Abstract
Objective
To profile clinically non‐aggressive and aggressive pituitary adenomas (PAs)/pituitary neuroendocrine tumours (PitNETs) and pituitary carcinomas for somatic mutations and epigenetic alterations of genes involved in cell proliferation/differentiation, microRNAs (miRNA)/long noncoding RNA (LncRNA)‐post‐transcriptional regulators and therapy targets.
Design
Retrospective observational study.
Patients and Measurements
A total of 64 non‐aggressive and 41 aggressive PAs/PitNETs and 6 pituitary carcinomas treated by endoscopic surgery with ≥1‐year follow‐up were included. Somatic mutations of 17 genes and DNA methylation of 22 genes were assessed. Ten normal pituitaries were used as control.
Results
We found at least one mutation in 17 tumours, including 6/64 non‐aggressive, 10/41 aggressive PAs/PitNETs, and 1/6 pituitary carcinoma. AIP (N = 6) was the most frequently mutated gene, followed by NOTCH (4), and TP53 (3). Hypermethylation of PARP15, LINC00599, ZAP70 was more common in aggressive than non‐aggressive PAs/PITNETs (p < .05). Lower levels of methylation of AIP, GNAS and PDCD1 were detected in aggressive PAs/PITNETs than non‐aggressive ones (p < .05). For X‐linked genes, males presented higher level of methylation of FLNA, UXT and MAGE family (MAGEA11, MAGEA1, MAGEC2) genes in aggressive vs. non‐aggressive PAs/PITNETs (p < .05).
In pituitary carcinomas, methylation of autosomal genes PARP15, LINC00599, MIR193 and ZAP70 was higher than in PAs/PITNETs, while X‐linked genes methylation level was lower.
Conclusions
Somatic mutations and methylation levels of genes involved in cell proliferation/differentiation, miRNA/LncRNA‐post‐transcriptional regulators and targets of antineoplastic therapies are different in non‐aggressive and in aggressive PAs/PitNETs. Methylation profile also varies according to gender. Combined genetic‐epigenetic analysis, in association with clinico‐radiological‐pathological data, may be of help in predicting PA/PitNET behaviour.
Keywords: adenoma, methylation profile, pituitary neuroendocrine tumours, prognosis, recurrence, somatic mutation
1. INTRODUCTION
Tumours arising from hormone‐secreting adenohypophyseal cells are a complex and heterogeneous group of lesions, with distinctive radiological, histological, and clinical features. Since the majority are benign and respond to surgery and/or medical therapy they have been traditionally termed ‘pituitary adenomas’ (PAs). 1 , 2
A small subset of PAs does not respond to standard treatments and presents with multiple local recurrences (aggressive pituitary tumours), and rarely, with metastases (pituitary carcinomas). 1 , 2 In 2018, The European Society of Endocrinology (ESE) provided the definition of aggressive pituitary tumours (i.e., radiological invasiveness and/or unusually rapid tumour growth rate, or clinically relevant tumour growth despite optimal standard surgery, radiation and medical therapies), and clinical guidelines for the diagnosis and management of aggressive pituitary tumours and carcinomas. 3
As per the other neuroendocrine neoplasms, the term pituitary neuroendocrine tumour (PitNET) has been placed alongside PA, resulting in the combined designation PA/PitNET. 4 , 5 PA/PitNET classification is primarily based on histological criteria and lineage differentiation, 6 and clinico‐radiological features. 2 , 3 A variety of somatic mutations and epigenetic alterations has been reported in PAs/PitNETs (see Tables S1 and S2), although their prevalence and significance are still debated. 7 , 8 , 9 , 10
Our study attempts to characterise surgically treated non‐aggressive and aggressive PAs/PitNETs, and pituitary carcinomas for somatic mutations and epigenetic alterations of selected autosomal and X‐linked genes.
