Abstract
SARS-CoV-2 has been shown to enter and infect human cells via interactions between spike protein (S glycoprotein), and angiotensin-converting enzyme 2 (ACE2). As such, it may be possible to suppress the infection of the virus via the blocking of this binding interaction, through the use of specific peptides that can mimic the human ACE 2 peptidase domain (PD) α 1-helix. Herein, we report the use of competitive assays along with surface plasmon resonance (SPR) to investigate the effect of peptide sequence and length on spike protein inhibition. The characterization of these binding interactions helps us understand the mechanisms behind peptide-based viral blockage and develops SPR methodologies to quickly screen disease inhibitors. This work not only helps further our understanding of the important biological interactions involved in viral inhibition but will also aid in future studies that focus on the development of therapeutics and drug options. Two peptides of different sequence lengths, [30–42] and [22–44], based on the α 1-helix of ACE2 PD were selected for this fundamental investigation. In addition to characterizing their inhibitory behavior, we also identified the critical amino acid residues of the RBD/ACE2-derived peptides by combining experimental results and molecular docking modeling. While both investigated peptides were found to effectively block the RBD residues known to bind to ACE2 PD, our investigation showed that the shorter peptide was able to reach a maximal inhibition at lower concentrations. These inhibition results matched with molecular docking models and indicated that peptide length and composition are key in the development of an effective peptide for inhibiting biophysical interactions. The work presented here emphasizes the importance of inhibition screening and modeling, as longer peptides are not always more effective.
Keywords: SARS-CoV-2, COVID-19, surface plasmon resonance, peptide inhibitors, molecular docking, biosensing
Graphical Abstract:

INTRODUCTION:
The recent coronavirus disease (COVID-19) outbreak caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has infected over 630 million individuals and has resulted in the death of over 6.6 million people as of November 2022, according to the World Health Organization (WHO).1 While a great deal of effort has gone into developing new ways to treat and prevent the onset of COVID-19,2–5 it is still affecting many people around the world, and it appears that it will continue to be present for the foreseeable future.6
At the molecular level, the infection of SARS-CoV-2 is directed via critical interactions between the viral spike protein and human ACE2 protein,7 whose normal function is catalyzing hydrolysis of the vasoconstrictor peptide angiotensin II.8, 9 To gain a deeper understanding of the interaction observed between SARS-CoV-2 and ACE2 one must first understand the structure of SARS-CoV-2. The overall virus structure consists of a single-stranded RNA genome made up of 30 kilobases that is encapsulated within a lipid bilayer. The virus also has three lipid membrane embedded structural proteins that are labeled as the envelope, membrane, and spike.10 The spike protein is a large (1208 residue), heavily glycosylated polypeptide that forms homotrimers which are what gives the Coronavirus its “corona” structure in electron micrographs. Each monomer consists of subunits (S1 and S2), with the receptor-binding domain (RBD) falling within the S1 subunit and corresponding to residues 319–541.11 In a demonstration of the high-resolution capabilities of CryoEM technologies, the full-length structure of the spike protein was determined at 2.8 Å only months after the global onset of COVID-19 (PDB 6VXX, 6VYB).12 From this work, the occurrence of an “open” and “closed” RBD configuration was identified, with only the “open” configuration being able to effectively bind to hACE2.12, 13 This binding induces dissociation, through prompting S2 transition to a more stable post-fusion phase that appears to be crucial for membrane fusion.14–16 Therefore it appears that binding to the ACE2 receptor is an imperative initial step for SARS-CoV-2 entry into human cells.17 It is important to note that other mechanisms of spike protein induced cellular damage have been proposed that do not rely on the ACE2 receptor of SARS-CoV-2.18, 19 Though disrupting the ACE2 binding site of SARS-CoV-2’s spike protein may also prevent these other interactions from having their deleterious effects.
Further information regarding the importance of the onset of COVD-19 and the structure of SARS-CoV-2 was obtained through X-ray crystallography. Most notably, the information related to the complex between spike protein RBD and hACE2 (PDB-ID: 6M0J, 6LZG, and 6VW1), which greatly aided in understanding the lethality of the virus.20–22 Based on crystal structures of the complex, various ACE2 and s-protein residues were identified as key parts of the ACE2/s-protein interaction. Using crystal structures of the ACE2/s-protein RBD complex, amino acids at the ACE2 motifs and the viral s-protein RBD in the interface core were defined. In the recognition of RBD, it was found that the protease domain (PD) of ACE2 primarily interacts with the α 1-helix (Ser19-Gln42).22 By deciphering which amino acids are in contact between the two proteins, the development of specific disruptors against the SARS-CoV-2/ACE2 protein-protein interaction (PPI) can be pursued.23, 24 This is significant as disrupting protein binding interfaces with small-molecule inhibitors is often considered to be less effective,25 whereas peptides appear as effective small molecules inhibitors due to their ability to bind at interface regions through multiple residue interactions.26
Peptides, in a similar fashion as COVID-19 monoclonal antibodies (mAbs), aim to abrogate the SARS-CoV-2/ACE2 interaction.27, 28 CoV mAbs predominantly target the trimeric S glycoprotein, and most achieve this through recognition of epitopes within the RBD ACE2 binding site.28–30 However, as RNA viruses accumulate mutations over time, antibody resistance can begin to arise which requires the use of antibody cocktails to maintain effectiveness.31 This has been shown in recent work where emerging SARS-CoV-2 mutants are no longer targeted by antibodies effective against the original strain.32 On the other hand, as human proteins evolve at a much slower rate peptides and proteins that mimic the human side of the binding interaction will not lose effectiveness and may facilitate the development of COVID-19 treatments or vaccines33 that are independent of further viral mutation. Overall, peptide and protein therapies have shown to be highly specific, display low interference with non-target biological systems, are excellently tolerated in humans, and can even enjoy faster FDA approval times.34 Though more research into the fundamental interactions between protein interactions where peptides act as blockers is clearly still needed.
Recent computational studies have attempted to show that small molecules and peptide-mimetic inhibitors can inhibit SARS-CoV-2 s-protein interactions.35 One study36 focused on the N-terminal helix of ACE2 and its 23 resides important in spike protein binding. Using MD simulations and free energy calculations they showed that the 23-residue peptide, as well as a mutant version, should bind with high affinity to the SARS-CoV2 s-protein RBD. Another study37 reported a minimum binding epitope from the ACE2 N-terminal helices identified using MD-based methods. With the smaller peptide retaining high affinity for the s-protein RBD. The combination of these computational studies, focused on peptides as potential inhibitors, could form the design basis for peptide-based SARS-CoV-2 therapeutics. This is significant as currently ACE2 recombinant protein is being considered as a COVID-19 treatment.38, 39 As ACE2 plays a pivotal role in many cellular processes, such as blood pressure regulation and various other cardiovascular functions,40 consideration of what side effects ACE2 treatment might cause is important. Therefore, here we investigate the potential of how small ACE2-derived peptides may function in interfering with SARS-CoV-2 binding, by blocking binding sites on the S glycoprotein without imparting the deleterious side effects expected from whole protein therapies.
