Abstract
Bacterial pathogens employ a plethora of virulence factors for host invasion, and their use is tightly regulated to maximize infection efficiency and manage resources in a nutrient-limited environment. Here we show that during Escherichia coli stationary phase the 3′ UTR-derived small non-coding RNA FimR2 regulates fimbrial and flagellar biosynthesis at the post-transcriptional level, leading to biofilm formation as the dominant mode of survival under conditions of nutrient depletion. FimR2 interacts with the translational regulator CsrA, antagonizing its functions and firmly tightening control over motility and biofilm formation. Generated through RNase E cleavage, FimR2 regulates stationary phase biology by fine-tuning target mRNA levels independently of the chaperones Hfq and ProQ. The Salmonella enterica orthologue of FimR2 induces effector protein secretion by the type III secretion system and stimulates infection, thus linking the sRNA to virulence. This work reveals the importance of bacterial sRNAs in modulating various aspects of bacterial physiology including stationary phase and virulence.
INTRODUCTION
The ability of bacterial pathogens to invade a host critically relies on the presence of adequate conditions promoting the survival of the pathogen and its arsenal of virulence factors. Several Gram-negative pathogens employ type III secretion systems (T3SS) to deliver a wide range of effector proteins directly into the cytoplasm of their target cells (1,2) and type 1 pili (T1P) to adhere to and invade host cells (1,2). Bacterial virulence factors are under tight regulation via multiple mechanisms to minimize the use of precious resources in an unpredictable environment (3). CsrA is a translational regulator that governs the expression of T3SS effectors in bacterial pathogens. For instance, in Salmonella enterica, CsrA downregulates hilD, the SPI-1 (Salmonella pathogenicity island 1) encoded regulator, and by doing so, inhibits the expression of multiple HilD-dependent T3SS effectors (4). In enteropathogenic Escherichia coli (EPEC), CsrA induces flagellar motility and the expression of some T3SS effectors involved in Attaching/Effacing (A/E lesions) (5). Both Salmonella and E. coli regulate the expression of T1P through phase variation (6), allowing some members of isogenic populations to display the pilus and others to retract it. In E. coli, the promoter of the fimAICDFGH operon that encodes T1P lies within an invertible DNA region (7) called fimS, and the recombinases FimB and FimE govern its orientation: FimB mediates the inversion in both ON (expression) and OFF (repression) orientations while FimE is biased to the OFF inversion (8). This complex mode of regulation in both pathogens is further controlled by additional transcriptional regulators (9,10).
Biofilm formation is another mode employed by bacterial pathogens to assist in host infection. This multicellular-like lifestyle allows resistance to environmental and cell-intrinsic stresses. Indeed, biofilms have been linked to the establishment of recurrent infections and bacterial persistence (11). Uropathogenic E. coli (UPEC) can form resilient biofilms through the coordinated expression of several outer membrane appendages, including pili, fimbriae, flagella, and curli (1,11). The T1P plays important roles in this context, allowing UPEC to form biofilms on both abiotic surfaces such as medical supplies and urinary catheters (12), and in host tissues, such as the bladder epithelium (11).
Recently, trans-encoded small non-coding RNAs (sRNAs) have emerged as potent regulators of gene expression in various bacterial species (13). By partially base pairing with various target mRNAs and regulating their expression at the post-transcriptional level, these non-coding RNAs (ncRNAs) govern various aspects of bacterial physiology (14), with various roles described in bacterial virulence and biofilm formation (3,15,16). For instance, EHEC GlmY and GlmZ sRNAs promote the formation of A/E lesions through induction of the T3SS effector EspFu, and diminish the expression of other T3SS effectors (17). In Salmonella, the sRNA PinT coordinates the expression of SPI-1 and SPI-2 (Salmonella pathogenicity islands 1 and 2) (18) allowing the pathogen to switch from its invasive to its persistent infective mode. Salmonella CpxQ, a 3′ UTR-derived sRNA, inhibits fimA expression through direct base-pairing with the fimA mRNA, controlling T1P expression under conditions of membrane stress (19). These sRNAs typically require the RNA chaperones Hfq or ProQ for generation, stability, and function (20,21). However, these chaperones may be dispensable for some regulatory circuits with the existence of chaperone-independent sRNAs in Gram-positive bacteria (22).
Previously, our deep sequencing analysis of growth phase-dependent sRNAs in E. coli revealed the abundant expression of sRNA_35 in the stationary phase (23), a ncRNA which was also recognized in previous studies (24,25). Here, we report the first functional characterization of this 35-nucleotide long sRNA that we rename to FimR2 for fim operon-derived sRNA as it originates from the 3′ UTR of fimA in the intergenic region between fimA-fimI in E. coli (Supplementary Figure S1) and S. enterica. This operon has been shown by others to encode another sRNA, dubbed fimR, which however derives from a different location within the fim operon and is thus completely independent of FimR2 (26). FimR2 overexpression in exponential phase, a growth phase during which this sRNA is normally undetectable, significantly causes biofilm formation, alters the outer membrane architecture, and switches the bacterial population to stationary phase. Through various genetic and biochemical approaches, we show that these phenotypes are due to the FimR2-mediated regulation of various targets involved in stationary phase biology and motility, directly through base-pairing with target mRNAs, and indirectly, through the sequestration of the translational regulator CsrA. Interestingly, overexpression of the Salmonella variant, FimR2S, in E. coli causes biofilm formation, whereas in S. enterica it potentiates invasiveness and promotes the expression of a T3SS-chaperone. Unlike most of the so far characterized bacterial trans-acting sRNAs, we show that the multifunctional regulator FimR2 acts independently of Hfq and ProQ and accumulates through the RNase E-dependent cleavage of fimAICDFGH. This work positions FimR2 as a master regulator of gene expression in stationary phase, employing multitasking to coordinate biofilm formation and virulence.
MATERIALS AND METHODS
Bacterial strains and growth conditions
E. coli K-12 strain and its derivatives (Supplementary Table S1) were grown at 37°C with shaking at 220 rpm in LB (10 g tryptone, 10 g NaCl, and 5 g yeast extract/l). Exponential phase samples were taken at OD600 = 0.4 or at indicated time points after reaching this OD600. Stationary phase samples were taken 24, 48 or 72 h after initial inoculation. Induction with IPTG (isopropyl β-d-1-thiogalactopyranoside) or tetracycline was done at OD600 = 0.4. All media were supplemented with kanamycin (25 μg/ml), ampicillin (100 μg/ml), streptomycin (100 μg/ml), IPTG (1mM), or tetracycline (25 nM) where indicated. For the RNase E inactivation experiment, K-12 and rne3071−ts (27) strains were grown at 30°C to stationary phase and then the cultures were transferred to 43°C. Salmonella serovar Typhimurium strain SL1344 sub-strain SB300 (28), M1525 (29), ATCC14028s (30), UK-1 (31) and SB245 ( sipABCDsptP::aphT) (Kaniga and Galan, unpublished) were grown at 37°C with shaking at 150 rpm in LB supplemented with 0.3 M NaCl. For northern blot analysis, Salmonella stationary phase samples were taken 48 and 72 h after initial inoculation.
sipABCDsptP::aphT) (Kaniga and Galan, unpublished) were grown at 37°C with shaking at 150 rpm in LB supplemented with 0.3 M NaCl. For northern blot analysis, Salmonella stationary phase samples were taken 48 and 72 h after initial inoculation.
Construction of strains and plasmids
To overexpress FimR2S-L and FimR2S-S from the expression plasmid, we included a portion of the fimA-fimI intergenic space downstream of the two variants, prompting RNAse E to generate the canonical 3′-end of both variants (32). Alternatively, the putative secondary structure of both variants would terminate expression. FimR2S-L and FimR2S-S were inserted, along with the aforementioned downstream intergenic space, at the transcriptional + 1 site under PlacO control in pBbE6k-RFP (33) by MEGAWHOP cloning (34). For this, a pair of primers (see Supplemental Table S2) were used to create a mega primer spanning the genomic FimR2S sequence by PCR using Phusion DNA Polymerase (NEB). The resulting PCR product was purified with Wizard® SV Gel and PCR Clean-Up (Promega) and used in excess to insert the sRNAs into the pBbE6k-RFP plasmid and replace the RFP sequence with PCR. The PCR reaction was digested with DpnI (New England Biolabs) for 2 h at 37°C and transformed into K-12 E. coli competent cells. Plasmids were extracted from positive clones by Wizard® Plus SV Minipreps DNA Purification System (Promega) and sequenced with HL0199 primer (Microsynth). FimR2 mutants (CU, AA, G24U, G23A-G24U, stem, loop and A-U stem) were generated by one-step cloning PCR (35) using pBbE6k-FimR2 plasmid as a template (23). PCR reactions were DpnI treated as previously mentioned, and subsequent transformation, plasmid extraction, and sequencing were carried out as described for pBbE6k-fimRS-L and pBbE6k-fimRS-S. Using one-step cloning PCR, 18 base-pairs of fliJ sequence corresponding to codons 6–12 were inserted between the first and second codon of the RFP sequence in pBbA2A-RFP. In a subsequent step, 12 additional base-pairs of the fliJ sequence corresponding to fliJ codons 2–5 were inserted upstream of the previous fliJ insert. To overexpress FimR2, FimR2-CU, FimR2-AA, and the control plasmid in the rpoS deletion strain, the kanamycin resistance cassette in the pBbE6k plasmid (backbone of each construct), was switched to an ampicillin resistance cassette using PIPE cloning (36). For this purpose, the ampicillin cassette was amplified from the pBbA2a plasmid and the backbones of pBbE6k-FimR2, pBbE6k-FimR2-CU, pBbE6k-FimR2-AA and pBbE6k-control were individually amplified by PCR. PCR reactions were DpnI-treated as standard and then co-transformed into bacterial strains. Subcloning thereafter was done as described.
CsrA was amplified from K-12 genomic DNA and SL1344/SB300 DNA by PCR with primer pairs containing BamH1 and NdeI restriction sites and the histidine tag (The same pair of primers was used for amplification of CsrA from the genomic DNA of both species as the annealing sequence is identical at the 5′- and 3′-end). The PCR product was then digested with both restriction enzymes as described and purified by size-exclusion electrophoresis. Ligation into BamH1/NdeI-cut backbone (pBbE6k-RFP with RFP excised) was done with T4 DNA Ligase (Promega). Subsequent subcloning was done as standard. Clones were confirmed by sequencing using HL0030 primer at Microsynth.
Genomic deletion strains were created as described (37). sRNA locus (FimR2, CsrB and CsrC) and gene (CsrD)- specific primers (Supplemental Table S2) were used to amplify a kanamycin cassette from pKD13 plasmid by PCR. The purified PCR product was electroporated into K-12 E. coli strain containing the pKD46 plasmid and expressing lambda red recombinase through induction with 1 mM Arabinose. Positive clones were recovered on LB agar plates supplemented with Kanamycin. Homologous recombination was confirmed by PCR as described in the PCR section. For the sRNA-deletion strains, the chromosomally-inserted Kanamycin cassette was removed by FLP recombination using pcp20 plasmid as previously described (38). Removal of the cassette was also confirmed by replica plating on LB agar plates with and without antibiotics, and by PCR.
RNA extraction and northern blotting
RNA extraction was performed with hot acidic phenol as described (39). For northern blot analyses, 5–10 μg of total RNA were separated on denaturing polyacrylamide gels (8% acrylamide M-Bis, 7 M Urea, 1 × TBE), and gels were run for 2 h at 250 V. RNA was transferred to a nylon membrane (Amersham Hybond-N+, GE Healthcare) using a semi-dry blotter (V20-SDB, Scie-Plas) and crosslinked to membranes using a microprocessor-controlled UV irradiation system (BLX-254, Vilber Lourmat). DNA oligonucleotides were end-labeled with [γ-32P]-ATP (Hartmann Analytic) using PNK (Thermo Fisher Scientific) and used for hybridization as described (40).
For agarose gel northern blot (Figure 1A, fimA), 20 μg total RNA were resolved on a denaturing agarose gel (1.2% agarose, 0.5% formaldehyde, 1× MOPS). RNA samples were transferred to a nylon membrane by capillary action in 20× SSC overnight at room temperature. After crosslinking RNA to membrane as mentioned, the membrane was incubated in 2× SSC buffer then in hybridization buffer at 65°C for 1 h. The template for fimA probe was amplified by PCR from K-12 genomic DNA then labelled with [α-32P]-dCTP (Hartmann Analytic) with the Klenow fragment (Thermo Fisher Scientific) for 1 h at 37°C. Hybridization was done overnight at 65°C. Two washes were carried out on the following day for 30 min each in wash buffer I (2× SSC, 0.1% SDS) and II (0.2× SSC, 0.1% SDS).
