Abstract
The viability of spatially structured populations depends on the abundance and connectivity between subpopulations of breeding adults. Yet, for many species, both are extremely difficult to assess. The speartooth shark is a critically endangered elasmobranch inhabiting tropical rivers with only three adults ever recorded in Australia. Close-kin mark-recapture models, informed by sibling pairs among 226 juveniles, were developed to estimate adult abundance and connectivity in two Australian river systems. Sixty-eight sibling pairs were found, and adult abundance was estimated at 892 for the Adelaide River and 1128 for the Alligator Rivers. We found strong evidence for female philopatry, with most females returning to the same river to pup. Adelaide River males appear largely philopatric, whereas Alligator Rivers males are highly connected to the Adelaide River. From only 4 years of sampling, our results demonstrate that juvenile-only kin pairs can inform simultaneous estimates of abundance and connectivity in a rare and threatened species.
Juveniles kin pairs provide estimates of key population parameters for the conservation of threatened species.
INTRODUCTION
Abundance and survival rates of breeding adults and how populations interbreed are crucially important quantities for effective conservation. Yet, their estimation is often exceedingly difficult, especially for species in which adults are difficult to sample. For many sharks and rays, one of the most threatened vertebrate groups, estimating both connectivity and abundance typically requires long-term monitoring and multiple data sources. Here, we simultaneously estimate breeding adult census abundance and within-generation, sex-specific spatial reproductive connectivity solely from genetic samples collected over a few years and without ever sampling or even encountering adults. We demonstrate this in a population of a critically endangered euryhaline elasmobranch, the speartooth shark (Glyphis glyphis), from two river systems in northern Australia. Our approach uses genotyping to detect juvenile sibling pairs across spatially separated subpopulations in a close-kin mark-recapture (CKMR) model estimating the number of breeding adults, their survival rate, and sex-specific connectivity.
Declines of a local population affect metapopulation viability (1, 2); therefore, understanding both local abundance and connectivity is key to predicting a population’s fate. Isolated populations without substantial reproductive connectivity cannot reverse declining abundance through immigration and are therefore more vulnerable than connected populations (3). Traditional indirect population genetic approaches based on differences in allele frequencies between populations measure connectivity integrated over evolutionary time scales (4), but they remain uninformative regarding demographic connectivity (5), which may be problematic for short-term conservation decision-making. Recent advances in population genomics allow inference of connectivity over demographic time scales, but they require vast genomic resources hardly obtainable for most wild species (6). Population assignment can, in theory, provide direct estimates of population connectivity but tend to perform poorly in high gene flow situations (7), which is precisely when they would be the most needed. Similarly, at generational time scales in long-lived species, mark-recapture methods using genetic of physical tags can be problematic if animals are hard to capture in sufficient numbers, do not retain marks, or are rarely recaptured.
The spatial distribution of close relatives within a metapopulation can provide direct estimates of population connectivity over generational time scales (8–10). However, for many species, adult abundance is required to quantify connectivity and must be estimated independently (11). Because close-kin relations must exist on generational, rather than evolutionary, time scales, these data provide a more direct and conservation-relevant understanding of spatial population dynamics (12). CKMR (13) has been applied to assess adult abundance from parent-offspring pairs (14, 15) and half-sibling pairs (HSPs) (16) in single, well-mixed populations. However, as we describe, CKMR can be extended to estimate both abundance and connectivity of breeding individuals within fragmented populations without having to sample them. Conceptually, we can see how the flow of information from the spatial distribution of HSPs informs whether a population is currently totally philopatric (Fig. 1A) or panmictic (Fig. 1B) or exhibits sex-linked reproductive connectivity/dispersal (Fig. 1, C and D).
Fig. 1. Concept diagram of hypothetical population connectivity scenarios.
Reproductive dispersal and spatial distribution of half-sibling pairs (HSPs) between two hypothetical rivers, denoted r1 and r2. Linked blue and orange dots represent HSPs that share and do not share the same haplotype, respectively. Scenarios (A) and (B) denote the extreme cases of complete philopatry and complete panmixia, respectively. In these cases, we expect to see no HSPs between rivers (philopatry) or uniformly likely (panmixia). In male-based dispersal (C), all HSPs between rivers are paternal and they only share the same haplotype by chance. In female-based dispersal (D), all HSPs between rivers are maternal and always share the same haplotype. In nature, sex-specific rates of dispersal would include potential intermediate scenarios between the cases shown here. The reproductive connectivity may or may not mean that the parent was physically present within a spatial location; mating may occur in a third unknown location, and then, mothers or young may physically move to the river.
Our aim was not only to determine abundance but also to simultaneously quantify the population’s sex-specific contemporary connectivity of G. glyphis, a rare and critically endangered euryhaline elasmobranch. Until the late 20th century, G. glyphis was known only from type specimens collected in the first half of the 19th century (17, 18) until its rediscovery in northern Australia and Papua New Guinea (19). The river sharks (genus Glyphis) as a whole are considered to be highly vulnerable (20). Because of their presence in only a few rivers (Fig. 2), population sizes were assumed small and G. glyphis is listed as critically endangered under Australia’s Environment Protection and Biodiversity Conservation (EPBC) Act 1999.
