Abstract
Tick-borne zoonotic diseases pose a threat to public health; hence, identifying the pathogenic agents associated with them is critical. The prevalence of Bartonella and Rickettsia in Iran is unknown. This study aimed to detect Rickettsia spp. and Bartonella species in ticks in northeast Iran and conduct phylogenetic analysis on these bacteria. Ticks from the sample bank in the Research Center for Emerging and Re-emerging Diseases were included in this study. The ticks were collected in 2017 and 2018 from domestic animals (sheep, goats, cows, camels, horses, dogs, and donkeys) and rodents in Golestan, Mazandaran, and Guilan provinces. Molecular methods were used to examine the DNA extracted from these samples to detect Rickettsia spp. and Bartonella species. The study examined a total of 3999 ticks. Ixodes ricinus (46.4%), Rhipicephalus turanicus (26.3%), and Rhipicephalus sanguineus (17.1%) were the most prevalent species. Among 638 DNA pools, real-time-PCR detected Rickettsia spp. in 161 (25.2%), mostly belonging to Rh. sanguineus (48.9%) and Rh. turanicus (41.9%). Golestan Province had the highest number of positive pools (29.7%). No positive samples for Bartonella were detected in a 638 pooled samples. Eight distinct Rickettsia species were detected in 65 sequenced samples, the majority of which were R. massiliae (n = 32, 49.2%) and R. sibirica (n = 20, 30.8%). Other species included R. rhipicephali (n = 3), R. aeschlimannii (n = 5), R. helvetica (n = 5), R. asiatica (n = 4), R. monacensis (n = 6), and R. raoultii (n = 1). The research findings may provide helpful information about tick-borne Rickettsiae in Iran and help to clarify the role of these arthropods in maintaining these agents. Rickettsia species were found to be circulating in three Northern provinces; thus, it is recommended that this disease be considered in the differential diagnosis of febrile diseases caused by tick bites and febrile diseases with skin rashes such as Crimean–Congo hemorrhagic fever (CCHF).
Introduction
Zoonotic diseases are a significant cause of infection-related mortality on a global scale and are critical as emerging infectious diseases in humans [1]. These diseases are a significant public health problem and a direct threat to human health, posing a risk of death. Climate change, travel and tourism, migration, animal trade, human factors, and natural factors significantly impact the spread of emerging and re-emerging infections [2].
Ticks are more likely than other blood-feeding arthropods to transmit pathogenic species, affecting humans and livestock [3]. Knowledge of the intricate relationship between ticks and their hosts, the environment, and the pathogens they transmit, is critical to understanding the epidemiology of tick-borne zoonotic diseases [4]. Ticks transmit various zoonotic diseases, including Lyme disease, Ehrlichiosis, and Rocky Mountain Spotted Fever (RMSF) [5]. Tick-borne diseases are a frequent occurrence in both human and veterinary medicine. The number of tick-borne diseases affecting domestic animals and humans has increased in recent years, necessitating additional research into the epidemiology, diagnosis, and ecology of these newly identified diseases [6].
Rickettsia infections transmitted by ticks are known as emerging and zoonotic diseases that affect human and livestock health. The distribution of these diseases worldwide is related to the vector [7]. Rickettsia, an obligate intracellular gram-negative bacterium, is an ancient vector-borne disease. This bacterium is transmitted via arthropods’ bites to animals and humans. Ticks are the primary vectors for this pathogen; however, other vectors, including fleas, lice, mites, and even mosquitoes, may contribute to transmission. These bacteria are classified into more than 30 species, comprising four subgenera: spotted fever, typhus fever, Rickettsia bellii, and Rickettsia canadensis. The typhus group comprises Rickettsia prowazekii and Rickettsia typhi, while the spotted fever group with more than 20 species [8, 9]. Spotted fever group (SFG) is subdivided into two distinct subgroups: Rocky Mountain Spotted Fever (RMSF) and Mediterranean Spotted Fever (MSF). The Mediterranean spotted fever is an endemic disease in the Mediterranean region (Africa, Southern Europe, and India) and is believed to be associated with global warming and increased tick infestation and bites [10]. Rickettsial infections are transmitted to animals via tick bites, and if humans are in the vicinity of tick-infected animals, be bitten by ticks accidentally, are unable to propagate the infection in nature, and are known as dead end host for this pathogen [11].
In humans, the symptoms of this disease are nonspecific and are not diagnosed in time; as a result, necessary treatments are not performed in the majority of cases. Early symptoms include fever, chills, headache, and myalgia, and additional symptoms such as nausea, vomiting, and diarrhea can be seen in the subsequent steps. In untreated patients, mortality rates ranged from 5% to 25% [7]. Eight tick-borne rickettsios including: Rickettsia rickettsii, Rickettsia sibirica, Rickettsia australis, Rickettsia honei, Rickettsia japonica, Rickettsia africae, Rickettsia conorii, and Rickettsia slovaca have been described in a worldwide In recent years, human infections caused by Rickettsia aeschlimannii, Rickettsia helvetica, Rickettsia mongolotimonae, and Rickettsia heilongjiangii have also been reported in various geographical regions [12].
