Abstract
A fibronectin (Fn)-binding adhesin of Staphylococcus aureus contains three tandem 37- or 38-amino-acid motifs (D1, D2, and D3), which function to bind Fn. Plasma from patients with S. aureus infections contain antibodies that preferentially recognize ligand induced binding sites in the D motifs and do not inhibit Fn binding (F. Casolini, L. Visai, D. Joh, P. G. Conaldi, A. Toniolo, M. Höök, and P. Speziale, Infect. Immun. 66:5433–5442, 1998). To eliminate the influence of Fn binding on antibody development, we used synthetic peptide immunogens D121–34 and D320–33, which each contain a conserved pattern of amino acids that is essential for Fn binding but which cannot bind Fn without N- or C-terminal extensions. The D320–33 immunogen promoted the production of polyclonal antibodies that were 10-fold more effective as inhibitors of Fn-binding to the D3 motif than antibodies obtained by immunizing with an extended peptide D316–36, which exhibits functional Fn binding. The D320–33 immunogen also facilitated the production of a monoclonal antibody, 9C3, which was highly specific for the epitope SVDFEED, and abolished Fn binding by the D3 motif. When mixed with polyclonal anti-D121–34 immunoglobulin G, 70% inhibition of Fn binding to the three tandem D motifs was achieved compared to no more than 30% inhibition with either antibody preparation alone. Therefore, by immunizing with short synthetic peptides that are unable to bind Fn, we have effectively stimulated the production of antibodies specific for epitopes comprised of amino acids that are essential for Fn binding. Although these epitopes occur within a conserved pattern of amino acids that is required for Fn binding, the antibodies recognized specific linear epitope sequences and not a conserved structure common to all repeated motifs.
Staphylococcus aureus is a persistent cause of infectious morbidity and mortality in community and hospital settings and is a leading cause of bacteremia and tissue infections in hospitalized patients (2). Efforts to prevent infection by stimulating protective immunity were initiated when S. aureus was first recognized as a significant human pathogen, but these efforts have generally not succeeded in identifying antigens that stimulate protective immunity (22). The success of antibiotics in the treatment of infection tended to discourage continued endeavors in this direction. However, the emergence of multiple-drug-resistant strains of methicillin-resistant S. aureus (MRSA) and recent reports of vancomycin intermediate-resistant strains (34) have promoted renewed interest in vaccine development (6, 22). Stimulation of antibodies specific for capsular polysaccharide antigens is one possibility (7, 23). Other targets include the RNA III-activating protein, which is involved in the regulation of S. aureus virulence factors (3) and the in vivo-expressed bacterial surface polysaccharide, N-succinyl β-1-6 glucosamine (28). The ideal vaccine would also induce antibodies able to prevent bacterial adherence. In this respect, our efforts have focused on interfering with the interaction between S. aureus and fibronectin (Fn) (36).
S. aureus cells express a cell surface Fn-binding protein (FnBP) belonging to a family of microbial adhesins known as MSCRAMMs, which mediate adherence to the tissue extracellular matrix (ECM) (32, 33). Fn-binding MSCRAMMs of S. aureus and group A, C, and G Streptococcus species promote the colonization of implanted medical devices and traumatized heart valves (9, 20, 39, 40) and adherence to epithelial cell surfaces, skin fibroblasts, cutaneous tissues, and exposed ECM proteins (17, 29–31, 37). Although there is little relatedness in terms of their primary amino acid sequences, these adhesins share a common mechanism of ligand binding. The most well-characterized mechanism involves repeated motifs of 37 to 42 amino acids in length, which bind a 29-kDa N-terminal fragment of Fn (16, 25). A second, less-well-characterized binding domain consists of a nonrepetitive element and is located N-terminal to the repeated motifs (14, 15, 31). This “upstream” domain has been reported to bind only intact Fn (15) or a larger N-terminal Fn fragment that also contains a collagen binding domain (31). With S. pyogenes, both domains are required for optimal adhesion to ECM (31). Therefore, antibodies specific for both functional domains will be required to effectively interfere with adhesion of S. aureus to Fn.
Our efforts have focused on developing antibodies specific for the repetitive binding domain (36). In S. aureus FnBP, the tandem Fn-binding repeats are designated D1, D2, and D3. Synthetic peptides corresponding to the C-terminal 20 to 21 amino acids of each motif are independently capable of binding Fn, and the D3 motif binds Fn with 5- to 10-fold-greater affinity than either D1 or D2 (11, 26). Plasma from patients diagnosed with S. aureus infections contain antibodies that preferentially recognize epitopes in the C-terminal 20 amino acids of the D3 motif. However, the antibodies only recognized the adhesin when it was complexed with Fn and did not inhibit Fn binding (5). Therefore, although the ligand-binding sequences are immunogenic during in vivo infections, their interaction with Fn appears to promote the formation of antibodies specific for ligand induced binding sites (LIBS), which have been known to stimulate Fn binding (35). While this represents a potential limitation to their use as vaccine components, the ligand-binding repeated motifs also offer a potential benefit of being able to elicit the formation of cross-reactive antibodies, which recognize a conserved sequence motif that is required for Fn binding. Specifically, the C-terminal 20 amino acids of each of the D motifs from S. aureus FnBP possess a pattern of amino acids defined by GG(X3,4)(I/V)DF, which is required for Fn binding (25) and is duplicated with minor variations in the Fn-binding repeated motifs from MSCRAMMs of group A, C, and G Streptococcus species (16, 17, 25).
The purpose of the present study was to develop antibodies specific for these conserved sequences and to evaluate their ability to inhibit Fn binding and to recognize the same conserved sequence motifs in other Fn-binding repeat domains. To achieve this objective, we employed a synthetic peptide strategy to direct the humoral immune response toward epitopes that contain critical ligand-binding sequences. The synthetic peptide immunogens D121–34, D320–33, and D316–36 each contain the conserved GG(X3)(I/V)DF pattern of amino acids that is essential for Fn binding (25). However, the former two peptides cannot bind Fn without additional N- or C-terminal extensions, while D316–36 is an extended version of D320–33 which exhibits functional Fn binding (26). We find that the ability of a synthetic peptide immunogen to bind Fn can influence both the epitope specificity and inhibition activity of the resulting polyclonal antibody preparations. Furthermore, the D320–33 immunogen which cannot bind Fn stimulated the production of monoclonal antibodies (MAbs) specific for a sequence in the D3 motif (SVDFEED) that is essential for Fn binding. The combined anti-D121–34 and D320–33 antibodies inhibited the binding of soluble Fn to the GSTD1-3 domain. However, the individual antibodies were highly specific for the repeated motif from which the respective synthetic peptide immunogens were derived, and there was no significant cross-reactivity toward repeated motifs from other Fn-binding MSCRAMM adhesins. Therefore, the antibodies are highly specific for linear epitopes that occur within a conserved pattern of amino acids that is essential for Fn binding.