To this end, we synthesized and tested short ACE2-derived peptides engineered to target the viral S glycoprotein, resulting in significant reduction in binding between the spike and ACE2 proteins. We accomplished this by utilizing the label-free approach surface plasmon resonance, which provides highly sensitive detection capabilities. The first peptide (Glu22-Ser44) was selected to mimic the ACE2 regions that interact with the S1 subunit based on results from previously discussed crystal structures.22 The second one is a truncated version of the first peptide (D30-Q42). Next, we performed molecular docking using the PatchDock program aimed at furthering our understanding of the interactions between SARS-Cov2 RBD and the ACE2-derived peptides. By gaining a deeper understanding of the interactions between peptides and viruses, such as ACE2 peptides’ ability to inhibit SARS-CoV-2 binding, our findings have the potential to open up new avenues of research related to ways to treat and investigate viral diseases such as COVID-19. This is of great interest as peptides are generally considered to be highly selective, effective, and safe, making them ideal for future therapeutic uses.41
EXPERIMENTAL:
Materials:
Recombinant SARS-CoV-2 spike protein, S1 subunit (Val16-Gln690) was purchased from RayBiotech. Human ACE2, His tag (E. coli) was obtained from MP Biomedicals, LLC. 11-mercaptoundecanoic acid (MUA). O-(2-Aminoethyl)-methylpoly-ethylene glycol (PEGamine), N-Hydroxysuccinimide (NHS), and 1-(3-dimethylaminopropyl)-3-ethylcarbodiimide hydrochloride (EDC) were purchased from Sigma-Aldrich (St. Louis, MO). All protein solutions were prepared in 20 mM phosphate buffered saline (containing 150 mM NaCl, pH 7.4). Rink amide MBHA resin was obtained from Aapptec (Louisville, KY). Fmoc-protected amino acids were obtained from Anaspec (Fremont, CA). Piperidine was purchased from Alfa Aesar (Ward Hill, MA).2-(1H-benzotriazol-1-yl)-1,1,3,3-tetramethyluronium hexafluorophosphate (HBTU, 99.6%) and diisopropylethylamine (DIEA, 99.5%) were purchased from Chem-Impex (Wood Dale, IL) and ACROS (Germany) respectively. Triisopropylsilane (TIPS) was obtained from TCI (Portland, OR). α-cyano-4-hydroxycinnamic acid (CHCA) was purchased from Sigma Aldrich (St. Louis, MO). Crystal structure of SARS-CoV-2 spike receptor-binding domain bound with ACE2 PD at 2.45 A° resolution with PDB ID: 6M0J was retrieved from RCSB PDB database (https://www.rcsb.org/).
Solid-phase peptide synthesis (SPPS) of linear peptides
CSBio CS336S peptide synthesizer (Menlo Park, CA) was used to synthesize the linear peptide sequences. 500 mg of Rink Amide resin with a loading capacity of 0.678 mmol/g was used for each synthesis. Fmoc-protected L-amino acids (1 mmol each), DIEA (0.8 M in DMF), HATU (0.4 M in DMF) were used for each coupling step, and 20% piperidine/DMF was used for Fmoc deprotection. At the end of synthesis, the resin was treated with a solution of TFA, tri-isopropyl silane, and water (95:2.5:2.5 by volume) to cleave the peptide from the resin. The cleavage solution was mixed with cold diethyl ether to obtain the crude peptide as precipitates.
Purification of the synthesized peptides
The crude peptide was purified on a Thermo Ultimate 3000BX HPLC, equipped with a preparative column (Kinetex 5 μm EVO, 250 Å~ 21.2 mm2). A gradient of 0–100% acetonitrile (with 0.1% TFA) in water (with 0.1% TFA) was used. The identity of each fraction was confirmed by MALDI-TOF on a SCIEX 5800 mass spectrometer. Fractions containing the product were pooled and lyophilized to obtain the pure peptide.
SPR analysis of ACE2-derived peptide inhibition of SARS-CoV-19 spike protein binding:
A two channel SPR spectrometer NanoSPR-6 (NanoSPR, Addison, IL) with a 670 nm laser light source was utilized for all SPR experiments. SPR gold chips were fabricated via e-beam evaporation onto cleaned BK-7 glass slides with a 2 nm thick chromium adhesion layer, followed by deposition of a 46 nm thick gold layer following previously published procedures.42
Surface interactions were monitored using angular scanning mode which tracks the angle of minimum reflectivity. The gold substrate was incubated for 18 h in 1 mM MUA ethanol solution to form a self-assembled monolayer of carboxyl functional groups. Following this the chip was extensively rinsed with ethanol and DI water, then dried under an air stream. To activate the carboxyl acid group, EDC (0.4 M)/NHS (0.1 M) solution was injected into the flow cell and allowed to interact for 30 min. After 10 min of rinsing, 0.5 μg/ml of spike protein in PBS at a pH of 7.4 was injected and incubated for 1 hr to allow the formation of covalent amide linkages. Followed by a 10 min rinse to eliminate any residual unbound spike protein. Passivation of unused activated carboxyl groups was performed by incubation with 10 mg/ml PEG amine solution for 1 hr. Then the inhibition assay was performed. All protein binding and inhibition studies were performed under identical buffer conditions with pH 7.4 PBS.
Preparation of both receptor and peptide molecules
The human coronavirus spike protein structures and ACE2 structures were downloaded from the RCSB protein data bank. ACE2 structures were modified manually to produce the derived peptide structures. Depending on the peptide needed, the appropriate section of ACE2 was isolated so as to run future docking simulations. In addition, the binding domain on the spike protein was separated from the rest of the structure to specify the interactions between the binding domain and ACE2-derived peptides. The structures of the peptides and spike protein binding domain were verified after editing using PyMOL.
Molecular Docking
The MD between human coronavirus spike protein and each peptide understudy were performed using PatchDock web server. PatchDock calculates the docking transformation between two molecules using a geometry-based MD algorithm to obtain the most complementary molecular interface. This ascertains the peptide position in relation to the receptor with maximal interface area coverage and minimal steric hindrance.43 Each ACE2-derived peptide was docked with SARS-CoV-2 RBD by uploading the molecules to the Patchdock server, which is an automated molecular docking server. Clustering RMSD was chosen as 4.0 Å. PyMOL was used to analyze the docking results of RBD/ACE2 derived peptide interaction by identifying the original binding residue between the RBD/ACE2 PD complex.
RESULT AND DISCUSSION:
The N-terminal region of the ACE2 PD is critical for binding to the SARS-CoV-2 spike protein
In order to investigate the best attributes to explore when designing a small peptide-based inhibitor that can block the interaction of SARS-CoV-2 spike protein with the ACE2 receptor, we investigated existing structures of known amino acid interactions that are critical for binding between SARS-CoV-2 and ACE2 (Table. 1). This includes the crystal structure of the ACE2 PD/RBD complex (PDB ID: 6M0J, 6VW1, and 6LZG) in addition to the full length of the ACE2 with SARS-CoV-2 complex (PDB ID: 6M17). Comparisons of the ACE2 interacting residues with SARS-CoV-2 spike protein according to the previous analysis of ACE2/RBD crystal structures20–22, 24 are shown in Table 2 and Table 3. It is clear from the table that the α 1 helix (S19-S44) of ACE2 provides the most contact with the SARs-CoV-2 RBD, as well as a small area on the α 2 helix, the short loop between α10/α11, and the linker between β3/β4. Therefore, selection of the peptide-based inhibitor was chosen based upon this insight and recent work published by Zhang et al.44 Where they suggested that the 23-mer peptide mimics the α−1 helix as a potential drug for SARS-CoV2 and demonstrated its affinity to the viral RBD. With the binding energies for interaction between the peptide and the viral RBD expected to be close to that of the naturally occurring RBD-ACE2 complex.
Table 1.
Crystal structure of ACE2/SARS-CoV-2 complex and the primary publication
| ID Structure | Paper Title | Experimental Method | Published Date | Resolution (A°) | Primary publication DIO | |
|---|---|---|---|---|---|---|
| 1 | 6M0J | Crystal structure of SARS-CoV-19 spike receptor-binding domain bound with ACE2 | X-Ray diffraction | 18/03/2020 | 2.45 | 22 |
| 2 | 6M17 | The 2019-nCov RBD/ACE2-B0AT1 complex | Electron microscopy | 11/03/2020 | 2.90 | 24 |
| 3 | 6VW1 | Structure of SARS-CoV-2 chimeric receptor-binding domain complexed with its receptor human ACE2 | X-Ray diffraction | 04/03/2020 | 2.68 | 21 |
| 4 | 6LZG | Structure of novel coronavirus spike receptor-binding domain complexed with its receptor ACE2 | X-Ray diffraction | 18/03/2020 | 2.50 | 20 |
Table 2.