2. MATERIALS AND METHODS
2.1. Patient selection
Adult patients with PAs/PitNETs of different histological subtypes, operated via endoscopic endonasal approach from 2003 to 2020 at the Pituitary Unit of the IRCCS Istituto delle Scienze Neurologiche di Bologna (Italy), for which clinico‐radiological and biochemical data of interest at diagnosis and follow‐up were available, were included provided (1) there was no history of prior surgery or radiation therapy; (b) there was no evidence of residual tumour in the 3‐month post‐surgical magnetic resonance imaging (MRI); and (c) post‐surgical follow‐up was ≥1 year. Patients who had received medical therapy before surgery were included. Fresh frozen and/or formalin‐fixed paraffin‐embedded (FFPE) tumour tissue of adequate size and quality to perform morphologic, immunohistochemical (IHC) and molecular characterisation were required.
According to preoperative MRI, PAs/PitNETs were classified as ‘micro‐’ (<10 mm) and ‘macro’ (≥10 mm). Histological subtype was defined according to the World Health Organization 2017 classification of endocrine tumours. 6 Tumour removal was considered ‘radical’ in the absence of a visible remnant at the MRI performed 3 months after surgery. Disease ‘cure’ was defined by radical tumour resection in non‐functioning PAs/PitNETs, and by its association with normalised hormone secretion in functioning tumours. Disease ‘recurrence’ corresponded to MRI tumour detection and/or hormone hypersecretion at follow‐up, in previously cured patients. The status of hormone secretion was defined according to the criteria reported in international consensus/guidelines for each tumour subtype. According to 2018 ESE guidelines, 3 pituitary tumours were divided in non‐aggressive pituitary tumours, aggressive pituitary tumours—presenting with size >10 mm, extra‐sellar invasion (defined by Knosp grades 3–4/Hardy‐Wilson stages D–E), high proliferation (ki 67 ≥3% and p53 > 10 strongly positive nuclei/10 HPF or the presence of >2/10 mitoses/HPF), and requiring multiple treatment to obtain disease remission or, at least, control—, and pituitary carcinoma, defined by the presence of craniospinal and/or systemic metastases. Six pituitary carcinomas—three diagnosed at the Pituitary Unit of the IRCCS Istituto delle Scienze Neurologiche di Bologna (Italy) and three at the University of Wisconsin (Madison, WI, USA)—, for which FFPE tissue of the pituitary and extra‐pituitary localisations was available, were also included.
2.2. Tumour sample collection and characterisation
All the original tumour slides were reviewed, and the diagnosis was confirmed independently by two pathologists (SA and AR). IHC staining for anterior pituitary hormones (ACTH, PRL, GH, FSH, LH and TSH), cytokeratin CAM5.2, Ki67 (MIB1) and p53, performed at the time of diagnosis, were reviewed. In hormone‐negative samples, transcription factors (SF1, T‐PIT and Pit‐1) 11 were performed using a fully automated IHC stainer (BenchMark XT, Roche Ventana Medical System Inc.). Main antibodies features are reported in Table S3.
All surgical FFPE specimens were cut for additional study. When available, fresh tumour samples were also analyzed. The pathologist selected the most representative tumour fragments for size and quality (i.e., maximum representation of pituitary cells and lowest portions of extra‐pituitary tissues and necrosis), thus selecting high quality cases for molecular analysis. 12 Ten normal pituitaries (seven from male and three from female patients) obtained to autopsies were used as reference control.
2.3. Mutational analysis
MasterPure Complete DNA extraction kit (Epicentre, code MC85200) and QuickExtract™ FFPE DNA extraction solution (Lucigen) were used to purify DNA from fresh frozen and FFPE tissues, respectively. Mutational analysis was performed using a next generation sequencing (NGS) approach for the following 17 genes: KRAS, BRAF, EGFR, TP53, NOTCH1, PIK3CA, CDKN2A, TERT, NRAS, HRAS, PTEN, FBXW7, EP300, CASP8, AIP, GNAS and USP8, known to be involved in the development of pituitary tumours (see Table S1). 13 Target enrichment of hot spot regions and generation of primer libraries was performed following previously reported protocols and methods. 14 FASTQ files, obtained after loading libraries onto MiSEQ (Illumina), were filtered (PHRED quality score >Q30; length >100 bp) and processed in a Galaxy Project environment, 15 using hg38 human reference genome with Bowtie2, GATK local realignment, HaplotypeCaller and Picard MarkDuplicates. Mutations were visualised using BAM files loaded onto the Integrative Genomic Viewer. Only mutations with a variant allele frequency (VAF) threshold >20% and a coverage depth of 50x in both strands were reported. The pathogenic score was calculated on Polyphen‐2 web tool (http://genetics.bwh.harvard.edu/pph2/), the in silico predictor of functional effects of gene variants, and verified on COSMIC database. 16 Each NGSs experiment was designed to allocate ≥1k reads/region, to obtain a depth of coverage ≥1000x.
