Abstract
Syphilis is a chronic infection with early relapses that are hypothesized to result from the emergence of phenotypic variants of Treponema pallidum. Recent studies demonstrated that TprK, a target of protective immunity, is heterogeneous in several T. pallidum strains, but not in Nichols strain Seattle (A. Centurion-Lara, C. Godornes, C. Castro, W. C. Van Voorhis, and S. A. Lukehart, Infect. Immun. 68:824–831, 2000). Analysis of PCR-amplified tprK from Nichols strain UNC and Street strain 14 treponemes showed that TprK has seven regions of intrastrain heterogeneity resulting from amino acid substitutions, insertions, and deletions. In contrast, analysis of PCR-amplified tprJ showed little intrastrain or interstrain heterogeneity. Reverse transcriptase PCR analysis demonstrated that mRNA transcripts representing unique polymorphic TprK proteins are present during syphilitic infection. Southern hybridization confirmed that Nichols strain UNC and Street strain 14 each contain a single copy of tprK, indicating that intrastrain heterogeneity is due to the presence of multiple treponemal subpopulations which contain a variant form of tprK.
Treponema pallidum subsp. pallidum is the agent of syphilis, a sexually transmitted disease that is a risk factor for the acquisition and transmission of human immunodeficiency virus (HIV) infection (8). The noncultivable nature of T. pallidum and the lack of a system for genetic manipulation have precluded standard approaches for identification of treponemal virulence factors (7). However, recent analysis of the complete genome of the T. pallidum Nichols strain has revealed some candidate virulence factors, including a polymorphic multigene family encoding 12 T. pallidum repeat (Tpr) proteins (6). TprA through TprL have amino acid sequence similarity to Msp, a cell surface adhesin of T. denticola, a periodontitis-associated spirochete (3, 6, 11). Based on the presence or absence of a putative N-terminal signal peptide, predicted molecular mass, and percent identity, the Tpr proteins form three subgroups: subgroup 1, comprising A, B, H, K, and L; subgroup 2, comprising C, D, F, and I; and subgroup 3, comprising E, G, and J (J. M. Hardham, M. P. McLeod, J. K. Howell, E. Sodergren, G. M. Weinstock, and S. J. Norris. Abstr. 98th Gen. Meet. Am. Soc. Microbiol., abstr. D-56, 1998). Proteins in subgroups 2 and 3 contain conserved N and C termini and central domains that vary in sequence and length. Proteins in subgroup 1 contain only small areas of conserved sequences in their N and C termini and have limited homology to other Tpr proteins (3, 6, 11).
Since the products of multigene families are often involved in bacterial pathogenesis (5), the Tpr proteins are being investigated as targets for vaccine development. Centurion-Lara et al. (3) reported that antibodies to a recombinant-expressed TprK fragment (amino acids 37 to 351 based on Nichols strain GenBank) were opsonic for T. pallidum Nichols strain Seattle. Additionally, immunization of rabbits with this fragment evoked significant, but not complete, protection against intradermal challenge with the homologous treponemal strain. Because the TprK protein is potentially important in protective immunity, we hypothesized that it might be subject to phenotypic (antigenic and/or phase) variation. Centurion-Lara et al. (4) recently reported intrastrain amino acid sequence heterogeneity in the central portion of the TprK hydrophilic domain in three T. pallidum strains. However, sequence heterogeneity was not observed in TprK of T. pallidum Nichols strain Seattle or GenBank. In this study, we demonstrate that intrastrain amino acid sequence heterogeneity is present in TprK of T. pallidum Nichols strain UNC and Street strain 14. This heterogeneity results from nucleotide sequence variability of the single-copy tprK gene in treponemal subpopulations. Additionally, we demonstrate that intrastrain or interstrain sequence heterogeneity is very limited in TprJ of these two treponemal strains.