Comparison of ACE2 interacting residues with SARS-CoV-2 spike protein of ACE2 amino acids provided by a 1 helix.
| α 1 Helix of ACE2 Residues Interreacting with RBD | ||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PDB | S 19 | T 20 | I 21 | E 22 | E 23 | Q 24 | A 25 | K 26 | T 27 | F 28 | L 29 | D 30 | K 31 | F 32 | N 33 | H 34 | E 35 | A 36 | E 37 | D 38 | L 39 | F 40 | Y 41 | Q 42 | S 43 | S 44 |
| 6M0J | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||||||||||
| 6M17 | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||||||||
| 6VW1 | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||||||||
| 6LZG | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | |||||||||||||
Table 3.
Comparison of ACE2 interacting residues with SARS-CoV-2 spike protein of ACE2 amino acids provided by α 2, the short loop between α10/α11, and the link between β3/β4.
| α 2, 10, 11, Helix and Loop β3/β4 of ACE2 Residues Interacting with RBD | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PDB | S 79 | T 80 | I 81 | E 82 | E 83 | A 325 | K 329 | T 330 | L 353 | D 354 | K 355 | K 357 | |
| 6M0J | ✓ | ✓ | |||||||||||
| 6M17 | ✓ | ✓ | ✓ | ✓ | ✓ | ||||||||
| 6VW1 | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||
| 6LZG | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ||
Consequently, the [22–44] peptide (Glu22-Ser44, wheat ribbon in fig 1B) has been synthesized to mimic the α−1 helix and tested for its ability to interfere with SARS-CoV-2-spike protein/ACE2 binding. In addition, a smaller peptide was formulated by eliminating the first 8 and last 2 histidine residues from the original [22–44] peptide. This 13-residue peptide was also synthesized and tested as an inhibitor of the ACE2/S1 complex. This small peptide (Asp30-Gln42, pink ribbon in fig 2B) contains most of the key contact with the RBD and represents the central and C-terminal of the isolated α−1 helix of ACE2. Here we aim to investigate the effect of changing the peptide length and residue coverage on the blocking efficiency of the spike protein.
Figure 1.
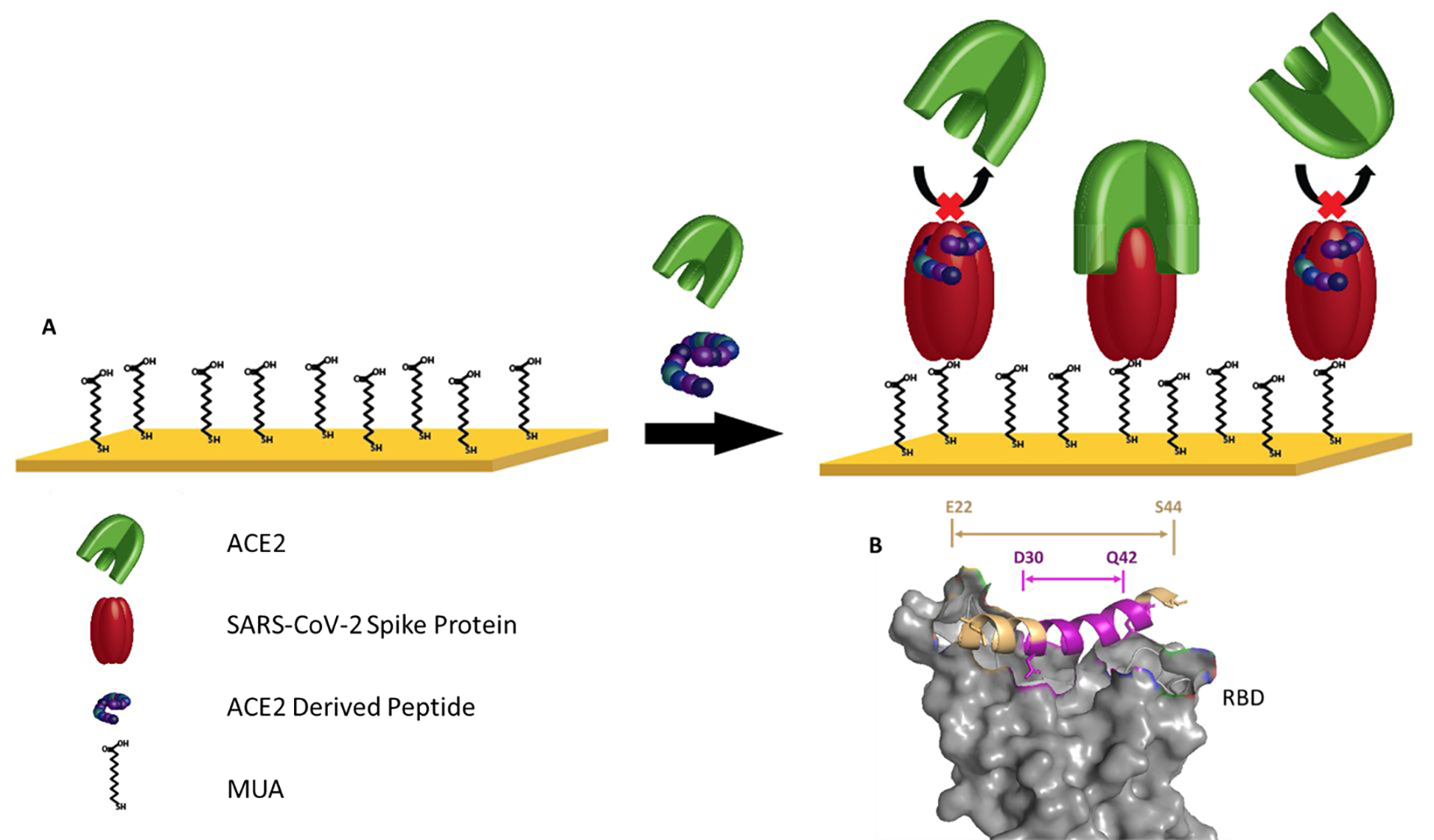
(A) Schematic of the competitive assay. (B) Relative location of [30–42] peptide (purple) and [22–44] peptide (tan) corresponding to PDB ID: 6m0j
Figure 2.
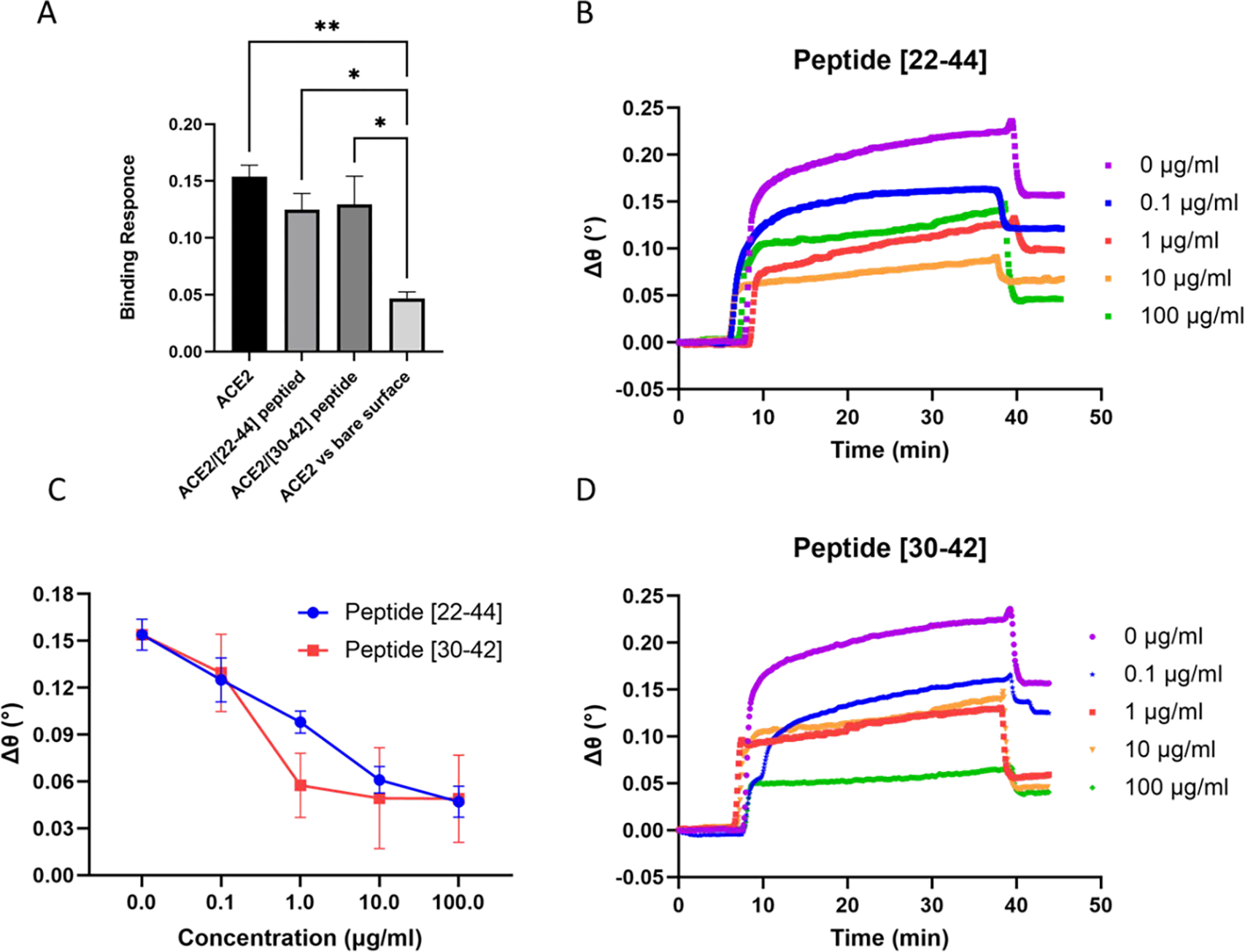
Evaluation of blocking efficiency of the ACE2-derived peptides to the S-protein using competitive SPR. A) Specific binding measured for the S-protein on the surface with ACE2. B) SPR sensorgrams with [22–44] peptide. C) The change of SPR binding signal as a function of peptide cocentration. D) SPR sensorgrams with [30–42] peptide.
