Abstract
Upon iron restriction, Actinobacillus pleuropneumoniae has been shown to express the transferrin-binding proteins TbpB and TbpA, both of which have been implied to be important virulence factors. In order to identify additional iron-regulated proteins, we cloned and analyzed the region upstream of the transferrin-binding protein genes in an A. pleuropneumoniae serotype 7 strain. We located immediately upstream of the tbpB gene two open reading frames which were 43% homologous to the neisserial ExbBD protein genes. By raising specific antibodies, we showed that ExbB is expressed under iron-limiting growth conditions only, and RT-PCR analysis revealed that the exbBD genes and the tbpB gene are transcribed on a single polycistronic mRNA. By constructing an isogenic and nonpolar exbBD mutant, we showed that the exbBD genes are required by A. pleuropneumoniae for utilization of transferrin-bound iron. Using PCR and Western blotting, we showed that the genetic organization found in A. pleuropneumoniae serotype 7 is similar in all 12 A. pleuropneumoniae serotype reference strains.
Iron is an essential factor for the growth of bacterial pathogens, as it plays an irreplaceable role in most oxidation and reduction processes. In the environment, the vast majority of iron occurs in the ferric state, which has an extremely low solubility. To overcome this problem, bacteria are able to take up iron in a variety of chelated and therefore more soluble forms (18, 30, 36). This acquisition is mediated by an energy-coupled process involving essentially an outer membrane receptor, the TonB protein, and inner membrane protein complexes, such as ExbBD, serving as energy couplers (1, 6, 11, 20, 22).
Actinobacillus pleuropneumoniae is the etiologic agent of porcine pleuropneumonia, a highly infectious disease of fattening pigs occurring worldwide (12). A. pleuropneumoniae, like other members of the families Neisseriaceae and Pasteurellaceae, has developed for iron assimilation a highly sophisticated system which allows the specific utilization of the transferrin-bound iron of the respective hosts (8, 18, 33, 34). The outer membrane proteins involved in transferrin binding have been designated TbpB and TbpA; they are highly immunogenic antigens able to induce a protective immune response (2, 23, 31, 38). In addition, it has been shown for Neisseria gonorrhoeae that transferrin-binding proteins are required for experimental infection (9). The encoding genes (tbpB and tbpA) have been cloned and characterized for N. meningitidis (21), Haemophilus influenzae (17), A. pleuropneumoniae (14, 15, 16, 38), Pasteurella haemolytica (26), and Moraxella catarrhalis (23). Both genes appear to be located in one operon with putative promoter and regulatory regions preceding the tbpB gene (4, 16, 17, 21). The acquisition of protein-bound iron has been shown to be tonB dependent in N. meningitidis (35) and N. gonorrhoeae (5), and in both organisms, the tonB gene is located immediately upstream of a set of exbBD genes. The exbBD genes appear not to be linked to the tbpBA operon, and iron-regulated expression of the ExbBD proteins has not been observed in members of the families Pasteurellaceae and Neisseriaceae. A functional requirement of both genes for the uptake of transferrin-bound iron has been shown for N. gonorrhoeae (5), whereas for N. meningitidis the possibility of functional complementation by Tol proteins is being discussed (35). For members of the family Pasteurellaceae, no experimental evidence on the function of ExbBD proteins is available.
In the present communication, we show that in A. pleuropneumoniae a set of functional exbB and exbD genes is transcriptionally linked to the tbpB gene. By constructing an isogenic and nonpolar A. pleuropneumoniae serotype 7 exbBD deletion mutant, we show that the exbBD genes are essential for the utilization of transferrin-bound iron.
MATERIALS AND METHODS
Bacterial strains, plasmids, and primers.
The strains, plasmids, and primers used in this work are listed in Table 1.
TABLE 1.
