Abstract
RNA-binding proteins (RBPs) have emerged as important players in multiple biological processes including transcription regulation, splicing, R-loop homeostasis, DNA rearrangement, miRNA function, biogenesis, and ribosome biogenesis. A large number of RBPs had already been identified by different approaches in various organisms and exhibited regulatory functions on RNAs’ fate. RBPs can either directly or indirectly interact with their target RNAs or mRNAs to assume a key biological function whose outcome may trigger disease or normal biological events. They also exert distinct functions related to their canonical and non-canonical forms. This review summarizes the current understanding of a wide range of RBPs’ functions and highlights their emerging roles in the regulation of diverse pathways, different physiological processes, and their molecular links with diseases. Various types of diseases, encompassing colorectal carcinoma, non-small cell lung carcinoma, amyotrophic lateral sclerosis, and Severe acute respiratory syndrome coronavirus 2, aberrantly express RBPs. We also highlight some recent advances in the field that could prompt the development of RBPs-based therapeutic interventions.
Keywords: RNA-binding proteins, Neurodegenerative diseases, RNA processing, Post-transcriptional gene regulation, Viral infection, Cancer, RNA
Introduction
RNA-binding proteins (RBPs) represent an important class of post-transcriptional regulators implicated in a plethora of biological processes in several organisms. It has become apparent that the binding function of RBPs is not limited to host survival but is implicated in abnormal cellular processes (e.g. cancer) and essential for some pathogens during infection of the host system. As almost every process in a cell is mediated by proteins and RBPs are posttranscriptional regulators of gene expression involved in stability, translation, or other biological events determining mRNAs fate, an approach focused on the deep characterization of RBPs for therapeutic drug discovery is required. RBPs have been prioritized in biomedical research due to their implication in gene expression and RNA processing, which are among the factors essential for gene function. Human RBPs are further classified in canonical RBPs containing RNA recognition motif (RRM), ribosomal and K-homology (KH) domains (600 structurally different RBPs, e.g. APOBEC3B, C9orf114, DDX58), and some non-canonical RBPs (Fig. 1) recognized by their intrinsically disordered regions (e.g. breast cancer type 1 susceptibility protein BRCA1, CDC40, SETD3) as not all IDR-containing RBPs are "non-canonical" [1]. These domains raised fundamental questions about the distinct role they might play in normal or altered biological events.
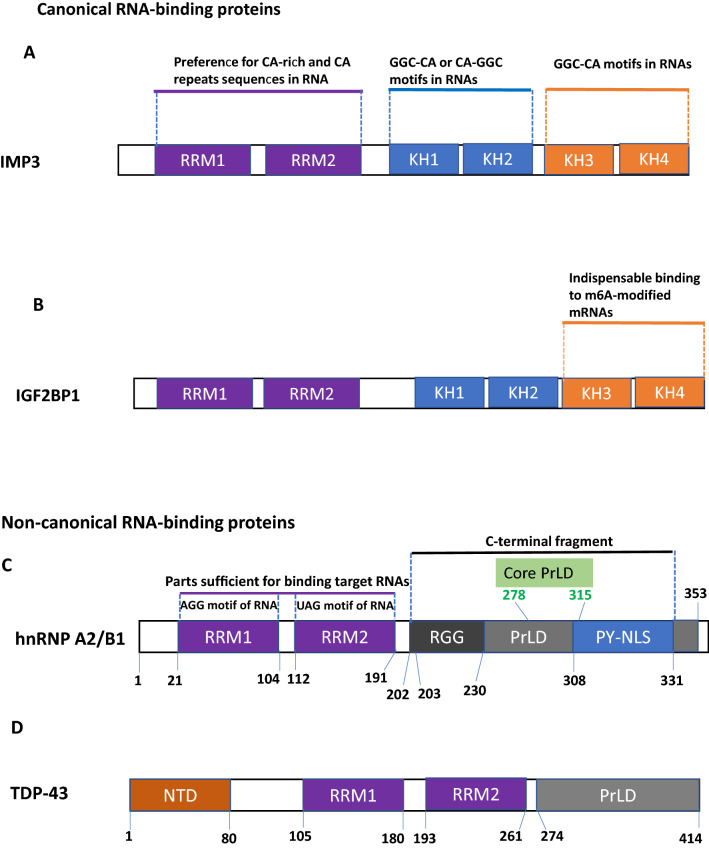
Fig. 1.
Typical domains observed in RNA-binding proteins. A Schematic representation of IMP3 domains. All tandem RBDs are endorsed with RNA-binding activities. RRM1 and RRM2 recognize CA-rich and CA repeats sequences in RNA. KH1-2 specifically recognizes GGC-CA or CA-GGC motifs in RNAs. KH3-4 identify GGC-CA motifs in RNAs [2]. B Schematic representation of IGF2BP domains. All tandem RBDs are identified but only KH domains are specifically used to recognizes m6A-modified mRNAs [3]. C Schematic representation of hnRNP A2/B1 domains with known residue numbers. RNA-binding domain (RBD) composed of tandem RRMs separated by a 15-aa linker, and a C-terminal Gly-rich low complexity (LC) region that includes a prion-like domain (PrLD), an RGG box: arginine-glycine-glycine box, and a PY-motif containing a M9 nuclear localization signal (PY-NLS). RRM1 specifically recognizes AGG motif in RNAs. RRM2 specifically recognizes UAG motif in RNAs but also purine-rich GAG, and pyrimidine-rich UU sequences [4]. D Schematic representation of TDP-43 RRM domains. 414-residue TDP-43 includes: N-terminal domain (NTD) over residues 1–80; two tandemly-tethered RNA recognition motifs (RRM1 and RRM2) over residues 105–261 connected by an unstructured linker over residues 181–192, and C-terminal prion-like domain over residues 274–414 [5]
RBPs can be classified based on their binding preferences [6], as single-stranded RNA (ssRNA) and double-stranded RNA (dsRNA). Amongst RBPs, the double-stranded RBPs TAO kinases (TAOK1, TAOK2, and TAOK3) are particularly critical for antiviral protein expression and/or antiviral signaling cascade regulation in human cells, with TAOK2 also being antiviral against Semliki Forest virus and Herpes-Simplex virus 1 [7]. The single-stranded RNA-binding protein coronaviral nonstructural protein 9 (Nsp9) contains a conserved self-associating GxxxG sequence motif within its C-terminal α-helix, the residues Phe-40, Val-41, as well as Ile-91 prone to perturbations, and N-terminus for high affinity for oligonucleotide [8]. For the ssRNA-binding protein, Nsp9, known to form an obligate homodimer via an essential GxxxG protein-interaction motif, in an unmodified state, it has been established that its ssRNA-binding mechanism used a pyrimidine RNA base-mimic compound [8].
The underlying mechanism implicated in the recruitment and binding of RBPs to the target mRNAs in several diseases is still under investigation. It is crucial to explore the regulation of RBP's interaction with the binding site of target mRNAs so that novel therapeutic strategies for the treatment of diverse RBPs-related diseases could be proposed.
Ribonucleic acid (RNA) therapeutics, exploiting messenger RNAs (mRNAs), small interfering RNAs (siRNAs), and micro RNAs (miRNAs), have garnered considerable interest as an approach to rapidly overcoming several diseases in modern medicine. Besides, recent reports have highlighted the promising role of mRNA vaccines in treating the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic [9, 10]. Further, the messenger RNA-based vaccine candidate encoding the SARS-CoV-2 spike glycoprotein receptor-binding domain (BNT162b1) has been revealed to display safety and immunogenicity results on vaccinated younger (aged 18–55 years) and older adults (aged 65–85 years) Chinese participants [11]. Vaccination with mRNA holds the potential to transform modern medicine by allowing it to overcome the existing bottleneck of conventional vaccines in the prevention of infectious diseases [12, 13]. Yet the reduced efficacy linked to poor RNA stability in solution is among the technical obstacles facing mRNA therapeutics [14, 15]. Recently some cases of infection with SARS-CoV-2 in vaccinated individuals have been reported [16], and post-SARS-CoV-2 mRNA vaccination responses exhibited a highly variable humoral and cell-mediated immunity across the range of immunodeficiency disorders [17].
Additionally, mRNAs have been identified to be presented as complexes with a diverse range of RNA-binding proteins (RBPs) where they contributed to the construction of messenger ribonucleoprotein (mRNP) complexes [18–20]. Therefore, efforts towards the production of therapeutic molecules in addition to vaccines (e.g. therapeutic antibodies) that would neutralize the target pathogen and impede its interaction with cellular receptors to gain entry into cells could be the object of much attention. Besides, it has already been envisioned for the treatment of COVID-19 [21]. RBPs represent an important class of post-transcriptional regulators implicated in a plethora of biological processes in several organisms. It has become apparent that the binding function of RBPs is not limited to host survival but is implicated in abnormal cellular processes (e.g. cancer) and essential for a diverse array of viruses during infection of the host system. The same RBP activity can be shared by at least two diseases. For example, beyond the role notified for the RBPs of Musashi (MSI) protein family in diverse cancer cells, spanning breast, colorectal, lung, and pancreatic cancers, as well as leukemia or glioblastoma [22], an additional role as host factors stimulating the replication of a mosquito-borne Flavivirus Zika virus (ZIKV) has been recognized for them [23]. The recent identification of 9 shared RBPs that connected COVID-19 to neurological disorders and diverse types of cancers at the molecular level prompted us to look into the role of RBPs in neurodegenerative diseases, cancer progression, and viral infection. For example, Ref. [24] unveiled that the RBPs RPS10, LIN28B, MOV10 (Moloney murine leukemia virus infection in mice), RPS3, EIF4B, EIF5A, EEF1A1, RPL18A, and Poly(A) Binding Protein Cytoplasmic 1 (PABPC1) are jointly involved in COVID-19, cancer, and neurological disorders [24]. Furthermore, it has been revealed that the RBP TDP-43 previously known for its importance in neurodegenerative disease progression, also played a key role in hepatocellular carcinoma progression. Indeed, TDP-43 binds to the UG-rich sequence1 of the 3’UTR of ABHD2 (abhydrolase domain containing 2) mRNA to improve the stability of ABHD2 mRNA stimulates the synthesis of free fatty acid and fatty acid oxidation-derived reactive oxygen species in an ABHD2-dependent manner, to oppose apoptosis and improve the proliferation of hepatocellular carcinoma cells [25]. Taking together, the aforementioned roles of RBPs turned them into essential targets to reveal the mechanism of a plethora of diseases.
This review is particularly important as it highlights the strong effect of RBPs in several pathways and mechanisms that can protect from or lead to disease, pinpoints the sequences from RNAs that can be monitored to find long-lasting treatments, offers strategies to control RBP activity in time and space, and reports methods to enrich subcompartment-specific RBPs.
General roles attributed to RNA-binding proteins
RBPs are of interest since their activities are involved in the direct monitoring of a diverse array of events spanning transcription, RNA splicing, polyadenylation, stabilization, localization, and translation [26], and their aberrant functions are implicated in a variety of diseases such as viral infection, cancer, and neurodegenerative disorders. They are among the diverse human innate immune response mechanisms that may trigger signaling cascades in the host to halt or allow pathogen establishment in the cell. A growing body of evidence supports the likelihood that RBPs exert a major influence on cellular events to drive cell function, and that a mechanistic understanding of RBP's function during disease progression is crucial for the development of new disease therapeutics. RBPs affected alternative splicing regulatory events (e.g. TDP-43-mediated mutually exclusive exons, skipped exons, alternative 5' and 3' splice sites, and intron retention [27], translational efficiency events, pre-mRNA processing, mRNA nuclear export and retinal development, sterility mechanisms, stress granules management, R loop formation and are identified in processes occurring in neurodegenerative disease, cancer, and viral infection. For example, in addition to their central role in the monitoring of gene expression through changes in RNA stability and degradation, RBPs are also implicated in DNA damage response regulation and one of them, HNRNPC (heterogeneous nuclear ribonucleoprotein C), is identified as a regulator of chemotherapeutic resistance in human lung cancer [28]. Furthermore, this latter finding has clear clinical relevance for the response of lung cancer to chemotherapy in human patients. The different roles attributed to RBPs are depicted in Fig. 2. Furthermore, in recent times there are many emerging microbial infections with obscure immune invasion properties and no effective therapeutic targets available to limit their fast expansion or outbreaks. Therefore, the design of strategies focusing on the proteins targeted by these pathogens in the host could bring more therapeutic solutions for their control.
Fig. 2.
Numerous cellular events and disease processes regulated by RNA-binding proteins. The RBP PUM1 binds the 3´-untranslated region of Toll-like receptor 4 (TLR4) to clear TLR4 mRNA translation and monitor the activity of nuclear factor-κB (NF-κB), a master regulator of the aging process in human mesenchymal stem cells. PUM1 also protects human chondrocytes to avoid chondrogenic phenotype loss and alleviates cellular osteoarthritis [29]. The loss of functional RBP muscle blind-like protein 1 (MBNL1) in response to its nuclear seclusion by mutant transcripts containing pathogenic expanded CUG repeats (CUGexp) leads to Myotonic dystrophy type 1 [30]. Mitochondrial RNA-binding protein tumor necrosis factor receptor-associated protein 1 (TRAP1)-associated, chaperone-mediated autophagy can stimulate ferroptosis [31]. The RBP human antigen R (HuR) translocation into the cytoplasm orchestrates by its interaction with circular RNAs circStag1 is involved in the regeneration of bone tissue [32]. Inclusion of the RBP TDP-43-regulated alternative exons is altered in skeletal muscles of patients with inclusion body myositis (IBM, [27]). RBP ZFP36 and ZFP36L1 bind transcripts encoding subunits of the NF-κB pathway, Notch1, Irf8 and Il2 to monitor the abundance of their protein products early after activation in CD8 + T cells [33]. Heterozygous frameshift variants in the RBP hnRNPA2/B1 do not augment the propensity of hnRNPA2 protein to fibrillize but diminished affinity for the nuclear import receptor karyopherin β2, leading to cytoplasmic accumulation of hnRNPA2 protein in cells and early-onset form of oculopharyngeal muscular dystrophy [34]. HNRNPC (heterogeneous nuclear ribonucleoprotein C) associated with the newly identified regulator of ferroptosis in Colorectal cancer, CUL9, for resistance to drug-induced ferroptosis [35]. The RBP heterogeneous nuclear ribonucleoprotein RBP (hnRNPH1) is essential for pre-mRNA alternative splicing, spermatogenesis and oogenesis [36]. RBP Tardbp is an important candidate monitoring alternative splicing and alternative polyadenylation to optimize clonal expansion and human CD8 + T cells effector function during an antigen-specific immune response [37]. In embryonic stem cells, RBPs represent half of the chromatin proteome, co-compartmentalize with RNA polymerase (Pol) II at promoters and enhancer, and connect RNA to the transcription machinery [38]. The RBP DDX41 is a tumor suppressor that opposes double-strand DNA breaks, genomic instability and R-loop-dependent replication stress by preferentially binding RNA–DNA hybrids and unwinding RNA–DNA hybrids in R-loops, and also reduces fragility of DNA in promoter regions [39]. RBP HNRNPC is demonstrated to interact with circXRCC5 to encourage circXRCC5 biogenesis that was proved to foster Gastric cancer progression [40]. By binding to hTERT (reverse transcriptase protein catalytic subunit hTERT) mRNAs in intron 8, the RBP NOVA1 (neuro-oncological ventral antigen 1) acts as a splicing enhancer that promotes the inclusion of exons 7 and 8 to enhance the production of full-length (FL) hTERT mRNAs in non-small cell lung cancer cells [41]. The RBP “partner of NOB1” (PNO1)-mediated ribosome biogenesis is negatively regulated by miR-340-5p directly binds the 3′-UTR of PNO1 and assumes an important role in LUAD progression via Notch signaling pathway [42]. RBP (ROD1, so-called PTBP3) interacts with Activation-induced cytidine deaminase (AID), which jointly bind bi-directionally transcribed RNAs to facilitate genome-wide AID targeting to immunoglobulin (Ig) loci in a way to induce DNA rearrangement during immune responses [43]. The RBP KHSRP regulates cell proliferation and cell cycle in Wilms tumor by modulating the expression of PPP2CA and p27 [44]
Human RBPs are reported to frequently contain specific low-complexity motifs and bind various coding along with non-coding RNAs (ncRNAs) types, including messenger RNA (mRNA), transfer RNA (tRNA), small nuclear RNAs (snRNAs), small nucleolar RNAs (snoRNAs), ribosomal RNA (rRNA), internal ribosome RNA (irRNA), the novel class of RNAs circular RNAs (circRNAs), RNA polymerase II [45]. CircRNAs are noticeable for their role in cancer progression. For example, RBP Polypyrimidine tract binding protein 1 (PTBP1) is reported to repress the expression of circ_0076611, whose interaction with diverse proliferation-related transcripts (MYC and VEGFA mRNAs) augments cell proliferation and migration in triple-negative breast cancer cells, the RBP oncogenic splicing factor SRSF1 (Serine And Arginine Rich Splicing Factor 1) encourages this expression after the inactivation of PTBP1 protein by the long non-coding RNAs (lncRNA) MALAT1 (metastasis-associated lung adenocarcinoma transcript 1 [46]. Several novel RBPs regulating circRNA functions in tumorigenesis, such as SRSF5, HNRNPA2B1, HNRNPF, KHSRP, or PSIP1, have also been unveiled [47].
