Abstract
Aims
Computer-aided detection systems for retinal fluid could be beneficial for disease monitoring and management by chronic age-related macular degeneration (AMD) and diabetic retinopathy (DR) patients, to assist in disease prevention via early detection before the disease progresses to a “wet AMD” pathology or diabetic macular edema (DME), requiring treatment. We propose a proof-of-concept AI-based app to help predict fluid via a “fluid score”, prevent fluid progression, and provide personalized, serial monitoring, in the context of predictive, preventive, and personalized medicine (PPPM) for patients at risk of retinal fluid complications.
Methods
The app comprises a convolutional neural network–Vision Transformer (CNN-ViT)–based segmentation deep learning (DL) network, trained on a small dataset of 100 training images (augmented to 992 images) from the Singapore Epidemiology of Eye Diseases (SEED) study, together with a CNN-based classification network trained on 8497 images, that can detect fluid vs. non-fluid optical coherence tomography (OCT) scans. Both networks are validated on external datasets.
Results
Internal testing for our segmentation network produced an IoU score of 83.0% (95% CI = 76.7–89.3%) and a DICE score of 90.4% (86.3–94.4%); for external testing, we obtained an IoU score of 66.7% (63.5–70.0%) and a DICE score of 78.7% (76.0–81.4%). Internal testing of our classification network produced an area under the receiver operating characteristics curve (AUC) of 99.18%, and a Youden index threshold of 0.3806; for external testing, we obtained an AUC of 94.55%, and an accuracy of 94.98% and an F1 score of 85.73% with Youden index.
Conclusion
We have developed an AI-based app with an alternative transformer-based segmentation algorithm that could potentially be applied in the clinic with a PPPM approach for serial monitoring, and could allow for the generation of retrospective data to research into the varied use of treatments for AMD and DR. The modular system of our app can be scaled to add more iterative features based on user feedback for more efficient monitoring. Further study and scaling up of the algorithm dataset could potentially boost its usability in a real-world clinical setting.
Supplementary information
The online version contains supplementary material available at 10.1007/s13167-022-00301-5.
Keywords: Retinal fluid; Ophthalmology; Optical coherence tomography; Deep learning; Computer-aided detection; Predictive, preventive, and personalized OCT; Anti-vascular endothelial growth factor; Age-related macular degeneration; Diabetic retinopathy; Diabetic macular edema
Introduction
Ocular disorders such as age-related macular degeneration (AMD), diabetic macular edema (DME), retinal vein occlusion (RVO), central serous chorioretinopathy (CSC), and uveitis can cause an accumulation of fluid exudates or cysts in the retinal layers, reducing visual acuity (VA), potentially leading to blindness if left untreated [1–5]. Currently, the most important routine examination for monitoring of fluid complications arising from these conditions is spectral-domain optical coherence tomography (SD-OCT), which has been the product of extensive development over the years since the inception of optical coherence tomography (OCT) in 1991 [6]. Compared to a regular fundus photograph which only shows the en-face view of the posterior pole of the eye, SD-OCT displays cross-sectional image scans (B-scans) of the macula region to allow clinicians to monitor fluid progression [7] and is currently recognized as a reference standard [8, 9] to diagnose macular edema and provide treatment. Treatment possibilities include the use of steroids [7], randomized controlled trial (RCT)-proven intravitreal injections (IVT) of anti-angiogenic medications (e.g., anti-VEGF) such as ranibizumab, bevacizumab, or aflibercept [10, 11], photodynamic therapy (PDT) with verteporfin [12], or submacular surgery [12].
The location and volume of fluid in the retina could affect visual acuity (VA) and disease progression differently and the explicit delineation of these regions could help in monitoring or quantifying the status of the condition, provide possible research avenues in precision medicine, avoid missed, or false negative fluid regions, or assist with adjusting anti-VEGF treatment requirements, such as frequency of injections via treat-and-extend (T&E) protocols, for instance [7]. In view of this, many research groups have used machine learning (ML) [13, 14] and deep learning (DL) [13–17] methods, or organized benchmark challenges such as RETOUCH [15] to develop methods for automatically segmenting the retina or retinal abnormalities, or classifying OCT B-scan images according to the absence or presence of several fluid types, such as intraretinal fluid (IRF), subretinal fluid (SRF), or pigment epithelial detachment (PED). We have also developed a DL algorithm to segment and produce an abnormality score for the outer retinal layer on OCT scans [18]. In addition, we have also demonstrated that a CNN can successfully identify, based on a heatmap, clinically relevant features of “wet” AMD in SD-OCT scans and was able to distinguish them from normal scans with an AUC of 95.2% on the individual level for an external test set [19]. Such DL approaches continue to advance and allow for the improved management of retinal abnormalities via computer-aided methods, which contributes to the information and communication (ICT) technologies that form the modernization of healthcare for predictive, preventive, and personalized medicine, PPPM [20].
Since the success of the AlexNet Convolutional Neural Network (CNN) in 2012 on images from ImageNet [21], CNNs have been progressively introduced into medical image analysis due to its effectiveness in imaging tasks compared to traditional ML methods [22]. With new bespoke network architectures being introduced frequently, CNNs are now one of the de facto image analysis methods for medical image segmentation and image classification tasks.
Transformers, currently one of the most effective DL software architectures used in language applications for machines, such as chatbots (including architectures such as Bidirectional Encoder Representations from Transformers (BERT) and Generative Pre-trained Transformer (GPT) [23]), have recently been introduced into the image analysis toolbox as Vision Transformers (ViTs) [24, 25] and could offer an alternative to CNN algorithms, each with different properties: CNNs offer the benefits of an inductive bias, local receptive fields, and hierarchical feature representation [22], while ViTs offer the benefit of global receptive fields [26] and inclusion of long-range dependencies [25]. Because of a lack of an inductive bias, transformers have been shown to be slightly less accurate in classification compared to CNNs even when trained with relatively large training datasets of 1.2 million images from ImageNet [24]. This hinders its use in DL in the medical domain, which commonly has very few or expensive training data. Despite this limitation, the use of ViTs alone or CNN-ViT architectures for DL on multimodal, 2D, and 3D medical images has been explored in several studies [25, 27–29]. We propose herein a new architecture which combines CNN and ViT (which we term CNN-ViT) that could work as a segmentation network for a computer-aided detection (CADe) system for retinal fluid detection. This system can leverage on the benefits of both CNNs and ViTs, can work with small training data, allowing CADe systems to better detect retinal fluid. In addition, the CADe has a CNN that could predict the chance of fluid on an OCT scan, via a probability score ranging from 0 to 1 – a “fluid score”—with a threshold point that indicates the presence of fluid when crossed.