Characteristics of bacterial strains, plasmids, and primers used in this study
| Strain, plasmid, or primer | Characteristics | Reference or source |
|---|---|---|
| Strains | ||
| E. coli DH5αF′ | F′/endA1 hsdR17 (rK− mK+) supE44 thi-1 recA1 gyrA (Nalr) relA1 Δ(lacZYA-argF)U169 deoR [φ80dlacΔ(lacZ)M15] | 29 |
| E. coli HB101 | F− Δ(gpt-proA)62 leuB6 supE44 ara-14 galK2 lacY1 Δ(mcrC-mrr) rpsL20 (Strr) xyl-5 mtl-1 recA13 | 32 |
| E. coli β2155 | thrB1004 pro thi strA hsdS lacZΔM15 (F′ lacZΔM15 lacIqtraD36 proA+proB+) ΔdapA::erm (Ermr) recA::RP4-2-tet (Tcr)::Mu-km (Kmr) λ pir | 10 |
| A. pleuropneumoniae AP76 | A. pleuropneumoniae serotype 7 strain kindly provided by the Western College of Veterinary Medicine, Saskatoon, Canada | 3 |
| A. pleuropneumoniae AP76Δexb | Unmarked ExbB-negative knockout mutant of A. pleuropneumoniae AP76 | This work |
| Plasmids | ||
| pBluescript SK | E. coli cloning vector carrying an ampicillin resistance determinant | Stratagenea |
| pGEX3x-5 | E. coli cloning vector carrying an ampicillin resistance determinant and devised to construct GST fusion proteins | Pharmacia |
| pGH432 | E. coli cloning vector carrying an ampicillin resistance determinant and a tac promoter | 14 |
| pJFF224 | E. coli-A. pleuropneumoniae shuttle vector | 13 |
| pBMK1 | Transconjugation vector based on pBluescript SK with mobRP4, polycloning site, Kmr, and transcriptional fusion of the omlA promoter with the sacB gene | 28 |
| pTF401 | A. pleuropneumoniae serotype 7 DNA from a λ2001 library subcloned into pGH432 | This work |
| pEXB1 | pGEX3x5 containing a BamHI-EcoRV fragment from a PCR product derived from pTF401 with primers BA9 and RE1 and expressing a GST-ExbB fusion protein | This work |
| pEXB10 | pBluescript SK carrying a BamHI-NsiI fragment from pTF401 | This work |
| pEXB10K | pBMK1 carrying the pEXB10 insert | This work |
| pEXB10Δexb | pEXB10 with a ScaI-PacI deletion abolishing the exbBD genes | This work |
| pEXB10ΔexbK | pBMK1 carrying the pEXB10Δexb insert | This work |
| pFOKE2 | pJFF224 carrying the BamHI fragment from pEXB10K | This work |
| pFOKE5 | pJFF224 carrying the BamHI fragment from pEXB10K controlled by the vector-derived T4 promoter | This work |
| Primers | ||
| BA6 | AGT GGG ATC CTG AAA GTT ACT ATT GGA G; primer (internal BamHI site) upstream of the exbB gene | This work |
| BA7 | CAA TGG ATC CAT TTT ATC TTC TTC AGG C; primer (internal BamHI site) upstream of the exbB gene | This work |
| BA9 | CAA GGG ATC CTA CTC TCG GTA TCT TTC T; primer (internal BamHI site) on 5′ end of the exbB gene | This work |
| BA23 | CAA CGG ATC CTG AAA GTC CTT GAA TTA G; primer (internal BamHI site) upstream of the tbpB gene | This work |
| RE1 | AAG TTT AAA ATG CAT ATT GC; primer overlapping the start codon of the tbpB gene | This work |
| TF6 | AGA ACT CAC CGG AAC T; primer ending at position 212 of the tbpB gene | This work |
| OF1f | CAA GAG ATC TTT CTT ATG CAT CTT ACT; primer overlapping the ATG codon of the rho gene | S. Thiedeb |
| OF1r | CAG AGT CCA TGG ATT TTT TAT GAA CGT TTC; primer overlapping the stop codon of the rho gene | S. Thiedeb |
Stratagene Europe, Amsterdam, The Netherlands.
EMBL accession no. Y17915.
Preparation of antisera and of porcine transferrin.
The serum raised against the A. pleuropneumoniae TbpB protein has been described previously (15). The serum directed against the ExbB protein was raised in rabbits by an initial intracutaneous injection and two subcutaneous boost injections of 100 μg of dissolved recombinant glutathione S-transferase (GST)–ExbB fusion protein in Emulsigen-Plus (MVP Inc., Ralston, Nebr.). Porcine transferrin was prepared by ammonium sulfate precipitation and subsequent column chromatography using DEAE–Sepharose CL-6B (Pharmacia, Freiburg, Germany) (24).
Media and growth conditions.
Escherichia coli strains were cultured in Luria-Bertani (LB) medium supplemented with the appropriate antibiotics (ampicillin, 100 μg/ml; kanamycin, 50 μg/ml); for cultivation of E. coli β2155 (ΔdapA), diaminopimelic acid (1 mM [Sigma Chemical Company, Deisenhofen, Germany]) was added. A. pleuropneumoniae strains were cultured in PPLO medium (Difco GmbH, Augsburg, Germany) supplemented with NAD (10 μg/ml [E. Merck AG, Darmstadt, Germany]), l-glutamine (100 μg/ml [Serva, Heidelberg, Germany]), l-cysteine hydrochloride (260 μg/ml [Sigma]), l-cystine dihydrochloride (10 μg/ml [Sigma]), dextrose (1 mg/ml), and Tween 80 (0.1%). Sucrose counterselection was performed as described previously (28). For the selection of A. pleuropneumoniae transconjugants, kanamycin (25 μg/ml) was added, and iron restriction was induced by the addition of 2,2-dipyridyl (Sigma) to a final concentration of 100 μM.
Plate bioassay testing of the utilization of transferrin-bound iron.
Brain heart infusion agar (Difco) was supplemented with 200 μM diethylenetriamine–penta-acetic acid calcium trisodium salt hydrate (Na3CaDTPA; Fluka Chemika and BioChemika, Deisenhofen, Germany) and NAD (10 μg/ml). A. pleuropneumoniae overnight broth cultures (supplemented PPLO medium) were diluted 1:10 in NaCl (150 mM), and 100 μl was evenly spread on a plate. Sterile filter paper disks were placed on the agar and loaded with 75 μl of porcine transferrin (500 μM) or ferric citrate (500 μM ferric nitrate, 1 mM sodium citrate), both in HEPES-NaCl-bicarbonate buffer (10 mM HEPES, 150 mM NaCl, 10 mM sodium bicarbonate [pH 7.4]); HEPES-NaCl-bicarbonate buffer alone was used as a negative control. Plates were incubated overnight at 37°C in a 5% CO2 atmosphere.