Figure 1.
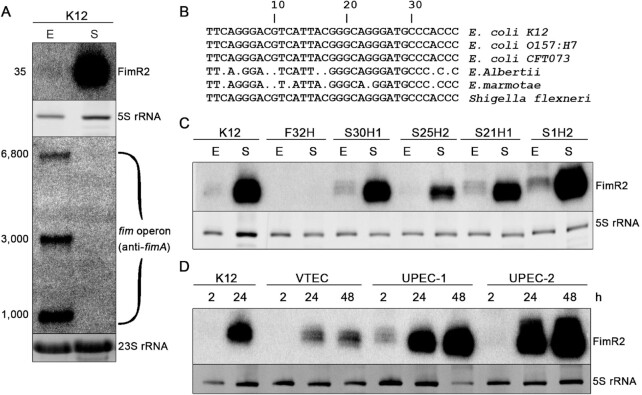
FimR2 is expressed in various E. coli strains. (A) Northern blot showing FimR2 and fimAICDFGH expression in the K-12 E. coli strain in exponential (E) and stationary (S) phase, respectively. Ethidium bromide staining of 5S rRNA and 23S rRNA are shown as loading controls on an 8% denaturing polyacrylamide gel and 1.2% agarose gel, respectively. Estimated transcript sizes are shown on the left in nucleotides. (B) Alignment of the FimR2 sequences from various enterobacterial strains. Sequential numbers indicate nucleotide positions. Dots indicate nucleotide mismatches compared to the K-12 strain. (C) Northern blot showing FimR2-phase dependent expression in ESBL E. coli strains. Total RNA samples from E (exponential phase) and S (stationary phase) are shown. Ethidium bromide staining of 5S rRNA is shown as a loading control. D. Northern blot analysis of FimR2 expression in VTEC, UPEC-1 and UPEC-2 strains. Total RNA samples from different time points of bacterial growth, 2, 24 and 48 h, are shown. Ethidium bromide staining of 5S rRNA is shown as a loading control.
Terminator exonuclease treatment
Terminator exonuclease treatment was done according to manufacturer manual. One microgram of total RNA from stationary phase was mixed with 1 unit TEX (Lucigen) and 4 units of RNasin Plus RNase inhibitor (Promega) in buffer A and incubated at 30°C for 60 min. Treated RNA was then phenol–chloroform extracted and precipitated in 2.5 volumes of 100% ethanol and 0.3 M sodium acetate.
In vitro transcription
FimR2, FimR2 mutants, and FimR2 precursors DNA templates were generated by PCR with primers containing the T7 promoter sequence (Supplemental Table S2). After PCR cleanup with Wizard® SV Gel and PCR Clean-Up (Promega), RNAs were transcribed by T7 polymerase as described (41). For crosslinking experiments, transcription was carried out using 4-thio-uridine (Jena Bioscience). All transcripts were purified by gel filtration with G-25 sephadex (Sigma). For 5′-end labelling for EMSA experiments and crosslinking experiments, in vitro transcribed FimR2 RNAs were dephosphorylated using CIP (New England Biolabs) at 37°C for 30 min. RNA was purified by phenol-chloroform extraction and precipitation in ethanol. RNA was then further purified by size-exclusion electrophoresis and 5′-end labelled with PNK (Thermo Fisher Scientific).
RNase E in vitro cleavage assay
The purified amino-terminal domain of RNase E was provided by B. F. Luisi (University of Cambridge). The following assay was carried out as described (42) with some modifications. Briefly, in vitro transcribed FimR2 precursors were treated with RppH (New England Biolabs) in NEB Buffer 2.0 to remove a pyrophosphate from the triphosphorylated 5′-end of the RNAs to generate RNase E cleavage templates containing a monophosphate at the 5′-end (43). The reaction was carried out at 37°C for 30 min then purified with a phenol-chloroform extraction. For in vitro cleavage, RNA was heat-denatured at 95°C for 1 min then incubated at room temperature for 10 min. Afterwards, 300 mM RNA was mixed with RNase E buffer (25 mM Tris pH 7.5, 50 mM NaCl, 50 mM KCl, 10 mM MgCl2, 1 mM DTT), 4 units of RNasin Plus RNase inhibitor (Promega), and with or without 300 mM CsrA-His10. The reactions were incubated at 30°C for 10 min then RNase E was added to a final concentration of 300 nM (+), 600nM (++) or 900 nM (+++). The reactions were then incubated at 30°C for an additional 30 min then phenol-chloroform extracted. RNA samples were precipitated at –20°C overnight in 2.5 volumes 100% ethanol, 0.3 M sodium acetate and glycoblue (Thermo Fisher Scientific). Precipitated RNA was resuspended in 2× RNA loading dye, resolved on denaturing polyacrylamide gels, and analyzed by northern blotting.
Co-immunoprecipitation and protein purification
CsrA-His10 was overexpressed in exponential phase using IPTG. For co-immunoprecipitation experiments, 50 ml cultures were harvested on the following day (24 h, stationary phase) from E. coli K-12 and K-12/pBbE6k-CsrA-His10 and from Salmonella serovar Typhimurium SL1344 sub-strain SB300 and SB300/pBbE6k-CsrA-His10. SB300 pellets were resuspended in phosphate buffered saline (1× PBS) and crosslinked in 0.37% formaldehyde for 15 min on ice. Quenching was done with 0.125 M glycine for 5 min at room temperature. Samples were briefly centrifuged to collect pellets. Thereafter, K-12 and SB300 pellets were treated similarly. Bacterial pellets were lysed in lysis buffer (50 mM NaH2PO4, 300 mM NaCl, 10 mM imidazole at pH 8.0), 1 mg/ml lysozyme and 10 mM VRC (New England Biolabs) for 30 min on ice. After sonication (6 rounds of 10 s, with 10 s pausing on ice in between), lysates were passed through a narrow-gauge needle to disrupt genomic DNA. Cell lysates were cleared through centrifugation at 10 000 × g for 30 min. Cleared lysates were incubated with Ni-NTA resin (Qiagen) at 4°C on a roller shaker. Lysates were loaded on Poly-Prep chromatography columns (BioRad) and flow-through fractions were collected. Columns were washed twice in wash buffer (50 mM NaH2PO4, 300 mM NaCl, 20 mM imidazole at pH 8.0). Beads were resuspended in elution buffer (50 mM NaH2PO4, 300 mM NaCl, 250 mM imidazole at pH 8.0). For all the collected fractions (input, flow-through, beads), samples were taken for western blot analysis and leftovers were used for RNA extraction using phenol-chloroform. Protein samples were analyzed by western blotting and RNA samples by northern blotting.
For CsrA-His10 purification, 100 ml cultures were harvested 4 h following induction in exponential phase. Pellets were lysed in native lysis buffer, 1 mg/ml lysozyme and 10 mM PMSF for 30 min on ice. After sonication and disruption of genomic DNA as described before, lysates were cleared by centrifugation. CsrA-His10 was then purified as described in the section on co-immunoprecipitation and using the Qiaexpressionist handbook. All fractions were analyzed by SDS-PAGE. Fractions containing CsrA-His10 were further confirmed by western blotting using mouse anti-His antibody (Protein Tech). Dialysis was performed overnight at 4°C in dialysis buffer (50 mM Tris–HCl pH 8.0, 150 mM NaCl, 10% glycerol, 1 mM DTT) and using Spectrum Spectra/Por 3 RC dialysis membrane tubing. Protein yield was quantified using Pierce BCA protein assay kit (Thermo Fisher Scientific).
Western blotting
For protein expression, purification and co-immunoprecipitation experiments, 5 μl of cell suspension or fractions were mixed with equal volume 2× Laemmli buffer and heated at 95°C for 5 min then loaded on 10–15% SDS-PAGE gels. For RFP assays, 30 μg total cleared cell lysates were mixed with 4× Laemmli buffer, boiled and loaded on SDS-gels. Protein samples were transferred to a nitrocellulose membrane (Sigma) using a semi-dry blotter (V20-SDB, Scie-Plas). Membranes were blocked in 1× PBSTM (1× PBS, 0.1% tween, 5% nonfat milk) for 1 h at room temperature on a shaker. Membranes were then incubated with the primary antibody prepared in 1× PBST in 1:5000 dilution, overnight at 4°C on a rotary shaker. Membranes were washed three times in 1× PBST then incubated for 1 h at room temperature with Licor anti-mouse antibody (1:20 000). After three washes, membranes were dried then scanned with a Licor scanner and visualized using ImageQuant software.
Crosslinking experiment
In vitro transcribed FimR2 sRNA containing 4-thio-Us was denatured at 95°C for 2 min then allowed to refold at room temperature for 10 min. The sRNA was then incubated with 30 μg total cell lysates extracted and quantified as described for protein extraction. Incubation was done at 37°C for 30 min. Samples destined for crosslinking were incubated on ice for 15 min under a UV lamp (365 nm). Control samples were incubated on ice without exposure to light. Proteinase K (Roth) treatment was done at 37°C for 15 min. All samples were heated at 95°C for 5 min then loaded on an SDS-PAGE for analysis. The gel was stained in a Coomassie solution, destained in destaining buffer (50% methanol, 10% acetic acid), and then dried on Whatman paper and exposed to a phosphor imaging screen overnight.
RNA EMSA
EMSA experiments were performed as described in Yakhnin et al. (44). 5′-end labelled RNAs of FimR2 or FimR2 mutants were heated at 85°C for 3 min then allowed to refold for 10 min at room temperature. RNAs were then incubated with or without CsrA-His10 in binding buffer (100 mM Tris–HCl pH 7.5, 100 mM MgCl2, 1 M KCl), and reaction mixture (7.5% glycerol, 0.2 μg yeast tRNA, 0.5% bromophenol native dye, 20 mM DTT, 0.04 U RNasin Plus RNase inhibitor). CsrA-His10 was diluted in dilution buffer (10 mM Tris–HCl pH 7.5, 2 mM DTT, 10% glycerol). Reactions were incubated at 37°C for 30 min then loaded on a 10% native gel (10% acrylamide, 2.5% glycerol, 0.5× TBE) and run for 1 h at 200 V. The gel was then wrapped in plastic foil and exposed to a phosphor imaging screen at –20°C.
RACE
5′- and 3′-RACE experiments were conducted using the FirstChoice™ RLM-RACE Kit (Invitrogen) and as per manual description. Twenty micrograms of total RNA were separated on an 8% denaturing polyacrylamide gel and using Riboruler LR RNA ladder (Thermo Fisher Scientific), gel pieces corresponding to RNA bands in the range of 20–50 nucleotides were cut, crushed, and eluted overnight at 4°C in elution buffer (0.3 M sodium acetate pH 5.5, 1 mM EDTA). On the following day, the supernatant was precipitated in 100% ethanol. RNA was then poly-adenylated using E. coli Poly(A)-Polymerase (New England Biolabs) as suggested at 37°C for 30 min. RNA was then purified from reaction components by a phenol–chloroform extraction. For 5′-RACE experiments, 5′-RACE adapter was annealed to template RNA using RNA Ligase (Thermo Fisher Scientific). cDNA synthesis was carried out using SuperScript IV reverse transcriptase (Thermo Fisher Scientific) with random hexamer primer (Thermo Fisher Scientific) with 10-min cycles at 23, 55 and 80°C. For 3′-RACE experiments, 3′-RACE adapter was used as a primer for cDNA synthesis using Superscript IV with 10-min cycles at 55°C and 80°C. FimR2S-specific primers were then used in combination with 5′-RACE and 3′-RACE outer primers in a first amplification step, and 5′-RACE and 3′-RACE inner primers in a second amplification step. PCR reactions were carried out using a homemade Taq Polymerase. PCR products were purified using Wizard® SV Gel and PCR Clean-Up (Promega) and ligated into the pGEM-T-Easy vector (Promega) using supplied DNA Ligase. Ligated products were transformed into XL1-Blue competent cells. Plasmid extraction was done with Wizard® Plus SV Minipreps DNA Purification System (Promega). Inserts were sequenced using T7 universal primer from Microsynth.