MATERIALS AND METHODS
Bacterial and keratinocyte cell cultures.
S. aureus 34, chosen for its high level of Fn binding and undetectable expression of protein A on Western blotting, is a sporadic MRSA isolate from the Sunnybrook and Women's College Health Science Centre. It was cultured for binding assays in brain heart infusion broth (Difco, Detroit, Mich.) as previously described (36). Escherichia coli TB1 was used as host for recombinant plasmids and cultured in Luria-Bertani broth (Gibco, Gaithersburg, Md.) containing 50 μg of ampicillin ml−1 when required. Media were supplemented with 15 g of agar liter−1 where appropriate for culturing on agar plates.
Normal human keratinocytes were obtained from neonatal foreskin as described earlier (18) and maintained in serum-free keratinocyte growth medium supplemented with bovine pituitary extract and recombinant epidermal growth factor (Gibco BRL, Burlington, Ontario, Canada). For subculturing, keratinocyte monolayers were treated with 0.05% trypsin–0.5 mM EDTA, and the trypsin was neutralized with 10% fetal bovine serum in phosphate-buffered saline (PBS). After the cell suspension was adjusted to a density of 2 × 105/ml, aliquots of 100 μl were transferred into individual wells of 96-well flat-bottom cell culture plates (Sarstedt, Newton, N.C.). When the cells had achieved confluent growth, ECM was exposed by treating the monolayers with 0.5% Triton X-100 for 30 min at 37°C, followed by treatment with 25 mM ammonium hydroxide for 10 min at 37°C as described elsewhere (19, 27).
Construction, expression, and purification of recombinant fusion proteins.
Purified rFNBD-P polypeptide consisting of the tandem P motifs of the S. pyogenes SfbI Fn-binding MSCRAMM, with an N-terminal polyhistidine tag (16), was generously provided by Magnus Höök. Plasmid pGEXD1-3 has been described previously (11) and directs the expression in E. coli of a glutathione S-transferase (GST) fusion protein (GSTD1-3) possessing the tandem Fn-binding D1, D2, and D3 motifs of the S. aureus FnBPA MSCRAMM. Plasmids pSDF100 and pSDF200 encoding the FnBA and FnBB adhesins of Streptococcus dysgalactiae (24) were provided by Martin Lindberg. These plasmids were used as template in PCR with Ultra-Therm DNA polymerase (BioCan Scientific, Mississauga, Ontario, Canada) and the oligonucleotide primers defined in Table 1 to generate PCR amplicons encoding different combinations of Fn-binding repeated motifs. The amplicons were then cloned in vector pGEX2T as described previously (11) to create vectors pGEXD1-2, pGEXD2-3, pGEXA1-3, and pGEXB1-3, which direct the expression in E. coli of the respective GST fusion proteins. Plasmid pGEXD3 was constructed by reverse PCR of plasmid pGEXD1-3, using a forward primer that anneals to a sequence encoding the N terminus of the D3 motif and a reverse primer spanning the C terminus of the pGEX2T-encoded GST. For this purpose, PCR was conducted with the Expand Long Template PCR reagent system (Roche Diagnostics, Laval, Quebec, Canada). E. coli TB1 was used as the host for the recombinant plasmids and the expression of fusion proteins. Fusion proteins were purified by affinity chromatography on glutathione agarose. The amino acid composition of each of the recombinant proteins was used to determine molar extinction coefficients at 280 nm, using the ProtParam program on the ExPASy molecular biology server from the Swiss Institute of Bioinformatics (1, 8). Protein concentrations were then determined by measuring the absorbance at 280 nm.
TABLE 1.
Sequence of forward (F) and reverse (R) primers used to amplify DNA for cloning in pGEX2T vector
| Plasmid construct | Orientation | Primer sequence (5′→3′)a | Nucleotide (coordinates) |
|---|---|---|---|
| pGEXD1-2 | F | cccGGATCCGAAGGTGGCCAAAAT | S. aureus fnbA (2338–2358) |
| R | cccgaattctcaTCCGTGAATATGTGGCACACT | S. aureus fnbA (2580–2560) | |
| pGEXD2-3 | F | cccggatccGGTCAAAATAAAGGTAATCAGTCATTC | S. aureus fnbA (2464–2484) |
| R | cgggaattctcaGCCGCTTACTTTTGGAAGTGT | S. aureus fnbA (2697–2677) | |
| pGEXD3 | F | cccggatccCACGGATTCAATAAGCACACTG | S. aureus fnbA (2575–2596) |
| R | cccGGATCCACGCGGAACCAGATC | pGEX2T (935–915) | |
| pGEXA1-3 | F | CCTGGATCCGGAAATGAAACA | S. dysgalactiae fnbA (2957–2977) |
| R | GGGAATTCTCGGTGATTGTT | S. dysgalactiae fnbA (3362–3343) | |
| pGEXB1-3 | F | cccggatccGACAAGCCGATTACTGAG | S. dysgalactiae fnbB (2975–2992) |
| R | cccgaattcTAGTTGTGGTTTGTCTTCC | S. dysgalactiae fnbB (3349–3331) |
Lowercase nucleotides indicate nontemplate 5′ additions to the PCR primers for efficient cleavage by restriction endonucleases (pGEXD1-2, F; pGEXD3, R), for incorporation of BamHI restriction endonuclease recognition sequences (pGEXD2-3, F; pGEXD3, F; and pGEXB1-3, F), for incorporation of both EcoRI restriction endonuclease recognition sequence and translation stop codon (pGEXD1-2, R; pGEXD2-3, R), or for incorporation of EcoRI restriction endonuclease recognition sequence (pGEXB1-3 R). Underlined nucleotides represent primer-directed modification of the original nucleotide sequence, for conversion to BamHI (pGEXD1-2, F; pGEXA1-3, F), or EcoRI restriction endonuclease recognition sequences (pGEXA1-3, R).
Synthetic peptides.
Synthetic peptides D121–34 and D320–33 corresponding to amino acids 21 to 34 and 20 to 33 of the D1 and D3 motifs, respectively, and overlapping biotinylated decapeptides spanning the D3 motif have been described previously (36). Synthetic peptide D316–36, corresponding to amino acids 16 to 36 of the D3 motif (KPSYQFGGHNSVDFEEDTLPK), was synthesized with an N-terminal cysteine by the University of Toronto peptide synthesis core facility. The peptides were coupled via the N-terminal cysteine to maleimide-activated keyhole limpet hemocyanin (KLH) according to the protocol provided by the manufacturer (Pierce, Rockford, Ill.). For finer specificity of epitope mapping within the D320–33 sequence, a series of biotinylated peptides were synthesized by Chiron Mimitopes (Clayton, Victoria, Australia), corresponding to amino acids 23 to 33 of the D3 motif (GHNSVDFEEDT), with an alanine substitution at each position. The peptides were synthesized with an N-terminal biotin and a 4-amino-acid spacer sequence, SGSG, to facilitate their capture on microtitre plates coated with 5 μg of streptavidin ml−1.