Manipulation of DNA.
DNA-modifying enzymes were purchased from New England Biolabs (Bad Schwalbach, Germany) and used according to the manufacturer's instructions. Taq polymerase was purchased from GIBCO-BRL Life Technologies (Karlsruhe, Germany). DNA for PCR and Southern blotting as well as plasmid DNA was prepared by standard protocols (32). Transformations, gel electrophoresis, PCR, and Southern blotting were done by standard procedures (32), and pulsed-field gel electrophoresis (PFGE) of A. pleuropneumoniae DNA was performed as described previously (27).
Cloning of the A. pleuropneumoniae tbpB upstream region.
A. pleuropneumoniae serotype 7 DNA was partially digested, and a λ2001 library was constructed. The library was screened using the tbpB gene (NsiI-KpnI-fragment) (15) as a probe, and a hybridizing clone was isolated. A BamHI-BglII-fragment was cloned into pGH432 cut with BamHI (resulting in plasmid pTF401), and a BamHI-EcoRV fragment was subcloned into M13mp18 and mp19 for DNA sequencing analysis.
Construction of recombinant plasmids.
To construct plasmid pEXB1 expressing a GST-ExbB fusion protein, a PCR fragment was obtained using primers BA9 and RE1 (Table 1 and Fig. 1). The fragment was restricted with BamHI and EcoRV and ligated into pGEX5x3 (Pharmacia) restricted with BamHI and SmaI. To construct plasmid pEXB10ΔexbK, used for transconjugation, several intermediate steps were required. First, the BamHI-NsiI fragment from pTF401 was cloned into pBluescript SK cut with BamHI and PstI, resulting in pEXB10. Plasmid pEXB10 was linearized with ScaI, completely digested with PacI, treated with E. coli DNA polymerase, and religated, resulting in pEXB10Δexb. The deletion obtained was characterized by nucleotide sequencing (Fig. 1). To construct pEXB10ΔexbK, the insert from pEXB10Δexb was removed with XbaI and SalI and ligated into the transconjugation vector pBMK1 (28) cut with XbaI and SalI. Plasmids pFOKE2 and pFOKE5 were constructed by ligating the XbaI-SalI fragment from pEXB10 into pBMK1. From here, a BamHI fragment was removed and ligated in either orientation into pJFF224 (13) with the vector-derived T4 promoter controlling exbBD transcription in pFOKE5.
FIG. 1.
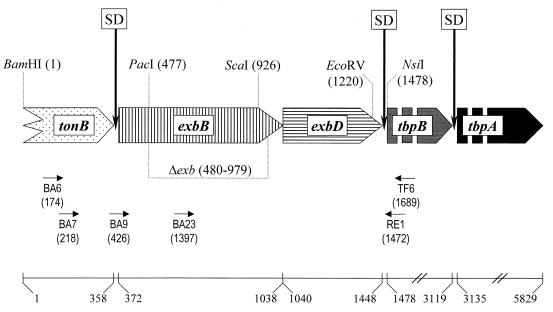
Organization of the A. pleuropneumoniae exbBD-tbpBA operon. The wide arrows indicate the relative locations, sizes, and directions of translation of the respective ORF. The zigzag line at the beginning of the tonB arrow indicates that the start of the ORF is not on the clones characterized to date. The gaps within the tbpB and tbpA arrows indicate that they are not drawn to scale. The vertical arrows marked with SD indicate putative Shine-Dalgarno sequences. The positions of restriction endonuclease sites used in this study are indicated by dotted lines, with the position of the first base (GenBank accession no. Y17916) given in parentheses. The dotted line underneath the exbB gene indicates the DNA region missing in the isogenic A. pleuropneumoniae mutant AP76Δexb. The small horizontal arrows indicate the relative positions of the primers used in the study; the number in parentheses indicates the position of the nucleotide at the 3′ end. The solid line on the bottom gives the scale in base pairs; the numbers indicate the position of the first base of the start and stop codons of the ORF with respect to the BamHI site at position 1.
Electroporation, transconjugation, and analysis of transconjugants and deletion mutants.
Electroporation and transconjugation were performed as described previously (28). Kanamycin-resistant colonies were analyzed by colony blotting using a 32P-dATP-labeled kanamycin resistance (Kmr) determinant. Counterselection to obtain unmarked deletion mutants was performed as previously described (28), and colonies were tested by PCR analysis using primers BA7 and RE1 (Table 1 and Fig. 1). Colonies with the correct PCR profile were confirmed by Southern blot analysis using the BamHI-EcoRV fragment of pEXB10 as a probe, by PFGE, by nucleotide sequence analysis, and by Western blotting.
Preparation of protein aggregates, electrophoresis, and Western blotting.
Protein aggregates were prepared as previously described (14). A. pleuropneumoniae whole-cell lysates and protein aggregates were analyzed by discontinuous sodium dodecyl sulfate-polyacrylamide gel electrophoresis and Western blotting as described earlier (15).
RNA isolation and RT-PCR.
