Abstract
To adapt to changes in environmental cues, Pseudomonas aeruginosa produces an array of virulence factors to survive the host immune responses during infection. Metabolic products contribute to bacterial virulence; however, only a limited number of these signaling receptors have been explored in detail for their ability to modulate virulence in bacteria. Here, we characterize the metabolic pathway of 2-methylcitrate cycle in P. aeruginosa and unveil that PmiR served as a receptor of 2-methylisocitrate (MIC) to govern bacterial virulence. Crystallographic studies and structural-guided mutagenesis uncovered several residues crucial for PmiR’s allosteric activation by MIC. We also demonstrated that PmiR directly repressed the pqs quorum-sensing system and subsequently inhibited pyocyanin production. Moreover, mutation of pmiR reduces bacterial survival in a mouse model of acute pneumonia infection. Collectively, this study identified P. aeruginosa PmiR as an important metabolic sensor for regulating expression of bacterial virulence genes to adapt to the harsh environments.
<span style="border: 0px; font-variant-numeric: inherit; font-variant-east-asian: inherit; font-stretch: inherit; font-size: 12pt; line-height: inherit; font-family: Calibri, Arial, Helvetica, sans-serif; margin: 0px; padding: 0px; vertical-align: baseline; color: rgb(0, 0, 0);">PmiR is a previously unidentified receptor of MIC that alters </span><i style="color: rgb(0, 0, 0); font-family: inherit; font-size: 12pt; font-variant-ligatures: inherit; font-variant-caps: inherit;">Pseudomonas aeruginosa </italic><span style="border: 0px; font-variant-numeric: inherit; font-variant-east-asian: inherit; font-stretch: inherit; font-size: 12pt; line-height: inherit; font-family: Calibri, Arial, Helvetica, sans-serif; margin: 0px; padding: 0px; vertical-align: baseline; color: rgb(0, 0, 0);">virulence and regulates the pqs system.</span>
INTRODUCTION
Propionate is one of the most abundant short-chain fatty acids and represents a rich potential source of carbon for the organisms to use for survival and metabolic processes (1). However, high concentrations of propionate or propionyl–coenzyme A (CoA) are lethally toxic to cells if they accumulate excessively (2). The 2-methylcitrate cycle (2-MCC) is a widely distributed carbon metabolic pathway playing a crucial role in maintaining a balance of propionate and propionyl-CoA in bacteria (3). To avoid excessive accumulation of propionate and propionyl-CoA, bacteria have evolved strategies to efficiently convert propionyl-CoA into pyruvate via the 2-MCC, which serves as a part of an extended tricarboxylic acid (TCA) cycle (4). Propionyl-CoA and oxaloacetate are converted to 2-methylcitrate by 2-methylcitrate synthase PrpC. This is followed by an isomerization event via dehydration and rehydration steps controlled by the 2-methylcitrate dehydratase PrpD and aconitase AcnB synthesizing 2-methyl-cis-aconitate and 2-methylisocitrate (MIC), respectively. Last, MIC lyase PrpB cleaves MIC into pyruvate and succinate, which serve as carbon and energy sources for enzymes in the central metabolism (5, 6). Although this pathway has been well studied in some bacteria, the role of 2-MCC in other bacterial virulence, such as Pseudomonas aeruginosa, remains largely unknown.
P. aeruginosa is an opportunistic pathogen that causes diseases in several clinic cases, such as pneumonia and urinary tract infections, particularly in burn patients and immunocompromised individuals (7). This versatility is thought to result from the strong capacity of this bacterium to adapt rapidly to environmental changes and to regulate the expression of a number of virulence factors in a coordinated manner (8). The classic model for an epidemiological triangle (host-pathogen-environment) posits that the progress and severity of an infection is influenced by a combination of factors, including the immune system of the host, virulence of the pathogen, and the environmental conditions governing the infection (9). Illustrating the role of each of these factors and their interactions will allow us to better understand the progression of infection. Similar to other bacteria, P. aeruginosa must adapt to changes in its environment by sensing environmental signals and regulating the expression of virulence factors (10). Some external signals, such as bacterial density, pH levels, ion levels, oxygen levels, and other cues, are mainly sensed and responded to directly by two-component systems (TCSs) (11, 12), which play an essential role in controlling virulence factors in response to environmental stimuli. However, other external signals that regulate the expression of virulence factors need a process of intracellular signal transduction before they are sensed by the transcriptional regulatory proteins. For example, the LysR-type transcriptional regulator, HinK, is required to evoke responses to histamine in P. aeruginosa. The histamine metabolite of imidazole 4–acetic acid induces HinK to activate transcription of the HinK regulon that predominantly consists of genes involved in histamine uptake and metabolism, iron acquisition, Pseudomonas quinolone signal (PQS) biosynthesis, and virulence (13).
To survive in the host, P. aeruginosa has evolved multiple means of scavenging nutrients and adjusting its metabolism to maintain growth while also coping with stress (14). Furthermore, P. aeruginosa secretes an array of surface-associated and extracellular virulence factors to cause disease in the host and interfere with host defense (15). The diverse arsenal of virulence factors includes pyocyanin, exotoxin, and protease, which are tightly regulated by the las, rhl, and pqs quorum sensing (QS) system (16, 17). In addition to QS systems, the GntR transcriptional factors are also able to regulate virulence. For example, we have recently reported that the GntR family protein MpaR coordinately regulates anthranilate metabolism and affects P. aeruginosa virulence (18). It is predicted that P. aeruginosa encodes more than 40 GntR family regulators, but only a few of them have been fully characterized. Moreover, the methods by which these regulators sense the metabolic intermediates to modulate the expression of virulence factors remain largely unknown in P. aeruginosa.
Here, we characterized the biosynthesis pathway of 2-MCC and identified the GntR family protein PmiR as an MIC sensor that regulates the expression of genes involved in both metabolism and virulence in P. aeruginosa. The crystal structure of PmiR/MIC complex and site mutagenesis data suggested that PmiR is critical for MIC-mediated expression of virulence in P. aeruginosa. In addition, deletion of pmiR promotes P. aeruginosa virulence in a mouse model, resulting in a worse phenotype. Together, this study points out that PmiR senses MIC to regulate P. aeruginosa pathogenesis.
RESULTS
Identification of PmiR as a regulator of virulence factors in P. aeruginosa
Because the transcriptional factors play critical roles in metabolic homeostasis to modulate virulence, we hypothesized that the GntR family proteins might contribute to the process of P. aeruginosa infection. To test this hypothesis, we generated deletion mutants for all 43 GntR family regulators and determined the pyocyanin production of each mutant. Pyocyanin is a blue redox-active secondary metabolite that is produced by P. aeruginosa and acts as an important virulence factor (19). Notably, the deletion of PA0797 (hereafter named pmiR, pqs regulation, and MIC receptor) exhibited a marked increase in pyocyanin compared to the wild-type PAO1 (WT PAO1) and its complemented strain (Fig. 1A). Bacterial motility as another virulence component is often associated with other phenotypic changes that alter bacterial behavior and virulence toward host (20). Critically, mutations of pmiR increased the bacterial swarming and swimming motility, and the phenotype was rescued by its complemented strain (fig. S1). Collectively, these results indicated that PmiR represses P. aeruginosa virulence.
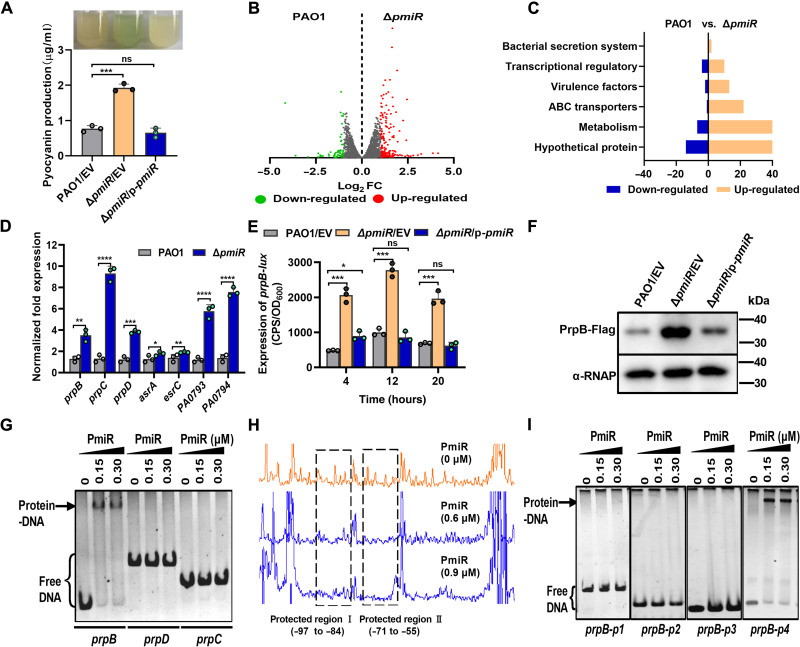
Fig. 1. PmiR controls pyocyanin production and regulates the genes expression of 2-MCC.
(A) Pyocyanin production of the wild-type PAO1 (WT PAO1), ΔpmiR mutant, and the complemented strain was detected in Luria-Bertani (LB) broth for 24 hours. (B) Volcano plot of the DEGs was analyzed between the WT PAO1 and ΔpmiR strain by RNA-seq. Red dots, up-regulated. Green dots, down-regulated. FC, fold change. (C) PmiR regulon categorized by Clusters of Orthologous Gene (COG). (D) qRT-PCR analysis of the indicated genes in WT PAO1 and ΔpmiR strain. ABC, adenosine triphosphate–binding cassette. (E) Expression of prpB was measured in the indicated strains. (A, D, and E) Error bars indicate the means ± SD of three independent experiments. *P <0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. ns, no significance. EV represents the empty vector pAK1900. OD600, optical density at 600 nm. CPS, Counts per second. (F) The expression of PrpB-Flag from the indicated strains was detected by Western blot with an α-Flag antibody. α–RNA polymerase (RNAP) was used as a loading control. (G) EMSA showing that PmiR binds to the promoter region of prpB but not prpC or prpD. Each reaction mixture contains PCR products (2.0 ng/μl). The protein concentrations were indicated above the lane. (H) DNAase I footpringing on the prpB promoter by PmiR showed two regions protected by PmiR. (I) EMSAs showing that PmiR binds to the promoter region of prpB-p4 (−97 to +100, with site I and site II) but not prpB-p1 (−71 to +100, which lacks site I), prpB-p2 (−217 to −84, which lacks site II), and prpB-p3 (−54 to +100, which lacks both site I and site II).
To understand how pmiR may affect gene expression at the transcriptional level, we compared the transcriptomics of WT PAO1 with the ΔpmiR mutant by means of RNA sequencing (RNA-seq). Mutational analyses of pmiR showed differential regulation of 167 genes in WT PAO1 during exponential growth. Among these, 139 genes were up-regulated, and 28 genes were down-regulated, belonging to differentially expressed genes (DEGs) (>2-fold change and P < 0.05) (Fig. 1B and table S1). Most of those genes encode hypothetical proteins with unknown functions. The remaining genes may play an important role in the bacterial system, for example, in metabolic processes, QS, pathogenesis, and regulation of transcription or as adenosine triphosphate–binding cassette transporters (Fig. 1C). To verify the reliability of the RNA-seq data, we randomly selected seven genes, and their expression was analyzed by quantitative reverse transcription polymerase chain reaction (qRT-PCR). Consistent with the RNA-seq result, the expression of these selected genes was differently regulated by pmiR (Fig. 1D).
Characterization and regulation of the gene clusters involved in 2-MCC
Notably, RNA-seq analysis identified the expression of the PA0792 to PA0796 cluster as among the most highly (20- to 60-fold) up-regulated genes (table S1). The database of P. aeruginosa genome showed that PA0793 and PA0794 share the same operon; it is likely that prpB, methylisocitrate lyase, is cotranscribed with pmiR (fig. S2A) (2). Moreover, it is predicted that these clusters encode enzymes that are required for 2-methylcitrate metabolism, which has been well established in other bacteria (6). To validate the metabolic pathway in P. aeruginosa, we expressed and purified the PrpB, PrpC, and PrpD proteins (fig. S2B), which are three key enzymes involved in 2-MCC. Metabolites analysis by high-performance liquid chromatography (HPLC) showed that the enzyme-catalyzed reaction was the same as in other bacteria (fig. S2C). To further assess the role of these genes in 2-methylcitrate metabolism, we generated three mutant strains, including ΔprpB (ΔPA0796), ΔprpC (ΔPA0795), and ΔprpD (ΔPA0792) and measured their growth in M9 medium supplemented with the corresponding substrate as a sole carbon source. Our data showed that the ΔprpB, ΔprpC, and ΔprpD mutants grew like WT PAO1 when cultured with M9 medium with sodium citrate (fig. S3A) but have notable growth defects when sodium propionate was used as the sole carbon source (fig. S3B). We also observed that the ΔprpB mutant grew slowly than WT PAO1 in a medium with MIC or succinate as the sole carbon source (fig. S3C). This growth defect was restored to the WT levels by introducing the corresponding complemented plasmid to these mutants. In addition, similar to pmiR, deletion of prpB, prpC, and prpD exhibited increased pyocyanin production and bacterial motility (fig. S4). Together, these results suggested that PrpB, PrpC, and PrpD play critical roles in 2-MCC and is related to P. aeruginosa virulence.
