Abstract
Identification of host transcriptional response signatures has emerged as a new paradigm for infection diagnosis. For clinical applications, signatures must robustly detect the pathogen of interest without cross-reacting with unintended conditions. To evaluate the performance of infectious disease signatures, we developed a framework that includes a compendium of 17,105 transcriptional profiles capturing infectious and non-infectious conditions and a standardized methodology to assess robustness and cross-reactivity. Applied to 30 published signatures of infection, the analysis showed that signatures were generally robust in detecting viral and bacterial infections in independent data. Asymptomatic and chronic infections were also detectable, albeit with decreased performance. However, many signatures were cross-reactive with unintended infections and aging. In general, we found robustness and cross-reactivity to be conflicting objectives, and we identified signature properties associated with this trade-off. The data compendium and evaluation framework developed here provide a foundation for the development of signatures for clinical application. A record of this paper’s transparent peer review process is included in the supplemental information.
Keywords: transcriptional host response signature, infection diagnosis, robustness, cross-reactivity, virus, bacteria, data compendium, influenza signature, non-infectious conditions, aging, signature evaluation framework
Graphical abstract

Host response signatures for infection diagnosis should robustly and specifically detect the intended pathogen. To evaluate signatures of infection, we develop a standardized framework and data compendium. Published viral and bacterial signatures robustly detect the intended pathogen yet cross-react with unintended conditions. We identify signature properties to optimize infection diagnosis.
Introduction
The ability to diagnose infectious diseases has a profound impact on global health. Most recently, diagnostic testing for SARS-CoV-2 infection has helped contain the COVID-19 pandemic, lessening the strain on healthcare systems. As a further example, diagnostic technologies that discriminate bacterial from viral infections can inform the prescription of antibiotics. This is a high-stakes clinical decision: if prescribed for bacterial infections, the use of antibiotics substantially reduces mortality,1 but if prescribed for viral infections, their misuse exacerbates antimicrobial resistance.2
Standard tests for infection diagnosis involve a variety of technologies including microbial cultures, PCR assays, and antigen-binding assays. Despite the diversity in technologies, standard tests generally share a common design principle, which is to directly quantify pathogen material in patient samples. As a consequence, standard tests have poor detection, particularly early after infection, before the pathogen replicates to detectable levels. For example, PCR-based tests for SARS-CoV-2 infection may miss 60%–100% of infections within the first few days of infection due to insufficient viral genetic material.3 , 4 Similarly, a study of community acquired pneumonia found that pathogen-based tests failed to identify the causative pathogen in over 60% of patients.5 To overcome these limitations, new tools for infection diagnosis are urgently needed.
Host transcriptional response assays have emerged as a new paradigm to diagnose infections.6 , 7 , 8 , 9 , 10 Research in the field has produced a variety of host response signatures to detect general viral or bacterial infections as well as signatures for specific pathogens such as influenza virus.6 , 11 , 12 , 13 , 14 , 15 Unlike standard tests that measure pathogen material, these assays monitor changes in gene expression in response to infection.16 For example, transcriptional upregulation of IFN response genes may indicate an ongoing viral infection because these genes take part in the host antiviral response.17 Host response assays have a major potential advantage over pathogen-based tests because they may detect an infection even when the pathogen material is undetectable through direct measurements.
Development of host response assays that can be implemented clinically poses new methodological problems. The most challenging problem is identifying the so-called “infection signature” for a pathogen of interest, that is, a set of host transcriptional changes induced in response to that pathogen. Signature performance is characterized along two axes, robustness and cross-reactivity. Robustness is defined as the ability of a signature to detect the intended infectious condition consistently in multiple independent cohorts. Cross-reactivity is defined as the extent to which a signature predicts any condition other than the intended one. To be clinically viable, an infection signature must simultaneously demonstrate high robustness and low cross-reactivity. A robust signature that does not demonstrate low cross-reactivity would detect unintended conditions, such as other infections (e.g., viral signatures detecting bacterial infections) and/or non-infectious conditions involving abnormal immune activation.
The clinical applicability of host response signatures ultimately depends on a rigorous evaluation of their robustness and cross-reactivity properties. Such an evaluation is a complex task because it requires integrating and analyzing massive amounts of transcriptional studies involving the pathogen of interest along with a wide variety of other infectious and non-infectious conditions that may cause cross-reactivity. Despite recent progress in this direction,10 , 18 , 19 a general framework to benchmark both robustness and cross-reactivity of candidate signatures is still lacking.
Here, we establish a general framework for systematic quantification of robustness and cross-reactivity of a candidate signature, based on a fine-grained curation of massive public data and development of a standardized signature scoring method. Using this framework, we demonstrated that published signatures are generally robust but substantially cross-reactive with infectious and non-infectious conditions. Further analysis of 200,000 synthetic signatures identified an inherent trade-off between robustness and cross-reactivity and determined signature properties associated with this trade-off. Our framework, freely accessible at https://kleinsteinlab.shinyapps.io/compendium_shiny_app/, lays the foundation for the discovery of signatures of infection for clinical application.
Results
A curated set of human transcriptional infection signatures
While many transcriptional host response signatures of infection have been published, their robustness and cross-reactivity properties have not been systematically evaluated. To identify published signatures for inclusion in our systematic evaluation, we performed a search of NCBI PubMed for publications describing immune profiling of viral or bacterial infections (Figure 1 A). We initially focused our curation on general viral or bacterial (rather than pathogen-specific) signatures from human whole-blood or peripheral-blood mononuclear cells (PBMCs). We identified 24 signatures that were derived using a wide range of computational approaches, including differential expression analyses,7 , 20 , 21 , 22 gene clustering,23 , 24 regularized logistic regression,19 , 20 , 25 and meta-analyses.8 , 11
Figure 1.
A curated set of human transcriptional infection signatures
(A) A standardized process was used to identify and curate published blood-based (whole blood or PBMC) transcriptional signatures of infection in humans from NCBI PubMed. Selection focused on signatures to detect general responses to viral (V) and bacterial (B) infections compared with control subjects. Signatures developed to differentiate viral from bacterial infections in a direct contrast (V/B) were also included. Signatures were parsed into positive (upregulated with respect to the intended contrast) and negative (downregulated) gene lists. Each signature was annotated with metadata including method of derivation, cohort details, and accessions for discovery datasets. Overall, this workflow produced 24 signatures curated for evaluation.
(B–D) The composition of each group of signatures (11 viral, 7 bacterial, and 6 V/B signatures) was characterized, including signature size, most frequently occurring genes and significantly enriched pathways (FDR < 0.05, selected examples are displayed). Frequency of occurrence for each gene is listed in parentheses. Enrichments were computed based on the total pool of genes in each signature group.
(E) Pairwise Jaccard similarity coefficients were computed between signatures using concatenated positive and negative gene lists.
The signatures were annotated with multiple characteristics that were needed for the evaluation of performance. The most important characteristic was the intended use of the signatures. The intended use of the included signatures was to detect viral infection (V), bacterial infection (B), or directly discriminate between viral and bacterial infections (V/B). For each signature, we recorded a set of genes and a group I versus group II comparison capturing the design of the signature, where group I was the intended infection type, and group II was a control group. For most viral and bacterial signatures, group II was comprised healthy controls; in a few cases, it was comprised of non-infectious illness controls. For signatures distinguishing viral and bacterial infections (V/B), we conventionally took the bacterial infection group as the control group.
We parsed the genes in these signatures as either “positive” or “negative” based on whether they were upregulated or downregulated in the intended group, respectively. We also manually annotated the PubMed identifiers for the publication in which the signature was reported, accession records to identify discovery datasets used to build each signature, association of the signature with either acute or chronic infection, and additional metadata related to demographics and experimental design (Table S1).
This curation process identified 11 viral (V) signatures intended to capture transcriptional responses that are common across many viral pathogens, 7 bacterial (B) signatures intended to capture transcriptional responses common across bacterial pathogens, and 6 viral versus bacterial (V/B) signatures discriminating between viral and bacterial infections. Viral signatures varied in size between 3 and 396 genes. Several genes appeared in multiple viral signatures. For example, OASL, an interferon-induced gene with antiviral function,26 appeared in 6 of 11 signatures. Enrichment analysis on the pool of viral signature genes showed significantly enriched terms consistent with antiviral immunity, including response to type I interferon (Figure 1B). Bacterial signatures ranged in size from 2 to 69 genes, and enrichment analysis again highlighted expected pathways associated with antibacterial immunity (Figure 1C). V/B signatures varied in size from 2 to 69 genes. The most common genes among V/B signatures were OASL and IFI27, both of which were also highly represented viral signature genes, and many of the same antiviral pathways were significantly enriched among V/B signature genes (Figure 1D). We further investigated the similarity between viral, bacterial, and V/B signatures and found that many viral signatures shared genes with each other and V/B signatures, but bacterial signatures shared fewer similarities with each other (Figure 1E). Overall, our curation produced a structured and well-annotated set of transcriptional signatures for systematic evaluation.
A compendium of human transcriptional infection datasets
To profile the performance of the curated infection signatures, we compiled a large compendium of datasets capturing host blood transcriptional responses to a wide diversity of pathogens. We carried out a comprehensive search in the NCBI Gene Expression Omnibus (GEO)27 selecting transcriptional responses to in vivo viral, bacterial, parasitic, and fungal infections in human whole blood or PBMCs. We screened over 8,000 GEO records and identified 136 transcriptional datasets that met our inclusion criteria (see STAR Methods). Furthermore, to evaluate whether infection signatures cross-react with non-infectious conditions with documented immunomodulating effects, we also compiled an additional 14 datasets containing transcriptomes from the blood of aged and obese individuals.28 , 29 All datasets were downloaded from GEO and passed through a standardized pipeline. Briefly, the pipeline included (1) uniform pre-processing of raw data files where possible, (2) remapping of available gene identifiers to Entrez Gene IDs, and (3) detection of outlier samples.30 In aggregate, we compiled, processed, and annotated 150 datasets to include in our data compendium (Figure 2 A; Table S2; see STAR Methods for details).
