Abstract
Background
There are still uncertainties in our knowledge of the amount of SARS-CoV-2 virus present in the environment – where it can be found, and potential exposure determinants – limiting our ability to effectively model and compare interventions for risk management.
Aim
This study measured SARS-CoV-2 in three hospitals in Scotland on surfaces and in air, alongside ventilation and patient care activities.
Methods
Air sampling at 200 L/min for 20 min and surface sampling were performed in two wards designated to treat COVID-19-positive patients and two non-COVID-19 wards across three hospitals in November and December 2020.
Findings
Detectable samples of SARS-CoV-2 were found in COVID-19 treatment wards but not in non-COVID-19 wards. Most samples were below assay detection limits, but maximum concentrations reached 1.7×103 genomic copies/m3 in air and 1.9×104 copies per surface swab (3.2×102 copies/cm2 for surface loading). The estimated geometric mean air concentration (geometric standard deviation) across all hospitals was 0.41 (71) genomic copies/m3 and the corresponding values for surface contamination were 2.9 (29) copies/swab. SARS-CoV-2 RNA was found in non-patient areas (patient/visitor waiting rooms and personal protective equipment changing areas) associated with COVID-19 treatment wards.
Conclusion
Non-patient areas of the hospital may pose risks for infection transmission and further attention should be paid to these areas. Standardization of sampling methods will improve understanding of levels of environmental contamination. The pandemic has demonstrated a need to review and act upon the challenges of older hospital buildings meeting current ventilation guidance.
Keywords: SARS-CoV-2, COVID-19, Environmental sampling
Introduction
Workers in healthcare settings have been at high risk of contracting COVID-19, particularly in the first phase of the pandemic [[1], [2], [3]]. While risks to health professionals (including doctors and nurses) decreased, the risks to health associate professionals (including technicians, personal care support staff) remained high even at later stages of the pandemic. Clinical and non-clinical staff may be exposed to the SARS-CoV-2 virus in the workplace through contact with infected patients, visitors and staff via airborne virus, or possibly droplets and/or contact with contaminated surfaces. In a systematic review of sampling studies, primarily in healthcare settings, Cherrie et al. reported a median detection rate of SARS-CoV-2 RNA of 6% for surface samples (range zero to 74%) and 7% for air samples (zero to 100%) from studies carried out in 2020 and 2021 [4]. However, sampling methodologies and exposure metrics varied greatly in these studies and contextual information that may influence exposure was often not reported. Measurement methods generally lack the sensitivity to accurately detect the low concentrations of SARS-CoV-2 RNA typically present in work environments. Thus, there are still many uncertainties in quantifying surface contamination and/or airborne concentration of SARS-CoV-2.
There is a need to better understand the concentrations of SARS-CoV-2 virus (or other pathogens) in workplace settings, along with relevant exposure determinants, by using comparable methods to evaluate human occupational exposure and infection risk. This information would allow consideration of risk management measures on workplace and community contamination. As part of a study to explore worker exposure to SARS-CoV-2 in healthcare settings, we conducted a screening campaign in three hospitals in Scotland. The aims of this work were to evaluate the presence of SARS-CoV-2 in COVID-19-positive patient wards and wards where patients were awaiting results of COVID-19 tests; and to examine contextual factors in these wards such as ventilation and worker activities.
Methods
Study design
Three hospitals in Scotland were enrolled in our study. These were NHS district general hospitals, of varying ages, each with more than 500 beds. We refer to a COVID-19-positive (COVID-19 treatment) ward as one used to exclusively treat COVID-19 patients (as determined by PCR test), and non-COVID-19 wards as wards not specifically set aside for COVID-19 patients. At hospital A, the study was conducted in a respiratory ward which was repurposed to treat COVID-19-positive patients. At hospital B, we investigated one COVID-19 treatment ward (B1) and another where patients awaited results of a COVID-19 test (B2) before being moved elsewhere. At hospital C, the study was conducted in a ward similar to B2. The patient rooms were either general mixed-gender wards with multiple beds or single rooms (Table A1), with ensuite bathrooms (one per room). Nurse stations were generally in the main thoroughfare of the ward and the waiting rooms did not include toilets. Site A was visited twice 12 days apart, while the other sites were visited once. More information on the areas visited is available in Table A1. Hospitals were visited between November 12th, 2020 and December 16th, 2020, when the Alpha SARS-CoV-2 strain was dominant.