Training for our DL segmentation was done on the Singapore Epidemiology of Eye Diseases (SEED) dataset and evaluated on an external dataset from Severance Eye Hospital. Inference of the trained segmentation model on publicly available OCT data [30] comprising diseased and healthy eyes produced segmentation masks, which were then subsequently used with the OCT image to train a classification algorithm for predicting the probability of fluid presence. The trained classification model was then externally validated on a dataset of DME patients from Westmead Hospital, Australia, to ensure that our model can predict well on data that has not been seen by the machine.
Previously, AI-based retinal fluid monitoring applications have been developed, such as the Vienna Fluid Monitor [31] as well as an effective computer-aided segmentation and classification algorithm for the diagnosis and referral of retinal disease [16]. These recent successes as well as further development of DL algorithms in the literature for ophthalmology-related tasks enhance their potential for deployment more widely as a clinical decision support tool. Herein, we propose a two-tier, DL segmentation and classification algorithm as a possible tool in the arsenal of AI algorithms in ophthalmology, with its development process outlined in Fig. 1.
Fig. 1.
Flowchart of the overall segmentation and classification algorithm development process. Larger images of the model architectures shown in this image are included in supplementary data (Supplementary Figures 1 and 2)
Similar tools and applications have also been released into the market for telemonitoring or home monitoring, such as the ForeseeHome by Notal Vision, which relies on the patented technology Preferential Hyperacuity Perimetry (PHP) to detect recent-onset choroidal neovascularization (CNV) by identifying and quantifying visual abnormalities such as scotomas or metamorphopsia [32]. First-of-its-kind home OCT devices such as the Notal Home OCT (Notal Vision) [33] with the Notal OCT Analyzer (NOA) machine learning algorithm are also available to allow patients to take daily OCT images of their eye, which are subsequently analyzed using an AI algorithm to be conveyed to the treating physician.
Working hypothesis and purpose of the study
We propose an alternative CNN-ViT segmentation algorithm and a browser-based app for use in a clinic-based setting. In the context of PPPM [20], a prediction of a “fluid score” probability, a targeted DL segmentation approach (compared to only a single DL classification score), preventive care via more precise screening, and a more personalized treatment regimen based on quantification of fluid levels over visits would be an advantageous and novel approach to manage patients with AMD or diabetic retinopathy (DR), as this is currently not widely practiced in the clinical setting. Usually, treatment is offered under the pro re nata (PRN, as needed) or treat-and-extend (T&E) model where treatment is given based on regular follow-ups. Often, these durations are not properly documented [34]. Thus a computer-aided PPPM approach is especially important since AMD [35] and diabetes [36] patients would benefit from an objective monitoring approach to prevent disease manifestation and deterioration. The app has a database that can store algorithm outputs of the scans such as the date of input, quantified total fluid levels of each batch of scans, “fluid score” probabilities, and classified categories. This could streamline the personalized management of chronic fluid conditions for both the patient and eye healthcare provider.
Methods
Segmentation dataset preparation
All anonymized, macula-centered B-scan images were extracted from Heidelberg Spectralis SD-OCT (Heidelberg Engineering, Germany). These scans are from the Singapore Epidemiology of Eye Diseases (SEED) study—a multi-ethnic, longitudinal, population-based study that evaluates the incidence, prevalence, risk factors, and novel biomarkers of age-related eye diseases for Singaporean adults of Malay, Indian, and Chinese descent [37]. After extraction from Heidelberg Spectralis .E2E files into the .png file format, they were subsequently processed via the image processing software GIMP (The GIMP Development Team. https://www.gimp.org) for histogram equalization and a subsequent adjustment of contrast and brightness as a pre-processing step. Training data included 75 Heidelberg Spectralis SD-OCT B-scans of size 496 × 768 pixels (9 mm width) from the Singapore Chinese Eye Study 2 (SCES 2) [37] and Singapore Indian Eye Study 2 (SINDI 2) [37] identified with subretinal fluid (SRF) and/or intraretinal fluid (IRF), 12 scans with epiretinal membrane (ERM), 10 scans with pigment epithelial detachment (PED), and 3 scans with macular hole. For validation during training, 5 B-scans with fluid were used, and for the internal testing set, 10 B-scans with fluid were used. The 25 B-scans with non-treatable fluid or non-fluid conditions (ERM, PED, macular hole) used for training were annotated without fluid ground truth—the inclusion of these cases was to enable the algorithm to better recognize the retinal structures to prevent incorrect annotations in a wider variety of pathologies, as we had seen in preliminary runs. Ground truth annotations were made by a retinal specialist (T.H.R.) with MIPAV software (National Institutes of Health (NIH), Center for Information Technology, https://mipav.cit.nih.gov/) and GIMP.
Image augmentation of the 100 SEED training images with ground truth was extensively applied to prevent overfitting to a small dataset and enable generalization to images with varying noise levels, image pre-processing, extent of fluid, and different scale and position, via the Albumentations package. The final training dataset has a total of 992 training images with their associated mask label of three classes—background, retinal region, and fluid region (if any).
100 SD-OCT scans (50 scans with 6 mm width and 50 scans with 9 mm width) from Severance Hospital, Yonsei University were annotated by a retinal specialist (K.T.) for fluid regions containing only SRF and/or IRF. The data was used as an external testing set for the evaluation of fluid segmentation performance via the metrics Intersection-Over-Union (IoU) and DICE Score. Segmentation annotations were made on MIPAV software. Description of the datasets used for segmentation can be found in Table 1.