Production and purification of antibodies.
The protocol for producing antisera against synthetic peptides coupled to maleimide-activated KLH using New Zealand White rabbits was described elsewhere (36). Peptide D320–33 coupled to KLH was also used to produce MAbs by using the University of Alabama hybridoma core facility. The procedure used BALB/c mice and the mouse myeloma line P3X63-Ag8.653. Hybridomas were screened 14 days postfusion by enzyme-linked immunosorbent assay (ELISA) for recognition of D320–33 coupled to bovine serum albumin (BSA), and positive hybridomas were rescreened for recognition of recombinant D1-3 peptide. A number of hybridomas that gave a strong ELISA response to both synthetic and recombinant antigens were cloned by limiting dilution, and hybridoma 9C3 was selected for ascites production. The antibody isotype was determined by using the reagents provided with the Bio-Rad (Hercules, Calif.) mouse hybridoma Isotyper Kit. Mouse monoclonal immunoglobulin G1 (IgG1) was purified from 9C3 ascites fluid by using the reagents provided with the Affi-Gel protein A MAPS kit (Bio-Rad), while polyclonal rabbit IgG was purified from sera by using UltraLink immobilized Protein A (Pierce, Rockford, Ill.).
Labeling of proteins and bacteria.
Human plasma Fn was obtained from Gibco (Gaithersburg, Md.). The 29-kDa N-terminal fragment of Fn (N29) was purified as described elsewhere (25). Where indicated, specific proteins (Fn, N29, or individual fusion proteins) or heat-killed exponential-phase cells of S. aureus 34 were labeled with biotinamidocaproate N-hydroxysuccinimide ester (Sigma, St. Louis, Mo.) as described previously (27).
ELISA protocols and inhibition assays.
ELISA was performed in Corning 96-well microtiter plates with wash buffer consisting of PBS containing 0.05% (vol/vol) Tween 20, a blocking solution consisting of 3% (wt/vol) BSA in PBS, and dilution buffer consisting of PBS supplemented with 0.05% Tween 20 and 0.1% BSA. Microtiter plates were coated overnight at 4°C with 100 μl of the indicated concentrations of antigens or ligands diluted in carbonate-bicarbonate buffer. After washing and blocking, wells were incubated with antibody, biotinylated fusion proteins, or biotin-labeled bacteria in 100 μl of dilution buffer for 60 min at room temperature. After an extensive washing, wells were incubated with 100-μl aliquots of alkaline phosphatase-conjugated secondary detection reagent consisting of either goat anti-mouse or anti-rabbit IgG (Jackson Immunoresearch, West Grove, Pa.) or streptavidin (Roche), diluted 5,000-fold in dilution buffer. After a 60-min incubation on an orbital mixer and extensive washing, the plates were developed for 60 min at 20°C with 1 mg of para-nitrophenyl phosphate (Sigma) per ml in 0.1 M diethanolamine buffer (pH 9.8). Absorbance was quantified on a Bio-Rad model 3550 MicroPlate reader equipped with a 405-nm filter. For testing the ability of antibodies to block Fn binding, wells coated with fusion proteins containing the Fn-binding motifs were preincubated for 60 min at room temperature with 50 μl of polyclonal or monoclonal IgG, diluted to the indicated concentrations, prior to the addition of biotin-labeled Fn. After an additional 60 min of incubation, the wells were washed extensively, and bound Fn was detected with alkaline phosphatase-conjugated streptavidin.
The method for quantifying the adhesion of S. aureus to ECM from human keratinocytes was an adaptation of the general ELISA protocol. Briefly, nonspecific binding sites were blocked by incubating ECM-coated wells with 3% BSA and then were washed and incubated with biotin-labeled S. aureus cells for 60 min. Wells were washed again, and bound bacteria was detected with alkaline phosphatase-conjugated streptavidin, as described above for ELISA. For testing the effect of antibodies on S. aureus binding, bacteria were preincubated for 1 h at room temperature with the combined polyclonal anti-D1 and monoclonal 9C3 IgG antibodies prior to incubation with ECM. Negative controls included preimmune polyclonal rabbit IgG and reagent-grade mouse IgG purchased from Sigma.
Western blotting techniques.
Purified fusion proteins were subjected to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) with 12% acrylamide resolving gels (21) and then either stained with Coomassie blue for protein detection or transferred to Immobilon-P membrane (Millipore, Bedford, Mass.) by using the Bio-Rad Trans-Blot apparatus and a standard transfer buffer (38). The immobilized fusion proteins were then probed either with monoclonal antibody specific for the D3 motif of S. aureus FnBP or with biotinylated N29 fragment of Fn as previously described (27). The secondary detection reagents consisted of the alkaline phosphatase conjugates as described for the general ELISA protocols, using nitroblue tetrazolium and 5-bromo-4-chloro-3-indolylphosphate substrates (Bio-Rad) according to the supplier's recommendations.
RESULTS
Antibody production and screening of hybridomas.
In an initial ELISA screen of hybridomas from mice immunized with the D320–33-KLH conjugate, eight exhibited a strong positive response towards both D320–33-BSA and recombinant D1-3 peptide. Additional screening was performed with a subset of overlapping decapeptides spanning the D3 motif (36). Seven hybridomas recognized the overlapping peptides GHNSVDFEED and NSVDFEEDTL, spanning amino acids 23 to 34 of the D3 motif. A single hybridoma showed weaker recognition of these same peptides and also the next consecutive peptide in the series, VDFEEDTLPK. However, it did not remain viable upon subculturing. On the basis of these initial observations, hybridoma 9C3 was selected for cloning by limiting dilution and ascites production. Antibody purified from ascites fluid was determined to be of the IgG1 subclass.
Antisera from rabbits immunized with KLH-coupled synthetic peptides D121–34, D320–33, and D316–36 yielded titers of between 3,000 and 5,000 when they were tested by ELISA using GSTD1-2 and GSTD1-3 as antigens. The titer values for purified IgG were 1 μg · ml−1 for anti-D121–31 when assayed with GSTD1-2 as antigen, while titers of 0.5 and 1.6 μg · ml−1 were obtained when anti-D316–36 and anti-D320–33, respectively, were assayed with GSTD1-3 antigen (data not shown). Therefore, each of the synthetic peptide immunogens produced similar titer values of polyclonal antisera and purified IgG.
Epitope specificity of anti-D320–33 and anti-D316–36 antibodies.