Site assessments
Measurements of room volume and ventilation, temperature and humidity, number of patients, and number of staff present during sampling were recorded. Airflow measurements were taken at air-inlet diffuser grilles within each ward/utility room using a TSI/Airflow PH731 electronic balometer (TSI, Shoreview, MN, USA). Pressure differentials between rooms were measured using a micromanometer (DPM RS323 Micromanometer, UK) with the inlet tube placed under the door between two rooms. The air change rate was then estimated based on the measured air flow volume (L/s) divided by the room volume (m3) multiplied by a conversion unit to obtain the air changes per hour.
Observational study
Staff were observed to determine how much time they spent with patients and the surfaces they touched during care activities. One member of staff was observed at a time, during specific patient care activities. The field worker would allow for 5–10 min of first standing in the same area as if they were observing the worker before recording information to allow the worker to get used to the field worker's presence. Surface touches were recorded for: bed, door, bed handrail, healthcare worker's notes, patient, sink, window. Observations were based on staff contact per patient. Ethical approval was obtained from ACCORD and the Research and Development Offices of each NHS Board we worked with, under Rec No. 20/NRS/0020.
Sample collection and analysis
Surface swab samples were taken using sterile nylon flocked swabs which were placed in 2 mL liquid Amies medium (Sterilin Ltd, Newport, UK) after collection. Surface area swabbed varied according to the object and ranged from 10 to 225 cm2. Locations swabbed included light switches to the toilet, sink taps, door handles, cot sides, patient bed tables, personal protective equipment (PPE) donning and doffing areas (biowaste disposal container and sink taps), waiting rooms (table and cupboard tops), and nurse workstations (keyboards, worktops). Sample blanks were taken into the field for each hospital surveyed. The surface sampling protocol was based on a study carried out in hospitals in England by Moore et al. [5].
Air samples were collected using the Coriolis micro sampler (Bertin, Montigny-le-Bretonneux, France) with an extended sampling attachment allowing fluid in the collection vessel to be replenished during sampling. Most samples were taken at 200 L/min for 20 min into 15 mL of RNAse-free phosphate-buffered saline. A pair of samples per visit was placed side-by-side to test whether sampling time (20 vs 60 min) might affect capture and recovery of virus RNA. Air samples were taken in the same rooms and bed areas as swab samples. For each hospital sample blanks using the same sample cones and media were taken into the field but no air was drawn through the media.
Samples were sent from Edinburgh to the UK Health Security Agency laboratory at Porton Down for analysis according to methods similar to those of Moore et al. [5]. Aliquots (140 μL) of each Coriolis air sample media and each surface swab sample were extracted using the QIAamp viral RNA Mini Kit (Qiagen Ltd, Manchester, UK) according to manufacturer's instruction. The remaining air sample volume was concentrated using a Vivaspin™ 20 centrifugal concentrator to ≤1 mL and 140 μL aliquots extracted using the same method. RNA extracts were eluted in 60 μL AVE buffer and stored at –80 °C until RT–PCR analysis. Each extraction event included two negative extractions containing 140 μL absolute ethanol which was analysed in the same RT–PCR run.
Environmental RNA extracts were tested for the presence of SARS-CoV-2 using the CE Viasure SARS-CoV-2 Real Time PCR Detection Kit (CerTest Biotec, Zaragoza) on an Applied Biosystems™ QuantStudio™ 5 Real-Time PCR System. The Viasure RT–PCR assay targets the Nucleoplasmid (N) and the ORF1ab genes, RNA extracts were tested in duplicate and recorded as ‘positive’ if at least one target amplified in both replicates. Samples were recorded as ‘weakly positive’ if one replicate amplified (in one or both targets). All weakly positive samples were reanalysed with the potential to be reclassified as ‘positive’ or remain ‘weakly positive’. This was done to ensure that the results were not due to experimental error, given that the level of RNA present was low. Quantification was carried out using the N gene. Each RT–PCR experiment contained an N gene standard curve (50,000 to 5 copies in 10-fold dilutions of in-vitro transcribed RNA) and two negative (no-template) and two positive control wells. The limit of detection of the Viasure assay reported by the manufacturer is 10 genomic copies per reaction, with the highest cycle threshold value as ≥39. Cycle threshold refers to the number of amplification cycles needed to detect genetic material. The theoretical detection limit, therefore, can be calculated by multiplying the assay LoD first by 12 (60/5) as extractions are eluted in 60 μL and 5 μL of sample was added to the reaction. This gives copies/extraction, which is then multiplied by the sample volume/140 (μL) as 140 μL of the sample is extracted. This results in a theoretical limit of detection of 3200 copies/m3 for air and 1700 copies/swab for surfaces. It should be noted, however, that the laboratory was able to reliably detect below the reported detection limit (to 5 genomic copies per reaction), resulting in a limit of detection of 1600 copies/m3 and 860 copies per swab for air and surfaces, respectively.