RT-qPCR
RT-qPCR was conducted as described (23). One microgram of total RNA from all experimental conditions were treated with DNase I (Thermo Fisher Scientific) to digest DNA, according to the manufacturer's protocol. Samples were then reverse transcribed into cDNA with SuperScript™ IV One-Step RT-PCR System (Invitrogen) and random primer hexamers (Thermo Fischer Scientific). After reverse transcription, cDNA samples were treated with RNase H (NEB) to hydrolyze leftover RNA. qPCR was done using GoTaq® qPCR Master Mix (Promega), 50-fold diluted cDNA, and a final concentration of 250 nM to 1 M of oligonucleotides. All primer pairs used for qPCR analysis were optimized using a standard curve. qPCR reactions were prepared by the CAS-1200 Corbett robot (Corbett Robotics) and analyzed using the Rotor Gene 6000, with suggested standard cycling conditions. recA was used as an internal control for the normalization of gene expression. All three biological replicates used for this analysis were run in duplicates. The 2−ΔΔCT method was used to calculate the fold-change relative to the control (45). The mean log2 fold-change and standard error of the mean were computed.
PCR
To confirm that the deletion strains acquired from the KEIO collection contain a kanamycin cassette instead of the ORF of the deleted locus, PCR amplification was done using oligonucleotides K1 and K2 combined with gene-specific primers (see Supplementary Table S2) as suggested in Datsenko and Wanner (37), and using home-made Taq Polymerase. For genomic DNA preparation, 1.5 mL of bacterial culture were lysed with 1.2 mg/ml Proteinase K (Roth) and 6% SDS in TE buffer (10 mM Tris–HCl pH 8.0 and 1 mM EDTA), at 37°C for 1 h. Genomic DNA was then extracted with basic phenol–chloroform extraction and precipitated in 2.5 volumes of 100% ethanol and 0.3 M sodium acetate.
Fluorescence assay
For RFP experiments, 500 μl of cultures, in two technical replicates and three biological replicates were pipetted into a 48-well plate (cellstar) following induction with IPTG and Tetracyclin. Fluorescence and OD600 were measured using a Tecan plate reader with parameters corresponding to an RFP excitation at 584 nm and emission at 607 nm. Background fluorescence from bacterial strains with sRNA overexpression plasmids and target-RFP expression plasmids were subtracted from experimental data.
For sicA-GFP, analysis was done similarly and GFP fluorescence was measured using Tecan plate reader with GFP excitation at 488 nm and emission at 525 nm. As FimR2S-S overexpression does not cause biofilm formation, fluorescence measurements were divided over OD600 of cultures (also measured using a Tecan plate reader) to obtain absolute fluorescence. Background fluorescence from SB300 cells normalized over OD600 was subtracted from experimental samples.
Biofilm assay
Biofilms assays were conducted as described in Merritt et al. (46) with minor modifications. Bacterial strains were diluted 1:100 from overnight cultures into fresh LB medium. At OD600 = 0.4, 200 μl of culture were inoculated in 96-well flat bottom microtiter plates (cellstar) for 2 h at 37°C (for exponential phase samples) and 22 h (for stationary phase samples). Planktonic bacteria were then removed by two washes with H2O. Biofilms were then stained with 0.1% crystal violet in water for 15 min at room temperature. Wells were washed three times with H2O and crystal violet was solubilized with a solution of 80% ethanol and 20% acetone. OD600 of solubilized crystal violet was measured using Vmax microplate reader (LabX Molecular Devices).
For micrographs, 500 μl of cultures were inoculated in 12-well flat bottom plates (TPP) mounted with a coverslip and incubated as indicated for liquid culture-based biofilm assays. Coverslips were then washed with H2O and stained with 0.1% crystal violet as described. Dried coverslips were mounted on glass microscope slides and visualized using a Leica Microscope.
Scanning electron microscopy
For visualizing bacterial cells in suspension by scanning electron microscopy, 150 μl of bacterial cells were taken from growing cultures 2 h after reaching OD600 = 0.4 (exponential phase) and 22 h post-inoculation (stationary phase). For inspecting biofilm potential, 1 ml of bacterial cells were inoculated in 24-well plate (TPP) mounted with 12 mm coverslips at OD600 = 0.4. Plates were incubated at 37°C for 2 h (exponential phase) or 22 h (stationary phase), under static conditions. Bacterial cell fixation was done as described in (47). Briefly, cell pellets and coverslips were washed with 0.1 M sodium cacodylate (pH 7.2) and fixed in 2% glutaraldehyde in sodium cacodylate buffer for 2 h at room temperature. Washes were done with 0.1 M sodium cacodylate and samples were further fixed with 2% osmium tetroxide in sodium cacodylate for 2 h at room temperature. After washing with water, the samples were dehydrated in increasing concentrations of ethanol (30%-100%). Samples were then resuspended or immersed in hexamethyl-disilazane and dried on metal SEM holders. Samples were sputter-coated with gold and visualized with a Zeiss Gemini 450 microscope operating at 5 kV.
HeLa infection assay
HeLa invasion assays were carried out as described in (48). Bacterial inocula for infection were cultured in LB medium with 0.3 M NaCl from overnight cultures at 37°C until OD550 = 0.6. Induction of SB300 strains containing pBbE6k-fimRS2-S was done with IPTG at OD550 = 0.4. Stationary phase samples were taken 24 h post-inoculation. HeLa (Kyoto) cells were seeded into 24-well plates (TPP) and grown in Dulbecco modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum (FBS) for 24 h at 37°C. Before infection with SB300 cells and derivatives, adherent cells were washed then incubated in Hank's buffered salt (HBSS) for 10 min. Infections were carried out using 3 × 106 CFU of bacteria per well, for 50 min at 37°C. Extracellular bacteria were inactivated by the addition of DMEM containing 10% FBS and gentamicin (400 μg/ml) and subsequent incubation at 37°C for 30 min. HeLa cells were then washed in PBS and lysed in 0.1% sodium-deoxycholate in PBS to release intracellular bacteria. The initial bacterial inocula and intracellular loads were quantified on LB/agar plates containing Streptomycin and/or Kanamycin.
CopraRNA analysis
The analysis was done using default parameters of CopraRNA (49) and with a search for target interactions 300 nucleotides upstream and downstream of mRNA start codons. As a search input, the first 18 nucleotides of the FimR2 sequence were used (5′-UUCAGGGACGUCAUUACG-3′).
MFold prediction
FimR2 secondary structure was predicted using MFold algorithm (50) with default values used for all parameters. The structure with the lowest predicted free energy is shown in this study.
Statistical analysis
Unpaired two-tailed t-test was used with Welsh's correction for all significance analysis and was conducted in Graphpad Prism. All analysis P-values are indicated in the figure legends.
RESULTS
FimR2 is abundantly expressed in various E. coli strains
We were intrigued by the abundant expression of FimR2 in stationary phase (23) knowing that the polycistronic mRNA fimAICDFGH is not expressed in this phase as previously reported (51). Indeed, we confirmed that the phase-dependent expression of FimR2 is coupled to the concomitant downregulation of fimAICDFGH expression (Figure 1A). FimR2 is conserved across various E. coli strains and within enteric bacteria (Figure 1B) suggesting an important role for this sRNA. Thus, we evaluated the expression of this sRNA by northern blotting in five extended spectrum beta-lactamases (ESBL)-producing recent human isolates of E. coli (52) (Figure 1C). All five isolates retain the genomic sequence of fimR2, and four of which expressed the sRNA in stationary phase to varying extents in comparison to the K-12 strain (Figure 1C). Extending our experiments also to pathogenic E. coli strains revealed an abundant expression of FimR2 in a verrotoxin-producing E. coli (VTEC) and two UPEC strains in a phase-dependent manner (Figure 1D).
FimR2 is an RNase E-dependent sRNA
The abundant expression of FimR2 in stationary phase of E. coli raises questions about the mechanism of its biogenesis. Deletion of the alternative sigma factor RpoS (53) did not abolish FimR2 expression (Supplementary Figure S2A). Furthermore, expression of FimR2 was not diminished by a fimA deletion (Supplementary Figure S2B) which abolishes a predicted internal transcription start site (TSS) (54). These findings suggested that FimR2 is not a primary transcript and indicated this sRNA as a processing product of fimAICDFGH. If this were the case, the expression of fimAICDFGH, which is governed by the two recombinases FimB and FimE (Supplementary Figure S1) (8), is a prerequisite for FimR2 biogenesis. Knockout of FimB, which allows both ON and OFF inversions to take place, abolished FimR2 expression in stationary phase (Supplementary Figure S2C). However, the knockout of the phase-OFF-specific recombinase FimE led to a prominent upregulation of FimR2 in both exponential and stationary phase (Supplementary Figure S2C). Additionally, FimR2 processing intermediates with various sizes accumulated exclusively in exponential phase upon fimE deletion (Supplementary Figure S2C). These experiments suggested that the biogenesis of FimR2 necessitates the prior expression of fimAICDFGH and that FimR2 is most likely cleaved from this transcript by an endonuclease specifically in stationary phase.
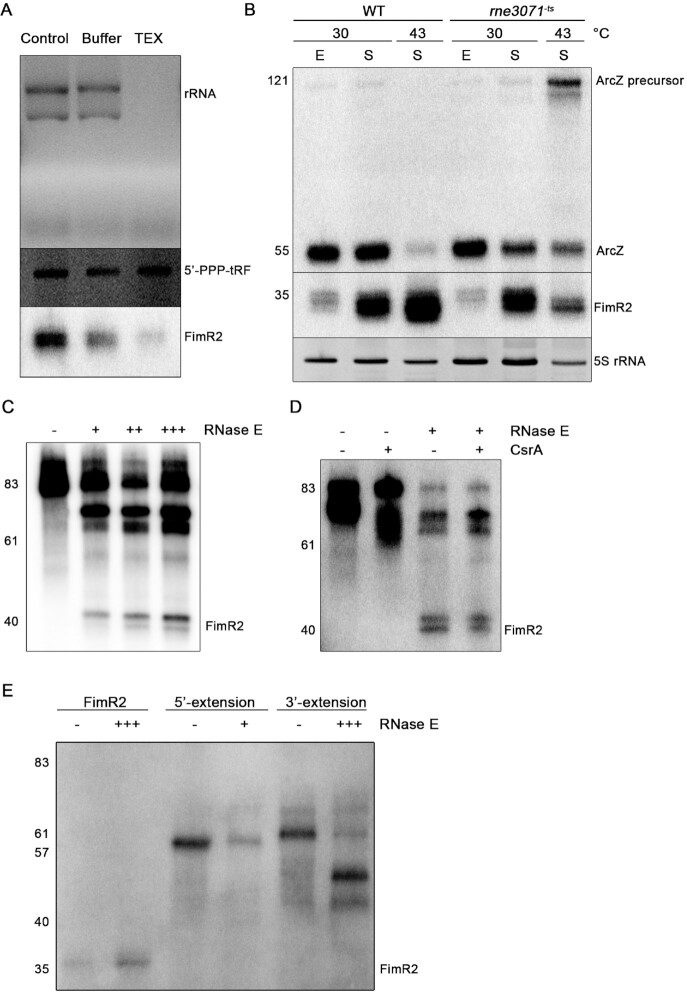
Knockout of RNase I, PNPase, RNase G, RNase R, RNase HI or RNase HII did not affect FimR2 levels during stationary phase (Supplementary Figure S2D). Similarly, a mutation that impairs RNase III function (55) did not change FimR2 abundance (Supplementary Figure S2E). A key endoribonuclease that is involved in mRNA turnover in stationary phase and in sRNA biogenesis is RNase E (13). RNase E cleavage generates RNA molecules with a monophosphate at the 5′-end (19). Similarly to rRNA, FimR2 was sensitive to TEX treatment, an exonuclease preferentially degrading RNA with a 5′ monophosphate, but not an in vitro transcribed tRNA fragment (tRF) with a 5′-triphosphate (Figure 2A). These findings suggested that FimR2 is processed and retains a 5′-end signature typical for RNase E cleavage. In support, FimR2 could not be efficiently processed in the RNase E temperature-sensitive strain rne3071−ts but was generated in the K-12 strain at the non-permissive temperature (Figure 2B). Simultaneously, the precursor of the RNase E-dependent sRNA ArcZ (42) accumulated in the rne3071−ts strain at the non-permissive temperature. This confirmed the role of RNase E in FimR2 processing. To further corroborate these findings, we used the purified catalytic domain of RNase E (56) in an in vitro cleavage assay with a FimR2 precursor containing extensions on both ends. Incubation of the FimR2 precursor with increasing amounts of RNase E led to the cleavage of the precursor to an intermediate size of about 40 nt (Figure 2C). Taken together, these findings suggest that RNase E governs FimR2 biogenesis from the parental fimAICDFGH transcript in stationary phase, adding FimR2 to the emerging class of 3′ UTR-derived sRNAs (57,58).