To further assess this regulation of PA0792-PA0797 clusters by pmiR, we constructed three promoter-lux fusions, including prpB-lux (prpB shares the same promoter with pmiR), prpC-lux, and prpD-lux and measured their activity in a WT PAO1 strain, a pmiR deletion mutant, and its complemented strain (ΔpmiR/p-pmiR). As expected, the promoter activity of these fusions was drastically increased in ΔpmiR mutant compared to the WT PAO1 strain. Introduction of the p-pmiR to ΔpmiR can restore activity to the WT levels (Fig. 1E and fig. S5A). To further confirm this result, we compared the protein level of prpB in WT PAO1 with ΔpmiR and with the complemented strain by Western blot analysis. Consistent with the transcription of prpB, the PrpB level was much higher in ΔpmiR than WT PAO1 and the complemented strain (Fig. 1F).
To determine whether this regulation is direct, electrophoretic mobility shift assays (EMSAs) were performed using the recombinant PmiR protein (fig. S5B) and the intergenic regions of prpB, prpC, and prpD. We found that PmiR efficiently bound only to the promoter region of prpB (Fig. 1G) but not to prpC and prpD. To ascertain the binding sites, we carried out deoxyribonuclease I (DNase I) footprinting assay. We found that PmiR protected prpB promoter region from DNase I digestion at two adjacent sites (site I: −97 to −83 and site II: −71 to −55; relative to the start codon) (Fig. 1H). To determine whether the two motifs were necessary for binding to PmiR, we performed EMSAs again using three truncated DNA fragments, which lacked the first (prpB-p1), the second (prpB-p2), or both of the two binding motifs (prpB-p3) (fig. S2A). Intriguingly, EMSAs showed that PmiR was unable to bind to the three truncated fragments but efficiently bound to the long palindromes (prpB-p4) with the two binding motifs (Fig. 1I). Collectively, these results suggested that PmiR was directly involved in regulation of the 2-methylcitrate pathway.
PmiR controls virulence expression through directly regulating the pqs system
Previous studies have shown that the pyocyanin production is tightly regulated by QS systems (20). As deletion of pmiR exhibited enhanced pyocyanin production (Fig. 1A), we next sought to determine whether the pyocyanin was controlled through regulating the QS system. To verify this hypothesis, we measured the promoter activity of lasI-lux, lasR-lux, rhlI-lux, rhlR-lux, pqsR-lux, and pqsH-lux in the WT PAO1 strain, ΔpmiR mutant, and complemented strain (ΔpmiR/p-pmiR). We found that the activity of pqsR-lux and pqsH-lux was strongly induced in ΔpmiR compared to the WT parent. Complementation of p-pmiR plasmid restored the expression of pqsR and pqsH to WT levels (Fig. 2A). Similarly, the transcription of pqsA was also increased in ΔpmiR (fig. S5C). However, lasI-lux, lasR-lux, rhlI-lux, and rhlR-lux showed no difference among the WT PAO1, ΔpmiR mutant, and the complemented strain (fig. S5, D to G). To further verify whether the increased expression of pqsA, pqsR, and pqsH leads to augmented PQS production, the PQS was extracted from WT PAO1 and the mutant strains and analyzed by thin-layer chromatography (TLC). These data showed that the production of PQS was much higher in the ΔpmiR mutant than that in WT PAO1 and the complemented strain (Fig. 2B).
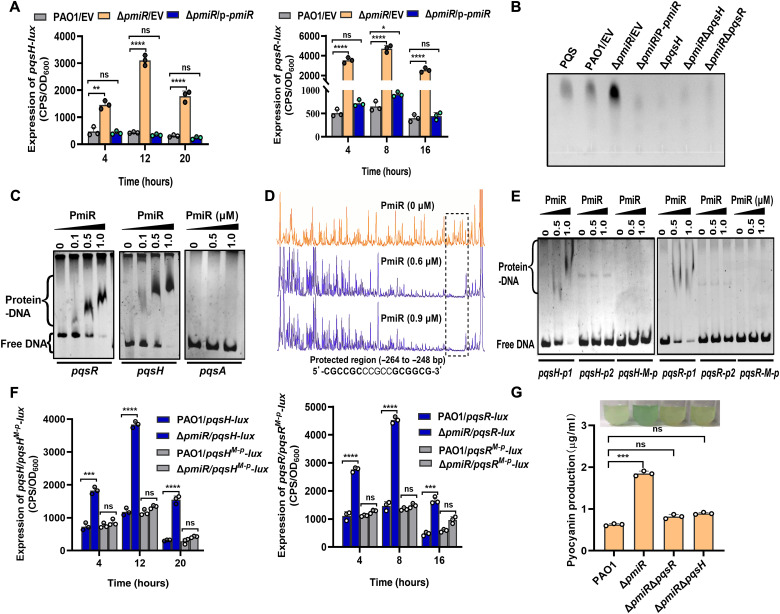
Fig. 2. PmiR controls the production of pyocyanin by directly regulating the pqs system.
(A) Expression of pqsR-lux and pqsH-lux was measured in WT PAO1, ΔpmiR, and the complemented strain. (B) TLC analysis of PQS produced by the indicated strains. PQS is a standard sample control. Data are representative of three independent replicates. (C) EMSAs showing that PmiR binds the promoter region of pqsR and pqsH but not pqsA. Each reaction mixture contains PCR products of pqsR, pqsH, and pqsA (2.0 ng/μl). The protein concentrations were indicated above the lane. (D) Electropherograms show the protection pattern of the pqsH promoter after digestion with DNase I following incubation in the absence and the presence of two different concentrations of PmiR. The PmiR-protected region (relative to the start codon) was indicated by a dotted line. (E) EMSAs showing that PmiR could bind to the truncated promoter region of pqsH-p1 and pqsR-p1 (with the protected region) but not pqsH-p2 and pqsR-p2 (without the protected region) and the mutated pqsH-M-p and pqsR-M-p. Each reaction mixture contains PCR products (2.0 ng/μl). The protein concentrations were indicated above the lane. (F) The promoter activity of pqsR-lux, pqsH-lux, pqsR-M-p, and pqsH-M-p was measured in WT PAO1 and ΔpmiR strain. (G) The production of pyocyanin was measured in PAO1, ΔpmiR, ΔpmiRΔpqsH, and ΔpmiRΔpqsR strains cultured overnight in LB medium. (A, F, and G) Error bars indicate the means ± SD of three independent experiments. Statistical significance was calculated using one-way analysis of variance (ANOVA) Dunnett’s multiple comparison test, * P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. EV represents the empty vector pAK1900.
To probe whether PmiR directly binds to the promoter regions of pqsA, pqsR, and pqsH, we carried out EMSAs using PmiR protein. We found that PmiR bound to the promoter regions of pqsR and pqsH but not pqsA (Fig. 2C). To further determine the function of PmiR on the regulation of pqsH, we searched for potential PmiR-binding sites in the promoter region of pqsH using a dye-based DNase I footprinting analysis. As shown in Fig. 2D, one region of the pqsH promoter was apparently protected against DNase I digestion in the presence of PmiR. The protected region (CGCCGCCCGCCGCGGCG) extended from nucleotides −264 to −248 relative to the pqsH start codon. We also noted that a certain degree of motif (CGCCGGCTCGCTTTCTGCGCGGCG, −552 to −529 from the pqsR translational start codon) was also present in the promoter regions of pqsR. To determine whether the motif was necessary for binding to PmiR, we performed EMSAs again using a pqsH-p1 DNA fragment (−249 to +20 relative to the start codon of pqsH, without the binding motif) and a pqsH-M-p DNA fragment (the motif CGCCGCCCGCCGCGGCG was mutated to CTCCTCCCGCCTCTTCT). Neither pqsH-p1 nor pqsH-M-p was able to bind to PmiR (Fig. 2E). Similarly, PmiR could not bind to the pqsR-p1 (−530 to +1 relative to the start codon of pqsR, without the binding motif) and pqsR-M-p (CGCCGGCTCGCTTTCTGCGCGGCG was mutated to CTCCTTCTCGCTTTCTGCTCTTCT) fragments (Fig. 2E). In addition, the transcriptional activity of both the pqsH-M-p and pqsR-M-p promoters in the ΔpmiR mutant was comparable to the WT parent (Fig. 2F), which contrasted with the observation that the expression of WT pqsR and pqsH was strongly induced in ΔpmiR strain. Collectively, these results suggested that PmiR directly bound to the promoters to repress the transcription of pqsH and pqsR.
As PmiR directly repressed the expression of the pqs system, we proposed that PmiR repressed pyocyanin production by means of regulating the pqs system. To test this hypothesis, the ΔpmiRΔpqsR and ΔpmiRΔpqsH double mutants were generated. In the ΔpmiR mutant, deletion of pqsR or pqsH restored the pyocyanin product to the WT levels (Fig. 2G). Together, these results demonstrated that PmiR controls pyocyanin production by directly repressing the pqs system.
MIC and its analogs appear to be the effector for PmiR
The GntR family proteins often bind DNA as homodimers, and that regulation of DNA binding is dependent on the binding of an effector molecule (21). For example, PrpR senses acyl-CoAs to regulate gene transcription in Mycobacterium tuberculosis (22). As aforementioned, PmiR participates in regulation of 2-MCC (Fig. 1). Deletion of pmiR caused a marked increase in prpB expression, which resulted in the accumulation of pyruvate and succinate and reduction of MIC (fig. S2C). Therefore, we were interested in determining whether the substrate MIC or the products, including pyruvate and succinate of the PrpB, could be a potential effector of PmiR. To address this, we performed EMSAs to evaluate whether these compounds affected PmiR binding to its targets. The addition of 1 mM MIC (Fig. 3A) or 4 mM succinate (fig. S6A) decreased the binding of PmiR to prpB promoter, whereas the addition of 10 mM citrate or pyruvate did not (fig. S6, B and C). In addition, the addition of isocitrate, a structural analog of MIC, was also able to prevent binding of DNA by PmiR (Fig. 3B).
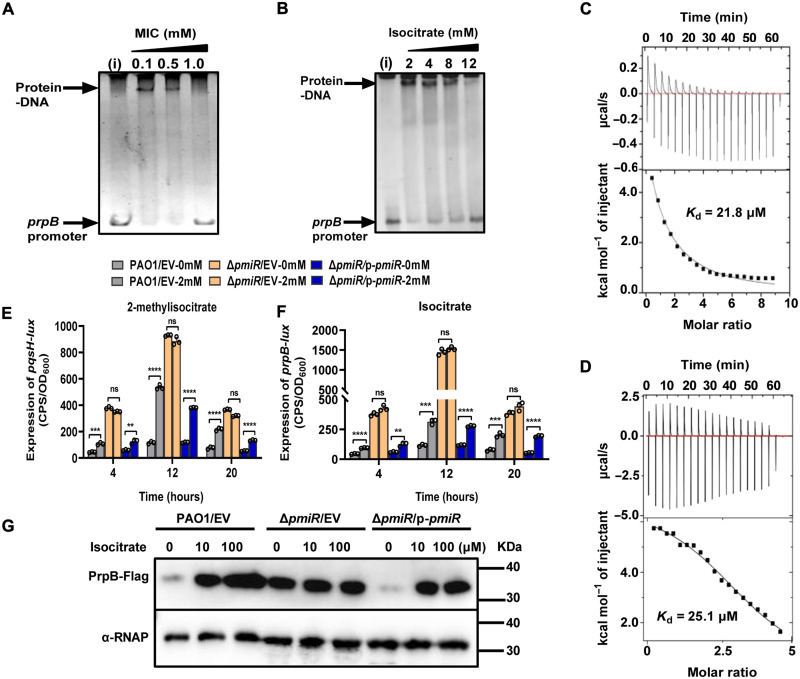
Fig. 3. PmiR senses MIC to regulate the expression of genes involved in virulence factors.