Figure 2.
A compendium of human transcriptional infection datasets
(A) A standardized procedure was used to build a compendium of human transcriptional infection datasets profiling PBMCs or whole blood. After a systematic search of NCBI GEO, 150 datasets were selected that profile in vivo responses to viral, bacterial, and parasitic infections, as well as immunomodulating non-infectious conditions. Datasets were passed through a standardized pre-processing pipeline. A total of 17,501 individual samples were annotated with condition type (e.g., infectious, non-infectious, or healthy control) as well as infection type (e.g., viral, bacterial, or parasitic) and the corresponding causative pathogen (e.g., influenza virus). Datasets were annotated with a study design (either cross-sectional or longitudinal).
(B) Datasets were labeled hierarchically by condition(s) profiled: infectious/non-infectious, viral/bacterial/other, and by unique pathogen. Within each layer of the hierarchy, bar heights correspond to the relative frequency of dataset labels.
(C–F) We evaluated technical characteristics of the viral and bacterial datasets within our compendium that may impact downstream analyses. We compared the number of samples per dataset (C), the number of datasets following each study design (D), the frequency of platform manufacturers (E), and the frequency of whole blood and PBMC samples (F). The number of studies refers to the number of datasets in each category.
The compendium datasets showed wide variability in study design, sample composition, and available metadata necessitating annotation both at the study level and at the finer-grained sample level. Datasets followed either cross-sectional study designs, where individual subjects were profiled once for a snapshot of their infection, or longitudinal study designs, in which individual subjects were profiled at multiple time points over the course of an infection. For longitudinal datasets, we also recorded subject identifiers and labeled time points. Many datasets contained multiple subgroups, each profiling infection with a different pathogen. Detailed review of the clinical methods and metadata for each study enabled us to annotate individual samples with infectious class (e.g., viral or bacterial) and causative pathogen. For clinical variables, we manually recorded whether datasets profiled acute or chronic infections according to the authors and annotated symptom severity when available. We further supplemented this information with biological sex, which we inferred computationally (see STAR Methods). In total, we annotated 16,173 infection and control samples in a consistent way, capturing host responses to viral, bacterial, and parasitic infections. We similarly annotated the additional 932 samples from aging and obesity datasets including young and lean controls, respectively. In aggregate we captured a broad range of more than 35 unique pathogens and non-infectious conditions (Figure 2B).
Most of our compendium datasets were composed of viral and bacterial infection response profiles. We examined several technical factors that may bias the signature performance evaluation between these pathogen categories. Datasets profiling viral infections and datasets profiling bacterial infections contained similar numbers of samples, with median samples sizes of 75.5 and 63.0, respectively, though the largest viral studies contained more samples than the largest bacterial studies (Figure 2C). The number of cross-sectional studies was also nearly identical for both viral and bacterial infection datasets, but our compendium contained 20 viral longitudinal datasets (35% of viral) compared with 6 bacterial longitudinal datasets (10% of bacterial) (Figure 2D). We also examined the distribution of platforms used to generate viral and bacterial infection datasets and found that gene expression was measured most commonly using Illumina platforms, followed by Affymetrix, for both viral and bacterial datasets (Figure 2E). We also examined the frequency of whole blood and PBMC samples in our compendium (Figure 2F). We did not identify systematic differences in the viral and bacterial datasets within our compendium, and therefore we do not expect these differences to impact the interpretation of our signature evaluations.
Establishing a general framework for signature evaluation
We sought to quantify two measures of performance for all curated signatures: (1) robustness, the ability of a signature to predict its target infection in independent datasets not used for signature discovery, and (2) cross-reactivity, which we quantify as the undesired extent to which a signature predicts unrelated infections or conditions. An ideal signature would demonstrate robustness but not cross-reactivity, e.g., an ideal viral signature would predict viral infections in independent datasets but would not be associated with infections caused by pathogens such as bacteria or parasites.
To score each signature in a standardized way, we leveraged the geometric mean scoring approach described in Haynes et al.31 For each signature (i.e., a set of positive genes and an optional set of negative genes), we calculated its sample score from log-transformed expression values by taking the difference between the geometric mean of positive signature gene expression values and the geometric mean of negative signature gene expression values. For cross-sectional study designs, this generates a single signature score for each subject, but for longitudinal study designs, this approach produces a vector of scores across time points for each subject. The scores at different time points can vary dramatically as the transcriptional program underlying an immune response changes over the course of an infection.11 , 16 , 32 In this case, we chose the maximally discriminative time point, so that a signature is considered robust if it can detect the infection at any time point but also considered cross-reactive if it would produce a false-positive call at any time point (see STAR Methods). These subject scores were then used to quantify signature performance as the area under a receiver operator characteristic curve (AUROC) associated with each group comparison. This approach is advantageous because it is computationally efficient and model-free. The model-free property presents an advantage over parameterized models because it does not require transferring or re-training model coefficients between datasets. Overall, this framework enables us to evaluate the performance of all signatures in a standardized and consistent way in any dataset (Figure 3 A).
Figure 3.
Establishing a general framework for signature evaluation
(A) Given a signature as input, a standardized evaluation framework was developed to calculate performance metrics across the data compendium. Signatures are scored for each subject in a target transcriptomic dataset using a geometric mean score approach that accommodates both cross-sectional and longitudinal study designs. The subject scores, paired with group labels, are used to compute an AUROC. AUROC statistics measuring performance for the intended and unintended conditions of a signature are reported as robustness and cross-reactivity, respectively.
(B) Performance of curated signatures was computed in their respective discovery datasets. Shading indicates increasing AUROC in arbitrary units. Signature B7 and V/B1 are not included here because discovery datasets were not available (see Dataset search and selection in STAR Methods).
We assessed the framework by computing each signature’s performance on the datasets used originally for its discovery. If this approach is valid, signatures evaluated in their own discovery datasets should perform well, generating AUROCs close to 1. Consistent with this reasoning, we found that each signature strongly predicted infections in its own discovery datasets: the lowest observed median AUROC was 0.78 among viral signatures, 0.82 among bacterial signatures, and 0.90 among V/B signatures (Figure 3B). We also specifically evaluated the choice of geometric mean scoring and found that the performance of this scoring method for all signatures was highly correlated with logistic regression,19 , 20 , 25 a popular alternative approach (Figure S1; see STAR Methods). These results highlighted that while individual signatures were developed using many different methods, signatures can be reliably evaluated using a standardized framework built on geometric mean scoring.
Existing signatures of bacterial and viral infection are generally robust
Having established a common framework for evaluating signatures, we next investigated the robustness of all curated signatures. Each signature in our compendium was first evaluated on every non-discovery (i.e., independent) dataset profiling intended pathogen responses and healthy controls. For example, all signatures of viral infection were evaluated on datasets that profiled viral pathogens and healthy controls. We use the median AUROC threshold of 0.7 for robustness determination (see STAR Methods). Overall, we found that 10 out of 11 viral signatures, 5 out of 7 bacterial signatures, and all 6 V/B signatures achieved a median AUROC greater than 0.70 in predicting infections in independent data (Figures 4 A–4C; Table S3). Additionally, because some signatures were derived using non-infectious illness controls (e.g., systemic inflammatory response syndrome), we characterized viral and bacterial signature performance in datasets that profiled this contrast.19 , 33 In this evaluation, 9 out of 11 viral signatures and 2 out of 7 bacterial signatures achieved a median AUROC greater than 0.70 (Figures S2A and S2B; Table S3), suggesting that bacterial but not viral signatures were sensitive to the control group used for signature evaluation. We categorized a signature as robust if its median AUROC in either set of independent data (i.e., versus healthy or non-infectious illness controls) was greater than 0.70, indicating strong predictive performance. Overall, we identified 10 viral, 6 bacterial, and all 6 V/B signatures that were robust.
Figure 4.
Existing signatures of bacterial and viral infection are generally robust when evaluated in independent data
(A and B) Viral (A) and bacterial (B) signature robustness was evaluated in independent datasets profiling intended infections and healthy controls. Ridge plots indicate AUROC distributions for each signature. Signatures with a median AUROC greater than 0.70 were considered robust. ‡ indicates a signature derived using non-infectious illness controls.
(C) V/B signature robustness was evaluated by computing AUROCs for distinguishing viral infections from bacterial infections in independent datasets profiling both infection types. ‡ indicates a signature derived using non-infectious illness controls.
(D and E) Signature robustness was also evaluated separately for selected pathogens that were not included during signature discovery. Viral signature performance was evaluated in HIV infection (D), where the only available datasets were those profiling HIV infected subjects and healthy controls. Bacterial signature performance was evaluated in B. pseudomallei infection compared with healthy controls (E) and with non-infectious illness controls (Figure S2C).
(F) One dataset in the compendium (GSE103119, median V/B signature AUROC < 0.50) was unique in its profiling of Mycoplasma infection. V/B signature AUROCs were compared for this dataset when including (+) or excluding (−) this pathogen (paired Wilcoxon signed-rank test, n = 6 V/B signature pairs). For (A)–(F), distributions shown in color indicate signature robustness.
(G) All 24 signatures were evaluated in male and female subjects separately.
(H) Viral signature performance was compared between acute and chronic infection datasets (Wilcoxon signed-rank test, n = 11 viral signature pairs).
(I) Viral signature performance was compared between symptomatic and asymptomatic subjects in a dataset profiling H3N2 influenza virus infections confirmed by viral shedding (GSE73072, paired Wilcoxon signed-rank test, n = 11 viral signature pairs).