RNA was prepared using the RNeasy Miniprep Kit (Qiagen, Hilden, Germany). RNA was treated for 30 min with RNase-free DNase (100 U; GIBCO-BRL) in the presence of RNase inhibitor (40 U; GIBCO-BRL), followed by phenol extraction and ethanol precipitation. RNA was resuspended in double-distilled water and stored at −70°C. For the reverse transcriptase (RT) PCR, primer TF6 (Table 1 and Fig. 1) was annealed to 20 μg of A. pleuropneumoniae RNA in a volume of 10 μl; the RT reaction was carried out in a total volume of 20 μl with Superscript II (GIBCO-BRL) according to the manufacturer's instructions. The resulting cDNA was treated with DNase-free RNase (2 U; Boehringer GmbH, Mannheim, Germany) for 20 min at 37°C, phenol extracted, ethanol precipitated, and resuspended in 10 μl of double-distilled water. The PCR was performed using the cDNA in a 1:1,000 dilution.
Nucleotide sequence accession number.
The DNA sequence containing the A. pleuropneumoniae exbBD genes has been assigned GenBank accession no. Y17916.
RESULTS
Cloning and transcriptional organization of the A. pleuropneumoniae exbBD genes.
In order to identify additional iron-regulated genes, we isolated a clone from a λ2001 library containing a 1,477-bp region upstream of the tbpB gene (Fig. 1). Sequence analysis of this region revealed the presence of two open reading frames (ORF) encoding products of 222 and 136 amino acids, the first one showing significant homologies of approximately 43% with the N. gonorrhoeae and N. meningitidis ExbB proteins and 29% with the E. coli TolQ protein and the second one showing similarities with the respective ExbD and TolR proteins (Fig. 2). Based on the very high degree of homology to the more recently accessible genes of Neisseria spp., the first gene was renamed from tolQ (27) to exbB, and the second one was designated exbD. The exbB ORF is preceded by a Shine-Dalgarno consensus sequence. The exbD ORF directly follows the exbB ORF, with the methionine codon of exbD overlapping the exbB stop codon, and the putative exbD ORF ends only 27 bp upstream of the tbpB methionine codon (Fig. 1). The transcriptional organization of the exbBD and tbpB genes, as assessed by RT-PCR analysis, revealed that all three ORF are located on a single polycistronic mRNA (Fig. 3).
FIG. 2.
Comparison of the ExbB (top) and ExbD (bottom) proteins of A. pleuropneumoniae (ACTPL), N. gonorrhoeae (neigo), and N. meningitidis (neime) as well as the TolQ and TolR proteins of E. coli (ecoli). Boxes indicate identity; dashes indicate differences.
FIG. 3.
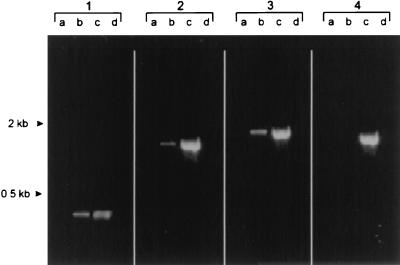
RT-PCR analysis of RNA from A. pleuropneumoniae grown under standard (lanes a) and iron-restricted (lanes b) conditions. Lanes C, A. pleuropneumoniae chromosomal DNA; lanes d, negative controls with no template DNA. Primer TF6 was paired with primers BA23 (panel 1), BA9 (panel 2), and BA6 (panel 3). In order to ensure the absence of chromosomal DNA in the RNA preparations, a pair of primers specific for the A. pleuropneumoniae rho gene was used as a control (panel 4).
Functional analysis of the exbBD genes.
In order to investigate translation of the putative exbBD genes in A. pleuropneumoniae, an GST-ExbB fusion protein was constructed, and antibodies raised against the fusion protein were used to determine expression in A. pleuropneumoniae grown under regular and iron-restricted conditions. A protein of the predicted size was expressed under iron-restricted growth conditions only (Fig. 4). In order to investigate a possible role of the ExbBD proteins in the utilization of transferrin-bound iron, an isogenic and nonpolar deletion mutant, designated AP76Δexb, was constructed (Fig. 1) and confirmed by PCR analysis, Southern blotting, PFGE, and Western blotting (Fig. 4) as well as by nucleotide sequence analysis of a PCR product obtained from AP76Δexb. Then, the A. pleuropneumoniae wild-type strain AP76, AP76Δexb, and AP76Δexb transformed with plasmid-carried exbBD genes were grown on porcine transferrin as the sole source of iron. The deletion mutant was unable to utilize transferrin-bound iron but could be complemented in trans by exbBD-carrying recombinant plasmids pFOKE2 and pFOKE5 (Fig. 5).
FIG. 4.