The discovery of different types of RNAs targeted by RBPs makes the overall investigation of RBP's target sites more challenging. It is established that RBPs regions that bind mRNA, rRNA, and snRNA are enriched in disordered proteins, while regions interacting with ncRNA and irRNA, and tRNA are depleted in disorder proteins [48].
Binding preference and activity of RNA-binding proteins
The function of RRMs as well as the residues they contain have been investigated. For example, the truncation in the C-terminal region identified as the RNA-binding domain of the RBP (Hel2), not only altered the ability of Hel2 to bind to 18S rRNA and translated mRNAs but also led to a loss of polysomal association and monitoring of both Ribosome-associated quality control (RQC) pathways and no-go decay [49].
Moreover, emerging pieces of evidence have shown that RBPs deployed their multidomain to recognize specific target sequences in RNAs. For example, the multidomain RBP, insulin-like growth factor 2 mRNA-binding protein 3 (IMP3, also known as IGF2BP3), used six RNA-binding domains (four KH and two RRM domains) to identify CA-rich and GGC-core elements in its targets RNA [2]. Furtherly, the RBP IGF2BP3 can directly augment the mRNA stability of E2F3 by binding the Long noncoding RNAs, LINC00958, and independently bind to the 3’UTR region of E2F3 to stimulate endometrial carcinoma progression [50]. Additionally, IGF2BP3 has the potential to post-transcriptionally control the oncogenic activity of MLL-AF4 protein; an oncogenic driver of MLL-rearranged leukemia [51].
RBPs preferentially bind sequences in 5′ or 3′ untranslated regions (UTRs) or RNA folds in the coding regions of mRNAs and orchestrate stability and translation efficiency [52]. It has been recently displayed that somatic 3′-UTR mutations (3′-UTR single nucleotide variants) in a given sequence motif (e.g. AACAAA) residing in mRNA could disrupt the binding between RBPs (e.g. SRSF1 and PTBP1) and their corresponding RNAs. The abnormal expression of RBPs resulting from this dysregulation is reported broadly in seven cancer types including biliary adenocarcinoma, breast adenocarcinoma, kidney renal cell carcinoma, liver hepatocellular carcinoma, prostate adenocarcinoma, stomach adenocarcinoma, as well as our in-house esophageal squamous cell carcinoma [53]. Mechanistically, the RBP La-related protein 6 (LARP6) is demonstrated to bind 5′ stem-loop (5′SL) of collagen mRNAs with its La domain-containing RNK-motif to stabilize the complex between La domain and 5′SL RNA using its juxtaposed RNA recognition motif and to stimulate the excessive synthesis of type I collagen leading to fibrotic diseases [54].
The binding properties and function of some RBPs on their targets can be sequence-specific. For example, the RBP DHX36 is an indispensable post-transcriptional regulator of stem cell proliferation and muscle regeneration acting via binding and unwinding G-quadruplex (rG4) structures at 5′ UTR of target mRNAs [55]. The identity of some RNAs' motifs has been determined for a subset of RBPs. A report elucidated that the RRM1 and RRM2 domains of the human RBP hnRNP A2/B1-mediating maturation or transport of mRNA as well as gene regulation of long noncoding RNA, recognized the AGG and UAG motifs, respectively, in their target RNAs [4].
Georgakopoulos-Soares and colleagues [56] demonstrated that the RBPs HNRNPK and HNRNPU displayed higher binding to G-quadruplex (G4) motif around splice sites, are positively connected to differentially included exons and have a direct impact on G4-mediated alternative splicing (AS) regulation. On the other side, RBPs such as AQR and RBM15 are depleted around splice sites flanked by G4 motifs and are negatively connected to differentially included exons, and the binding and impact of these regulatory factors over AS are suggested to be repressed by G4 formation [56]. The same authors further revealed that exon–intron junctions in humans exhibited an acute enrichment for G4s motifs, both at the 3′ss and the 5′ss, 31% of human genes displayed a G4 motif near at least one splice site within a distance of 100 bp, alternative splicing modulation by G4s is constricted to mammals and birds, but absent in plants, other tetrapods or fish. This data highlights the key regulatory roles exerted by RBPs on alternative splicing events known as a pivotal step of eukaryotic mRNA processing during gene expression.
Besides binding to nucleic acids [57] identified that the RRM of RBPs can bind to proteins. For example, the RRM2 of the pre-mRNA processing hnRNP proteins member, hnRNP L, forms a ternary complex with RNA as well as with the methyltransferase protein SETD2. For the interaction with SEDT2 to take place, leucine pair within a highly conserved stretch of SETD2 insert their side chains in hydrophobic pockets formed by hnRNP L RRM2. The same authors highlighted how specific interactions between RBPs and other proteins allowed the coupling of transcription and splicing. Indeed, they uncovered that the histone methyltransferase SETD2 collaborates with Pol II and interacts with RRM2 domains of the RBP hnRNP L via its SETD2 hnRNP interaction (SHI) domain to allow the deposition of the histone mark H3K36me3-regulated splicing, thereby mediating the coupling of splicing and transcription in mammalian cells [57].
Additionally, mutations in RNA-binding motifs drive disease progression such as cancer progression. The RBP U2AF1 carries hotspot mutations in its RNA-binding motifs that create new 3′ splice site contacts (at − 3 nucleotides) altering RNA splicing, illuminating how mutations in RBPs’ motifs affect splicing and promote cancer [58].
Some RBPs required a certain modification (e.g. post-transcriptional mRNA modification N6-methyladenosine m6A) of their target mRNAs to act properly. N6-methyladenosine (m6A) is commonly detected methyl-marks on RNA (e.g. mRNA, rRNA) and its deposition is catalyzed by RNA methylation writer enzymes, so-called RNA methyltransferases (e.g. complex METTL3-METTL14 complex acting on m6A deposition; [59]. A study revealed that the RBPs IGF2BPs preferentially recognize m6A-modified mRNAs, promote the stability of thousands of potential mRNA targets in an m6A-dependent manner under normal as well as stress conditions, and their oncogenic roles in cervical, as well as liver cancers, depending on their role as m6A readers [60]. It has been also evidenced that the RBP, Fragile X mental-retardation protein (FMR1), preferentially binds m6A-modified “AGACU”-containing mRNAs. The presence of m6A-modified RNA augments the size of FMR1 granules, improves the partitioning of non-methylated RNA into the granules, and facilitates the role of FMR1 in maternal mRNA decay [61]. On the other hand, in human cell lines, the DEAH-box helicase DHX36/RHAU is reported to preferentially bind the G-rich and G4-forming sequences in mRNAs and unwinds the typically formed rG4s to increase translational efficiency [62].
All the aforementioned events prompt the detailed analysis of RBPs' actions in mammals, especially from a disease development viewpoint.
RBPs assume diverse types of functions in mammalian cells
A thorough report on the role of RBPs in humans has been associated with neurodegenerative disease, viral infection, and cancer.
Contribution of dysregulated RBPs to neurodegenerative diseases
RBPs are involved in the metabolism of neurodegenerative diseases demonstrated recently to share a feature with the neurological effects of COVID-19; the occurrence of cytotoxic aggregated amyloid protein or peptides [63]. RBPs notified in cellular mechanisms altered in neurodegenerative diseases (mislocalization of RBPs), played a role in multiple RNA processing stages spanning transcription (FUS), turnover (MATR3), alternative splicing (TDP-43, FUS, hnRNPA1, hnRNPA2/B1), and stress granules; TDP-43 [64–66].
In addition to the commonly possessed RNA interacting motifs, RBPs often harbor IDRs that have been noticed to be involved in forming membrane-less organelles (such as stress granules SGs), which assemble via liquid–liquid phase separation (LLPS). SGs have a physiologically relevant role as many disease-linked RBPs (FUS, hnRNPA1, and hnRNPA2) identified inside contained intrinsically disordered low-complexity domains that can be prion-like domains (PrLDs) whose mutations accelerated fibrillization and caused disease [67]. For example, the RBP TDP-43 whose activity is dysregulated in neurodegenerative disease contained the two highly conserved RNA recognition motifs (RRM1 and RRM2), a short polyalanine tract and an intrinsically disordered region (IDR) harboring an “AMMAAAQAALQ” amino acid motif prone to form an α-helix promoting condensation. Its IDR differs from hnRNP A1 or A2’s as it is prion-like and possesses a short α-helical region promoting self-association and LLPs. The LLPS is driven by just a few aromatic residues, a feature that has been attributed to the presence of the α-helix stimulating inter-molecular contacts [68]. In healthy cells, TDP-43 proteins normally egress nuclei by passive diffusion, independent of facilitated mRNA export, and their nuclear localization relies on binding to GU-rich nuclear RNAs [69]. Table 1 highlighted some RBPs connected to neurodegenerative disease and the effect of their inhibitors. On the other hand, Fig. 3 shows illustrative examples of representative diseases implicating intrinsically disordered regions.
Table 1.
Contribution of dysregulated RNA-binding proteins to neurodegenerative diseases
| Dysregulated RBPs | Biological events | Disease case | References | |
|---|---|---|---|---|
| hnRNPA2/B1 | Missense variants in the LCD | MSP or Paget’s disease of bone | [70] | |
| Heterozygous frameshift variants in the LCD partially impairing nuclear import and augmenting cytoplasmic concentration of protein without changing its intrinsic propensity to fibrillate | Early-onset oculopharyngeal muscular dystrophy | |||
| TDP-43 | Mislocalization to the cytoplasm due to the impaired structural integrity of the nucleus connected to the translation of repeat expansion in C9orf72-linked neurodegeneration disease into dipeptide repeat proteins | ALS or FTD | [71] | |
| [72] | ||||
| Cytoplasmic mislocalization and LLPS due to RNA depletion | ALS | [73] | ||
| FUS | Oligomerize into condensates | [74] | ||
| Synaptic localization of FUS, in absence of aggregation, triggered synaptic impairment | ALS | [75] | ||
| MATR3 | Prone to phase separation into droplet-like structures | [76] | ||
| Inhibitors of dysregulated RBPs | Anti-TDP-43 aggregation activity of TDP-43 bearing a missense mutation A315T | Acridine derivative (AIM4) | [73] | |
| Impairment of the TDP-43 prion-like domain proneness to aggregation | ATP with concentrations > mM | [5] | ||
| Suppression of aberrant phase separation and solidification of TDP-43 | ALS and FTD | C-terminal hyperphosphorylation of TDP-43 | [77] | |
| Inhibition of toxic misfolding effects of both TDP-43 and FUS | Single missense mutation in nucleotide-binding domain 1 (NBD1) or NBD2 of the AAA + protein disaggregate, Hsp104 | [78] | ||
| Targeted cNLS (a canonical NLS) to repress and reverse | Nuclear-import receptor Karyopherin-β1 cooperating with Importin-α | [67] | ||
| TDP-43 fibrillization | ||||
| Targeted PY-NLSs (PY-nuclear localization signal (NLS)) to repress and reverse TAF15, EWSR1, hnRNPA1, and hnRNPA2 fibrillization | Karyopherin-β2 | [67] | ||
| Binding to the C-terminal proline-tyrosine nuclear localization signal (PY-NLS) of FUS and impairment of FUS self-association, blocking LLPS | Karyopherin-β2/transportin-1 | [79] | ||
| Other inhibition method | Depletion of cytosolic RBP (ATXN2) | ALS and FTLD | Protection against TDP-43 neurotoxicity | [80] |
Fig. 3.
Architecture of typical RNA-binding proteins with low complexity domain localized in intrinsically disordered proteins. The figure highlighted RNA recognition motifs 1 and 2 (RRM1 and RRM2), a low complexity domain (LCD), and a nuclear localization signal (NLS) within the LCD. Missense mutations (G287S, A315T, G348C, R361S, A382T, N390D, N390S) in LCD are responsible for amyotrophic lateral sclerosis [81]. Pathogenic mutations (c.785/941A.T, p.D262V/D314V in hnRNPA1 and c.869/905A.T, p.D290V/D302V in hnRNPA2B1) in prion-like domains of hnRNPA2B1 and hnRNPA1 are responsible for multisystem proteinopathy (MSP; [34]. Missense mutations (P362L, A381T, and E384K) impacting the LCD of the RNA-binding protein T cell-restricted intracellular antigen-1 (TIA1) drive ALS and Frontotemporal Dementia (FTD) [82]. RBMX’s low-complexity region consists of a serine and arginine-rich region (SRR) followed by a tyrosine-rich region (TRR) is required for monitoring the expression of CBX5 mRNA encoding CBX5 protein in the hematological malignancy acute myeloid leukemia (AML) cells [83]
Dysregulated role of RBPs in viral replication and infection
Certain viruses are a serious public health threat responsible for frequent morbidity and mortality in humankind. The antiviral activity of RBPs is stimulated by both RNA viruses and RNAs transcribed from DNA viruses [84]. Elucidating the molecular interplay between viral RNAs and host cell proteins during infection can enhance our understanding of viral RNA functions and the host's innate immune defenses. A virus can remodel host metabolism for successful viral infection and transient stimulation of cancer in cancer virus conditions. A set of RBPs involved in viral metabolism has been gathered in Table 2. Based on these data, deploying programmable RBPs is conceivable to strengthen human immune responses.