Estimated distribution of concentrations and surface loadings
The R software package EnvStats was used to carry out the statistical analysis of the virus RNA air concentrations and surface loadings. Due to the large number of non-detected samples, the elnorm function was used to extrapolate the geometric mean and standard deviation for a lognormal function. For air samples, the lowest reported concentration in copies/m3 was used as the limit of detection, while the lowest reported copies/swab was used as the limit of detection for surface samples.
The sample size was not extensive enough for a detailed statistical analysis, due to limited availability of staff and access to hospital wards during the pandemic, and we qualitatively explored the relationship between ventilation rates, occupancy, and concentration of viral RNA contamination.
Results
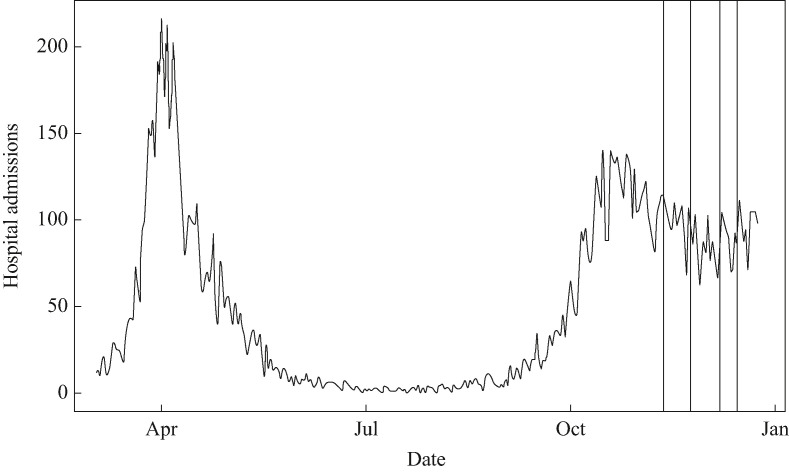
This study occurred at a time when it was estimated that around 1% of the population in Scotland would have tested positive for COVID-19 [6]. It was during the second peak of COVID-19 in the UK, just after the B1.1.7 strain had been first detected in the UK (see Figure 1 ).
Figure 1.
COVID-19 hospital admissions in Scotland in 2020. Vertical lines indicate sampling dates.
In total, 186 samples (127 surface swabs and 59 Coriolis air samples), including blank and control samples, were sent for analysis. The number of samples and their locations are shown in Table A1. In hospital A (COVID-19 treatment ward), 59 surface swabs and 25 air samples were taken; in hospital B1 (hospital B, COVID-19 treatment ward), 23 surface swabs and nine air samples; in hospital B2 (hospital B, non-COVID ward), 13 surface swabs and eight air samples; and in hospital C, 27 surface swabs and 11 air samples. Hospital A was visited on two occasions and more bedrooms were sampled compared with the other hospitals' wards. Additionally, hospital A's ward had more beds per room than the other two, and only rooms with one bed were sampled in hospital C (non-COVID-19) (Table A1).
Table A.2 shows the C T-values and air concentrations or surface loadings for positive and weakly positive samples, all of which were found in the COVID-19 treatment wards, but not in any of the assessment wards. In hospital A, 19% of air samples and 23% of surface samples had detectable levels of SARS-CoV-2 RNA. Of the positive samples, C T-values ranged from 32 to 39, with the lowest measurable air sample concentration at 30 copies/m3 and the highest 4.2×102 copies/m3. The surface loadings ranged from 3 to 3.8×102 copies/cm2. Both of the visits to one of the ward rooms in which continuous positive airway pressure (CPAP) was being delivered showed the highest proportion of positive surface samples although there were no detectable positive (or weakly positive) air samples. Hospital ward B1 had 44% detectable positive air samples (or weakly positive) and 17% detectable positive surface (or weakly positive) samples. C T-values had the same range as ward A, and measurable air concentrations ranged from 35 to 1.7×103 copies/m3 while surface loadings ranged from 3 to 3.2×102 copies/cm2. Table I shows the estimated geometric mean and standard deviation for each COVID-19 hospital ward, assuming a lognormal distribution of the measured concentrations. We assume that the amount of virus material present is a continuum and have extrapolated values below the estimated detection limits based on methods used for developing distributions of environmental contaminants.