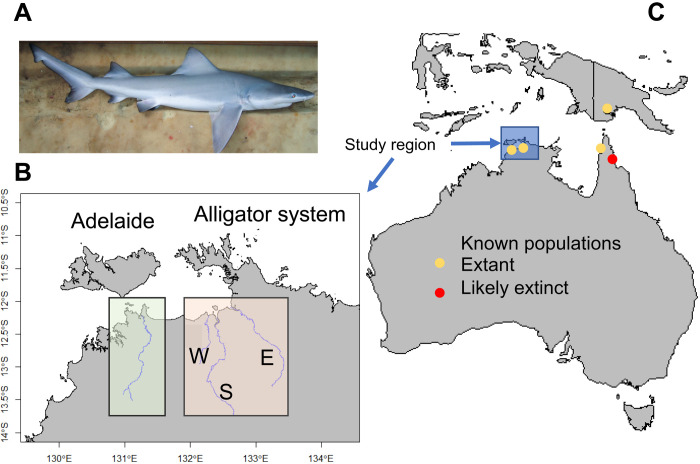
Fig. 2. G. glyphis distribution and sampling locations.
(A) Juvenile G. glyphis. (B) Study area in the Northern Territory, Australia, with the Adelaide River and the Alligator rivers, composed of the west, south, and east Alligator rivers. (C) Regional map with the approximate location of the documented populations of G. glyphis in Australia and Papua New Guinea. The population in the Bizant River, Queensland, is likely extinct as no sightings of the species have been recorded since the 1990s.
Only neonate and juvenile sharks were typically encountered in surveys (21, 22). The capture of one mature individual from the Wenlock River, Queensland, in 2015 was the first time this species had been encountered as an adult, with only two other records in Australian waters since (23). Thus, expectedly, the abundance of G. glyphis was unknown (24, 25).
Previous research found strong support for river fidelity in juveniles combined with strong female philopatry in rivers separated by >150 km. For rivers <150 km apart, males sired offspring between rivers (12), although we note that spawning may well not occur within the rivers but in some currently unknown location offshore from both river mouths. In addition, the other distant population to the east in the Wenlock River, Queensland, was not included as it has been demonstrated to be isolated from these Northern Territory populations (12). These are important results, which were used to guide the construction of the spatial and sex-specific model used here for estimating abundance and connectivity.
In this study, single-nucleotide polymorphism (SNP) data from sampled juveniles were used to infer whether a pair of individuals were either an unrelated pair (UP), a full-sibling pair (FSP) sharing both mother and father, or an HSP that shares a single parent. In addition, using full mitogenome haplotypes, we determined whether HSPs were likely maternal (mHSP) or paternal (pHSP). As we show, these data were combined via a population model to determine abundance of breeding adults by river and the degree of reproductive connectivity by sex. CKMR also allows us to estimate important population parameters that are typically very difficult, or even impossible, to observe. Specifically, the population model estimates the following (see Materials and Methods and the Supplementary Materials for details): abundance N0, r by river r, adult survival rate ψ, sex ratio (in terms of proportion of females) ζ, and several important biological parameters that are estimated by CKMR: ν, the potential for overrepresentation of within-cohort maternal kin (FSPs and mHSPs) given high heterogeneity in very early-life survival, termed the “lucky litter” effect following (13); θ, probability that a female will mate with multiple males; γ, effective number of female mates per male; ωs, r, r′, the probability that a shark of sex s originating in river r will reproduce in river r′. Again, these quantities pertaining to the breeding population are obtained without ever sampling any adult individuals. The results we present demonstrate a powerful technique for estimating adult abundance from juveniles while simultaneously establishing the connectivity of breeding populations.
RESULTS
Field sampling programs collecting tissue samples operated between 2012 and 2016 in the Adelaide River and the west, south, and east Alligator Rivers (collectively referred to as the “Alligator rivers”), yielding 221 samples from individuals that passed quality control checks. This resulted in 24,310 comparisons between individuals to establish whether two given individuals were UP, FSP, or HSP. Each comparison is scored using a pseudo-log odds (PLOD) value that describes the likelihood of the pair being one of the kin types (UP, FSP, or HSP) (see text S3 for full details).
Kin-pair data
The PLOD scores for the FSP comparisons (based on 226 sharks and 1400 loci) indicated the clear presence of both types of related kin pairs from both river systems. There were 21 FSPs found: 12 from the Adelaide River and 9 from the Alligator rivers (Table 1).
Table 1. Observed FSPs and HSPs across the two river systems.