T. pallidum Nichols strain UNC (obtained from J. Baseman via the Centers for Disease Control and Prevention, Atlanta, Ga.) and Street strain 14 (an erythromycin-resistant 1977 clinical isolate obtained from the Centers for Disease Control and Prevention) were cultivated by intratesticular infection of adult New Zealand White rabbits as previously described (12). The spirochetes were extracted and genomic DNA was isolated as previously described (11). Nichols strain Seattle genomic DNA was provided by S. Lukehart and A. Centurion-Lara. The entire tprK (AE001259) and tprJ (AE001237) genes were PCR amplified from treponemal genomic DNA using oligonucleotide primer pairs that were based on the nucleotide sequence of regions flanking these genes (for tprK, forward primer 5′-TTTGGAAGCGTTAGAATCTC-3′ and reverse primer 5′-ACAATACCAAAAACGCTAAAG-3′; for tprJ, forward primer 5′-GAAGCGTTGTTGTCAGTGC-3′ and reverse primer 5′-GGGTGTGGGTGTGGTTTG-3′). The information for primer design was obtained from the complete genome sequence of T. pallidum Nichols strain GenBank (6). The central region of the tprK gene (corresponding to amino acids 218 to 361) was PCR amplified from treponemal genomic DNA using the oligonucleotide primers of Centurion-Lara et al. (3). PCR products were gel purified, cloned into a plasmid vector, and transformed into Escherichia coli as previously described (11). Sequences of DNA inserts were determined at the University of North Carolina Automated DNA Sequencing Facility as previously described (11). Both DNA strands were sequenced to ensure accuracy. DNA and amino acid sequence analyses were performed using the AssemblyLIGN program 1.0.9b (MacVector 6.5.1; Oxford Molecular Group, Inc., Campbell, Calif.) and the PileUp and PlotSimilarity programs (Wisconsin Genetics Computer Group, version 10.0, University of Wisconsin Biotechnology Center, Madison). Total treponemal RNA was isolated from Nichols strain UNC treponemes as previously described (11). Reverse transcriptase PCRs (RT-PCRs) were performed using the primers of Centurion-Lara et al. (3) for the central region of tprK as described above. Southern hybridizations of restriction endonuclease-digested treponemal genomic DNA were performed as previously described (9) with an oligonucleotide probe (5′-CAGCGGGTAGGGCACAGG-3′) that is based on sequence present in the conserved 5′ end of each tprK gene examined in this study.
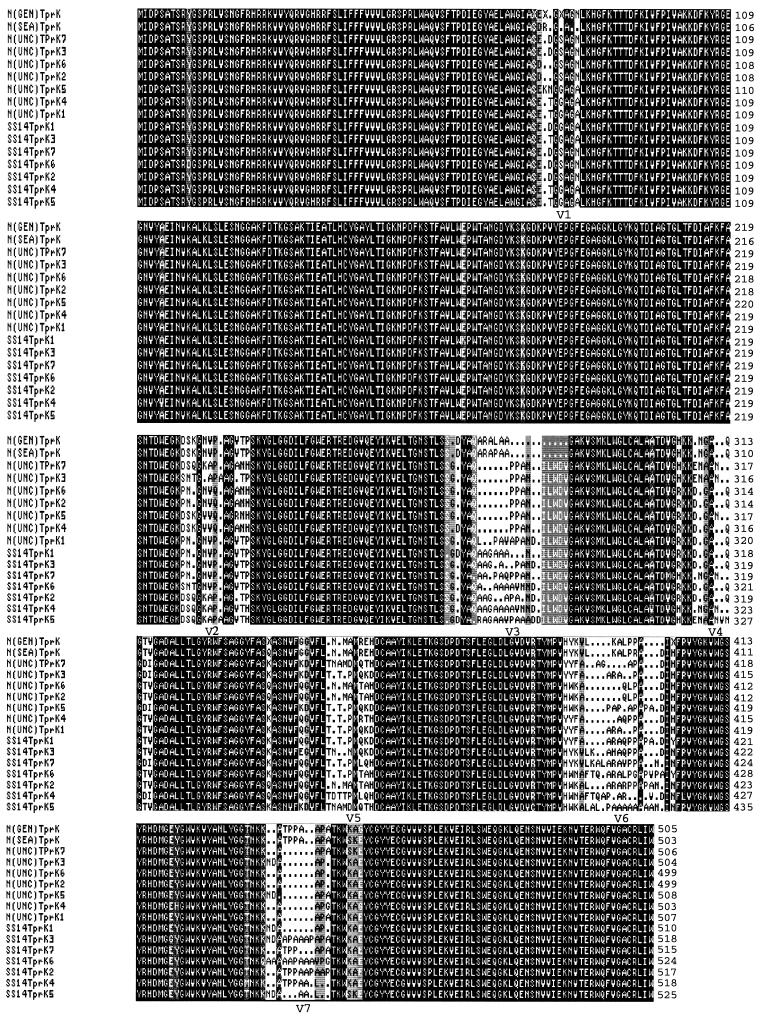
An amino acid sequence alignment of the full-length TprK protein of T. pallidum Nichols strain GenBank, Seattle, and UNC and Street strain 14 is shown in Fig. 1. Amino acid sequences were deduced from nucleotide sequences of seven E. coli clones containing the PCR-amplified tprK gene of Nichols strain UNC or Street strain 14. There was virtually complete identity in the first N-terminal 73 amino acids and the last C-terminal 54 amino acids of all the TprK sequences, regardless of the treponemal strain or clone. However, between the conserved N- and C-terminal regions, seven regions of