Table I.
Geometric mean, standard deviation, and confidence intervals of the mean estimate, assuming that the distributions are lognormal
| Parameter | No. of samples | Geometric mean | Geometric SD | 95% CI |
|---|---|---|---|---|
| Hospital A | ||||
| Air (copies/m3) | 25 | 0.89 | 51 | 0.10–69 |
| Surface (copies/swab) | 59 | 10 | 24 | 1.9–34 |
| Surface (copies/cm2)a | 0.12 | 0.28 | 0.02–0.41 | |
| Hospital B1 | ||||
| Air (copies/m3) | 9 | 16 | 24 | 1.0–560 |
| Surface (copies/swab) | 23 | 12 | 9.7 | 1.0–160 |
| Surface (copies/cm2)a | 0.15 | 0.12 | 0.01–1.9 | |
SD, standard deviation; CI, confidence interval.
Below-detection-limit values have been probabilistically estimated based on the assumption of lognormality.
Surface loading distributions were estimated by dividing the copies/swab parameters by the mean of surface areas swabbed in the location reported.
Two of the co-located air samples, testing the effect of sample time (and hence volume of air sampled) in hospital A, were not detectable for the shorter samples (20 min) but weakly positive for the longer samples (60 min).
Ward volumes ranged from 113 to 161 m3 for multiple bed rooms and from 43 to 48 m3 for single or treatment rooms (Table A3). Most of the bedrooms did not have mechanical supply and extract ventilation, save one treatment room for a single patient in hospital A. The ensuite toilets and shower rooms (WC) were mechanically ventilated and the extract airflows were measured. In hospital A, additional extract fans beside the bedroom windows were included, which were estimated to add an additional 68 L/s airflow. This would result in approximately two additional air changes per hour above the ventilation rate without the fans. Extract air changes per hour (ach) for toilets ranged from 3 to 10 ach. Both supply and extract could only be measured in one room in hospital A (a treatment room) and bedroom 05 in hospital B2, as those were the only ones with mechanical supply and extract (i.e. provided through an air handling unit as part of the original design).
In the COVID-19-positive wards, 20 staff were observed and in the COVID-19-assessment wards, 13 staff were observed. These included 25 nurses, three doctors, and two domestic or portering staff. Staff contact with the patients during care was very varied and ranged from under 1 min to 45 min of contact, with longer time spent with patients who were elderly or who had comorbidities needing assistance (tasks observed are detailed in the Appendix). The most frequently touched surfaces are shown in Table II . These generally also correspond with the surface touch results finding that the beds were the most likely to have detectable levels of SARS-CoV-2, as well as the sink taps. The distribution of the touches to surfaces was positively skewed. We estimated that total contact with surfaces within 1 m of a patient per hour was: mode 1.38 (range: 0–240); and contact >1 m from the patient was: mode 23.1 (range: 0–600).
Table II.
Surfaces recorded for observation study, based on 33 observed workers
| Surface touched | Frequency |
|---|---|
| Patient | 57 |
| Notes | 36 |
| Bed | 20 |
| Door | 20 |
| Sink | 18 |
| Window | 4 |
| Handrail | 3 |
Discussion
SARS-CoV-2 RNA was measured in air and on surfaces in patient rooms and nurse work areas in wards with and without COVID-19 patients across three Scottish hospitals during a period of high levels of community prevalence in 2020. Whereas SARS-CoV-2 RNA was not detected in the non-COVID-19 wards, viral RNA was detected in both COVID-19 treatment wards. This suggests that the hospitals were successful in identifying and isolating COVID-19 patients. SARS-CoV-2 detection rates varied from room to room but were about 20%, which is higher than the 6% median detection rate found in an earlier review for healthcare settings [4]. The same review estimated the geometric mean for air concentrations of SARS-CoV-2 as 0.014 (95% confidence interval (CI): 0.003–0.047) RNA copies/m3, as imputed from the pooled study data. For comparison, in our study, hospital A's geometric mean was 0.89 (95% CI: 0.10–69) copies/m3, and for hospital B1 the geometric mean was 16 (95% CI: 1.0–560). These are higher than the estimated geometric means from Cherrie et al., and despite the very wide CIs in this estimate, the lower range of the interval implies a statistical difference between the review and our study [4]. One should interpret these results with caution, however, as we are estimating distributions with a large number of non-detects (>50%) and the sample values span a wide range. Sampling methods across studies also varied. Our study sampled larger volumes of air than, e.g. two other studies using a similar sampler in the UK [5,7]. Additionally, we cannot draw any conclusions about the representativeness of our data for the hospitals over time, particularly as the patient population and predominant SARS-CoV-2 strains change.