Evaluation of Blocking Efficiency Using Competition Surface Plasmon Resonance (SPR)
SPR is a well-established technique that has been utilized to investigate various biophysical interactions, especially those targeting proteins.45–48 However, using SPR for direct analysis of interactions between small molecules, such as small peptides, and proteins has commonly been a difficulty, largely due to the low signal change caused from small molecule interactions.49 We, therefore, utilized a competition assay to evaluate the binding of the peptide to the SARS-CoV-2-spike protein using SPR spectroscopy. With Figure 1 providing a schematic of this competition assay. 50 μg/ml of SARS-CoV-2-spike protein was immobilized on a self-assembled monolayer via carboxyl functional groups linked to the gold chip substrate. An SPR competitive assay was then utilized to determine the ability of the ACE2-derived peptides to abrogate interactions between human ACE2 and SARS-CoV-2 spike protein. Peptides with different concentrations were mixed with 30 μg/ml of human ACE2 protein and injected onto the chip coated with SARS-CoV-2-spike protein. Then, as a control, free human ACE2 solution (30 μg/ml) was injected. To confirm the specificity of ACE2/spike protein interaction, we also conducted an additional independent control experiment using a -COOH terminated alkanethiol surface without the spike protein.
Figure 2A depicts the SPR binding response of ACE2/spike protein with no inhibition, ACE2/spike protein in the presence of 0.1 μg/ml of each peptide, and the independent control which represents ACE2 binding to S1 surface free. The SPR binding shift observed during the independent control experiment is significantly lower, thereby demonstrating the specificity of the ACE2/spike protein complex under our experimental conditions. Figure 2B and 2D demonstrate that increasing the concentration of both peptides results in a decrease in the binding signal for the ACE2 protein binding to SARS-CoV-2-spike protein on the chip, indicating a concentration-dependent inhibition of the spike protein.
To visually compare binding signal between the S1 subunit and ACE2 in the absence of the peptide (0 μg/ml) with signal after adding the peptides at different concentrations (0.1, 1, 10, and 100 μg/ml), we plotted the SPR angle response versus the peptide’s concentration (fig 2C). We found that the ACE2/spike protein binding caused an angle shift of 0.147±0.009 degrees. While the [22–44] peptide showed a maximum SPR angle inhibition of the ACE2/S1 complex formation with a 0.047±0.009-degree reduction in SPR signal. It is of note that the small peptide, [30–42], shows a similar maximum inhibition potential at (0.049±0.027 degrees). This result suggests that both peptides are able to disrupt the ACE2/spike protein interactions. In addition, this could indicate that the central and C-terminal region of the isolated α−1 helix of ACE2 is the essential motif important for disrupting the ACE2/S1 interaction since both peptides show similar max inhibition effects.
Based on the crystal structure of the SARS-CoV-2 spike protein and ACE2 receptor complex (PDB ID: 6m0j) solved by Lan et al.22 the polar residues [24, 30, 35, 37, 38, 41, 42] of ACE2 helix-1 are found to be the key interfacial interactions and are able to establish a network of hydrogen bonds with the SARS-CoV-2 spike protein. Accordingly, we can exhibit 6 critical amino acids that are part of both peptides under our study (at the central and C-terminal region). While [22–44] peptide has only one extra critical residue at the N-terminal region of the peptide. Moreover, it has been highlighted in the modeling study of RBD/ACE2 that the residues 37, 38, 41, and 42 are the key interfacial interactions between ACE2 and the RBD/spike protein.20 The combination of these structural insights and MD study lend support to our result of the [22–44] and [30–42] peptides as disruptors of the ACE2/S1 interaction and that the central and C-terminal of the isolated α−1 helix of ACE2 contain more critical residue compared to the N-terminal of the isolated α 1 helix of ACE2 PD.
When examining Figures 2 B, C, and D in more detail, it was found that the [30–42] peptide reached higher blocking efficiency and achieved saturation of the spike protein at lower concentrations compared to the [22–44] peptide. In the case of the [30–42] peptide, a 0.057±0.02-degree reduction of SPR signal was observed at 1 μg/ml. At the same concentration, the [22–44] peptide shows only 0.098±0.007-degree reduction of the SPR signal. In addition, at 10 μg/ml the [30–42] peptide showed a maximum neutralization to the spike protein. On the other hand, the [22–44] peptide indicates similar maximum neutralization, however, at a higher concentration (100 μg/ml). This is possibly attributed to the existence of two consecutive serine residues in the [22–44] peptide, which likely affect the peptides flexibility. As it has been shown that the presence of two consecutive serine residues affects the [37–45] peptide binding affinity to spike protein.50
Next, we determined the half-maximal inhibitory concentration (IC50) of each peptide. Figures 3A and 3B show that the [22–44] and [30–42] peptides blocked the SARS-CoV-2 spike protein/ACE2 interaction with an IC50 values of 2.00 and 0.65 μg/ml, respectively. This implies that the two peptides exhibit a strong affinity to the spike protein, although the [22–44] peptide has extra amino acid residues. This is of interest as Yang et al.51 suggested that additional amino acids do not necessarily improve the overall affinity of the peptide for the SARS-CoV-2 S1 subunit. Their experimental study used two peptides [22–44] and [22–57], which showed similar inhibition potential of the ACE2/spike complex. This was also supported by their MD simulation, which showed the [22–57] peptide established fewer hydrogen bonds when compared to that of the shorter one, [22–44] peptide. Moreover, it was reported that residues 21–43 of the human ACE2 α 1-helix (similar to [22–44] peptide understudy) could strongly bind to SARS-CoV-2 RBD with micromolar affinity (KD= 1.3μM)13 which is comparable to that of the full-length ACE2 binding to RBD23.
Figure 3.