Bartonella is a vector-borne bacterium linked to increased emerging zoonotic infections in humans and animals. The infections range from mild or self-limiting to severe and life-threatening diseases in human. The bacterium is widespread among small mammals and may threaten human health [13]. Bartonella spp. are gram-negative, slow-growing, intracellular pleomorphic bacillus and includes at least 35 species and three subspecies. Numerous animals are considered hosts and reservoirs for this disease, primarily transmitted through flea and lice feces, sandflies, and possibly tick bites [14]. These bacteria cause chronic infections by infecting red blood cells and attacking endothelial cells, CD34+ progenitor cells, and host dendritic cells [15].
Bartonella infects humans and a wide variety of animal species. Bartonella DNA has been detected in ticks suspected of being a source of animal-associated Bartonella infection in humans. Bartonella spp. cause a variety of diseases in humans, including cat-scratch disease (Bartonella henselae), Carrion’s disease (CD) (Bartonella bacilliformis), trench fever (Bartonella quintana), endocarditis (B. quintana and B. henselae), bacillary angiomatosis (B. quintana and B. henselae), and hepatic peliosis (B. henselae) [5, 16]. B. quintana has emerged in homeless individuals who were in poor health [14]. The zoonotic pathogen B. henselae, which causes cat scratch disease, is probably the most prevalent in developed countries and is strongly associated with other syndromes, particularly eye infections and endocarditis. Cats are the main reservoirs; cat fleas (Ctenocephalides felis) are considered the primary vector, but other arthropods, e.g., ticks, may contribute to transmission [16].
Given the threat to public health posed by tick-borne zoonotic diseases, it is critical to study pathogens associated with ticks. There is limited information on the prevalence of Bartonella and Rickettsia in Iran and the prevalence of Bartonella and Rickettsia in human and vector infections. Thus, this study aimed to detect Rickettsia spp. and Bartonella species in ticks in northern Iran and conduct phylogenetic analysis.
Methods
Ethics statement
The Ethics Committee for National Institute for Medical Research Development "NIMAD" approved this study (Ethic Code: IR.NIMAD.REC.1398.373).
Sample collection
Ticks from the Research Center for Emerging and Re-emerging Diseases’ samples bank were used for analysis. The ticks were collected in 2017 and 2018 from domestic animals (sheep, goats, cows, camels, horses, dogs, and donkeys) and rodents in Golestan, Mazandaran, and Guilan provinces in northern Iran. The study area is bounded on the north by the Caspian Sea and the south by the Alborz mountain range. These provinces are covered with forests, mountains, and coasts and, have a temperate climate with three distinct geographical regions including temperate plains, mountainous and semi-arid regions.
Ticks samples were previously identified using morphological keys [17]. After the identification of collected ticks, the ticks were pooled for DNA extraction, based on the same tick’s species, collected locations, hosts, tick sex, and the growth stage of ticks. Finally, the pools of ticks included 1 to 22 ticks based on the above criteria.
DNA extraction from ticks
Pooled ticks were homogenized using liquid nitrogen and sterile PBS, and DNA was extracted using the potassium acetate method. Initially, 500 μl lysis buffer (0.1 M Tris-HCl, 0.05 M EDTA, 0.2 M sucrose, and 0.5% SDS) and 20 μl proteinase K (10 mg/ml) were added to the homogenized samples, and incubated at 56°C for one day. Amounts of 120 μl of 5 M sodium acetate were added to the samples and kept on ice for 10 min. After centrifuging the samples at 9660 RCF for 10 minutes, the supernatant was recovered, and 35 μl of 4 M sodium acetate and 1 ml of pure ethanol were added. The samples were thoroughly mixed and kept on ice for 10 minutes. The supernatant was discarded after centrifuging at 9660 RCF for 20 min, and the precipitates were washed with 500 μl of 70% ethanol, and the remaining alcohol was allowed to dry completely at room temperature. Finally, 200 μl of elution buffer (1 M Tris-HCl, 1 M EDTA) was added to the completely dried precipitates and kept at -20°C until the test [18].
Detection of the Rickettsia genus
Using the TaqMan Real-Time PCR method, DNA extracted from ticks was analyzed for Rickettsia spp. The 20 μl reactions contained 10 μl commercial master mix (RealQ Plus 2x Master Mix Ampliqon, Denmark), 4 μl template DNA, 900 nmol of forward and reverse primers, 200 nmol of a probe (marked with 6-Carboxyfluorescein (6-Fam) fluorescent dye as a reporter dye and TAMRA as a quencher) [19], and sterile distilled water to final volume. R. conorii DNA (Amplirun, Vircell) and distilled water were included in all assays as positive and negative controls, respectively. The amplification was performed in a Corbett 6000 Rotor-Gene system (Corbett, Victoria, Australia) programmed for a 10-min activation at 95°C, followed by 45 cycles at 95°C for 15 sec, and 60°C for 60 sec. The quantitative analysis was performed using the Rotor-Gene Q Series software and reading was taken at the end of each cycle in green at 60°C (Table 1).