Viral and bacterial signatures also robustly detected infections caused by pathogens in the same class (i.e., viral or bacterial) that were not included among discovery datasets. For example, all 10 robust viral signatures detected infections caused by HIV (median AUROC > 0.8, Figure 4D), while this pathogen was not included among the discovery datasets. Similarly, all robust bacterial signatures detected infections caused by B. pseudomallei (Figures 4E and S2C), while this pathogen was not included among the discovery datasets. These results suggest strong conservation of transcriptional programs underlying immune responses against a broad array of viruses and bacteria.
While signatures were discovered using different blood subsets and transcriptional profiling platforms, signature robustness was not strongly influenced by these factors. Signature performance in datasets profiling whole blood was strongly correlated with performance in datasets profiling PBMCs (r = 0.96, Figure S2D). Similarly, signature performance in datasets generated using Illumina microarray platforms was strongly correlated with performance in datasets generated using Affymetrix platforms (r = 0.91, Figure S2E).
There were a few datasets in our compendium where most signatures performed poorly, generating a dataset median AUROC less than or equal to 0.50. For viral signatures, we observed 2 such outlier datasets (Gene Expression Omnibus: GSE85599 and Gene Expression Omnibus: GSE59312) characterizing immune responses to acute and chronic Epstein-Barr virus (EBV) infection and chronic hepatitis C virus (HCV) infection, respectively (Figures S3A and S3B). For bacterial signatures, we observed one such outlier dataset (Gene Expression Omnibus: GSE625625), characterizing the response to Mycobacterium tuberculosis infection (TB, Figure S3C). We considered whether performance loss in the outlier datasets was due to the causative pathogens or to technical factors of the datasets. To address this, we analyzed signature performance in additional datasets profiling the same causative pathogens (EBV, HCV, and TB). We found that both viral and bacterial signatures showed robust performance in these additional datasets, demonstrating that performance loss in the outlier datasets was likely due to technical factors rather than the pathogen (Figures S3A–S3C). V/B signatures performed poorly in one outlier dataset profiling viral and bacterial pediatric pneumonia (Gene Expression Omnibus: GSE103119). While multiple datasets in the compendium profiled pediatric pneumonia, this outlier dataset was unique in its inclusion of Mycoplasma, a bacterium that lacks a cell wall, as the causative pathogen. To assess whether V/B signatures perform poorly for Mycoplasma infection specifically, we removed this pathogen from the dataset and repeated our signature assessment. The performance of signatures for non-Mycoplasma bacterial infections was significantly improved (p = 0.031, Figure 4F). Thus, the poor V/B signature performance in this case likely reflects a unique biological response for Mycoplasma that more closely resembles a viral infection.
Multiple infection characteristics modulate signature robustness
A number of factors can influence transcriptional profiles of infection and therefore signature performance, including subject demographics and clinical state.11 , 16 Although the availability and structure of metadata fields varied between datasets, we were able to systematically evaluate three variables of interest: sex, acute versus chronic infection characterization, and disease severity. Due to data limitations, the analysis of infection characterization and severity was conducted only for viral signatures. In particular, to address the effect of severity, we analyzed a single challenge study with three datasets, each of which profiled symptomatic and asymptomatic viral infections by human rhinovirus (hRV) or influenza virus H1N1 (Gene Expression Omnibus: GSE73072). In the analysis of these datasets, we only included subjects with evidence of viral shedding, to ensure that results were not due to lack of productive infection.
We found that signature performance was (1) strongly correlated between males and females (Figure 4G) and not significantly different in the two groups (paired t test, p > 0.05, Figure S4A), (2) robust for both acute and chronic infections albeit significantly lower in the latter case (Figure 4H, p < 0.001), and (3) robust for both symptomatic and asymptomatic infections but significantly lower in the second group (p = 0.0024, Figures 4I and S4B). Taken together, this analysis identified chronic versus acute characterization and severity, but not sex, as determinants that significantly affect signature performance.
Infection timing may also play an important role in modulating signature robustness.32 While time of pathogen exposure relative to sample collection is unknown for nearly all subjects in our compendium, eight datasets profiled healthy volunteers who were challenged with exposure to live respiratory viruses.12 , 34 To investigate the effect of timing on signature robustness in these datasets, we treated each time point post-infection as an independent cross-sectional study and computed signature AUROCs (Figure S5). Robust viral signatures achieved median AUROCs greater than 0.70 between 34 and 77 h post-infection and remained robust for several days through the end of each study. This analysis identified an initial, undetectable infection period followed by a prolonged, robust detection window for acute viral infection.
Nearly all infection signatures are cross-reactive with infectious and non-infectious conditions
We next assessed the cross-reactivity of the 22 infection signatures that were found to be robust. Signature cross-reactivity was estimated with the same evaluation framework used to assess robustness but now applied to data from infectious and non-infectious conditions for which the signature was not intended (see STAR Methods). A signature was considered cross-reactive with an unintended condition if the corresponding median AUROC was greater than 0.60 (see STAR Methods).
First, we examined the cross-reactivity of viral signatures with respect to bacterial infections and found that V2 and V8 were cross-reactive (Figure 5 A). All viral signatures showed a wide range of cross-reactivity values. We explored whether this variability might reflect the variability in the classes of bacterial pathogens. We classified the bacterial pathogens based on cell wall characteristics as Gram-positive, Gram-negative, and acid-fast bacteria and quantified cross-reactivity separately for these classes (Figure 5B). Most signatures cross-reacted with acid-fast bacteria (8 out of 10 signatures, median AUROCs between 0.66 and 0.92), a class that includes Mycobacterium tuberculosis. In contrast, only 3 of 10 signatures cross-reacted with Gram-negative bacteria, and only one with Gram-positive bacteria. Overall, only two signatures, V9 and V10, did not cross-react with any bacterial pathogen class. These results demonstrate that viral signatures’ cross-reactivity depends on pathogen cell wall characteristics and is highest for acid-fast bacteria. Despite this, it is possible to generate viral signatures that are not cross-reactive with any bacterial class.
Figure 5.
Nearly all infection signatures are cross-reactive with unintended infections or non-infectious conditions
(A) Robust viral signatures were evaluated for cross-reactivity in datasets profiling bacterial infections and healthy controls. Signatures with median AUROCs greater than 0.60 were considered cross-reactive.
(B) This cross-reactivity was further separated by bacterial class, using datasets in the compendium where this information was available.
(C) Robust bacterial signatures were evaluated for cross-reactivity in datasets profiling viral infections and healthy controls.
(D–F) All 22 robust signatures were evaluated for cross-reactivity in parasitic infection (D), obesity (E), and aging (F) datasets. V/B signatures were considered cross-reactive if they had a median AUROC greater than 0.60 or less than 0.40 (‡). This latter condition reflects that the designation of positive and negative genes in V/B signatures is arbitrary, and prediction in either direction is relevant to cross-reactivity. Signatures indicated in bold lettering were derived from discovery cohorts containing both pediatric and adult subjects. For (A)–(F), distributions shown in color indicate a lack of signature cross-reactivity.
Second, we examined the extent to which bacterial signatures predicted viral infections. We found that 4 of the 6 robust bacterial signatures were cross-reactive with viral infections (Figure 5C). Similarly, we observed a wide range of cross-reactivity values. To better understand this variability, we classified viral pathogens based on the presence or absence of a viral envelope and on viral genome characteristics, and quantified cross-reactivity separately for these groups. Cross-reactivity did not vary based on the presence of a viral envelope (Figure S6A). In contrast, we observed a trend indicating greater cross-reactivity among single-stranded RNA viruses compared with double-stranded DNA viruses (Figure S6B).
As a final test of cross-reactivity with infectious conditions, we quantified signature cross-reactivity with parasitic infections. In addition to the viral and bacterial signatures, it was also possible to measure the cross-reactivity of the V/B signatures in this evaluation because parasites were an unintended pathogen class for these signatures. Most bacterial signatures, but few viral or V/B signatures, showed cross-reactivity with parasitic infections (Figure 5D).
Non-infectious factors, such as obesity and aging, are associated with altered immune states that may produce false-positive signals for infectious signatures.28 , 29 We evaluated the cross-reactivity of viral, bacterial, and V/B signatures with these non-infectious conditions (see STAR Methods for clinical definitions; cohort accessions in Table S2). We found that viral, bacterial, and V/B signatures did not cross-react with obesity (Figure 5E; Table S3). In contrast, 6 of 10 viral, 2 of 6 bacterial, and 4 of 6 V/B signatures were cross-reactive with aging (Figure 5F; Table S3). In these cases, the signatures falsely detected an infection signal in healthy, older adults relative to young adults. Among the 10 signatures, which were not cross-reactive with aging, 7 were derived from cohorts containing both pediatric and adult subjects spanning an age range greater than 50 years (Figure 5F; Table S1).