heterogeneity, flanked by regions of high conservation, were apparent in all the TprK sequences. This heterogeneity, which resulted from amino acid substitutions, insertions and minor deletions, did not introduce any premature stop codons or frameshifts. None of the clones for either Nichols strain UNC or Street strain 14 contained sequences identical to those of the other clones or those of Nichols strain GenBank or Seattle. Additionally, in contrast to the data of Centurion-Lara et al. (4) for Nichols strain Seattle, we did not observe a 67-bp deletion in the nucleotide sequence 38 bp downstream of the Nichols strain UNC tprK gene. Although TprK of Nichols strain Seattle and TprK of Nichols strain GenBank are virtually identical, the latter strain also does not contain a deletion downstream of tprK (4). Since the sequence heterogeneity observed with Nichols strain UNC was unanticipated, a control PCR was performed using Nichols strain Seattle DNA and our primers for the amplification of full-length tprK. Analysis of PCR products from four clones indicated that the deduced amino acid sequences of Nichols strain Seattle TprK were essentially identical (data not shown). These results, which are in agreement with those of Centurion-Lara et al. (4), confirmed the specificity of the PCR products obtained with our primers.
FIG. 1.
Deduced amino acid sequence alignment of the full-length TprK protein of T. pallidum Nichols (N) strain GenBank (GEN), Nichols strain Seattle (SEA), Nichols strain UNC (UNC), and Street strain 14 (SS14). Designations 1 through 7 represent TprK sequences obtained from seven E. coli clones for Nichols strain UNC and Street strain 14. Identical residues are shown on a black background. Gaps in the sequence are indicated by dots.
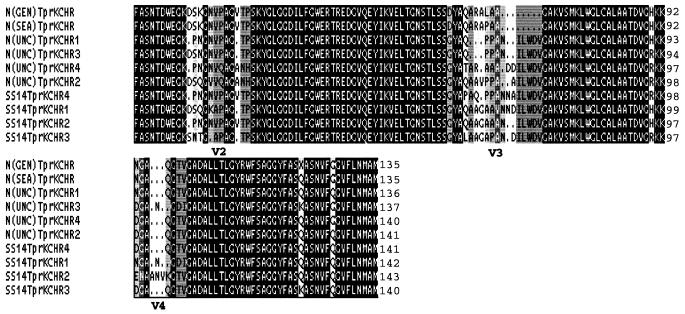
To further investigate the TprK variability of Nichols strain UNC and Street strain 14, we PCR amplified, cloned, and sequenced treponemal DNA encoding the TprK central hydrophilic region (CHR), which contains three of the seven heterogeneous regions that were observed in the full-length TprK. An amino acid sequence alignment of the TprK CHR is shown in Fig. 2. This alignment contains the sequences from four Nichols strain UNC clones and four Street strain 14 clones and the corresponding sequences of the Nichols strain GenBank and Seattle. Comparison of the Nichols strain UNC and Street strain 14 CHR sequences either within or between strains showed that none was completely identical in the heterogeneous regions of TprK. Since all of the Nichols strain UNC and Street strain 14 CHR sequences analyzed to date are unique, the scope of TprK variability in these treponemes is unknown. Centurion-Lara et al. (4) reported that approximately 50 to 80% of TprK clones from three T. pallidum strains (Bal 7, Bal 73-1, and Sea 81-4) contained unique CHR sequences. The heterogeneity of TprK is in contrast to several other T. pallidum proteins that exhibit remarkable amino acid sequence conservation within and between treponemal strains (1, 2, 9, 10).