Table 1. The primers and the probe used for the detection of Rickettsia spp. & Bartonella spp.
| Genus | Gene target | Sequence (5´ to 3´) | Amplicon size (bp) |
| Rickettsia spp. | Rsp | 5′- CGCAACCCTYATTCTTATTTGC -3′ | 149 |
| 5′- CCTCTGTAAACACCATTGTAGCA -3′ | |||
| 6- FAM-TAAGAAAACTGCCGGTGATAAGCCGGAG–TAMRA | |||
| Bartonella spp. | 16S-23S rRNA | 5′-GGGGAAGGTTTTCCGGTTTATC-3′ | 92 |
| 5′-GAGGACTTGAACCTCCGACC-3′ |
Amplification of gltA gene
According to the result of Real-Time PCR, positive sample for Rickettsia spp. with a CT ≤ 30 were further analyzed to identify the Rickettsia species by amplifying the gltA gene. The 20 μl reactions contained 10 μl of 2X commercial master mix (2x TEMPase Master Mix RED A Ampliqon, Denmark), 4 μl of DNA template, 500 nmol of g1tA-F primer (GCTCTTCTCATCCTATGGCTATTAT), and 500 nmol of g1tA-R primer (CAGGGTCTTCRTGCATTTCTT), and distilled water to the final volume [20]. The amplification program included an initial denaturation at 95°C followed by 40 cycles at 94°C for 30 sec, 58°C for 30 sec, 72°C for 60 sec, and final annealing at 72°C for 7 min.
Phylogenetic analysis
PCR products were electrophoresed on 1% agarose gel, and samples with specific bands (834 bp) were sent for sequencing (Genomin Co., Tehran, Iran). Chromas 2.6.6 software was used to evaluate the sequences. Final nucleotide sequences were compared with those available in the GenBank® database using the Basic Local Alignment Search Tool (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The gltA gene sequences for various Rickettsia species were extracted from the GenBank database and phylogenetically analyzed using MEGA software in conjunction with the sequences obtained from the samples in this study (MEGA version X 10.1). Phylogenetic relationships were inferred using the Neighbor-Joining method based on the Kimura 2-parameter model for Rickettsia spp. Gamma distribution (+G) was used to model evolutionary rate differences among sites selected by the best-fit model. Evolutionary analyses were conducted on 1000 bootstrap replications using the MEGA X software.
Bartonella detection
The 16S-23S rRNA gene was targeted in ticks’ DNA using the SYBR Green Real-time PCR method. The 20 μl reactions contained 10 μl of Plus 2x Master Mix Green Low ROXTM (AmpliQon, Denmark), 700 nmol of the forward primer and reverse primers (Table 1), 4 μl of template DNA, and distilled water to the final volume. The amplification was programmed in a Corbett 6000 Rotor-Gene system (Corbett Victoria, Australia) for an initial denaturation at 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec, 60°C for 20 sec, and 72°C for 20 sec with a melting step in between. Bartonella henselae DNA (Amplirun, Vircell) and distilled water were included in all assays as positive and negative control.
Results
In this study, 3999 ticks were examined by molecular method. After morphological examinations, ticks were pooled according to species, location, sex, and growth stage. There were 1415 male ticks (35.38%), 2542 female ticks (63.56%), and 42 nymphs (1.05%). A total of 1926 ticks were collected from cattle (48.16%), 1458 ticks were collected from sheep (36.45%), 383 ticks were collected from goats (9.57%), 50 ticks were collected from dogs (1.25%), 128 ticks were collected from camels (3.20%), five ticks were collected from horses (0.12%), two ticks were collected from donkeys (0.05%), and 47 ticks were collected from hedgehogs (1.17%). The study identified 11 tick species using morphological keys. They were classified into four genera: Ixodes, Haemaphysalis, Hyalomma, and Rhipicephalus, with the highest percentage belonging to Ixodes ricinus (46.4%), followed by Rhipicephalus turanicus (26.3%) and Rhipicephalus sanguineus (17.1%) (Table 2). The most numerous species in the prepared pools were I. ricinus (n = 259, 40.59%), Rh. sanguineus (n = 145, 22.73%), and Rh. turanicus (n = 131, 20.53%), respectively (Table 2).
Table 2. The population of ticks collected in the studied provinces, and the prevalence of positive tick pools for Rickettsia spp.es in 2017–18.
| Genus | Species | No. of collected ticks in each province | Total N (%) | Number of tested pools | No. of positive pools for Rickettsia sp. (%) | |||
|---|---|---|---|---|---|---|---|---|
| Mazandaran N (%) | Golestan N (%) | Guilan N (%) | Per tick species | Per tick genus (%) | ||||
| Ixodes | I. ricinus | 1805(76.6) | 6(0.41) | 140(74.8) | 1951(46.4) | 259 | 13(5) | 13(5) |
| Haemaphysalis | H. concinna | 6(0.25) | - | 1(0.53) | 7(0.16) | 4 | 0(0) | 1(11.1) |
| H. inermis | 3(0.12) | - | 1(0.53) | 4(0.09) | 4 | 1(25) | ||
| H. punctata | - | - | 1(0.53) | 1(0.02) | 1 | 0 (0) | ||
| Hyalomma | H. anatolicum | - | 40(2.74) | - | 40(1.0) | 10 | 1(10) | 15(25.8) |
| H. marginatum | 56(2/37) | 65(4.46) | 16(8.5) | 137(3.42) | 44 | 13(29.5) | ||
| H. dromedarii | - | 18(1.23) | - | 18(0.45) | 4 | 1(25) | ||
| Rhipicephalus | Rh. bursa | 48(2.03) | 1(0.06) | - | 49(1.22) | 13 | 4(3.07) | 132(42.3) |
| Rh. sanguineus | 343(14.5) | 343(23.5) | 1(0.53) | 687(17.1) | 145 | 71(48.9) | ||
| Rh. turanicus | 61(2.58) | 983(67.5) | 9(4.81) | 1053(26.3) | 131 | 55(41.9) | ||
| Rh. annulatus | 34(1.4) | - | 18(9.6) | 52(1.3) | 23 | 2(8.69) | ||
| Total | 2356(100) | 1456(100) | 187 (100) | 3999(100) | 638 | 161 (25.23) | 161 (25.23) | |
In this study, 638 pools of ticks were prepared for DNA extraction and molecular analysis. Finally, 59.2%(n = 378), 31.6% (n = 202), and 9.09% (n = 58) of the prepared pools belonged to Mazandaran, Golestan, and Guilan provinces, respectively (Table 3).