(A and B) EMSA showing the effect of different concentrations of MIC and isocitrate on the DNA binding ability of PmiR. Each reaction mixture contains DNA (2 ng/μl), 0.3 μM PmiR, and the indicated concentrations of MIC and isocitrate. Data are representative of three independent replicates. (C and D) ITC experiments of PmiR toward MIC and isocitrate. The top panel shows heat differences upon injection of MIC (C) or isocitrate (D), and the bottom panel shows integrated heats of injection with the best fit (solid line) to a single binding model using Microcal ORIGIN. (E) Expression of pqsH-lux was tested in WT PAO1, ΔpmiR, and the complemented strain cultured in M9 minimal medium (MM) with or without 2 mM MIC. (F) Expression of prpB-lux was measured in WT PAO1, ΔpmiR, and the complemented strain cultured in M9 MM with or without 2 mM isocitrate. Error bars indicate the means ± SD of three independent experiments. Statistical significance was calculated using one-way ANOVA Dunnett’s multiple comparison test, **P < 0.01; ***P < 0.001; ****P < 0.0001. EV represents the empty vector pAK1900. (G) The protein levels of PrpB-Flag were detected in the indicated strains cultured in M9 medium with or without isocitrate by Western blot analysis. The tagged proteins were detected using a Flag antibody. α-RNAP antibody was used as a loading control. Data are representative of three independent replicates.
Because the MIC and isocitrate can alter the DNA binding affinity of PmiR, we next sought to determine whether these two compounds bind to PmiR. To this end, the interaction between PmiR and MIC or isocitrate was confirmed by isothermal titration calorimetry (ITC), and the binding affinities were compared. We found that PmiR could bind to MIC with a dissociation constant (Kd) of 21.8 μM (Fig. 3C). Similarly, isocitrate also interacts with PmiR with a Kd of 64.1 μM (Fig. 3D). No interaction was observed between PmiR and citrate (fig. S6D), which is consistent with the results from EMSAs.
We reasoned that binding of these compounds to PmiR would affect PmiR-mediated regulation of gene expression. To verify this, we monitored the transcription activity of pqsH-lux in the ΔpmiR mutant in the presence or absence of MIC and compared to WT PAO1. We observed that in WT PAO1 and the complemented strain, MIC strongly induced the activity of pqsH. No significant difference in pqsH expression could be found in the ΔpmiR mutant strain (Fig. 3E). We also examined the activity of prpB-lux in WT PAO1, ΔpmiR, and its complemented strain cultured in M9 medium with or without isocitrate. Similar to pqsH, the activity of prpB was strongly stimulated by isocitrate in WT PAO1 and the complemented strain but not in the ΔpmiR mutant (Fig. 3F). These results were further verified by monitoring the protein level of PrpB using Western blot analysis. In line with the transcription of prpB gene, the PrpB levels were drastically increased in WT PAO1 and the complemented strain by the addition of different concentrations of isocitrate. In contrast, no changes were observed in the ΔpmiR mutant (Fig. 3G). Collectively, these results suggested that MIC and its analog isocitrate were involved in PmiR-mediated regulation of gene expression.
Structural characterization of Zn2+ ion and MIC binding by PmiR
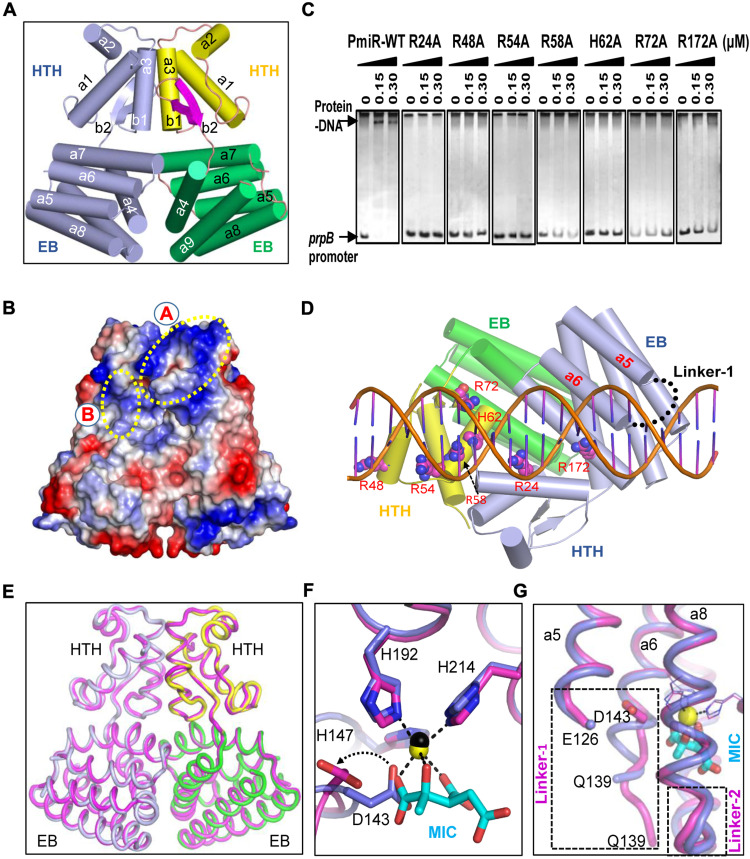
To reveal the structural basis for effector binding by PmiR, we performed extensive crystallization trials. Although no crystal grew for the full-length protein, we successfully solved the structures of one truncated PmiR protein (amino acids 15 to 232) in the absence or presence of MIC. In vitro binding assays confirmed that N-terminal residue truncation had no obvious impacts on DNA binding by PmiR (fig. S7A). ITC analysis showed that the truncated PmiR could also bind to MIC with a Kd value of 62.9 μM (fig. S7B). Both the apo form and the MIC-complexed crystals belong to the I4122 space group (table S2) and contain one PmiR monomer per asymmetric unit. Each PmiR monomer (Fig. 4A and fig. S7C) can be divided into two domains: Helix-Turn-Helix (HTH) domain (amino acids 17 to 81) and effector binding (amino acids 85 to 223) domain. The HTH domain is composed of three α helices (α1 to α3) and two β strands (β1 and β2); the effector binding domain is composed of six α helices (α4 to α9) that are arranged in two layers (Fig. 4A). The orientations between the two domains are mainly stabilized by the hydrophobic residues located at the interface, such as V31, P37, L69, V80, and V81 of the HTH domain and L170, R172, and M173 of the effector binding domain (fig. S7, D and E).
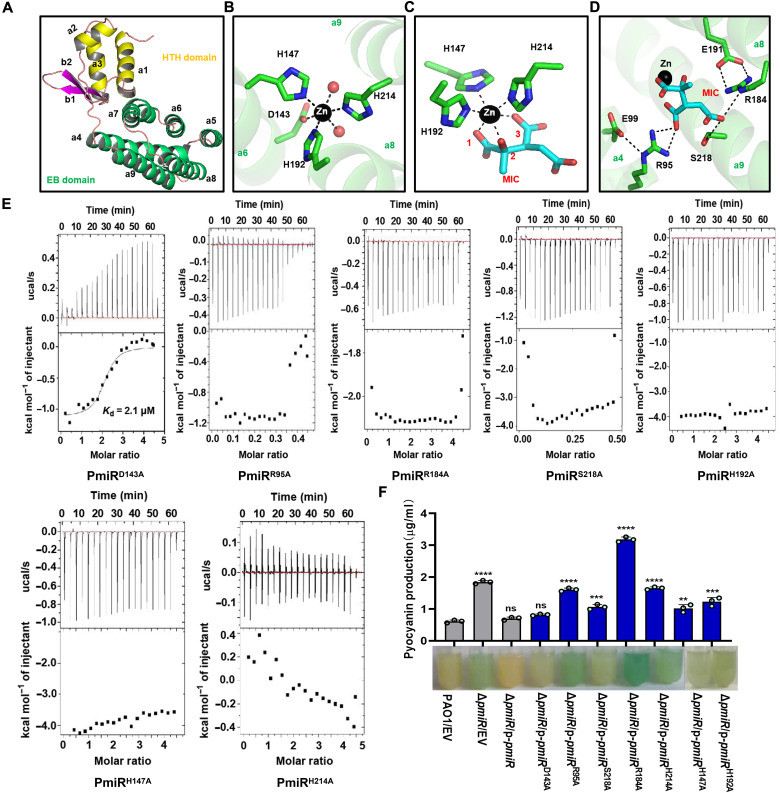
Fig. 4. Crystal structure of PmiR-MIC complex.
(A) The overall fold of PmiR monomer. (B) Zn2+ coordination observed in the apo-form PmiR structure. (C) Zn2+ and MIC interactions observed in the MIC-complexed structure. (D) Interactions between MIC and other residues of PmiR. In (B) to (D), the Zn2+ ion is shown as sphere; MIC, MIC-interacting, and Zn2+-coordinating residues are shown as sticks. (E) ITC analysis showing the impacts of Zn2+ binding and other key residues on MIC binding by PmiR. For each figure, the top panel shows heat differences upon injection of MIC, and the bottom panel shows integrated heats of injection with the best fit (solid line) to a single binding model using Microcal ORIGIN. (F) The production of pyocyanin was measured in WT PAO1, ΔpmiR, and the different complemented strains. Lower photos show strains cultured in LB broth for 24 hours. Data are representative of three independent replicates. Error bars indicate the means ± SD of three independent experiments. **P < 0.01; ***P < 0.001; ****P < 0.0001 compared to WT PAO1 by Student’s t test. EV represents the empty vector pAK1900.
The effector binding domain captured one well-defined Zn2+ ion in both structures (fig. S7, F and G). Zn2+ is copurified with PmiR and is six-coordinated. In the apo-form structure, the Zn2+ ion coordinates with two water molecules and the side chains of D143, H147, H192, and H214 (Fig. 4B). D143 and H147 belong to the α5 helix, whereas H192 and H214 are located in the middle of the α8 and α9 helices, respectively (Fig. 4C). The complex structure was refined at 2.5-Å resolution, which produced clear electron density maps for MIC (fig. S7G). Like H147, H192, and H214, MIC also coordinates with the Zn2+ ion through the carboxylate groups at positions 1 and 3 and the hydroxyl group at position 2. The conformation of MIC is further stabilized by its hydrogen binding (H-bond) interactions with the side chains of R95, R184, and S218 (Fig. 4D). Because of stable H-bond interactions with E99 or E191, the conformations of R95 and R184 are well conserved in the two structures.
The crystal structures led us to further analyze the function of several conserved residues in PmiR in vivo and in vitro. To this end, we constructed and purified several PmiR mutants with either Zn2+-coordinating (D143, H147, H192, and H214) or MIC-binding (R95, R184, and S218) residue substituted with Ala (fig. S7H). We next examined the binding ability of PmiR to MIC by ITC (Fig. 4E). The mutants, including PmiRH147A, PmiRH192A, PmiRH214A, PmiRR95A, PmiRR184A, and PmiRS218A could not efficiently bind to MIC. However, the PmiRD143A mutant displayed higher binding affinity (Kd = 2.1 μM) than WT PmiR, likely due to the removal of competition between D143 site chain and MIC for Zn2+ coordination (Fig. 4, B and C). To further investigate the function of MIC and the conserved residues of PmiR, we performed EMSAs. Our data revealed that the addition of 1 mM MIC decreased the binding of PmiRD143A to the prpB probe, whereas MIC did not affect the binding of PmiRH147A, PmiRH192A, PmiRH214A, PmiRR95A, PmiRR184A, and PmiRS218A to the same DNA probe (fig. S8A). Collectively, these results strongly suggested that these conserved residues, including H147, H192, H214, R95, R184, and S218 play, a critical role in the binding of MIC, which in turn inhibits DNA binding by PmiR.
As mentioned above, pyocyanin production is increased in ΔpmiR as compared to WT PAO1 (Fig. 1A). Therefore, we used this phenotype to compare the in vivo activity of the WT and mutated variants of PmiR by means of a complementation assay. The site-mutated coding sequence of pmiR was cloned into plasmid pAK1900 and independently introduced into the ΔpmiR strain. As shown in Fig. 4G, complementation of this mutant with p-pmiR or p-pmiRD143A restored the pyocyanin production to WT levels. Conversely, the pyocyanin production was not complemented by introducing plasmids containing p-pmiRH147A, p-pmiRH192A, p-pmiRH214A, p-pmiRR95A, p-pmiRR184A, or p-pmiRS218A. We also confirmed the function of these conserved residues using the phenotype that the expression of pqsH was increased in the ΔpmiR strain (fig. S8B). Combined with the in vitro results, these data clearly demonstrated that PmiR residues H147, H192, H214, R95, R184, and S218 are essential for PmiR activity in the cell.