Hit rates (number of kin pair per comparison is also given).
| Kin-type | Adelaide | Alligators | Cross | Total |
|---|---|---|---|---|
| FSP | 12 | 9 | 0 | 21 |
| (FSP hit rate) | (0.0012) | (0.0) | (0.0028) | |
| HSP | 28 | 7 | 6 | 41 |
| (HSP hit rate) | (0.0028) | (0.0022) | (0.0005) |
None of the FSPs were found among juveniles in different rivers, and none of the FSPs were found to be across juvenile cohorts (i.e., in all cases, both had the same birth year). There were 41 HSPs—28 came from within the Adelaide River, 7 came from within the Alligator rivers, and 6 were across-river (Fig. 3). Of these 41 HSPs, 6 were within cohort HSP, 5 from the Adelaide, and 1 from the Alligator.
Fig. 3. Spatial linkages of HSPs between river systems.
(A) The vertical axis is the cohort of the sampled juvenile, indicating that the parent bred in that river system in that year. Orange indicates that the two half-sibling juveniles share the same haplotype, and blue lines depict an HSP that do not, and hence whether the movement of reproductive allocation was likely via the maternal or paternal parent. The data (see Table 1) on cross-river HSPs (N = 6) and whether they did (N = 4) or did not share (N = 2) haplotypes indicate that males and possibly some females move from the Alligator rivers to the Adelaide River with only one male reproductive transition in the opposite direction. While the sample size creates uncertainty, the close-kin model (see Table 2 and the Supplementary Materials) estimates of probability of reproductive connectivity accounting for the observed mtDNA haplotype distribution indicate male linked connectivity being far more likely than female. (B) Numbers of samples collected in each river system through time. Note that these are assigned to an earlier cohort year based on estimated age given size.
The within-river “hit rates” (HSPs per comparison) for the Adelaide River was 0.0027, and for the Alligator rivers, it was 0.0022 (higher hit rate is suggestive of a smaller population). The across-river HSP was 0.0005, far smaller than the within-river hit rate, which indicates more limited connectivity.
Mean time between birth years of HSPs were both just over 3 years for within-river comparisons and 2.9 for the across-river HSPs, suggestive of a high survival rate of breeding adults. Of the six within-cohort HSPs, five came from the Adelaide River, and only one came from the Alligator rivers (close to the 4:1 ratio seen in the overall comparison and when looking only at cross-cohort HSPs). None of the within-cohort HSPs were across rivers, supporting our model structure where breeding can occur across-rivers but not within a single year. This suggests that adults breed in one river or other, but not both, in a given year. There were clear differences in the hit rate of FSPs (FSPs per comparison) for the two rivers: For the Adelaide River, it was 0.01, but for the Alligator rivers, it was 0.016—over 50% higher. However, within-rivers, the within and cross-cohort HSP hit rates were very similar between rivers. This indicated that the Alligator rivers’ litter effect is likely larger than that of the Adelaide River (see below), which could indicate river-specific differences in the early-life environment for neonates.
Mitochondrial DNA
A total of 11 mitochondrial DNA (mtDNA) haplotypes were found, with one particular haplotype being very dominant in the population (a frequency of occurrence of 78%). Very low haplotype diversity has been documented in G. glyphis (26), which intrinsically limits the amount of detailed information that can be extracted from these data with respect to both within-river sex ratio and across-river reproductive transfer. Nevertheless, there is clear qualitative information in the data that we can summarize succinctly: All of the FSPs shared a haplotype (which they should since mtDNA is maternally inherited); for the within-cohort case (c1 = c2), all but one of the six HSPs shared a haplotype (the most common one), and none of the within-cohort HSPs were detected in different rivers (as above, we consider this to be an impossible event); for the 35 cross-cohort HSPs, 31 shared a haplotype, while 4 did not; of those 31 that shared a haplotype, 28 shared the most common haplotype. The remaining three HSPs shared a rare haplotype [p(h) = 0.04] and were almost certainly mHSPs.
For the cross-river HSPs (the information source on the connectivity parameters ωs, r, r′), 6 of the 35 cross-cohort HSPs (17%) were cross-river, and of those 6 HSPs, 4 shared a haplotype (the dominant one) and 2 did not. The sample sizes are small, but this nevertheless constitutes clear information on reproductive exchange between adults with different rivers of origin and, furthermore, the 4:2 cross-river shared/nonshared haplotype ratio compared to the 2:29 within-river ratio provides clear indication that this transfer is likely to be more male dominated rather than female.
Abundance and connectivity estimates
Total adult abundance (Adelaide and Alligator rivers) was estimated at N = 2020 [coefficient of variation (CV), 0.28; Table 2A]. Abundance of the Adelaide River of origin was N = 892 (CV, 0.21) and that of the Alligator rivers of origin was N = 1128 (CV, 0.47).
Table 2. Abundance and connectivity parameters for G. glyphis.