Rickettsia detection by Real-Time PCR
Of 638 DNA pooled samples tested by the Real-Time PCR, 161 pools (25.2%) were positive Rickettsia. Rhipicephalus genus had the highest percentage of positive pools (42.3%), and also, the highest percentage of positive pools belongs to Rh. sanguineus (48.9%), Rh. turanicus (41.9%), Hyalomma marginatum (29.5%), and Haemaphysalis inermis (25%) respectively(Table 2). Rickettsia infection was highest (42.3%) in the Rhipicephalus genus and lowest (5%) in the Ixodes genus (P<0.001). According to the number of positive pools in each province, 29.7% of Golestan province, 25.9% of Mazandaran province, and 17.5% of Guilan province were positive (Table 3). Rickettsial infection was significantly higher in ticks from Mazandaran and Golestan provinces than in ticks from Guilan (P<0.001). There was no statistically significant difference in Rickettsia infection between ticks from Mazandaran and Golestan provinces (P = 0.33).
Table 3. Prevalence of Rickettsia spp. in ticks based on counties and provinces.
| Counties | provinces | Number of pools tested | The number of positive pools for Rickettsia spp. per genus (%) N |
|---|---|---|---|
| Golestan | Aq Qala | 67 | 16(23.8) |
| Bandar Torkaman | 56 | 11(19.6) | |
| Aliabad | 8 | 3(37.5) | |
| Gorgan | 39 | 19(48.7) | |
| Gomishan | 8 | 1(12.5) | |
| Aliabad-e-Katul | 23 | 10(43.4) | |
| Gonbad Kavus | 1 | 0(0) | |
| Total | 202 | 60(29.7) | |
| Mazandaran | Nur | 62 | 17(27.4) |
| Amol | 78 | 18(23) | |
| Babol | 45 | 13(28.8) | |
| Sari | 71 | 20(28.1) | |
| Qaemshahr | 55 | 22(40) | |
| Mahmudabad | 16 | 2(12.5) | |
| Savadkuh | 31 | 3(9.67) | |
| Unknown | 20 | 3(15) | |
| Total | 378 | 98(25.9) | |
| Guilan | Talesh | 46 | 2(4.34) |
| Masal | 5 | 1(20) | |
| Rudsar | 1 | 0(0) | |
| Lahijan | 6 | 0(0) | |
| Total | 58 | 3(5.17) |
Identification and phylogenetic analysis of Rickettsia species
A total of 65 pool samples positive for Rickettsia were selected for species identification in such a way that the selected samples included different tick species, and from all the studied counties and from different hosts. Also, the Load of Rickettsia DNA (CT ≤30) was considered in sample selection for the phylogeny survey. Based on results of sequences Blast in GenBank® and phylogenetic analysis, eight distinct species were identified from a total of 65 sequenced Rickettsia gltA samples, the majority of which were R. massiliae (49.2%) (n = 32) and R. sibirica (30.8%) (n = 20). Other species of Rickettsia identified in present study included R. rhipicephali (n = 3), R. aeschlimannii (n = 5), R. helvetica (n = 5), R. asiatica (n = 4), R. monacensis (n = 6), and R. raoultii (n = 1) (Table 4 and Fig 1).
Table 4. Distribution of different species of Rickettsia spp. identified in tick species and provinces in this study.
| Tick specie | Province | Host | No. of positive sample typed | Rickettsia spp (N) |
|---|---|---|---|---|
| Rh. turanicus | Golestan | Cattle | 1 | R. massiliae (1) |
| Dog | 3 | R. massiliae (3) | ||
| Goat | 2 | R. massiliae (2) | ||
| Sheep | 4 | R. massiliae (4) | ||
| Mazandaran | Cattle | 1 | R. sibirica (1) | |
| Dog | 4 | R. sibirica (2), R. massiliae (2) | ||
| Goat | 2 | R. sibirica (1), R. massiliae (1) | ||
| Sheep | 7 | R. sibirica (4), R. massiliae (3), R. rhipicephali (1) | ||
| Rh. sanguineous | Golestan | Goat | 1 | R. massiliae (1) |
| Sheep | 4 | R. massiliae (4) | ||
| Mazandaran | Cattle | 2 | R. sibirica (1), R. massiliae (1) | |
| Goat | 8 | R. sibirica (2), R. massiliae (4), R. rhipicephali (2) | ||
| Sheep | 13 | R. sibirica (8), R. massiliae (6), R. aeschlimannii (1) | ||
| Rh. bursa | Mazandaran | ND | 1 | R. sibirica (1) |
| I. ricinus | Guilan | Cattle | 1 | R. monacensis (1) |
| Mazandaran | Cattle | 6 | R. helvetica (5), R. asiatica (4), R. monacensis (5) | |
| H. marginatum | Guilan | Cattle | 1 | R. aeschlimannii (1) |