FIG. 2.
Deduced amino acid sequence alignment of the TprK CHR of T. pallidum Nichols (N) strain GenBank (GEN), Nichols strain Seattle (SEA), Nichols strain UNC (UNC), and Street strain 14 (SS14). Designations CHR 1 through 4 represent the TprK CHR sequences obtained from four additional E. coli clones for Nichols strain UNC and Street strain 14. Identical residues are shown on a black background. Gaps in the sequence are indicated by dots.
Transcription of the tprK gene of T. pallidum Nichols strain UNC was examined by RT-PCR analysis. PCR products that represented amplified cDNA reverse transcribed from treponemal mRNA were cloned and sequenced. Data from four clones indicated that transcripts with heterogeneous tprK sequences were present in treponemes isolated from infected rabbit testes (data not shown). Interestingly, all of the tprK sequences are unique. Our results support the contention that heterogeneous TprK proteins are synthesized during syphilitic infection. These findings have implications from the standpoint of vaccine development, since they suggest that TprK is likely to be a challenging target due to its variability.
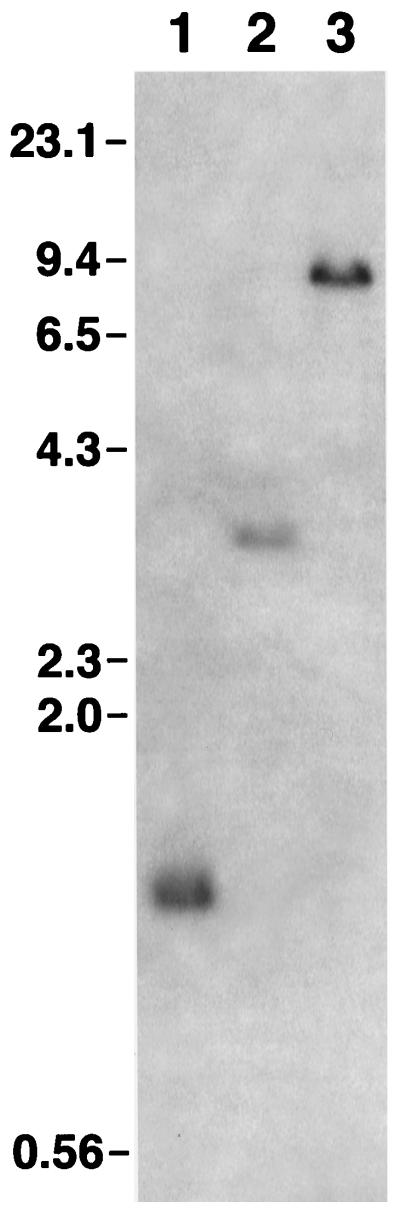
The intrastrain heterogeneity of T. pallidum Nichols strain UNC and Street strain 14 TprK could be due to the presence of multiple copies of the tprK gene in a single spirochete or to subpopulations of treponemes that contain variations in the nucleotide sequence of tprK. In the absence of Southern hybridization data, Centurion-Lara et al. (4) proposed that their results with the non-Nichols strains could be accounted for by the latter. To address this, we performed Southern hybridizations of Sau3AI-, MseI-, or HindIII-digested Nichols strain UNC and Street strain 14 genomic DNA with an oligonucleotide probe that recognized an 18-bp sequence in a DNA restriction fragment containing the conserved 5′ end of tprK and extragenic flanking sequence. The probe hybridized to a single DNA fragment of the predicted size in each of the genomic DNA digests of Nichols strain UNC (Fig. 3) and Street strain 14 (data not shown), indicating that tprK is present in a single copy. Identical results were observed with an oligonucleotide probe that recognized a sequence in a DNA restriction fragment containing the conserved 3′ end of tprK and extragenic flanking sequence (data not shown). Thus, intrastrain heterogeneity of TprK in T. pallidum Nichols strain UNC and Street strain 14 is not due to the presence of additional copies of tprK. These results are in agreement with those of Fraser et al. (6) for the complete genome sequence of Nichols strain GenBank.