Single-pathogen influenza signatures are robust but cross-reactive
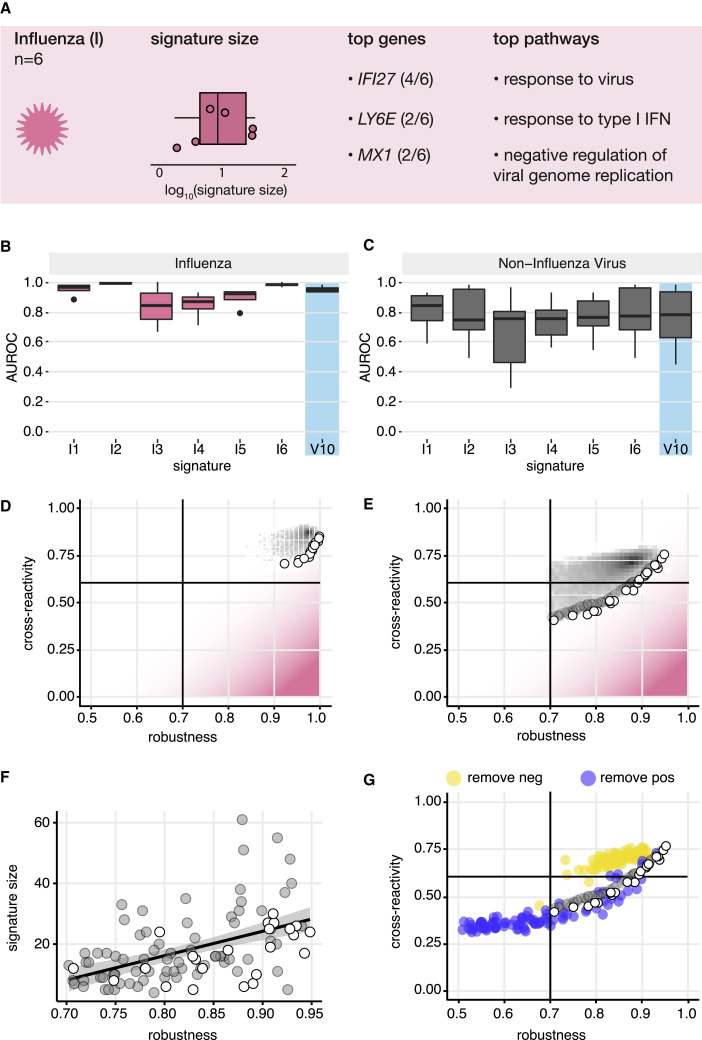
Our previous analyses focused on generic signatures of infection by a pathogen class, such as viral signatures. We next focused on signatures of infection by a single pathogen and chose to study the influenza virus because influenza causes a large, worldwide morbidity and mortality burden.35 Influenza was also the most abundant viral pathogen in our data compendium, with profiles from infected subjects reported in more than 30 datasets. A targeted search of NCBI PubMed identified 6 published signatures (I1–I6, Table S4) containing between 1 and 27 genes. These signatures included many interferon response genes that were also found in generic viral signatures (Figure 6 A; Table S4) and were significantly enriched for terms such as “response to type I interferon” and “response to virus.” Unlike general viral signatures, none of the curated influenza signatures were derived using non-infectious illness controls (e.g., sterile inflammatory response syndrome). Our evaluation therefore focused on discriminating influenza virus infection from healthy control samples.
Figure 6.
Analysis of influenza signatures demonstrates a trade-off between robustness and cross-reactivity
(A) A targeted literature search for influenza signatures was performed as a case study of single-pathogen signatures.
(B and C) Robustness (B) and cross-reactivity (C) of influenza signatures were evaluated using healthy control samples. General viral signature V10 was included as a positive control for viral detection.
(D) A meta-analysis procedure (see STAR Methods) was adapted to generate a pool of 124 candidate genes that are differentially expressed between influenza infection and healthy control samples. 100,000 synthetic signatures were generated by randomly sampling these candidate genes. For consistency with the meta-analysis procedure, performance was characterized using a weighted average AUROC (<AUROC>) across validation datasets. Robustness (<AUROC> in influenza datasets) and cross-reactivity (<AUROC> in non-influenza datasets) were evaluated for all candidate signatures (gray shading depicting density). Signatures comprising the Pareto front (white points) were identified to define signatures with locally optimal robustness and cross-reactivity characteristics. Pink shading indicates proximity to an ideal influenza signature with perfect robustness and no cross-reactivity.
(E) A similar analysis was carried out using a new set of candidate genes generated from the results of a meta-analysis directly contrasting influenza infection with non-influenza viral infection samples.
(F) A local neighborhood along the Pareto front in (E) was defined (gray points), and the relationship between signature size and signature robustness was examined.
(G) Each synthetic signature was separated into two signatures by removing either its positive (blue points) or negative (yellow points) gene sets. Performance was evaluated independently for each of these signatures.
To evaluate the performance of influenza signatures, we used as reference the best performing generic viral signature (V10). Compared with this generic viral signature, we expected influenza signatures to be at least as robust, but substantially less cross-reactive with viral infections not caused by influenza. We found that all influenza signatures robustly discriminated influenza infection from healthy controls, with median AUROCs ranging from 0.82 to 0.99, comparable with V10 (Figure 6B). However, all influenza signatures cross-reacted with non-influenza respiratory viral infections (such as hRV and respiratory syncytial virus [RSV] infection) with median AUROCs between 0.74 and 0.84 (Table S4). These values were comparable to those observed with the generic viral signature V10, confirming that influenza signatures lack influenza specificity (Figure 6C). Further evaluation of cross-reactivity showed that only two of six influenza signatures (I3 and I5) did not cross-react with bacterial infections, parasite infections, or aging (Figures S7A–S7D). Thus, while these influenza signatures were robust, they were cross-reactive with both infectious and non-infectious conditions.
Analysis of influenza signatures demonstrates a trade-off between robustness and cross-reactivity
We next investigated whether it was possible to reduce the cross-reactivity of influenza infection signatures. We followed the meta-analysis-based signature derivation approach used to develop V10, a general viral signature that was not cross-reactive. We performed a meta-analysis of 10 datasets profiling influenza infection and healthy control samples to identify an initial set of 124 differentially expressed candidate genes (data accessions in Table S5; gene IDs in Table S6; see STAR Methods). To characterize the performance space of signatures derived using these candidate genes, we generated a set of 100,000 synthetic signatures through random sampling (see STAR Methods). For each generated signature we assessed robustness using four independent influenza infection datasets and cross-reactivity using 12 datasets profiling other non-influenza respiratory viruses (Figure 6D; data accessions in Table S5). While most synthetic signatures were robust, they were also cross-reactive (Figure 6D), likely reflecting shared biology between respiratory virus infection responses.
Previous works have suggested that the inclusion of infectious controls may reduce signature cross-reactivity.8 , 19 , 33 Following this approach, we performed a meta-analysis of four datasets profiling both influenza and non-influenza viral infections, and identified 179 candidate genes that were differentially expressed between these two groups (Tables S6 and S7; see STAR Methods). We again generated a set of 100,000 synthetic signatures by randomly sampling from these genes. In this case, the signatures spanned a much wider range of robustness and cross-reactivity values, and we could identify a subset of signatures that were robust without being cross-reactive (Figure 6E). These results showed that single-pathogen signatures can satisfy both objectives through the inclusion of targeted infections as control groups.
Examining the performance of all signatures in Figure 6E, we found that robustness and cross-reactivity were positively correlated (r = 0.69). This suggested that maximizing robustness and minimizing cross-reactivity are conflicting objectives. A salient way to study such a trade-off is to analyze the Pareto-efficient solutions, a concept developed in multi-objective optimization.36 In the general case, Pareto-efficient solutions to a multi-objective optimization problem are the ones for which no individual objective (e.g., robustness) can be improved without impairing the other objective (e.g., cross-reactivity). Over the space of candidate signatures, the set of Pareto-efficient solutions, called the Pareto front, corresponds to signatures with locally optimal robustness and cross-reactivity characteristics (Figures 6D and 6E, white points).
To explore factors that may influence the trade-off between robustness and cross-reactivity of influenza signatures, we analyzed the properties of signatures along the Pareto front (Figure 6E). To increase the number of observations for this analysis, we augmented the Pareto front with signatures in a proximal neighborhood, for a total of 100 signatures (Figure 6E, gray points; see STAR Methods). Looking along the augmented Pareto front, we found that the signature size was positively correlated with robustness (r = 0.50, Figure 6F). However, signatures with larger size also suffered from higher cross-reactivity (r = 0.52, Figure S7E). Furthermore, we found that positive and negative genes in the signatures played different roles. After removing negative genes, signatures composed only of positive genes had better robustness but higher cross-reactivity. Conversely, after removing positive genes, signatures composed only of negative genes had worse robustness but reduced cross-reactivity (Figure 6G). These results combined suggest that signature size and inclusion of both positive and negative genes are critical factors in designing optimal single-pathogen signatures.
A web application to assess signature performance
To make our dataset curations available to the wider research community, we have created a web application that allows users to upload gene signatures and evaluate their performance in viral infection, bacterial infection, parasitic infection, aging, and obesity datasets. This tool is available at https://kleinsteinlab.shinyapps.io/compendium_shiny_app/.
Discussion
In this study, we established a framework for benchmarking the performance of host transcriptional response signatures of infection. The scope of our study was fundamentally different from the scope of previous studies that focused on deriving potential signatures of viral or bacterial infections. Going beyond initial efforts to compare the robustness of existing signatures,10 , 18 , 19 our evaluation framework is the first to provide a reference space where any arbitrary signature of infection can be rigorously assessed along two equally critical axes, robustness, and cross-reactivity. The framework is based on an extensive data curation of 17,105 blood transcriptional profiles from infectious and non-infectious conditions combined with a universal, model-free signature scoring method. By evaluating the robustness and cross-reactivity of 30 published and 200,000 synthetic signatures, we gained new insight toward the implementation of host response assays for clinical infection diagnosis.
Our evaluation found that most signatures were remarkably robust in detecting their intended conditions, consistent with previous work.18 Signatures generalized well to independent cohorts, and signatures intended to broadly detect viral or bacterial infection even generalized to pathogens not included in their discovery data. Signatures were also robust to varying infection severity and clinical phase, albeit with reduced performance. Viral signatures also remained robust for several days post-infection, suggesting that signatures are capturing sustained biological processes. These findings raise the question as to what biological underpinnings make the signatures of infection so robust. In the case of viral infections, we observed that all robust signatures included members of the type I IFN pathway, a highly conserved antiviral mechanism. Generally, signatures of infection may be more robust if they capture immunological pathways conserved across a pathogen class. Consistent with this hypothesis, we envisage that signatures of infection that explicitly include relevant immunological pathways would provide further gains in robustness.