Table 2.
Contribution of dysregulated RNA-binding proteins to viral infection
| Dysregulated RBPs/Inhibitors | Biological events | Virus type | References |
|---|---|---|---|
| Interferon-induced protein with tetratricopeptide repeats 2 (IFIT2) | Targeted by virus to sustain viral gene expression and viral replication | Influenza virus | [85] |
| Heterogeneous nuclear ribonucleoprotein AB (hnRNPAB) | Impaired influenza A virus replication via the inhibition of the nuclear export of viral mRNA | Influenza A virus | [86] |
| EIF4G1, EIF4G3, EIF4A1, EIF4A2, EIF4B, PABPC1, HNRNPA1, LSM14A, PATL1, DDX6, DDX60, DHX57, HSP90AA1 | Interactors of SARS-CoV-2 RNA | Severe acute respiratory syndrome coronavirus 2 | [87] |
| PPIA or cyclophilin A, PA2G4, ZC3H11A, DDX3, and HSP90AB1 | Highly upregulated by SARS-CoV-2 as well as Sindbis infections | SARS-CoV-2 and Sindbis (SINV) | [87] |
| IGF2BP1 | Binds the SARS-CoV-2 RNA genome, stabilizes RNA, encourages the translation of SARS-CoV-2 RNA, and regulates SARS-CoV-2 infection | SARS-CoV-2 | [88] |
| Promotes the translation of Zika virus RNA | |||
| Heat shock protein HSP90AB1 | Binds SARS-CoV-2 RNA and monitors infection of SARS-CoV-2 | SARS-CoV-2 | [88] |
| KSHV-encoded viral interferon regulatory factor 1 (vIRF1) | Triggered by KS-associated herpesvirus (KSHV) infection, upregulated and recruited E3 ubiquitin ligase Kelch-like 3 (KLHL3) to decompose heterogeneous nuclear ribonuclear protein Q1 (hnRNP Q1) via a ubiquitin–proteasome pathway, which destabilized GDPD1 mRNA | KS-associated herpesvirus | [89] |
| Poly (rC) binding protein 2 (PCBP2) | Targeted by microRNA-like small RNA (miR-HA-3p) encoded by the H5N1 influenza virus that suppressed its expression and promoted cytokine production in human macrophages infected with the H5N1 virus | H5N1 influenza virus | [90] |
| Ebola virus viral protein 35 (VP35) | Contained interferon inhibitory domain (IID) and a cryptic pocket allosterically coupled to a key dsRNA-binding interface involved in Ebola infection | Ebola Virus | [91] |
| GRSF1 | Allowed efficient cytosolic accumulation and translation of viral mRNAs | Avian influenza A virus | [92] |
| DEAD-box helicase 6 | Interactor of Zika virus–derived subgenomic flavivirus RNA for compromising host response to viral infection | Zika virus | [93] |
| The isoform of the canonical translation factor eukaryotic translation initiation factor 4γI (DAP5) | Essential for the initial round of translation of the enteroviral (Coxsackievirus B3) RNA | Coxsackievirus B3 | [94] |
| EB2 | Essential for stabilizing viral mRNAs generated from intronless genes in the nucleus and trigger their export to the cytoplasm along with their translation | Virus | [95] |
| TRIM25, TRIM56, ZC3HAV1/ZAP, DHX36, and GEMIN5 | Involved in the anti-viral response against both SARS-CoV-2 and SINV | SARS-CoV-2 and SINV | [87] |
| HSP90 and IGF2BP1 | Drugs targeting these RBPs significantly repress SARS-CoV-2 protein production and infection | SARS-CoV-2 and SINV | [87] |
| Nuclear Factor 90 (NF90) | Bound to the 5’UTR and regulated the abundance of the Interferon Regulatory Factors (IRFs) IRF3 and IRF9 mRNAs encoding IRF3 and IRF9 proteins that are found vital for the monitoring of the interferon pathway implicated in the immunological response to the SARS-CoV-2 infection | SARS-CoV-2 | [96] |
| La-related protein 1 (LARP1) | Bound to the 5′-leader containing TOP-like sequence motif in SARS-CoV-2 mRNA and impeded replication in infected human cells under dynamic phosphorylation by the mTORC1 complex | SARS-CoV-2 | [97] |
| Interferon-induced zinc-finger antiviral protein (ZAP-S) | Interacted with the SARS-CoV-2 RNA through the SL2 and SL3 regions of the SARS-CoV-2 pseudoknot and interfered with the folding of the frameshift RNA element to repress programmed ribosomal frameshifting and impair viral replication | SARS-CoV-2 | [98] |
| Peptidyl-prolyl cis–trans isomerase A, sodium/potassium -transporting ATPase subunit alpha-1 and actin-related protein 2 complex (ARP2/3 complex) | Decreased viral replication in human cell lines | Virus | [97] |
| ZFP36 and ZFP36L1 | Their defects led to superior cell-intrinsic cytotoxicity and conferred the greatest protection against influenza A virus infection | Influenza A virus | [33] |
| Human Silencing Hub complex (TASOR) | Facilitating the association of RNA degradation proteins with RNA polymerase II is implicated in the human immunodeficiency virus proviral expression repression | Human immunodeficiency virus | [99] |
Furthermore, a recent role demonstrated for RBPs in defending the host system against viral infection and promoting the expression of genes (IFIT1, IFIT2, IFTI3, IFI44, HERC4, OASL) endowed with antitumoral effects (e.g. multifunctional RBP Raly; [100]), led us to highlight the diverse impact of RBPs in tumorigenesis.
Crucial role played by dysregulated RNA-binding proteins in tumorigenesis and cancer progression
Cancers are often characterized by frequent genetic abnormalities, inhibition of tumor suppressor genes, molecular alterations, high resistance to existing therapies, and recently by the overwhelming presence of 3′-UTR posttranscriptional impairment-related single nucleotide variants (3′-UTR piSNVs) that disturbed RBPs binding, APA and m6A RNA modification [53]. Some cancers are associated with the worst prognosis with decreased overall survival rates. For example, it has been demonstrated that left-sided breast cancer is linked to aggressive biology and worse sequels as opposed to right-sided breast cancer [101]. Further investigations revealed that glioblastoma is a highly aggressive brain tumor displaying poor outcomes with a median survival time of approximately 1-year, a 5-year survival rate of only 5.8%, and a major therapeutic challenge [102]. An ideal medical treatment for a given cancer should accurately address its complexity and its resistance to standard therapies. In line with this statement, cancer interacting partners (e.g. RBPs) may represent a therapeutic opportunity for identifying sustainable solutions for patients with advanced or aggressive diseases. RBPs have emerged as important contributors to tumorigenesis and tumor progression. For example, the RBPs PUMILIO, comprising the two human homologs PUM1 and PUM2, have been demonstrated to suppress the function of the cell cycle regulators p21 and to exert a critical role in human colorectal cancer progression. Their inactivation resulted in colorectal tumor growth reduction and a delay in G1/S transition [103]. These data illustrated the oncogenic roles of some RBPs and opened new therapeutic perspectives for targeting cancers in humans.
It has been provided evidence that the RBP ADAR3 (Adenosine deaminases that act on RNA) increased Nuclear factor-κB (NF-κB) activation and a gene expression program favorable for glioblastoma tumors progression [104]. The molecular mechanism underlying the role of other tumor-promoting RBPs has also been established. Indeed, it has been unveiled that the RBP zinc finger CCHC domain-containing protein 4 (ZCCHC4) exerted its antitumor effect by impeding DNA-damage-induced apoptosis in hepatocellular carcinoma (HCC) cells through the inhibition of the pro-apoptotic function of the new long noncoding RNA (AL133467.2) and also interrupts the interaction between AL133467.2 and γH2AX upon DNA-damage agent (DDA) treatment to repress apoptotic signaling and stimulate chemoresistance to DDAs. Its defects stimulate AL133467.2 and γH2AX interaction for improving chemosensitivity in HCC cells [105]. Another study pointed out that the RBP Paraspeckle component 1 (PSPC1), exerted a crucial role in the proliferation of estrogen receptor (ER)-positive breast cancer cells by interacting with the RBP belonging to Drosophila behavior human splicing family, PSF, and affecting the post-transcriptional regulation of PSF target genes, ESR1 and SCFD2 [106].
Additional roles have been attributed to RBPs in tumorigenesis where they exerted multiple functions spanning tumor initiation, chemoresistance, tumor suppression, R-loop clearance, and circular RNAs biogenesis. Yet, the RBPs monitoring the activity of the tumor suppressors (e.g. p53) is particularly important to analyze.
The RBP named RBM28, known as a nucleolar component of spliceosomal small nuclear ribonucleoproteins, promotes the survival and growth of cancer cells by both translocating from the nucleolus to the nucleoplasm upon phosphorylation and interacting with the DNA-binding domain of tumor suppressor p53 to inhibit p53 transcriptional activity [107]. Moreover, the direct binding of the RBP, LIN28B, to the 5′-UTR of p53 mRNA is reported to impede the translation of p53 by preventing the binding of the translation enhancer protein, ribosomal protein L26 (RPL26), to 5′UTR of TP53 mRNA, thereby facilitating the resistance of cancer cells to p53–MDM2 protein–protein interaction inhibitors (Nutlin3a and MI773, [108]. These two RBPs can be therefore the perfect potential biomarkers and therapeutic targets for cancer inhibition. The tumor-suppressive RBP, hnRNP-E1, is known to modulate splicing and translation of the epithelial to mesenchymal transition (EMT)-associated transcripts and to play a major role in the control of epithelial cell plasticity during cancer progression. The ARIH1, an E3 ubiquitin ligase, can target and interfere with the activity of this RBP by degrading it and promoting breast cancer progression [109]. The RBP DDX41 is a tumor suppressor that opposes double-strand DNA breaks, genomic instability, and R-loop-dependent replication stress by preferentially binding RNA–DNA hybrids and unwinding RNA–DNA hybrids in R-loops, and also reduces the fragility of DNA in promoter regions [39]. It joins therefore the repertoire of RBPs capable of suppressing R-loop formation in human cells.
In the disease suppression pathways, RBPs can regulate microRNA’s (miRNA) function in cancer cells. Indeed, the RBP, Heterogeneous nuclear ribonucleoprotein A2/B1 (HNRNPA2B1), binds to the UUAGG motif of the stem structure, formed by the conserved cis-elements of miR-506 (UGCCUUA) and HNRNPA2B1 binding sites (UUAGG) in the CDK6 3′-UTR, to denature it and recruit the RNA helicase DExH-box helicase 9 (DHX9) to the 3’-UTR, which ultimately facilitates miRNAs-mediated suppression of CDK6 protein expression in lung cancer cells [110].
In addition to their involvement in pathways leading to disease suppression, RBPs are found in pathways resulting in disease activation. An intact RNA-binding motif is necessary for the EXOSC9 protein (RNA exosome constituent) to promote tumor growth and mediate stress resistance as well as P-bodies formation in cancer cells [111].
Although RBPs have been linked to different human cancers, the activities of some of them can be cancer-specific or patient-specific. For example, the higher expression of the RBP KHDRBS1 (KH domain-containing, RNA-binding, signal transduction-associated protein 1)/Sam68 (Src substrate associated in mitosis 68 kDa) leads to reduced survival of the patient in kidney renal papillary cell carcinoma and lung adenocarcinoma but not in acute myeloid leukemia and ovarian cancer [112]. This data displayed its critical role in cancer-specific events.
RBPs are implicated in the early stage of cancer manifestation. For example, in the work of [113], the increase in mitochondrial fatty acid beta-oxidation (FAO) and oxidative phosphorylation (OXPHOS) allows adenosine triphosphate (ATP) production and cellular lamellipodia formation during the initial step of High-grade serous ovarian cancer (HGSOC) cell migration and invasion. FAO and OXPHOS activities are linked to the improvement of mRNA stability and expression of carnitine palmitoyltransferase 1A (CPT1A) as well as NADH: Ubiquinone Oxidoreductase Subunit A2 (NDUFA2) resulting from the direct interaction between the Cancer-type organic anion transporting polypeptide 1B3 (Ct-OATP1B3) and the RBP insulin-like growth factor 2 mRNA-binding protein 2 (IGF2BP2). IGF2BP2 binding to N(6)-methyladenosine (m6A) methyltransferase of polo-like kinase 1 (PLK1) 3′UTR, initiated by the methyltransferase-like 3 (METTL3) activity, is also revealed to be essential for maintaining the cell cycle in pancreatic adenocarcinoma [114].
RBPs can be an important cause of chemoresistance in cancer. For example, in colorectal cancer (CRC), an investigation by [115] demonstrated that the RBP, leucine-rich pentatricopeptide repeat-containing protein (LRPPRC) specifically bound to the mRNA of multidrug resistance 1 (MDR1), augmenting MDR1 mRNA stability and protein expression. Fortunately, LRPPRC is more recently found to be impaired by gossypol-acetic acid [115]. RBPs activities have been noticed in autophagosome processes in human cancer. The RBP SQSTM1 is responsible for transporting the lncRNAs ARHGAP5-AS1 for autophagic degradation. The impaired autophagic degradation of ARHGAP5-AS1 is found to stimulate chemoresistance in human gastric cancer [116].
RBPs have been found in breast cancer (e.g. the RBPs RBM20 and PCDH20, [117], and their roles in cancer progression have also been proven. It has been shown that the RBP nucleolin (NCL) bound with Long non-coding RNA LINC01296, mainly located in the nuclei of Neuroblastoma (NB) cells, and formed a complex that triggered SRY-box transcription factor 11 (SOX11) gene transcription and accelerated the tumorigenesis of NB, a known common extracranial solid tumor in childhood [118]. Puf-A, a novel member of Puf-family RBPs, encourages non-small cell lung cancer (NSCLC) and colorectal cancer progression by interacting with nucleolar factors nucleophosmin-mediated ribosome biogenesis in nucleolus via the residues 478RRR480 of Puf-A [119]. This RBP may be useful for an effective mRNA-based treatment that will significantly suppress tumor growth.
RBPs are the targets of proteins promoting cancer invasion and metastasis. In the case of human Gastric cancer (GC), ecdysoneless (ECD) protein promotes cancer invasion and metastasis by binding the RBP hnRNP F via the N-terminal STG1 domain (13-383aa), preventing hnRNP F ubiquitination by the E3 ubiquitin ligase ZFP91, and proteasomal degradation. This effect of ECD turned it into an attractive marker candidate and a potential therapeutic target for GC patients [120].