Basis for DNA binding and regulation by MIC
Although only one PmiR monomer presents in the asymmetric unit of the crystal lattice, the in vitro size exclusion chromatographic profile indicated that PmiR exists as homodimer in solution (fig. S9A). PmiR homodimer (Fig. 5A) can be easily generated through symmetric operation. Dimerization is mediated by various interactions between the two PmiR monomers. As depicted in fig. S9B, the side chain of R161 forms H-bond interaction with the main chain O atom of G66, whereas it forms salt-bridge interaction with the side chain of E166. Salt-bridge interaction also forms between the side chains of E93 and R109; besides the main chain O atom of E90, the side chain of N156 also forms H-bond interaction with the side chain of E93 (fig. S9C). Notably, there are many hydrophobic residues (fig. S9, D and E) located at the dimerization interface, such as I94 and L98 of the α4 helix and L159, M162, and L167 of the α7 helix, forming extensive hydrophobic interactions. In addition to the WT protein, we also constructed two PmiR mutants, I94E/L98E and L159E/M162E. Unlike the WT protein, both mutants are very unstable and quickly precipitate after the first HisTrap column purification, suggesting that the hydrophobic interactions are important for the stability and function of PmiR.
Fig. 5. Basis for DNA binding regulation by MIC.
(A) The cartoon presentation of the apo-form PmiR homodimer. The HTH and effector binding domains are colored differently in one monomer, whereas both are colored in light-blue in the partner molecule. (B) The surface presentations of PmiR homodimer. The positive and negative residues are colored in blue and red, respectively. (C) EMSAs showing the DNA binding abilities of WT and mutated-PmiR proteins. Each reaction mixture contains PCR products of prpB (2.0 ng/μl). The protein concentrations are indicated above the lane. Data are representative of three independent replicates. (D) The proposed DNA binding model of PmiR homodimer, which is colored as in (A). Residues important for DNA binding are highlighted as spheres. The disordered linker-1 region is indicated by black dots. (E) Comparison of the overall structures of the apo-form and MIC-complexed PmiR. (F and G) Superposition showing the conformational changes of D143 and the 139QQEGD143 fragment in the apo-form and MIC-complexed PmiR structures. In (E) to (G), the C atoms of the MIC-complexed PmiR proteins are colored in magenta. Zn2+ ion and MIC are shown as yellow sphere and sticks in atomic color (C, cyan; O, red), respectively. The apo-form PmiR structure is colored similarly as in (A). EB, Effector binding
PmiR can bind the promoter and regulate the transcription of various genes. As demonstrated by many reported structures, transcription factors mainly use their positively charged residues for interacting with DNA. Of the total 240 residues of PmiR, 39 are positively charged (either Arg, Lys, or His) and concentrate on several different regions of the surface (Fig. 5B). To map the residues important for DNA binding by PmiR, we performed systematic mutations (fig. S9F) and in vitro DNA binding assays. The results showed that single Ala substitution of R24, R48, R54, R58, H62, R72, or R172 all lowered the DNA binding abilities of the protein, compared to that of WT PmiR (Fig. 5C). R48, R54, R58, H62, and R72 all belong to the HTH domain and are located at the A region of the surface (Fig. 5B). R24 from the same HTH domain resides on the back and is far away from any of the A-region residues. However, we found that R24 and R172 from the partner molecule HTH and effector binding domains are located at the B region, which is very close to the A region in space (Fig. 5B). On the basis of the structural analysis and in vitro binding assay results, we proposed one plausible DNA binding model of PmiR (Fig. 5D).
The DNA binding ability of PmiR is suppressed by MIC. To understand the underlying basis, we compared the apo-form and MIC-bound PmiR structures. As supported by the low root mean square deviation value (0.6 Å), the overall folds of the two structures are very similar (Fig. 5E). However, binding of MIC does cause some conformational changes of local residues. In the apo-form structure, the side chain of D143 coordinates with the Zn2+ ion (Fig. 4B), whereas it rotates about 120° around the C─C bond to avoid clash with the carboxylate group at position 1 of MIC in the MIC-complexed structure (Fig. 5F). Unlike the apo-form structure, 139QQEGD143 does not form continuous helix (α6) with residues 144 to 153 in the MIC-complexed structure; instead, it adopts an extended conformation.
PmiR α5 and α6 helices are connected by one long linker, linker-1 (amino acids 127 to 138). Linker-1 is disordered in the apo-form structure and is very close to DNA in the proposed model (Fig. 5G). The high flexibility may allow linker-1 to undergo certain conformational arrangement to interact with DNA. Although it is also disordered in the MIC-complexed structure, connecting with the extended 139QQEGD143 fragment may prevent linker-1 from large conformational arrangement, leading to the lower DNA binding ability of PmiR.
Comparison with homologous proteins
PmiR belongs to the GntR superfamily. The structures of many GntR superfamily proteins have been reported, but they all share low (<40%) sequence similarity with PmiR. To better understand the structural and functional relationship of PmiR, we compared PmiR with FadR, one of the most studied members of the GntR superfamily. FadR shares 25% sequence similarity with PmiR (fig. S10A). Of the four Zn2+ ion-coordinating residues, only one is conserved in FadR, which may contribute to their different effector preference. Besides the apo-form structure, the ligand-bound and DNA-bound structures of FadR have also been reported. Superposition showed that the overall folds of the HTH and EB domains are similar in the PmiR and FadR structures (fig. S10, B and C), but the linker-1 region of FadR is much shorter (fig. S10A). Besides the orientations of the HTH domains (fig. S10C), the structural analysis and binding assay results (Fig. 5C) indicated that residues involved in DNA binding are not identical for PmiR and FadR.
Very recently, the structures of Atu1419 from Agrobacterium fabrum were reported (23). Atu1419 shares 36% sequence similarity with PmiR (fig. S11A). The overall folds of the effector binding domains are similar in the PmiR and Atu1419 structures, but they showed clear differences at two regions (fig. S11B), namely, the linker-1 region and the linker-2 region (which connects the α7 and α8 helices). The effector binding domain of Atu1419 also captured one Zn2+ ion, which forms similar coordination as the one in the apo-form PmiR structure. The crystallization conditions of several Atu1419 crystals contained high-concentration citric acid (CIT). In the structures, well-defined CIT was observed, coordinating with the Zn2+ ion. CIT adopts two different conformations (fig. S11C), but neither changes the conformations of N137 (corresponding to D143 of PmiR) and the linker-1 region residues. Conformational changes can be observed for the linker-2 region residues (fig. S11D); however, comparison with the ternary Atu1419-DNA-CIT complex (fig. S11E) indicated that CIT has no effects on DNA binding and is not a natural effector of Atu1419. Upon effector binding, conformational changes also occur to the linker-2 region of the FadR structure.
FadR and Atu1419 bind DNA by their HTH domains in a symmetric manner (figs. S11B and S12E). The effector-induced conformational changes of the effector binding domains do not directly block target DNA recognition by either FadR or Atu1419; instead, they mainly follow the allosteric mechanism in suppressing target DNA transcription. The recognized DNA sequences are 5′-ATGTATACAT-3′ and 5′-TGGTNxACCA-3′ for Atu1419 and FadR, respectively. PmiR regulates the expression of various genes. Although it also belongs to the GntR superfamily, it recognizes very different DNA sequences. Unique effectors and conformational changes (fig. S11F) all suggested that PmiR is different from Atu1419, FadR, and many other GntR members. Rather than indirect allosteric mechanism, the effectors of PmiR directly compete with DNA binding in an asymmetric manner (Fig. 5D). The HTH domains adopt different orientations in the Atu1419-DNA-CIT complex and the PmiR structures (fig. S11G). At this stage, we could not fully rule out the possibility that the PmiR HTH domain undergoes conformational rearrangement and binds DNA in a symmetric manner. However, the biological relevance of such symmetric binding mode needs to be further investigated in the future.
Loss of pmiR promotes mouse mortality and lung injury following P. aeruginosa infection
The increased pyocyanin production and bacterial motility in the ΔpmiR strain led us to investigate the role of PmiR in pathogenesis using a well-established mouse model of acute pneumonia with which we have years’ experience (24). To this end, the WT PAO1, ΔpmiR, the complemented strain ΔpmiR/p-pmiR and ΔpmiRΔpqsR double mutant were intranasally inoculated into C57BL/6J mice, respectively. Survival studies showed that the loss of pmiR significantly decreased mouse survival compared to the WT PAO1, in which ΔpmiR caused mouse death up to 33.33% by 24 hours. By contrast, the WT PAO1–infected mice showed lowered death rates with 61.11% survival by 24 hours (Fig. 6A). The survival experiments in mice have been vigorously performed with three repetitions containing six mice per group and analyzed by Kaplan-Meier (log-rank test) assays. This phenotype of survival was restored by infection with the complemented strain ΔpmiR/p-pmiR or ΔpmiRΔpqsR double mutant strain. This observation indicates that PmiR plays a role in virulence in mouse pneumonia models, which may be also exerted by pqsR, as ΔpmiRΔpqsR double deletion exhibited a similar phenotype of WT PAO1 strain. In addition, the colony-forming units (CFUs) in lungs and bronchoalveolar lavage fluid (BALF) from the ΔpmiR-infected mice at 24 hours were significantly higher than those of mice challenged with WT PAO1 or the complemented strain (Fig. 6, B and C). Moreover, there were increased lung injury and inflammatory cells in the mice infected with ΔpmiR strain compared to those infected by WT PAO1 (Fig. 6E). Quantitative scoring of histological lung injury demonstrated that ΔpmiR-infected mice appeared significantly aggravated compared to WT PAO1-infected mice (Fig. 6D). Furthermore, cell viability of alveolar macrophages (AMs) isolated from mice was enumerated using a cell counting kit-8 (CCK-8) kit, which decreased evidently after ΔpmiR strain infection compared to WT PAO1 infection (Fig. 6F).
Fig. 6. PmiR plays an important role in P. aeruginosa pathogenesis.
(A) Deletion of pmiR increased the virulence of P. aeruginosa. C57BL/6J mice were intranasally challenged with WT PAO1, ΔpmiR, ΔpmiR/p-pmiR, or ΔpmiRΔpqsR at 2 × 107 CFU in 50 μl of phosphate-buffered saline (PBS), and moribund mice were euthanized for survival data, which are presented with Kaplan-Meier survival curves (n = 6 for each group). The mouse survival assay was measured three times, and data shown are representative of these assays. Statistical significance was calculated using log-rank test. (B and C) Mice were infected with bacteria as (A) or a PBS control. At 12 and 24 hours, the lungs (B) and BALF (C) from infected mice were collected. Bacterial loads were enumerated through serial dilution and plating. (D) Evaluation of lung injury. (E) Lung injury assessed by hematoxylin and eosin staining at 24 hours after infection with the indicated P. aeruginosa strains. Scale bars, 1000 μm (top; original magnification, ×20) and 1000 μm (bottom; original magnification, ×40). The boxes at the top were enlarged in insets below. (F) Cell viability of AM was evaluated through the CCK-8 assay. (G and H) ROS production in AMs was detected by an H2DCF assay (G) and NBT assay (H). RFU, Relative Fluorescence Unit; RLU, Relative Light Unit. (I) The measurements of MPO activity in the lung. Error bars indicate the SD from three independent replicates. Groups were compared with each other in statistics. **P < 0.01, ***P < 0.001, ****P < 0.0001 based on one-way ANOVA Dunnett’s multiple comparison test.
Bacterial infection is consistently shown to promote reactive oxygen species (ROS) induction, and excessive production of ROS can cause organelle disruption and cell malfunction, consequently leading to tissue injury unless immediate and effective removal of ROS (25, 26). To assess the levels of ROS in the infected mouse cells, we isolated AMs from the mice infected with above strains, which showed a ∼2-fold increase in oxidative levels of the ΔpmiR strain–infected mice compared to the PAO1 strain–infected mice (Fig. 6G), using a dihydrodichlorofluorescein diacetate (H2DCF) assay, which is a commonly used sensitive fluorescence method for quantifying superoxide. To ascertain the data of ROS analysis, the ROS levels following deletion of pmiR was also determined using a nitroblue tetrazolium (NBT) assay, and similar results to the H2DCF assay were attained (Fig. 6H). It would be a merited approach to differentiate the source of ROS from two major acute inflammatory phagocytes, AMs and polymorphonulcear neutrophils (PMNs). As myeloperoxidase (MPO) activity reflects the main contribution of ROS of PMNs, we carried out an MPO assay for the homogenate of lung tissues following infection and observed an increase in MPO in the lungs infected by the pmiR-deletion strain versus the WT strain or complemented strain (Fig. 6I). Together, these results clearly demonstrate that PmiR plays a pivotal role in regulating bacterial virulence in mice with increased death rates, bacterial burdens, and tissue injury by augmenting oxidative stress and inflammatory responses.