Additional curation and analysis are required to verify that our results are consistent for RNA-seq data, as all compendium data and signatures, with the exception of B7, were derived from microarray platforms. Given the enhanced sensitivity of RNA-seq measurements over those from microarrays, we might expect our framework to systematically underestimate the performance of signature B7. Despite this, we found B7 to be robust, even in bacterial pathogens for which it was not explicitly designed (Figures 4B and 4E). B7 does not contain negative genes, and we therefore do not expect that improved detection would suppress the cross-reactivity demonstrated in Figure 5. Technical differences between transcriptional profiling technologies are thus unlikely to change the conclusions of our analysis.
While the evaluated signatures were robust, we found that they suffered from substantial cross-reactivity in two important ways. First, likely due to conservation in immune responses, signatures of infection cross-reacted with unintended pathogen classes (e.g., viral signatures detected bacterial infections, and vice versa). Viral signatures were especially cross-reactive with infections caused by acid-fast bacteria such as Mycobacterium tuberculosis, which may reflect the strong type I IFN response induced by this pathogen.37 This pathogen was the most abundant acid-fast bacterial pathogen, and so we are uncertain whether this is a pathogen-specific effect or if it applies to all bacteria with this cell wall characteristic. Bacterial signatures were slightly more cross-reactive with infections caused by viruses with single-stranded genomes, which suggests conserved immune response mechanisms that require further investigation. Second, both viral and bacterial signatures cross-reacted with aging. To our knowledge, this is the first demonstration that signatures of infection can cross-react with non-infectious conditions. From a diagnostic perspective, this in-depth analysis emphasizes the need for infection signatures to undergo extensive cross-reactivity testing before clinical implementation. The cross-reactivity testing should include unintended pathogens, both viral and bacterial, but also aging, and possibly other non-infectious inflammatory conditions.
In-depth analysis of published and synthetic-influenza-specific signatures identified an inherent trade-off between robustness and cross-reactivity. We also identified several signature properties associated with this trade-off, such as size and the inclusion of both positively and negatively regulated genes. Larger signatures may be more robust but are also less suitable for clinical application: PCR-based diagnostic platforms impose a ceiling on the number of genes that can be measured,38 and our results demonstrate that larger signatures are generally more cross-reactive.
Although discovering robust signatures with limited cross-reactivity was beyond the scope of this work, our results are useful to guide the derivation of future signatures. For example, our results suggest that including both young and elderly subjects during signature discovery may improve cross-reactivity with aging. Additionally, we demonstrate that inclusion of unintended infections as targeted contrasts during signature discovery can greatly reduce cross-reactivity. Because robustness and cross-reactivity are conflicting objectives, pathogen-specific signatures could be identified as solutions of a multi-objective optimization problem, with appropriate constraints on the signature that reflect the desired properties. Developing methods to discover robust signatures that do not cross-react with unintended infections or non-infectious conditions is of great interest, which we explore further in our companion manuscript.39
In summary, our framework lays the foundation for the discovery of signatures of infection for clinical application. We implemented the framework as a publicly accessible, user-friendly resource (https://kleinsteinlab.shinyapps.io/compendium_shiny_app/).
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Deposited Data | ||
| Data compendium | Gene Expression Omnibus (GEO) | GSE122737 |
| Data compendium | GEO | GSE122737 |
| Data compendium | GEO | GSE117827 |
| Data compendium | GEO | GSE117613 |
| Data compendium | GEO | GSE117613 |
| Data compendium | GEO | GSE116149 |
| Data compendium | GEO | GSE116149 |
| Data compendium | GEO | GSE116014 |
| Data compendium | GEO | GSE114466 |
| Data compendium | GEO | GSE111368 |
| Data compendium | GEO | GSE110551 |
| Data compendium | GEO | GSE110106 |
| Data compendium | GEO | GSE103842 |
| Data compendium | GEO | GSE103119 |
| Data compendium | GEO | GSE101710 |
| Data compendium | GEO | GSE101702 |
| Data compendium | GEO | GSE97741 |
| Data compendium | GEO | GSE97298 |
| Data compendium | GEO | GSE94916 |
| Data compendium | GEO | GSE94916 |
| Data compendium | GEO | GSE85599 |
| Data compendium | GEO | GSE83456 |
| Data compendium | GEO | GSE83223 |
| Data compendium | GEO | GSE82050 |
| Data compendium | GEO | GSE81926 |
| Data compendium | GEO | GSE81246 |
| Data compendium | GEO | GSE77939 |
| Data compendium | GEO | GSE77528 |
| Data compendium | GEO | GSE73464 |
| Data compendium | GEO | GSE73462 |
| Data compendium | GEO | GSE73461 |
| Data compendium | GEO | GSE73072 |
| Data compendium | GEO | GSE72810 |
| Data compendium | GEO | GSE72809 |
| Data compendium | GEO | GSE69606 |
| Data compendium | GEO | GSE69528 |
| Data compendium | GEO | GSE69039 |
| Data compendium | GEO | GSE68310 |
| Data compendium | GEO | GSE68004 |
| Data compendium | GEO | GSE67059 |
| Data compendium | GEO | GSE65682 |
| Data compendium | GEO | GSE65219 |
| Data compendium | GEO | GSE64456 |
| Data compendium | GEO | GSE64338 |
| Data compendium | GEO | GSE64338 |
| Data compendium | GEO | GSE63990 |
| Data compendium | GEO | GSE63117 |
| Data compendium | GEO | GSE62525 |
| Data compendium | GEO | GSE61821 |
| Data compendium | GEO | GSE61754 |
| Data compendium | GEO | GSE60244 |
| Data compendium | GEO | GSE59743 |
| Data compendium | GEO | GSE59654 |
| Data compendium | GEO | GSE59635 |
| Data compendium | GEO | GSE59312 |
| Data compendium | GEO | GSE57065 |
| Data compendium | GEO | GSE56960 |
| Data compendium | GEO | GSE56619 |
| Data compendium | GEO | GSE56153 |
| Data compendium | GEO | GSE55205 |
| Data compendium | GEO | GSE54992 |
| Data compendium | GEO | GSE53232 |
| Data compendium | GEO | GSE51808 |
| Data compendium | GEO | GSE50834 |
| Data compendium | GEO | GSE47199 |
| Data compendium | GEO | GSE47172 |
| Data compendium | GEO | GSE45919 |
| Data compendium | GEO | GSE45918 |
| Data compendium | GEO | GSE44228 |
| Data compendium | GEO | GSE43777 |
| Data compendium | GEO | GSE42830 |
| Data compendium | GEO | GSE42826 |
| Data compendium | GEO | GSE42825 |
| Data compendium | GEO | GSE42026 |
| Data compendium | GEO | GSE41233 |
| Data compendium | GEO | GSE41055 |
| Data compendium | GEO | GSE40586 |
| Data compendium | GEO | GSE40396 |
| Data compendium | GEO | GSE40223 |
| Data compendium | GEO | GSE40184 |
| Data compendium | GEO | GSE40012 |
| Data compendium | GEO | GSE39940 |
| Data compendium | GEO | GSE38900 |
| Data compendium | GEO | GSE38246 |
| Data compendium | GEO | GSE37250 |
| Data compendium | GEO | GSE36539 |
| Data compendium | GEO | GSE36238 |
| Data compendium | GEO | GSE35860 |
| Data compendium | GEO | GSE35860 |
| Data compendium | GEO | GSE35859 |
| Data compendium | GEO | GSE35859 |
| Data compendium | GEO | GSE34404 |
| Data compendium | GEO | GSE34404 |
| Data compendium | GEO | GSE34205 |
| Data compendium | GEO | GSE31348 |
| Data compendium | GEO | GSE30385 |
| Data compendium | GEO | GSE30310 |
| Data compendium | GEO | GSE30119 |
| Data compendium | GEO | GSE29536 |
| Data compendium | GEO | GSE29429 |
| Data compendium | GEO | GSE29385 |
| Data compendium | GEO | GSE29366 |
| Data compendium | GEO | GSE29333 |
| Data compendium | GEO | GSE29161 |
| Data compendium | GEO | GSE28988 |
| Data compendium | GEO | GSE28750 |
| Data compendium | GEO | GSE28623 |
| Data compendium | GEO | GSE28405 |
| Data compendium | GEO | GSE28177 |
| Data compendium | GEO | GSE27990 |
| Data compendium | GEO | GSE25504 |
| Data compendium | GEO | GSE25001 |
| Data compendium | GEO | GSE23140 |
| Data compendium | GEO | GSE22098 |
| Data compendium | GEO | GSE21802 |
| Data compendium | GEO | GSE20346 |
| Data compendium | GEO | GSE19444 |
| Data compendium | GEO | GSE19442 |
| Data compendium | GEO | GSE19439 |
| Data compendium | GEO | GSE18897 |
| Data compendium | GEO | GSE18090 |
| Data compendium | GEO | GSE16129 |
| Data compendium | GEO | GSE13015 |
| Data compendium | GEO | GSE11908 |
| Data compendium | GEO | GSE11755 |
| Data compendium | GEO | GSE9960 |
| Data compendium | GEO | GSE7000 |
| Data compendium | GEO | GSE7000 |
| Data compendium | GEO | GSE6377 |
| Data compendium | GEO | GSE6269 |
| Data compendium | GEO | GSE5808 |
| Data compendium | GEO | GSE5418 |
| Data compendium | GEO | GSE5418 |
| Data compendium | GEO | GSE4124 |
| Data compendium | GEO | GSE2729 |
| Data compendium | GEO | GSE2171 |
| Data compendium | GEO | GSE1739 |
| Data compendium | GEO | GSE1124 |
| Data compendium | GEO | GSE1124 |
| Software and algorithms | ||
| Original source code | This paper | https://doi.org/10.5281/zenodo.7145903 |
| tidyverse v1.3.1 | Wickham et al.40 | https://www.tidyverse.org/ |
| limma v3.42.2 | Ritchie et al.41 | https://bioconductor.org/packages/release/bioc/html/limma.html |
| affy v1.64.0 | Bolstad et al.42 | https://bioconductor.org/packages/release/bioc/html/affy.html |
| GEOquery v2.54.1 | Davis and Meltzer43 | https://bioconductor.org/packages/release/bioc/html/GEOquery.html |
| AnnotationDbi v1.52.0 | Pagès et al.44 | https://bioconductor.org/packages/release/bioc/html/AnnotationDbi.html |
| ArrayQualityMetrics v3.42.0 | Kauffmann et al.30 | https://www.bioconductor.org/packages/release/bioc/html/arrayQualityMetrics.html |
| MetaIntegrator v2.3.1 | Haynes et al.31 | https://cran.r-project.org/web/packages/MetaIntegrator/index.html |
| enrichR v3.0 | Kuleshov et al.45 | https://cran.r-project.org/web/packages/enrichR/index.html |
Resource availability
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Steven H. Kleinstein (steven.kleinstein@yale.edu).