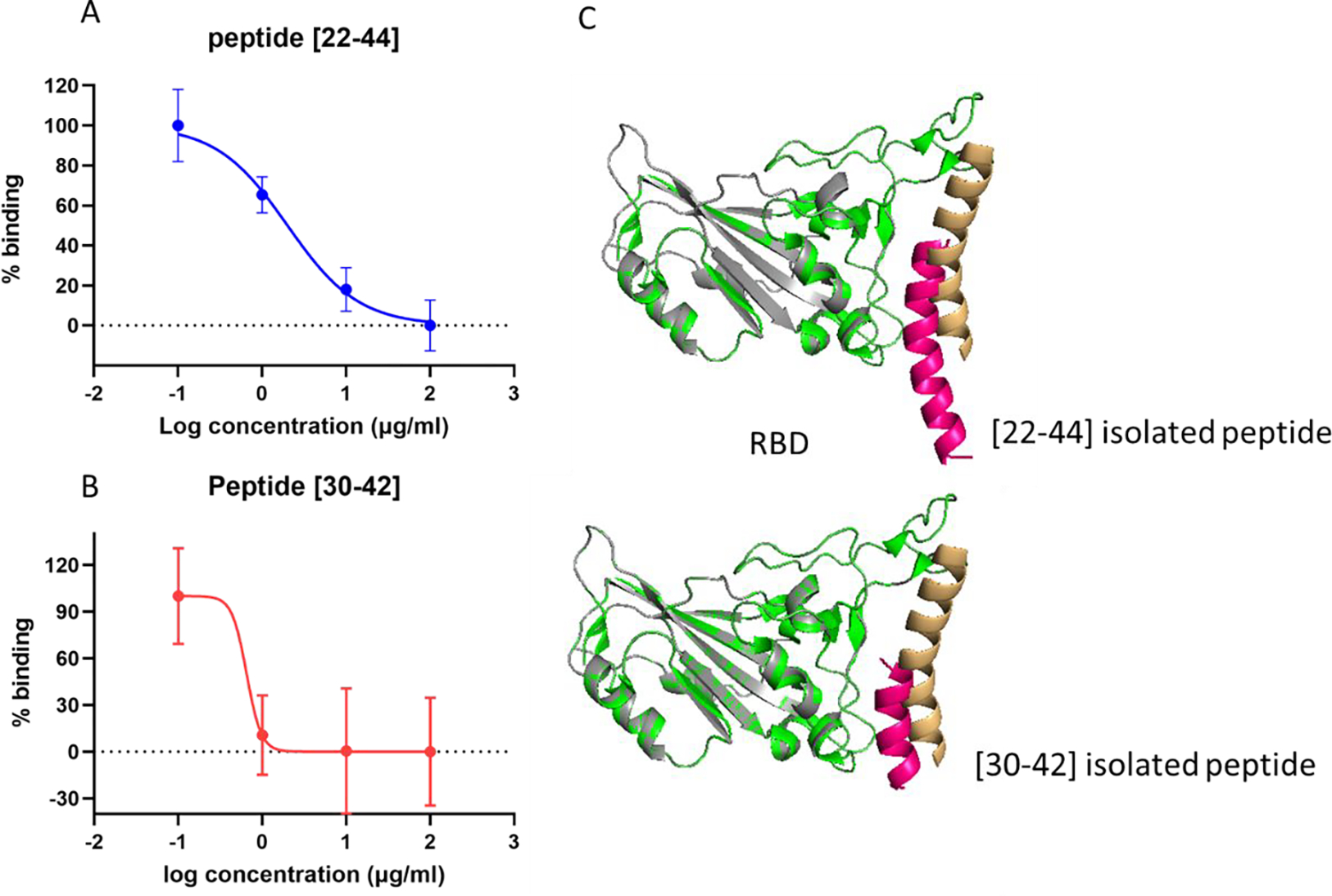
A) and B) Dose-dependent blocking of the S-protein with [22–44] and [30–42] peptides. C) structural alignment of both the docked result of the peptide/RBD complex and the α 1 helix/RBD complex, constructed from the ACE2/RBD complex (PDB ID 6m0j).
Molecular Docking of ACE2-derived peptide/RBD interaction
In order to evaluate and compare the two peptides as inhibitors of the ACE2/S1 complex, we combined molecular docking with the experimental study. Molecular docking was performed through the PatchDock server to study the peptides binding efficiency and to identify the amino acid residues that contribute most to the binding of the RBD/ACE2-derived peptide complex.
We evaluated the binding structure of the 23 and 13-amino acid peptides, [22–44] and [30–42] respectively, to the SARS-CoV-2 spike protein without the remainder of the ACE2 PD domain. To ensure a non-biased analysis, we performed a blind docking run where the binding site was not specified during the docking simulations. To analyze the obtained results, we compared the arrangement of each peptide within the SARS-CoV-2 spike protein binding site. For this, any RBD atom that fell within 3 A° of any peptide atom was considered to be a contact.
At the same time, we retrieved the SARS-CoV-2 spike protein and ACE2 PD domain crystal structure (PDB ID: 6M0J) and explored it as a further comparison point with the peptide models. The interfacial residues between the protein complex were viewed and interpreted using Pymol software. After a detailed analysis of interface residues, the α 1 helix, which occupies a concave groove formed by β5 and β6 sheets of the RBD, was found to provide the majority of the interaction between the ACE2/RBD interface (fig.4). Specifically, 8 residues (N487, K417, Q493, Y505, Y449, T500, N501, G446) in RBD provide contact via hydrogen bond with ACE2 (fig. 4A and 4B).
Figure 4.
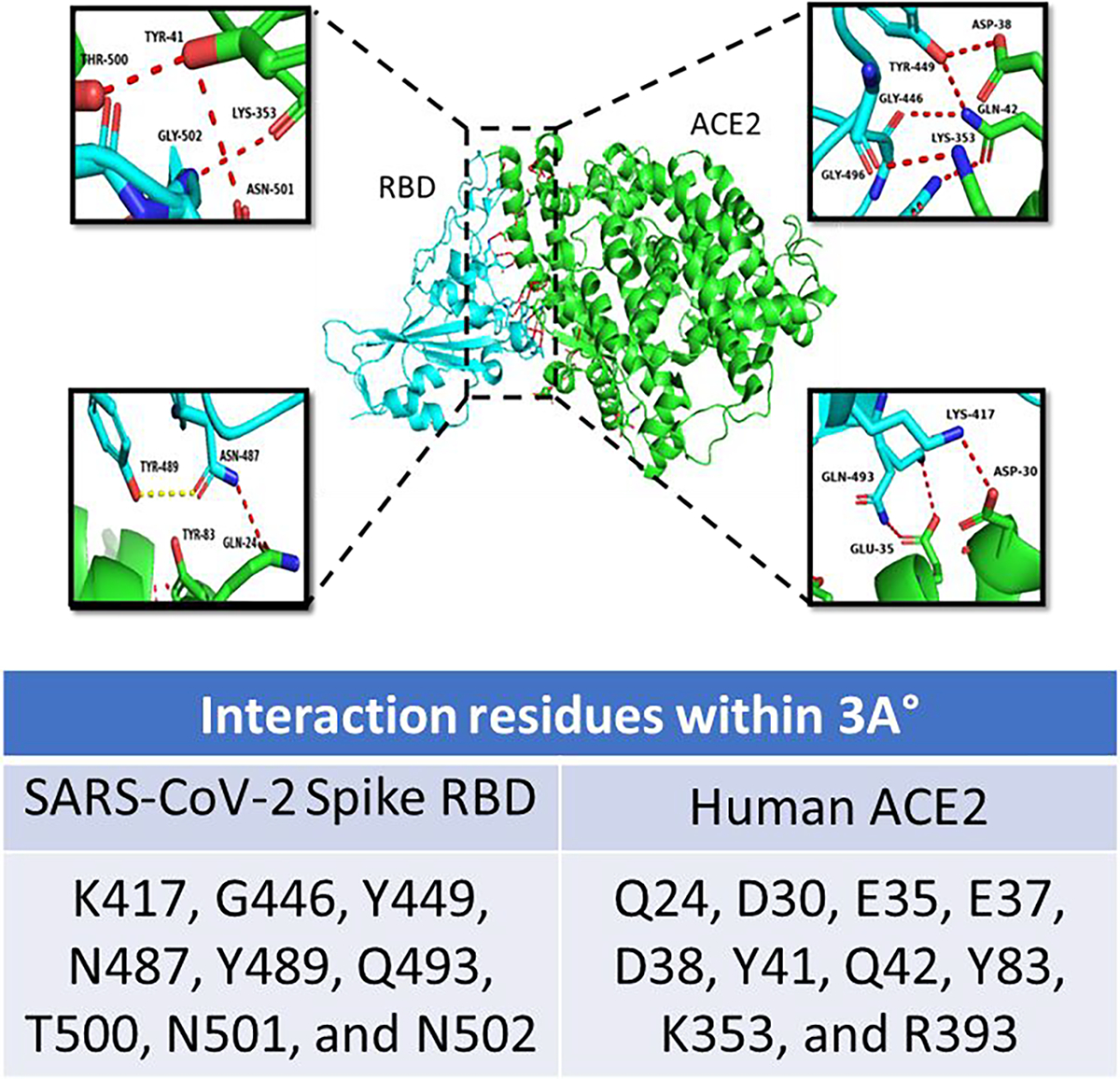
An illustration of the interacting interface of the SARS-CoV-2 receptor-binding domain (RBD) (cyan) and hACE2 (green) from PDB-ID: 6M0J. The key interacting residues are shown in close-up as insets. The table shows the interacting residues within a 3A° region analyzed using the PyMOL tool.