Our results also show similarities with other studies (Table III ) that focused on characterizing SARS-CoV-2 in hospital environments in the UK with one involving eight hospitals, including 12 non-intensive care unit (ICU) cohort bays with COVID-19 patients, and the other focused on seven clinical areas and one public area in a hospital in London [5,7]. Although these other studies were done during the first peak of COVID-19 in the UK in contrast to ours, our study found similar air and surface concentrations in non-ICU wards. It appears that ICUs do not necessarily have higher levels of SARS-CoV-2, despite being areas with aerosol-generating procedures. This may be because, in critical care, patients are generally not in the acute phase of their infection where they are much more likely to be shedding virus, and even if the patients were contagious, patients were usually ventilated with an enclosed system preventing environmental shedding. Our findings also suggest that increased amounts of virus may settle on to surfaces around patients who undergo CPAP, which occurred in general COVID-19 wards. This is also reflected in Zhou et al.’s study, which found that a temporary CPAP ward also had statistically significantly higher surface concentrations than the ICU, although no CPAP occurred during sampling. CPAP is not a closed system, so virus-loaded aerosol could leak from masks which generally do not form a close seal with the patient's face.
Table III.
Comparison of studies on environmental SARS-CoV-2 done in UK hospitals
| Study | Location | Air concentration (copies/m3) | Surface levels |
|---|---|---|---|
| This study | Three hospitals in Scotland | <30 to 1717 copies/m3 15% positive |
<160 to 19,168 copies/swab <1.49 to 319 copies/cm2 16% positive |
| Moore et al. [5] | Eight hospitals in England | <10 to 460 copies/m3 7% positive |
59 to 2.2 × 105 copies/swab 9% positive |
| Zhou et al. [7] | One hospital in London, UK | 31 to 7048 copies/m3 11% positive |
∼101 to 104 copies/swab ∼0.4 to 400a copies/cm2 52% positive |
Surface loading estimated by dividing copies/swab by 25 cm2, which was the approximate area of surfaces swabbed reported by authors.
At the time of the study, the relevant guidance for recommended air exchange rates and pressure differentials for various types of hospital areas receiving mechanical ventilation was Scottish Health Technical Memorandum (SHTM) 03-01 Part A (the SHTM was updated in 2022) [8]. Due to the age of the buildings studied (all built prior to 2014), ventilation guidance at time of build would have been Health Technical Memorandum 2025 (HTM 2025, 1994), which did not advise that ward bedrooms should have mechanical ventilation and therefore an exemption would have been agreed upon at the time of inspection. We observed some interventions in place to improve the ventilation, such as window extract fans, which we estimated to provide up to 2 ach when in use in these otherwise naturally ventilated rooms. However, due to the noise, these fans were not always in use. During the study period, we were told that the fans were generally used when CPAP was in process. Other interventions, such as air cleaning units or opening of windows, were also used in hospitals B and C if COVID-19 outbreaks were found to occur [9]. Regular opening of windows was instituted as an intervention in January 2021, given that most of the ward bedrooms relied on natural ventilation, but compliance was not formally audited [10]. Air extract for most rooms occurred through the ensuite toilet rather than within the room itself, which was often the only area where there was mechanical ventilation. The corridors have supplied filtered air resulting in positive pressure with respect to the wards. Pressures across the rooms were slightly negative, although opening the windows tends to cause pressure cascades to become variable and occasionally positive with respect to the corridor. Although the SHTM does not prescribe pressure requirements for general wards it does for single rooms, they should not be positive with respect to the corridor.