An expanding set of RBPs is also involved in multiple malignancy events. The frequently overexpressed cancer-associated protein, death-associated protein 3 (DAP3), is an RBP demonstrated to act as a splicing regulatory RBP in cancer via its ability: (i) to mediate the formation of ribonucleoprotein complexes to trigger substrate-specific splicing changes, and (ii) to modulate the splicing of numerous splicing factors (RBM6, HNRNPH1, AKAP17A, and TIA1) to cause an indirect effect on splicing. This RBP triggers the nonsense-mediated decay of RBP (RBM6) and allows the association of splicing factor proline and glutamine-rich (SFPQ) and Non-POU domain-containing octamer binding (NONO) with target RNAs for splicing modulation. Its defects impair tumorigenesis at least partially via stimulating alternative splicing coupled nonsense-mediated decay (AS-NMD) of the WSB1 (WD repeat and SOCS box-containing protein 1) gene [121].
Colorectal cancer (CRC) cell proliferation relies on the activity of the RBP PTBP3 that monitors the expression of the E3 ubiquitin ligase UBE4A by binding its 3′UTR to impair UBE4A mRNA degradation and allows UBE4A to mediate P53 decomposition and stimulate CRC progression [122]. This process exemplifies the oncogenic function of PTBP3 and turns it into a promising therapeutic target for CRC in humans.
The two RNA/DNA-binding proteins, DHX9 helicase known to be overexpressed in cancer, are required for transcriptional termination and are involved in genome stability maintenance, interact with PARP-1 helicase to inhibit R-R-loop accumulation, which triggers DNA damage in human cells [123]. As these two proteins exerted a role in the cancer transcriptional program, they can be a promising target in a cancer treatment program. Given the compelling genetic evidence that RBPs exert a critical role in human disease, approaches using RBPs as a biomarker are of significant therapeutic interest, especially in the diagnosis or treatment of human cancer.
Furthermore, current evidence suggests that RBPs in various cell types serve as platforms for lncRNAs to regulate the cognate signaling pathways. The long non-coding RNA Urothelial carcinoma-associated 1 (UCA1) physically interacts with RBP (PTBP1)-regulated erythroid maturation and recruits PTBP1 to ALAS2 (5-aminolevulinic acid synthase 2) mRNA to stimulate ALAS2 mRNA stability and monitor heme biosynthesis as well as human erythroid maturation [124]. Defects in the regulation of RBPs activities by lncRNA drive genome instability in mammalian cells. The nuclear lncRNA MIR31HG interacts with the RBP YBX1 to allow its phosphorylation at serine 102 by the kinase RSK resulting in the induction of IL1A translation which stimulates the expression and secretion of a subset of senescence-associated secretory phenotype (SASP) components (interleukins and chemokines such as interleukin-6 (IL6), IL8 and CXCL1) during BRAF-induced senescence. Therefore, YBX1 can be used as a potential therapeutic target to promote senescence in cancer cells to repress cancer progression [125].
Interestingly, recent pieces of evidence have shown that RBPs are also responsible for the biogenesis circRNAs (circular RNAs) in cancer. In the line with this evidence, the role of the RBP (RBM3), acting as an oncogene critical for the formation of circRNA derived from the 3’UTR of the stearoyl-CoA desaturase (SCD) gene (so-called SCD-circRNA 2 stearoyl-CoA desaturase (SCD)-circRNA 2) in hepatocellular carcinoma (HCC) cells, has been unveiled [126]. It is worth noting that circRNAs such as hsa_circ_0001394 are suggested to be the key players in the proliferation, migration, and invasion of HCC cells [127].
Tools for identifying and analyzing RNA-interactome of RBPs
In recent years many RBPs have been identified and the mechanism regulating the activity of some of them has been elucidated, yet their precise function in the interactome involving RNAs is still under investigation. The complete elucidation of key elements or structures underlying protein-RNA interactions has the potential to greatly aid in analyzing the functionality of both the RNA and RNA-binding protein implicated as a large proportion of RBPs are still of unknown function.
Conventional approaches for elucidating RNA-interactome of RBPs or motifs in the RNAs and sequence-specific binding by a given RBP included a range of crosslinking and immunoprecipitations (eCLIP-seq) followed by RNA-sequencing [128]. To determine the functional relevance of RNA binding, a UV RNA crosslinking (CRAC) for a target protein and RNA in parallel with ChIP-seq experiments can be performed [129]. For example, individual-nucleotide resolution ultraviolet (UV) crosslinking and immunoprecipitation (iCLIP) are coupled to examine the potential role of RNA binding in CDK11 (cyclin-dependent kinase 11) functions. Crucially, the data revealed that this non-canonical RBP (CDK11) specifically monitors the expression of Replication-dependent histone (RDH)-coding genes. It binds to RDH RNAs and FLASH-containing RDH chromatin via its N-terminal RNA-binding region and associates with the chromatin of RDH genes in a cell-cycle-dependent manner. CDK11 is further demonstrated to phosphorylate Ser2 in the CTD of RNAPII positioned on the RDH genes to specifically control their transcriptional elongation and recruitment of 3′-end processing factors [130].
RBPs can be recognized by combining eCLIP with orthogonal approaches including in vitro assessment of RNA affinity for the same RBPs, chromatin association by ChIP–seq, and functional evaluation of transcriptome changes by RBP depletion and RNA-seq [131]. To identify the complete set of RBPs from yeast to humans, some new high-throughput technologies or models have been employed to detect RBPs during interactions between proteins and nucleic acids. For example, a tool combining peroxidase-catalyzed proximity labeling (PL) with organic-aqueous phase separation of crosslinked protein-RNA complexes (so-called APEX-PS) is employed to identify RBPs with novel functions in specific subcellular compartments. Notably, using this tool, it has been found in an outer mitochondrial membrane (OMM) that the RBP, SYNJ2BP, retains specific nuclear-encoded mitochondrial mRNAs at the OMM during translation stress, facilitating their local translation and channeling of protein products into the mitochondrion during stress recovery [132]. A method named STAMP (Surveying Targets by APOBEC-Mediated Profiling), which efficiently detects RBP–RNA interactions and does not rely on ultraviolet cross-linking or immunoprecipitation, is coupled with single-cell capture to identify RBP-specific and cell-type-specific RNA–protein interactions for multiple RBPs and cell types in single, pooled experiments [133]. It has been reported the use of post-transcriptional regulatory element sequencing (PTRE-seq) to examine post-transcriptional regulation by 3′ untranslated regions (UTRs) elements such as RBPs that often bind sequences in 3′UTR of mRNAs, and are known to regulate different aspects of the RNA lifecycle: RNA stability, translation efficiency, and translation initiation [134]. A deep learning tool, NucleicNet, complementing structural biology experiments, serves to provide quantitative fitness of RNA sequences for given binding pockets or to predict potential binding pockets as well as binding RNAs for previously unknown RBPs. For example, its application identified a diverse set of challenging RNA-binding proteins, including Fem-3-binding-factor 2, Argonaute 2, and Ribonuclease III [135]. A recent 3D2D model to predict binding sites of protein–RNA interaction named PRIME-3D2D has been also reported. It predicts binding sites on PDB data and transcription-wide [136]. In addition, photoswitchable RBP LicV responsive to blue light irradiation has been designed for controlling the spatiotemporal activity of RBP [137]. Additional tools for examining RBPs are highlighted in Table 3. To further comprehend protein-RNA interactions, a method for purifying protein-crosslinked RNA as a physical entity that may serve as a universal starting point for diverse downstream applications has been developed under the name XRNAX. This tool (XRNAX) can be combined with CLIP-seq to validate proteomic data, allow the discovery of the WKF RNA-binding domain, and helped both to pinpoint more than 700 proteins interacting with non-polyadenylated RNA and to observe the dynamic changes to the RNA-bound proteome [45]. A model for constructing protein-binding RNA sequences and motifs using a long short-term memory (LSTM) neural network has been employed to generate protein-binding RNA motifs comparable to the motifs from experimentally validated protein-binding RNA sequences [138]. A tool employing the ultraviolet cross-linking method and named kinetic cross-linking and analysis of cDNAs (χCRAC) has been used to quantitatively measure the global RNA-binding dynamics of the yeast transcription termination factor Nab3 in response to stress (glucose starvation) in vivo on a minute time-scale [139]. The post-transcriptional regulatory element sequencing (PTRE-seq) tool has been applied to assay the target sequence preferences and interactions between miRNAs and RBPs. It highlighted diverse facets of the RNA lifecycle encompassing RNA stability, translation efficiency, and translation initiation [134]. As most of the methods used so far to identify RBPs and their RNA-binding sites took advantage of proteins connected to polyadenylated RNAs and ignored RBPs bound to non-adenylate RNA classes (tRNA, rRNA, pre-mRNA) as well as species lacking poly-A tails in their mRNAs, a method free from specific RNA sequence or motif constraints and relying on the purification of cross-linked ribonucleoproteins (RNPs) based entirely on their physicochemical properties, and named (PTex) protocol has also been designed. It allowed capturing RBPs that bind to RNA as short as 30 nucleotides [140]. All the mentioned tools for RNA biology can serve as a universal starting point for the understanding of post-transcriptional regulatory processes in diverse cell types and organisms.
Table 3.
Additional tools for exploring proteins-RNAs interactions
| Tools | Advantages | Targets components | References |
|---|---|---|---|
| Deep-RBPPred | Needs few physicochemical properties based on protein sequences | RNA-binding protein (RBP) | [141] |
| Runs much faster | |||
| Good generalization ability | |||
| rec-YnH | Detects interactions within protein libraries or between protein libraries and RNA fragment pools | Screen protein libraries against protein libraries or RNA fragment libraries | [142] |
| Generate bait–prey fusion libraries | |||
| Maps interactions between multiple RNA-binding proteins and RNAs | |||
| NucleicNet | Binding preference of RNA backbone constituents and different bases | Local physicochemical characteristics of protein | [135] |
| Identify new RBPs and their binding pockets/preferences | Structure surface | ||
| Predict potential binding pockets | Predict interaction modes for differents RNAs | ||
| Predict binding RNAs for previously unknown RNA-binding proteins | constituents: Phosphate, Ribose, Adenine, | ||
| Accurately recover interaction modes discovered by structural | Guanine, Cytosine, Uracil, and non-site | ||
| Biology experiments | |||
| Score the binding potential of individual RNA sequences | |||
| Gemini-561–o-Coral | Improved brightness and photostability | A bright and photostable fluorogen for | [143] |
| High affinity | RNA imaging in cells | ||
| Direct fluorescence imaging in live mammalian cells | RNA in living mammalian cells | ||
| Sensitive detection of RNA inside live cells | |||
| RNA-based fluorogenic module | |||
| Spliceosome iClip | Identifies crosslinks of endogenous, untagged spliceosomal factors on pre-mRNAs at nucleotide resolution | RBPs | [144] |
| Peaks of spliceosomal crosslinking | |||
| Interactions of diverse RBPs on pre-mRNAs during spliceosomal assembly | Around branchpoints (BPs) | ||
| Splice sites | |||
| vIPR (in vivo Interactions | In vivo RNA–protein crosslinking and pulldown approach | Common and specific RNA-binding proteins | [145] |
| by ulldown of RNA) | Enables highly specific and efficient enrichment of an RNA of interest | Identify factors binding to selected mRNAs | |
| Enables identification of protein interactors of both transgenic and endogenous transcripts and endogenous transcripts | microRNAs (miRNAs) binding to mRNAs | ||
| Enhanced RNA interactome | Greater specificity and increased signal-to-noise ratios | RBPs | [146] |
| capture (eRIC) | Empowers comparative analyses of changes of RNA-bound proteomes | ||
| Identifies m6A-responsive RNA-binding proteins that escape RNA interactome capture (RIC) | |||
| RBS-ID | Fully cleave RNA into mono-nucleosides | RNA-binding sites (RBSs) | [147] |
| Replace enzymatic digestion and drastically improves the coverage (~ 2000 sites) and resolution (single amino acid level) of RNA-binding sites identification | |||
| Reach single amino acid resolution | |||
| View of residues important for genome editing | |||
| RNA-specific chemical cleavage | |||
| Quantitative SILAC-based | Associate individual RNA structures within messenger RNA with their interacting proteins | RNA-binding proteins | [52] |
| RNA pull-down | Proteins deprive of canonical RNA-binding domains | ||
| STAMP(Surveying Targets by | Efficiently detects RBP–RNA interactions | RBP–RNA interactomes and translational landscapes | [133] |
| APOBEC-Mediated Profiling) | Identify RBP-specific and cell-type–specific RNA–protein interactions for multiple RBPs and cell types in single, pooled experiments | ||
| Identify full-length RBPs that bind both polyadenylated mRNAs (RBFOX2 and TIA1) and non-polyadenylated mRNAs (SLBP | |||
| Identifies RBP–RNA sites at isoform-specific and single-cell resolution | |||
| Allows deconvolution of targets for multiplexed RBPs | |||
| Uncovers cell type–specific binding of an RBP in a heterogeneous mixture | |||
| of cell types | |||
| Allows single-cell detection of ribosome association while simultaneously measuring gene expression | |||
| pRBS-ID | Offering direct RNA-binding evidence and identifying RBSs at single amino acid-resolution with base-specificity (U or G) | Human RBSs | [147] |
| Profile uridine-contacting RNA-binding sites (RBSs) | |||
| Discovering guanosine-contacting RBSs | |||
| System-wide profiling of PAR-RBSs with a robust identification rate and accurate label-free quantification | |||
| RNA interactome capture (RNA-IC) and orthogonal organic phase separation (OOPS) | A global understanding of post-transcriptional gene regulation | Identify canonical and non-canonical RBPs from mouse and human CD4 + T cells RBPomes Elucidate higher-order RBP Roquin-mediated mRNA regulation | [1] |
| Identify how the cellular RBPs participate or intervene with post-transcriptional control of target mRNAs | |||
|
CAPRI (Crosslinked and Adjacent Peptides-based RNA-binding domain Identification) |
Simultaneously identifying both crosslinked peptides (in the immediate vicinity of the RNA–protein interface) and adjacent peptides (next to the crosslinked peptide) | Map RNA-binding domains (RBDs) | [148] |
| Identification of evolutionary conserved RBDs in globular domains and intrinsically disordered regions (IDRs) | |||
| FOREST (folded RNA element profiling with structure library) | Identify functional interactions from transcriptome-wide RNA structure datasets | RNA–protein (RNP) interactions | [149] |
| Structure-binding preferences of RBPs | |||
| Reveals different binding landscapes of RNA G-quadruplex (rG4) | |||
| structures-binding proteins and | |||
| Discovery of rG4 structures in the terminal loops of precursor microRNAs | |||
| Discovers translational regulatory elements that function in living cells | |||
| Evaluate a variety of RNA structures regardless of the organism | |||
| Eliminate reverse transcription (RT) and polymerase chain reaction (PCR) amplification bias | |||
| Quantitative RNA-interactome capture (qRIC) | Combination with phosphoproteomics allowed systematic comparison of pull-down efficiencies of phosphorylated and nonphosphorylated forms of mRBPs Capture known regulatory phosphorylation sites in ELAVL1, SF3B1, and UPF1 and identify potential regulatory sites Reveal multiple phosphorylation sites in the C-terminal disordered region in the splicing regulator RBM20 affecting nucleocytoplasmic localization, association with cytoplasmic, ribonucleoprotein granules and alternative splicing | Quantify the fraction of mRNA-binding proteins (mRBPs) pulled down with polyadenylated mRNAs | [150] |
Conclusion
Collectively, our review advanced our mechanistic understanding of RNA-binding protein diversity, provided a rich resource for functional studies, and identified therapeutic targets. A more comprehensive understanding of the molecular mechanism underlying the activity of the interactors of cancer progression, viral replication, and neurodegenerative disease is pivotal for the design of novel biomarkers and effective therapeutic targets for these diseases. While the biological implications of some RBPs domains remain to be explored, almost all domains reported herein are associated with important and various cellular processes. Furthermore, our review provides the basis for measuring the physiological impact of RBPs. The outcome of this review could aid to identify novel drugs for better treatment of RBPs-related disease.