| Mazandaran | Cattle | 1 | R. aeschlimannii (1) | |
| Goat | 1 | R. aeschlimannii (1) | ||
| Sheep | 2 | R. aeschlimannii (1), R. raoultii (1) | ||
| Total | 65 | R. sibirica (20), R. massiliae (32), R. rhipicephali (3), R. aeschlimannii (5), R. helvetica (5), R. asiatica (4), R. monacensis (6), R. raoultii (1) | ||
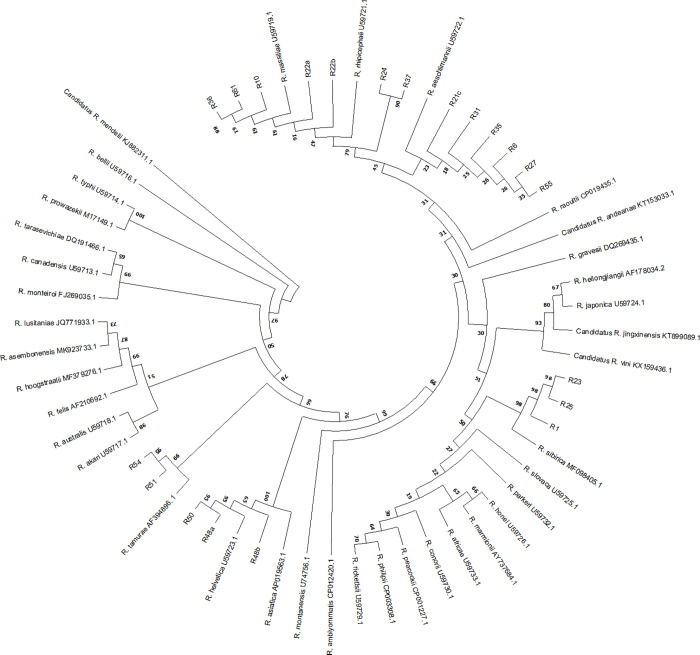
Fig 1. Phylogenetic analysis based on Rickettsia gltA gene sequencing and Neighbor-Joining method algorithm (Kimura 2-parameter model).
The test was performed with bootstrap (1000 repetitions) by MEGA X 10.1 software. Sample ID included R_number were the studied samples in this study.
R. massiliae infection was detected in ticks (Rh. turanicus and Rh. sanguineous)from different hosts (cattle, sheep, goats, and dogs) in Mazandaran and Golestan provinces. This species was identified in all Rh. turanicus ticks collected in Golestan province. Additionally, R. massiliae infection was detected in Rh. sanguineus ticks isolated from dogs and sheep in Golestan province and ticks isolated from dogs, sheep, and goats in Mazandaran province. Thirty samples (99.75%) of R. massiliae detected in this study were identical in gltA gene sequence, and matched 100% to the reference strain R. massiliae MTU5 (CP000683.1) in BLAST analysis. Two samples (R33 and R22a) were identical (100% and 99.87%, respectively) to the Genbank reference strain R. massiliae AZT80 (CP003319.1) and R. massiliae MTU5 (CP000683.1), respectively. No R. massiliae DNA was detected in R. bursa, I. ricinus, and H. marginatum ticks.
Rh. sanguineus, Rh. turanicus, and Rh. bursa ticks showed R. sibirica infection in Mazandaran province. According to gltA gene blast analysis in the Genbank, the majority (80%) of R. sibirica sequencesgenerated in the study were identical to R. sibirica strain (MF098405.1), and three R. sibirica identified (15%) also had 99.78% similarity with the above reference strain. Furthermore, a sample was matched 100% with another R. sibirica strain (acc. no. MF098404.1). There were no positive samples for R. sibirica in I. ricinus or H. marginatum ticks. Moreover, there were no positive samples for R. sibirica in ticks from Guilan and Golestan provinces.
Only H. marginatum and Rh. sanguineus ticks in the Mazandaran and Guilan provinces were infected with R. aeschlimannii, the most frequently isolated Rickettsia from H. marginatum ticks collected from cattle (Mazandaran, Guilan), sheep (Mazandaran), and goats (Mazandaran). Based on the gltA gene blast in the GenBank, two R. aeschlimannii detected in the present study (R6 and R21c) had a 99.88% similarity to a R. aeschlimannii strain (KY411135.1) belonging to the H. marginatum, and Rh. sanguineus ticks in Mazandaran province. Also three samples R31 (Mazandaran), R35 (Mazandaran), and R55 (Guilan) which were detected in H. marginatum had a 99.78% similarity to a R. aeschlimannii strain (KU961540.1), 99.78% similarity to R. aeschlimannii (HQ335513.1), and a 100% similarity to R. aeschlimannii (MH932014.1) in the GenBank for gltA gene sequence blast.
In Mazandaran province, R. helvetica was detected only in I. ricinus ticks isolated from cattle. Four R. helvetica samples (R48a, R49a, R52a, and R53a) were 100% identical to the R. helvetica sequence (KU310588.1) in the GenBank, and a sample (R50) was 99.88% identical to the recorded sequence in GanBanke database. Additionally, no infection with R. helvetica was detected in ticks collected from the other provinces.
R. asiatica infection was detected in four samples of I. ricinus isolated from cattle in Mazandaran province. When the gltA gene sequences of all samples were compared to the sequence of R. asiatica (AP019563.1) in the GenBank, 99.74% similarity was observed. Other tick species collected from other provinces were not infected with R. asiatica.