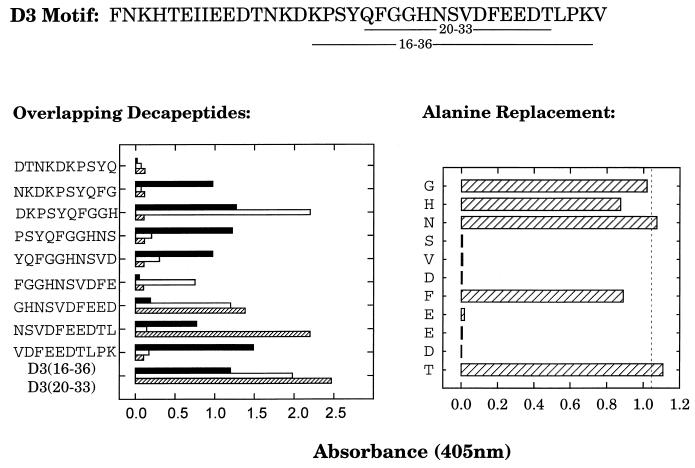
Synthetic peptide D320–33 contains a conserved pattern of amino acids that is critical for Fn-binding activity, but it cannot itself bind Fn without the N- or C-terminal extensions that are present on D316–36, which exhibits functional Fn binding (26). Polyclonal IgG obtained with these two immunogens exhibited different profiles of epitope specificity when assayed for recognition of a panel of overlapping decapeptides spanning the D3 motif (Fig 1, left panel). Comparatively, the anti-D320–33 IgG displayed a narrower range of epitope specificities, exhibiting the greatest response toward the sequence DKPSYQFGGH, and lesser recognition of two overlapping peptides spanning the sequence FGGHNSVDFEED. In contrast, the polyclonal D316–36 IgG exhibited equivalent recognition of several overlapping peptides but displayed poor recognition of the secondary epitope of polyclonal anti-D320–33 IgG (FGGHNSVDFEED). The MAb 9C3 IgG1 recognized two peptides that overlapped with the secondary epitope of the polyclonal D320–33 IgG, defined by the sequence GHNSVDFEEDTL. Additional mapping of the epitope specificity of MAb 9C3 was obtained by scanning alanine substitution of the sequence GHNSVDFEEDT (Fig. 1, right panel). Alanine substitution at Gly23, His24, Asn25, Phe29, or Thr33 resulted in little or no loss of recognition by MAb 9C3. In contrast, peptides with alanine substitutions at Ser26, Val27, Asp28, Glu30, Glu31, or Asp32 were not recognized by MAb 9C3. Therefore, the epitope for MAb 9C3 is localized to SVDFEED (D326–32), in which alanine substitution at any position other than the central phenylalanine results in loss of recognition.
FIG. 1.
Epitope mapping of anti-D316–36 and anti-D320–33 antibodies. Biotin-labeled synthetic peptides (2 μg · ml−1) were captured on streptavidin-coated microtiter plates, followed by incubation with 1 μg of anti-D316–36 or anti-D320–33 polyclonal IgG per ml or 50 ng of monoclonal anti-D320–33 IgG (MAb 9C3) per ml. Plates were developed as in the standard ELISA protocol. The sequence of the intact D3 motif, with the underlined amino acids representing the D316–36 and D320–33 sequences is displayed at the top of the figure. The left panel displays the comparison of reactivity between the polyclonal anti-D316–36 (■) and anti-D320–33 (□) IgG and the MAb 9C3 (▨), toward a series of overlapping decapeptides spanning amino acids 11 to 37 of the D3 motif. The right panel displays the reactivity of MAb 9C3 toward a series of peptides containing a single alanine substitution at each of the indicated amino acids within the sequence D323–33. The dashed line represents the response to D323–33 with no alanine substitutions. Each value represents the average of triplicate determinations.
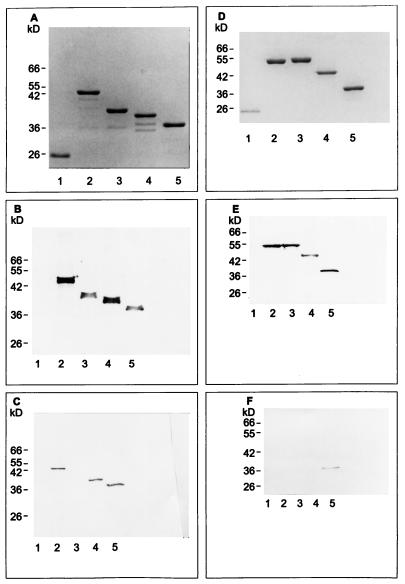
Within the D1 and D2 motifs, the epitope that is homologous to SVDFEED is represented by DIDFDSV. Within these sequences, the underlined amino acids indicate a conserved acidic amino acid flanked by hydrophobic residues, which is highly conserved in Fn-binding repeated motifs from adhesins of S. aureus, S. dysgalactiae, and S. pyogenes (16). However, consistent with the low tolerance of MAb 9C3 for amino acid substitution within the SVDFEED epitope, the antibody was unable to recognize a fusion protein that contained only the D1 and D2 motifs (Fig. 2C). Additional Western blotting (Fig. 2F) and ELISA experiments (data not shown) revealed that MAb 9C3 also did not recognize fusion proteins containing the Fn-binding A motifs (VIDFTED) or B motifs (IVDIEEN or VVDIEES) from the S. dysgalactiae FnBA and FnBB MSCRAMMs or the P-motifs (SVEFTKD) of S. pyogenes SfbI. Although each of these heptapeptides contain four amino acids (underlined) that match with the epitope for MAb 9C3, this homology was not sufficient to promote cross-reactivity. Therefore, MAb 9C3 was highly specific for the epitope SVDFEED. Additional ELISA experiments with polyclonal antibody specific for D320–33 again demonstrated a lack of cross-reactivity toward repeated motifs from other Fn-binding MSCRAMMs (data not shown). Therefore, this lack of cross-reactivity was observed with both polyclonal and MAb preparations.
FIG. 2.
Recombinant Fn-binding peptides separated by SDS-PAGE and stained with Coomassie blue (A and D) or transferred to Immobilon-P membrane, followed by detection with 1 μg of biotin-labeled N29 fragment of Fn (B and E) or MAb 9C3 (C and F) per ml. Lanes (A, B, and C): 1, GST protein; 2, GSTD1-3; 3, GST-D1-2; 4, GSTD2-3; 5, GSTD3. Lanes (D, E, and F): 1, GST protein; 2, GSTA1-3, and 3, GSTB1-3 (both from S. dysgalactiae); 4, rFNBD-P (from S. pyogenes); 5, GSTD3. Protein loading was 1 μg per lane in panels A, D, E, and F, except for lane 5 in panels E and F, which contained 25 and 100 ng, respectively. In panels B and C, 25 and 100 ng of protein per lane, respectively, was used.
Inhibition of Fn-binding by polyclonal antibodies and MAbs specific for individual D motifs.