Table 1.
Description of the datasets used in this study
| Category | Dataset source | Use | Diseases | Modality | Testing |
|---|---|---|---|---|---|
| Training and internal validation | SEED Study (SCES 2, SINDI 2) (Singapore) [37] | Training and internal test for segmentation | Patients identified with retinal fluid; and patients with non-fluid conditions (ERM, PED, macular hole) | Heidelberg Spectralis SD-OCT | Segmentation only |
| Kermany et. al [27], Kaggle OCT Image dataset | Training and internal test for classification | Patients with choroidal neovascularization (CNV), DME, or AMD; and Normal patients | Heidelberg Spectralis SD-OCT | Segmentation inference; classification test | |
| External validation | Severance Eye Hospital (South Korea) | External validation to test segmentation quality | AMD or DME patients identified with retinal fluid | Heidelberg Spectralis SD-OCT | Segmentation only |
| Westmead Hospital (Sydney, Australia) | External validation to test classification CNN | DME patients and normal patients | Heidelberg Spectralis SD-OCT | Segmentation inference; classification test |
Classification dataset preparation
Training data for classification was obtained from Kermany et al. [30], which is also released publicly as a Kaggle dataset (URL: https://www.kaggle.com/datasets/paultimothymooney/kermany2018). Images were selected and sorted by two retinal specialists (K.T. and T.H.R.) into normal scans, abnormal with/without inactive fluid, and abnormal scans with fluid suitable for IVT treatment, resulting in a total of 11,330 B-scans. As a pre-processing step, areas that did not belong to the OCT scan (white areas from image pre-processing) were processed to be zero intensity (pixel value 0) using the image processing software GIMP. There was a total of 3212 images showing retina with active fluid, 3138 images showing abnormal retina without active fluid, and 5000 images showing normal retina. Images with normal retina and the abnormal retina without active fluid were combined and set as one category, while the other category was images with abnormal retina with active fluid—resulting in a binary classification task. The images were randomly and uniformly split into training, tuning, and testing set, in the ratio 75%: 5%: 20% respectively as the data was not initially organized at the patient level. Training data consisted of 8497 B-scans with their generated segmentation masks; tuning data consisted of 566 images with generated masks; internal testing data consisted of 2267 scans with generated masks.
The external testing dataset from Westmead Hospital consisted of 29 patients of varied ethnicity, with a total of 204 eyes (5043 B-scans) with or without diabetic macular edema (DME). They were first extracted from.E2E files and converted to.png files. Subsequent pre-processing steps involved a cropping to 496 × 768 pixels centered to the macular region, followed by histogram equalization and adjustment of brightness and contrast via GIMP software. The images were classified at the B-scan level into fluid and non-fluid class by a retinal specialist (K.T. and T.H.R.) resulting in 932 scans with fluid (positive) class and 4111 scans with no fluid (negative) class. Description of the datasets used for classification can be found in Table 1.
CNN-ViT segmentation
We utilized a CNN-ViT with 3 large kernel blocks and 12 ViT layers; with a modified decoder consisting of feature fusion blocks that incorporate large kernel blocks, squeeze-excitation, and gated-attention mechanisms (Supplementary Figure 1)—to segment three categories of our OCT scan: (1) background, (2) retinal “focus” region bounded by the Bruch’s membrane/retinal pigment epithelium (BM/RPE) layer and the internal limiting membrane (ILM) layer, and (3) fluid region (IRF and/or SRF), as a 3-channel output. The optimizer used was AdaBelief with an initial learning rate of 4e-5 and a fixed weight decay of 1e-4 for regularization of the model; the learning rate was gradually decreased to 2e-5 and 1e-5 upon a stasis in training loss; weight decay was decreased till a fixed value of 1e-5 to 1e-6 with each iteration to enable smoother learning. The objective function minimized was a combination of Cross Entropy Loss and Generalized Dice Loss (via MONAI package). The model has approximately 269 million parameters. It was trained with a batch size of 3 for 103 epochs.
Stacked large kernel squeeze-excite CNN classification
We utilized a handcraft CNN (Supplementary Figure 2) that contains stacked large kernel blocks and squeeze-excite mechanism, and two output nodes (one for the probability of no-fluid, one for the probability of fluid). We used a learning rate of 3e-8 with a fixed weight decay of 1e-5 and an AdamW optimizer, trained with a batch size of 10 for 25 epochs; the learning rate was decreased to 1e-8 upon loss plateau with a fixed weight decay value of 1e-5. The objective function minimized was the Cross Entropy Loss with weighting for the class imbalance between the non-fluid and fluid classes based on the proportion of positive and negative class samples in the training set. The model has approximately 120 million parameters. The input was the OCT B-scan and generated mask with only the fluid region segmented.
All models were made using Python version 3.8 (Python software foundation), PyTorch deep learning framework, and Anaconda (Anaconda Inc., https://www.anaconda.com). Image pre-processing and augmentation were mainly done via Numpy, OpenCV, Scikit-image, and Albumentations. Results were analyzed using Matplotlib, Pandas, and Scikit-learn packages.
Shiny browser-based app
We developed a browser-based app with the Shiny framework (https://shiny.rstudio.com/py/) with an SQLite database backend to store predictions on the local server. Packages used include PyTorch, OpenCV, Numpy, Scikit-image, Matplotlib, and Pillow. The app needs to run on sufficiently powered hardware for the AI algorithm to generate predictions—minimum requirements include an NVIDIA graphics processing unit (GPU) with 12 GB of memory or higher, with the installation of NVIDIA CUDA software.
Results
Segmentation test
For the SEED clinical dataset, internal testing produced an IoU score of 83.0% (95% CI = 76.7–89.3%) and a DICE score of 90.4% (86.3–94.4%). For the Korean clinical dataset, we obtained an IoU score of 66.7% (63.5–70.0%) and a DICE Score of 78.7% (76.0–81.4%) (Fig. 2).
Fig. 2.