Furthermore, R. monacensis was isolated from I. ricinus ticks collected from cattle in Mazandaran and Guilan provinces. All five R. monacensis samples detected in the study were identical to the R. monacensis (LN794217.1) reference strains.
Only in Mazandaran province were R. rhipicephali infections observed in Rh. sanguineus (isolated from goats) and Rh. turanicus (isolated from sheep). When the gltA gene sequences of samples R22b, R24, and R37 were compared, they matched the R. rhipicephali (CP001313.1) reference strain in the GenBank by 97.87%, 99.5%, and 99.48%, respectively.
Moreover, we had a positive sample for R. raoultii in a H. marginatum tick isolated from sheep in Mazandaran province. The sample’s gltA gene sequence was 99.83% identical to the sequence of R. raoultii (MT178338.1) in the GenBank.
Five samples contained three different Rickettsia species, four of which were I. ricinus isolates from cattle in Mazandaran province co-infected with R. helvetica, R. asiatica, and R. monacensis. Infections with R. sibirica, R. massiliae, and R. aeschlimannii were detected in a sample of Rh. sanguineus ticks isolated from sheep in Mazandaran province. Furthermore, an Rh. turanicus isolate from sheep in Mazandaran province was found to be co-infected with R. massiliae and R. sibirica.
Bartonella detection
Based on the results of Real-time PCR, no positive samples for Bartonella spp. were detected among 638 tick pools.
Discussion
Zoonotic diseases are a significant threat to public health. Here, we examined Rickettsia and Bartonella pathogens in ticks from three northern Iranian provinces using Real-time PCR to better understand their epidemiology and possible transmission via vectors. According to the results, Rickettsia spp. was detected in 161 of 638 pools (25.2%).
Rickettsial pathogens have received significant attention over the last 25 years, as several tick-borne species previously considered non-pathogenic are now associated with human infections [21]. There are few studies for detection of Rickettsia spp on ticks in Iran. However, numerous studies have been conducted in neighboring countries of Iran on this pathogen as an infectious and high-risk agent for public health [22]. According to the results of gltA gene sequencing on 65 positive samples, the majority of species were identified as belonging to R. massiliae (49.2%) and R. sibirica (30.8%). R. massiliae was initially isolated from the Rhipicephalus genus, and the majority of reports in France, Greece, and Spain have been associated with Rh. turanicus and Rh. sanguineus [23]. The first human infection with this species was reported in 2005 in Italy. In humans, R. massiliae causes a disease similar to MSF [24].
R. massiliae was detected in ticks Rh. turanicus and Rh. sanguineus isolated from cattle, sheep, goats, and dogs in the provinces of Golestan, Mazandaran, and Guilan. In Palestine, a study was conducted to detect Rickettsial SFG in 867 ticks collected from various hosts. In this study, 148 samples were identified as positive for Rickettsia infection R. massiliae was the most frequently identified Rickettsia in this study. Twenty-eight ticks were positive for this species, with the most positive samples belonging to Rh. turanicus. Rh. sanguineus ticks were collected from dogs and sheep, corroborating the current study [25].
In 1991, R. sibirica was isolated for the first time from H. asiaticum ticks in Mongolia. This pathogen was subsequently isolated from members of the genus Hyalomma, particularly H. truncatum and H. excavatum, worldwide. The first human infection with R. sibirica was reported in 1996 in France, and the patient presented with a mild illness that included fever, skin rashes, and scarring R. sibirica was identified in Rh. sanguineus, Rh. turanicus, and Rh. bursa, but not in any of the Hyalomma genera in this study [26]. R. aeschlimannii was initially isolated in 1997 from Hy. marginatum ticks on cattle in Hungary. Furthermore, this species infects H. marginatum in Niger, Zimbabwe, and Mali. In humans, R. aeschlimannii causes a febrile illness similar to MSF, and the first case was diagnosed in a tourist to Hungary in 2000 [27]. As with the previous studies, most positive samples for R. aeschlimannii were detected in H. marginatum ticks isolated from cattle, sheep, and goats. Moreover, it was detected in the Rh. sanguineus ticks isolated from sheep in Mazandaran.
According to a study in Turkey, from 322 ticks were isolated from humans Rickettsia spp. was detected in 100 samples, and following species were identified: R. aeschlimannii, R. slovaca, R. raoultii, R. hoogstraalii, R. sibirica, and R. monacensis. R. aeschlimannii was the most frequently detected species (19.5%), in Hy. marginatum ticks. After R. massiliae, the most positive samples in present study belonged to R. sibirica, but in Turkey, only 0.31% of samples were infected with R. sibrica [28].
R. monacensis, a member of the spotted fever group, was isolated for the first time using molecular methods from I. ricinus ticks in Germany. It has also been isolated from I. ricinus ticks in Hungary and Portugal. The pathogenesis of R. monacensis is unknown, but human infections with this species have been reported in Spain, Italy, and the Netherlands [29]. In this study, R. monacensis was also detected in I. ricinus ticks in Mazandaran.
In a study conducted in Turkey, 25 out of 1019 ticks collected from various country regions were found to be positive for Rickettsia infection. Four of the 25 samples were reported positive for R. monacensis, only in I. ricinus ticks, and was consistent with the findings of this study. Additionally, R. raoultii was detected in Dermacentor marginatus ticks in the study in Turkey [30], but only positive samples for R. raoultii were detected in H. marginatum isolated from sheep in Mazandaran province. R. raoultii was first discovered in 1999 in Rhipicephalus pomilius and Dermacentor nuttalli ticks in the former Soviet Union and was later named R. raoultii in 2008 as a member of the Rickettsia family. Moreover, this pathogen was identified in D. marginatum isolated from a patient in France, demonstrating the pathogen’s pathogenicity in humans [31].