pmiR deficiency aggravated inflammatory response and pyroptosis through the signal transducer and activator of transcription 3/nuclear factor κB signal pathway after P. aeruginosa infection
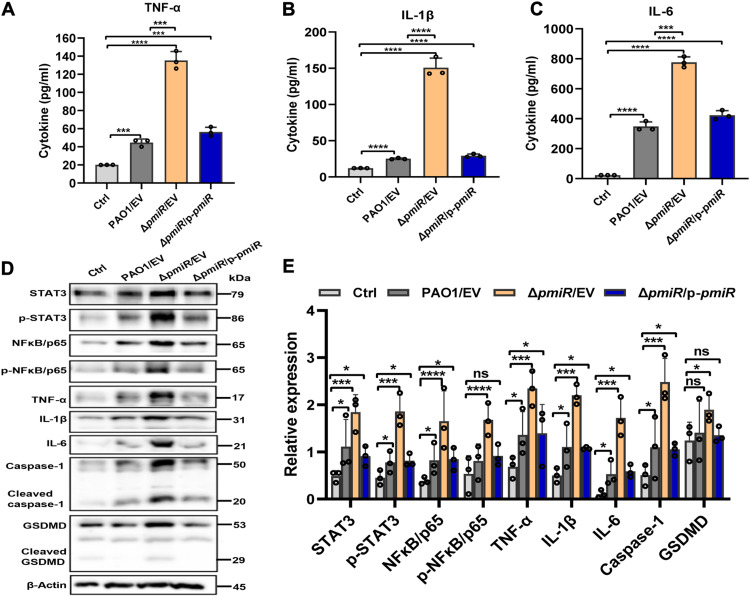
To investigate whether PmiR influences host inflammatory responses, we measured critical inflammatory cytokines [tumor necrosis factor–α (TNF-α), interleukin-1β (IL-1β), and IL-6] in BALF 24 hours after infection as inflammatory situations are critical for host defense and survival against infectious diseases. We found that these inflammatory cytokines were significantly increased in BALF from ΔpmiR strain–infected mice compared to those from WT PAO1–infected mice (Fig. 7, A to C), indicating that pmiR deletion intensified inflammatory response. In addition, immunoblotting analysis demonstrated increased expression of inflammatory cytokines (TNF-α, IL-1β, and IL-6). Recent studies have identified a role of inflammasome components in bacterial infection, which is tightly associated with pyroptosis (27). To evaluate the importance of PmiR for inflammasome pathways, we performed Western blotting analysis and found increased caspase-1 and gasdermin-D (GSDMD) in ΔpmiR strain–infected mice compared to WT PAO1–infected mice (Fig. 7, D and E), which is consistent with the results of the inflammatory cytokine profiles of the infected mouse lungs.
Fig. 7. pmiR deficiency aggravated inflammatory response and pyroptosis through an STAT3/NF-кB signaling pathway after P. aeruginosa infection.
(A to C) Inflammatory cytokines in BALF were assessed by enzyme-linked immunosorbent assay. (D) Western blot detected the expression of inflammatory cytokines (TNF-α, IL-1β, and IL-6), pyroptosis markers (caspase-1 and GSDMD), STAT3, p-STAT3, NF-кB/p65, and p-NF-кB/p65. (E) The densitometry of the data shown in (D). Error bars indicate the SD from three independent replicates. Groups were compared with each other in statistics. *P < 0.05, ***P < 0.001, ****P < 0.0001 based on one-way ANOVA Dunnett’s multiple comparison test.
Our previous studies reported that the signal transducer and activator of transcription 3 (STAT3)/nuclear factor κB (NF-κB) signal pathway in the host was activated to promote proinflammatory cytokine expression and alter pyroptosis against P. aeruginosa infection (27). To further understand the underlying mechanisms for PmiR-mediated virulence alteration, we again determined expression levels of STAT3 and NF-κB using Western blotting assays. The protein levels of STAT3 and NF-κB/p65 in ΔpmiR-infected mice were markedly increased compared to PAO1-infected mice (Fig. 7, D and E). Together, these findings indicated that PmiR may down-regulate pyroptosis and inflammatory responses in mice through the STAT3/NF-κB signaling pathway, which may be associated with the regulation of pathogenic virulence.
DISCUSSION
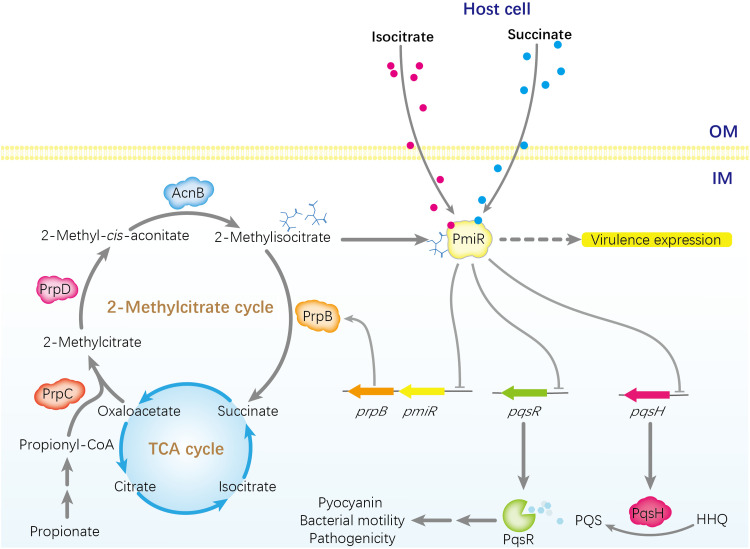
The metabolic status of P. aeruginosa may have a profound impact on the sensing of the environmental signals or favorable changes to fight against the host immune system. P. aeruginosa is capable of transforming from a commensal organism into a virulent pathogen (28). Here, we identified a GntR family protein, PmiR, which was involved in regulation of the pqs system and virulence factors in P. aeruginosa. We resolved the crystal structure of PmiR/MIC binary complex and found that PmiR sensed the metabolic intermediates, including MIC and its analogs, to regulate the 2-MCC metabolism and the expression of virulence factors. On the basis of these data, we propose a model in which PmiR plays a critical role in the regulation of pathogenesis by sensing metabolites in P. aeruginosa (Fig. 8). In summary, to our knowledge, we report the first example of an MIC receptor involved in regulating virulence in bacteria.
Fig. 8. Proposed model of PmiR sensing the metabolic intermediates to regulate expression of genes involved in pqs system and virulence factors in P. aeruginosa.
In this study, we defined the pathway of 2-MCC and demonstrated that PmiR represses the expression of genes involved in 2-MCC and pqs system directly in P. aeruginosa. PmiR as a receptor could bind and sense MIC and its analogs, such as succinate and isocitrate (produced by P. aeruginosa or come from cell host), to regulate virulence factors. Arrows indicate positive regulation, and T bars present negative regulation. OM, outer membrane; IM, inner membrane.
As a metabolic intermediate of 2-MCC, the MIC is converted from 2-methylcitrate by PrpD and AcnB. PrpB, methylisocitrate lyase, was the last but critical enzyme to convert MIC to succinate and pyruvate, and many studies unraveled that the deletion of prpB gene significantly promoted the virulence to the host (4, 29), suggesting that the activated or full-fledged MIC is closely related to the bacterial virulence. PmiR was characterized as an MIC-dependent regulator, which can directly suppress the enzymatic activity of PrpB. Presumably, the accumulation of MIC induced the virulence via PmiR-mediated PrpB inactivation. Moreover, the prpB deletion increased the pyocyanin production and bacterial motility seen as the virulent phenotype, which was in agreement with that of pmiR mutant. To our delight, the in vivo studies showed that the ΔpmiRΔpqsR double mutant reversed the disease phenotype of ΔpmiR single mutant infection (Fig. 6), which led to better mouse survival and worse patterns in bacterial burdens of lungs and BALF, tissue injury, and intense inflammatory responses. These critical findings reveal that PmiR plays key roles in virulence, which affects host responses through a potential PmiR-PqsR axis. Our studies not only dissect the biochemical mechanism of PmiR involved in 2-MCC metabolic pathways but also indicate the potential significance in mammalian immunity and clinical sciences.
Recently, we have identified another GntR family regulator, MpaR, which coordinately controls anthranilate metabolism and P. aeruginosa virulence by directly regulating pqsR (18). Phylogeny of GntR family proteins suggests that PmiR and MpaR were found to localize on distinct subclades, indicating that they might have experienced different evolutionary scenarios (fig. S12). However, the pqs system is simultaneously regulated by MpaR and PmiR, although each of these is involved in modulating two different metabolic pathways. In addition, these two proteins have been shown to play critical roles in P. aeruginosa pathogenicity in a mouse model of acute pneumonia (24). Therefore, these observations provide insights into the dual functionality of these metabolic regulators, which indicates a need to fully elucidate the other uncharacterized GntR family proteins.
Bacteria have evolved different types of signal transduction systems, including the transcriptional regulators, TCSs, and chemosensory pathways, that enable them to adapt to a wide range of environmental changes (30, 31). The interaction between metabolic sensing and bacterial responses has been proposed in many bacterial species. In Enterohemorrhagic Escherichia coli, the Cra regulator senses succinic acid to activate expression of virulence factors (32). The PrpR protein has been identified as a receptor of 2-methylcitrate to regulate the propionate catabolism in Salmonella enterica (33). P. aeruginosa exhibits chemotaxis toward histamine via TlpQ, PctA, and PctC chemoreceptors (34). In addition, a recent study has revealed that histamine promotes the virulence of P. aeruginosa by activating HinK (13). However, the receptors of MIC and response patterns to this signal remain elusive. PmiR belongs to the GntR superfamily. Although it shares similarity with FadR, Atu1419, and many other GntR superfamily members in their overall folding, our structural studies showed that PmiR was unique in the EB region, especially the linker-1 and linker-2 regions (Fig. 5G). PmiR was different from the homology proteins in the detailed EB mode, the associated conformational changes, and the mechanisms by which DNA binding was inhibited (Fig. 3, A and B, and fig. S7). In addition to MIC, our ITC binding assays showed that PmiR also recognized other effectors, such as isocitric acid and succinic acid. In comparison with MIC, isocitric acid lacks a methyl group at position 2, which was not involved in MIC binding by PmiR (Fig. 3D). Apart from this, the compositions and geometries of isocitric acid and MIC were identical. The similar isocitric acid and MIC binding affinities suggested that the presence or absence of the position-2 methyl group did not affect the EB by PmiR. In all probability, PmiR uses an identical mode of isocitric acid and MIC binding. Succinic acid is composed of two carboxyl groups and two methylene groups. Such structural elements are conserved in both isocitric acid and MIC; however, succinic acid is more flexible in conformational changes because of the high flexibility of the methylene groups. Although it remains to be verified, we reasoned that PmiR has the ability to bind and interact with both carboxyl groups of succinic acid. Considering limited impact of citrate on the PmiR-DNA interaction, and citrate is also the substrate of TCA and easily consumed, this interaction may be unlikely to act as the potential signals for PmiR system. Although methylcitrate is the first intermediate of 2-MCC, the methyl group appeared to have limited influence on PmiR binding ability, and noticeably, prpCD mutation of M. tuberculosis suggested no further degradation and accumulation of methylcitrate, which cannot cause distinguishable changes from the WT in terms of bacterial growth and persistence and, especially, the virulence (35). Henceforth, methylcitrate seems not to be the candidate signals of PmiR.
Most pathogenic bacteria have evolved sophisticated systems to detect hostile environments and trigger compensatory gene expression to survive within the host (8). The metabolic cross-talk between the host cell and the pathogen and the response of the pathogen to a milieu often filled with immunosignaling metabolites have been documented previously (36). In most bacteria, 2-MCC is the main pathway used for the detoxification of propionate and propionyl-CoA (2, 6, 37). Research reported that PrpB and PrpD are localized in the cytoplasm, and Acn are localized in both the cytoplasm and mitochondria in Toxoplasma gondii (5). However, mitochondria are not present in bacteria. Thus, MIC may accumulate in the cytoplasm and is a bacterial product. Our data indicate that PmiR could sense MIC or isocitrate signal molecule to regulate the expression of virulence factors, which temper P. aeruginosa pathogenesis and coordinate its continual integration with environmental signals. Bacteria also use the host immunometabolites as signaling cues to induce expression of virulence factors (38). For example, host succinate is an activation signal for Salmonella virulence during intracellular infection (39). Sensing of host-derived pyruvate and lactate by the TCS CreBC results in the induction of the SPI-2 regulon (40). Host-derived glutathione was shown to regulate PrfA activation through allosteric binding in Listeria monocytogenes (41). During infection with P. aeruginosa, succinate is released to airways (38). P. aeruginosa isolates from patients with cystic fibrosis have been shown to become adapted to using succinate as the primary carbon source and has substantial metabolic flexibility to adapt to the environment within the host often by changing their transcriptional profiles (42). Here, we revealed that PmiR binds not only to MIC but also to succinate and isocitrate. Therefore, it is important to investigate how PmiR senses the host-derived succinate/isocitrate and the interaction between P. aeruginosa and its host after sensing these immunometabolites in the future.
In conclusion, this study unveils that PmiR senses MIC to regulate the expression of virulence-associated genes in P. aeruginosa. These findings promote a better understanding of the interaction between the environmental signals sensed by bacteria and the subsequent host responses. Future investigations should determine whether other bacterial species share such a mechanism. Moreover, the binding of PmiR to isocitrate or succinate could be considered for developing a biosensor or therapeutic target, which could have applications in the diagnosis and treatment of 2-MCC–related diseases.