Segmentation results of internal and external validation cohorts (SEED and Korean (Severance Hospital)) with 95% confidence intervals (CI) shown in brackets
The segmentation visualizations of samples from the internal and external testing datasets are shown in Fig. 3, together with color-coded regions for true positive pixels, false negative pixels, and false positive pixels.
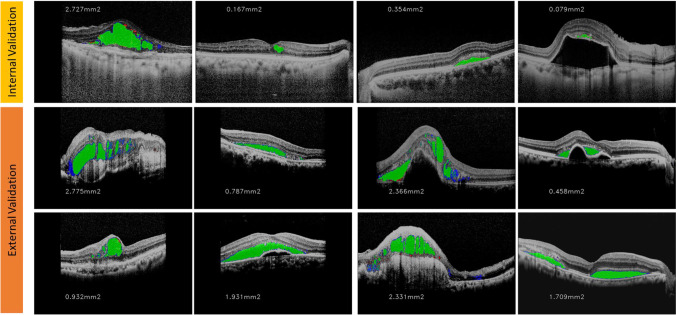
Fig. 3.
Segmentation result visualization of samples from the internal and external validation cohorts. Green is true positive regions, red is false negative regions, and blue is false positive regions. Area of fluid (in mm2) for the B-scan is shown on the left-hand side of the image
The external testing results are similar to RFS-Net for joint segmentation and characterization for anti-VEGF treatment by Hassan et al. [38], whereby a combined IoU of multiple fluids (IRF, SRF, and PED) of 65.1% and an F1 score (equivalent to DICE) of 78.8% was attained. Mantel et al. [39] also reported a Dice score of 72.8% for IRF, 67.5% for SRF, and 81.9% for PED. Wilson et al. [40] have also reported a Dice score of 0.43 (95% CI = 0.29–0.66) to 0.78 (0.57–0.85) for segmentation of fluid. The metrics are not directly comparable with other studies due to the differences in the size of the testing set, as well as the training set; however, the encouraging performance of our CNN-ViT in the segmentation task could be a good indication of improved performance with more varied and larger training dataset sizes.
Classification test
For the internal validation dataset of images from publicly available Kaggle data (from Kermany et al.), we observed an area under the receiver operating characteristics curve (AUC) of 99.18%, and a Youden Index of 0.3806, the cut-off probability at which there is maximum sensitivity and specificity of the model. For the external validation dataset of images from Westmead Hospital, Australia, we observed an AUC of 94.55%, and an accuracy of 94.98% and an F1 score of 85.73% when the Youden Index threshold was applied (Fig. 4).
Fig. 4.
Metrics for classification performance on internal and external testing datasets of the Kaggle dataset and Westmead Hospital dataset (numbers displayed in the table are in percentages). Receiver Operating Characteristics (ROC) curves of the internal validation testing set and the external validation testing set are as shown
Discussion
Output of CADe algorithm
The final output of the combined segmentation and classification algorithm is presented in an easy-to-read, clinically approachable form, suitable for both the patient and the physician or reader. Included is information on the area of fluid segmented in mm2 on the left-hand side, as well as the probability of fluid for the B-scan on the right-hand side along with a color-coded circle indicator (red for fluid, green for no fluid, amber for borderline/just-below threshold), and a colorized visualization of the segmented retinal layers in green, and fluid region in red, if any (Figs. 5, 6, 7).
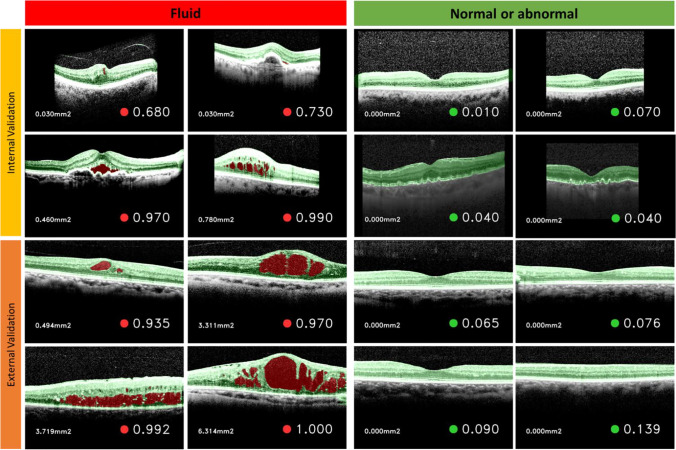
Fig. 5.
Segmentation visualization (retinal layers in green, fluid areas in red) and classification result (“fluid score”) of the CADe fluid detector on the internal and external validation sets with representative B-scan outputs from both the segmentation and classification algorithms (CADe algorithm)
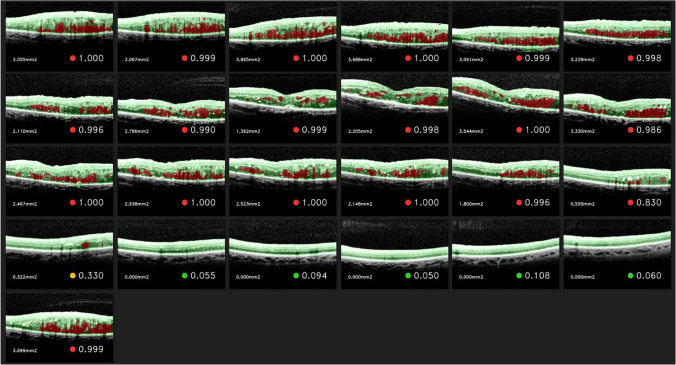
Fig. 6.
Example output of the segmentation and classification algorithm for a DME patient OCT volume of 25 B-scans from the external validation dataset. Segmented retinal layers are in green and segmented fluids are in red
Fig. 7.