Materials availability
This study did not generate new unique reagents.
Method details
Curation of infection signatures
We performed NCBI PubMed searches to identify published signatures of infection. We searched for: ‘viral transcriptional signature’, ‘bacterial transcriptional signature’, ‘infection transcriptional signature’, and ‘influenza transcriptional signature’’. Our inclusion criteria for signatures were that they (1) contain gene lists that describe in vivo responses to general viral or general bacterial infections in humans; (2) were derived from analyses of PBMCs/whole blood. We performed a separate search for influenza virus infection signatures. We screened the first 200 hits for each search to create a seed pool of papers. We then screened the references of these papers, as well as the ‘cited by’ publication results from Google Scholar, for additional signatures that met our inclusion criteria. Signatures published as differentially expressed genes were curated as sets of positive genes (up-regulated in the intended condition) and negative genes (down-regulated in the intended condition). Signatures published as classifiers with coefficients were discretized into positive and negative gene sets based on the sign of the coefficients. The identified signatures were grouped in the following four categories: generic viral (n=11), generic bacterial (n=7), viral versus bacterial (n=6), and influenza-specific (n=6). Enrichment terms were identified using EnrichR.45
Building a data compendium
Dataset search and selection
We searched NCBI GEO for public human expression datasets using an approach modeled after Sweeney et al.8 Infectious exposures were searched in August 2019 with the following keywords: ‘infection’, ‘bact∗’, ‘vir∗’, ‘fung∗’, ‘fever’, ‘sepsis’, ‘pneumonia’, ‘nosocomial’, ‘ICU’, and ‘SIRS’. Non-infectious exposures were searched in January 2020 with keywords ‘age’ and ‘(obesity | BMI)’. For both searches, filters were set to limit results to ‘Homo sapiens’ and ‘high throughput expression profiling by microarray’. For over 8,000 resulting dataset accessions, we screened associated abstracts and included studies that profiled in vivo infections and non-infectious conditions in PBMCs or whole blood. For infectious exposures we included studies that contained at least two conditions and at least 5 samples per condition. These two conditions could be, for example, bacterial and healthy, bacterial and viral, bacterial and non-infectious, bacterial and convalescent, or other permutations of these labels. Studies that compared two pathogens of the same condition type, e.g., comparing one virus against another virus without a non-viral comparison group did not meet this criterion. For non-infectious exposures we included studies that profiled the condition of interest and healthy controls. We also searched the recount2 database for RNA-seq datasets, but no datasets met our inclusion criteria.46
Dataset processing
Datasets from studies that met our inclusion criteria were passed through a standardized pre-processing pipeline designed to handle the most common Illumina (GPL10558, GPL6102, GPL6883, GPL6884, GPL6947) and Affymetrix (GPL11532, GPL6244, GPL13158, GPL13667, GPL201, GPL5175, GPL96, GPL97, GPL570, GPL571) platforms. While each accession generally contained a single dataset, a small number of accessions contained multiple independent cohorts that were treated as separate datasets for processing and analysis. Each dataset was passed through a standardized processing pipeline. Our pipeline for Illumina platforms utilized the neqc function with background correction from the limma package (v3.42.2),41 and the rma function from the affy package (v1.64.0) for Affymetrix arrays.42 Datasets from Illumina and Affymetrix platforms were quantile normalized. Datasets from other platforms, datasets that did not contain raw data, or datasets with incomplete raw data (e.g. Illumina probe intensities without p-values) were taken in their processed form from the GEO series accession using GEOquery (v2.54.1).43 Datasets were log2 transformed where appropriate and shifted to prevent negative expression values. Gene identifiers for all datasets were remapped to ENTREZIDs using AnnotationDbi (v1.52.0) and the latest platform annotation files.44 Outlier detection was performed using the ArrayQualityMetrics package (v3.42.0) with default parameters and thresholds.30 Briefly, samples were removed if identified as outliers satisfying the following 3 criteria: (1) a large sum of pairwise distances to other samples, (2) a significantly different intensity distribution compared to a pooled distribution from the remainder of the dataset, and (3) a strong trend on an MA plot comparing each sample to a pseudo-sample of dataset median expression values.
Dataset annotation
Infection types were manually annotated for each sample using metadata from GEO and methods from each associated publication. Infections were labeled ‘bacterial’, ‘viral’, ‘other non-infectious’, or ‘parasitic’ based on the exposures or pathogens within each dataset. Samples from subjects coinfected with both bacterial and viral pathogens were removed. No fungal infections were identified, despite explicitly including this in our search terms. The causative pathogen was identified for each sample where possible. For longitudinal datasets, subject IDs and time points were collected.
Sex imputation
To predict male and female labels for subjects across the compendium, we utilized the imputeSex function from the MetaIntegrator package with default genes, which clusters subjects according to the expression of several genes with sexually dimorphic expression patterns.31
Signature scoring framework
The evaluation of signature performance was based on the geometric mean scoring.31 This was defined for each sample i as:
where is the expression of gene g in sample i, Np and Nn are the number of positive and negative genes in the signature, respectively. The signature score for a sample is the difference between the geometric mean of the expression of the up-regulated genes and the geometric mean of the expression of the down-regulated genes.
For cross-sectional studies, subject scores were determined by the single sample score. For longitudinal studies, subject scores were summarized by taking the maximally discriminative score per subject. The most typical longitudinal design included profiling of multiple time points for the infected group and a single reference time point for the control group. In this case, the subject score for an infected subject is determined by the maximum sample score over time. For designs that included multiple sampling in the control group, the subject score for a control subject is determined by the minimum sample score over time.
Given a signature and a transcriptional contrast, the performance metric is defined as the resulting area under the ROC curve (AUROC). To calculate this metric, subject scores for this contrast were calculated and ranked. The resulting ranking, paired with the binary labels annotating the subjects (e.g. virus-infected or healthy), were then used to compute the study AUROC. AUROCs were computed only for datasets containing 50% or more of both positive and negative signature genes.
Evaluation of robustness and cross-reactivity
Robustness was evaluated using conditions that match the signature contrast: e.g., evaluating viral signatures in viral datasets. Signatures that generated median AUROCs greater than 0.7 in independent datasets profiling these intended pathogens were considered robust. This threshold roughly corresponds to the AUROC = 0.68 using the critical value for a one-sided Mann-Whitney U test where α = 0.05, assuming both group sample sizes were equal to 15.47 Median sample sizes in our compendium were 75.5 and 63 for datasets profiling viral and bacterial infections respectively, and so these conditions reflect a lower bound above which performance is considered robust.
Cross-reactivity was evaluated using unintended conditions that do not match the signature contrast: e.g., evaluating viral signatures in bacterial datasets. Viral and bacterial signatures that generated median AUROCs greater than 0.6 for profiling unintended conditions were considered cross-reactive. This cross-reactivity threshold was selected as a compromise between (1) absolute lack of signal and (2) an overly stringent cutoff. While a perfectly non-cross-reactive signature would generate an AUROC less than or equal to 0.5, human cohorts can be highly variable and an AUROC slightly above 0.5 does not necessarily indicate biologically meaningful differences between cases and controls. Conversely, an AUROC threshold of 0.7, as used for determining robustness, would reflect an overly stringent condition for determining whether a signature generates signal for an unintended condition. V/B signatures were considered cross-reactive if they generated a median AUROC greater than 0.6 or less than 0.4. This latter condition reflects that the designation of positive and negative genes in V/B signatures is arbitrary (i.e., these signatures could have been recorded with a bacterial vs. viral contrast), and therefore prediction in either direction is relevant to cross-reactivity.
Comparison of logistic regression scoring and geometric mean scoring
We compared the performance of signatures using geometric mean scoring with logistic regression scoring. Logistic regression scoring was only applied to datasets that met specific criteria: (1) cross-sectional study design, (2) measurements for ≥50% of positive and ≥50% of negative signature genes, (3) a greater number of samples than the number of model features, and (4) ≥15 cases and ≥15 controls. Signature V8 was omitted from evaluation because this signature contained many more genes than samples in all datasets.
Logistic regression models were trained using leave-one-out cross-validation with the caret package (v6.0).48 Subject scores were defined as the held-out sample prediction probability. As with geometric mean scoring, these scores were paired with the binary subject labels (e.g. infected or control) to compute the study AUROC. The geometric mean and logistic regression AUROCs were compared using Pearson correlation.