Figure. 3C shows the structural alignment of both the docked result of the peptide/RBD complex and the α 1 helix/RBD complex, which were constructed from the ACE2/RBD complex (PDB ID 6m0j). It is clear from the docking result that the [30–42] peptide binds to the concave groove of the RBD that the original ACE2 PD domain also binds. The [22–44] peptide laid on the RBD groove as well; however, only the central and the C-terminus of the peptide showed binding. This result aligns very well with the experimental results that the two peptides independently have the potential to inhibit the SARS-CoV-2 spike protein and ACE2 complex interaction but the shorter peptide results in more efficient inhibition.
By analyzing the docking results, the critical interacting amino acids of RBD/ACE2 were identified. Examination of Figures 5A and 5B presents that the RBD residues (449, 496, 493,494 500, 502, 505) were blocked by the [30–42] peptide. Similarly, the 35, 37, 38, and 41 residues of the [22–44] peptide have the ability to occupy the RBD residues (449, 498, 496, 494, 493, 502) by making 6 Hydrogen bonds within 3°A. From the PatchDock analysis it is clear that most of the RBD-interacting amino acidic residues were within the prominent binding sites. This finding was consistent with the crystal structure of the ACE2/RBD complex (fig. 4A and 4B). The docking position of the two peptides within the RBD pocket ensured a high possibility of blocking spike protein interactions from occurring with the ACE2 receptor, which is in alignment with our experimental results (fig. 2).
Figure 5.
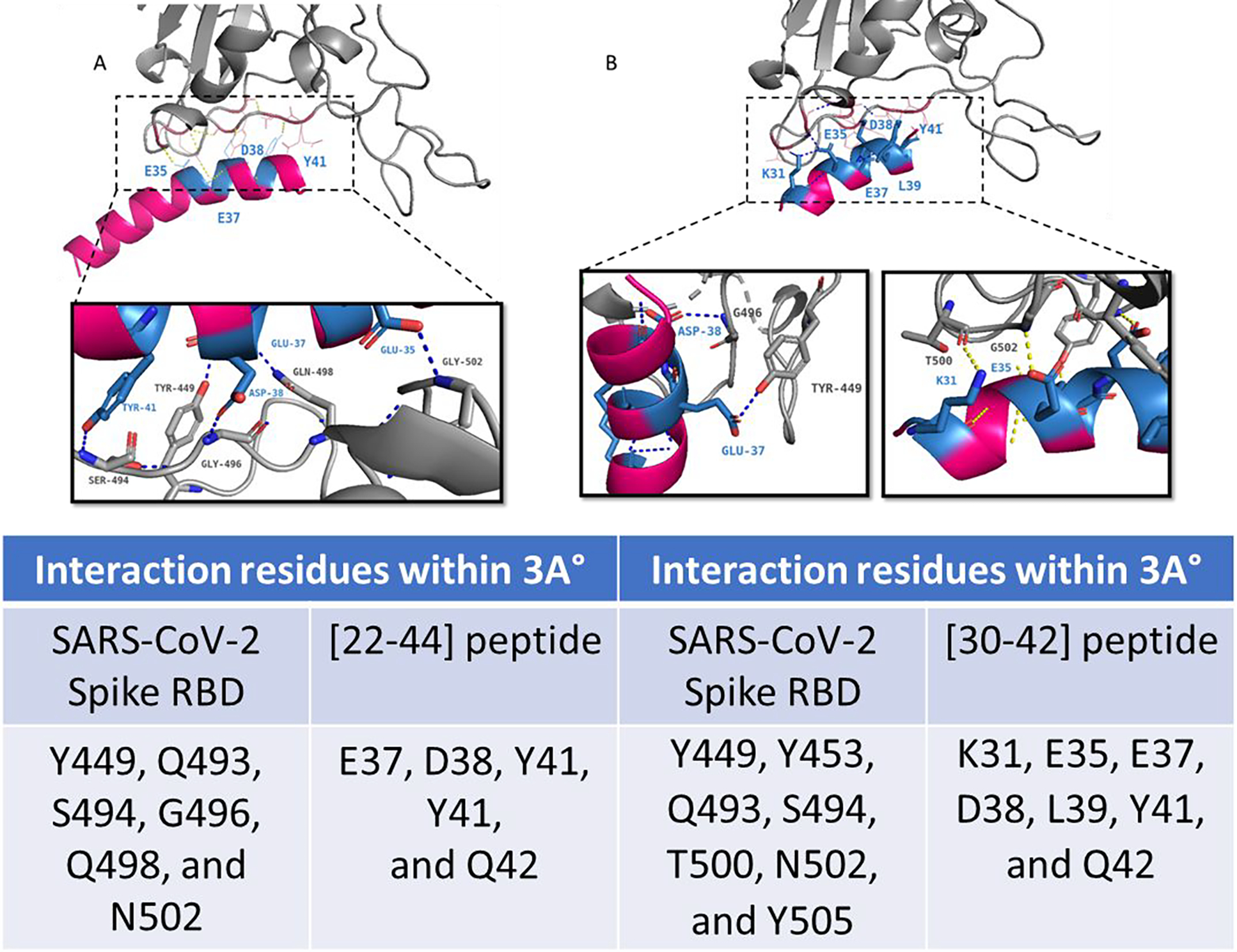
Interaction of the A) [22–44] peptide B) [30–42] peptide (pink) with the SARS-CoV-2 receptor-binding domain (RBD) (gray). Molecular docking complex obtained with PachDock. The key interacting residues are shown in close-up as insets. The table shows the interacting residues within a 3A° region analyzed using the PyMOL tool.
In addition, an inspection of the binding between each peptide and the RBD shows that fewer hydrogen bonds formed between the RBD/ peptide [22–44] complex compared with the RBD/peptide [30–42] complex, which makes 8 hydrogen bonds. This result is in line with our experimental results in which the [30–42] peptide shows higher inhibition of the ACE2/spike protein interaction for 0.1 to 1 μg/ml concentrations (fig. 2C) compared to the [22–44] peptide. It may be recalled that this result is in synergy with another study which suggests that the addition of extra amino acids does not necessarily increase the hydrogen bond or the binding efficiency of the peptide toward the spike protein.51 Combining this data together indicates that 7 residues blocked by the smaller peptide are crucial targets for blocking the ACE2/RBD binding interaction.