Example output of the segmentation and classification algorithm for a healthy patient OCT Volume of 25 B-scans from the external validation dataset. Segmented retinal layers are in green and segmented fluids are in red
Network architectures
For segmentation, we combined mechanisms from several networks, namely Dense Prediction Transformer [41], Effective Squeeze-Excitation [42, 43], Stacked Dilated U-Net [44] for our large-kernel blocks, and Gated Attention [45]. For classification, we handcrafted a CNN with a maximum filter size of 2048 for extraction of features as well as stacked large-kernel CNN blocks and Squeeze-excite blocks. Mantel et al. [43] have reported success with squeeze-excite blocks and larger kernels at the beginning of the neural network to capture a larger context for OCT fluid identification, which we have added in and found to have been beneficial for our purposes; the larger receptive fields provided by the large kernels and ViTs could have contributed to better segmentation performance by providing more context of surrounding areas to improve comprehension of the overall OCT image by the neural network.
Segmentation performance
Segmentation of the background, retinal “focus” region, and fluid region as three categories is strongly preferred over a binary segmentation that only segments fluid region because we found that especially for a small-to-medium training data size, including the background regions and retinal layer (non-fluid) regions along with the fluid regions as ground truth can aid in segmentation. This is supported by Mantel et al. who hypothesized that the segmentation of structures into different classes offered the model a benefit of context of the surrounding regions [39]; other ML and DL algorithms have also used the delineation of the area between the ILM and RPE as a focal “region of interest” for identification of fluid [38, 46].
We also found that our network is generally good at delineating SRF, a possible reason being its proximity to the hyper-reflective RPE layer which gives rise to a larger numerical input signal to the DL network or a large contrast in signal with surrounding areas.
Suboptimal resolution and contrast of the OCT image could also have negatively affected the segmentation of structures such as IRF, which has been shown in a study by Wilson et al. [40]. Low signal-to-noise ratio (SNR) also prevented robust segmentation of both retinal and fluid regions in some OCT scans. Otherwise, images with acceptable SNR, little artifacts, shadowing, or noise and retinal layers that are not severely deformed attained relatively clear segmentations of retinal boundaries and fluid region. Transformer-based or semi-transformer-based software architectures could potentially improve segmentation ability by allowing the neural network to “see” the image better. Also, we add that the measurement of fluid area in mm2 that is produced via the segmentation algorithm can be used as an image biomarker for disease progression and assessment of intervention efficacy over time.
It is encouraging that a DL system trained on 100 unique B-scans with the aid of data augmentation can segment retinal regions and fluid with a reasonable performance. With more varied scans and more data included into the training set, we feel that it will be possible to further improve segmentation as well as subsequent classification performance, especially with our novel CNN-ViT network. A limitation of our work is the availability of scans with large areas of edema or large deformities in the segmentation training dataset, which could have affected the ability of our network to recognize very diverse and abnormal scans.
Classification performance
An output probability score generated based on input parameters (in our case an image), which we use in our CNN and termed it as a “fluid score,” is an accepted metric used in medical ML/AI systems such as the Triage and Acuity Score (TAS) system [47]. The probability score is a metric for indication of how the model makes a classification decision for each image after extraction of features via the DL network, aiding its use as a clinical decision support tool in addition to the segmentation visualization. The Youden Index (point at which there is highest sensitivity and specificity) for the internal testing set was noted and applied to our external testing set—a value of 0.3806 for the cut-off threshold was found after training for 25 epochs with our CNN, applied via an early stopping call-back when no improvement was made to the highest validation accuracy after 3 epochs. Classification metrics such as accuracy, precision, recall, sensitivity, and specificity of the internal and external testing sets were obtained based on this cut-off threshold value.
Only segmentation masks of fluid regions were included as input together with the OCT scan into the classification DL network as we found that the inclusion of segmented retinal layers was detrimental to classification performance. Another group has recently developed an AI-based anti-VEGF response predictor called the Treatment Response Analyzer System (TRAS) with the same mechanism of inputting an SD-OCT image along with DL-segmented fluid regions to classify into a binary - poor, or good - anti-VEGF response outcome [48], which further supports the validity of our method.
Particularly of note is that the performance on the external validation dataset attained a higher specificity (98.03% vs. 96.44%) than the internal validation dataset, even though the external validation dataset contains a larger number of B-scans belonging to the no-fluid class (4111 vs. 1000). The DL algorithm appears to be able to detect true negative cases well, which could possibly be due to the larger number of no-fluid examples used for training the classifier (3212 positive class vs. 8138 negative class). There was a drop in sensitivity (i.e., recall) when the algorithm was applied onto the external validation dataset (98.12% vs. 81.55%)—a larger number of false negatives. However, the precision (i.e., positive predictive value) was comparably slightly lower for the external validation (90.37%) compared to internal validation (91.53%), suggesting that there is a comparable percentage of false positive picked up, which would be beneficial in preventing unnecessary referrals or alarm. The threshold could also be adjusted as well for different specificities and sensitivities, lending itself to a more personalized medicine approach.
We have packaged our AI algorithms in the form of an offline browser-based app with the framework Shiny (https://shiny.rstudio.com/py/), originally made for the R language but recently developed for the Python ecosystem. The app (as shown in Fig. 8 and Supplementary Figures 3 and 4) uses an SQLite database to store the name (or id) of the patient, date of visit, number of scans uploaded per batch, amount of quantified fluid for each batch of images, prediction scores for each image, classification category for each image (fluid or no fluid), and file path for each batch of images generated with the algorithm. All data is stored on a local server and can be used offline, which preserves patient privacy and security. Additional features can potentially be implemented to improve the model predictions, the user interface (UI), as well as general security of the data and software.
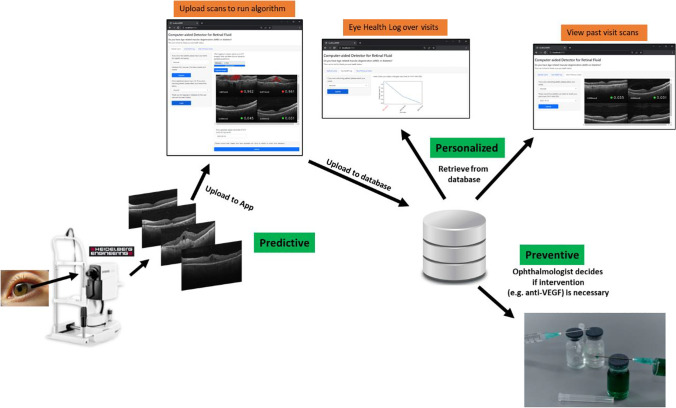
Fig. 8.