MATERIALS AND METHODS
Ethical statement
All mouse experiments were performed in accordance with the U.S. National Institutions of Health Animal Care and Institutional Guidelines. Animal handling protocols were approved by the University of North Dakota Institutional Animal Care and Use Committee. The University of North Dakota animal facilities were accredited by the Association for Assessment and Accreditation of Laboratory Animal Care International (assurance number: A3917-01).
Bacterial strains and culture conditions
Bacterial strains, plasmids, and primers used in this study are listed in tables S3 and S4. P. aeruginosa PAO1 and its derivatives were grown in Luria-Bertani (LB) or M9 minimal medium with different carbon sources (10 mM sodium propionate and 2 mM MIC or succinate). When needed, appropriate antibiotics were used at the following concentrations: for E. coli, ampicillin (100 μg/ml), kanamycin (50 μg/ml), and tetracycline (at 10 μg/ml in LB); for P. aeruginosa PAO1, trimethoprim (300 μg/ml), tetracycline (100 μg/ml), and carbenicillin (300 μg/ml in LB or 150 μg/ml in Pseudomonas isolate agar).
Construction of plasmids
To construct p-pmiR plasmid, the DNA fragments containing pmiR promoter were generated by PCR using pmiR-com-S and pmiR-com-R primer pairs. The DNA fragments were then cloned into pAK1900 vector (43). Accordingly, the coding sequences containing the prpC, prpB, and prpD were amplified with the corresponding primer pairs and cloned into the pAK1900 vector after enzymatic digestion, yielding p-prpB, p-prpC, and p-prpD, respectively. For construction of mini–CTX-prpB-Flag plasmid, the DNA fragments encoding prpB gene driven by pmiR promoter were generated by overlap PCR using the mini–prpB-F and mini–prpB-R primer pairs. The digested PCR products were cloned into the mini–CTX-lacZ-Flag plasmid (44).
The plasmid pMS402 with a promoterless luxCDABE reporter gene cluster was used to construct promoter-luxCDABD reporter fusion of the prpB, as previously described (45, 46). The prpB promoter region was amplified by PCR using the primers prpB-lux-F and prpB-lux-R (table S4). The PCR products were cloned into the PMS402, yielding p-prpB-lux. A similar strategy was used to construct the p-prpC-lux p-prpD-lux, p-asrA-lux, and p-esrC-lux plasmids. To generate mini–CTX-pqsH-M-p-lux (CGCCGCCCGCCGCGGCG was mutated to CTCCTCCCGCCTCTTCT), the two fragments were amplified by PCR using pqsH-M-F1/pqsH-M-R1 and pqsH-M-F2/pqsH-M-R2 primer pairs, respectively (table S4). Subsequently, the mutated sequences were obtained by PCR using the pqsH-M-F1/pqsH-M-R2 primers and the above two PCR products as template. The overlap PCR products were digested with the corresponding enzymes and ligated into CTX6.1 vector (47). A similar strategy was used to construct the mini–CTX-pqsR-M-p-lux. All the constructs were sequenced to verify that no mutations had occurred.
To construct the pET-Sumo-PmiR plasmid, genomic DNA from P. aeruginosa was used as the template for PCR with pET-pmiR-F and pET-pmiR-R primer pairs (table S4). The PCR products were resolved on agarose gel and digested with corresponding enzymes. The target fragment was cloned into the pET28a-Sumo vector. The truncated protein (amino acids 15 to 232) of PmiR was generated by PCR using the pET28a-Sumo-PmiR plasmid as the template. The PCR fragments were separated by agarose gel and recombined into the digested pET28a-Sumo vector. A similar strategy was used to construct the pET-PmiR, pET-PrpB, pET-PrpC, and pET-PrpC plasmids, except for the pET28a-Sumo, which was replaced by pET28a vector. For construction of the pET28a-pmiRD143A, the two fragments were PCR using pET-pmiR-F/p-pmiRD143A-overlap-R and p-pmiRD143A-overlap-R/pET-pmiR-R primer pairs. Subsequently, the mutated sequence was PCR using the pET-pmiR-F/ pET-pmiR-R primers and the above two PCR products as the template. Then, the mutated PCR products were digested with the corresponding enzymes and cloned into pET28a vector. The PmiRH192A, PmiRH214A, PmiRH147A, PmiRD143A, PmiRR95A, PmiRS218A, PmiRR184A, PmiRR24A, PmiRR48A, PmiRR54A, PmiRH62A, PmiRR72A, PmiRR127A, PmiRR135A, and PmiRR172A mutants were constructed by the same strategy. All constructs were confirmed by DNA sequencing. The recombinant plasmid was transferred into E. coli BL21 (DE3) competent cells for protein expression.
Construction of deletion mutants
For construction of gene knockout mutants, a sacB-based strategy was used, as previously described (44). To construct the pmiR null mutant (ΔpmiR), PCRs were performed to amplify sequences upstream [1521 base pairs (bp)] and downstream (1515 bp) of the intended deletion. The upstream was amplified from PAO1 genomic DNA using pEX-pmiR-up-F/R and pEX-pmiR-down-F/R primer pairs (table S4). The two PCR products were digested and then cloned into the Hind III/Xba I– and Xba I/Bam HI–digested gene replacement vector pEX18Ap (48), yielding pEX18Ap-pmiR. The resultant plasmids were electroporated into PAO1 with selection for carbenicillin resistance. Colonies showing both carbenicillin susceptibility and sucrose resistance were selected on LB agar plates containing carbenicillin (300 μg/ml) and 10% sucrose, which typically indicates a double-crossover event and thus of gene replacement occurrence. A similar strategy was used to construct the ΔprpB, ΔprpC, and ΔprpD. For generating ΔpmiRΔpqsH and ΔpmiRΔpqsR double-deletion strains, the pmiR gene in ΔpqsH and ΔpqsR mutant was deleted by a similar strategy with plasmid pEX18Ap-pmiR. These mutants were verified by PCR.
Measurement of pyocyanin production
Pyocyanin was extracted from culture supernatants and measured using previously reported methods (18). Briefly, 3 ml of chloroform was added to 5 ml of culture supernatant and vortex mix. After extraction, the chloroform layer was transferred to a fresh tube and mixed with 1 ml of 0.2 M HCl, and the top layer was removed by centrifugation. Measuring its absorption at 520 nm, the pyocyanin production (microgram)/culture supernatant (milliliter) was determined by multiplying the optical density at 520 nm (OD520) by 17.072.
Bacterial motility assay
The motility assay was carried out as previously described with minor modification (49). The swarming medium was 0.8% nutrient broth, 0.5% glucose, and 0.5% agar. Swarming agar plates were spot-inoculated with bacteria as 2 μl of aliquots taken directly from overnight LB cultures. The plates were incubated for 16 hours at 37°C. The swimming medium was 1% tryptone, 0.5% NaCl, and 0.3% agar. The maximum diameter of the motility zone was recorded for three replicates per strain.
RNA extraction and real-time quantitative PCR
Overnight bacterial cultures were subcultured in fresh LB medium to OD600 of 0.6. The bacterial strains were collected by centrifugation at 12,000g for 2 min. Total RNA was isolated with an RNAprep pure cell/bacteria kit (Tiangen Biotech, Beijing, China). Complementary DNA (cDNA) was synthesized from each RNA sample using a PrimerScript Reverse Transcriptase (TaKaRa, Dalian, China) with random primer and then subjected to qRT-PCR using SYBR Premix Ex Taq II (TaKaRa). The 30S ribosomal protein gene rpsL was used as an internal control.
RNA-seq and data analysis
The WT PAO1 and ΔpmiR strains were grown in LB medium at 37°C until OD600 reached approximately 0.6. Total RNA was extracted immediately upon cell harvest using TRIzol reagent according to the manufacturer’s instructions (Invitrogen), and genomic DNA was removed using DNase I (TaKaRa). After removing ribosomal RNA, mRNA was used to generate the cDNA library according to the TruSeq RNA sample preparation kit from Illumina (San Diego, CA). Thereafter, the cDNA libraries were sequenced on an Illumina HiSeq 2000 system. Each sample in RNA-seq assay was repeated twice. RNA-seq reads were mapped to the P. aeruginosa genomes (NC_002516.2) provided by National Center for Biotechnology Information using Bowtie2, and only the uniquely mapped reads were kept for the subsequent analyses. DEGs were identified using DESeq2 (Benjamini-Hochberg–adjusted P < 0.05 and |log2 fold change| > 1) (50). The data have been uploaded to BioProject and accession number is PRJNA837700.
Luminescence screening assays
Expression of lux-based reporters from cells grown in liquid culture was measured as counts per second of light production in a Synergy 2 plate reader (BioTek), as previously described (18, 45). Briefly, overnight cultures of the reporter strains were diluted to an OD600 of 0.2 with fresh LB medium and then cultured for another 2 hours at 37°C with shaking. Subsequently, the cultures were diluted 1:20 with 100 μl of fresh LB and then added to a black 96-well plate with a transparent bottom. Fifty microliters of sterilized mineral oil were added to the culture. Luminescence intensity and bacterial growth (OD600) were continuously measured every 30 min for 24 hours in a Synergy 2 plate reader.
Western blot analysis
Overnight LB cultures were washed three times using M9 minimal medium, and the cultures were adjusted to OD600 of 0.01. Subsequently, the cultures were cultured for 5 hours, and the isocitrate was added to M9 minimal medium. After incubation for 4 hours, samples from equivalent numbers of bacterial cells were loaded and separated on 12% SDS–polyacrylamide gel electrophoresis (SDS-PAGE) gels. The proteins were transferred to a polyvinylidene difluoride membrane and hybridized with a mouse monoclonal Flag antibody (Ab, Sigma-Aldrich). The signal was detected using an ECL Plus kit (Amersham Biosciences).
PQS extraction and analysis by TLC
The extraction and analysis of PQS were performed as previously described (45). Briefly, P. aeruginosa PAO1 and its derivatives were grown in 5 ml of LB overnight at 37°C. The cultures were centrifuged and resuspended to inoculate 5 ml of subcultures to OD600 of 0.05. Subcultures were grown for 24 hours, and about 0.5 ml of each culture was mixed with 1 ml of acidified ethyl acetate and vortexed vigorously, and the water phase was removed by centrifugation. The organic phase was collected and air-dried using a vacuum freeze drier and dissolved in 50 μl of sample buffer (1:1 of acidified ethyl acetate and acetonitrile). TLC was performed on a silica gel plate that had been soaked in 5% KH2PO4 and activated at 100°C for 1 hour before use. Extracts were separated using 17:2:1 methylene chloride/acetonitrile/dioxane as the solvent. When the solvent front reached the top of the plate, the plate was immediately photographed under ultraviolet (UV) light.
Bacterial growth assay
The chemical sodium propionate, MIC, sodium citrate, and succinate were dissolved in double-distilled water, which were filter-sterilized through a 0.22-μm filter membrane and added into M9 MM at indicated final concentrations. Overnight LB cultures were washed three times with fresh M9 minimal medium and then adjusted to OD600 of 1.0. Subsequently, the bacterial cells were subcultured with 100-fold dilution in M9 minimal medium supplemented with 10 mM sodium propionate and 2.0 mM MIC or succinate as the sole carbon source. The absorption of A600 (absorbance at 600 nm) was detected at the indicated time points by a Synergy 2 Multi-Mode Microplate Reader (BioTek). To calculate the generation time, two points were selected (t = 4 hours, t = 12 hours) and using the formula: T = t × 0.301/(lg Nt-lg N0), where Nt = higher CFU/ml, N0 = lower CFU/ml, and t = time interval (in hours) between the points.
Protein expression and purification
All proteins were expressed and purified using similar procedures. The cells were cultured at 37°C in LB medium. When the OD600 reached 0.6 to 0.8, isopropyl-d-1-thiogalacto-pyranoside (final concentration of 0.2 mM) was added to induce the expression of the protein. The cultures were incubated at 18°C for an additional 18 to 20 hours. The cells were collected via centrifugation and resuspended in buffer A [20 mM tris-HCl (pH 8.0), 300 mM NaCl, 200 mM (NH4)2SO4, and 25 mM imidazole]. The cells were lysed by high-pressure homogenization and centrifuged at 17,000 rpm for 1 hour at 4°C. The supernatant was applied to a 5-ml HisTrap HP column (Cytiva), and the target protein was eluted via AKTA purifier (GE Healthcare) using buffer B [20 mM tris-HCl (pH 8.0), 300 mM NaCl, 200 mM (NH4)2SO4, and 500 mM imidazole]. The target protein was pooled, treated with ULP1 protease, and dialyzed against buffer C [20 mM tris-HCl (pH 8.0), 300 mM NaCl, and 200 mM (NH4)2SO4] at 4°C for 3 hours. The sample was reloaded onto the HisTrap HP column. The target protein was collected and applied to a HiLoad 16/600 Superdex 75-pg gel filtration column (Cytiva) equilibrated with buffer composed of 20 mM tris-HCl (pH 8.0), 500 mM NaCl, 60 mM (NH4)2SO4, and 2 mM dithiothreitol (DTT). The purified protein was concentrated to 15 mg/ml and stored at −80°C.