There was a high number of samples where the measured concentration was below the detection limit of the Viasure assay, even in COVID-19-positive wards. Whereas Cherrie et al. found a positive correlation between number of positive surface samples and air samples, this was not clearly seen in our study [4]. Measurements showed that both air and surface concentrations were low, and levels in other studies of SARS-CoV-2 in healthcare environments also found similar levels of viral material. It is possible that existing sampling methods are not able to successfully capture viable virus, or adequate amounts of virus material – two samples in one of the COVID-19 treatment wards of our study found that a longer sample time and larger sample volume could be more effective at capturing virus material, but this finding is not enough to draw any conclusions. Swabbed surface areas are also relatively small. It should also be noted that we were not able to account for any losses which may occur in the sampling or transport process.
Our results show that SARS-CoV-2 RNA can be present on surfaces and in air around infected patients, and that touching the patient and nearby surfaces could potentially transmit virus elsewhere. Healthcare worker's notes and the sink are frequently touched during patient care, and could lead to transmission of the virus to these areas. We did not take samples of the worker's notes, but the sink taps in the rooms with COVID-19 patients had detectable levels of virus, although we cannot say whether this is due to touches by the HCWs or the patients. The levels found in this and other studies of environmental SARS-CoV-2 have frequently not been culturable, therefore it is not possible to say anything about the infectiousness of the RNA detected. Results here may not be reflective of later, potentially more transmissible variants of SARS-CoV-2.
SARS-CoV-2 is chiefly transmitted by airborne exposure although viral shedding varies hugely between individuals. It is difficult to rely on the randomized clinical trial to evaluate the real-world effectiveness of filtering facepiece (FFP) vs surgical masks for protecting HCWs from infection, as it is impossible to control all confounding factors. Some evidence indicates potential benefits for HCWs and patients with use of FFPs, but successful infection control will include a bundle of measures, not just respiratory protective equipment [11,12]. Current guidance on use of respiratory protection for HCWs suggests use of FFP3 respirators for working with patients with known or suspected infectious diseases transmitted ‘wholly by the airborne route’ or when carrying out aerosol-generating procedures on patients with an infectious disease spread at least partially by droplet or airborne routes, or as a precautionary measure if the hazard is unknown [13].
Much literature on COVID-19 and the SARS-CoV-2 virus in healthcare settings focuses on patient wards or ICUs. Other areas, such as staff break rooms, changing rooms, or public areas have not been well studied, but may be areas for transmission, especially as respiratory protective equipment use may be relaxed in these settings. We also found positive air and surface samples in the PPE changing area of one of the hospitals, which was a repurposed bathroom area. Changing areas may therefore present a high exposure risk for staff, given that staff may be removing their PPE and are therefore vulnerable to virus exposure [14]. The repurposed toilet was supposedly not used as a toilet, but we do not know if it might have been during sampling days. Additionally, we detected SARS-CoV-2 RNA in air and surface samples taken from waiting and meeting rooms in the wards. This is an indication that ventilation in those rooms may be insufficient to remove airborne virus particles, which may have been generated by an earlier occupant and spread throughout the room. Furthermore, cleaning in these areas is unlikely to be as rigorous or as frequent as in patient areas, so any surface contamination may not be as well controlled. There is a need to further evaluate controls for these non-clinical areas, given that people may feel more relaxed and be less likely to take precautions in these places. Studies on behaviours in various parts of hospitals, particularly non-patient areas, related to PPE use and surface touches compared to patient areas, could help further define predictors of risk in different areas of the hospitals. These, along with environmental surface contamination levels from sampling studies such as ours, can also be used to inform quantitative microbial risk assessment models. These models could help decision-makers in these hospitals evaluate risks and compare the efficacy of different control measures.
Since this study, population levels of infection have been low, and, although infectivity and health impacts have been reduced by immunization, further more-infective variants are possible. Lessons from the COVID-19 pandemic need to be translated to improve resilience of health and social care for the future.
The implications of our study for policymakers are as follows.
-
–
Regular environmental sampling for SARS-CoV-2 RNA should be used to inform the effectiveness of control measures where there is evidence of occupationally acquired transmission in healthcare settings and elsewhere.
-
–
The protocol for such surveys should be standardized to facilitate comparability and inform improvement.
-
–
The widespread non-compliance with healthcare ventilation standards in older buildings justifies a national review and action plan.
Acknowledgements
We would like to thank all the hospital staff who have helped facilitate or who participated in our study.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jhin.2022.11.019.
Conflict of interest statement
None declared.