R. helvetica, a member of the spotted fever group, was first detected in many European and Asian countries from I. ricinus ticks. Although this species is generally considered non-pathogenic, mild to severe disease cases, have been reported in humans [32]. R. helvetica was isolated from I. ricinus ticks collected from cattle in this study. in Turkey, 69 of 167 ticks isolated from humans hospitalized across the country tested positive for Rickettsia infection. According to the findings, this study found that R. monacensis (70%) was the most frequently detected species in I. ricinus ticks. Moreover, 2 (3%) I. ricinus ticks tested positive for R. helvetica, consistent with our findings in present study [33].
R. rhipicephali was isolated for the first time from Rh. sanguineus ticks in Mississippi but has not been associated with human infections [34]. R. rhipicephali was also found in Rh. sanguineus and Rh. turanicus ticks in this study. R. asiatica, originally designated Rickettsia IO-1T, was isolated from Ixodes ticks in Japan in 1993 [35]. R. asiatica was detected in I. ricinus ticks isolated from cattle in this study. The R. rhipicephali and R. asiatica species appeared to have not been isolated in neighboring countries and were described for the first time in this study.
In this study, the collected ticks were identified using only morphological keys. It is recommended to do the molecular identification of ticks as a confirmatory in future studies. Another limitation was that the main focus of the current study was on the detection of Bartonella spp that they are pathogenic in humans and animals. Therefore, it is possible that we could not detect some non-pathogenic Bartonella.
Conclusion
According to the result of our study, Rickettsia species were found to be circulating in three Northern provinces. Based on data available in the world, a number of these identified pathogens in this study are capable of causing disease in humans. This is a cautionary tale for the health care system. As a result, it is recommended that these diseases be considered in the differential diagnosis of febrile diseases caused by tick bites, skin rashes, or Crimean–Congo hemorrhagic fever (CCHF) and that the healthcare system be educated about the disease’s critical importance and symptoms.
Acknowledgments
We would also express our gratitude to Dr. Ahmad Mahmoudi, Hamed Hanifi, Amir Hesam Neamati, Alireza Mordadi, Ali Mohammadi, and Seyyed Adel Hosseini from Department of Epidemiology and Biostatistics of Pasteur Institute of Iran, who helped us in tick’s collection and some of the laboratory process.
Data Availability
All relevant data are within the paper.
Funding Statement
The study was supported by a grant from the National Institute for Medical Research Development (NIMAD) (grant no: 987978). Also supported by Pasteur Institute of Iran (grant no: 1744) and Prof. Mostafavi received the fund.
References
- 1.Slingenbergh J., et al., Ecological sources of zoonotic diseases. Revue scientifique et technique-Office international des épizooties, 2004. 23(2): p. 467–484. doi: 10.20506/rst.23.2.1492 [DOI] [PubMed] [Google Scholar]
- 2.Rahman M., et al., Zoonotic diseases: etiology, impact, and control. Microorganisms, 2020. 8(9): p. 1405. doi: 10.3390/microorganisms8091405 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Durden L.A., Taxonomy, host associations, life cycles and vectorial importance of ticks parasitizing small mammals, in Micromammals and Macroparasites. 2006, Springer. p. 91–102. [Google Scholar]
- 4.Pfäffle M., et al., The ecology of tick-borne diseases. International journal for parasitology, 2013. 43(12–13): p. 1059–1077. [DOI] [PubMed] [Google Scholar]
- 5.Chang C., et al., Molecular evidence of Bartonella spp. in questing adult Ixodes pacificus ticks in California. Journal of Clinical Microbiology, 2001. 39(4): p. 1221–1226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dantas-Torres F., Chomel B.B., and Otranto D., Ticks and tick-borne diseases: a One Health perspective. Trends in parasitology, 2012. 28(10): p. 437–446. [DOI] [PubMed] [Google Scholar]
- 7.Sosa-Gutiérrez C.G., et al., Tick-borne rickettsial pathogens in rodents from Mexico. Journal of Biomedical Science and Engineering, 2014. 7(11): p. 884. [Google Scholar]
- 8.Blanda V., et al., New real-time PCRs to differentiate Rickettsia spp. and Rickettsia conorii. Molecules, 2020. 25(19): p. 4431. doi: 10.3390/molecules25194431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Pacheco R.C., et al., Rickettsial infection in ticks (Acari: Ixodidae) collected on birds in southern Brazil. Journal of medical entomology, 2012. 49(3): p. 710–716. doi: 10.1603/me11217 [DOI] [PubMed] [Google Scholar]