Acknowledgements
The authors would like to thank all persons who provided the proofreading format of this publication.
Abbreviations
- ALS
Amyotrophic lateral sclerosis
- ATP
Adenosine triphosphate
- CRC
Colorectal carcinoma
- DDX
DEAD-box RNA helicase
- lncRNAs
Long non-coding RNAs
- PRC2
Polycomb repressive complex 2
- FUS
Fused in sarcoma
- hnRNP-K
Heterogeneous nuclear ribonucleoprotein K
- HCC
Hepatocellular carcinoma
- RBP
RNA-binding protein
- NSCLC
Non-small cell lung cancer
- LCD
Intrinsically disordered low-complexity domain
- miRNAs
MicroRNAs
- MLL
Mixed lineage leukemia
- ZFP36
Zinc finger protein 36
- eIF
Eukaryotic initiation factor
- PARP-1
Poly(ADP-ribose) polymerase-1
- IGF2BP
Insulin-like growth factor 2 mRNA-binding proteins
- RRM
RNA recognition motif
- RNP
Ribonucleoprotein
- METTL3
Methyltransferase 3, N6-adenosine-methyltransferase complex catalytic subunit
- TDP-43
Transactive response DNA-binding protein 43 kDa
Author contributions
DRAS conceived, designed and wrote the review paper. CC and DO analyzed the data and made the proofread of the manuscript. All authors read and approved the manuscript.
Funding
This work received no external funding.
Availability of data and materials
Not applicable.
Declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Hoefig KP, Reim A, Gallus C, Wong EH, Behrens G, Conrad C, Xu M, Kifinger L, Ito-Kureha T, Defourny KAY, Geerlof A, Mautner J, Hauck SM, Baumjohann D, Feederle R, Mann M, Wierer M, Glasmacher E, Heissmeyer V. ‘Defining the RBPome of primary T helper cells to elucidate higher-order Roquin-mediated mRNA regulation. Nat Commun. 2021;12(1):5208. doi: 10.1038/s41467-021-25345-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schneider T, Hung L-H, Aziz M, Wilmen A, Thaum S, Wagner J, Janowski R, Müller S, Schreiner S, Friedhoff P, Hüttelmaier S, Niessing D, Sattler M, Schlundt A, Bindereif A. Combinatorial recognition of clustered RNA elements by the multidomain RNA-binding protein IMP3. Nat Commun. 2019;10(1):2266. doi: 10.1038/s41467-019-09769-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, Hu Y-C, Hüttelmaier S, Skibbe JR, Su R, Deng X, Dong L, Sun M, Li C, Nachtergaele S, Wang Y, Hu C, Ferchen K, Greis KD, Jiang X, Wei M, Qu L, Guan J-L, He C, Yang J, Chen J. Recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20(3):285–295. doi: 10.1038/s41556-018-0045-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wu B, Su S, Patil DP, Liu H, Gan J, Jaffrey SR, Ma J. Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1. Nat Commun. 2018;9(1):420. doi: 10.1038/s41467-017-02770-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Dang M, Lim L, Kang J, Song J. ATP biphasically modulates LLPS of TDP-43 PLD by specifically binding arginine residues. Commun Biol. 2021;4(1):714. doi: 10.1038/s42003-021-02247-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sanchez de Groot N, Armaos A, Graña-Montes R, Alriquet M, Calloni G, Vabulas RM, Tartaglia GG. RNA structure drives interaction with proteins. Nat Commun. 2019;10(1):3246. doi: 10.1038/s41467-019-10923-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Pennemann FL, Mussabekova A, Urban C, Stukalov A, Andersen LL, Grass V, Lavacca TM, Holze C, Oubraham L, Benamrouche Y, Girardi E, Boulos RE, Hartmann R, Superti-Furga G, Habjan M, Imler J-L, Meignin C, Pichlmair A. Cross-species analysis of viral nucleic acid interacting proteins identifies TAOKs as innate immune regulators. Nat Commun. 2021;12(1):7009. doi: 10.1038/s41467-021-27192-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Littler DR, Mohanty B, Lowery SA, Colson RN, Gully BS, Perlman S, Scanlon MJ, Rossjohn J. ‘Binding of a pyrimidine RNA base-mimic to SARS-CoV-2 nonstructural protein 9. J Biol Chem. 2021;297(3). [DOI] [PMC free article] [PubMed]
- 9.Baden LR, El Sahly HM, Essink B, Kotloff K, Frey S, Novak R, Diemert D, Spector SA, Rouphael N, Creech CB, McGettigan J, Khetan S, Segall N, Solis J, Brosz A, Fierro C, Schwartz H, Neuzil K, Corey L, Gilbert P, Janes H, Follmann D, Marovich M, Mascola J, Polakowski L, Ledgerwood J, Graham BS, Bennett H, Pajon R, Knightly C, Leav B, Deng W, Zhou H, Han S, Ivarsson M, Miller J, Zaks T. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N Engl J Med. 2020;384(5):403–416. doi: 10.1056/NEJMoa2035389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Walsh EE, Frenck RW, Falsey AR, Kitchin N, Absalon J, Gurtman A, Lockhart S, Neuzil K, Mulligan MJ, Bailey R, Swanson KA, Li P, Koury K, Kalina W, Cooper D, Fontes-Garfias C, Shi P-Y, Türeci Ö, Tompkins KR, Lyke KE, Raabe V, Dormitzer PR, Jansen KU, Şahin U, Gruber WC. Safety and immunogenicity of two RNA-based COVID-19 vaccine candidates. N Engl J Med. 2020;383(25):2439–2450. doi: 10.1056/NEJMoa2027906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Li J, Hui A, Zhang X, Yang Y, Tang R, Ye H, Ji R, Lin M, Zhu Z, Türeci Ö, Lagkadinou E, Jia S, Pan H, Peng F, Ma Z, Wu Z, Guo X, Shi Y, Muik A, Şahin U, Zhu L, Zhu F. Safety and immunogenicity of the SARS-CoV-2 BNT162b1 mRNA vaccine in younger and older Chinese adults: a randomized, placebo-controlled, double-blind phase 1 study. Nat Med. 2021;27(6):1062–1070. doi: 10.1038/s41591-021-01330-9. [DOI] [PubMed] [Google Scholar]
- 12.Mascola JR, Fauci AS. Novel vaccine technologies for the 21st century. Nat Rev Immunol. 2020;20(2):87–88. doi: 10.1038/s41577-019-0243-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gebre MS, Brito LA, Tostanoski LH, Edwards DK, Carfi A, Barouch DH. Novel approaches for vaccine development. Cell. 2021;184(6):1589–1603. doi: 10.1016/j.cell.2021.02.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Crommelin DJA, Anchordoquy TJ, Volkin DB, Jiskoot W, Mastrobattista E. Addressing the cold reality of mRNA vaccine stability. J Pharm Sci. 2021;110(3):997–1001. doi: 10.1016/j.xphs.2020.12.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Jackson NAC, Kester KE, Casimiro D, Gurunathan S, DeRosa F. The promise of mRNA vaccines: a biotech and industrial perspective. npj Vaccines. 2020;5(1):11. doi: 10.1038/s41541-020-0159-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hacisuleyman E, Hale C, Saito Y, Blachere NE, Bergh M, Conlon EG, Schaefer-Babajew DJ, DaSilva J, Muecksch F, Gaebler C, Lifton R, Nussenzweig MC, Hatziioannou T, Bieniasz PD, Darnell RB. Vaccine breakthrough infections with SARS-CoV-2 variants. N Engl J Med. 2021;384(23):2212–2218. doi: 10.1056/NEJMoa2105000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gao Y, Cai C, Wullimann D, Niessl J, Rivera-Ballesteros O, Chen P, Lange J, Cuapio A, Blennow O, Hansson L, Mielke S, Nowak P, Vesterbacka J, Akber M, Perez-Potti A, Sekine T, Müller TR, Boulouis C, Kammann T, Parrot T, Muvva JR, Sobkowiak M, Healy K, Bogdanovic G, Muschiol S, Söderdahl G, Österborg A, Hellgren F, Grifoni A, Weiskopf D, Sette A, Loré K, Sällberg Chen M, Ljungman P, Sandberg JK, Smith CIE, Bergman P, Ljunggren H-G, Aleman S, Buggert M. Immunodeficiency syndromes differentially impact the functional profile of SARS-CoV-2-specific T cells elicited by mRNA vaccination. Immunity. 2022;55(9):1732–1746.e1735. doi: 10.1016/j.immuni.2022.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Singh G, Pratt G, Yeo GW, Moore MJ. The clothes make the mRNA: past and present trends in mRNP fashion. Annu Rev Biochem. 2015;84:325–354. doi: 10.1146/annurev-biochem-080111-092106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Castello A, Fischer B, Eichelbaum K, Horos R, Beckmann BM, Strein C, Davey NE, Humphreys DT, Preiss T, Steinmetz LM, Krijgsveld J, Hentze MW. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell. 2012;149(6):1393–1406. doi: 10.1016/j.cell.2012.04.031. [DOI] [PubMed] [Google Scholar]
- 20.Hentze MW, Castello A, Schwarzl T, Preiss T. A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol. 2018;19(5):327–341. doi: 10.1038/nrm.2017.130. [DOI] [PubMed] [Google Scholar]
- 21.Taylor PC, Adams AC, Hufford MM, de la Torre I, Winthrop K, Gottlieb RL. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat Rev Immunol. 2021;21(6):382–393. doi: 10.1038/s41577-021-00542-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kudinov AE, Karanicolas J, Golemis EA, Boumber Y. Musashi RNA-binding proteins as cancer drivers and novel therapeutic targets. Clin Cancer Res. 2017;23(9):2143–2153. doi: 10.1158/1078-0432.CCR-16-2728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Chavali PL, Stojic L, Meredith LW, Joseph N, Nahorski MS, Sanford TJ, Sweeney TR, Krishna BA, Hosmillo M, Firth AE, Bayliss R, Marcelis CL, Lindsay S, Goodfellow I, Woods CG, Gergely F. Neurodevelopmental protein Musashi-1 interacts with the Zika genome and promotes viral replication. Science. 2017;357(6346):83–88. doi: 10.1126/science.aam9243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Prasad K, Gour P, Raghuvanshi S, Kumar V. The SARS-CoV-2 targeted human RNA binding proteins network biology to investigate COVID-19 associated manifestations. Int J Biol Macromol. 2022;217:853–863. doi: 10.1016/j.ijbiomac.2022.07.200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu B-W, Wang X-Y, Cao J-L, Chen L-L, Wang Y-L, Zhao B-Q, Zhou J, Shen Z-F. TDP-43 upregulates lipid metabolism modulator ABHD2 to suppress apoptosis in hepatocellular carcinoma. Commun Biol. 2022;5(1):816. doi: 10.1038/s42003-022-03788-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tauber D, Tauber G, Parker R. Mechanisms and regulation of RNA condensation in RNP granule formation. Trends Biochem Sci. 2020;45(9):764–778. doi: 10.1016/j.tibs.2020.05.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Šušnjar U, Škrabar N, Brown A-L, Abbassi Y, Phatnani H, Phatnani H, Fratta P, Kwan J, Sareen D, Broach JR, Simmons Z, Arcila-Londono X, Lee EB, Van Deerlin VM, Shneider NA, Fraenkel E, Ostrow LW, Baas F, Berry JD, Butovsky O, Baloh RH, Shalem O, Heiman-Patterson T, Stefanis L, Chandran S, Pal S, Smith C, Malaspina A, Hammell MG, Patsopoulos NA, Dubnau J, Poss M, Zhang B, Zaitlen N, Hornstein E, Miller TM, Dardiotis E, Bowser R, Menon V, Harms M, Atassi N, Lange DJ, MacGowan DJ, McMillan C, Aronica E, Harris B, Ravits J, Crary J, Thompson LM, Raj T, Paganoni S, Adams DJ, Babu S, Drory V, Gotkine M, Broce I, Phillips-Cremins J, Nath A, Finkbeiner S, Cox GA, Cortese A, Cereda C, Bugiardini E, Cardani R, Meola G, Ripolone M, Moggio M, Romano M, Secrier M, Fratta P, Buratti E. Cell environment shapes TDP-43 function with implications in neuronal and muscle disease. Commun Biol. 2022;5(1):314. doi: 10.1038/s42003-022-03253-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Krismer K, Bird MA, Varmeh S, Handly ED, Gattinger A, Bernwinkler T, Anderson DA, Heinzel A, Joughin BA, Kong YW, Cannell IG, Yaffe MB. Transite: a computational motif-based analysis platform that identifies RNA-binding proteins modulating changes in gene expression. Cell Rep. 2020;32(8):108. doi: 10.1016/j.celrep.2020.108064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yoon DS, Lee K-M, Choi Y, Ko EA, Lee N-H, Cho S, Park KH, Lee J-H, Kim H-W, Lee JW. Correction: TLR4 downregulation by the RNA-binding protein PUM1 alleviates cellular aging and osteoarthritis. Cell Death Differ. 2022;29(5):1077–1077. doi: 10.1038/s41418-022-00945-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Arandel L, Matloka M, Klein AF, Rau F, Sureau A, Ney M, Cordier A, Kondili M, Polay-Espinoza M, Naouar N, Ferry A, Lemaitre M, Begard S, Colin M, Lamarre C, Tran H, Buée L, Marie J, Sergeant N, Furling D. Reversal of RNA toxicity in myotonic dystrophy via a decoy RNA-binding protein with high affinity for expanded CUG repeats. Nat Biomed Eng. 2022;6(2):207–220. doi: 10.1038/s41551-021-00838-2. [DOI] [PubMed] [Google Scholar]
- 31.Zhou B, Liu J, Kang R, Klionsky DJ, Kroemer G, Tang D. Ferroptosis is a type of autophagy-dependent cell death. Semin Cancer Biol. 2020;66:89–100. doi: 10.1016/j.semcancer.2019.03.002. [DOI] [PubMed] [Google Scholar]