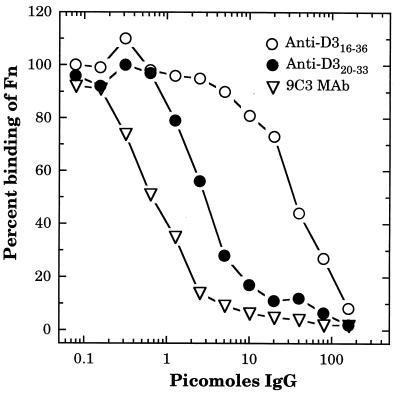
Of the three antibody preparations specific for the D3 motif, the MAb 9C3 IgG1 was the most effective inhibitor of Fn binding to the GSTD3 fusion protein, achieving 50% inhibition at less than 1 pmol of IgG1 (Fig. 3). Of the two polyclonal preparations, the anti-D320–33 IgG was 10-fold more effective as an inhibitor relative to anti-D316–36, with 50% inhibition occurring at approximately 3- and 30-pmol, respectively (Fig. 3). Therefore, although both preparations of polyclonal D3-specific antibody were capable of achieving 90% inhibition of Fn binding to the D3 motif, the antibodies obtained by immunizing with D320–33 were much more effective at low concentrations.
FIG. 3.
Inhibition of Fn binding to GSTD3 by anti-D316–36 and anti-D320–33 antibodies. GSTD3-coated microtiter plates were incubated for 60 min with 50 μl of various concentrations of anti-D316–36 or anti-D320–33 polyclonal IgG or anti-D320–33 monoclonal IgG (MAb 9C3), followed by the addition of 28 ng in 50 μl per well of biotinylated Fn. After incubation for an additional 60 min, bound Fn was detected with streptavidin-alkaline phosphatase conjugate. Absorbance values (A405) were determined after 60 min of development and then converted into percent values, with 100% representing the A405 value in the absence of added IgG. Each point represents the average from triplicate determinations.
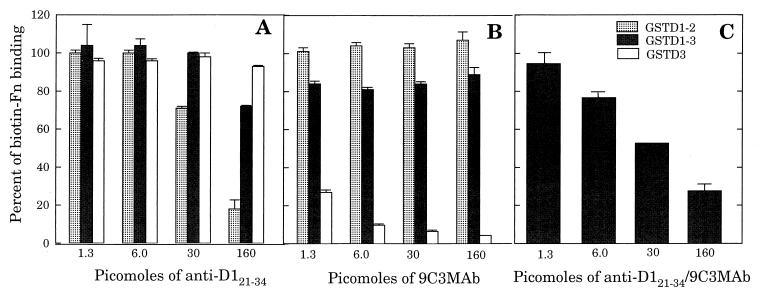
Within the D1 motif, the amino acid sequence 21QGGNIVDIDFDSVP34 is functionally equivalent to the region D320–33 and is highly homologous to the same region of D2, 21HGGNIIDIDFDSVP34. Consistent with these sequences representing functional Fn-binding sites, polyclonal anti-D121–34 IgG provided a maximum of 85% inhibition of Fn binding to fusion protein GSTD1-2, containing the tandem D1 and D2 motifs (Fig. 4A). These data establish that antibodies obtained by immunizing with D121–34 are able to recognize and inhibit binding of Fn to a nearly identical sequence in the D2 motif. However, the same antibody preparation exhibited less than 10% inhibition of Fn binding to GSTD3 and achieved only 30% inhibition of Fn binding to GSTD1-3. Similarly, although MAb 9C3 effectively inhibited Fn binding to GSTD3, it provided no more than 20% inhibition of Fn binding to GSTD1-3 and caused a slight enhancement of Fn binding to GSTD1-2 (Fig. 4B). When both anti-D121–31 polyclonal IgG and MAb 9C3 were combined, concentration-dependent inhibition of Fn binding to GSTD1-3 was observed, to a maximum of 70% inhibition with 160 pmol of total IgG (Fig. 4C). Therefore, greater inhibition could be achieved with the mixture of antibodies than could be obtained with either antibody preparation alone.
FIG. 4.
Effect of specific antibodies on the binding of biotinylated Fn to different Fn-binding GST fusion proteins. Wells of microtiter plates were coated with 500 ng of GSTD3, GSTD1-2, or GSTD1-3 and then incubated for 60 min with 50 μl of various concentrations of anti-D121–34 polyclonal IgG (A), anti-D32033 MAb 9C3 IgG (B), or combined anti-D121–34 polyclonal IgG and MAb 9C3 (C), followed by incubation for an additional 60 min with 28 ng (in 50 μl) of biotinylated Fn per well. Bound Fn was detected with streptavidin-alkaline phosphatase conjugate. Each point represents the average from triplicate determinations.
While these assays duplicate an in vivo situation where S. aureus cells may bind soluble plasma Fn, it is also desirable that antibodies should block adherence of bacteria to ECM and Fn deposited on biomaterials. These situations were duplicated by assessing the ability of anti-D121–34 and anti-D320–33 antibodies to inhibit the adhesion of biotinylated fusion proteins and biotinylated S. aureus cells to solid-phase Fn and to ECM. Although MAb 9C3 interfered with the binding of GSTD3 to Fn-coated microtitre plates (Fig. 5A), the polyclonal anti-D121–34 did not block attachment of GSTD1-2 to Fn, and the combination of both antibodies also did not inhibit binding of GSTD1-3 (Fig. 5A). To determine if this was an artifact due to an unnatural conformation of Fn absorbed on plastic, the assay was repeated with ECM synthesized by keratinocyte cell culture. The D motif-specific antibodies were also unable to block the binding of biotin-labeled S. aureus to ECM. Furthermore, an increase in bacterial attachment to both Fn and ECM is induced by preincubating S. aureus cells with combined anti-D121–34 and 9C3 IgG (Fig. 5B). When this assay was repeated with the individual antibodies, stimulation of adherence was observed with MAb 9C3 IgG but not with polyclonal D121–34 IgG or with normal mouse or preimmune rabbit IgG (data not shown). Therefore, the slight stimulation of adhesion is attributed to MAb 9C3.
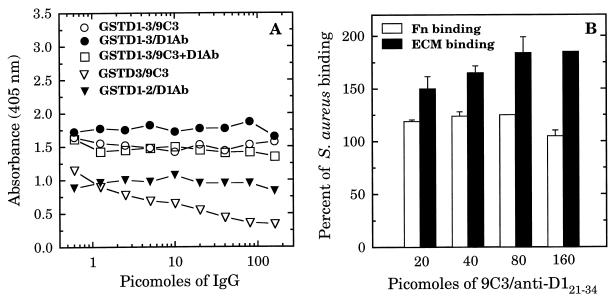
FIG. 5.