(A) Estimates of population abundance and kinship parameters. (B) Estimated connectivity parameters. The connectivity values give the probability an animal with origin river r′ reproduces in r. For example, the model estimates that only 1% of male sharks originating in the Adelaide River reproduce in the Alligator rivers.
| (A) Estimated parameters | Joint | Adelaide | Alligators |
|---|---|---|---|
| Abundance N0 | 2020 | 892 | 1128 |
| CV (N0) | (0.28) | (0.21) | (0.47) |
| Adult annual survival, ϕ | 0.88 | ||
| Litter effect, ν | – | 3.42 | 7.28 |
| Multiple paternity θ | 0.00043 | – | – |
| Multiple females/male γ | 1.83 | – | – |
| Fixed population parameters | |||
| Sex ratio (% female), ζ | 0.4 | ||
| Population growth rate, λ | 0 | ||
| (B) Estimated connectivity parameters | River | Adelaide | Alligators |
| Sex | |||
| Female | Adelaide | 0.99 | 0.01 |
| Alligators | 0.88 | 0.12 | |
| Male | Adelaide | 0.99 | 0.01 |
| Alligators | 0.55 | 0.45 |
Within-cohort parameters
Lucky litter effects, ν, were estimated to be 3.42 and 7.28 (Table 2A) for the Adelaide and Alligator rivers, respectively (as suspected given the exploratory data analysis of the SNP data), and suggest that there are potentially strong differences in neonate survival between the rivers. The θ parameter (probability that a female will mate with multiple males, assumed to be shared across rivers) was estimated to be very small (0.00043), and γ (effective number of female mates per male) was estimated to be 1.83. This indicates that females are unlikely to have multiple fathers within a litter (although does not totally rule it out) and that, on average, males will mate with about two females.
Connectivity and sex ratio
Strong philopatry was indicated in females in both rivers, with only 1% of Adelaide River females expected to breed in the Alligator rivers (Table 2B) and higher (12%) in the opposite direction (Alligator females breeding in Adelaide). Adelaide River–of–origin males were estimated to display a high degree of philopatry, with less than 1% siring offspring in the Alligator rivers. However, our model indicated a net “flow” of male reproductive allocation from the Alligator rivers to the Adelaide River (55%).
Sex-specific connectivity of adults should be considered alongside likely within-river sex ratio. This is informed by the ratio of cross-cohort HSPs, within river, that do or do not share haplotypes. When estimating connectivity, very few cross-river HSPs not sharing a haplotype constitutes strong evidence of significant male reproductive connectivity. However, with a single dominant haplotype, the information on within-river sex ratio will be very diffuse—even given the number of HSPs we found. The dominance of one haplotype in both populations is why connectivity and sex ratio were not jointly estimable from the current dataset.
While not directly estimable, in the Adelaide River specifically, the ratios of HSP that do or do not share a haplotype (table S1) suggest a male-biased sex ratio in the adult population reproducing in each river (eq. S2 in text S2). However, the movement rates through the ω parameters suggest that 50% of the males from the Alligator rivers are also present in the Adelaide River, but almost all the Adelaide River males breed locally. This suggests a possible male bias in the Adelaide River even for a 50:50 sex ratio in each river of origin.
DISCUSSION
This study has obtained adult census abundance estimates for the breeding population of a critically endangered euryhaline elasmobranch along with estimates of sex-specific connectivity, adult survival rates, and insights into the breeding dynamics of the population. These results were based purely from juvenile cross-cohort sibling data. Therefore, the study presents an extension to the previous applications of CKMR for single, mixed populations (14–16, 27).
The other feature of our study is that results were obtained on a long-lived species from sampling almost entirely over a 4-year sampling period. This is a crucial acceleration of our ability to rigorously assess these populations, going as it were from minimal knowledge (presence only) to abundance and connectivity. Given the wide acknowledgement that elasmobranchs are among the most threatened vertebrates (28), methods to rapidly assess these long-lived and biologically unproductive species (29) from a range of taxa in rapid time frames are urgently required. Theoretically, abundance can be obtained from a single sampling event (13), if samples adequately cover several cohorts as information on adult abundance and survival derives from the fact that the unseen adult must have survived over the period spanned by the birthdates of two half-sibling juveniles. We note that a source of uncertainty for the current study arises in assignment of birth cohort from a small sample of direct vertebral ages (30). However, the animals’ status as critically endangered means that sacrificing large numbers of animals to obtain vertebral samples is problematic (30). Addressing this uncertainty is likely to involve further development and refinement of nonlethal age estimation (31–33).
A correlate of the adult census abundance estimated here, effective number of breeders, can be obtained from within-cohort sibling data (34) but does not inform on generational connectivity. By resolving sex-specific differences in spatial allocation of reproductive output (subject to conditioning on plausible sex ratios), we can assess generational connectivity, as opposed to evolutionary mixing rates. Note that this would be impossible to obtain in this species using other methods, such as tagging, as adults are exceedingly difficult to encounter. This is important information for management of these and similar populations, which are currently prioritized using scant data and expert opinion (35). In contrast, our method generates directly estimated quantities required in conservation listing and management. In this case, we found low abundance (around ~1000 adults in each population) and a complex spatial structure requiring careful management. Continued low-cost monitoring will refine these results and reveal population trends. Understanding connectivity will be crucial to ongoing management; the loss of one documented Australian G. glyphis population in the Bizant River, Queensland (Fig. 2C) (24) shows that isolated populations of this species are apparently easily removed by anthropogenic activities, such as fishing pressure or alterations to flow regimes.