- 32.Chen G, Long C, Wang S, Wang Z, Chen X, Tang W, He X, Bao Z, Tan B, Zhao J, Xie Y, Li Z, Yang D, Xiao G, Peng S. Circular RNA circStag1 promotes bone regeneration by interacting with HuR. Bone Res. 2022;10(1):32. doi: 10.1038/s41413-022-00208-x. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 33.Petkau G, Mitchell TJ, Chakraborty K, Bell SE, D’Angeli V, Matheson L, Turner DJ, Saveliev A, Gizlenci O, Salerno F, Katsikis PD, Turner M. The timing of differentiation and potency of CD8 effector function is set by RNA binding proteins. Nat Commun. 2022;13(1):2274. doi: 10.1038/s41467-022-29979-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kim HJ, Kim NC, Wang Y-D, Scarborough EA, Moore J, Diaz Z, MacLea KS, Freibaum B, Li S, Molliex A, Kanagaraj AP, Carter R, Boylan KB, Wojtas AM, Rademakers R, Pinkus JL, Greenberg SA, Trojanowski JQ, Traynor BJ, Smith BN, Topp S, Gkazi A-S, Miller J, Shaw CE, Kottlors M, Kirschner J, Pestronk A, Li YR, Ford AF, Gitler AD, Benatar M, King OD, Kimonis VE, Ross ED, Weihl CC, Shorter J, Taylor JP. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature. 2013;495(7442):467–473. doi: 10.1038/nature11922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Yang L, WenTao T, ZhiYuan Z, Qi L, YuXiang L, Peng Z, Ke L, XiaoNa J, YuZhi P, MeiLing J, QingYang F, GuoDong H, YueXiang W, JianMin X. Cullin-9/p53 mediates HNRNPC degradation to inhibit erastin-induced ferroptosis and is blocked by MDM2 inhibition in colorectal cancer. Oncogene. 2022;41(23):3210–3221. doi: 10.1038/s41388-022-02284-z. [DOI] [PubMed] [Google Scholar]
- 36.Feng S, Li J, Wen H, Liu K, Gui Y, Wen Y, Wang X, Yuan S. hnRNPH1 recruits PTBP2 and SRSF3 to modulate alternative splicing in germ cells. Nat Commun. 2022;13(1):3588. doi: 10.1038/s41467-022-31364-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Karginov TA, Ménoret A, Vella AT. Optimal CD8+ T cell effector function requires costimulation-induced RNA-binding proteins that reprogram the transcript isoform landscape. Nat Commun. 2022;13(1):3540. doi: 10.1038/s41467-022-31228-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Shao W, Bi X, Pan Y, Gao B, Wu J, Yin Y, Liu Z, Peng M, Zhang W, Jiang X, Ren W, Xu Y, Wu Z, Wang K, Zhan G, Lu JY, Han X, Li T, Wang J, Li G, Deng H, Li B, Shen X. Phase separation of RNA-binding protein promotes polymerase binding and transcription. Nat Chem Biol. 2022;18(1):70–80. doi: 10.1038/s41589-021-00904-5. [DOI] [PubMed] [Google Scholar]
- 39.Mosler T, Conte F, Longo GMC, Mikicic I, Kreim N, Möckel MM, Petrosino G, Flach J, Barau J, Luke B, Roukos V, Beli P. R-loop proximity proteomics identifies a role of DDX41 in transcription-associated genomic instability. Nat Commun. 2021;12(1):7314. doi: 10.1038/s41467-021-27530-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Liu Z-L, Wang S-K, Pang L, Meng X-W. circXRCC5 foster gastric cancer growth and metastasis by the HNRNPC/circXRCC5/miR-655-3p/RREB1/UBA2 positive feedback loop. Cancer Gene Ther. 2022;29(11):1648–1661. doi: 10.1038/s41417-022-00482-1. [DOI] [PubMed] [Google Scholar]
- 41.Ludlow AT, Wong MS, Robin JD, Batten K, Yuan L, Lai T-P, Dahlson N, Zhang L, Mender I, Tedone E, Sayed ME, Wright WE, Shay JW. NOVA1 regulates hTERT splicing and cell growth in non-small cell lung cancer. Nat Commun. 2018;9(1):3112. doi: 10.1038/s41467-018-05582-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Liu D, Lin L, Wang Y, Chen L, He Y, Luo Y, Qi L, Guo Y, Chen L, Han Z, Li G, Li Q, Liu Z, Chen P, Guo H. PNO1, which is negatively regulated by miR-340–5p, promotes lung adenocarcinoma progression through Notch signaling pathway. Oncogenesis. 2020;9(5):58. doi: 10.1038/s41389-020-0241-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chen J, Cai Z, Bai M, Yu X, Zhang C, Cao C, Hu X, Wang L, Su R, Wang D, Wang L, Yao Y, Ye R, Hou B, Yu Y, Yu S, Li J, Xue Y. The RNA-binding protein ROD1/PTBP3 cotranscriptionally defines AID-loading sites to mediate antibody class switch in mammalian genomes. Cell Res. 2018;28(10):981–995. doi: 10.1038/s41422-018-0076-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Cheng C, Cai Y, Liu X, Wu Y, Cheng Q, Wu Y, Wu Z. KHSRP modulated cell proliferation and cell cycle via regulating PPP2CA and p27 expression in Wilms tumor. Cell Signal. 2022;100:110447. doi: 10.1016/j.cellsig.2022.110447. [DOI] [PubMed] [Google Scholar]
- 45.Trendel J, Schwarzl T, Horos R, Prakash A, Bateman A, Hentze MW, Krijgsveld J. The human RNA-binding proteome and its dynamics during translational arrest. Cell. 2019;176(1):391–403.e319. doi: 10.1016/j.cell.2018.11.004. [DOI] [PubMed] [Google Scholar]
- 46.Turco C, Esposito G, Iaiza A, Goeman F, Benedetti A, Gallo E, Daralioti T, Perracchio L, Sacconi A, Pasanisi P, Muti P, Pulito C, Strano S, Ianniello Z, Fatica A, Forcato M, Fazi F, Blandino G, Fontemaggi G. MALAT1-dependent hsa_circ_0076611 regulates translation rate in triple-negative breast cancer. Commun Biol. 2022;5(1):598. doi: 10.1038/s42003-022-03539-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Shao M, Hao S, Jiang L, Cai Y, Zhao X, Chen Q, Gao X, Xu J. CRIT: Identifying RNA-binding protein regulator in circRNA life cycle via non-negative matrix factorization. Mol Ther Nucleic Acids. 2022;30:398–406. doi: 10.1016/j.omtn.2022.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Zhao B, Katuwawala A, Oldfield CJ, Hu G, Wu Z, Uversky VN, Kurgan L. Intrinsic disorder in human RNA-binding proteins. J Mol Biol. 2021;433(21):167229. doi: 10.1016/j.jmb.2021.167229. [DOI] [PubMed] [Google Scholar]
- 49.Winz M-L, Peil L, Turowski TW, Rappsilber J, Tollervey D. Molecular interactions between Hel2 and RNA supporting ribosome-associated quality control. Nat Commun. 2019;10(1):563. doi: 10.1038/s41467-019-08382-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Wang C, Kong F, Ma J, Miao J, Su P, Yang H, Li Q, Ma X. IGF2BP3 enhances the mRNA stability of E2F3 by interacting with LINC00958 to promote endometrial carcinoma progression. Cell Death Discov. 2022;8(1):279. doi: 10.1038/s41420-022-01045-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Okuda H, Miyamoto R, Takahashi S, Kawamura T, Ichikawa J, Harada I, Tamura T, Yokoyama A. RNA-binding proteins of KHDRBS and IGF2BP families control the oncogenic activity of MLL-AF4. Nat Commun. 2022;13(1):6688. doi: 10.1038/s41467-022-34558-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Casas-Vila N, Sayols S, Pérez-Martínez L, Scheibe M, Butter F. The RNA fold interactome of evolutionary conserved RNA structures in S. cerevisiae. Nat Commun. 2020;11(1):2789. doi: 10.1038/s41467-020-16555-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Wei W, Gao W, Li Q, Liu Y, Chen H, Cui Y, Sun Z, Liu Z. Comprehensive characterization of posttranscriptional impairment-related 3′-UTR mutations in 2413 whole genomes of cancer patients. npj Genom Med. 2022;7(1):34. doi: 10.1038/s41525-022-00305-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Stefanovic L, Gordon BH, Silvers R, Stefanovic B. Characterization of sequence-specific binding of LARP6 to the 5′ stem-loop of type I collagen mRNAs and implications for rational design of antifibrotic drugs. J Mol Biol. 2022;434(2):167394. doi: 10.1016/j.jmb.2021.167394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Chen X, Yuan J, Xue G, Campanario S, Wang D, Wang W, Mou X, Liew SW, Umar MI, Isern J, Zhao Y, He L, Li Y, Mann CJ, Yu X, Wang L, Perdiguero E, Chen W, Xue Y, Nagamine Y, Kwok CK, Sun H, Muñoz-Cánoves P, Wang H. Translational control by DHX36 binding to 5′UTR G-quadruplex is essential for muscle stem-cell regenerative functions. Nat Commun. 2021;12(1):5043. doi: 10.1038/s41467-021-25170-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Georgakopoulos-Soares I, Parada GE, Wong HY, Medhi R, Furlan G, Munita R, Miska EA, Kwok CK, Hemberg M. Alternative splicing modulation by G-quadruplexes. Nat Commun. 2022;13(1):2404. doi: 10.1038/s41467-022-30071-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Bhattacharya S, Levy MJ, Zhang N, Li H, Florens L, Washburn MP, Workman JL. The methyltransferase SETD2 couples transcription and splicing by engaging mRNA processing factors through its SHI domain. Nat Commun. 2021;12(1):1443. doi: 10.1038/s41467-021-21663-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Biancon G, Joshi P, Zimmer JT, Hunck T, Gao Y, Lessard MD, Courchaine E, Barentine AES, Machyna M, Botti V, Qin A, Gbyli R, Patel A, Song Y, Kiefer L, Viero G, Neuenkirchen N, Lin H, Bewersdorf J, Simon MD, Neugebauer KM, Tebaldi T, Halene S. Precision analysis of mutant U2AF1 activity reveals deployment of stress granules in myeloid malignancies. Mol Cell. 2022;82(6):1107–1122.e1107. doi: 10.1016/j.molcel.2022.02.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Tzelepis K, Rausch O, Kouzarides T. RNA-modifying enzymes and their function in a chromatin context. Nat Struct Mol Biol. 2019;26(10):858–862. doi: 10.1038/s41594-019-0312-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, Zhao BS, Mesquita A, Liu C, Yuan CL, Hu Y-C, Hüttelmaier S, Skibbe JR, Su R, Deng X, Dong L, Sun M, Li C, Nachtergaele S, Wang Y, Hu C, Ferchen K, Greis KD, Jiang X, Wei M, Qu L, Guan J-L, He C, Yang J, Chen J. Publisher correction: recognition of RNA N6-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2020;22(10):1288–1288. doi: 10.1038/s41556-020-00580-y. [DOI] [PubMed] [Google Scholar]
- 61.Zhang G, Xu Y, Wang X, Zhu Y, Wang L, Zhang W, Wang Y, Gao Y, Wu X, Cheng Y, Sun Q, Chen D. Dynamic FMR1 granule phase switch instructed by m6A modification contributes to maternal RNA decay. Nat Commun. 2022;13(1):859. doi: 10.1038/s41467-022-28547-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Sauer M, Juranek SA, Marks J, De Magis A, Kazemier HG, Hilbig D, Benhalevy D, Wang X, Hafner M, Paeschke K. DHX36 prevents the accumulation of translationally inactive mRNAs with G4-structures in untranslated regions. Nat Commun. 2019;10(1):2421. doi: 10.1038/s41467-019-10432-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Charnley M, Islam S, Bindra GK, Engwirda J, Ratcliffe J, Zhou J, Mezzenga R, Hulett MD, Han K, Berryman JT, Reynolds NP. Neurotoxic amyloidogenic peptides in the proteome of SARS-COV2: potential implications for neurological symptoms in COVID-19. Nat Commun. 2022;13(1):3387. doi: 10.1038/s41467-022-30932-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Gerstberger S, Hafner M, Tuschl T. A census of human RNA-binding proteins. Nat Rev Genet. 2014;15(12):829–845. doi: 10.1038/nrg3813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Nussbacher JK, Batra R, Lagier-Tourenne C, Yeo GW. RNA-binding proteins in neurodegeneration: Seq and you shall receive. Trends Neurosci. 2015;38(4):226–236. doi: 10.1016/j.tins.2015.02.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Montalbano M, McAllen S, Cascio FL, Sengupta U, Garcia S, Bhatt N, Ellsworth A, Heidelman EA, Johnson OD, Doskocil S, Kayed R. TDP-43 and tau oligomers in Alzheimer's disease, amyotrophic lateral sclerosis, and frontotemporal dementia. Neurobiol Dis. 2020;146:105130. doi: 10.1016/j.nbd.2020.105130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Guo L, Kim HJ, Wang H, Monaghan J, Freyermuth F, Sung JC, O’Donovan K, Fare CM, Diaz Z, Singh N, Zhang ZC, Coughlin M, Sweeny EA, DeSantis ME, Jackrel ME, Rodell CB, Burdick JA, King OD, Gitler AD, Lagier-Tourenne C, Pandey UB, Chook YM, Taylor JP, Shorter J. Nuclear-import receptors reverse aberrant phase transitions of RNA-binding proteins with prion-like domains. Cell. 2018;173(3):677–692.e620. doi: 10.1016/j.cell.2018.03.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Chiu S-H, Ho W-L, Sun Y-C, Kuo J-C, Huang J-r. Phase separation driven by interchangeable properties in the intrinsically disordered regions of protein paralogs. Commun Biol. 2022;5(1):400. doi: 10.1038/s42003-022-03354-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Duan L, Zaepfel BL, Aksenova V, Dasso M, Rothstein JD, Kalab P, Hayes LR. Nuclear RNA binding regulates TDP-43 nuclear localization and passive nuclear export. Cell Rep. 2022;40(3):111106. doi: 10.1016/j.celrep.2022.111106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Kim HJ, Mohassel P, Donkervoort S, Guo L, O’Donovan K, Coughlin M, Lornage X, Foulds N, Hammans SR, Foley AR, Fare CM, Ford AF, Ogasawara M, Sato A, Iida A, Munot P, Ambegaonkar G, Phadke R, O’Donovan DG, Buchert R, Grimmel M, Töpf A, Zaharieva IT, Brady L, Hu Y, Lloyd TE, Klein A, Steinlin M, Kuster A, Mercier S, Marcorelles P, Péréon Y, Fleurence E, Manzur A, Ennis S, Upstill-Goddard R, Bello L, Bertolin C, Pegoraro E, Salviati L, French CE, Shatillo A, Raymond FL, Haack TB, Quijano-Roy S, Böhm J, Nelson I, Stojkovic T, Evangelista T, Straub V, Romero NB, Laporte J, Muntoni F, Nishino I, Tarnopolsky MA, Shorter J, Bönnemann CG, Taylor JP. Heterozygous frameshift variants in HNRNPA2B1 cause early-onset oculopharyngeal muscular dystrophy. Nat Commun. 2022;13(1):2306. doi: 10.1038/s41467-022-30015-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Jafarinia H, van der Giessen E, Onck PR. Phase separation of toxic dipeptide repeat proteins related to C9orf72 ALS/FTD. Biophys J. 2020;119(4):843–851. doi: 10.1016/j.bpj.2020.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Ryan S, Rollinson S, Hobbs E, Pickering-Brown S. C9orf72 dipeptides disrupt the nucleocytoplasmic transport machinery and cause TDP-43 mislocalisation to the cytoplasm. Sci Rep. 2022;12(1):4799. doi: 10.1038/s41598-022-08724-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Girdhar A, Bharathi V, Tiwari VR, Abhishek S, Deeksha W, Mahawar US, Raju G, Singh SK, Prabusankar G, Rajakumara E, Patel BK. Computational insights into mechanism of AIM4-mediated inhibition of aggregation of TDP-43 protein implicated in ALS and evidence for in vitro inhibition of liquid-liquid phase separation (LLPS) of TDP-432C-A315T by AIM4. Int J Biol Macromol. 2020;147:117–130. doi: 10.1016/j.ijbiomac.2020.01.032. [DOI] [PubMed] [Google Scholar]