FIG. 3.
Southern blot analysis of restriction endonuclease-digested T. pallidum Nichols strain UNC genomic DNA with an oligonucleotide probe that recognized a DNA restriction fragment containing the conserved 5′ end of tprK and extragenic flanking sequence. Genomic DNA was digested with Sau3AI (lane 1), MseI (lane 2), or HindIII (lane 3). Size standards are shown on the left in kilobases. Identical results were obtained with Street strain 14 genomic DNA digests (data not shown).
Our observations with TprK prompted us to determine if heterogeneity is present in other Tpr proteins. We had previously identified and sequenced the tprJ gene of T. pallidum Nichols strain UNC and found that the deduced amino acid sequence of TprJ is identical to that predicted from the Nichols strain GenBank sequence (11). We PCR amplified, cloned, and sequenced the full-length tprJ gene from Street strain 14. Figure 4 shows the deduced amino acid sequence of TprJ from three E. coli clones. Only four single-amino-acid changes were observed between the Street strain 14 clones. The Street strain 14 TprJ amino acid sequence from the clones is 99.5 to 99.6% identical to that of the Nichols strain. Our results suggest that there is little heterogeneity of TprJ within or between these two strains. Studies are currently in progress to evaluate the potential heterogeneity of additional Tpr proteins.
FIG. 4.
Deduced amino acid sequence alignment of the full-length TprJ protein of T. pallidum Nichols (N) strain GenBank (GEN) and Street strain 14 (SS14). Designations 1 through 3 represent TprJ sequences obtained from three E. coli clones for Street strain 14. Identical residues shown on a black background.
Phenotypic variation of proteins encoded by multigene families is a common strategy used by bacterial pathogens for evading host clearance and adapting to host microenvironments (5). It is likely that the intrastrain heterogeneity of TprK is similarly beneficial to T. pallidum. However, such heterogeneity is not required for syphilis pathogenesis, since Nichols strain GenBank and Nichols strain Seattle produce a chronic, disseminated infection in the rabbit model. Centurion-Lara et al. (4) proposed that multiple testicular passages of Nichols strain GenBank and Seattle have led to the selection of a rabbit-adapted clonal population of treponemes that express only one TprK. It should be noted that both of these organisms were originally obtained in the late 1970s from J. N. Miller (University of California, Los Angeles), who had serially passaged the treponemes in rabbit testes every 7 to 10 days for more than 25 years (4). In contrast, the non-Nichols strains that exhibit intrastrain heterogeneity of TprK were isolated more recently and have been passaged in rabbits only a few times (4). Although Nichols strain UNC has been passaged multiple times during the 20 years that it has been in our laboratory, we routinely administered hydrocortisone acetate to infected rabbits to promote higher yields of treponemes. We hypothesize that this protocol altered the selective immunologic pressure of the host, thus allowing for the maintenance of TprK variants that arise during infection. Studies are ongoing in our laboratory to elucidate genetic events (such as mutation and recombination) that have resulted in the generation of these variants.
Nucleotide sequence accession numbers.
The GenBank accession numbers for the sequences of the tprK genes of Nichols strain UNC and Street strain 14 are AF227219 through AF227232 and AF250503 through 250510. The GenBank accession numbers for the sequences of the tprJ gene of Street strain 14 are AF261075 through AF261077.
Acknowledgments
We thank S. Greene for reviewing the manuscript prior to submission, D. Fenstermacher and the UNC-CH Center for BioInformatics for assistance with figure production, and S. Lukehart and A. Centurion-Lara for providing Nichols strain Seattle DNA.
This research was supported by National Institutes of Health grant U19-AI31496.