MATERIALS AND METHODS
Field sampling
Capture and sampling of G. glyphis was conducted under Northern Territory Fisheries Special Permits S17/3251, S17/3252, and S17/3364; Kakadu National Park Research Permit RK805; and Charles Darwin University Animal Ethics Committee A11041. Juvenile G. glyphis [52 to 160 cm total length (TL)] were captured with rod and line or gill net in the brackish riverine and estuarine reaches of the Adelaide and west, south, and east Alligator Rivers (collectively referred to as the Alligator rivers) (Fig. 2). Neonate sharks were easily recognized by the presence of an umbilical scar, and otherwise, animals were assigned to an estimated age class based on length and date of capture. Given the need to assign animals to a likely cohort, estimates of length at age were obtained from N = 10 G. glyphis from the Adelaide River, which were aged from vertebral sections and used to estimate a model of age at length (30). Age at length is typically inferred on the basis of data from many specimens, but given the rarity of this species, sample size was deliberately kept to an absolute minimum (30).
Genetics and kin-finding
SNP genotyping was done using the DArTseq restriction-associated DNA approach exactly as described by (12) and resulted in precisely 1400 high-quality SNPs for kin-finding. mtDNA haplotypes were defined using targeted sequencing of all the variable sites found by sequencing the entire mitogenome of 93 G. glyphis from the same region (26, 36). The SNP genotyping methodology exactly followed the process described in a previous study of G. glyphis (12). Following the methods for kin-finding used for other sharks using DArTseq (37), a two-step process was used to identify HSPs and FSPs used in the CKMR model, which is highly informative on both within-cohort and across-river breeding dynamics (see text S1). The first step is quality control on the genetic samples themselves. This requires checking that the allele frequencies of sampled animals are in approximate Hardy-Weinberg equilibrium and that other statistical properties of the allele frequencies are acceptable [see text S1 and details in (16)]. Loci and sharks that did not meet these assumptions were removed. In addition, in the case where repeat samples (i.e., the same individual) were detected, a single random shark in the pair was removed.
In the second step, allele frequencies were used to identify FSP, HSP, and UPs. UPs include all weaker than FSP and HSP types of kin, that is, separated by three or more meioses. Specifically, we used a (pseudo) likelihood of difference (PLOD) statistic (13, 16) that, for each unique pairwise comparison, posits a specific kin relationship (e.g., UP versus HSP or UP versus FSP). The FSP analysis was conducted first, and given the numbers of SNPs retained after quality control processing (N = 1400), FSPs are unambiguously identifiable from the vast majority of combinations, which are UPs. In cases with a single FSP (i.e., two fish that share both parents), one of the fish is randomly removed from the dataset, avoiding false-positive HSPs that are actually FSPs. In cases with groups of FSPs, all but one of the fish are removed from the dataset. We then performed UP versus HSP comparisons on the FSP-filtered data, again based on PLOD statistic values. For identifying HSPs, given the number of SNPs we use, there is no clear threshold of the PLOD statistic, above which we would be 100% certain that we have only HSPs, and below which they are either UPs or more distant relations (such as half-cousins). Therefore, we identified a critical level, ηcrit, of the HSP PLOD statistic, above which we would statistically expect there to be less than one non-HSP. For this critical false-positive threshold of the HSP PLOD, we calculated the distribution of the PLOD around the expected value for HSPs (13, 16). Using this distribution, we calculated the fraction of true HSPs that are expected to be above the false-positive threshold value, πη. This gives us the false-negative retention rate of true HSPs that is used later in constructing the kinship probabilities in the population model (see text S1).
Population model
Full details of the model and statistical estimation procedures are provided in the Supplementary Materials (text S2). Here, we provide an overview of the major details of the model structure and parameters. Population dynamics were given by an exponential growth model for Nt the number of breeding adults in year t, split by sex s ∈ {m, f} within river r
where λs, r is the population growth rate for sex s in river r. Note that given potential for connectivity, the abundance in river r is also a function of input across rivers r′. Hence, total abundance is
where ωs, r, r′ is the probability an animal with origin river r′ reproduces in r such that
For example, in the case of exclusive and obligatory female philopatry, ωf,r,r′ → 1 for r = r′ and ωf,r,r′ → 0 for r ≠ r′. The abundance Ns,r,t is related to the kin pair data via equations that link the probability of finding HSP to the abundance and other demographic parameters.