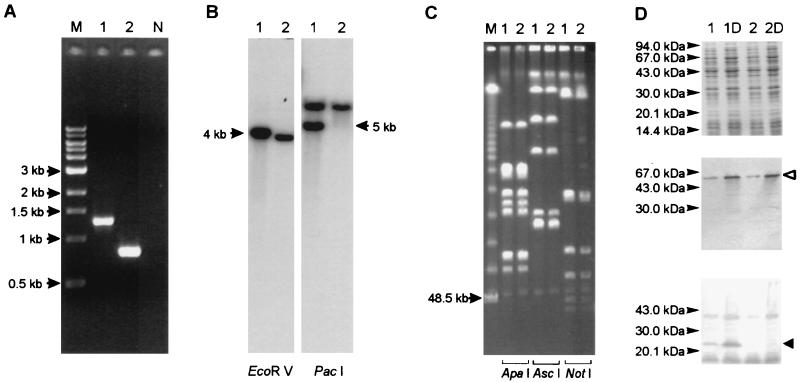
Analysis of A. pleuropneumoniae AP76 (lanes 1) and mutant strain AP76Δexb (lanes 2) and a negative control for PCR (lane N). Lanes M, size markers. (A) PCR using primers RE1 and BA7. (B) Southern blot analysis with EcoRV- and PacI-digested DNA and the exbB gene as a probe. (C) PFGE of ApaI-, AscI, and NotI-digested DNA. (D) Coomassie blue-stained gel (top) and Western blots developed with serum directed against the TbpB protein (middle) and the ExbB protein (bottom). Growth under iron-restricted conditions was achieved by the addition of dipyridyl (lanes 1D and 2D). The open arrowhead indicates the position of the TbpB protein; the solid arrowhead indicates the position of the ExbB protein.
FIG. 5.
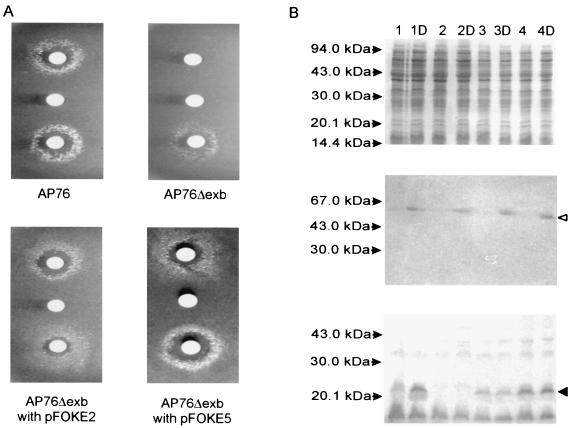
ExbBD function is required for the utilization of transferrin-bound iron and can be complemented in trans. (A) Plate bioassay with strains cultured on iron-depleted brain heart infusion agar supplemented via paper disks with porcine transferrin (top), dilution buffer (middle), and ferric citrate (bottom). (B) Coomassie blue-stained gel (top) and Western blots developed with serum directed against the TbpB protein (middle) and the ExbB protein (bottom) of A. pleuropneumoniae AP76 (lane 1), AP76Δexb (lane 2), and AP76Δexb transformed with pFOKE2 (lane 3) or pFOKE5 (lane 4) in the absence and presence of dipyridyl (D). The openarrow head indicates the position of the TbpB protein; the solid arrowhead indicates the position of the ExbB protein.
Localization of the exbBD genes in A. pleuropneumoniae serotype reference strains.
In order to investigate whether the linkage of the exbBD and tbpB genes was unique to A. pleuropneumoniae serotype 7, DNAs of the 12 A. pleuropneumoniae serotype reference strains were investigated by PCR using exbB- and tbpB-specific primers BA7 and RE1; in addition, whole-cell lysates of these strains grown under iron-limiting conditions were tested in a Western blot for the presence of ExbB protein. The close linkage of the exbBD and tbpB genes was present in all A. pleuropneumoniae serotype reference strains, and all strains expressed the ExbB protein upon iron restriction (Fig. 6).
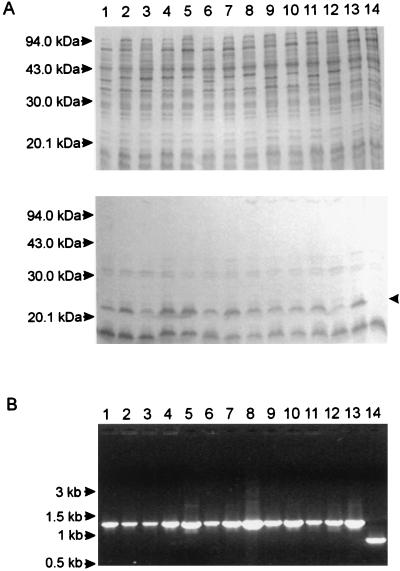
FIG. 6.
ExbB expression and localization of the exbBD genes in A. pleuropneumoniae serotype reference strains 1 to 12 (lanes 1 to 12), AP76 (lane 13), and AP76Δexb (lane 14). (A) Coomassie blue-stained gel (top) and Western blot developed with serum directed against the ExbB protein (bottom) of A. pleuropneumoniae strains grown under iron-restricted conditions. The arrowhead indicates the position of ExbB. (B) Ethidium bromide-stained gel of a PCR analysis using primers RE1 and BA7.
DISCUSSION
In this report, we describe the cloning and molecular analysis of a 1.5-kb region upstream of the tbpB gene of A. pleuropneumoniae serotype 7. We show that in A. pleuropneumoniae, two ORF whose products have more than 40% homology to neisserial ExbBD proteins (5, 35) are located immediately upstream of the tbpBA genes. This localization of exbBD genes in a single operon with the tbpBA genes has not been observed for other bacterial species. Thus, in Neisseria spp., in H. influenzae, and in P. haemolytica, a putative promoter region is located immediately in front of the tbpB gene (4, 17, 21); this arrangement had also been proposed for A. pleuropneumoniae (16). In H. influenzae and in P. haemolytica, the accessible genomic sequence data reveal no homology of DNA upstream of tbpB to exbBD sequences. In Neisseria spp., the exbBD genes have been cloned (5, 35), and no genetic linkage to the tbpBA genes has been observed. These results further support previous observations that despite significant homology between single genes, genomic organization significantly differs among the members of the family Pasteurellaceae (23, 27).