Figure 2.
FimR2 is processed by RNase E. (A) Ethidium bromide staining of ribosomal RNA and 5′-triphosphorylated tRNA fragment, and northern blot analysis of FimR2 upon TEX treatment. Total RNA samples were untreated (control), incubated with buffer, or with TEX. (B) Northern blot analysis of FimR2 and ArcZ sRNAs expression in WT (wild-type) and rne3071−ts (RNase E temperature-sensitive) strains. E and S denote total RNA samples extracted from exponential phase and stationary phase, respectively, and from incubations at the indicated temperatures. Ethidium bromide staining of 5S rRNA is shown as a loading control. (C) Northern blot analysis of FimR2 following in vitro cleavage of a 5′- and 3′-extended precursor with (+) and without (–) increasing amounts of RNase E. (D) Northern blot analysis of FimR2 following in vitro cleavage of a 5′- and 3′-extended precursor with (+) or without (–) RNase E and CsrA. (E) Northern blot analysis of FimR2 following in vitro cleavage of the sRNA and its precursors carrying a 5′-extension or a 3′-extension with (+++) or without (–) RNase E. In panels (B–E), nucleotide sizes are indicated in numbers on the left.
FimR2 promotes stationary phase-dependent biofilm formation and alters bacterial morphology
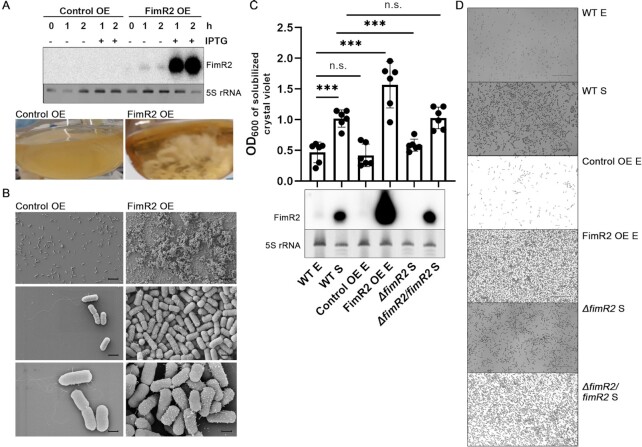
To gain insight into possible FimR2 functions, we ectopically expressed this sRNA in an inducible manner during exponential phase (23). Strikingly, FimR2 expressing strains aggregated in culture, a phenotype with no counterpart in the control strain (Figure 3A). Furthermore, the aggregated cellular suspension forms strongly adhering biofilms at the bottom and edges of the flask, suggesting that FimR2 overexpression causes biofilm formation. To inspect this phenotype further, we performed scanning electron microscopy (SEM) of the FimR2 overexpressing strain grown in microtiter plates mounted with coverslips. SEM revealed significant crowding of the coverslip with multiple embedded cells for the FimR2 overexpression strain, and only a modest bacterial population for the control strain, which overexpressed an unrelated RNA sequence (Figure 3B). We further corroborated these findings using quantitative biofilm assays and a gentler qualitative biofilm assay of bacterial cells grown on coverslips. Both approaches confirmed that FimR2 overexpression in exponential phase causes biofilm formation (Figure 3C, D). Interestingly, both wild-type (WT) and fimR2 genomic deletion (ΔfimR2) strains formed biofilms in stationary phase (Figure 3C, D) while ΔfimR2 showed markedly reduced biofilm formation potential compared to the WT strain. Genetic complementation of fimR2 deletion (ΔfimR2/fimR2) through ectopic expression of this sRNA restores biofilm formation to WT levels (Figure 3C, D). Interestingly, the fimE deletion strain, a strain transcribing the fimAICDFGH operon constitutively, phenocopied FimR2 overexpression in exponential phase and caused prominent biofilm formation (Supplementary Figure S2F, G).
Figure 3.
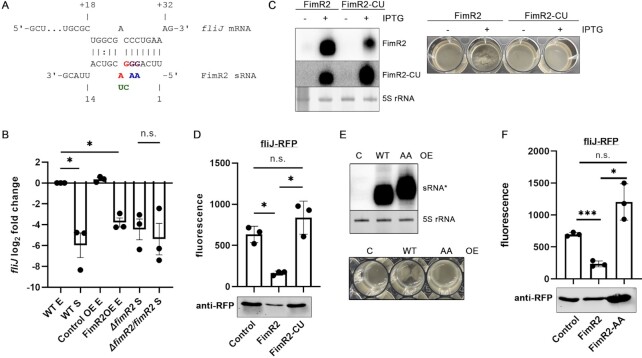
FimR2 regulates biofilm formation. (A) Northern blot (top) of FimR2 expression in exponential phase following overexpression (OE) of the sRNA via IPTG induction. Pictures of resulting cultures are shown on the bottom. Control OE refers to the expression of a random short RNA sequence. Ethidium bromide staining of 5S rRNA is shown as a loading control. (B) Scanning electron micrographs of coverslip-formed biofilms of Control and FimR2 OE strains. Scale bars = 10 μm (top), 1 μm (middle), and 500 nm (bottom). (C) Quantitative biofilm assay (top) showing mean ± SD OD600 of solubilized crystal violet-staining from six biological replicates from WT (wild-type), Control OE, FimR2 OE (FimR2 overexpression), ΔfimR2 (fimR2 deletion), and ΔfimR2/fimR2 (fimR2 complementation) strains. Samples are shown from exponential (E) and stationary (S) phase. Unpaired two-tailed t-test with Welch's correction was used to determine significance with n.s and *** indicating not significant and significant results, respectively. The P-values are from bottom to top 0.0001, 0.6369, 0.0003, 0.0002 and 0.9202, respectively. Northern blot of FimR2 expression (bottom) in the same strains as those in the upper panel. Ethidium bromide staining of 5S rRNA is shown as a loading control. (D) Micrographs of air-liquid phase biofilms stained with crystal violet, under conditions mentioned in (C). Scale bar = 25 μm.
To further validate the involvement of FimR2 in this phenotype, we carried out mutagenesis studies. MFold analysis predicted FimR2 to contain a single-stranded region and a stem-loop secondary structure (Supplementary Figure S3A). Mutations that individually disrupt the stem or the loop of this hairpin destabilized the sRNA and consequently eliminated the biofilm formation (Supplementary Figure S3B–D). A FimR2 compensatory mutant that allows the re-establishment of the stem–loop structure was stably expressed and functional in eliciting biofilm formation albeit to a lesser extent than the wild-type sequence (Supplementary Figure S3B-D). We conclude that this secondary structure likely exists in vivo and contributes to FimR2 stability, and thus possesses functional relevance to propagate biofilm formation.
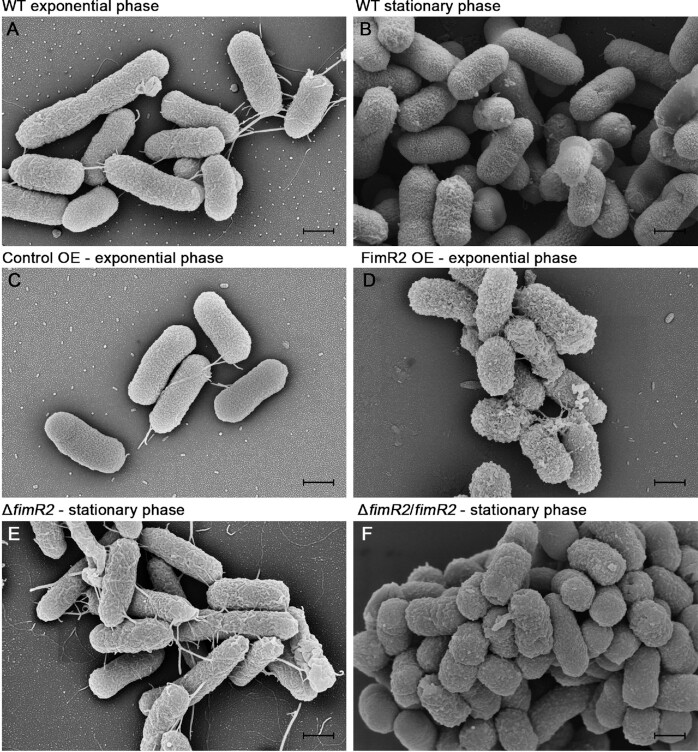
We next evaluated bacterial morphology and outer membrane architecture using SEM. These experiments revealed that bacterial cells derived from exponential growth phase were elongated and displayed flagella, fimbriae, and pili (Figure 4A). In contrast, bacterial cells in stationary phase were more ovoid and naked, displaying curli fimbriae (59) and PGA (poly-β-1,6-N-acetyl-d-glucosamine) (60) (Figure 4B). Strikingly, FimR2 overexpressing strains in exponential phase displayed pronounced aggregation and PGA synthesis and adopted a stationary phase-like morphology while the control overexpressing strains resembled exponentially growing cells (Figure 4C,D). ΔfimR2 and ΔfimR2/fimR2 strains significantly diverged in their morphologies from one another, although both strains were in stationary phase. ΔfimR2 exhibited exponential phase-like morphologies with extensive flagellation, while ΔfimR2/fimR2 mimics wild-type stationary phase physiology (Figure 4E, F). These SEMs thus showed that FimR2 expression and FimR2 deletion resulted in opposing morphologies on the extreme ends of a spectrum that separates stationary from exponentially growing cells, with control overexpressing and FimR2 compensation strains lying in the middle and behaving like their wild-type counterparts in their respective stages. In this manner, FimR2 appeared to trigger a significant outer membrane architecture remodeling, a critical process in the transition to stationary phase.
Figure 4.
FimR2 alters bacterial outer membrane architecture. Scanning electron micrographs of E. coli K-12 strains in (A) WT exponential phase, (B) WT stationary phase, (C) Control OE (Control overexpression) in exponential phase, (D) FimR2 OE (FimR2 overexpression) in exponential phase, (E) ΔfimR2 in stationary phase and (F) ΔfimR2/fimR2 in stationary phase. Scale bar = 500 nm.
FimR2 regulates cell motility and stationary phase biology
As bacterial sRNAs regulate mRNA targets at the post-transcriptional level (13), we searched for potential FimR2 sequence complementary targets using CopraRNA (49). Guided by the observed FimR2-dependent phenotype a subset of the predicted 200 targets were selected for experimental validation. Multiple predicted targets (fliJ, fliG, fliI and fliR) are mRNAs coding for flagellar biosynthesis proteins (61) (Figure 5A; Supplementary Figure S4A–C) three of which (fliJ, fliG and fliI) are part of the same operon (62). Additionally, one of the top candidates, hofQ (Supplementary Figure S4D), is a poorly characterized importer of exogenous DNA to be used in catabolic reactions in stationary phase (63). RT-qPCR revealed the flagellar biosynthesis operon and fliR to be downregulated upon FimR2 overexpression during exponential growth, a pattern resembling the canonical expression of these messages in stationary phase (Figure 5B; Supplementary Figure S4B, C). hofQ was upregulated upon FimR2 overexpression during exponential phase (Supplementary Figure S4D). ΔfimR2 perturbed the stationary phase expression pattern of all candidates while in the ΔfimR2/fimR2 strain the stationary phase-specific expression was restored (Figure 5B; Supplementary Figure S4A–C).
Furthermore, reporter assays showed that FimR2 overexpression, and not that of the stably expressed but non-functional FimR-CU mutant (Figure 5A, C), or the FimR2-AA mutant (Figures 5A,E), downregulated fliJ-RFP expression by almost 80% (Figure 5D, F). As FimR2 overexpression caused cell aggregation and biofilm formation (Figure 3A), fluorescence measurements were additionally validated by western blotting (Figure 5D, F), and these experiments fully confirmed the fluorescence data.