Effect of specific antibodies on binding of biotin-labeled GST D-motif fusion proteins to the wells of microtiter plates coated with 500 ng of Fn per well (A) and on the binding of biotin-labeled S. aureus MRSA strain 34 to the wells of microtiter plates coated with 500 ng per well of Fn or ECM deposited by keratinocyte cell culture (B). In panel A, 1 μg of biotin-labeled fusion proteins was preincubated for 60 min with the indicated amount of polyclonal D121–34 IgG, MAb 9C3 IgG, or both combined in a volume of 100 μl prior to addition to Fn-coated microtiter plates. In panel B, 107 biotinylated S. aureus cells were preincubated for 60 min with a mixture of D121–34 IgG and MAb 9C3 prior to addition to microtiter plates coated with Fn or keratinocyte ECM. Binding was detected with streptavidin-alkaline phosphatase conjugate. Each point represents the average of duplicate determinations.
DISCUSSION
In using the Fn-binding D motifs of S. aureus FnBP as an immunogen for the production of antibodies, it is necessary to consider the high affinity of the receptor-ligand interaction, the structure of the Fn-binding domain, and the certainty that immunogen will complex with Fn in vivo. Furthermore, the Fn-binding D motifs are primarily unstructured in solution but undergo a dramatic structural rearrangement upon binding of Fn (10). One consequence of this rearrangement is evident from the recent finding that blood plasma from patients diagnosed with S. aureus infections contained antibodies that preferentially recognized LIBS within amino acids 15 to 36 of the D3 motif (5). Although this same sequence contains the major Fn-binding determinant of the D3 motif (26), these anti-LIBS antibodies did not inhibit Fn binding (5). In the present study, our goal was to negate the deleterious consequence of this structural rearrangement by immunization with a synthetic peptide D320–33, which contains a core sequence of amino acids that are critical for Fn binding but cannot alone bind Fn without short N- or C-terminal extensions that are present on D316–36.
Our work demonstrates that both peptides are immunogenic and elicit a similar titer of antibodies specific for the D1-3 domain. However, antibodies generated using the peptide which binds Fn (D316–36) were 10-fold less effective as an inhibitor of Fn binding to the D3 motif. Comparatively, these antibodies also exhibited a broader spectrum of epitope specificity but were least reactive toward the sequence FGGHNSVDFEED, which contains a conserved pattern of amino acids that is essential for Fn binding (25). As the D316–36 peptide binds Fn with much lower affinity than the intact tandem D motifs (11), it is likely that this effect is much more pronounced when native Fn-binding proteins are used as immunogens, thereby favoring the development of antibodies specific for LIBS epitopes which have been reported previously (5, 35).
The ability of the D320–33 immunogen to stimulate the production of antibodies specific for a critical Fn-binding sequence was confirmed through the production of MAbs. MAb 9C3 completely blocked binding of Fn to the D3 motif and was highly specific for the amino acid sequence SVDFEED. This epitope contains four acidic amino acids, and replacement of any single acidic amino acid with alanine resulted in the loss of recognition by MAb 9C3. Chemical modification experiments have previously established that acidic amino acids are essential for binding of Fn by the D3 motif of S. aureus FnBPA, and the A2 motif of S. dysgalactiae FnBA (25, 26). In addition, each of the Fn-binding D and A motifs of S. aureus FnBPA and S. dysgalactiae FnBA share a conserved sequence, GG(X3,4)(I/V)DF. Amino acid substitutions at either the GG or IDF elements within the A2 motif of the S. dysgalactiae FnBA adhesin resulted in loss of Fn binding (25), and the epitope recognized by MAb 9C3 contains the (I/V)DF element of this conserved pattern. Therefore, our use of a synthetic peptide immunogen that could not bind Fn effectively stimulated the production of antibodies specific for an epitope that is comprised of amino acids which are essential for Fn binding.
Although the C-terminal 20 amino acids of each D motif share a conserved pattern of amino acids that is required for Fn binding, our data indicate that antibodies specific for these epitopes recognize a specific primary sequence and not a conserved structure that is common to all Fn-binding repeats. First, although D121–34 and D320–33 both contain conserved amino acids that are required for Fn binding, antibodies obtained with the respective immunogens were highly specific for the D motif from which the immunogen was derived. Since the D1 and D2 motifs are identical at 33 of 37 amino acids, antibodies specific for D121–34 achieved greater than 80% inhibition of Fn binding to the GSTD1-2 fusion protein. However, the D3 motif is quite divergent from D1 and D2, and this antibody preparation did not inhibit Fn binding to a fusion protein that contained only the D3 motif. MAb 9C3 recognized the epitope SVDFEED in the D3 motif but did not recognize the D1 or D2 motifs and also failed to recognize a closely related sequence SVEFTKD present in the repeated motifs of the Fn-binding adhesins SfbI and PrtF expressed by S. pyogenes. This same lack of cross-reactivity was also observed with polyclonal antibody specific for D320–33.
Surprisingly, an MAb specific for an anti-LIBS epitope in the FnBA adhesin of S. dysgalactiae was cross-reactive on Western blots with a related sequence in the Fn-binding Sfb adhesin from S. pyogenes, and stimulated binding of Fn by S. pyogenes cells (35). Therefore, anti-LIBS antibodies which stimulate Fn binding do not exhibit the same restricted specificity as that observed with both the polyclonal antibodies and MAbs described in the present study. This may be due to the finding that the repeated Fn-binding motifs are unstructured in solution but undergo a dramatic rearrangement upon contact with the N-terminal fragment of Fn (10). Consequently, anti-LIBS antibodies may recognize conformational epitopes, representing a specific structure that is acquired as a consequence of Fn binding. In contrast, our use of short synthetic peptide immunogens that are unable to bind Fn has promoted the formation of antibodies that recognize linear epitopes. Although these epitopes contain amino acids that are conserved in other Fn-binding adhesins, epitope recognition is highly dependent on the context of the surrounding amino acids, and cross-reactivity is less likely to occur.
In this respect, our results appear to differ from a recent report in which amino acids 1 to 30 of the D2 motif or the entire D2 motif were expressed on cowpea mosaic virus and potato virus, respectively, for use as immunogens (4). The resulting sera completely blocked binding of Fn to a GSTD1-3 fusion protein and also blocked adherence of S. aureus to solid-phase Fn. These findings imply that this immunization protocol has generated antibodies that are also cross-reactive with the D3 motif, as our work has demonstrated that failure to recognize just one of the three tandem D motifs renders an antibody preparation ineffective as an inhibitor of Fn binding. Although our data indicate that such cross-reactive antibodies are unlikely to occur, it is possible that expressing the D2 motif on a viral particle promotes a defined conformation that permits the production of antibodies specific for a conserved structural feature rather than a linear sequence of amino acids.