An exploratory analysis was performed on 25 fresh frozen samples and repeated on FFPE samples of the same tumours, confirming the same mutation profile. Therefore, 86 FFPE (81 PAs/PitNETs and 6 carcinomas) samples were considered in the subsequent analyses.
2.4. DNA methylation analysis
Bisulfite treatment of genomic DNA (50–500 ng) was performed using the EZ DNA Methylation‐Lightning Kit (Zymo Research Europe) according to the manufacturer's protocol. DNA methylation was evaluated using targeted bisulfite NGS in 16 target genes mapped on autosomes (i.e., MIR137HG, LRRTM1, ZAP70, ITGA4, KIF1A, PDCD1, PARP15, TERT, Linc00599, AIP, FLI1, NTM, CDH1, MIR193a, GNAS, GP1BB), and 6 genes mapped on chromosome X (i.e., UXT, HDAC6, MAGEC2, MAGEA11, MAGEA1, FLNA), for which altered methylation had been previously reported in various types of tumour (see Table S2).
Genomic sequences stored in the Ensembl genome browser (http://www.ensembl.org/index.html) were employed as query sequences to identify putative CpG islands in gene promoter regions. MethPrimer (http://www.urogene.org/cgi-bin/methprimer/methprimer.cgi) designing was applied to identify CpGs and the best primers of choice. 17 Overall, 460 CpG were analyzed quantitatively by bisulfite sequencing. The list of genomic regions, primer sequences and interrogated mapping coordinates are reported in Table S4. Locus‐specific bisulfite amplicon libraries were generated with tagged primers using Phusion U DNA polymerase (Thermo Fisher Scientific, cod. F555L). To prepare libraries, a first multiplex PCR amplification for target enrichment and a second round of amplification for the barcoding were performed, using the Nextera index kit, as previously described. 18 Sequencing was conducted on MiSeq sequencer (Illumina), according to the manufacturer's protocol. Each NGS experiment was designed to allocate ≥1k reads/region, to obtain a depth of coverage ≥1000x.
Methylation ratio for each CpG was calculated by BWAmeth in a Galaxy Project environment followed by the MethylDackel tool and, in parallel, by EPIC‐TABSAT. 19
As per mutation analysis, an exploratory methylation profile was performed on 25 fresh frozen samples and repeated on FFPE samples of the same tumours were assessed, showing similar methylation profile. Therefore, 86 FFPE (81 PAs/PitNETs and 6 carcinomas) samples were included in subsequent analyses.
For recurrent pituitary tumours, mutation and methylation analyses were performed on the specimens retrieved at the first endoscopic surgery, while for carcinomas, analyses were performed on pituitary tumour.
2.5. Statistical analysis
Mutational and DNA methylation differences among non‐aggressive and aggressive PAs/PitNETs and carcinomas were determined by Fisher's exact test. For all tests, significance was defined as p < .05. Kruskal–Wallis test was used to define the informative CpGs for each group of genes to which they belong. Statistical analysis was performed using IBM SPSS version 23.0 and Stata 12.
3. RESULTS
3.1. Patients features
A total of 111 patients (59 M:52 F), 105 with PA/PitNET and 6 with pituitary carcinoma, were included in the study. A total of 41 cases (39%) were aggressive PAs/PitNETs. The PA/PitNET group included 49 FSH‐/LH‐, 22 ACTH, 19 GH‐ (8 sparsely and 8 densely granulated, 3 mixed GH‐/PRL‐), 12 PRL‐secreting, 1 null cell and 2 plurihormonal PIT‐1‐positive tumours. Carcinomas included 3 ACTH‐, 2 PRL‐ and 1 FSH/LH‐secreting tumours. Overall, 52 tumours (48 PAs/PitNETs and 4 carcinomas) were functioning, while 59 (57 PAs/PitNETs and 2 carcinomas) were non‐functioning (Table 1). Clinico‐radiological and IHC data of 33 patients with PAs/PitNETs had been previously reported. 11
Table 1.