REFERENCES
- 1.Cameron C E, Castro C, Lukehart S A, Van Voorhis W C. Sequence conservation of glycerophosphodiester phosphodiesterase among Treponema pallidum strains. Infect Immun. 1999;67:3168–3170. doi: 10.1128/iai.67.6.3168-3170.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Centurion-Lara A, Arroll T, Castillo R, Schaffer J M, Castro C, Van Voorhis W C, Lukehart S A. Conservation of the 15-kilodalton lipoprotein among Treponema pallidum subspecies and strains and other pathogenic treponemes: genetic and antigenic analyses. Infect Immun. 1997;65:1440–1444. doi: 10.1128/iai.65.4.1440-1444.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Centurion-Lara A, Castro C, Barrett L, Cameron C, Mostowfi M, Van Voorhis W C, Lukehart S A. Treponema pallidum major sheath protein homologue TprK is a target of opsonic antibodies and the protective immune response. J Exp Med. 1999;189:647–656. doi: 10.1084/jem.189.4.647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Centurion-Lara A, Godornes C, Castro C, Van Voorhis W C, Lukehart S A. The tprK gene is heterogeneous among Treponema pallidum strains and has multiple alleles. Infect Immun. 2000;68:824–831. doi: 10.1128/iai.68.2.824-831.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Deitsch K W, Moxon E R, Wellems T E. Shared themes of antigenic variation and virulence in bacterial, protozoal, and fungal infections. Microbiol Mol Biol Rev. 1997;61:281–293. doi: 10.1128/mmbr.61.3.281-293.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Fraser C M, Norris S J, Weinstock G M, White O, Sutton G G, Dodson R, Gwinn M, Hickey E K, Clayton R, Ketchum K A, Sodergren E, Hardham J M, McLeod M P, Salzberg S, Peterson J, Khalak H, Richardson D, Howell J K, Chidambaram M, Utterback T, McDonald L, Artiach P, Bowman C, Cotton M D, Fujii C, Garland S, Hatch B, Horst K, Roberts K, Sandusky M, Weidman J, Smith H O, Venter J C. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science. 1998;281:375–388. doi: 10.1126/science.281.5375.375. [DOI] [PubMed] [Google Scholar]
- 7.Norris S J the Treponema pallidum Polypeptide Research Group. Polypeptides of Treponema pallidum: progress toward understanding their structural, functional, and immunological roles. Microbiol Rev. 1993;57:750–779. doi: 10.1128/mr.57.3.750-779.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Stamm L V. Biology of Treponema pallidum. In: Holmes K K, Mardh P A, Sparling P F, Lemon S M, Stamm W E, Piot P, Wasserheit J N, editors. Sexually transmitted diseases. 3rd ed. New York, N.Y: McGraw-Hill; 1999. pp. 467–472. [Google Scholar]
- 9.Stamm L V, Greene S R, Barnes N Y. Identification and characterization of the gyrB gene from Treponema pallidum subsp. pallidum. FEMS Microbiol Lett. 1997;153:129–134. doi: 10.1111/j.1574-6968.1997.tb10473.x. [DOI] [PubMed] [Google Scholar]
- 10.Stamm L V, Greene S R, Barnes N Y, Bergen H L, Hardham J M. Identification and characterization of a Treponema pallidum subsp. pallidum gene encoding a DNA adenine methyltransferase. FEMS Microbiol Lett. 1997;155:115–119. doi: 10.1111/j.1574-6968.1997.tb12694.x. [DOI] [PubMed] [Google Scholar]
- 11.Stamm L V, Greene S R, Bergen H L, Hardham J M, Barnes N Y. Identification and sequence analysis of Treponema pallidum tprJ, a member of a polymorphic multigene family. FEMS Microbiol Lett. 1998;169:155–161. doi: 10.1111/j.1574-6968.1998.tb13312.x. [DOI] [PubMed] [Google Scholar]
- 12.Stamm L V, Kerner T C, Jr, Bankaitis V A, Bassford P J., Jr Identification and preliminary characterization of Treponema pallidum protein antigens expressed in Escherichia coli. Infect Immun. 1983;41:709–721. doi: 10.1128/iai.41.2.709-721.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]