Figure 5.
FimR2 regulates fliJ mRNA at the post-transcriptional level. (A) FimR2-fliJ predicted interaction. Positions of mutated nucleotides are indicated in red or purple and the introduced nucleobase changes in green (FimR2-CU mutant) or blue (FimR2-AA mutant). (B) RT-qPCR analysis of fliJ in WT, Control OE, FimR2 OE, ΔfimR2 and ΔfimR2/fimR2 strains. Total RNA samples from E (exponential phase) and S (stationary phase) are shown. Mean log2 fold change ± SEM are shown from 3 biological replicates. log2 fold change was based on comparison with WT E samples. Unpaired two-tailed t-test with Welch's correction was used to determine significance with n.s and * indicating not significant and significant results, respectively. The P-values are 0.0373, 0.0128 and 0.6379, respectively. (C) Northern blot analysis of FimR2 and FimR2-CU (left) showing the expression of the sRNAs upon induction with IPTG, using two distinct probes that anneal differently to each sRNA. Ethidium bromide staining of 5S rRNA is shown as a loading control. Pictures of bacterial cultures from the same conditions (right) showing the aggregation upon FimR2 overexpression. (D) Mean ± SD of fluorescence of the fliJ-RFP fusion protein following control, FimR2 or FimR2-CU overexpression (top) and RFP western-blot of the same samples (bottom). Three biological replicates were used for these experiments and background fluorescence from individual sRNA overexpression strains were subtracted from experimental values. Unpaired two-tailed t-test with Welch's correction was used to determine significance with n.s. and * indicating not significant and significant results, respectively. The P-values are as follows: 0.0112, 0.0268 and 0.2153. (E) Northern blot analysis of FimR2 and FimR2-AA (top) showing the expression of the sRNAs upon overexpression of either sRNA or a control plasmid. Ethidium bromide staining of 5S rRNA is shown as a loading control. The asterisk (*) refers to both FimR2 and FimR2-AA (a single probe that anneals to both sRNAs was used). Pictures of bacterial cultures from the same conditions (bottom) showing the aggregation upon FimR2 overexpression. (F) Mean ± SD of fluorescence of the fliJ-RFP fusion protein following control, FimR2 or FimR2-AA overexpression (top) and RFP western-blot of the same samples (bottom). Three biological replicates were used for these experiments and background fluorescence from individual sRNA overexpression strains were subtracted from experimental values. Unpaired two-tailed t-test with Welch's correction was used to determine significance with n.s. and * indicating not significant and significant results, respectively. The P-values from bottom to top are as follows: 0.0003, 0.0250 and 0.0892.
Taken together, these findings showed that FimR2 acts as a trans-acting sRNA, inhibiting the expression of fliJ and potentially other transcripts involved in motility (fliG, fliI and fliR), thereby eliciting biofilm formation. Concomitantly, FimR2 promotes the expression of hofQ, providing means for the stressed bacterium to import foreign DNA for catabolic reactions.
FimR2 interacts with CsrA
We next evaluated the dependence of FimR2 on known RNA chaperones by monitoring its stability following hfq and proQ deletions. FimR2 abundance was unaffected in stationary phase in the absence of either chaperone Hfq and ProQ (Supplementary Figure S5A). Furthermore, FimR2 was not recovered in RIL-seq analyses of both chaperones (64). Taken together, these observations suggested that, unlike most trans-encoded sRNAs, FimR2 acts independently of both Hfq and ProQ. Another prominent bacterial RNA-binding protein (RBP) is the translational regulator CsrA that is involved in a plethora of cellular activities (65). Interestingly, a recent CLIP-seq analysis revealed that CsrA interacts with both the fimA sequence and the intergenic fimA-fimI region (66). Co-immunoprecipitation of ectopically expressed his-tagged CsrA in stationary phase revealed a significant enrichment of FimR2 in the eluted fractions (Figure 6A). The known CsrA-interacting sRNA CsrB (67) was also recovered in the same fractions. However, full-length tRNAGly and its 5′-tRNA fragment, a stationary-phase enriched sRNA that we previously recovered (23), were both largely depleted from these fractions. Taken together, these pulldown data suggested FimR2 and CsrA to interact in vivo.
Figure 6.
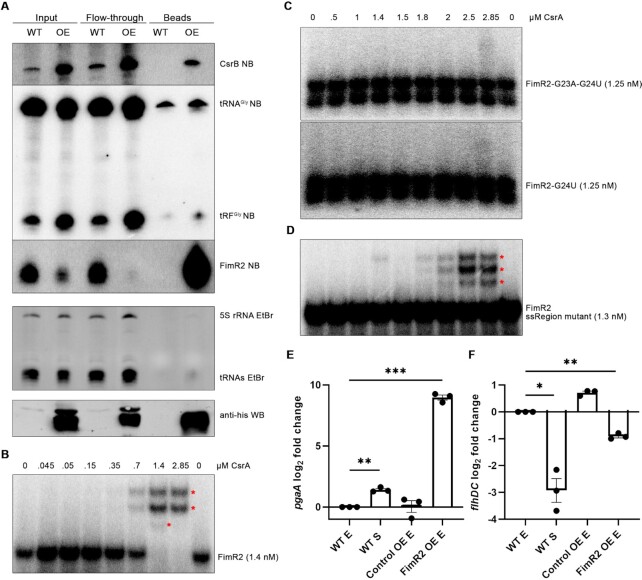
FimR2 sequesters CsrA from its targets. (A) Northern blot of CsrB, 5′-tRFGly and FimR2 (top) from CsrA-his10 CoIP fractions (top) and western blot of CsrA-his10 from the same samples (bottom). Samples shown are taken from the stationary phase of growth of WT and OE (CsrA-his10 overexpression) strains. Ethidium bromide staining of 5S rRNA and tRNAs are shown as loading controls. (B) EMSA of radioactively labelled FimR2 with increasing concentrations of purified CsrA-his10. (C) EMSA of radioactively labelled FimR2-G23A-G24U (top) and FimR2-G24U (bottom) with increasing concentrations of purified CsrA-his10. (D) EMSA of radioactively labelled FimR2 ssRegion mutant with increasing concentrations of purified CsrA-his10 (as in C). Upshifts in (B–D) are marked with red asterisks. (E) RT-qPCR analysis of pgaA and (F) flhDC expression from WT, Control OE, and FimR2 OE. Samples from E (exponential phase) and S (stationary phase) are shown. Mean log2 fold change ± SEM are shown for both transcripts from three biological replicates. log2 fold change was based on comparison with exponential phase samples. Unpaired two-tailed t-test with Welch's correction was used to determine significance with * indicating significant results. The P-values are (E) 0.0058 and 0.005, (F) 0.0221 and 0.0046.
To corroborate these findings, we carried out electrophoretic mobility shift assays (EMSA) with in vitro transcribed and 5′-end labelled FimR2 and purified CsrA. These EMSA experiments showed that FimR2 upshifts with increasing concentrations of purified CsrA (Figure 6B). CsrA favorably binds GGA motifs, with a preference to those located in secondary structures (66). The FimR2 sequence contains two GGA motifs with the first located in the single-stranded region, and the second in the loop structure (Supplementary Figure S5B), suggesting that the sRNA could use both motifs to interact with a CsrA dimer. Mutations of the GGA motif in the FimR2 stem-loop region, but not in its single-stranded region (ssRegion), abolished association with CsrA (Figure 6C, D). Notably, mutations in the FimR2 loop region did not destabilize the mutants (Supplementary Figure S5C) despite their loss of interactions with CsrA (Figure 6C). Importantly, the overexpression of these loop mutants in exponential phase still caused biofilm formation (Supplementary Figure S5D). Taken together, these experiments suggested that CsrA binds FimR2 at the stem-loop region and that this interaction is dispensable for biofilm formation.
We next hypothesized that CsrA may be stabilizing FimR2 and thus we altered cellular CsrA availability to test FimR2 dependence on this RBP. CsrA is sequestered from its targets through its interaction with sRNAs CsrB and CsrC (67,68). These sRNAs are degraded by RNase E through the adaptor activity of CsrD (69). As such, deletion of CsrD upregulated CsrB and CsrC expression but did not affect FimR2 levels (Supplementary Figure S5E), suggesting that the FimR2 interaction with CsrA is not critical for its stability. Subsequently, deletion of either CsrB or CsrC sRNAs or both simultaneously did not boost FimR2 expression (Supplementary Figure S5F). Therefore CsrB/CsrC and FimR2 are functionally independent sRNAs and their binding to and modulations of CsrA are distinct. Furthermore, the addition of purified CsrA to in vitro cleavage reactions did not enhance the RNase E-dependent processing of the FimR2 precursor (Figure 2D). We thus concluded that despite the FimR2/CsrA interaction, CsrA does not stabilize FimR2 in vivo and thus this interaction must be fulfilling another distinct function.
FimR2 has the potential for sequestering CsrA from its targets
The interactions of CsrA with sRNA partners have largely been associated with antagonizing mechanisms with the sRNAs sequestrating the translational regulator from its mRNA targets (67,68,70). We thus considered the hypothesis that FimR2 is sequestering CsrA from its targets. Interestingly, SEM showed depletion of T1P under conditions giving rise to prominent FimR2 expression (Figure 4B, D, F). Under similar conditions, RT-qPCR analysis revealed significant downregulation of fimA and other individual members of the fim polycistron (Supplementary Figure S6). CsrA has been shown to stabilize fimAICDFGH (71) and to interact with the fim operon (66). In this manner, the interaction of FimR2 with CsrA could be antagonizing this stabilization of the fim transcript, suggesting that FimR2 is sequestering CsrA and antagonizing its activity.
Prominent morphological features seen following FimR2 overexpression on SEM include increased PGA synthesis and a dramatic loss of flagellation (Figure 4D). CsrA is known to modulate both phenotypes by allowing premature rho-dependent transcription termination of the pgaABCD transcript (72) and protecting the flhDC transcript from RNase E degradation (73). RT-qPCR analysis showed dramatic upregulation of pgaA upon FimR2 overexpression (Figure 6E), suggesting that binding of FimR2 to CsrA prevented the latter from downregulating pgaA. In parallel, FimR2 overexpression downregulates flhDC (Figure 6F) presenting the opposite scenario of the regulation by CsrA. These two experiments present so far unknown examples of CsrA antagonisms by FimR2 and suggest that FimR2 sequesters CsrA away from its targets in similar fashions as reported for CsrB, CsrC, and other sRNAs (65). Furthermore, this sequestration model is compatible with the observed prominent biofilm formation upon FimR2 overexpression (Figure 3A), resembling those seen upon CsrA deletion (74). Importantly, FimR2 appears to regulate biofilm formation in a dual mode since removing the capability for sequestrating CsrA (by mutating the loop of the hairpin structure; Figure 6C) still triggered biofilm formation (Supplementary Figure S5D) by utilizing its single stranded domain for mRNA target regulation.
FimR2 promotes S. enterica cell invasion
As the sequence of the sRNA is conserved in many Escherichia and Shigella species (Figure 1B), we wondered if FimR2 is present in pathogens from the Salmonella genus. RNase E was shown to cleave near the fimA stop codon in S. enterica generating a small transcript (32) whose genomic locus is similar to that of E.coli FimR2 (Figure 7A). We refer to this putative sRNA as FimR2S to denote it as the FimR2 variant in Salmonella. Northern blot analyses demonstrated the expression of a ∼30-nt RNA in late stationary phase in the assayed Salmonella strains (Figure 7B). S. enterica often express T1P under laboratory growth conditions (75). This suggests that the lower abundance seen for FimR2S as compared to E. coli is due to the continuous T1P expression in these Salmonella strains.
Figure 7.