Although a mixture of polyclonal and monoclonal antibodies could block binding of soluble Fn to GSTD1-3, the same antibodies did not inhibit attachment of the fusion protein to immobilized Fn and exhibited slight stimulation of S. aureus adherence to ECM synthesized by a keratinocyte cell culture. These findings may reflect the presence of a less-well-characterized binding site in the FnBP adhesin that is upstream or N-terminal to the D motifs (12, 13, 15). These “upstream” binding sites have also been identified in Fn-binding MSCRAMMs of S. pyogenes and S. dysgalactiae (31, 35). In addition, the D motifs could also interact with solid phase Fn through a mechanism that differs from binding of soluble Fn. The N-terminal domain of Fn which is bound by the D motifs is comprised of five tandem finger-like structures known as type I modules. The C-terminal half of most MSCRAMM motifs bind to the type-I module pair 4 and 5 (11, 15). However, N-terminal sequences in some of the D motifs of S. aureus FnBPA exhibit secondary interactions with the type-1 module pairs 1-2 and 2-3 (15). These secondary interactions could be of greater significance in adhesion to ECM compared to the binding of soluble plasma Fn.
In view of these considerations, it is apparent that the receptor-mediated interaction of S. aureus with both soluble and ECM forms of Fn is a complex interaction, which involves multiple binding sites and specificities. The present work represents a significant advance in terms of directing the humoral immune response toward sequences that are critical to ligand binding, while still avoiding the deleterious effect of anti-LIBS antibodies. Although these antibodies are highly specific for a linear sequence of amino acids, they effectively inhibit the interaction of Fn with the sequence toward which they are targeted. Therefore, additional characterization of the FnBP adhesin is necessary to identify the full complement of sequences that are capable of interacting with both soluble Fn and fibrillar Fn incorporated as a component of the ECM.
ACKNOWLEDGMENTS
This work was supported by an operating grant (MT-12669) and Scholarship award to M.J.M. from the Medical Research Council of Canada and a Fellowship award to M.H. from the Sunnybrook and Women's College Health Sciences Centre Foundation.
REFERENCES
- 1.Appel R D, Bairoch A, Hochstrasser D F. A new generation of information retrieval tools for biologists: the example of the ExPASy WWW server. Trends Biochem Sci. 1994;19:258–260. doi: 10.1016/0968-0004(94)90153-8. [DOI] [PubMed] [Google Scholar]
- 2.Archer G L. Staphylococcus aureus: a well-armed pathogen. Clin Infect Dis. 1998;26:1179–1181. doi: 10.1086/520289. [DOI] [PubMed] [Google Scholar]
- 3.Balaban N, Goldkorn T, Nhan R T, Dang L B, Scott S, Ridgley R M, Rasooly A, Wright S C, Larrick J W, Rasooly R, Carlson J R. Autoinducer of virulence as a target for vaccine and therapy against Staphylococcus aureus. Science. 1998;280:438–440. doi: 10.1126/science.280.5362.438. [DOI] [PubMed] [Google Scholar]
- 4.Brennan F R, Jones T D, Longstaff M, Chapman S, Bellaby T, Smith H, Xu F, Hamilton W D, Flock J I. Immunogenicity of peptides derived from a fibronectin-binding protein of S. aureus expressed on two different plant viruses. Vaccine. 1999;17:1846–1857. doi: 10.1016/s0264-410x(98)00485-x. [DOI] [PubMed] [Google Scholar]
- 5.Casolini F, Visai L, Joh D, Conaldi P G, Toniolo A, Hook M, Speziale P. Antibody response to fibronectin-binding adhesin FnbpA in patients with Staphylococcus aureus infections. Infect Immun. 1998;66:5433–5442. doi: 10.1128/iai.66.11.5433-5442.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Fattom A I, Naso R. Staphylococcal vaccines: a realistic dream. Ann Med. 1996;28:43–46. doi: 10.3109/07853899608999073. [DOI] [PubMed] [Google Scholar]
- 7.Fattom A I, Sarwar J, Ortiz A, Naso R. A Staphylococcus aureus capsular polysaccharide (CP) vaccine and CP-specific antibodies protect mice against bacterial challenge. Infect Immun. 1996;64:1659–1665. doi: 10.1128/iai.64.5.1659-1665.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gill S C, von Hippel P H. Calculation of protein extinction coefficients from amino acid sequence data. Anal Biochem. 1989;182:319–326. doi: 10.1016/0003-2697(89)90602-7. [DOI] [PubMed] [Google Scholar]
- 9.Greene C, McDevitt D, Francois P, Vaudaux P E, Lew D P, Foster T J. Adhesion properties of mutants of Staphylococcus aureus defective in fibronectin-binding proteins and studies on the expression of fnb genes. Mol Microbiol. 1995;17:1143–1152. doi: 10.1111/j.1365-2958.1995.mmi_17061143.x. [DOI] [PubMed] [Google Scholar]
- 10.House-Pompeo K, Xu Y, Joh D, Speziale P, Hook M. Conformational changes in the fibronectin binding MSCRAMMs are induced by ligand binding. J Biol Chem. 1996;271:1379–1384. doi: 10.1074/jbc.271.3.1379. [DOI] [PubMed] [Google Scholar]
- 11.Huff S, Matsuka Y V, McGavin M J, Ingham K C. Interaction of N-terminal fragments of fibronectin with synthetic and recombinant D motifs from its binding protein on Staphylococcus aureus studied using fluorescence anisotropy. J Biol Chem. 1994;269:15563–15570. [PubMed] [Google Scholar]
- 12.Jacobsson K, Frykberg L. Cloning of ligand-binding domains of bacterial receptors by phage display. BioTechniques. 1995;18:878–885. [PubMed] [Google Scholar]
- 13.Jacobsson K, Frykberg L. Phage display shot-gun cloning of ligand-binding domains of prokaryotic receptors approaches 100% correct clones. BioTechniques. 1996;20:1070–1078. doi: 10.2144/96206rr04. [DOI] [PubMed] [Google Scholar]
- 14.Jaffe J, Natanson-Yaron S, Caparon M G, Hanski E. Protein F2, a novel fibronectin-binding protein from Streptococcus pyogenes, possesses two binding domains. Mol Microbiol. 1996;21:373–384. doi: 10.1046/j.1365-2958.1996.6331356.x. [DOI] [PubMed] [Google Scholar]
- 15.Joh D, Speziale P, Gurusiddappa S, Manor J, Hook M. Multiple specificities of the staphylococcal and streptococcal fibronectin-binding microbial surface components recognizing adhesive matrix molecules. Eur J Biochem. 1998;258:897–905. doi: 10.1046/j.1432-1327.1998.2580897.x. [DOI] [PubMed] [Google Scholar]
- 16.Joh H J, House-Pompeo K, Patti J M, Gurusiddappa S, Hook M. Fibronectin receptors from gram-positive bacteria: comparison of active sites. Biochemistry. 1994;33:6086–6092. doi: 10.1021/bi00186a007. [DOI] [PubMed] [Google Scholar]