In the simplest nonspatial case, an mHSP in a population with a 50:50 sex ratio is given by Pr(mHSP) = (4/Nt)ϕ∣∣, where t is the birth year of the youngest member of an HSP pair (16) and ϕ is adult survival. The spatiotemporal case requires specific probabilities (see text S3.1) for the cases of within-cohort FSP, mHSP, and pHSPs and across-cohort mHSP and pHSP, which are used depending on the nature of the comparison being evaluated. The cross-cohort probabilities include ωf,r,r′ (capturing the spatial connectivity) and the river- and sex-specific Ns,r′,t .
Chance events, such as predation of juveniles, could induce heterogeneity in neonate survival rates, which may strongly influence the probability of finding HSPs. Following (16), we estimate a case that deals with lucky litter θ for females and multiple partner γ for males. In addition, the observed mtDNA data (sharing/not sharing haplotypes) depend on haplotype frequencies, sex ratio (and survival if sex-dependent), and spatial reproductive dynamics. Note that the model used here implicitly assumes that fecundity is not strongly age or size dependent. This is reasonable for many elasmobranchs, which often have relatively constant litter size after maturity. This would not be the case in other taxa, such as teleost fish, where reproductive output is strongly size dependent.
In the model, we set λs,r = 0, and survival probability ϕ was aggregated across sexes and rivers. On the basis of likelihood profiling, we used the lower approximate 95th percentile of ϕ = 0.88 as a conservative estimate of adult survival. Again, cross-river HSPs are the source of information regarding spatial allocation of reproduction. However, limited mtDNA information content (due to the limited number of haplotypes and dominance of one of them) meant that estimates of sex ratio ζ and movement rate could not be estimated jointly. We therefore assumed a male-biased sex ratio of 40:60 (female/male), as has been documented in other G. glyphis (38) populations and elasmobranch species (39, 40). The model was implemented and fitted with the R library TMB (41) by minimization of the negative log-likelihood
where θ = [N0,r, ωr,r′, ν, γ ] and ℙ(kij) are the kinship probabilities, which are described in detail in the Supplementary Materials (see text S3.1).
Acknowledgments
We thank M. Bravington for CKMR development and detailed comments on an early version of this manuscript. We are also grateful to two anonymous referees whose input led to significant improvements. P. Grewe, N. Bax, M. Douglas, and T. Saunders provided project development and direction. We thank Kakadu National Park staff, rangers, and Traditional Owners, and all volunteers who assisted in the field.
Funding: This work was undertaken for the Marine Biodiversity Hub, a collaborative partnership supported through funding from the Australian Government’s National Environmental Research Program (NERP) and National Environmental Science Program (NESP).
Author contributions: T.A.P. and R.M.H. conceived the population model for the study designed by R.D.P., P.F., and P.M.K. Fieldwork and sample collection were by P.M.K., R.D.P., G.J.J., and P.F. R.M.H. and T.A.P. implemented the population model code and conducted analysis. P.F., J.R.M., and R.M.G. carried out the molecular work. P.F. and R.M.H. analyzed genotype data. T.A.P. wrote the manuscript with input from all other authors.
Competing interests: The authors declare that they have no competing interests.
Data and materials availability: All data needed to evaluate the conclusions in the paper are present in the paper and/or the Supplementary Materials.
Supplementary Materials
This PDF file includes:
Supplementary Text
Figs. S1 to S5
Tables S1 and S2
Other Supplementary Material for this : manuscript includes the following:
Data S1
REFERENCES AND NOTES
- 1.M. Gyllenberg, I. Hanski,Habitat deterioration, habitat destruction, and metapopulation persistence in a heterogenous landscape. Theor. Popul. Biol. 52,198–215 (1997). [DOI] [PubMed] [Google Scholar]
- 2.H. D. Bowlby, A. J. F. Gibson,Evaluating whether metapopulation structure benefits endangered diadromous fishes. Can. J. Fish. Aquat. Sci. 77,388–400 (2020). [Google Scholar]
- 3.K. R. Crooks, M. Sanjayan, Connectivity Conservation (Cambridge Univ. Press, 2006), vol. 14. [Google Scholar]
- 4.J. T. Kool, A. Moilanen, E. A. Treml,Population connectivity: Recent advances and new perspectives. Landsc. Ecol. 28,165–185 (2013). [Google Scholar]
- 5.W. H. Lowe, F. W. Allendorf,What can genetics tell us about population connectivity? Mol. Ecol. 19,3038–3051 (2010). [DOI] [PubMed] [Google Scholar]
- 6.M. Leitwein, M. Duranton, Q. Rougemont, P.-A. Gagnaire, L. Bernatchez,Using haplotype information for conservation genomics. Trends Ecol. Evol. 35,245–258 (2020). [DOI] [PubMed] [Google Scholar]
- 7.P. Saenz-Agudelo, G. P. Jones, S. R. Thorrold, S. Planes,Estimating connectivity in marine populations: An empirical evaluation of assignment tests and parentage analysis under different gene flow scenarios. Mol. Ecol. 18,1765–1776 (2009). [DOI] [PubMed] [Google Scholar]