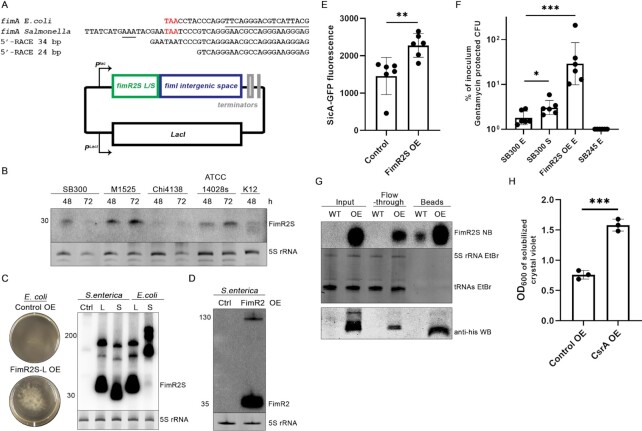
Salmonella FimR2S is involved in infection. (A) 5′-RACE results mapped to S. enterica fimA sequence (top) and FimR2S expression plasmid (bottom). fimA sequences from E. coli and Salmonella are shown with the stop codons indicated in red. The first 18 nucleotides of E. coli FimR2 are underlined. The predicted sequence for RNase E cleavage (AAA) in the SalmonellafimA locus is underlined (32). Sizes of recovered transcripts are indicated in bp (base pairs). (B) Northern blot analysis of FimR2S in four Salmonella strains and the E. coli K-12 strain. Total RNA samples from different time points of bacterial growth, 48 and 72 h (hours), are shown. Ethidium bromide staining of 5S rRNA is shown as a loading control. (C) Pictures of Control and FimR2S-L OE in E. coli (left) and northern blot analysis of FimR2S expression in S. enterica and E. coli upon overexpression of FimR2S-L and FimR2S-S, the long and short FimR2S variants, respectively. Ctrl designate overexpression of the control. Ethidium bromide staining of 5S rRNA is shown as a loading control. The size of the different RNA molecules is indicated on the left. (D) Northern blot analysis of FimR2 expression in S. enterica under control and FimR2 OE conditions. Ethidium bromide staining of 5S rRNA is shown as a loading control. (E) Mean ± SD of sicA-GFP fluorescence without (Control) or with FimR2S OE from six biological replicates. Unpaired two-tailed t-test with Welch's correction was used to determine significance with ** indicating significant results and P-value of 0.0086 as compared to the control samples. (F) Mean ± SD of percentage of inoculum protected from gentamicin treatment following infection of HeLa cells with an initial inoculum of SL1344 (SB300) from E (exponential phase) and S (stationary phase), and FimR2S overexpression strains, from six replicate infections. Strain SB245 served as a non-invasive negative control. Calculations were done following counting of colony forming units (CFU) of protected cells and initial inocula. Unpaired two-tailed t-test with Welch's correction was used to determine significance with * indicating significant results. The P-values are, in order, 0.0280 and 0.0010. (G) Northern blot of FimR2S (top) from CsrA-his10 CoIP fractions and western blot of CsrA-his10 from the same samples (bottom). Samples shown are taken from the stationary phase of growth of WT (Control OE) and OE (CsrA-his10 overexpression) strains. Ethidium bromide staining of 5S rRNA and tRNAs are shown as loading controls. (H) Quantitative biofilm assay showing mean ± SD OD600 of solubilized crystal violet-staining from three biological replicates from Control OE and CsrA OE strains. Unpaired two-tailed t-test with Welch's correction was used to determine significance with *** indicating significant results (P-value = 0.005).
5′-RACE experiments (Figure 7A) showed that this putative sRNA exists as a lower abundance, longer transcript (FimR2S-L) that retains part of the fimA coding sequence, and a shorter more abundant form (FimR2S-S) solely generated from the fimA-fimI intergenic space. To understand the potential functions of FimR2S, we overexpressed both variants in S. enterica and E. coli (Figure 7A) Strikingly, overexpression of FimR2S-L, but not FimR2S-S in E. coli caused biofilm formation (Figure 7C). Ectopic overexpression of neither of the two FimR2S variants in S. enterica promoted biofilm formation. Similarly, the overexpression of the E. coli FimR2 in S. enterica (Figure 7D) did not promote biofilm formation. Taken together, these experiments suggested that Salmonella FimR2S mimics FimR2 functions in E. coli but FimR2S is not functional in S. enterica. Alternatively, FimR2S may have a different function during stationary phase in S. enterica than in E. coli.
S. enterica infectivity largely relies on the secretion of several effectors by the T3SS (2). Considering the effects of FimR2 on biofilm formation (Figure 3A) and outer membrane architecture (Figure 4D) in E. coli, we hypothesized that the sRNA is involved in virulence gene regulation. SicA is a critical T3SS chaperone that promotes the expression of cell invasion-mediating effector proteins (76). Significantly, FimR2S overexpression in Salmonella induced sicA-GFP fusion protein (77) in a reporter assay (Figure 7E), suggesting that the Salmonella sRNA is functional in promoting SPI-1 T3SS expression. Next, we hypothesized that the sRNA could regulate invasiveness. To test this, we infected HeLa cells with wild-type S. enterica from both exponential and stationary phase and with the FimR2S overexpression strain (48). Using the isogenic mutant strain SB245, which is invasion deficient as a negative control, we quantified the intracellular bacterial load following treatment with gentamicin, which is membrane impermeable and consequently kills non-invaded extracellular bacteria selectively. According to this invasion assay, the FimR2S overexpressing strain was markedly more invasive than either wild-type strain (Figure 7F). To investigate possible mechanistic similarities between FimR2 and FimR2S functions in respect to the involvement of CsrA, we overexpressed a His-tagged version of CsrA in Salmonella and used it for a co-immunoprecipitation assay. Figure 7G clearly confirms that, analogously to the situation in E. coli, also in Salmonella FimR2S associates with CsrA in vivo. Interestingly, ectopic overexpression of CsrA in Salmonella led to a simultaneous upregulation of FimR2S from its chromosomal locus (Figure 7G) with the concomitant induction of biofilm formation (Figure 7H). Therefore, these findings revealed a further functional similarity of this sRNA in E. coli and Salmonella in respect to biofilm formation. Moreover, the results suggested that FimR2S is a functional sRNA in S. enterica where it promotes invasion and internalization of bacterial cells into human cells.
DISCUSSION
In this study, we functionally characterized FimR2, an abundant and conserved RNase E-dependent sRNA that is specifically expressed in stationary phase of bacterial growth. 3′ UTR-derived sRNAs have only been recently recognized as a novel group of regulatory molecules in various bacterial species where they have been functionally linked to adaptation processes during challenging environmental cues (78). With the generated knowledge we confidently ascribe a master regulatory status to FimR2, a so far uncharacterized 3′ UTR-derived sRNA that coordinates motility, biofilm formation, stationary phase biology, and even virulence in E. coli and S. enterica, apparently without the assistance of the two so far best characterized RNA chaperones Hfq or ProQ (Figure 8).
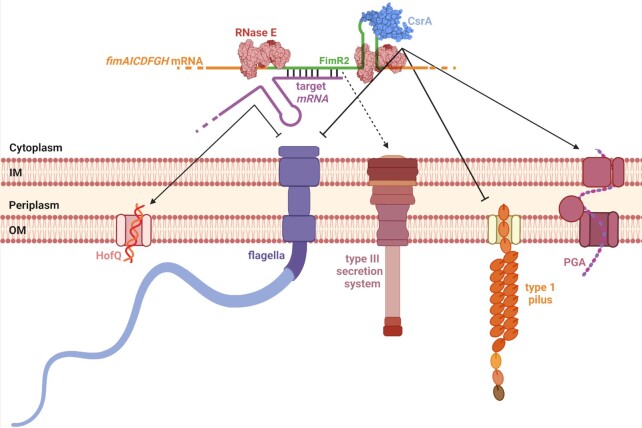
Figure 8.
Model of FimR2 function. FimR2 sRNA is processed from the fimAICDFGH transcript in stationary phase by RNase E. On the one hand, the sRNA (green) interacts with the translational regulator CsrA and antagonizes its effects, upregulating PGA synthesis, and downregulating type 1 pilus and flagellar synthesis. FimR2 upregulates type III secretion likely through the sequestration of CsrA. On the other hand FimR2 regulates in parallel several transcripts through direct base-pairing, inhibiting flagellar synthesis and upregulating HofQ-mediated import of extracellular DNA for use in catabolic reactions. The crystal structure of RNase E catalytic domain (2C4R) and the NMR structure of CsrA (1Y00) were used (56,85). This figure was created with www.biorender.com.
The simultaneous modulation of biofilm formation, motility, stationary phase biology, and even virulence is not a trivial task. The success of FimR2 in these strenuous activities and its ability to navigate several branches within a complex regulatory network can be attributed to two distinct features: FimR2 dual mode of function and its abundance. Bacterial trans-encoded sRNAs are efficient and potent regulatory molecules as they typically employ their single-stranded regions to base-pair and regulate various targets mRNAs at the post-transcriptional level (14). Indeed, the range of FimR2 mRNA targets is very diverse (Figure 5A, Supplementary Figure S4A–D) and includes transcripts coding for flagellar proteins and porins, among many others. Another mode by which FimR2 exhibits utmost efficiency is through its proposed sequestration of the translational regulator CsrA. CsrA itself is a master regulator of gene expression, governing diverse cellular processes including motility, biofilm formation, virulence, quorum sensing, metabolism, and oxidative stress (65). By sequestering CsrA from mediating its cellular functions, FimR2 automatically gains regulatory access to the CsrA-specific expression network and navigates its intricate web, remodeling its fibers in an antagonistic manner. Such a dual regulatory function has also been attributed to other sRNAs such as McaS (70). FimR2 is enriched in stationary phase, the phase in which both well-characterized CsrA-binding sRNAs CsrB and CsrC are abundantly expressed (Supplementary Figure S5E, F) and have been described to be induced (68,79). However, FimR2 binds CsrA with an apparently weaker affinity than both CsrB and CsrC as FimR2 failed to interact with CsrA when EMSA experiments were performed with protein concentrations in the nanomolar range as described by Weilbacher et al. (68) (data not shown). This suggests that the abundance of the FimR2 sRNA plays an important role in the sequestration of the CsrA. Additionally, FimR2 overexpression caused the upregulation of rpoS, the stationary phase-dependent RNA polymerase sigma factor (23). While this inductive effect is likely indirect, as we do not observe a putative base-pairing site for FimR2 within the rpoS mRNA sequence, it explains the propensity of bacterial cells to form biofilms (Figure 3A) and to adopt stationary phase-like morphology (Figure 4B, D, F). Interestingly, overexpression of FimR2, but not of the ssRegion mutants, in the rpoS deletion strain caused biofilm formation (Supplementary Figure S7), suggesting that FimR2 may be regulating stationary phase biology through its interplay with RpoS, but not biofilm formation. Future work is needed for a better understanding of the mechanism by which FimR2 affects rpoS expression. With the existence of these various parallel examples, it is now more evident that bacterial species have evolved efficient and elegant regulatory modes to rapidly adapt to unpredictable environments.
FimR2 abundance is another feature contributing to its potent effect. While the expression of such an abundant RNA molecule would normally entail overwhelming energy expenditure, FimR2 biogenesis constitutes another efficient mechanism. As the accumulation of this sRNA occurs in stationary phase, the RNase E-dependent processing event likely overlaps with fimAICDFGH mRNA turnover during the same growth phase. In vitro cleavage experiments of a FimR2 precursor containing extensions at both the 5′- and the 3′-ends generated a 40-nt intermediate RNA (Figure 2C, D). Furthermore, in vitro cleavage assays with total cell lysates generated the same 40-nt intermediate (data not shown) suggesting the in vitro precursor transcript lacking crucial determinants for complete processing, such as post-transcriptional modifications or the establishment of correct three-dimensional architecture. Experiments with FimR2 transcripts containing an extension either at the 5′- or the 3′-ends demonstrated cleavage thus indicating RNase E processing of FimR2 precursors from both ends (Figure 2E).
That FimR2 is a functional sRNA independently of the known RNA chaperones Hfq and ProQ is surprising as most trans-encoded regulatory sRNAs in bacterial species require a chaperone to maintain their stabilities or to anneal to their target mRNAs (13). In variance, FimR2 interacts with the bacterial translational regulator CsrA. Compatible with this finding, CsrA and FimR2 (termed sRNA_00370 in the study of Hör et al.) have been shown to co-migrate in the same glycerol gradient fractions in E. coli (80). However, it is also possible that FimR2 associates with a so far uncharacterized RNA chaperone in vivo, a hypothesis in line with the observed upshift patterns seen in crosslinking experiments (Supplementary Figure S5G).