Funding sources
This work was supported by the Chief Scientist's Office of Scotland, reference number COV/IOM/Portfolio.
Appendix A. Supplementary data
The following is the Supplementary data to this article.
References
- 1.Mutambudzi M., Niedwiedz C., Macdonald E.B., Leyland A., Mair F., Anderson J., et al. Occupation and risk of severe COVID-19: prospective cohort study of 120 075 UK Biobank participants. Occup Environ Med. 2020;78:307–314. doi: 10.1136/oemed-2020-106731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Nafilyan V., Pawelek P., Ayoubkhani D., Rhodes S., Pembrey L., Matz M., et al. Occupation and COVID-19 mortality in England: a national linked data study of 14.3 million adults. Occup Environ Med. 2022;79:433–441. doi: 10.1136/oemed-2021-107818. [DOI] [PubMed] [Google Scholar]
- 3.Shah A.S.V., Wood R., Gribben C., Caldwell D., Bishop J., Weir A., et al. Risk of hospital admission with coronavirus disease 2019 in healthcare workers and their households: nationwide linkage cohort study. BMJ. 2020;371 doi: 10.1136/bmj.m3582. m3582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Cherrie J.W., Cherrie M.P.C., Smith A., Holmes D., Semple S., Steinle S., et al. Contamination of air and surfaces in workplaces with SARS-CoV-2 virus: a systematic review. Ann Work Expo Health. 2021;65:879–892. doi: 10.1093/annweh/wxab026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Moore G., Rickard H., Stevenson D., Aranega-Bou P., Pitman J., Crook A., et al. Detection of SARS-CoV-2 within the healthcare environment: a multi-centre study conducted during the first wave of the COVID-19 outbreak in England. J Hosp Infect. 2021;108:189–196. doi: 10.1016/j.jhin.2020.11.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Scottish Government. Coronavirus (COVID-19): ONS Infection Survey – headline results – 4 December 2020. Available at: https://www.gov.scot/publications/coronavirus-covid-19-ons-infection-survey-results--4-december-2020 [last accessed January 2022].
- 7.Zhou J., Otter J.A., Price J.R., Cimpeanu C., Meno Garcia D., Kinross J., et al. Investigating SARS-CoV-2 surface and air contamination in an acute healthcare setting during the peak of the COVID-19 pandemic in London. Clin Infect Dis. 2021;73:e1870–e1877. doi: 10.1093/cid/ciaa905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Scottish Health Technical Memorandum 03-01 Ventilation for healthcare premises Part A – Design and validation. https://www.nss.nhs.scot/media/2373/shtm-03-01-part-a-v20-feb-2014-archived.pdf.
- 9.Conway Morris A., Sharrocks K., Bousfield R., Kermack L., Maes M., Higginson E., et al. The removal of airborne severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and other microbial bioaerosols by air filtration on coronavirus disease 2019 (COVID-19) surge units. Clin Infect Dis. 2022;75:e97–e101. doi: 10.1093/cid/ciab933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Dancer S.J., Cormack K., Loh M., Coulombe C., Thomas L., Pravinkumar S.J., et al. Healthcare-acquired clusters of COVID-19 across multiple wards in a Scottish health board. J Hosp Infect. 2022;120:23–30. doi: 10.1016/j.jhin.2021.11.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Haller S., Güsewell S., Egger T., Scanferla G., Thoma R., Leal-Neto O.B., et al. Impact of respirator versus surgical masks on SARS-CoV-2 acquisition in healthcare workers: a prospective multicentre cohort. Antimicrob Resist Infect Control. 2022;11:27. doi: 10.1186/s13756-022-01070-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lawton T., Butler M., Peters C. Airborne protection for staff is associated with reduced hospital-acquired COVID-19 in English NHS trusts. J Hosp Infect. 2022;120:81–84. doi: 10.1016/j.jhin.2021.11.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.NHS Scotland. National Infection Prevention and Control Manual: Chapter 2 – Transmission Based Precautions (TBPs). Available at: https://www.nipcm.hps.scot.nhs.uk/chapter-2-transmission-based-precautions-tbps/#a1091 [last accessed June 2022].
- 14.Birgand G., Peiffer-Smadja N., Fournier S., Kerneis S., Lescure F.X., Lucet J.C. Assessment of air contamination by SARS-CoV-2 in hospital settings. JAMA Netw Open. 2020;3 doi: 10.1001/jamanetworkopen.2020.33232. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.