case control study, Pediatr Neurosurg. 48 (2012) 152–156. 10.1159/000346261. [DOI] [PubMed] [Google Scholar]
- [66].Carmichael SL, Ma C, Rasmussen SA, Cunningham ML, Browne ML, Dosiou C, Lammer EJ, Shaw GM, Craniosynostosis and Risk Factors Related to Thyroid Dysfunction, Am J Med Genet A. 0 (2015) 701–707. 10.1002/ajmg.a.36953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [67].Chawla R, Alden TD, Bizhanova A, Kadakia R, Brickman W, Kopp PA, Squamosal Suture Craniosynostosis Due to Hyperthyroidism Caused by an Activating Thyrotropin Receptor Mutation (T632I), Thyroid. 25 (2015) 1167–1172. 10.1089/thy.2014.0503. [DOI] [PubMed] [Google Scholar]
- [68].Hashmi SS, Canfield MA, Marengo L, Moffitt KB, Belmont JW, Freedenberg D, Tanksley SM, Lupo PJ, The association between neonatal thyroxine and craniosynostosis, Texas, 2004–2007, Birth Defects Research Part A: Clinical and Molecular Teratology. 94 (2012) 1004–1009. 10.1002/bdra.23077. [DOI] [PubMed] [Google Scholar]
- [69].McNab T, Ginsberg J, Use of anti-thyroid drugs in euthyroid pregnant women with previous Graves’ disease, Clin Invest Med. 28 (2005) 127–131. [PubMed] [Google Scholar]
- [70].Radetti G, Zavallone A, Gentili L, Beck-Peccoz P, Bona G, Foetal and neonatal thyroid disorders, Minerva Pediatr. 54 (2002) 383–400. [PubMed] [Google Scholar]
- [71].Rasmussen SA, Yazdy MM, Carmichael SL, Jamieson DJ, Canfield MA, Honein MA, Maternal thyroid disease as a risk factor for craniosynostosis, Obstet Gynecol. 110 (2007) 369–377. 10.1097/01.AOG.0000270157.88896.76. [DOI] [PubMed] [Google Scholar]
- [72].O’Shea PJ, Bassett JHD, Sriskantharajah S, Ying H, Cheng S, Williams GR, Contrasting Skeletal Phenotypes in Mice with an Identical Mutation Targeted to Thyroid Hormone Receptor α1 or β, Molecular Endocrinology. 19 (2005) 3045–3059. 10.1210/me.2005-0224. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The datasets generated for this study can be found in the NCBI GEO under accession GSE66149.