Flowchart of the CADe algorithm and app in the context of PPPM. Examples of the web pages are provided in Supplementary Figures 3 and 4
Expert recommendations and impact of the CADe in the context of PPPM
Python shiny app for deployment of AI CADe algorithm
Prediction
The predicted “fluid score” probability of the CNN is an indicator of how likely the OCT scan, along with its segmented fluid mask, contains fluid, with a cut-off score of 0.380 providing a threshold that when crossed, signals the need for intervention. Total fluid levels in the B-scan are also predicted and shown in mm2, as well as providing a visual representation of the segmented fluid area. The promising performance of our network signals potential substantial improvement with a larger dataset with varied pathology. This CADe system is also more informative than the current modality inbuilt into OCT machines, which can only provide a reading for layer thickness. Within the context of PPPM, it would be advantageous to strive to incorporate this new modality into current clinical workflows due to its potential for providing a more personalized and quantifiable approach for retinal fluid conditions, but at the same time requires further research into its feasibility and reliability.
Prevention
Within the context of prevention, we can prevent unnecessary treatment based solely on the thickness of the retina (as current guidelines are based only on thickness [7]). In addition, the “Eye health log” tab in the CADe app can also alert the patient and ophthalmologist of any dangerous spike in total fluid levels thus requiring intervention (e.g., via anti-VEGF), therefore preventing further deterioration of the patient’s condition—thereby preventing further progression. The segmentation visualization can help to highlight missed areas as well, especially for IRF, which appears as round, dark cystoid spaces within the retinal layers of varied size, with persistent cystoid spaces representing an irreversible degenerative nature [9], further underscoring the advantage of detecting or highlighting small abnormalities in the retina via segmentation and measuring its trend or persistence over time. The CADe system could potentially provide an indication of fluid presence, and a segmentation visualization opens the “black box” of deep learning classification-only models by providing insight into where the model is making its decisions, and any errors if evident.
Personalized medicine
Another advantage of the OCT CADe system for retinal fluid is the ability for ophthalmologists to titrate treatment based on the response, as well as providing a retrospective database or dataset for research on treatment decisions and outcomes. This retrospective data includes “fluid score” as well as quantified fluid levels, location/type of fluid, or any segmentation or classification errors, all accomplished via a computer-aided approach that minimizes human labor to collect and consolidate annotations, saving time, effort, and resources. Monitoring of disease before and after treatment via the database will be possible over time (serial monitoring) as part of a standard of care. Prescription can be stopped if the intervention is not working based on the levels of total quantified fluid over several treatments—thus saving resources and costs both for the healthcare provider and patient. Anti-VEGF injection dosage could be given differently based on the quantified fluid levels and previous response patterns (currently, fixed-dose pre-packaged injections are given [9, 12]). Monthly changes or reductions in fluid levels can be monitored over time by the patient in the medical records system, and telemonitoring with home OCT devices could incorporate this feature as well.
Deployment at several levels can be possible—screening in primary care settings and for people with AMD or DME; referral setups with ophthalmologists in secondary care; a more personalized approach of dose titration; and treatment and retreatment decisions [49] in tertiary care, research into using microgram titrations of anti-VEGF, or research into switching of anti-VEGF treatment regimens [50].
We have bundled our CADe for retinal fluid within a Python Shiny app (https://shiny.rstudio.com/py/). The Python Shiny framework, similar to available open-source data visualization software such as Streamlit (https://streamlit.io/) and Dash (http://dash.plotly.com/), can integrate seamlessly with Python’s data science and scientific analysis technology stack, which will be very beneficial in terms of streamlining technology integration and development resources. This can produce a result containing segmentation of fluid, area of fluid as well as probability of fluid from a 496 × 768 pixels (9 mm) Heidelberg Spectralis OCT Scan. Results are also logged into a database for each user that logs in.
The potential inclusion of OCT scans, especially with AI algorithms, in yearly health check-ups in primary care settings (e.g., private hospitals and GP clinics) could help to facilitate the early detection of fluid conditions such as AMD or DME. Our model could potentially help to provide an early indicator of potential disease and provide early intervention to prevent progression. This could be useful for general practitioners and for screening programs, where a suite of AI tools, such as for retinal fluid, outer retinal layer abnormalities [18], or glaucomatous optic neuropathy (GON) [51] comprising various algorithms can help to provide an indicator of metrics that can act as a decision support tool for the physician and patient, providing a personalized medicine approach that will help to improve patient disease management.
Finally, we add that the applicability of the generalization of OCT scans for both segmentation and classification to different geographic regions or ethnicity shown in this study also suggests that it is possible to pool OCT retinal fluid datasets from different cohorts or ethnicities for model development in OCT fluid identification. This has been supported by another study carried out by our group that showed that a DL algorithm trained on a Korean OCT dataset could be generalized to an American OCT dataset [19].
We introduce a POC CNN-ViT-based segmentation and CNN classification DL network for computer-aided detection (CADe) of retinal fluid, validated on external testing sets for segmentation and classification performance, bundled within a Shiny app that can be used for serial monitoring, in line with a PPPM approach, especially for personalized medicine. Training of CADe systems of larger scale can bolster the potential usability of AI algorithms for fluid detection—and coupled with an efficient and customizable app, can facilitate the ease of monitoring eye health, to provide better clinical outcomes for patients with AMD or DME via a PPPM approach.
Supplementary information
Below is the link to the electronic supplementary material.
Author contribution
T.H.R. conceptualized the study. T.C.Q. developed the algorithms and application, pre-processed images with associated data, and edited all manuscript versions. K.T. and T.H.R. contributed equally to the annotation/labelling of ground truth and sorted image data into their classes for training. H.G.K., S.S.K., H.N., and G.L. provided external validation data for the validation of the deep learning algorithm. S.T. provided comments on the potential operational implementation of the algorithm and app in clinical practice, in the context of PPPM. M.D., R.T., K.T., and T.Y.C. provided comments to improve the overall manuscript.