- 10.Parola P., Paddock C.D., and Raoult D., Tick-borne rickettsioses around the world: emerging diseases challenging old concepts. Clinical microbiology reviews, 2005. 18(4): p. 719–756. doi: 10.1128/CMR.18.4.719-756.2005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Kuo C.-C., et al., High prevalence of Rickettsia spp. infections in small mammals in Taiwan. Vector-Borne and Zoonotic Diseases, 2015. 15(1): p. 13–20. doi: 10.1089/vbz.2014.1584 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Parola P., et al., Detection of Ehrlichia spp., Anaplasma spp., Rickettsia spp., and other eubacteria in ticks from the Thai-Myanmar border and Vietnam. Journal of clinical microbiology, 2003. 41(4): p. 1600–1608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gundi V.A., et al., Bartonella spp. in rats and zoonoses, Los Angeles, California, USA. Emerging infectious diseases, 2012. 18(4): p. 631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Oteo J.A., et al., Prevalence of Bartonella spp. by culture, PCR and serology, in veterinary personnel from Spain. Parasites & vectors, 2017. 10(1): p. 1–9. doi: 10.1186/s13071-017-2483-z [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Billeter S., et al., Vector transmission of Bartonella species with emphasis on the potential for tick transmission. Medical and veterinary entomology, 2008. 22(1): p. 1–15. [DOI] [PubMed] [Google Scholar]
- 16.Cotté V., et al., Transmission of Bartonella henselae by Ixodes ricinus. Emerging infectious diseases, 2008. 14(7): p. 1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Walker A.R., Ticks of domestic animals in Africa: a guide to identification of species. 2003: Bioscience Reports Edinburgh. [Google Scholar]
- 18.Rodríguez I., et al., An alternative and rapid method for the extraction of nucleic acids from ixodid ticks by potassium acetate procedure. Brazilian Archives of Biology and Technology, 2014. 57(4): p. 542–547. [Google Scholar]
- 19.Giulieri S., et al., Development of a duplex real time PCR for the detection of Rickettsia spp. and typhus group rickettsia in clinical samples. FEMS Immunology & Medical Microbiology, 2012. 64(1): p. 92–97. [DOI] [PubMed] [Google Scholar]
- 20.Labruna M.B., et al., Molecular evidence for a spotted fever group Rickettsia species in the tick Amblyomma longirostre in Brazil. Journal of Medical Entomology, 2004. 41(3): p. 533–537. [DOI] [PubMed] [Google Scholar]
- 21.Parola P., et al., Update on tick-borne rickettsioses around the world: a geographic approach. Clinical microbiology reviews, 2013. 26(4): p. 657–702. doi: 10.1128/CMR.00032-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Khamesipour F., et al., Tick-borne zoonoses in the Order Rickettsiales and Legionellales in Iran: A systematic review. PLoS neglected tropical diseases, 2018. 12(9): p. e0006722. doi: 10.1371/journal.pntd.0006722 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Cardeñosa N., Segura F., and Raoult D., Serosurvey among Mediterranean spotted fever patients of a new spotted fever group rickettsial strain (Bar29). European journal of epidemiology, 2003. 18(4): p. 351–354. doi: 10.1023/a:1023654400796 [DOI] [PubMed] [Google Scholar]
- 24.Vitale G., et al., Rickettsia massiliae human isolation. Emerging infectious diseases, 2006. 12(1): p. 174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ereqat S., et al., Molecular detection and identification of spotted fever group rickettsiae in ticks collected from the West Bank, Palestinian territories. PLoS neglected tropical diseases, 2016. 10(1): p. e0004348. doi: 10.1371/journal.pntd.0004348 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.De Sousa R., et al., Rickettsia sibirica isolation from a patient and detection in ticks, Portugal. Emerging infectious diseases, 2006. 12(7): p. 1103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Raoult D., et al., First documented human Rickettsia aeschlimannii infection. Emerging infectious diseases, 2002. 8(7): p. 748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Karasartova D., et al., Bacterial and protozoal pathogens found in ticks collected from humans in Corum province of Turkey. PLoS neglected tropical diseases, 2018. 12(4): p. e0006395. doi: 10.1371/journal.pntd.0006395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kim Y.S., et al., First isolation of Rickettsia monacensis from a patient in South Korea. Microbiology and immunology, 2017. 61(7): p. 258–263. [DOI] [PubMed] [Google Scholar]
- 30.Orkun Ö., et al., Turkey tick news: A molecular investigation into the presence of tick-borne pathogens in host-seeking ticks in Anatolia; Initial evidence of putative vectors and pathogens, and footsteps of a secretly rising vector tick, Haemaphysalis parva. Ticks and tick-borne diseases, 2020. 11(3): p. 101373. doi: 10.1016/j.ttbdis.2020.101373 [DOI] [PubMed] [Google Scholar]
- 31.Jia N., et al., Human infections with Rickettsia raoultii, China. Emerging infectious diseases, 2014. 20(5): p. 866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Nilsson K., Elfving K., and Påhlson C., Rickettsia helvetica in patient with meningitis, Sweden, 2006. Emerging infectious diseases, 2010. 16(3): p. 490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Gargili A., et al., Rickettsia species in ticks removed from humans in Istanbul, Turkey. Vector-Borne and zoonotic diseases, 2012. 12(11): p. 938–941. doi: 10.1089/vbz.2012.0996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Labruna M.B., et al., Isolation of Rickettsia rhipicephali and Rickettsia bellii from Haemaphysalis juxtakochi ticks in the state of São Paulo, Brazil. Applied and Environmental Microbiology, 2007. 73(3): p. 869–873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Thu M.J., et al., Complete Genome Sequence of Rickettsia asiatica Strain Maytaro1284, a Member of Spotted Fever Group Rickettsiae Isolated from an Ixodes ovatus Tick in Japan. Microbiology resource announcements, 2019. 8(37): p. e00886–19. doi: 10.1128/MRA.00886-19 [DOI] [PMC free article] [PubMed] [Google Scholar]