Clinical definitions of aging and obesity
For datasets profiling aging, healthy subjects over the age of 64 were considered aged. Young controls were healthy individuals under the age of 36. Obesity definitions were taken from the publication associated with each obesity dataset, often but not always corresponding to a BMI greater than or equal to 30.
Deriving candidate influenza signature genes
To derive a set of candidate signature genes that distinguished influenza infection from healthy control samples, we identified all datasets in our compendium containing (1) individuals that could be identified as exclusively influenza infected (removing subjects with co-infections) and (2) profiles from healthy control subjects. Datasets were split 70/30 for discovery and validation purposes, and metadata was used to ensure balanced representation of different platform manufacturers, age groups, tissue types, and sample sizes (Table S5). For each longitudinal dataset, a single acute time point was selected for analysis. This was the time point closest to hospital admission or median time of peak symptoms for outpatient cohorts. We adapted the meta-analysis procedure described in Sweeney et al.8 and Haynes et al.31 A leave-one-dataset-out round-robin meta-analysis was performed. Genes with an absolute effect size cutoff ≥ 1.75 and an FDR cutoff ≤ 0.01 in all rounds were selected. To account for differences between PBMCs and whole blood, separate meta-analyses were then performed for PBMC and whole blood datasets in the training set. Our final selection filtered the first pool of 148 genes to 124 candidate influenza signature genes that showed an absolute effect size ≥ 1.25 in both PBMC and whole blood analyses.
To derive a signature that distinguishes influenza infection from non-influenza viral infections (such as those caused by hRV and RSV), we identified all datasets in our compendium profiling subjects infected exclusively with influenza virus as well as subjects infected with non-influenza viruses (Table S5). A round-robin meta-analysis was performed as described above with an absolute effect size cutoff ≥ 0.80 and an FDR cutoff ≤ 0.01. The effect size cutoff was adjusted to generate a pool of candidate genes of similar size to the previous analysis. All but one training datasets profiled whole blood, so candidate genes were not further filtered. 179 candidate influenza signature genes were identified. Predictive performance of these genes for discriminating influenza infection from non-influenza infection was validated in held-out datasets (Table S5, AUROCs > 0.87).
Sampling synthetic influenza signatures
We generated 100,000 synthetic signatures from the influenza vs. healthy candidate gene pool and an additional 100,000 synthetic signatures from the influenza vs. non-influenza virus candidate gene pool, using a common approach. To generate each synthetic signature, a signature size was randomly sampled from a discrete uniform distribution ranging from a minimum of 3 and a maximum corresponding to the pool size minus 3. This range was selected to reduce the number of identical synthetic signatures. A synthetic signature of the selected size was then randomly sampled from the corresponding pool of candidate genes.
Evaluating synthetic signatures
Synthetic signatures were evaluated for robustness in validation datasets profiling influenza infection and healthy controls, as well as for cross-reactivity in datasets profiling non-influenza infection and healthy controls (Table S5). For each synthetic signature, we computed an AUROC in each validation dataset. While we reported median AUROCs in other analyses, here we computed a weighted average AUROC (<AUROC>). This was done for consistency with the validation procedure of Sweeney et al.8 the study that proposed the meta-analysis approach we used to derive the initial gene pool. Weights were determined by dataset sample sizes for robustness and cross-reactivity computation.
Defining the augmented Pareto front set
A local polynomial function was fitted to determine the relationship between cross-reactivity and robustness for the set of Pareto front signatures. Residuals from this fitted model were calculated for all synthetic signatures. Signatures were filtered to those with robustness greater than 0.7 and binned into 5 groups with equal robustness bin widths. The signatures corresponding to the 20 smallest residuals per bin were identified. This set of 100 signatures defines the augmented Pareto front, which contains the Pareto front set as well as additional points from its neighborhood.
Quantification and statistical analysis
Analyses were conducted using R. Statistical tests and related details are listed in figure captions.
Acknowledgments
We would like to thank Prithvi Parthasarathy for his help generating and checking labels during the curation of our data compendium. This work was supported by Defense Advanced Research Projects Agency contract N6600119C4022.
Author contributions
Conceptualization, S.C.S., E.Z., and S.H.K.; data curation, D.G.C.; software, D.G.C. and A.C.; formal analysis, D.G.C., A.C., and A.T.; methodology, D.G.C., A.C., S.C.S., E.Z., and S.H.K.; writing – original draft, D.G.C., A.C., S.C.S., E.Z., and S.H.K.; writing – review & editing, all authors; visualization, D.G.C., A.C., S.C.S., E.Z., and S.H.K.
Declaration of interests
Icahn School of Medicine at Mount Sinai has submitted a provisional patent related to this work. A.C., D.G.C., S.C.S., S.H.K., and E.Z. are inventors of the technology filed through ISMMS related to this manuscript. S.H.K. receives consulting fees from Peraton.
Published: December 21, 2022
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.cels.2022.11.007.
Supplemental information
Data and code availability
Transcriptional datasets were curated from GEO and are publicly available at the accession numbers provided in the key resources table.
All original code has been deposited in Zenodo and is publicly available as of the date of publication. DOIs are listed in the key resources table.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
References
- 1.Ferrer R., Martin-Loeches I., Phillips G., Osborn T.M., Townsend S., Dellinger R.P., Artigas A., Schorr C., Levy M.M. Empiric antibiotic treatment reduces mortality in severe sepsis and septic shock from the first hour: results from a guideline-based performance improvement program. Crit. Care Med. 2014;42:1749–1755. doi: 10.1097/CCM.0000000000000330. [DOI] [PubMed] [Google Scholar]
- 2.CDC . Centers for Disease Control and Prevention; 2020. Antibiotic Resistance is a National Priority.https://www.cdc.gov/drugresistance/us-activities.html [Google Scholar]
- 3.Killingley B., Mann A.J., Kalinova M., Boyers A., Goonawardane N., Zhou J., Lindsell K., Hare S.S., Brown J., Frise R., et al. Safety, tolerability and viral kinetics during SARS-CoV-2 human challenge in young adults. Nat. Med. 2022;28:1031–1041. doi: 10.1038/s41591-022-01780-9. [DOI] [PubMed] [Google Scholar]
- 4.Kucirka L.M., Lauer S.A., Laeyendecker O., Boon D., Lessler J. Variation in false-negative rate of reverse transcriptase polymerase chain reaction-based SARS-CoV-2 tests by time since exposure. Ann. Intern. Med. 2020;173:262–267. doi: 10.7326/M20-1495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Self W.H., Balk R.A., Grijalva C.G., Williams D.J., Zhu Y., Anderson E.J., Waterer G.W., Courtney D.M., Bramley A.M., Trabue C., et al. Procalcitonin as a marker of etiology in adults hospitalized with community-acquired pneumonia. Clin. Infect. Dis. 2017;65:183–190. doi: 10.1093/cid/cix317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ramilo O., Allman W., Chung W., Mejias A., Ardura M., Glaser C., Wittkowski K.M., Piqueras B., Banchereau J., Palucka A.K., et al. Gene expression patterns in blood leukocytes discriminate patients with acute infections. Blood. 2007;109:2066–2077. doi: 10.1182/blood-2006-02-002477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Suarez N.M., Bunsow E., Falsey A.R., Walsh E.E., Mejias A., Ramilo O. Superiority of transcriptional profiling over procalcitonin for distinguishing bacterial from viral lower respiratory tract infections in hospitalized adults. J. Infect. Dis. 2015;212:213–222. doi: 10.1093/infdis/jiv047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sweeney T.E., Wong H.R., Khatri P. Robust classification of bacterial and viral infections via integrated host gene expression diagnostics. Sci. Transl. Med. 2016;8:346ra91. doi: 10.1126/scitranslmed.aaf7165. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Tsalik E.L., Henao R., Montgomery J.L., Nawrocki J.W., Aydin M., Lydon E.C., Ko E.R., Petzold E., Nicholson B.P., Cairns C.B., et al. Discriminating bacterial and viral infection using a rapid Host Gene Expression Test. Crit. Care Med. 2021;49:1651–1663. doi: 10.1097/CCM.0000000000005085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Warsinske H., Vashisht R., Khatri P. Host-response-based gene signatures for tuberculosis diagnosis: a systematic comparison of 16 signatures. PLoS Med. 2019;16:e1002786. doi: 10.1371/journal.pmed.1002786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Andres-Terre M., McGuire H.M., Pouliot Y., Bongen E., Sweeney T.E., Tato C.M., Khatri P. Integrated, multi-cohort analysis identifies conserved transcriptional signatures across multiple respiratory viruses. Immunity. 2015;43:1199–1211. doi: 10.1016/j.immuni.2015.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Davenport E.E., Antrobus R.D., Lillie P.J., Gilbert S., Knight J.C. Transcriptomic profiling facilitates classification of response to influenza challenge. J. Mol. Med. (Berl.) 2015;93:105–114. doi: 10.1007/s00109-014-1212-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Parnell G.P., McLean A.S., Booth D.R., Armstrong N.J., Nalos M., Huang S.J., Manak J., Tang W., Tam O.-Y., Chan S., et al. A distinct influenza infection signature in the blood transcriptome of patients with severe community-acquired pneumonia. Crit. Care. 2012;16:R157. doi: 10.1186/cc11477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Tang B.M., Shojaei M., Parnell G.P., Huang S., Nalos M., Teoh S., O’Connor K., Schibeci S., Phu A.L., Kumar A., et al. A novel immune biomarker IFI27 discriminates between influenza and bacteria in patients with suspected respiratory infection. Eur. Respir. J. 2017;49:1602098. doi: 10.1183/13993003.02098-2016. [DOI] [PubMed] [Google Scholar]