Funding
This study was funded by SingHealth Duke-NUS AM (AM-NHIC/JMT010/2020/SRDUKAMR20M0).
Data availability
Not applicable.
Code Availability
Not applicable.
Declarations
Ethics approval
This retrospective study was deemed exempt from institutional review board (IRB) review by the SingHealth Centralised Institutional Review Board (CIRB). This study was conducted in adherence to the tenets of the Declaration of Helsinki.
Consent to participate
Written informed consent was obtained from the participants of the original studies involved.
Consent for publication
This article has been approved for publication by the principal investigator (PI) of this study, T.H.R.
Competing interests
T.H.R. was a former scientific adviser and owns stock of Medi Whale. All other authors declare no competing interests.
Footnotes
Gerald Liew and Ching-Yu Cheng are co-last authors.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Ten Cheer Quek and Kengo Takahashi contributed equally to this work.
References
- 1.Mitchell P, et al. Age-related macular degeneration. Lancet. 2018;392(10153):1147–1159. doi: 10.1016/S0140-6736(18)31550-2. [DOI] [PubMed] [Google Scholar]
- 2.Cheung N, Mitchell P, Wong TY. Diabetic retinopathy. Lancet. 2010;376(9735):124–136. doi: 10.1016/S0140-6736(09)62124-3. [DOI] [PubMed] [Google Scholar]
- 3.Daruich A, et al. Central serous chorioretinopathy: Recent findings and new physiopathology hypothesis. Prog Retin Eye Res. 2015;48:82–118. doi: 10.1016/j.preteyeres.2015.05.003. [DOI] [PubMed] [Google Scholar]
- 4.Hayreh SS, Zimmerman MB. Branch retinal vein occlusion: Natural history of visual outcome. JAMA Ophthalmol. 2014;132(1):13–22. doi: 10.1001/jamaophthalmol.2013.5515. [DOI] [PubMed] [Google Scholar]
- 5.Pichi, F. and P. Neri, Complications in uveitis. p. X, 288 p. 89 illus., 73 illus. in color. online resource.
- 6.Huang D, et al. Optical coherence tomography. Science. 1991;254(5035):1178–1181. doi: 10.1126/science.1957169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Schmidt-Erfurth U, et al. AI-based monitoring of retinal fluid in disease activity and under therapy. Prog Retin Eye Res. 2022;86:100972. doi: 10.1016/j.preteyeres.2021.100972. [DOI] [PubMed] [Google Scholar]
- 8.Virgili, G., et al., Optical coherence tomography (OCT) for detection of macular oedema in patients with diabetic retinopathy. Cochrane Database Syst Rev, 2015. 1: p. CD008081. [DOI] [PMC free article] [PubMed]
- 9.Schmidt-Erfurth U, et al. Guidelines for the management of neovascular age-related macular degeneration by the European Society of Retina Specialists (EURETINA) Br J Ophthalmol. 2014;98(9):1144–1167. doi: 10.1136/bjophthalmol-2014-305702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Group CR et al. Ranibizumab and bevacizumab for neovascular age-related macular degeneration. N Engl J Med. 2011;364(20):1897–1908. doi: 10.1056/NEJMoa1102673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Silva R, et al. Treat-and-extend versus monthly regimen in neovascular age-related macular degeneration: Results with ranibizumab from the TREND study. Ophthalmology. 2018;125(1):57–65. doi: 10.1016/j.ophtha.2017.07.014. [DOI] [PubMed] [Google Scholar]
- 12.Flaxel CJ, et al. Age-related macular degeneration preferred practice pattern(R) Ophthalmology. 2020;127(1):P1–P65. doi: 10.1016/j.ophtha.2019.09.024. [DOI] [PubMed] [Google Scholar]
- 13.Keenan TDL, et al. Retinal specialist versus artificial intelligence detection of retinal fluid from OCT: Age-related eye disease study 2: 10-Year follow-on study. Ophthalmology. 2021;128(1):100–109. doi: 10.1016/j.ophtha.2020.06.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Sarhan MH, et al. Machine learning techniques for ophthalmic data processing: A review. IEEE J Biomed Health Inform. 2020;24(12):3338–3350. doi: 10.1109/JBHI.2020.3012134. [DOI] [PubMed] [Google Scholar]
- 15.Bogunovic H, et al. RETOUCH: The retinal OCT fluid detection and segmentation benchmark and challenge. IEEE Trans Med Imaging. 2019;38(8):1858–1874. doi: 10.1109/TMI.2019.2901398. [DOI] [PubMed] [Google Scholar]
- 16.De Fauw J, et al. Clinically applicable deep learning for diagnosis and referral in retinal disease. Nat Med. 2018;24(9):1342–1350. doi: 10.1038/s41591-018-0107-6. [DOI] [PubMed] [Google Scholar]
- 17.Lin, M., et al., Recent advanced deep learning architectures for retinal fluid segmentation on optical coherence tomography images. Sensors (Basel), 2022. 22(8). [DOI] [PMC free article] [PubMed]
- 18.Rim TH, et al. Computer-aided detection and abnormality score for the outer retinal layer in optical coherence tomography. Br J Ophthalmol. 2022;106(9):1301–1307. doi: 10.1136/bjophthalmol-2020-317817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Rim TH, et al. Detection of features associated with neovascular age-related macular degeneration in ethnically distinct data sets by an optical coherence tomography: Trained deep learning algorithm. Br J Ophthalmol. 2021;105(8):1133–1139. doi: 10.1136/bjophthalmol-2020-316984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Golubnitschaja O, et al. Medicine in the early twenty-first century: Paradigm and anticipation - EPMA position paper 2016. EPMA J. 2016;7:23. doi: 10.1186/s13167-016-0072-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Krizhevsky A, Sutskever I, Hinton GE. ImageNet classification with deep convolutional neural networks. Commun ACM. 2017;60(6):84–90. doi: 10.1145/3065386. [DOI] [Google Scholar]
- 22.LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521(7553):436–444. doi: 10.1038/nature14539. [DOI] [PubMed] [Google Scholar]
- 23.Vaswani, A., et al., Attention is all you need, in Proceedings of the 31st International Conference on Neural Information Processing Systems. 2017, Curran Associates Inc.: Long Beach, California, USA. p. 6000–6010.