- 17.Kline J B, Xu S, Bisno A L, Collins C M. Identification of a fibronectin-binding protein (GfbA) in pathogenic group G streptococci. Infect Immun. 1996;64:2122–2129. doi: 10.1128/iai.64.6.2122-2129.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kondo S, Kono T, Sauder D N, McKenzie R C. IL-8 gene expression and production in human keratinocytes and their modulation by UVB. J Investig Dermatol. 1993;101:690–694. doi: 10.1111/1523-1747.ep12371677. [DOI] [PubMed] [Google Scholar]
- 19.Kruk P A, Auersperg N. A line of rat ovarian surface epithelium provides a continuous source of complex extracellular matrix. In Vitro Cell Dev Biol Anim. 1994;30A:217–225. doi: 10.1007/BF02632043. [DOI] [PubMed] [Google Scholar]
- 20.Kuypers J M, Proctor R A. Reduced adherence to traumatized rat heart valves by a low-fibronectin-binding mutant of Staphylococcus aureus. Infect Immun. 1989;57:2306–2312. doi: 10.1128/iai.57.8.2306-2312.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Laemmli U K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 22.Lee J C. The prospects for developing a vaccine against Staphylococcus aureus. Trends Microbiol. 1996;4:162–166. doi: 10.1016/0966-842x(96)10021-4. [DOI] [PubMed] [Google Scholar]
- 23.Lee J C, Park J S, Shepherd S E, Carey V, Fattom A. Protective efficacy of antibodies to the Staphylococcus aureus type 5 capsular polysaccharide in a modified model of endocarditis in rats. Infect Immun. 1997;65:4146–4151. doi: 10.1128/iai.65.10.4146-4151.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lindgren P E, McGavin M J, Signas C, Guss B, Gurusiddappa S, Hook M, Lindberg M. Two different genes coding for fibronectin-binding proteins from Streptococcus dysgalactiae. The complete nucleotide sequences and characterization of the binding domains. Eur J Biochem. 1993;214:819–827. doi: 10.1111/j.1432-1033.1993.tb17985.x. [DOI] [PubMed] [Google Scholar]
- 25.McGavin M J, Gurusiddappa S, Lindgren P E, Lindberg M, Raucci G, Hook M. Fibronectin receptors from Streptococcus dysgalactiae and Staphylococcus aureus. Involvement of conserved residues in ligand binding. J Biol Chem. 1993;268:23946–23953. [PubMed] [Google Scholar]
- 26.McGavin M J, Raucci G, Gurusiddappa S, Hook M. Fibronectin binding determinants of the Staphylococcus aureus fibronectin receptor. J Biol Chem. 1991;266:8343–8347. [PubMed] [Google Scholar]
- 27.McGavin M J, Zahradka C, Rice K, Scott J E. Modification of the Staphylococcus aureus fibronectin binding phenotype by V8 protease. Infect Immun. 1997;65:2621–2628. doi: 10.1128/iai.65.7.2621-2628.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.McKenney D, Pouliot K L, Wang Y, Murthy V, Ulrich M, Doring G, Lee J C, Goldmann D A, Pier G B. Broadly protective vaccine for Staphylococcus aureus based on an in vivo-expressed antigen. Science. 1999;284:1523–1527. doi: 10.1126/science.284.5419.1523. [DOI] [PubMed] [Google Scholar]
- 29.Mempel M, Schmidt T, Weidinger S, Schnopp C, Foster T, Ring J, Abeck D. Role of Staphylococcus aureus surface-associated proteins in the attachment to cultured HaCaT keratinocytes in a new adhesion assay. J Investig Dermatol. 1998;111:452–456. doi: 10.1046/j.1523-1747.1998.00293.x. [DOI] [PubMed] [Google Scholar]
- 30.Okada N, Pentland A P, Falk P, Caparon M G. M protein and protein F act as important determinants of cell-specific tropism of Streptococcus pyogenes in skin tissue. J Clin Investig. 1994;94:965–977. doi: 10.1172/JCI117463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ozeri V, Tovi A, Burstein I, Natanson-Yaron S, Caparon M G, Yamada K M, Akiyama S K, Vlodavsky I, Hanski E. A two-domain mechanism for group A streptococcal adherence through protein F to the extracellular matrix. EMBO J. 1996;15:989–998. [PMC free article] [PubMed] [Google Scholar]
- 32.Patti J M, Allen B L, McGavin M J, Hook M. MSCRAMM-mediated adherence of microorganisms to host tissues. Annu Rev Microbiol. 1994;48:585–617. doi: 10.1146/annurev.mi.48.100194.003101. [DOI] [PubMed] [Google Scholar]
- 33.Patti J M, Hook M. Microbial adhesins recognizing extracellular matrix macromolecules. Curr Opin Cell Biol. 1994;6:752–758. doi: 10.1016/0955-0674(94)90104-x. [DOI] [PubMed] [Google Scholar]
- 34.Perl T M. The threat of vancomycin resistance. Am J Med. 1999;106:26S–37S. doi: 10.1016/s0002-9343(98)00354-4. [DOI] [PubMed] [Google Scholar]
- 35.Speziale P, Joh D, Visai L, Bozzini S, House-Pompeo K, Lindberg M, Hook M. A monoclonal antibody enhances ligand binding of fibronectin MSCRAMM (adhesin) from Streptococcus dysgalactiae. J Biol Chem. 1996;271:1371–1378. doi: 10.1074/jbc.271.3.1371. [DOI] [PubMed] [Google Scholar]
- 36.Sun Q, Smith G M, Zahradka C, McGavin M J. Identification of D motif epitopes in Staphylococcus aureus fibronectin-binding protein for the production of antibody inhibitors of fibronectin binding. Infect Immun. 1997;65:537–543. doi: 10.1128/iai.65.2.537-543.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Talay S R, Valentin-Weigand P, Jerlstrom P G, Timmis K N, Chhatwal G S. Fibronectin-binding protein of Streptococcus pyogenes: sequence of the binding domain involved in adherence of streptococci to epithelial cells. Infect Immun. 1992;60:3837–3844. doi: 10.1128/iai.60.9.3837-3844.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Towbin H, Staehelin T, Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci USA. 1979;76:4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Vaudaux P, Pittet D, Haeberli A, Lerch P G, Morgenthaler J J, Proctor R A, Waldvogel F A, Lew D P. Fibronectin is more active than fibrin or fibrinogen in promoting Staphylococcus aureus adherence to inserted intravascular catheters. J Infect Dis. 1993;167:633–641. doi: 10.1093/infdis/167.3.633. [DOI] [PubMed] [Google Scholar]
- 40.Vaudaux P E, Francois P, Proctor R A, McDevitt D, Foster T J, Albrecht R M, Lew D P, Wabers H, Cooper S L. Use of adhesion-defective mutants of Staphylococcus aureus to define the role of specific plasma proteins in promoting bacterial adhesion to canine arteriovenous shunts. Infect Immun. 1995;63:585–590. doi: 10.1128/iai.63.2.585-590.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]