CONCLUSION:
Here we reported the fundamental investigation of two peptides’ abilities to inhibit the ACE/SARS-CoV-2 interaction. The two inhibitors of different sequence lengths based on the α 1 helix of ACE2 PD showed similar blocking efficiency, with the shorter peptide reaching maximal blocking efficiency at a lower concentration. This demonstrates the feasibility of targeting the ACE2/spike protein binding pocket with peptide-based inhibitors to inhibit viral infection. We observed a progressive reduction of the SPR binding signal as a function of peptide concentration confirming that specific inhibition was achieved. The [30–42] peptide, which is a truncated version of the longer peptide, highlights the importance of the amino acid residues at the central and C-terminus of the isolated α 1-helix of ACE2 for interaction with the spike protein. Moreover, we identified the critical residues of the RBD/ACE2 derived peptide interface using molecular docking, PatchDock. Analyzing the docking results revealed that the peptide inhibitors block most of the RBD residues that bind with ACE2, as predicted by analyzing the crystal structure of the ACE2/RBD complex. The results of our molecular docking and experimental inhibition assay were in alignment, indicating that small inhibitory peptides can effectively be used to block interactions between ACE/SARS-CoV-2 spike protein complex. However, substantial work will be necessary to ensure effectiveness of an inhibitory peptide in vivo such as introduction of D-amino acids at N- and C- terminal regions to reduce proteolytic degradation.52 This small peptide inhibition assay with SPR demonstrates its potential as a platform for screening potential small molecule and peptide inhibitors to aid in the future investigation of drug discovery and development focused on peptides. The presented information and approach can be used to gain a deeper understanding of the RBD/ACE2 binding interaction, as well as aid in the development of an anti-SARS-CoV-2 treatment of the viral infection without the adverse side effects that exist for many other small molecules or recombinant protein therapeutic avenues.
Acknowledgements
QC would like to acknowledge the financial support from NSF (CHE-2109042). MX would like to acknowledge the financial support from NIH (R35 GM138214).
REFERENCES
- 1.WHO COVID-19 Dashboard. Geneva: World Health Organization 2020: Available online: https://covid19.who.int/ Sept. 9th 2022. [Google Scholar]
- 2.Mahmud N; Anik MI; Hossain MK; Khan MI; Uddin S; Ashrafuzzaman M; Rahaman MM, Advances in Nanomaterial-Based Platforms to Combat COVID-19: Diagnostics, Preventions, Therapeutics, and Vaccine Developments. ACS Applied Bio Materials 2022. 5, 6, 2431–2460. [DOI] [PubMed] [Google Scholar]
- 3.Karmacharya M; Kumar S; Gulenko O; Cho Y-K, Advances in facemasks during the COVID-19 pandemic era. ACS Applied Bio Materials 2021, 4 (5), 3891–3908. [DOI] [PubMed] [Google Scholar]
- 4.Yadav S; Sadique MA; Ranjan P; Kumar N; Singhal A; Srivastava AK; Khan R, SERS Based Lateral Flow Immunoassay for Point-of-Care Detection of SARS-CoV-2 in Clinical Samples. ACS Appl Bio Mater 2021, 4 (4), 2974–2995. [DOI] [PubMed] [Google Scholar]
- 5.Pramanik A; Mayer J; Sinha SS; Sharma PC; Patibandla S; Gao Y; Corby LR; Bates JT; Bierdeman MA; Tandon R; Seshadri R; Ray PC, Human ACE2 Peptide-Attached Plasmonic-Magnetic Heterostructure for Magnetic Separation, Surface Enhanced Raman Spectroscopy Identification, and Inhibition of Different Variants of SARS-CoV-2 Infections. ACS Appl Bio Mater 2022. 5, 9, 4454–4464. [DOI] [PubMed] [Google Scholar]
- 6.Parihar A; Ranjan P; Sanghi SK; Srivastava AK; Khan R, Point-of-care biosensor-based diagnosis of COVID-19 holds promise to combat current and future pandemics. ACS applied bio materials 2020, 3 (11), 7326–7343. [DOI] [PubMed] [Google Scholar]
- 7.Kim J; Jeong S; Sarawut S; Kim H; Son SU; Lee S; Rabbani G; Kwon H; Lim E-K; Ahn S. N. J. L. o. a. C., An immunosensor based on a high performance dual-gate oxide semiconductor thin-film transistor for rapid detection of SARS-CoV-2. 2022, 22 (5), 899–907. [DOI] [PubMed] [Google Scholar]
- 8.Li WH; Moore MJ; Vasilieva N; Sui JH; Wong SK; Berne MA; Somasundaran M; Sullivan JL; Luzuriaga K; Greenough TC; Choe H; Farzan M, Angiotensin-converting enzyme 2 is a functional receptor for the SARS coronavirus. Nature 2003, 426 (6965), 450–454. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Li W; Zhang C; Sui J; Kuhn JH; Moore MJ; Luo S; Wong SK; Huang IC; Xu K; Vasilieva N; Murakami A; He Y; Marasco WA; Guan Y; Choe H; Farzan M, Receptor and viral determinants of SARS-coronavirus adaptation to human ACE2. EMBO J 2005, 24 (8), 1634–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Chen N; Zhou M; Dong X; Qu J; Gong F; Han Y; Qiu Y; Wang J; Liu Y; Wei Y; Xia J. a.; T. Yu; Zhang X; Zhang L, Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. The Lancet 2020, 395 (10223), 507–513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ke Z; Oton J; Qu K; Cortese M; Zila V; McKeane L; Nakane T; Zivanov J; Neufeldt CJ; Cerikan B; Lu JM; Peukes J; Xiong X; Krausslich HG; Scheres SHW; Bartenschlager R; Briggs JAG, Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588 (7838), 498–502. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Walls AC; Park YJ; Tortorici MA; Wall A; McGuire AT; Veesler D, Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181 (2), 281–292 e6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Xiong X; Qu K; Ciazynska KA; Hosmillo M; Carter AP; Ebrahimi S; Ke Z; Scheres SHW; Bergamaschi L; Grice GL; Zhang Y; Collaboration C-NC-B; Nathan JA; Baker S; James LC; Baxendale HE; Goodfellow I; Doffinger R; Briggs JAG, A thermostable, closed SARS-CoV-2 spike protein trimer. Nat Struct Mol Biol 2020, 27 (10), 934–941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gui M; Song W; Zhou H; Xu J; Chen S; Xiang Y; Wang X, Cryo-electron microscopy structures of the SARS-CoV spike glycoprotein reveal a prerequisite conformational state for receptor binding. Cell Res 2017, 27 (1), 119–129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Song W; Gui M; Wang X; Xiang Y, Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog 2018, 14 (8), e1007236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kirchdoerfer RN; Wang N; Pallesen J; Wrapp D; Turner HL; Cottrell CA; Corbett KS; Graham BS; McLellan JS; Ward AB, Stabilized coronavirus spikes are resistant to conformational changes induced by receptor recognition or proteolysis. Sci Rep 2018, 8 (1), 15701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Zamzami MA; Rabbani G; Ahmad A; Basalah AA; Al-Sabban WH; Ahn SN; Choudhry H, Carbon nanotube field-effect transistor (CNT-FET)-based biosensor for rapid detection of SARS-CoV-2 (COVID-19) surface spike protein S1. Bioelectrochemistry 2022, 143, 107982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Asandei A; Mereuta L; Schiopu I; Park J; Seo CH; Park Y; Luchian T, Non-receptor-mediated lipid membrane permeabilization by the SARS-CoV-2 spike protein S1 subunit. J ACS Applied Materials Interfaces 2020, 12 (50), 55649–55658. [DOI] [PubMed] [Google Scholar]
- 19.Biancatelli RMLC; Solopov PA; Gregory B; Khodour Y; Catravas JD, HSP90 Inhibitors Modulate SARS-CoV-2 Spike Protein Subunit 1-Induced Human Pulmonary Microvascular Endothelial Activation and Barrier Dysfunction. J Frontiers in Physiology 2022, 13:812199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Wang Q; Zhang Y; Wu L; Niu S; Song C; Zhang Z; Lu G; Qiao C; Hu Y; Yuen KY; Wang Q; Zhou H; Yan J; Qi J, Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181 (4), 894–904 e9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shang J; Ye G; Shi K; Wan Y; Luo C; Aihara H; Geng Q; Auerbach A; Li F, Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581 (7807), 221–224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lan J; Ge J; Yu J; Shan S; Zhou H; Fan S; Zhang Q; Shi X; Wang Q; Zhang L; Wang X, Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581 (7807), 215–220. [DOI] [PubMed] [Google Scholar]