FimR2 promotes stationary phase biology, biofilm formation and suppresses bacterial motility (Figures 3A, 5B, D, 6E, F). All three processes have been shown to be important for bacterial virulence (11,81,82). Indeed, S. enterica FimR2S promotes pathogen invasiveness (Figure 7F), suggesting that the sRNA may play similar functions in the pathogenic E. coli strains through its phase-dependent expression. The mechanism by which this sRNA promotes S. enterica virulence is unclear. However, we hypothesize that in S. enterica, FimR2S can function to sequester CsrA from its targets in a comparable manner as in E. coli. The upregulation of SicA and the increased invasiveness of S. enterica upon FimR2S overexpression (Figure 7E, F) are two phenotypes also displayed by S. enterica upon mutation of CsrA (83). Like FimR2, FimR2S also retains two GGA motifs that could be employed to interact with CsrA. Furthermore, the 5′ UTR of the Salmonella fim transcript has been reported previously to interact with CsrA (84). Similarly to the suggested scenario in E. coli, Salmonella FimR2S could be sequestering the RBP from the 5′ UTR of its parental transcript. Support for this scenario comes from co-immunoprecipitation experiments demonstrating interaction of FimR2S with CsrA in Salmonella (Figure 7G). Another functional similarity between E. coli FimR2 and S. enterica FimR2S is the fact that both sRNAs trigger biofilm formation under overexpression condition (Figures 3 and 7G, H).
It appears that FimR2 is a conserved Enterobacterales sRNA regulating diverse and biologically important phenotypes relevant for stationary phase biology and/or infectivity. FimR2 might play an even more intricate role as global regulator on the population level during infections, as it has been demonstrated to be one of the most abundant secreted RNA molecules in outer membrane vesicles (25). Therefore our study can be regarded as steppingstone for future dedicated work on FimR2 for elucidating even more elaborate mechanisms by which this tiny RNA molecule weaves its regulatory network.
DATA AVAILABILITY
All relevant data are within the paper and its supplementary data.
Supplementary Material
ACKNOWLEDGEMENTS
We would like to thank Marc Landolfo for experimental help, Alexander Mankin (UIC, Chicago) for providing KEIO strains, Eric Massé (University of Sherbrooke) for sharing the rne3071−ts strain, Ben Luisi (University of Cambridge) for sharing active RNE-NTD fractions, and Markus Hilty (IFIK, Bern) for providing E. coli human and pathogenic strains. We would also like to thank Andrew Hemphill (VetSuisse, Bern) for providing expertise and reagents for SEM, and Beatrice Frey (DCBP, Bern) for assisting with the Zeiss Microscope.
Author contribution: Conceptualization, N.R. and N.P.; Methodology, N.R., T.D. and S.H.; Formal Analysis, N.R.; Investigation, N.R.; Resources, N.R.; N.P., T.D. and S.H.; Writing – Original Draft, N.R.; Writing – Review and Editing, N.P., N.R. and S.H.; Supervision, N.P.; Funding Acquisition, N.P.
Notes
Present address: Nicole Raad, Department of Molecular Biology, Science III, University of Geneva, 1211 Geneva, Switzerland.
Contributor Information
Nicole Raad, Department of Chemistry, Biochemistry, and Pharmaceutical Sciences, University of Bern, Bern, Switzerland; Graduate School for Cellular and Biomedical Sciences, Bern, Switzerland.
Disha Tandon, Graduate School for Cellular and Biomedical Sciences, Bern, Switzerland; Institute for Infectious Diseases, University of Bern, Bern, Switzerland.
Siegfried Hapfelmeier, Institute for Infectious Diseases, University of Bern, Bern, Switzerland.
Norbert Polacek, Department of Chemistry, Biochemistry, and Pharmaceutical Sciences, University of Bern, Bern, Switzerland.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Swiss National Science Foundation [310030-197515 to N.P.]; NCCR ‘RNA& Disease’ (205601 to N.P.); NCCR ‘Microbiomes’ (to S.H.) funded by the Swiss National Science Foundation; S.H. and D.T. were additionally supported by a grant of the Helmut-Horten-Stiftung (2018–2021). Funding for open access charge: Swiss National Science Foundation [310030-197515].
Conflict of interest statement. None declared.
REFERENCES
- 1. Kaper J.B., Nataro J.P., Mobley H.L.T.. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004; 2:123–140. [DOI] [PubMed] [Google Scholar]
- 2. Galán J.E. Salmonella typhimurium and inflammation: a pathogen-centric affair. Nat. Rev. Microbiol. 2021; 19:716–725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Sy B.M., Tree J.J.. Small RNA regulation of virulence in pathogenic Escherichia coli. Front. Cell. Infect. Microbiol. 2021; 10:622202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Martínez L.C., Yakhnin H., Camacho M.I., Georgellis D., Babitzke P., Puente J.L., Bustamante V.H.. Integration of a complex regulatory cascade involving the SirA/BarA and csr global regulatory systems that controls expression of the salmonella SPI-1 and SPI-2 virulence regulons through hilD. Mol. Microbiol. 2011; 80:1637–1656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Bhatt S., Edwards A.N., Nguyen H.T., Merlin D., Romeo T., Kalman D. The RNA binding protein CsrA is a pleiotropic regulator of the locus of enterocyte effacement pathogenicity island of enteropathogenic Escherichia coli. Infect. Immun. 2009; 77:3552–3568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Henderson I.R., Owen P., Nataro J.P.. Molecular switches–the ON and OFF of bacterial phase variation. Mol. Microbiol. 1999; 33:919–932. [DOI] [PubMed] [Google Scholar]
- 7. Abraham J.M., Freitag C.S., Clements J.R., Eisenstein B.I.. An invertible element of DNA controls phase variation of type 1 fimbriae of Escherichia coli. Proc. Nat. Acad. Sci. U.S.A. 1985; 82:5724–5727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. McClain M.S., Blomfield I.C., Eisenstein B.I.. Roles of fimB and fimE in site-specific DNA inversion associated with phase variation of type 1 fimbriae in Escherichia coli. J. Bacteriol. 1991; 173:5308–5314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kolenda R., Ugorski M., Grzymajlo K.. Everything you always wanted to know about salmonella type 1 fimbriae, but were afraid to ask. Front. Microbiol. 2019; 10:1017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Schwan W.R. Regulation of fim genes in uropathogenic Escherichia coli. World J. Clin. Infect. Dis. 2011; 1:17–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Naziri Z., Kilegolan J.A., Moezzi M.S., Derakhshandeh A.. Biofilm formation by uropathogenic escherichia coli: a complicating factor for treatment and recurrence of urinary tract infections. J. Hosp. Infect. 2021; 117:9–16. [DOI] [PubMed] [Google Scholar]
- 12. Wang L., Keatch R., Zhao Q., Wright J.A., Bryant C.E., Redmann A.L., Terentjev E.M.. Influence of type i fimbriae and fluid shear stress on bacterial behavior and multicellular architecture of early Escherichia coli biofilms at single-cell resolution. Appl. Environ. Microbiol. 2018; 84:e02343-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Hör J., Matera G., Vogel J., Gottesman S., Storz G.. Trans-acting small RNAs and their effects on gene expression in Escherichia coli and Salmonella enterica. EcoSal Plus. 2020; 9: 10.1128/ecosalplus.ESP-0030-2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Nitzan M., Rehani R., Margalit H.. Integration of bacterial small RNAs in regulatory networks. Annu. Rev. Biophys. 2017; 46:131–148. [DOI] [PubMed] [Google Scholar]
- 15. Chambers J.R., Sauer K.. Small RNAs and their role in biofilm formation. Trends Microbiol. 2013; 21:39–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Westermann A.J. Regulatory RNAs in virulence and host-microbe interactions. Microbiol. Spectrum. 2018; 6: 10.1128/microbiolspec.RWR-0002-2017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Gruber C.C., Sperandio V., McFall-Ngai M.J.. Posttranscriptional control of microbe-induced rearrangement of host cell actin. Mbio. 2014; 5:e01025-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Westermann A.J., Förstner K.U., Amman F., Barquist L., Chao Y., Schulte L.N., Müller L., Reinhardt R., Stadler P.F., Vogel J.. Dual RNA-seq unveils noncoding RNA functions in host–pathogen interactions. Nature. 2016; 529:496–501. [DOI] [PubMed] [Google Scholar]
- 19. Chao Y., Vogel J.. A 3′ UTR-derived small RNA provides the regulatory noncoding arm of the inner membrane stress response. Mol. Cell. 2016; 61:352–363. [DOI] [PubMed] [Google Scholar]
- 20. Santiago-Frangos A., Woodson S.A.. Hfq chaperone brings speed dating to bacterial sRNA. WIREs RNA. 2018; 9:e1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Smirnov A., Wang C., Drewry L.L., Vogel J.. Molecular mechanism of mRNA repression in trans by a proq-dependent small RNA. EMBO J. 2017; 36:1029–1045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Mai J., Rao C., Watt J., Sun X., Lin C., Zhang L., Liu J.. Mycobacterium tuberculosis 6C sRNA binds multiple mRNA targets via C-rich loops independent of RNA chaperones. Nucleic Acids Res. 2019; 47:4292–4307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Raad N., Luidalepp H., Fasnacht M., Polacek N.. Transcriptome-wide analysis of stationary phase small ncRNAs in E. coli. Int. J. Mol. Sci. 2021; 22:1703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Kawano M., Reynolds A.A., Miranda-Rios J., Storz G.. Detection of 5′- and 3′-UTR-derived small RNAs and cis-encoded antisense RNAs in escherichia coli. Nucleic Acids Res. 2005; 33:1040–1050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Ghosal A., Upadhyaya B.B., Fritz J.V., Heintz-Buschart A., Desai M.S., Yusuf D., Huang D., Baumuratov A., Wang K., Galas D.et al.. The extracellular RNA complement of Escherichia coli. Microbiol Open. 2015; 4:252–266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Pichon C., Merle du, Caliot L., Trieu-Cuot M.E., Le Bouguénec C. An in silico model for identification of small RNAs in whole bacterial genomes: characterization of antisense RNAs in pathogenic Escherichia coli and Streptococcus agalactiae strains. Nucleic Acids Res. 2012; 40:2846–2861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. McDowall K.J., Hernandez R.G., Lin-Chao S., Cohen S.N.. The ams-1 and rne-3071 temperature-sensitive mutations in the ams gene are in close proximity to each other and cause substitutions within a domain that resembles a product of the Escherichia coli mre locus. J. Bacteriol. 1993; 175:4245–4249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Hoiseth S.K., Stocker B.A.. Aromatic-dependent Salmonella typhimurium are non-virulent and effective as live vaccines. Nature. 1981; 291:238–239. [DOI] [PubMed] [Google Scholar]
- 29. Suar M., Jantsch J., Hapfelmeier S., Kremer M., Stallmach T., Barrow P.A., Hardt W.-D.. Virulence of Broad- and Narrow-Host-Range salmonella enterica serovars in the Streptomycin-pretreatedmouse model. Infect. Immun. 2006; 74:632–644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Fields P.I., Swanson R.V., Haidaris C.G., Heffron F.. Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Nat. Acad. Sci. U.S.A. 1986; 83:5189–5193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Dieye Y., Ameiss K., Mellata M., Curtiss R.. The salmonella pathogenicity island (SPI) 1 contributes more than SPI2 to the colonization of the chicken by Salmonella enterica serovar typhimurium. BMC Microbiol. 2009; 9:3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Chao Y., Li L., Girodat D., Förstner K.U., Said N., Corcoran C., Śmiga M., Papenfort K., Reinhardt R., Wieden H.-J.et al.. In vivo cleavage map illuminates the central role of RNase e in coding and non-coding RNA pathways. Mol. Cell. 2017; 65:39–51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Lee T.S., Krupa R.A., Zhang F., Hajimorad M., Holtz W.J., Prasad N., Lee S.K., Keasling J.D.. BglBrick vectors and datasheets: a synthetic biology platform for gene expression. J. Biol. Eng. 2011; 5:12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Miyazaki K. MEGAWHOP cloning: a method of creating random mutagenesis libraries via megaprimer PCR of whole plasmids. Methods Enzymol. 2011; 498:399–406. [DOI] [PubMed] [Google Scholar]