In order to investigate whether the exbBD ORF were translated and coregulated with the tbpBA genes, a specific antiserum was raised against a GST-ExbB fusion protein and used in Western blot analyses. This investigation proved a clearly increased expression of ExbB protein under iron-restricted conditions (Fig. 4D). Due to the close linkage and coordinated expression of exbBD and tbpBA genes, it was hypothesized that the ExbBD proteins of A. pleuropneumoniae might be required as energy couplers for the utilization of transferrin-bound iron, as has been described for N. gonorrhoeae (5). This hypothesis was further supported by a recently reported linkage of a set of exbBD genes to heme transport genes of Vibrio cholerae (25) and, in addition, by the observation that the exbBD genes are essential for the uptake of ferric iron in Xanthomonas campestris (37).
In order to prove this hypothesis, a newly developed method of constructing nonpolar deletion mutants of A. pleuropneumoniae (28) was used to interrupt the exbB ORF without interfering with the expression of the transferrin-binding proteins (Fig. 4). Growth experiments with wild-type and isogenic mutant strains showed that the exbBD genes were in fact required for the utilization of transferrin-bound iron in A. pleuropneumoniae. This result was further confirmed by restoring the wild-type phenotype of the mutant in trans by transforming it with exbBD-expressing plasmids. This dependence on exbB showed that the A. pleuropneumoniae uptake of transferrin-bound iron cannot be supplemented by a backup system, such as tolQR. However, the unaffected growth in vitro as well as the continuous utilization of ferric citrate supports the presence of a second locus involved in the uptake of ferric iron by A. pleuropneumoniae, as has been described previously (7).
Since a putative promoter sequence upstream of the tbpB gene of A. pleuropneumoniae had been reported previously (16), the transcriptional organization of the exbBD and tbpB genes was investigated; these genes are located on a single polycistronic mRNA. In addition, the Western blot investigation of A. pleuropneumoniae AP76Δexb transformed with pFOKE2 and pFOKE5 implied that the iron-regulated promoter of the operon is located upstream of the DNA fragment investigated in this study. Thus, no increase of ExbB expression could be demonstrated in A. pleuropneumoniae pFOKE2 transformants containing the exbBD genes, although these genes are not controlled by the vector-derived T4 promoter (Fig. 5). Further, this finding suggests that an incomplete ORF whose product has a significant degree of homology to the carboxy-terminal end of the neisserial TonB protein and which is located immediately upstream of the exbB ORF might also be located on the same transcript driven by a single iron-regulated promoter even further upstream.
An investigation of the A. pleuropneumoniae serotype reference strains further revealed that this genomic organization was not unique to the A. pleuropneumoniae serotype 7 strain used in this study (Fig. 6). Thus, a PCR fragment of identical size was amplified from all A. pleuropneumoniae serotype reference strains with primers located upstream of the exbB gene and at the beginning of the tbpB gene. In addition, the antibody directed against the ExbB protein detected a protein of identical size in all lysates. The variable intensities seen could have been due to the presence of various amounts of protein; alternatively, slight antigenic differences might have been responsible.
In conclusion, the results described here imply that a nonpolar deletion of exbBD might present a suitable way to attenuate A. pleuropneumoniae strains for use as a live vaccine. Thus, this mutation does not interfere with routine in vitro culturing or with the expression of protective iron-regulated proteins and, at the same time, completely blocks an iron uptake mechanism considered to be of prime importance during infection (19). In addition, the possibility of restoring the ability of using transferrin-bound iron in trans might allow the use of the exbBD genes as a nonantibiotic selection marker for maintaining recombinant plasmids in A. pleuropneumoniae.
ACKNOWLEDGMENTS
This work was supported by grant GE522/3-1 from the Deutsche Forschungsgemeinschaft, Bonn, Germany. W.T. is a fellow of the Mahanakorn University of Technology, Bangkok, Thailand. S.T. is a fellow of the Graduiertenkolleg Zell- und Molekularbiologie in der Tiermedizin of the Deutsche Forschungsgemeinschaft, Bonn, Germany.