- 74.Rhine K, Dasovich M, Yoniles J, Badiee M, Skanchy S, Ganser LR, Ge Y, Fare CM, Shorter J, Leung AKL, Myong S. Poly(ADP-ribose) drives condensation of FUS via a transient interaction. Mol Cell. 2022;82(5):969–985.e911. doi: 10.1016/j.molcel.2022.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Sahadevan S, Hembach KM, Tantardini E, Pérez-Berlanga M, Hruska-Plochan M, Megat S, Weber J, Schwarz P, Dupuis L, Robinson MD, De Rossi P, Polymenidou M. Synaptic FUS accumulation triggers early misregulation of synaptic RNAs in a mouse model of ALS. Nat Commun. 2021;12(1):3027. doi: 10.1038/s41467-021-23188-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Gallego-Iradi MC, Strunk H, Crown AM, Davila R, Brown H, Rodriguez-Lebron E, Borchelt DR. N-terminal sequences in matrin 3 mediate phase separation into droplet-like structures that recruit TDP43 variants lacking RNA binding elements. Lab Invest. 2019;99(7):1030–1040. doi: 10.1038/s41374-019-0260-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Gruijs da Silva LA, Simonetti F, Hutten S, Riemenschneider H, Sternburg EL, Pietrek LM, Gebel J, Dötsch V, Edbauer D, Hummer G, Stelzl LS, Dormann D. Disease-linked TDP-43 hyperphosphorylation suppresses TDP-43 condensation and aggregation. EMBO J. 2022;41(8):e108443. doi: 10.15252/embj.2021108443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Tariq A, Lin J, Jackrel ME, Hesketh CD, Carman PJ, Mack KL, Weitzman R, Gambogi C, Hernandez Murillo OA, Sweeny EA, Gurpinar E, Yokom AL, Gates SN, Yee K, Sudesh S, Stillman J, Rizo AN, Southworth DR, Shorter J. Mining disaggregase sequence space to safely counter TDP-43, FUS, and α-synuclein proteotoxicity. Cell Rep. 2019;28(8):2080–2095.e2086. doi: 10.1016/j.celrep.2019.07.069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Yoshizawa T, Ali R, Jiou J, Fung HYJ, Burke KA, Kim SJ, Lin Y, Peeples WB, Saltzberg D, Soniat M, Baumhardt JM, Oldenbourg R, Sali A, Fawzi NL, Rosen MK, Chook YM. Nuclear import receptor inhibits phase separation of FUS through binding to multiple sites. Cell. 2018;173(3):693–705.e622. doi: 10.1016/j.cell.2018.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Canet-Pons J, Sen N-E, Arsović A, Almaguer-Mederos L-E, Halbach MV, Key J, Döring C, Kerksiek A, Picchiarelli G, Cassel R, René F, Dieterlé S, Fuchs NV, König R, Dupuis L, Lütjohann D, Gispert S, Auburger G. Atxn2-CAG100-KnockIn mouse spinal cord shows progressive TDP43 pathology associated with cholesterol biosynthesis suppression. Neurobiol Dis. 2021;152:105289. doi: 10.1016/j.nbd.2021.105289. [DOI] [PubMed] [Google Scholar]
- 81.Kabashi E, Valdmanis PN, Dion P, Spiegelman D, McConkey BJ, Velde CV, Bouchard J-P, Lacomblez L, Pochigaeva K, Salachas F, Pradat P-F, Camu W, Meininger V, Dupre N, Rouleau GA. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet. 2008;40(5):572–574. doi: 10.1038/ng.132. [DOI] [PubMed] [Google Scholar]
- 82.Mackenzie IR, Nicholson AM, Sarkar M, Messing J, Purice MD, Pottier C, Annu K, Baker M, Perkerson RB, Kurti A, Matchett BJ, Mittag T, Temirov J, Hsiung G-YR, Krieger C, Murray ME, Kato M, Fryer JD, Petrucelli L, Zinman L, Weintraub S, Mesulam M, Keith J, Zivkovic SA, Hirsch-Reinshagen V, Roos RP, Züchner S, Graff-Radford NR, Petersen RC, Caselli RJ, Wszolek ZK, Finger E, Lippa C, Lacomis D, Stewart H, Dickson DW, Kim HJ, Rogaeva E, Bigio E, Boylan KB, Taylor JP, Rademakers R. TIA1 mutations in amyotrophic lateral sclerosis and frontotemporal dementia promote phase separation and alter stress granule dynamics. Neuron. 2017;95(4):808–816.e809. doi: 10.1016/j.neuron.2017.07.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Prieto C, Nguyen DTT, Liu Z, Wheat J, Perez A, Gourkanti S, Chou T, Barin E, Velleca A, Rohwetter T, Chow A, Taggart J, Savino AM, Hoskova K, Dhodapkar M, Schurer A, Barlowe TS, Vu LP, Leslie C, Steidl U, Rabadan R, Kharas MG. Transcriptional control of CBX5 by the RNA binding proteins RBMX and RBMXL1 maintains chromatin state in myeloid leukemia. Nat Cancer. 2021;2:741–757. doi: 10.1038/s43018-021-00220-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Zhao Y, Karijolich J. Know thyself: RIG-I-like receptor sensing of DNA virus infection. J Virol. 2019;93(23):e01085–e11019. doi: 10.1128/JVI.01085-19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Tran V, Ledwith MP, Thamamongood T, Higgins CA, Tripathi S, Chang MW, Benner C, García-Sastre A, Schwemmle M, Boon ACM, Diamond MS, Mehle A. Influenza virus repurposes the antiviral protein IFIT2 to promote translation of viral mRNAs. Nat Microbiol. 2020;5(12):1490–1503. doi: 10.1038/s41564-020-0778-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Wang X, Lin L, Zhong Y, Feng M, Yu T, Yan Y, Zhou J, Liao M. Cellular hnRNPAB binding to viral nucleoprotein inhibits flu virus replication by blocking nuclear export of viral mRNA. iScience. 2021;24(3):102160. doi: 10.1016/j.isci.2021.102160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Kamel W, Noerenberg M, Cerikan B, Chen H, Järvelin AI, Kammoun M, Lee JY, Shuai N, Garcia-Moreno M, Andrejeva A, Deery MJ, Johnson N, Neufeldt CJ, Cortese M, Knight ML, Lilley KS, Martinez J, Davis I, Bartenschlager R, Mohammed S, Castello A. Global analysis of protein-RNA interactions in SARS-CoV-2-infected cells reveals key regulators of infection. Mol Cell. 2021;81(13):2851–2867.e2857. doi: 10.1016/j.molcel.2021.05.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Zhang S, Huang W, Ren L, Ju X, Gong M, Rao J, Sun L, Li P, Ding Q, Wang J, Zhang QC. Comparison of viral RNA–host protein interactomes across pathogenic RNA viruses informs rapid antiviral drug discovery for SARS-CoV-2. Cell Res. 2022;32(1):9–23. doi: 10.1038/s41422-021-00581-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Qi X, Yan Q, Shang Y, Zhao R, Ding X, Gao S-J, Li W, Lu C. A viral interferon regulatory factor degrades RNA-binding protein hnRNP Q1 to enhance aerobic glycolysis via recruiting E3 ubiquitin ligase KLHL3 and decaying GDPD1 mRNA. Cell Death Differ. 2022;29(11):2233–2246. doi: 10.1038/s41418-022-01011-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Li X, Fu Z, Liang H, Wang Y, Qi X, Ding M, Sun X, Zhou Z, Huang Y, Gu H, Li L, Chen X, Li D, Zhao Q, Liu F, Wang H, Wang J, Zen K, Zhang C-Y. H5N1 influenza virus-specific miRNA-like small RNA increases cytokine production and mouse mortality via targeting poly(rC)-binding protein 2. Cell Res. 2018;28(2):157–171. doi: 10.1038/cr.2018.3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Cruz MA, Frederick TE, Mallimadugula UL, Singh S, Vithani N, Zimmerman MI, Porter JR, Moeder KE, Amarasinghe GK, Bowman GR. A cryptic pocket in Ebola VP35 allosterically controls RNA binding. Nat Commun. 2022;13(1):2269. doi: 10.1038/s41467-022-29927-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Lutz M, Schmierer J, Takimoto T. Host adaptive mutations in the 2009 H1N1 pandemic influenza A virus PA gene regulate translation efficiency of viral mRNAs via GRSF1. Commun Biol. 2022;5(1):1102. doi: 10.1038/s42003-022-04082-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Michalski D, Ontiveros JG, Russo J, Charley PA, Anderson JR, Heck AM, Geiss BJ, Wilusz J. Zika virus noncoding sfRNAs sequester multiple host-derived RNA-binding proteins and modulate mRNA decay and splicing during infection. J Biol Chem. 2019;294(44):16282–16296. doi: 10.1074/jbc.RA119.009129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Dave P, George B, Raheja H, Rani P, Behera P, Das S. The mammalian host protein DAP5 facilitates the initial round of translation of Coxsackievirus B3 RNA. J Biol Chem. 2019;294(42):15386–15394. doi: 10.1074/jbc.RA119.009000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Mure F, Corbin A, Benbahouche NEH, Bertrand E, Manet E, Gruffat H. The splicing factor SRSF3 is functionally connected to the nuclear RNA exosome for intronless mRNA decay. Sci Rep. 2018;8(1):12901. doi: 10.1038/s41598-018-31078-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Lodde V, Floris M, Munk R, Martindale JL, Piredda D, Napodano CMP, Cucca F, Uzzau S, Abdelmohsen K, Gorospe M, Noh JH, Idda ML. Systematic identification of NF90 target RNAs by iCLIP analysis. Sci Rep. 2022;12(1):364. doi: 10.1038/s41598-021-04101-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Schmidt N, Lareau CA, Keshishian H, Ganskih S, Schneider C, Hennig T, Melanson R, Werner S, Wei Y, Zimmer M, Ade J, Kirschner L, Zielinski S, Dölken L, Lander ES, Caliskan N, Fischer U, Vogel J, Carr SA, Bodem J, Munschauer M. The SARS-CoV-2 RNA–protein interactome in infected human cells. Nat Microbiol. 2021;6(3):339–353. doi: 10.1038/s41564-020-00846-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Zimmer MM, Kibe A, Rand U, Pekarek L, Ye L, Buck S, Smyth RP, Cicin-Sain L, Caliskan N. The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting. Nat Commun. 2021;12(1):7193. doi: 10.1038/s41467-021-27431-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Matkovic R, Morel M, Lanciano S, Larrous P, Martin B, Bejjani F, Vauthier V, Hansen MMK, Emiliani S, Cristofari G, Gallois-Montbrun S, Margottin-Goguet F. TASOR epigenetic repressor cooperates with a CNOT1 RNA degradation pathway to repress HIV. Nat Commun. 2022;13(1):66. doi: 10.1038/s41467-021-27650-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Liang Z, Rehati A, Husaiyin E, Chen D, Jiyuan Z, Abuduaini B. RALY regulate the proliferation and expression of immune/inflammatory response genes via alternative splicing of FOS. Genes Immunity. 2022. [DOI] [PMC free article] [PubMed]
- 101.Abdou Y, Gupta M, Asaoka M, Attwood K, Mateusz O, Gandhi S, Takabe K. ‘Left sided breast cancer is associated with aggressive biology and worse outcomes than right sided breast cancer. Sci Rep. 2022;12(1):13377. doi: 10.1038/s41598-022-16749-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Tan AC, Ashley DM, López GY, Malinzak M, Friedman HS, Khasraw M. ‘Management of glioblastoma: state of the art and future directions. CA Cancer J Clin. 2020;70(4):299–312. doi: 10.3322/caac.21613. [DOI] [PubMed] [Google Scholar]
- 103.Gong Y, Liu Z, Yuan Y, Yang Z, Zhang J, Lu Q, Wang W, Fang C, Lin H, Liu S. ‘PUMILIO proteins promote colorectal cancer growth via suppressing p21. Nat Commun. 2022;13(1):1627. doi: 10.1038/s41467-022-29309-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Raghava Kurup R, Oakes EK, Vadlamani P, Nwosu O, Danthi P, Hundley HA. ADAR3 activates NF-κB signaling and promotes glioblastoma cell resistance to temozolomide. Sci Rep. 2022;12(1):13362. doi: 10.1038/s41598-022-17559-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Zhu H, Chen K, Chen Y, Liu J, Zhang X, Zhou Y, Liu Q, Wang B, Chen T, Cao X. ‘RNA-binding protein ZCCHC4 promotes human cancer chemoresistance by disrupting DNA-damage-induced apoptosis. Signal Transduct Target Ther. 2022;7(1):240. doi: 10.1038/s41392-022-01033-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Takeiwa T, Ikeda K, Suzuki T, Sato W, Iino K, Mitobe Y, Kawabata H, Horie K, Inoue S. PSPC1 is a potential prognostic marker for hormone-dependent breast cancer patients and modulates RNA processing of ESR1 and SCFD2. Sci Rep. 2022;12(1):9495. doi: 10.1038/s41598-022-13601-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Lin X, Zhou L, Zhong J, Zhong L, Zhang R, Kang T, Wu Y (2022) RNA-binding protein RBM28 can translocate from the nucleolus to the nucleoplasm to inhibit the transcriptional activity of p53. J Biol Chem. 2022;298(2). [DOI] [PMC free article] [PubMed]