- 8.P. J. Palsbøll, M. Zachariah Peery, M. Berube,Detecting populations in the ‘ambiguous’ zone: Kinship-based estimation of population structure at low genetic divergence. Mol. Ecol. Resour. 10,797–805 (2010). [DOI] [PubMed] [Google Scholar]
- 9.S. Planes, G. P. Jones, S. R. Thorrold,Larval dispersal connects fish populations in a network of marine protected areas. Proc. Natl. Acad. Sci. U.S.A. 106,5693–5697 (2009). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.C. Schunter, M. Pascual, J. Garza, N. Raventós, E. Macpherson,Kinship analyses identify fish dispersal events on a temperate coastline. Proc. Biol. Sci. 281,20140556 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.M. Bode, J. M. Leis, L. B. Mason, D. H. Williamson, H. B. Harrison, S. Choukroun, G. P. Jones,Successful validation of a larval dispersal model using genetic parentage data. PLOS Biol. 17,3000380 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.P. Feutry, O. Berry, P. M. Kyne, R. D. Pillans, R. M. Hillary, P. M. Grewe, J. R. Marthick, G. Johnson, R. M. Gunasekera, N. J. Bax, M. Bravington,Inferring contemporary and historical genetic connectivity from juveniles. Mol. Ecol. 26,444–456 (2017). [DOI] [PubMed] [Google Scholar]
- 13.M. V. Bravington, H. J. Skaug, E. C. Anderson,Close-kin mark-recapture. Stat. Sci. 31,259–274 (2016). [Google Scholar]
- 14.M. V. Bravington, P. M. Grewe, C. R. Davies,Absolute abundance of southern bluefin tuna estimated by close-kin mark-recapture. Nat. Commun. 7,13162 (2016). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.D. E. Ruzzante, G. R. McCracken, B. Førland, J. MacMillan, D. Notte, C. Buhariwalla, J. Mills Flemming, H. Skaug,Validation of close-kin mark–recapture (CKMR) methods for estimating population abundance. Methods Ecol. Evol. 10,1445–1453 (2019). [Google Scholar]
- 16.R. Hillary, M. Bravington, T. Patterson, P. Grewe, R. Bradford, P. Feutry, R. Gunasekera, V. Peddemors, J. Werry, M. Francis, C. A. J. Duffy, B. D. Bruce,Genetic relatedness reveals total population size of white sharks in eastern Australia and New Zealand. Sci. Rep. 8,2661 (2018). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.L. J. Compagno, W. T. White, P. R. Last, Glyphis garricki sp. nov., a new species of river shark (Carcharhiniformes: Carcharhinidae) from northern Australia and Papua New Guinea, with a redescription of Glyphis glyphis (Müller & Henle, 1839), in Descriptions of New Australian Chondrichthyans (CSIRO Marine and Atmospheric Research, 2008), pp. 203–225. [Google Scholar]
- 18.C. Li, S. Corrigan, L. Yang, N. Straube, M. Harris, M. Hofreiter, W. T. White, G. J. Naylor,DNA capture reveals transoceanic gene flow in endangered river sharks. Proc. Natl. Acad. Sci. U.S.A. 112,13302–13307 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.W. T. White, S. A. Appleyard, B. Sabub, P. M. Kyne, M. Harris, R. Lis, L. Baje, T. Usu, J. J. Smart, S. Corrigan,Rediscovery of the threatened river sharks, Glyphis garricki and G. glyphis, in Papua New Guinea. PLOS ONE 10,e0140075 (2015). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.A. B. Haque, S. A. Das,New records of the critically endangered Ganges shark Glyphis gangeticus in Bangladeshi waters: Urgent monitoring needed. Endanger. Species Res. 40,65–73 (2019). [Google Scholar]
- 21.I. C. Field, B. J. Tillett, R. Charters, G. J. Johnson, R. C. Buckworth, M. G. Meekan, C. J. A. Bradshaw,Distribution, relative abundance and risks from fisheries to threatened Glyphis sharks and sawfishes in northern Australia. Endanger. Species Res. 21,171–180 (2013). [Google Scholar]
- 22.B. Lyon, R. G. Dwyer, R. D. Pillans, H. Campbell, C. E. Franklin,Distribution, seasonal movements and habitat utilisation of an endangered shark, Glyphis glyphis, from northern Australia. Mar. Ecol. Prog. Ser. 573,203–213 (2017). [Google Scholar]
- 23.R. Pillans, G. C. Fry, G. Carlin, T. Patterson,Bycatch of a critically endangered shark Glyphis glyphis in a crab pot fishery: Implications for management. Front. Mar. Sci. 9,787634 (2022). [Google Scholar]
- 24.R. D. Pillans, J. D. Stevens, P. M. Kyne, J. Salini,Observations on the distribution, biology, short-term movements and habitat requirements of river sharks Glyphis spp. in northern Australia. Endanger. Species Res. 10,321–332 (2009). [Google Scholar]
- 25.J. Stevens, R. Pillans, J. Salini, Conservation assessment of Glyphis sp A (speartooth shark), Glyphis sp C (northern river shark), Pritis microdon (freshwater sawfish) and Pritis zijsron (green sawfish) (Final report to Department of Environment and Heritage, 2005). [Google Scholar]
- 26.P. Feutry, P. M. Kyne, R. D. Pillans, X. Chen, G. J. Naylor, P. M. Grewe,Mitogenomics of the speartooth shark challenges ten years of control region sequencing. BMC Evol. Biol. 14,232 (2014). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.S. Prystupa, G. R. McCracken, R. Perry, D. E. Ruzzante,Population abundance in arctic grayling using genetics and close-kin mark-recapture. Ecol. Evol. 11,4763–4773 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.IUCN, The IUCN Red List of Threatened Species. Version 2021-2 (2021); https://iucnredlist.org.