To analyze the oligomerization state of PmiR, WT (500 μl) or mutated PmiR proteins (1 mg) were applied to a Superdex 200 10/300 GL column (GE Healthcare) equilibrated with buffer containing 20 mM tris-HCl (pH 8.0), 500 mM NaCl, 60 mM (NH4)2SO4, and 2 mM DTT. The flow rate was set at 0.5 ml/min.
The enzymatic activity test by HPLC
HPLC was applied for analysis of the enzymatic reaction (51). To measure the PrpB activity, 1.0 mM commercial DL-thero-MIC (MedChemExpress LLC) and PrpB protein (20 μg) were mixed in 1.0 ml of reaction buffer [1 mM tris-HCl (pH 7.5)]. The A214 was monitored by HPLC-UV. To measure the PrpC activity, 1-ml mixture containing 10 mM propionyl-CoA, 20 mM oxaloacetate, and 50 μg of PrpC protein in 10 mM Hepes (pH 7.2) was incubated overnight. The A220 was monitored by HPLC-UV. To measure the PrpD activity, 1-ml samples containing 0.5 mM commercial (2RS, 3RS)-2-methylcitrate and 50-μg PrpD protein in reaction buffer [20 mM tris-HCl (pH 7.5)] were incubated overnight. All the reactions were quenched with 100 μl of 1 M sodium phosphate buffer (pH 2.9). The A240 was monitored by HPLC-UV. Peaks were correlated by retention time and mass/charge ratio with commercially standards of MIC, succinate, pyruvate, propionyl-CoA, oxaloacetate, and 2-methylcitrate (Sigma-Aldrich). Unfortunately, the standard sample of (Z)- or (E)-2-methylaconitate was not commercially available. The HPLC settings were followed as previously described (51).
Electrophoretic mobility shift assay
Purified proteins were mixed with DNA probe (2 ng/μl) in 20 μl of the gel shift loading buffer [20 mM Hepes (pH 8.0), 100 mM NaCl, 1 mM DTT, sheared salmon sperm DNA (3 ng/μl), and 1% glycerol]. After incubation at 25°C for 20 min, the reaction mixtures were separated on 6% nondenaturing acrylamide gels in 0.5 × tris-borate-EDTA buffer. The gels were stained with GelRed staining solution (TransGen Biotech) and subjected to a screen by a Tanon 5200 Image analysis system (Tanon Technologies, Shanghai, China).
Dye primer–based DNase I footprinting assay
The DNA footprinting assay was followed as previously described (13). For the DNase I footprinting assay of prpB, 370 bp-5-carboxyfluorescein–labeled fragment of the prpB promoter region (nucleotide −370 to −1) was generated using the primers prpB-FT-F/prpB-FT-R (table S4). PCR products were purified with a QIA quick gel purification kit. Subsequently, a 50-μl reaction mixture containing 300 ng of prpB promoter DNA and 2 μM (or indicated) PmiR protein and binding buffer [20 mM Hepes (pH 8.0), 100 mM NaCl, 1 mM DTT, 10% glycerol, and EDTA (1 mM/liter)] up to a volume of 50 μl was incubated at room temperature for 15 min. Subsequently, 0.05 U of DNase I (Promega, Madison, USA) was added, and the reaction mixture was incubated for an additional 5 min. The DNase I digestion was terminated using quenching solution (200 mM NaCl, 30 mM EDTA, and 1% SDS), and then the mixture was extracted with 200 μl of phenol/chloroform/isoamyl alcohol (25:24:1). The digested DNA fragments were isolated by ethanol precipitation, dried under vacuum, and resuspended in ribonuclease-free water. Then, 5 μl of digested DNA was mixed with 4.9 μl of Hi-Di formamide and 0.1 μl of GeneScan-500 LIZ size standards (Applied Biosystems). A 3730 × l DNA analyzer was used to detect the sample, and the result was analyzed using GeneMapper software (Applied Biosystems, USA). The same method was used for the pqsH promoter footprinting.
ITC measurements
Isothermal titration calorimeter experiments were performed using an ITC200 isothermal titration calorimeter from MicroCal (Malvern). The experiments were carried out at 25°C. To determine the interaction between PmiR and MIC, the syringe was loaded with of 60 μl of MIC (45 μM), and the cell was filled with 300 μl of PmiR (1.0 mM). The titration curve was obtained by injecting 2 μl × 20 aliquots of MIC into the cell at intervals of 270 s while stirring at 500 rpm. The experimental data were fitted to theoretical titration curves with software supplied by MicroCal (ORIGIN). The same strategy was used to study the binding of the truncated PmiR with other molecules and the PmiR mutants with MIC.
Crystallization and data collection
The initial crystallization conditions were identified at 16°C using the Gryphon crystallization robot system from Art Robbins Instrument company and crystallization kits from Hampton Research company. Sitting-drop vapor diffusion method was used during the initial screen and optimization process. The final condition is composed of 0.1 M MES/sodium hydroxide (pH 6.5) and 1.6 M magnesium sulfate for the apo-form PmiR crystals. The MIC-complexed crystals were grown in buffer containing 0.1 M MES/sodium hydroxide (pH 5.8) and 1.2 M magnesium sulfate. The drop contains 0.3 μl of crystallization buffer and 0.6 μl of protein sample. The final concentration of the protein is 10 mg/ml. The concentration of MIC (if present) is 5.0 mM.
All crystals were cryo-protected by their mother liquid supplemented with 20% glycerol and flash-frozen using liquid nitrogen. The diffraction data were collected on beamline BL18U1 and BL19U1 at the Shanghai Synchrotron Radiation Facility. Data processing was carried out using the HKL3000 program (52). The data collection and processing statistics are summarized in table S2.
Structure determination and refinement
The apo-form PmiR structure was solved by molecular replacement (MR) method with the Phaser program embedded in the Ccp4i suite (53). Although it is not very accurate, the model generated by the AlphaFold2 program can serve as the search model to solve the PmiR structure. The resulting model was refined against the diffraction data using the Refmac5 program of the CCP4 suite. The 2Fo-Fc and Fo-Fc electron density maps were regularly calculated and used as guide for model building using COOT (54). The MIC-complex PmiR structure was solved by MR method using the apo-form structure as the search model. The Zn2+ ion and MIC molecule were all built manually using COOT. The final refinement of the two structures was done using the phenix.refine program of the Phenix suit (55). The structural refinement statistics are summarized in table S2.
Mouse pneumonia model and analysis
Overnight cultures of bacteria were subcultured 1:100 into LB medium and grown with 200 rpm shaking at 37°C to an OD600 of 0.6 to 0.8. Then, 2 × 107 CFUs of P. aeruginosa were intranasally instilled into 6 to 8 weeks old C57BL/6J mice (both male and female are randomly assigned; the Jackson Laboratory, Bar Harbor, ME), and survival rates were calculated for each bacterial strain, including deletion variants. The mice were euthanized by inhalation of CO2 after 12 hours of infection. BALF and homogenized lungs in phosphate-buffered saline were assessed for CFUs by serial dilution and plating. Cell viability was measured using CCK-8 assay (APExBIO). ROS were determined using H2DCF dye (Life Technologies) and NBT dye (Sigma-Aldrich). MPO activity in lung homogenates was measured according to our previously described methods (56). Tissue morphological damage and inflammatory response were examined using hematoxylin and eosin staining according to the standard staining procedure. Lung injury score was evaluated according to previous methods with some modification (57). Alveolar and interstitial inflammation, alveolar and interstitial hemorrhage, atelectasis, necrosis, and edema were each scored on a 0- to 4-point scale: no injury = score of 0; injury in 25% of the field = score of 1; injury in 50% of the field = score of 2; injury in 75% of the field = score of 3; and injury throughout the field = score of 4. The lung injury was then calculated on the basis of the mean score of the above parameters.
Immunoblotting
Homogenized lung tissues were lysed in ice-cold radioimmunoprecipitation assay (Thermo Fisher Scientific, Waltham, MA) supplemented with protease inhibitor cocktail (Thermo Fisher Scientific), separated by electrophoresis on SDS-PAGE gels, and transferred to nitrocellulose transfer membranes (GE Amersham Biosciences, Pittsburgh, PA). Membranes were incubated with primary Abs overnight and then were incubated with corresponding secondary Abs conjugated to horseradish peroxidase (Santa Cruz Biotechnology). Signals were visualized using an enhanced chemiluminescence detection kit (GE Health). The Abs were used from different vendors as stated below: mouse Abs against p-NF-κB/p65 (sc-136548), caspase-1 (sc-514), TNF-α (sc52746), IL-6 (28343), and β-actin (sc-47778) were obtained from Santa Cruz Biotechnology (Dallas, TX). Rabbit Abs against NF-κB/p65 (no. 8242), STAT3 (no. 12640), p-STAT3 (no. 9145), and IL-1β (no. 12703) were obtained from Cell Signaling Technology (Danvers, MA). Rabbit Abs against GSDMD (anti-GSDMD, ab209845) were obtained from Abcam.
Acknowledgments
Funding: This study was supported by grants from the National Natural Science Foundation of China (32170188 and 31870060) and ShaanXi Science and Technology Innovation Team (2019TD-016). The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Author contributions: H.L. and J.G supervised and designed the study. G.C. and Y.Z. performed most of the experiments. X.X. generated the genetic materials and performed RNA-seq experiments. Y.L. performed animal experiments. Z.L. and M.W. analyzed the data. H.L., J.G., and J.L. analyzed the results. G.C., H.L., and J.G. wrote the manuscript.
Competing interests: The authors declare that they have no competing interests.
Data and materials availability: All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials.