- 35. Qi D., Scholthof K.B.. A one-step PCR-based method for rapid and efficient site-directed fragment deletion, insertion, and substitution mutagenesis. J. Virol. Methods. 2008; 149:85–90. [DOI] [PubMed] [Google Scholar]
- 36. Klock H.E., Lesley S.A.. The polymerase incomplete primer extension (PIPE) method applied to high-throughput cloning and site-directed mutagenesis. Methods Mol. Biol. 2009; 498:91–103. [DOI] [PubMed] [Google Scholar]
- 37. Datsenko K.A., Wanner B.L.. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. U.S.A. 2000; 97:6640–6645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Cherepanov P.P., Wackernagel W.. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene. 1995; 158:9–14. [DOI] [PubMed] [Google Scholar]
- 39. Luidalepp H., Berger S., Joss O., Tenson T., Polacek N.. Ribosome shut-down by 16S rRNA fragmentation in stationary-phase Escherichia coli. J. Mol. Biol. 2016; 428:2237–2247. [DOI] [PubMed] [Google Scholar]
- 40. Gebetsberger J., Zywicki M., Künzi A., Polacek N.. tRNA-derived fragments target the ribosome and function as regulatory non-coding RNA in haloferax volcanii. Archaea (Vancouver, B.C.). 2012; 2012:260909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Erlacher M.D., Chirkova A., Voegele P., Polacek N.. Generation of chemically engineered ribosomes for atomic mutagenesis studies on protein biosynthesis. Nat. Protoc. 2011; 6:580–592. [DOI] [PubMed] [Google Scholar]
- 42. Updegrove T.B., Kouse A.B., Bandyra K.J., Storz G.. Stem-loops direct precise processing of 3′ UTR-derived small RNA micL. Nucleic Acids Res. 2018; 47:1482–1492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Jiang X., Belasco J.G.. Catalytic activation of multimeric RNase e and RNase g by 5′-monophosphorylated RNA. Proc. Natl. Acad. Sci. U.S.A. 2004; 101:9211–9216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Yakhnin A.V., Yakhnin H., Babitzke P.. Gel mobility shift assays to detect protein-RNA interactions. Methods Mol. Biol. 2012; 905:201–211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Rao X., Huang X., Zhou Z., Lin X.. An improvement of the 2ˆ(-delta delta CT) method for quantitative real-time polymerase chain reaction data analysis. Biostat. Bioinform. Biomath. 2013; 3:71–85. [PMC free article] [PubMed] [Google Scholar]
- 46. Merritt J.H., Kadouri D.E., O’Toole G.A. Growing and analyzing static biofilms. Curr. Protoc. Microbiol. 2005; Chapter 1:Unit 1B.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Winzer P., Müller J., Imhof D., Ritler D., Uldry A.-C., Braga-Lagache S., Heller M., Ojo K.K., Van Voorhis W.C., Ortega-Mora L.-M.et al.. Neospora caninum: differential proteome of multinucleated complexes induced by the bumped kinase inhibitor BKI-1294. Microorganisms. 2020; 8:801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Pfister S.P., Schären O.P., Beldi L., Printz A., Notter M.D., Mukherjee M., Li H., Limenitakis J.P., Werren J.P., Tandon D.et al.. Uncoupling of invasive bacterial mucosal immunogenicity from pathogenicity. Nat. Commun. 2020; 11:1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Wright P.R., Georg J., Mann M., Sorescu D.A., Richter A.S., Lott S., Kleinkauf R., Hess W.R., Backofen R.. CopraRNA and intarna: predicting small RNA targets, networks and interaction domains. Nucleic Acids Res. 2014; 42:W119–W123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Zuker M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003; 31:3406–3415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Dove S.L., Smith S.G., Dorman C.J.. Control of Escherichia coli type 1 fimbrial gene expression in stationary phase: a negative role for rpoS. Mol. Gen. Genet. 1997; 254:13–20. [DOI] [PubMed] [Google Scholar]
- 52. Moor J., Aebi S., Rickli S., Mostacci N., Overesch G., Oppliger A., Hilty M.. Dynamics of extended-spectrum cephalosporin-resistant Escherichia coli in pig farms: a longitudinal study. Int. J. Antimicrob. Agents. 2021; 58:106382. [DOI] [PubMed] [Google Scholar]
- 53. Hengge-Aronis R. Signal transduction and regulatory mechanisms involved in control of the sigma(S) (RpoS) subunit of RNA polymerase. Microbiol. Mol. Biol. Rev. 2002; 66:373–395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54. Thomason M.K., Bischler T., Eisenbart S.K., Förstner K.U., Zhang A., Herbig A., Nieselt K., Sharma C.M., Storz G.. Global transcriptional start site mapping using differential RNA sequencing reveals novel antisense RNAs in Escherichia coli. J. Bacteriol. 2015; 197:18–28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Dasgupta S., Fernandez L., Kameyama L., Inada T., Nakamura Y., Pappas A., Court D.L.. Genetic uncoupling of the dsRNA-binding and RNA cleavage activities of the Escherichia coli endoribonuclease RNase III–the effect of dsRNA binding on gene expression. Mol. Microbiol. 1998; 28:629–640. [DOI] [PubMed] [Google Scholar]
- 56. Callaghan A.J., Marcaida M.J., Stead J.A., McDowall K.J., Scott W.G., Luisi B.F.. Structure of Escherichia coli RNase e catalytic domain and implications for RNA turnover. Nature. 2005; 437:1187–1191. [DOI] [PubMed] [Google Scholar]
- 57. Miyakoshi M., Chao Y., Vogel J.. Regulatory small RNAs from the 3′ regions of bacterial mRNAs. Curr. Opin. Microbiol. 2015; 24:132–139. [DOI] [PubMed] [Google Scholar]
- 58. Adams P.P., Storz G.. Prevalence of small base-pairing RNAs derived from diverse genomic loci. Biochim. Biophys. Acta. 2020; 1863:194524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Serra D.O., Richter A.M., Klauck G., Mika F., Hengge R., Kolter R.. Microanatomy at cellular resolution and spatial order of physiological differentiation in a bacterial biofilm. Mbio. 2013; 4:e00103-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Boehm A., Steiner S., Zaehringer F., Casanova A., Hamburger F., Ritz D., Keck W., Ackermann M., Schirmer T., Jenal U.. Second messenger signalling governs Escherichia coli biofilm induction upon ribosomal stress. Mol. Microbiol. 2009; 72:1500–1516. [DOI] [PubMed] [Google Scholar]
- 61. Minamino T., Namba K.. Self-assembly and type III protein export of the bacterial flagellum. Microb. Physiol. 2004; 7:5–17. [DOI] [PubMed] [Google Scholar]
- 62. Fitzgerald D.M., Bonocora R.P., Wade J.T.. Comprehensive mapping of the Escherichia coli flagellar regulatory network. PLoS Genet. 2014; 10:e1004649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Palchevskiy V., Finkel S.E.. Escherichia coli competence gene homologs are essential for competitive fitness and the use of DNA as a nutrient. J. Bacteriol. 2006; 188:3902–3910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Melamed S., Adams P.P., Zhang A., Zhang H., Storz G.. RNA-RNA interactomes of ProQ and hfq reveal overlapping and competing roles. Mol. Cell. 2020; 77:411–425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Pourciau C., Lai Y.-J., Gorelik M., Babitzke P., Romeo T.. Diverse mechanisms and circuitry for global regulation by the RNA-Binding protein csrA. Front. Microbiol. 2020; 11:601352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Potts A.H., Vakulskas C.A., Pannuri A., Yakhnin H., Babitzke P., Romeo T.. Global role of the bacterial post-transcriptional regulator CsrA revealed by integrated transcriptomics. Nat. Commun. 2017; 8:1596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Liu M.Y., Gui G., Wei B., Preston J.F., Oakford L., Yüksel Ü., Giedroc D.P., Romeo T. The RNA molecule CsrB binds to the global regulatory protein CsrA and antagonizes its activity in Escherichia coli. J. Biol. Chem. 1997; 272:17502–17510. [DOI] [PubMed] [Google Scholar]
- 68. Weilbacher T., Suzuki K., Dubey A.K., Wang X., Gudapaty S., Morozov I., Baker C.S., Georgellis D., Babitzke P., Romeo T.. A novel sRNA component of the carbon storage regulatory system of Escherichia coli. Mol. Microbiol. 2003; 48:657–670. [DOI] [PubMed] [Google Scholar]
- 69. Suzuki K., Babitzke P., Kushner S.R., Romeo T., RNase E.. Identification of a novel regulatory protein (CsrD) that targets the global regulatory RNAs CsrB and CsrC for degradation by. Genes Dev. 2006; 20:2605–2617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Jørgensen M.G., Thomason M.K., Havelund J., Valentin-Hansen P., Storz G.. Dual function of the McaS small RNA in controlling biofilm formation. Genes Dev. 2013; 27:1132–1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Mitra A., Palaniyandi S., Herren C.D., Zhu X., Mukhopadhyay S.. Pleiotropic roles of uvrY on biofilm formation, motility and virulence in uropathogenic Escherichia coli CFT073. PLoS One. 2013; 8:e55492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Figueroa-Bossi N., Schwartz A., Guillemardet B., D’Heygère F., Bossi L., Boudvillain M.. RNA remodeling by bacterial global regulator CsrA promotes Rho-dependent transcription termination. Genes Dev. 2014; 28:1239–1251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Yakhnin A.V., Baker C.S., Vakulskas C.A., Yakhnin H., Berezin I., Romeo T., Babitzke P.. CsrA activates flhDC expression by protecting flhDC mRNA from RNase E-mediated cleavage. Mol. Microbiol. 2013; 87:851–866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Jackson D.W., Suzuki K., Oakford L., Simecka J.W., Hart M.E., Romeo T.. Biofilm formation and dispersal under the influence of the global regulator CsrA of Escherichia coli. J. Bacteriol. 2002; 184:290–301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Hansmeier N., Miskiewicz K., Elpers L., Liss V., Hensel M., Sterzenbach T.. Functional expression of the entire adhesiome of Salmonella enterica serotype typhimurium. Sci. Rep. 2017; 7:10326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Tucker S.C., Galán J.E.. Complex function for SicA, a Salmonella enterica serovar typhimurium type III secretion-associated chaperone. J. Bacteriol. 2000; 182:2262–2268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Sturm A., Heinemann M., Arnoldini M., Benecke A., Ackermann M., Benz M., Dormann J., Hardt W.-D.. The cost of virulence: retarded growth of Salmonella typhimurium cells expressing type III secretion system 1. PLoS Pathog. 2011; 7:e1002143. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78. Ponath F., Hör J., Vogel J.. An overview of gene regulation in bacteria by small RNAs derived from mRNA 3' ends. FEMS Microbiol Rev. 2022; 46:fuac017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Jonas K., Melefors Ö.. The Escherichia coli CsrB and CsrC small RNAs are strongly induced during growth in nutrient-poor medium. FEMS Microbiol. Lett. 2009; 297:80–86. [DOI] [PubMed] [Google Scholar]
- 80. Hör J., Di Giorgio S., Gerovac M., Venturini E., Förstner KonradU., Vogel J.. Grad-seq shines light on unrecognized RNA and protein complexes in the model bacterium Escherichia coli. Nucleic Acids Res. 2020; 48:9301–9319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Navarro Llorens J.M., Tormo A., Martínez-García E.. Stationary phase in gram-negative bacteria. FEMS Microbiol. Rev. 2010; 34:476–495. [DOI] [PubMed] [Google Scholar]
- 82. Josenhans C., Suerbaum S.. The role of motility as a virulence factor in bacteria. Int. J. Med. Microbiol. 2002; 291:605–614. [DOI] [PubMed] [Google Scholar]
- 83. Hung C.-C., Eade C.R., Betteken M.I., Pavinski Bitar P.D., Handley E.M., Nugent S.L., Chowdhury R., Altier C.. Salmonella invasion is controlled through the secondary structure of the hilD transcript. PLoS Pathog. 2019; 15:e1007700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Sterzenbach T., Nguyen K.T., Nuccio S.P., Winter M.G., Vakulskas C.A., Clegg S., Romeo T., Bäumler A.J.. A novel CsrA titration mechanism regulates fimbrial gene expression in Salmonella typhimurium. EMBO J. 2013; 32:2872–2883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Gutiérrez P., Li Y., Osborne M.J., Pomerantseva E., Liu Q., Gehring K.. Solution structure of the carbon storage regulator protein CsrA from Escherichia coli. J. Bacteriol. 2005; 187:3496–3501. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All relevant data are within the paper and its supplementary data.