- 23.Daniel Wrapp NW, Kizzmekia S Corbett Jory A. Goldsmith, Ching-Lin Hsieh, Olubukola Abiona, Barney S. Graham, Jason S. McLellan, Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Renhong Yan YZ, Yaning Li, Lu Xia, Yingying Guo, Qiang Zhou, Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Smith MC; Gestwicki JE, Features of protein-protein interactions that translate into potent inhibitors: topology, surface area and affinity. Expert Rev Mol Med 2012, 14, e16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Josephson K; Ricardo A; Szostak JW, mRNA display: from basic principles to macrocycle drug discovery. Drug Discov Today 2014, 19 (4), 388–99. [DOI] [PubMed] [Google Scholar]
- 27.Tai W; He L; Zhang X; Pu J; Voronin D; Jiang S; Zhou Y; Du L, Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol 2020, 17 (6), 613–620. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhe Lv Y-QD, Qing Ye, Lei Cao, Chun-Yun Sun, Changfa Fan, Weijin Huang, Shihui Sun, Yao Sun, Zhu Ling, Qi Chen, Nan Wang, Jianhui Nie, Zhen Cui, Dandan Zhu, Neil Shaw, Xiao-Feng Li, Qianqian Li, Liangzhi Xie, Youchun Wang, Zihe Rao, Cheng-Feng Qin, Xiangxi Wang, Structural basis for neutralization of SARS-CoV-2 and SARS-CoV by a potent therapeutic antibody. Science 2020, 369, 1505–1509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sui J; Deming M; Rockx B; Liddington RC; Zhu QK; Baric RS; Marasco WA, Effects of human anti-spike protein receptor binding domain antibodies on severe acute respiratory syndrome coronavirus neutralization escape and fitness. J Virol 2014, 88 (23), 13769–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang C; Li W; Drabek D; Okba NMA; van Haperen R; Osterhaus A; van Kuppeveld FJM; Haagmans BL; Grosveld F; Bosch BJ, A human monoclonal antibody blocking SARS-CoV-2 infection. Nat Commun 2020, 11 (1), 2251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Alina Baum BOF, Elzbieta Wloga, Richard Copin, Pascal Kristen E., Vincenzo Russo, Stephanie Giordano, Kathryn Lanza, Nicole Negron, Min Ni, Yi Wei, Atwal Gurinder S., Andrew J. Murphy, Neil Stahl, George D. Yancopoulos, Christos A Kyratsous, Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies. Science 2020, 369, 1014–1018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Wibmer CK; Ayres F; Hermanus T; Madzivhandila M; Kgagudi P; Oosthuysen B; Lambson BE; de Oliveira T; Vermeulen M; van der Berg K; Rossouw T; Boswell M; Ueckermann V; Meiring S; von Gottberg A; Cohen C; Morris L; Bhiman JN; Moore PL, SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. Nat Med 2021, 27 (4), 622–625. [DOI] [PubMed] [Google Scholar]
- 33.Hamley IW, Peptides for Vaccine Development. ACS Appl Bio Mater 2022, 5 (3), 905–944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Benjamin Leader Q. J. B. a. D. E. G., Protein therapeutics: a summary and pharmacological classification. Naturere reviews | drug discovery 2008, 7, 21–39. [DOI] [PubMed] [Google Scholar]
- 35.Rabbani G; Ahn SN; Kwon H; Ahmad K; Choi I, Penta-peptide ATN-161 based neutralization mechanism of SARS-CoV-2 spike protein. Biochemistry and Biophysics Reports 2021, 28, 101170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Saurabh S; Purohit SS, A Modified ACE2 peptide mimic to block SARS-CoV2 entry. bioRxiv 2020. [Google Scholar]
- 37.Renzi F; Ghersi D, ACE2 fragment as a decoy for novel SARS-Cov-2 virus. bioRxiv 2020, 1–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Zhang H; Penninger JM; Li Y; Zhong N; Slutsky AS, Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Med 2020, 46 (4), 586–590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Monteil V; Kwon H; Prado P; Hagelkruys A; Wimmer RA; Stahl M; Leopoldi A; Garreta E; Hurtado Del Pozo C; Prosper F; Romero JP; Wirnsberger G; Zhang H; Slutsky AS; Conder R; Montserrat N; Mirazimi A; Penninger JM, Inhibition of SARS-CoV-2 Infections in Engineered Human Tissues Using Clinical-Grade Soluble Human ACE2. Cell 2020, 181 (4), 905–913 e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Rabbani G; Ahn SN, Roles of human serum albumin in prediction, diagnoses and treatment of COVID-19. International Journal of Biological Macromolecules 2021, 193, 948–955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Cao B; Wang Y; Wen D; Liu W; Wang J; Fan G; Ruan L; Song B; Cai Y; Wei M; Li X; Xia J; Chen N; Xiang J; Yu T; Bai T; Xie X; Zhang L; Li C; Yuan Y; Chen H; Li H; Huang H; Tu S; Gong F; Liu Y; Wei Y; Dong C; Zhou F; Gu X; Xu J; Liu Z; Zhang Y; Li H; Shang L; Wang K; Li K; Zhou X; Dong X; Qu Z; Lu S; Hu X; Ruan S; Luo S; Wu J; Peng L; Cheng F; Pan L; Zou J; Jia C; Wang J; Liu X; Wang S; Wu X; Ge Q; He J; Zhan H; Qiu F; Guo L; Huang C; Jaki T; Hayden FG; Horby PW; Zhang D; Wang C, A Trial of Lopinavir-Ritonavir in Adults Hospitalized with Severe Covid-19. N Engl J Med 2020, 382 (19), 1787–1799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hinman SS; Ruiz CJ; Drakakaki G; Wilkop TE; Cheng Q, On-Demand Formation of Supported Lipid Membrane Arrays by Trehalose-Assisted Vesicle Delivery for SPR Imaging. ACS Appl Mater Interfaces 2015, 7 (31), 17122–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Schneidman-Duhovny D; Inbar Y; Nussinov R; Wolfson HJ, PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res 2005, 33 (Web Server issue), W363–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zhang G; Pomplun S; Loftis AR; Tan X; Loas A; Pentelute BL, Investigation of ACE2 N-terminal fragments binding to SARS-CoV-2 Spike RBD. bioRxiv 2020. [Google Scholar]
- 45.Chang CC, Recent Advancements in Aptamer-Based Surface Plasmon Resonance Biosensing Strategies. Biosensors (Basel) 2021, 11 (7), 233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Azzouz A; Hejji L; Kim KH; Kukkar D; Souhail B; Bhardwaj N; Brown RJC; Zhang W, Advances in surface plasmon resonance-based biosensor technologies for cancer biomarker detection. Biosens Bioelectron 2022, 197, 113767. [DOI] [PubMed] [Google Scholar]
- 47.Wang W; Thiemann S; Chen Q, Utility of SPR technology in biotherapeutic development: Qualification for intended use. Anal Biochem 2022, 654, 114804. [DOI] [PubMed] [Google Scholar]
- 48.Harbour V; Casillas C; Siddiqui Z; Sarkar B; Sanyal S; Nguyen P; Kim KK; Roy A; Iglesias-Montoro P; Patel S, Regulation of lipoprotein homeostasis by self-assembling peptides. ACS Applied Bio Materials. 2020, 3 (12), 8978–8988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Schuck P; Zhao H, The role of mass transport limitation and surface heterogeneity in the biophysical characterization of macromolecular binding processes by SPR biosensing. Methods Mol Biol 2010, 627, 15–54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Larue RC; Xing E; Kenney AD; Zhang Y; Tuazon JA; Li J; Yount JS; Li PK; Sharma A, Rationally Designed ACE2-Derived Peptides Inhibit SARS-CoV-2. Bioconjug Chem 2021, 32 (1), 215–223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yang J; Petitjean SJL; Koehler M; Zhang Q; Dumitru AC; Chen W; Derclaye S; Vincent SP; Soumillion P; Alsteens D, Molecular interaction and inhibition of SARS-CoV-2 binding to the ACE2 receptor. Nat Commun 2020, 11 (1), 4541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bruno BJ; Miller GD; Lim CS, Basics and recent advances in peptide and protein drug delivery. Ther Deliv 2013, 4 (11), 1443–67. [DOI] [PMC free article] [PubMed] [Google Scholar]