- 29.S. A. Pardo, H. K. Kindsvater, J. D. Reynolds, N. K. Dulvy,Maximum intrinsic rate of population increase in sharks, rays, and chimaeras: The importance of survival to maturity. Can. J. Fish. Aquat. Sci. 73,1159–1163 (2016). [Google Scholar]
- 30.P. M. Kyne, J. J. Smart, G. Johnson, Extremely low sample size allows age and growth estimation in a rare and threatened shark. bioRxiv 2022.09.26.509619 [Preprint]. 28 September 2022. 10.1101/2022.09.26.509619. [DOI]
- 31.B. Mayne, W. Mustin, V. Baboolal, F. Casella, K. Ballorain, M. Barret, M. A. Vanderklift, A. D. Tucker, D. Korbie, S. Jarman, O. Berry,Age prediction of green turtles with an epigenetic clock. Mol. Ecol. Resour. 22,2275–2284 (2022). [DOI] [PubMed] [Google Scholar]
- 32.B. Mayne, D. Korbie, L. Kenchington, B. Ezzy, O. Berry, S. Jarman,A DNA methylation age predictor for zebrafish. Aging 12,24817–24835 (2020). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.B. Mayne, T. Espinoza, D. Roberts, G. L. Butler, S. Brooks, D. Korbie, S. Jarman,Nonlethal age estimation of three threatened fish species using DNA methylation: Australian lungfish, Murray cod and Mary River cod. Mol. Ecol. Resour. 21,2324–2332 (2021). [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 34.R. S. Waples, P. Feutry,Close-kin methods to estimate census size and effective population size. Fish Fish. 23,273–293 (2022). [Google Scholar]
- 35.K. Friedman, M. Braccini, M. Bjerregaard-Walsh, R. Bonfil, C. J. Bradshaw, S. Brouwer, I. Campbell, R. Coelho, E. Cortés, W. Dimmlich, M. G. Frisk, I. Kingma, S. R. McCully Phillips, C. O’Criodain, D. Parker, S. Shephard, J. Tovar-Ávila, K. Yokawa,Informing CITES Parties: Strengthening science-based decision-making when listing marine species. Fish Fish. 21,13–31 (2020). [Google Scholar]
- 36.X. Chen, M. Liu, P. M. Grewe, P. M. Kyne, P. Feutry, Complete mitochondrial genome of the critically endangered speartooth shark Glyphis glyphis (Carcharhiniformes: Carcharhinidae). Mitochondrial DNA 25, 431–432 (2014). [DOI] [PubMed] [Google Scholar]
- 37.B. Bruce, R. Bradford, M. Bravington, P. Feutry, P. Grewe, R. Gunasekera, D. Harasti, R. Hillary, T. Patterson, A national assessment of the status of white sharks (National Environmental Science Program Marine Biodiversity Hub, 2018). [Google Scholar]
- 38.T. Patterson, R. Hillary, P. Feutry, R. Gunasakera, J. Marthick, R. Pillans, Rapid estimation of cryptic adult abundance and breeding dynamics in a critically endangered elasmobranch from close-kin mark recapture. bioRxiv 2022.02.24.481858 [Preprint]. 27 February 2022. 10.1101/2022.02.24.481858. [DOI]
- 39.V. J. Wearmouth, E. J. Southall, D. Morritt, R. C. Thompson, I. C. Cuthill, J. C. Partridge, D. W. Sims,Year-round sexual harassment as a behavioral mediator of vertebrate population dynamics. Ecol. Monogr. 82,351–366 (2012). [Google Scholar]
- 40.V. J. Wearmouth, D. W. Sims,Chapter 2 sexual segregation in marine fish, reptiles, birds and mammals: Behaviour patterns, mechanisms and conservation implications. Adv. Mar. Biol. 54,107–170 (2008). [DOI] [PubMed] [Google Scholar]
- 41.K. Kristensen, A. Nielsen, C. Berg, H. Skaug, B. Bell,Template model builder TMB. J. Stat. Softw. 70,1–21 (2015). [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Text
Figs. S1 to S5
Tables S1 and S2
Data S1