REFERENCES
- 1.Ahmer B M, Thomas M G, Larsen R A, Postle K. Characterization of the exbBD operon of Escherichia coli and the role of ExbB and ExbD in TonB function and stability. J Bacteriol. 1995;177:4742–4747. doi: 10.1128/jb.177.16.4742-4747.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ala'Aldeen D A. Transferrin receptors of Neisseria meningitidis: promising candidates for a broadly cross-protective vaccine. J Med Microbiol. 1996;44:237–243. doi: 10.1099/00222615-44-4-237. [DOI] [PubMed] [Google Scholar]
- 3.Anderson C, Potter A A, Gerlach G F. Isolation and molecular characterization of spontaneously occurring cytolysin-negative mutants of Actinobacillus pleuropneumoniae serotype 7. Infect Immun. 1991;59:4110–4116. doi: 10.1128/iai.59.11.4110-4116.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Anderson J A, Sparling P F, Cornelissen C N. Gonococcal transferrin-binding protein 2 facilitates but is not essential for transferrin utilization. J Bacteriol. 1994;176:3162–3170. doi: 10.1128/jb.176.11.3162-3170.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Biswas G D, Anderson J E, Sparling P F. Cloning and functional characterization of Neisseria gonorrhoeae tonB, exbB and exbD genes. Mol Microbiol. 1997;24:169–179. doi: 10.1046/j.1365-2958.1997.3421692.x. [DOI] [PubMed] [Google Scholar]
- 6.Braun V, Killmann H. Bacterial solutions to the iron-supply problem. Trends Biochem Sci. 1999;24:104–109. doi: 10.1016/s0968-0004(99)01359-6. [DOI] [PubMed] [Google Scholar]
- 7.Chin N, Frey J, Chang C F, Chang Y F. Identification of a locus involved in the utilization of iron by Actinobacillus pleuropneumoniae. FEMS Microbiol Lett. 1996;143:1–6. doi: 10.1111/j.1574-6968.1996.tb08452.x. [DOI] [PubMed] [Google Scholar]
- 8.Cornelissen C N, Sparling P F. Iron piracy: acquisition of transferrin-bound iron by bacterial pathogens. Mol Microbiol. 1994;14:843–850. doi: 10.1111/j.1365-2958.1994.tb01320.x. [DOI] [PubMed] [Google Scholar]
- 9.Cornelissen C N, Kelley M, Hobbs M M, Anderson J E, Cannon J G, Cohen M S, Sparling P F. The transferrin receptor expressed by gonococcal strain FA1090 is required for the experimental infection of human male volunteers. Mol Microbiol. 1998;27:611–616. doi: 10.1046/j.1365-2958.1998.00710.x. [DOI] [PubMed] [Google Scholar]
- 10.Dehio C, Meyer M. Maintenance of broad-host-range incompatibility group P and group Q plasmids and transposition of Tn5 in Bartonella henselae following conjugal plasmid transfer from Escherichia coli. J Bacteriol. 1997;179:538–540. doi: 10.1128/jb.179.2.538-540.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Eick-Helmerich K, Braun V. Import of biopolymers into Escherichia coli: nucleotide sequences of the exbB and exbD genes are homologous to those of the tolQ and tolR genes, respectively. J Bacteriol. 1989;171:5117–5126. doi: 10.1128/jb.171.9.5117-5126.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fenwick B, Henry S. Porcine pleuropneumonia. J Am Vet Med Assoc. 1994;204:1334–1340. [PubMed] [Google Scholar]
- 13.Frey J. Construction of a broad host range shuttle vector for gene cloning and expression in Actinobacillus pleuropneumoniae and other Pasteurellaceae. Res Microbiol. 1992;143:263–269. doi: 10.1016/0923-2508(92)90018-j. [DOI] [PubMed] [Google Scholar]
- 14.Gerlach G-F, Anderson C, Potter A A, Klashinsky S, Willson P J. Cloning and expression of a transferrin-binding protein from Actinobacillus pleuropneumoniae. Infect Immun. 1992;60:892–898. doi: 10.1128/iai.60.3.892-898.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gerlach G-F, Klashinsky S, Anderson C, Potter A A, Willson P J. Characterization of two genes encoding distinct transferrin-binding proteins in different Actinobacillus pleuropneumoniae isolates. Infect Immun. 1992;60:3253–3261. doi: 10.1128/iai.60.8.3253-3261.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gonzalez G C, Yu R-H, Rosteck P R, Schryvers A B. Sequence, genetic analysis, and expression of Actinobacillus pleuropneumoniae transferrin receptor genes. Microbiology. 1995;141:2405–2416. doi: 10.1099/13500872-141-10-2405. [DOI] [PubMed] [Google Scholar]
- 17.Gray-Owen S D, Loosmore S, Schryvers A B. Identification and characterization of genes encoding the human transferrin-binding proteins from Haemophilus influenzae. Infect Immun. 1995;63:1201–1210. doi: 10.1128/iai.63.4.1201-1210.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Guerinot M L. Microbial iron transport. Annu Rev Microbiol. 1994;48:743–772. doi: 10.1146/annurev.mi.48.100194.003523. [DOI] [PubMed] [Google Scholar]
- 19.Hennig I, Teutenberg-Riedel B, Gerlach G-F. Downregulation of a protective Actinobacillus pleuropneumoniae antigen during the course of infection. Microb Pathog. 1999;26:53–63. doi: 10.1006/mpat.1998.0249. [DOI] [PubMed] [Google Scholar]