- 24.Dosovitskiy, A., et al., An image is worth 16x16 words: Transformers for image recognition at scale. 2021.
- 25.Han, K., et al., A survey on visual transformer. ArXiv, 2020. abs/2012.12556.
- 26.Raghu, M., et al. Do vision transformers see like convolutional neural networks? 2021. arXiv:2108.08810.
- 27.Karimi, D., S. Vasylechko, and A. Gholipour Convolution-free medical image segmentation using transformers. 2021. arXiv:2102.13645.
- 28.Chen, J., et al. TransUNet: Transformers make strong encoders for medical image segmentation. 2021. arXiv:2102.04306.
- 29.Luo, X., et al. Semi-supervised medical image segmentation via cross teaching between CNN and transformer. 2021. arXiv:2112.04894.
- 30.Kermany DS, et al. Identifying medical diagnoses and treatable diseases by image-based deep learning. Cell. 2018;172(5):1122–1131 e9. doi: 10.1016/j.cell.2018.02.010. [DOI] [PubMed] [Google Scholar]
- 31.Fuchs P, et al. Artificial intelligence in the management of anti-VEGF treatment: The Vienna fluid monitor in clinical practice. Ophthalmologe. 2022;119(5):520–524. doi: 10.1007/s00347-022-01618-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Yu HJ, et al. Home monitoring of age-related macular degeneration: Utility of the ForeseeHome device for detection of neovascularization. Ophthalmol Retina. 2021;5(4):348–356. doi: 10.1016/j.oret.2020.08.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Liu Y, Holekamp NM, Heier JS. Prospective, longitudinal study: Daily self-imaging with home OCT for neovascular age-related macular degeneration. Ophthalmol Retina. 2022;6(7):575–585. doi: 10.1016/j.oret.2022.02.011. [DOI] [PubMed] [Google Scholar]
- 34.Skelly, A., et al., Treat and extend treatment interval patterns with anti-VEGF therapy in nAMD patients. Vision (Basel), 2019. 3(3). [DOI] [PMC free article] [PubMed]
- 35.Hasler PW, Flammer J. Predictive, preventive and personalised medicine for age-related macular degeneration. EPMA J. 2010;1(2):245–251. doi: 10.1007/s13167-010-0017-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Golubnitschaja O. Time for new guidelines in advanced diabetes care: Paradigm change from delayed interventional approach to predictive, preventive & personalized medicine. EPMA J. 2010;1(1):3–12. doi: 10.1007/s13167-010-0014-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Majithia S, et al. Cohort profile: The Singapore epidemiology of eye diseases study (SEED) Int J Epidemiol. 2021;50(1):41–52. doi: 10.1093/ije/dyaa238. [DOI] [PubMed] [Google Scholar]
- 38.Hassan B, et al. Deep learning based joint segmentation and characterization of multi-class retinal fluid lesions on OCT scans for clinical use in anti-VEGF therapy. Comput Biol Med. 2021;136:104727. doi: 10.1016/j.compbiomed.2021.104727. [DOI] [PubMed] [Google Scholar]
- 39.Mantel I, et al. Automated quantification of pathological fluids in neovascular age-related macular degeneration, and its repeatability using deep learning. Translational Vision Science & Technology. 2021;10(4):17–17. doi: 10.1167/tvst.10.4.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Wilson M, et al. Validation and clinical applicability of whole-volume automated segmentation of optical coherence tomography in retinal disease using deep learning. JAMA Ophthalmol. 2021;139(9):964–973. doi: 10.1001/jamaophthalmol.2021.2273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ranftl, R., A. Bochkovskiy, and V. Koltun Vision transformers for dense prediction. 2021. arXiv:2103.13413.
- 42.Hu, J., L. Shen, and G. Sun. Squeeze-and-excitation networks. in 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition. 2018.
- 43.Lee, Y. and J. Park CenterMask : Real-time anchor-free instance segmentation. 2019. arXiv:1911.06667.
- 44.Wang, S., et al., U-Net using stacked dilated convolutions for medical image segmentation. ArXiv, 2020. abs/2004.03466.
- 45.Oktay, O., et al. Attention U-net: Learning where to look for the pancreas. 2018. arXiv:1804.03999.
- 46.Chakravarthy U, et al. Automated identification of lesion activity in neovascular age-related macular degeneration. Ophthalmology. 2016;123(8):1731–1736. doi: 10.1016/j.ophtha.2016.04.005. [DOI] [PubMed] [Google Scholar]
- 47.Kwon JM, et al. Validation of deep-learning-based triage and acuity score using a large national dataset. PLoS ONE. 2018;13(10):e0205836. doi: 10.1371/journal.pone.0205836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Alryalat, S.A., et al., Deep learning prediction of response to anti-VEGF among diabetic macular edema patients: Treatment response analyzer system (TRAS). Diagnostics (Basel), 2022. 12(2). [DOI] [PMC free article] [PubMed]
- 49.Lanzetta P, Loewenstein A, Vision Academy Steering Fundamental principles of an anti-VEGF treatment regimen: Optimal application of intravitreal anti-vascular endothelial growth factor therapy of macular diseases. Graefes Arch Clin Exp Ophthalmol. 2017;255(7):1259–1273. doi: 10.1007/s00417-017-3647-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Granstam E, et al. Switching anti-VEGF agent for wet AMD: evaluation of impact on visual acuity, treatment frequency and retinal morphology in a real-world clinical setting. Graefes Arch Clin Exp Ophthalmol. 2021;259(8):2085–2093. doi: 10.1007/s00417-020-05059-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Ran AR, et al. Three-dimensional multi-task deep learning model to detect glaucomatous optic neuropathy and myopic features from optical coherence tomography scans: A retrospective multi-centre study. Front Med (Lausanne) 2022;9:860574. doi: 10.3389/fmed.2022.860574. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Not applicable.
Not applicable.