- 15.Zaas A.K., Chen M., Varkey J., Veldman T., Hero A.O., Lucas J., Huang Y., Turner R., Gilbert A., Lambkin-Williams R., et al. Gene expression signatures diagnose influenza and other symptomatic respiratory viral infections in humans. Cell Host Microbe. 2009;6:207–217. doi: 10.1016/j.chom.2009.07.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Huang Y., Zaas A.K., Rao A., Dobigeon N., Woolf P.J., Veldman T., Øien N.C., McClain M.T., Varkey J.B., Nicholson B., et al. Temporal dynamics of host molecular responses differentiate symptomatic and asymptomatic influenza A infection. PLOS Genet. 2011;7:e1002234. doi: 10.1371/journal.pgen.1002234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.McNab F., Mayer-Barber K., Sher A., Wack A., O’Garra A. Type I interferons in infectious disease. Nat. Rev. Immunol. 2015;15:87–103. doi: 10.1038/nri3787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Bodkin N., Ross M., McClain M.T., Ko E.R., Woods C.W., Ginsburg G.S., Henao R., Tsalik E.L. Systematic comparison of published host gene expression signatures for bacterial/viral discrimination. Genome Med. 2022;14:18. doi: 10.1186/s13073-022-01025-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Tsalik E.L., Henao R., Nichols M., Burke T., Ko E.R., McClain M.T., Hudson L.L., Mazur A., Freeman D.H., Veldman T., et al. Host gene expression classifiers diagnose acute respiratory illness etiology. Sci. Transl. Med. 2016;8:322ra11. doi: 10.1126/scitranslmed.aad6873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Herberg J.A., Kaforou M., Wright V.J., Shailes H., Eleftherohorinou H., Hoggart C.J., Cebey-López M., Carter M.J., Janes V.A., Gormley S., et al. Diagnostic test accuracy of a 2-transcript Host RNA Signature for Discriminating Bacterial vs Viral Infection in Febrile Children. JAMA. 2016;316:835–845. doi: 10.1001/jama.2016.11236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Smith S.B., Dampier W., Tozeren A., Brown J.R., Magid-Slav M. Identification of common biological pathways and drug targets across multiple respiratory viruses based on human host gene expression analysis. PLoS One. 2012;7:e33174. doi: 10.1371/journal.pone.0033174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Smith S.B., Magid-Slav M., Brown J.R. Host response to respiratory bacterial pathogens as identified by integrated analysis of human gene expression data. PLoS One. 2013;8:e75607. doi: 10.1371/journal.pone.0075607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Statnikov A., McVoy L., Lytkin N., Aliferis C.F. Improving development of the molecular signature for diagnosis of acute respiratory viral infections. Cell Host Microbe. 2010;7:100–101. doi: 10.1016/j.chom.2010.01.003. author reply 102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hu X., Yu J., Crosby S.D., Storch G.A. Gene expression profiles in febrile children with defined viral and bacterial infection. Proc. Natl. Acad. Sci. USA. 2013;110:12792–12797. doi: 10.1073/pnas.1302968110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Bhattacharya S., Rosenberg A.F., Peterson D.R., Grzesik K., Baran A.M., Ashton J.M., Gill S.R., Corbett A.M., Holden-Wiltse J., Topham D.J., et al. Transcriptomic biomarkers to discriminate bacterial from nonbacterial infection in adults hospitalized with respiratory illness. Sci. Rep. 2017;7:6548. doi: 10.1038/s41598-017-06738-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhu J., Zhang Y., Ghosh A., Cuevas R.A., Forero A., Dhar J., Ibsen M.S., Schmid-Burgk J.L., Schmidt T., Ganapathiraju M.K., et al. Antiviral activity of human OASL protein is mediated by enhancing signaling of the RIG-I RNA sensor. Immunity. 2014;40:936–948. doi: 10.1016/j.immuni.2014.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Barrett T., Wilhite S.E., Ledoux P., Evangelista C., Kim I.F., Tomashevsky M., Marshall K.A., Phillippy K.H., Sherman P.M., Holko M., et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 2013;41:D991–D995. doi: 10.1093/nar/gks1193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Frasca D., Blomberg B.B. Adipose tissue inflammation induces B cell inflammation and decreases B cell function in aging. Front. Immunol. 2017;8:1003. doi: 10.3389/fimmu.2017.01003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Pereira B.I., Akbar A.N. Convergence of innate and adaptive immunity during human aging. Front. Immunol. 2016;7:445. doi: 10.3389/fimmu.2016.00445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kauffmann A., Gentleman R., Huber W. arrayQualityMetrics--a bioconductor package for quality assessment of microarray data. Bioinformatics. 2009;25:415–416. doi: 10.1093/bioinformatics/btn647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Haynes W.A., Vallania F., Liu C., Bongen E., Tomczak A., Andres-Terrè M., Lofgren S., Tam A., Deisseroth C.A., Li M.D., et al. Empowering multi-cohort gene expression analysis to increase reproducibility. Pac. Symp. Biocomput. 2017;22:144–153. doi: 10.1142/9789813207813_0015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sweeney T.E., Shidham A., Wong H.R., Khatri P. A comprehensive time-course-based multicohort analysis of sepsis and sterile inflammation reveals a robust diagnostic gene set. Sci. Transl. Med. 2015;7:287ra71. doi: 10.1126/scitranslmed.aaa5993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sampson D.L., Fox B.A., Yager T.D., Bhide S., Cermelli S., McHugh L.C., Seldon T.A., Brandon R.A., Sullivan E., Zimmerman J.J., et al. A four-biomarker blood signature discriminates systemic inflammation due to viral infection versus other etiologies. Sci. Rep. 2017;7:2914. doi: 10.1038/s41598-017-02325-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Liu T.-Y., Burke T., Park L.P., Woods C.W., Zaas A.K., Ginsburg G.S., Hero A.O. An individualized predictor of health and disease using paired reference and target samples. BMC Bioinformatics. 2016;17:47. doi: 10.1186/s12859-016-0889-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Iuliano A.D., Roguski K.M., Chang H.H., Muscatello D.J., Palekar R., Tempia S., Cohen C., Gran J.M., Schanzer D., Cowling B.J., et al. Estimates of global seasonal influenza-associated respiratory mortality: a modelling study. Lancet. 2018;391:1285–1300. doi: 10.1016/S0140-6736(17)33293-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Emmerich M.T.M., Deutz A.H. A tutorial on multiobjective optimization: fundamentals and evolutionary methods. Nat. Comput. 2018;17:585–609. doi: 10.1007/s11047-018-9685-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Berry M.P.R., Graham C.M., McNab F.W., Xu Z., Bloch S.A.A., Oni T., Wilkinson K.A., Banchereau R., Skinner J., Wilkinson R.J., et al. An interferon-inducible neutrophil-driven blood transcriptional signature in human tuberculosis. Nature. 2010;466:973–977. doi: 10.1038/nature09247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Holcomb Z.E., Tsalik E.L., Woods C.W., McClain M.T. Host-Based Peripheral Blood Gene Expression Analysis for Diagnosis of Infectious Diseases. J. Clin. Microbiol. 2017;55:360–368. doi: 10.1128/JCM.01057-16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Cappuccio A., Chawla D.G., Chen X., Rubenstein A.B., Mao W., Burke T.W., Tsalik E.L., Petzold E., Henao R., McClain M.T., et al. Multi-objective optimization identifies a specific and interpretable COVID-19 host response signature. Cell Syst. 2022 doi: 10.1016/j.cels.2022.11.008. [DOI] [PubMed] [Google Scholar]
- 40.Wickham H., Averick M., Bryan J., Chang W., McGowan L.D., François R., Grolemund G., Hayes A., Henry L., Hester J., et al. Welcome to the tidyverse. J. Open Source Software. 2019;4:1686. doi: 10.21105/joss.01686. [DOI] [Google Scholar]
- 41.Ritchie M.E., Phipson B., Wu D., Hu Y., Law C.W., Shi W., Smyth G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43:e47. doi: 10.1093/nar/gkv007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Bolstad B.M., Irizarry R.A., Astrand M., Speed T.P. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics. 2003;19:185–193. doi: 10.1093/bioinformatics/19.2.185. [DOI] [PubMed] [Google Scholar]
- 43.Davis S., Meltzer P.S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics. 2007;23:1846–1847. doi: 10.1093/bioinformatics/btm254. [DOI] [PubMed] [Google Scholar]
- 44.Pagès H., Carlson M., Falcon S., Li N. AnnotationDbi: manipulation of SQLite-based annotations in bioconductor. 2020. https://aur.archlinux.org/r-annotationdbi.git
- 45.Kuleshov M.V., Jones M.R., Rouillard A.D., Fernandez N.F., Duan Q., Wang Z., Koplev S., Jenkins S.L., Jagodnik K.M., Lachmann A., et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016;44:W90–W97. doi: 10.1093/nar/gkw377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Collado-Torres L., Nellore A., Kammers K., Ellis S.E., Taub M.A., Hansen K.D., Jaffe A.E., Langmead B., Leek J.T. Reproducible RNA-seq analysis using recount2. Nat. Biotechnol. 2017;35:319–321. doi: 10.1038/nbt.3838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mason S.J. Areas beneath the Relative Operating Characteristics (ROC) and Relative Operating Levels (ROL) Curves: Statistical Significance and Interpretation. Quarterly Journal of the Royal Meteorological Society128. 2002;((584):):2145–2166. doi: 10.1256/003590002320603584. [DOI] [Google Scholar]
- 48.Kuhn M. Building Predictive Models in R Using the caret Package. J. Stat. Software. 2008;28:1–26. doi: 10.18637/jss.v028.i05. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Transcriptional datasets were curated from GEO and are publicly available at the accession numbers provided in the key resources table.
All original code has been deposited in Zenodo and is publicly available as of the date of publication. DOIs are listed in the key resources table.
Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.