- 108.Shi J, Jin X, Wang Y, Zhu T, Zhang D, Li Q, Zhong X, Deng Y, Shen J, Fan X. LIN28B inhibition sensitizes cells to p53-restoring PPI therapy through unleashed translational suppression. Oncogenesis. 2022;11(1):37. doi: 10.1038/s41389-022-00412-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Howley BV, Mohanty B, Dalton A, Grelet S, Karam J, Dincman T, Howe PH. The ubiquitin E3 ligase ARIH1 regulates hnRNP E1 protein stability, EMT and breast cancer progression. Oncogene. 2022;41(12):1679–1690. doi: 10.1038/s41388-022-02199-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Yin M, Cheng M, Liu C, Wu K, Xiong W, Fang J, Li Y, Zhang B. HNRNPA2B1 as a trigger of RNA switch modulates the miRNA-mediated regulation of CDK6. iScience. 2021;24(11):103345. doi: 10.1016/j.isci.2021.103345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Yoshino S, Matsui Y, Fukui Y, Seki M, Yamaguchi K, Kanamori A, Saitoh Y, Shimamura T, Suzuki Y, Furukawa Y, Kaneko S, Seiki M, Murakami Y, Inoue J-I, Sakamoto T. EXOSC9 depletion attenuates P-body formation, stress resistance, and tumorigenicity of cancer cells. Sci Rep. 2020;10(1):9275. doi: 10.1038/s41598-020-66455-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Sumithra B, Saxena U, Das AB. A comprehensive study on genome-wide coexpression network of KHDRBS1/Sam68 reveals its cancer and patient-specific association. Sci Rep. 2019;9(1):11083. doi: 10.1038/s41598-019-47558-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Huang Y, Du Y, Zheng Y, Wen C, Zou H, Huang J, Zhou H, Zhao H, Wu L. Ct-OATP1B3 promotes high-grade serous ovarian cancer metastasis by regulation of fatty acid beta-oxidation and oxidative phosphorylation. Cell Death Dis. 2022;13(6):556. doi: 10.1038/s41419-022-05014-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Tatekawa S, Tamari K, Chijimatsu R, Konno M, Motooka D, Mitsufuji S, Akita H, Kobayashi S, Murakumo Y, Doki Y, Eguchi H, Ishii H, Ogawa K. N(6)-methyladenosine methylation-regulated polo-like kinase 1 cell cycle homeostasis as a potential target of radiotherapy in pancreatic adenocarcinoma. Sci Rep. 2022;12(1):11074. doi: 10.1038/s41598-022-15196-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Yang Y, Yuan H, Zhao L, Guo S, Hu S, Tian M, Nie Y, Yu J, Zhou C, Niu J, Wang G, Song Y. Targeting the miR-34a/LRPPRC/MDR1 axis collapse the chemoresistance in P53 inactive colorectal cancer. Cell Death Differ. 2022;29(11):2177–2189. doi: 10.1038/s41418-022-01007-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Zhu L, Zhu Y, Han S, Chen M, Song P, Dai D, Xu W, Jiang T, Feng L, Shin VY, Wang X, Jin H. Impaired autophagic degradation of lncRNA ARHGAP5-AS1 promotes chemoresistance in gastric cancer. Cell Death Dis. 2019;10(6):383. doi: 10.1038/s41419-019-1585-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Cava C, Armaos A, Lang B, Tartaglia GG, Castiglioni I. Identification of long non-coding RNAs and RNA binding proteins in breast cancer subtypes. Sci Rep. 2022;12(1):693. doi: 10.1038/s41598-021-04664-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Wang J, Wang Z, Lin W, Han Q, Yan H, Yao W, Dong R, Jia D, Dong K, Li K. LINC01296 promotes neuroblastoma tumorigenesis via the NCL-SOX11 regulatory complex. Mol Ther Oncolytics. 2022;24:834–848. doi: 10.1016/j.omto.2022.02.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Cho H-C, Huang Y, Hung J-T, Hung T-H, Cheng K-C, Liu Y-H, Kuo M-W, Wang S-H, Yu AL, Yu J. Puf-A promotes cancer progression by interacting with nucleophosmin in nucleolus. Oncogene. 2022;41(8):1155–1165. doi: 10.1038/s41388-021-02138-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Xu S-H, Zhu S, Wang Y, Huang J-Z, Chen M, Wu Q-X, He Y-T, Chen D, Yan G-R. ECD promotes gastric cancer metastasis by blocking E3 ligase ZFP91-mediated hnRNP F ubiquitination and degradation. Cell Death Dis. 2018;9(5):479. doi: 10.1038/s41419-018-0525-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Han J, An O, Ren X, Song Y, Tang SJ, Shen H, Ke X, Ng VHE, Tay DJT, Tan HQ, Kappei D, Yang H, Chen L. Multilayered control of splicing regulatory networks by DAP3 leads to widespread alternative splicing changes in cancer. Nat Commun. 2022;13(1):1793. doi: 10.1038/s41467-022-29400-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Xie C, Long F, Li L, Li X, Ma M, Lu Z, Wu R, Zhang Y, Huang L, Chou J, Gong N, Hu G, Lin C. PTBP3 modulates P53 expression and promotes colorectal cancer cell proliferation by maintaining UBE4A mRNA stability. Cell Death Dis. 2022;13(2):128. doi: 10.1038/s41419-022-04564-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Cristini A, Groh M, Kristiansen MS, Gromak N. RNA/DNA hybrid interactome identifies DXH9 as a molecular player in transcriptional termination and R-loop-associated DNA damage. Cell Rep. 2018;23(6):1891–1905. doi: 10.1016/j.celrep.2018.04.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124.Liu J, Li Y, Tong J, Gao J, Guo Q, Zhang L, Wang B, Zhao H, Wang H, Jiang E, Kurita R, Nakamura Y, Tanabe O, Engel JD, Bresnick EH, Zhou J, Shi L. Long non-coding RNA-dependent mechanism to regulate heme biosynthesis and erythrocyte development. Nat Commun. 2018;9(1):4386. doi: 10.1038/s41467-018-06883-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Liu L, Zhang P, Dong X, Li H, Li S, Cheng S, Yuan J, Yang X, Qian Z, Dong J. Circ_0001367 inhibits glioma proliferation, migration and invasion by sponging miR-431 and thus regulating NRXN3. Cell Death Dis. 2021;12(6):536. doi: 10.1038/s41419-021-03834-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126.Dong W, Dai ZH, Liu FC, Guo XG, Ge CM, Ding J, Liu H, Yang F. The RNA-binding protein RBM3 promotes cell proliferation in hepatocellular carcinoma by regulating circular RNA SCD-circRNA 2 production. EBioMedicine. 2019;45:155–167. doi: 10.1016/j.ebiom.2019.06.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127.Yan Y, Nie Y, Peng C, Xing F, Ji S, Liu H, Zhu C. Correction to: The circular RNA hsa_circ_0001394 promotes hepatocellular carcinoma progression by targeting the miR-527/UBE2A axis. Cell Death Discov. 2022;8(1):145. doi: 10.1038/s41420-022-00955-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Kaiser RWJ, Ignarski M, Van Nostrand EL, Frese CK, Jain M, Cukoski S, Heinen H, Schaechter M, Seufert L, Bunte K, Frommolt P, Keller P, Helm M, Bohl K, Höhne M, Schermer B, Benzing T, Höpker K, Dieterich C, Yeo GW, Müller R-U, Fabretti F. A protein-RNA interaction atlas of the ribosome biogenesis factor AATF. Sci Rep. 2019;9(1):11071. doi: 10.1038/s41598-019-47552-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Luciano P, Jeon J, El-kaoutari A, Challal D, Bonnet A, Barucco M, Candelli T, Jourquin F, Lesage P, Kim J, Libri D, Géli V. Binding to RNA regulates Set1 function. Cell Discov. 2017;3(1):17040. doi: 10.1038/celldisc.2017.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Gajdušková P, Ruiz de los Mozos I, Rájecký M, Hluchý M, Ule J, Blazek D. CDK11 is required for transcription of replication-dependent histone genes. Nat Struct Mol Biol. 2020;27(5):500–510. doi: 10.1038/s41594-020-0406-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.Van Nostrand EL, Freese P, Pratt GA, Wang X, Wei X, Xiao R, Blue SM, Chen J-Y, Cody NAL, Dominguez D, Olson S, Sundararaman B, Zhan L, Bazile C, Bouvrette LPB, Bergalet J, Duff MO, Garcia KE, Gelboin-Burkhart C, Hochman M, Lambert NJ, Li H, McGurk MP, Nguyen TB, Palden T, Rabano I, Sathe S, Stanton R, Su A, Wang R, Yee BA, Zhou B, Louie AL, Aigner S, Fu X-D, Lécuyer E, Burge CB, Graveley BR, Yeo GW. Author correction: a large-scale binding and functional map of human RNA-binding proteins. Nature. 2021;589(7842):E5–E5. doi: 10.1038/s41586-020-03067-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132.Qin W, Myers SA, Carey DK, Carr SA, Ting AY. Spatiotemporally-resolved mapping of RNA binding proteins via functional proximity labeling reveals a mitochondrial mRNA anchor promoting stress recovery. Nat Commun. 2021;12(1):4980. doi: 10.1038/s41467-021-25259-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Brannan KW, Chaim IA, Marina RJ, Yee BA, Kofman ER, Lorenz DA, Jagannatha P, Dong KD, Madrigal AA, Underwood JG, Yeo GW. Robust single-cell discovery of RNA targets of RNA-binding proteins and ribosomes. Nat Methods. 2021;18(5):507–519. doi: 10.1038/s41592-021-01128-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134.Cottrell KA, Chaudhari HG, Cohen BA, Djuranovic S. PTRE-seq reveals mechanism and interactions of RNA binding proteins and miRNAs. Nat Commun. 2018;9(1):301. doi: 10.1038/s41467-017-02745-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Lam JH, Li Y, Zhu L, Umarov R, Jiang H, Héliou A, Sheong FK, Liu T, Long Y, Li Y, Fang L, Altman RB, Chen W, Huang X, Gao X. A deep learning framework to predict binding preference of RNA constituents on protein surface. Nat Commun. 2019;10(1):4941. doi: 10.1038/s41467-019-12920-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136.Xie J, Zheng J, Hong X, Tong X, Liu S. PRIME-3D2D is a 3D2D model to predict binding sites of protein–RNA interaction. Commun Biol. 2020;3(1):384. doi: 10.1038/s42003-020-1114-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137.Liu R, Yang J, Yao J, Zhao Z, He W, Su N, Zhang Z, Zhang C, Zhang Z, Cai H, Zhu L, Zhao Y, Quan S, Chen X, Yang Y. Optogenetic control of RNA function and metabolism using engineered light-switchable RNA-binding proteins. Nat Biotechnol. 2022;40(5):779–786. doi: 10.1038/s41587-021-01112-1. [DOI] [PubMed] [Google Scholar]
- 138.Park B, Han K. Discovering protein-binding RNA motifs with a generative model of RNA sequences. Comput Biol Chem. 2020;84:107171. doi: 10.1016/j.compbiolchem.2019.107171. [DOI] [PubMed] [Google Scholar]
- 139.van Nues R, Schweikert G, de Leau E, Selega A, Langford A, Franklin R, Iosub I, Wadsworth P, Sanguinetti G, Granneman S. Kinetic CRAC uncovers a role for Nab3 in determining gene expression profiles during stress. Nat Commun. 2017;8(1):12. doi: 10.1038/s41467-017-00025-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Urdaneta EC, Vieira-Vieira CH, Hick T, Wessels H-H, Figini D, Moschall R, Medenbach J, Ohler U, Granneman S, Selbach M, Beckmann BM. Purification of cross-linked RNA-protein complexes by phenol-toluol extraction. Nat Commun. 2019;10(1):990. doi: 10.1038/s41467-019-08942-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141.Zheng J, Zhang X, Zhao X, Tong X, Hong X, Xie J, Liu S. Deep-RBPPred: Predicting RNA binding proteins in the proteome scale based on deep learning. Sci Rep. 2018;8(1):15264. doi: 10.1038/s41598-018-33654-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 142.Yang J-S, Garriga-Canut M, Link N, Carolis C, Broadbent K, Beltran-Sastre V, Serrano L, Maurer SP. rec-YnH enables simultaneous many-by-many detection of direct protein–protein and protein–RNA interactions. Nat Commun. 2018;9(1):3747. doi: 10.1038/s41467-018-06128-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143.Bouhedda F, Fam KT, Collot M, Autour A, Marzi S, Klymchenko A, Ryckelynck M. A dimerization-based fluorogenic dye-aptamer module for RNA imaging in live cells. Nat Chem Biol. 2020;16(1):69–76. doi: 10.1038/s41589-019-0381-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144.Briese M, Haberman N, Sibley CR, Faraway R, Elser AS, Chakrabarti AM, Wang Z, König J, Perera D, Wickramasinghe VO, Venkitaraman AR, Luscombe NM, Saieva L, Pellizzoni L, Smith CWJ, Curk T, Ule J. Publisher Correction: A systems view of spliceosomal assembly and branchpoints with iCLIP. Nat Struct Mol Biol. 2021;28(5):455–455. doi: 10.1038/s41594-021-00595-5. [DOI] [PubMed] [Google Scholar]
- 145.Theil K, Imami K, Rajewsky N. Identification of proteins and miRNAs that specifically bind an mRNA in vivo. Nat Commun. 2019;10(1):4205. doi: 10.1038/s41467-019-12050-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Perez-Perri JI, Rogell B, Schwarzl T, Stein F, Zhou Y, Rettel M, Brosig A, Hentze MW. Discovery of RNA-binding proteins and characterization of their dynamic responses by enhanced RNA interactome capture. Nat Commun. 2018;9(1):4408. doi: 10.1038/s41467-018-06557-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 147.Bae JW, Kim S, Kim VN, Kim J-S. Photoactivatable ribonucleosides mark base-specific RNA-binding sites. Nat Commun. 2021;12(1):6026. doi: 10.1038/s41467-021-26317-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 148.Panhale A, Richter FM, Ramírez F, Shvedunova M, Manke T, Mittler G, Akhtar A. CAPRI enables comparison of evolutionarily conserved RNA interacting regions. Nat Commun. 2019;10(1):2682. doi: 10.1038/s41467-019-10585-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 149.Komatsu KR, Taya T, Matsumoto S, Miyashita E, Kashida S, Saito H. RNA structure-wide discovery of functional interactions with multiplexed RNA motif library. Nat Commun. 2020;11(1):6275. doi: 10.1038/s41467-020-19699-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150.Vieira-Vieira CH, Dauksaite V, Sporbert A, Gotthardt M, Selbach M. Proteome-wide quantitative RNA-interactome capture identifies phosphorylation sites with regulatory potential in RBM20. Mol Cell. 2022;82(11):2069–2083.e2068. doi: 10.1016/j.molcel.2022.03.024. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Not applicable.