- 20.Higgs P I, Myers P S, Postle K. Interactions in the TonB-dependent energy transduction complex: ExbB and ExbD form homomultimers. J Bacteriol. 1998;180:6031–6038. doi: 10.1128/jb.180.22.6031-6038.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Legrain M, Jacobs E, Irwin S W, Schryvers A B, Quentin-Millet M J. Molecular cloning and characterization of Neisseria meningitidis genes encoding the transferrin binding proteins Tbp1 and Tbp2. Gene. 1993;130:73–80. doi: 10.1016/0378-1119(93)90348-7. [DOI] [PubMed] [Google Scholar]
- 22.Moeck G S, Coulton J W. TonB-dependent iron acquisition: mechanisms of siderophore-mediated active transport. Mol Microbiol. 1998;28:675–681. doi: 10.1046/j.1365-2958.1998.00817.x. [DOI] [PubMed] [Google Scholar]
- 23.Myers L E, Yang Y P, Du R P, Wang Q, Harkness R E, Schryvers A B, Klein M H, Loosmore S M. The transferrin binding protein B of Moraxella catarrhalis elicits bactericidal antibodies and is a potential vaccine antigen. Infect Immun. 1998;66:4183–4192. doi: 10.1128/iai.66.9.4183-4192.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Niven D F, Donga J, Archibald F S. Responses of Haemophilus pleuropneumoniae to iron restriction: changes in the outer membrane protein profile and the removal of iron from porcine transferrin. Mol Microbiol. 1989;3:1083–1089. doi: 10.1111/j.1365-2958.1989.tb00258.x. [DOI] [PubMed] [Google Scholar]
- 25.Occhino D A, Wyckoff E E, Henderson D P, Wrona T J, Payne S M. Vibrio cholerae iron transport: haem transport genes are linked to one of two sets of tonB, exbB, exbD genes. Mol Microbiol. 1998;29:1493–1507. doi: 10.1046/j.1365-2958.1998.01034.x. [DOI] [PubMed] [Google Scholar]
- 26.Ogunnariwo J A, Woo T K W, Lo R Y C, Gonzalez G C, Schryvers A B. Characterization of the Pasteurella haemolytica transferrin receptor genes and the recombinant receptor proteins. Microb Pathog. 1997;23:273–284. doi: 10.1006/mpat.1997.0156. [DOI] [PubMed] [Google Scholar]
- 27.Oswald W, Konine D V, Rohde J, Gerlach G-F. First chromosomal restriction map of Actinobacillus pleuropneumoniae and localization of putative virulence-associated genes. J Bacteriol. 1999;184:4161–4169. doi: 10.1128/jb.181.14.4161-4169.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Oswald W, Tonpitak W, Ohrt G, Gerlach G-F. A single-step transconjugation system for the introduction of unmarked deletions into Actinobacillus pleuropneumoniae serotype 7 using a sucrose sensitivity marker. FEMS Microbiol Lett. 1999;179:153–160. doi: 10.1111/j.1574-6968.1999.tb08721.x. [DOI] [PubMed] [Google Scholar]
- 29.Raleigh F A, Lech K, Brent R. Selected topics from classical bacterial genetics. In: Ausubel F M, et al., editors. Current protocols in molecular biology. New York, N.Y.: Publishing Associates and Wiley Interscience; 1989. pp. 1.4.1–1.4.14. [DOI] [PubMed] [Google Scholar]
- 30.Raymond K N, Müller G I, Matzanke B F. Complexation of iron by siderophores. A review of their solution and structural chemistry and biological functions. Top Curr Chem. 1989;123:49–58. [Google Scholar]
- 31.Rossi-Campos A, Anderson C, Gerlach G-F, Klashinsky S, Potter A A, Willson P J. Immunization of pigs against Actinobacillus pleuropneumoniae with two recombinant protein preparations. Vaccine. 1992;10:512–518. doi: 10.1016/0264-410x(92)90349-o. [DOI] [PubMed] [Google Scholar]
- 32.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1989. [Google Scholar]
- 33.Schryvers A B, Gonzalez G C. Receptors for transferrin in pathogenic bacteria are specific for the host's transferrin. Can J Microbiol. 1990;36:145–147. doi: 10.1139/m90-026. [DOI] [PubMed] [Google Scholar]
- 34.Schryvers A B, Gray-Owen S. Iron acquisition in Haemophilus influenzae: receptors for human transferrin. J Infect Dis. 1992;165:S103–S104. doi: 10.1093/infdis/165-supplement_1-s103. [DOI] [PubMed] [Google Scholar]
- 35.Stojiljkovic I, Srinivasan N. Neisseria meningitidis tonB, exbB, and exbD genes: Ton-dependent utilization of protein-bound iron in neisseriae. J Bacteriol. 1997;179:805–812. doi: 10.1128/jb.179.3.805-812.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Weinberg E D. Iron and infection. Microbiol Rev. 1978;42:45–66. doi: 10.1128/mr.42.1.45-66.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wiggerich H G, Klauke B, Köplin R, Priefer U B, Pühler A. Unusual structure of the tonB-exb DNA region of Xanthomonas campestris pv. campestris: tonB, exbB, and exbD1 are essential for ferric iron uptake, but exbD2 is not. J Bacteriol. 1997;179:7103–7110. doi: 10.1128/jb.179.22.7103-7110.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Wilke M, Franz B, Gerlach G-F. Characterization of a large transferrin-binding protein from Actinobacillus pleuropneumoniae serotype 7. Zentbl Veterinaermed Reihe B. 1997;44:73–86. doi: 10.1111/j.1439-0450.1997.tb00953.x. [DOI] [PubMed] [Google Scholar]