Supplementary Materials
This PDF file includes:
Figs. S1 to S12
Tables S1 to S4
REFERENCES AND NOTES
- 1.Horswill A. R., Dudding A. R., Escalante-Semerena J. C.,Studies of propionate toxicity in Salmonella enterica identify 2-methylcitrate as a potent inhibitor of cell growth. J. Biol. Chem. 276,19094–19101 (2001). [DOI] [PubMed] [Google Scholar]
- 2.Dolan S. K., Wijaya A., Geddis S. M., Spring D. R., Silva-Rocha R., Welch M.,Loving the poison: The methylcitrate cycle and bacterial pathogenesis. Microbiology (Reading) 164,251–259 (2018). [DOI] [PubMed] [Google Scholar]
- 3.Eoh H., Rhee K. Y.,Methylcitrate cycle defines the bactericidal essentiality of isocitrate lyase for survival of Mycobacterium tuberculosis on fatty acids. Proc. Natl. Acad. Sci. U.S.A. 111,4976–4981 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Upton A. M., McKinney J. D.,Role of the methylcitrate cycle in propionate metabolism and detoxification in Mycobacterium smegmatis. Microbiology (Reading) 153,3973–3982 (2007). [DOI] [PubMed] [Google Scholar]
- 5.Limenitakis J., Oppenheim R. D., Creek D. J., Foth B. J., Barrett M. P., Soldati-Favre D.,The 2-methylcitrate cycle is implicated in the detoxification of propionate in Toxoplasma gondii. Mol. Microbiol. 87,894–908 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zheng C., Yu Z., Du C., Gong Y., Yin W., Li X., Li Z., Römling U., Chou S.-H., He J.,2-methylcitrate cycle: A well-regulated controller of Bacillus sporulation. Environ. Microbiol. 22,1125–1140 (2020). [DOI] [PubMed] [Google Scholar]
- 7.Deretic V., Schurr M. J., Yu H.,Pseudomonas aeruginosa, mucoidy and the chronic infection phenotype in cystic fibrosis. Trends Microbiol. 3,351–356 (1995). [DOI] [PubMed] [Google Scholar]
- 8.Duprey A., Reverchon S., Nasser W.,Bacterial virulence and Fis: Adapting regulatory networks to the host environment. Trends Microbiol. 22,92–99 (2014). [DOI] [PubMed] [Google Scholar]
- 9.Corral-Lugo A., Matilla M. A., Martín-Mora D., Jiménez H. S., Torres N. M., Kato J., Hida A., Oku S., Conejero-Muriel M., Gavira J. A., Krell T.,High-affinity chemotaxis to histamine mediated by the TlpQ chemoreceptor of the human pathogen Pseudomonas aeruginosa. MBio 9,e01894-18 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Valentini M., Gonzalez D., Mavridou D. A., Filloux A.,Lifestyle transitions and adaptive pathogenesis of Pseudomonas aeruginosa. Curr. Opin. Microbiol. 41,15–20 (2018). [DOI] [PubMed] [Google Scholar]
- 11.Mitrophanov A. Y., Groisman E. A.,Signal integration in bacterial two-component regulatory systems. Gene Dev. 22,2601–2611 (2008). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Francis V. I., Stevenson E. C., Porter S. L.,Two-component systems required for virulence in Pseudomonas aeruginosa. FEMS Microbiol. Lett. 364,fnx104 (2017). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wang Y., Cao Q., Cao Q., Gan J., Sun N., Yang C.-G., Bae T., Wu M., Lan L.,Histamine activates HinK to promote the virulence of Pseudomonas aeruginosa. Sci. Bull. 66,1101–1118 (2021). [DOI] [PubMed] [Google Scholar]
- 14.Tomlinson K. L., Prince A. S., Wong T., Lung F.,Immunometabolites drive bacterial adaptation to the airway. Front. Immunol. 12,790574 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Malhotra S., Hayes D. Jr., Wozniak D. J.,Cystic fibrosis and Pseudomonas aeruginosa: The host-microbe interface. Clin. Microbiol. Rev. 32,e00138-18 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Smith R. S., Iglewski B. H.,P. aeruginosa quorum-sensing systems and virulence. Curr. Opin. Microbiol. 6,56–60 (2003). [DOI] [PubMed] [Google Scholar]
- 17.Schuster M., Sexton D. J., Diggle S. P., Greenberg E. P.,Acyl-homoserine lactone quorum sensing: From evolution to application. Annu. Rev. Microbiol. 67,43–63 (2013). [DOI] [PubMed] [Google Scholar]
- 18.Wang T., Qi Y., Wang Z., Zhao J., Ji L., Li J., Cai Z., Yang L., Wu M., Liang H.,Coordinated regulation of anthranilate metabolism and bacterial virulence by the GntR family regulator MpaR in Pseudomonas aeruginosa. Mol. Microbiol. 114,857–869 (2020). [DOI] [PubMed] [Google Scholar]
- 19.Lau G. W., Hassett D. J., Ran H., Kong F.,The role of pyocyanin in Pseudomonas aeruginosa infection. Trends Mol. Med. 10,599–606 (2004). [DOI] [PubMed] [Google Scholar]
- 20.Kazmierczak B. I., Schniederberend M., Jain R.,Cross-regulation of Pseudomonas motility systems: The intimate relationship between flagella, pili and virulence. Curr. Opin. Microbiol. 28,78–82 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Dubern J. F., Diggle S. P.,Quorum sensing by 2-alkyl-4-quinolones in Pseudomonas aeruginosa and other bacterial species. Mol. Biosyst. 4,882–888 (2008). [DOI] [PubMed] [Google Scholar]
- 22.Bandera A. M., Bartho J., Lammens K., Drexler D. J., Kleinschwärzer J., Hopfner K.-P., Witte G.,BusR senses bipartite DNA binding motifs by a unique molecular ruler architecture. Nucleic Acids Res. 49,10166–10177 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Horne C. R., Venugopal H., Panjikar S., Wood D. M., Henrickson A., Brookes E., North R. A., Murphy J. M., Friemann R., Griffin M. D. W., Ramm G., Demeler B., Dobson R. C. J.,Mechanism of NanR gene repression and allosteric induction of bacterial sialic acid metabolism. Nat. Commun. 12,1988 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li X., He S., Li R., Zhou X., Zhang S., Yu M., Ye Y., Wang Y., Huang C., Wu M.,Pseudomonas aeruginosa infection augments inflammation through miR-301b repression of c-Myb-mediated immune activation and infiltration. Nat. Microbiol. 1,16132 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Heid M. E., Keyel P. A., Kamga C., Shiva S., Watkins S. C., Salter R. D.,Mitochondrial reactive oxygen species induces NLRP3-dependent lysosomal damage and inflammasome activation. J. Immunol. 191,5230–5238 (2013). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.West A. P., Brodsky I. E., Rahner C., Woo D. K., Erdjument-Bromage H., Tempst P., Walsh M. C., Choi Y., Shadel G. S., Ghosh S.,TLR signalling augments macrophage bactericidal activity through mitochondrial ROS. Nature 472,476–480 (2011). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gao P., Guo K., Pu Q., Wang Z., Lin P., Qin S., Khan N., Hur J., Liang H., Wu M.,oprC impairs host defense by increasing the quorum-sensing-mediated virulence of Pseudomonas aeruginosa. Front. Immunol. 11,1696 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nguyen T.-K., Selvanayagam R., Ho K. K. K., Chen R., Kutty S. K., Rice S. A., Kumar N., Barraud N., Duong H. T. T., Boyer C.,Co-delivery of nitric oxide and antibiotic using polymeric nanoparticles. Chem. Sci. 7,1016–1027 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Dubey M. K., Broberg A., Jensen D. F., Karlsson M.,Role of the methylcitrate cycle in growth, antagonism and induction of systemic defence responses in the fungal biocontrol agent Trichoderma atroviride. Microbiology 159,2492–2500 (2013). [DOI] [PubMed] [Google Scholar]
- 30.Jacob-Dubuisson F., Mechaly A., Betton J. M., Antoine R.,Structural insights into the signalling mechanisms of two-component systems. Nat. Rev. Microbiol. 16,585–593 (2018). [DOI] [PubMed] [Google Scholar]
- 31.Wuichet K., Zhulin I. B.,Origins and diversification of a complex signal transduction system in prokaryotes. Sci. Signal. 3,ra50 (2010). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Curtis M. M., Hu Z., Klimko C., Narayanan S., Deberardinis R., Sperandio V.,The gut commensal bacteroides thetaiotaomicron exacerbates enteric infection through modification of the metabolic landscape. Cell Host Microbe 16,759–769 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Palacios S., Escalante-Semerena J. C.,2-Methylcitrate-dependent activation of the propionate catabolic operon (prpBCDE) of Salmonella enterica by the PrpR protein. Microbiology (Reading) 150,3877–3887 (2004). [DOI] [PubMed] [Google Scholar]
- 34.Kyriakidis D. A., Theodorou M. C., Tiligada E.,Histamine in two component system-mediated bacterial signaling. Front. Biosci. 17,1108–1119 (2012). [DOI] [PubMed] [Google Scholar]
- 35.Munoz-Elias E. J., Upton A. M., Cherian J., McKinney J. D.,Role of the methylcitrate cycle in Mycobacterium tuberculosis metabolism, intracellular growth, and virulence. Mol. Microbiol. 60,1109–1122 (2006). [DOI] [PubMed] [Google Scholar]
- 36.Riquelme S. A., Wong T., Lung F., Prince A.,Pulmonary pathogens adapt to immune signaling metabolites in the airway. Front. Immunol. 11,385 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Yan Y., Wang H., Zhu S., Wang J., Liu X., Lin F., Lu J.,The methylcitrate cycle is required for development and virulence in the rice blast fungus Pyricularia oryzae. Plant. Microbe Interact 32,1148–1161 (2019). [DOI] [PubMed] [Google Scholar]
- 38.Rosenberg G., Riquelme S., Prince A., Avraham R.,Immunometabolic crosstalk during bacterial infection. Nat. Microbiol. 7,497–507 (2022). [DOI] [PubMed] [Google Scholar]
- 39.Rosenberg G., Yehezkel D., Hoffman D., Mattioli C. C., Fremder M., Ben-Arosh H., Vainman L., Nissani N., Hen-Avivi S., Brenner S., Itkin M., Malitsky S., Ohana E., Ben-Moshe N. B., Avraham R.,Host succinate is an activation signal for Salmonella virulence during intracellular infection. Science 371,400–405 (2021). [DOI] [PubMed] [Google Scholar]
- 40.Jiang L., Wang P., Song X., Zhang H., Ma S., Wang J., Li W., Lv R., Liu X., Ma S., Yan J., Zhou H., Huang D., Cheng Z., Yang C., Feng L., Wang L.,Salmonella Typhimurium reprograms macrophage metabolism via T3SS effector SopE2 to promote intracellular replication and virulence. Nat. Commun. 12,879 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Reniere M. L., Whiteley A. T., Hamilton K. L., John S. M., Lauer P., Brennan R. G., Portnoy D. A.,Glutathione activates virulence gene expression of an intracellular pathogen. Nature 517,170–173 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Riquelme S. A., Lozano C., Moustafa A. M., Liimatta K., Tomlinson K. L., Britto C., Khanal S., Gill S. K., Narechania A., Azcona-Gutiérrez J. M., Mango E. D., Saénz Y., Planet P., Prince A.,CFTR-PTEN-dependent mitochondrial metabolic dysfunction promotes Pseudomonas aeruginosa airway infection. Sci. Transl. Med. 11,eaav4634 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jansons I., Touchie G., Sharp R., Almquist K., Farinha M. A., Lam J. S., Kropinski A. M.,Deletion and transposon mutagenesis and sequence analysis of the pRO1600 OriR region found in the broad-host-range plasmids of the pQF series. Plasmid 31,265–274 (1994). [DOI] [PubMed] [Google Scholar]
- 44.Wang T., du X., Ji L., Han Y., Dang J., Wen J., Wang Y., Pu Q., Wu M., Liang H.,Pseudomonas aeruginosa T6SS-mediated molybdate transport contributes to bacterial competition during anaerobiosis. Cell Rep. 35,108957 (2021). [DOI] [PubMed] [Google Scholar]
- 45.Haihua L., Xin D., Xuefeng L., Yan Y., Min W.,Molecular mechanisms of master regulator VqsM mediating quorum-sensing and antibiotic resistance in Pseudomonas aeruginosa. Nucleic Acids Res. 42,10307–10320 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Duan C. D. K., Stein J., Rabin H., Surette M. G.,Modulation of Pseudomonas aeruginosa gene expression by host microflora through interspecies communication. Mol. Microbiol. 50,1477–1491 (2010). [DOI] [PubMed] [Google Scholar]
- 47.Zhao J., Yu X., Zhu M., Kang H., Ma J., Wu M., Gan J., Deng X., Liang H.,Structural and molecular mechanism of CdpR involved in quorum-sensing and bacterial virulence in Pseudomonas aeruginosa. PLoS Biol. 14,e1002449 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hoang T. T., Karkhoff-Schweizer R. R., Kutchma A. J., Schweizer H. P.,A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: Application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212,77–86 (1998). [DOI] [PubMed] [Google Scholar]
- 49.Harunur Rashid M., Kornberg A.,Inorganic polyphosphate is needed for swimming, swarming, and twitching motilities of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 97,4885–4890 (2000). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Huang H., Shao X., Xie Y., Wang T., Deng X.,An integrated genomic regulatory network of virulence-related transcriptional factors in Pseudomonas aeruginosa. Nat. Commun. 10,2931 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Reddick J. J., Sirkisoon S., Dahal R. A., Hardesty G., Hage N. E., Booth W. T., Quattlebaum A. L., Mills S. N., Meadows V. G., Adams S. L. H., Doyle J. S., Kiel B. E.,First biochemical characterization of a methylcitric acid cycle from Bacillus subtilis Strain 168. Biochemistry 56,5698–5711 (2017). [DOI] [PubMed] [Google Scholar]
- 52.Minor W., Cymborowski M., Otwinowski Z., Chruszcz M.,HKL-3000: The integration of data reduction and structure solution--from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62,859–866 (2006). [DOI] [PubMed] [Google Scholar]
- 53.Potterton E., Briggs P., Turkenburg M., Dodson E.,A graphical user interface to the CCP4 program suite. Acta Crystallogr. D Biol. Crystallogr. 59,1131–1137 (2003). [DOI] [PubMed] [Google Scholar]
- 54.Emsley P., Cowtan K.,Coot: Model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60,2126–2132 (2004). [DOI] [PubMed] [Google Scholar]
- 55.Adams P. D., Grosse-Kunstleve R. W., Hung L. W., Ioerger T. R., McCoy A. J., Moriarty N. W., Read R. J., Sacchettini J. C., Sauter N. K., Terwilliger T. C.,PHENIX: Building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58,1948–1954 (2002). [DOI] [PubMed] [Google Scholar]
- 56.Zhou X., Li X., Ye Y., Zhao K., Zhuang Y., Li Y., Wei Y., Wu M.,MicroRNA-302b augments host defense to bacteria by regulating inflammatory responses via feedback to TLR/IRAK4 circuits. Nat. Commun. 5,3619 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Jin X., Zhang W., Wang Y., Liu J., Hao F., Li Y., Tian M., Shu H., Dong J., Feng Y., Wei M.,Pyruvate kinase M2 promotes the activation of dendritic cells by enhancing IL-12p35 expression. Cell Rep. 31,107690 (2020). [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Figs. S1 to S12
Tables S1 to S4