Main clinical, radiological and histological patient data at first surgery
| Total | PA/PitNETs | Carcinoma | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Non‐aggressive | Aggressive | |||||||||||
| Number of cases | 111 | 64 | 41 | 6 | ||||||||
| Gender (F:M) | 53:57 | 31:33 | 18:23 | 3:3 | ||||||||
| Age at surgery (mean ± SD; years) | 51.5 ± 15.7 | 54.5 ± 15.3 | 48.4 ± 15.3 | 54.7 ± 7.3 | ||||||||
| Size (micro; macro) | 21:90 | 21:43 | 0:41 | 0:6 | ||||||||
| Pre‐surgical medical therapy | 24 | 5 | 15 | 4 | ||||||||
| 4 ACTH‐; 1 FSH/LH‐; 8 GH‐; 11 PRL‐secreting) | (1 ACTH‐; 2 GH‐; 2 PRL‐secreting) | (1 ACTH‐; 1 FSH/LH‐; 6 GH‐; 7 PRL‐secreting) | (2 ACTH‐; 2 PRL‐secreting) | |||||||||
| Tumour type (total; F:M; functioning) | ||||||||||||
| Corticotroph | 25 | 15:10 | 15 | 13 | 9:4 | 10 | 9 | 4:5 | 5 | 3 | 2:1 | 2 |
| Somatotroph | 19 | 14:5 | 19 | 8 | 5:3 | 8 | 11 | 9:2 | 11 | – | – | – |
| Lactotroph | 14 | 6:8 | 14 | 3 | 1:2 | 3 | 9 | 4:5 | 9 | 2 | 1:1 | 2 |
| Gonadotroph | 50 | 18:32 | 2 | 38 | 16:22 | 1 | 11 | 1:10 | 1 | 1 | 1:0 | – |
| Null cell | 1 | 0:1 | – | – | – | – | 1 | 0:1 | – | – | – | – |
| Plurihormonal Pit1+ | 2 | 0:2 | – | 2 | 0:2 | – | – | – | – | – | – | – |
| Post‐surgical treatment | ||||||||||||
| Endoscopic pituitary surgery | 25 | 4 | 15 | 6 | ||||||||
| Medical therapy | 27 | 3 | 23 | 1 | ||||||||
| Radiation therapy | 9 | – | 7 | 2 | ||||||||
| ADRENALEctomya | 2 | – | 2 | – | ||||||||
Abbreviations: PA, pituitary adenoma; PitNET, pituitary neuroendocrine tumour.
% Has been calculated over the 21 corticotroph PAs/PitNETs.
Mean age at first surgery was 52.1 ± 16.1 years in patients with PAs/PitNETs, and 54.5 ± 7.3 years for those with carcinomas. Follow‐up was 49.8 ± 31.7 months for PAs/PitNETs and 46.3 ± 21.5 months for carcinomas (Table 1).
Overall, 24 (of 111; 21.6%) patients, of whom 5 (of 64; 7.8%; 1 ACTH‐; 2 GH‐; 2 PRL‐secreting) with non‐aggressive, and 15 (of 41) with aggressive (36.6%; 1 ACTH‐; 1 FSH/LH‐; 6 GH‐; 7 PRL‐secreting) PAs/PitNETs, and 4 (of 6; 66.7%) with pituitary carcinomas had received medical therapy before surgery. The rate of patients who had received medical therapy was higher in carcinomas than in PAs/PitNETs (p = .011), and in aggressive vs. non‐aggressive PAs/PitNETs (p = .0002); the distribution for histotype was similar among groups.
No significant differences were detected in terms of gender, age at diagnosis and duration of follow‐up between non‐aggressive and aggressive PAs/PitNETs, and carcinomas.
Data on plurihormonal Pit1‐positive and null cells tumours could not be further analyzed since they were too few (Table 1).
Patients with pituitary carcinomas presented intracranial/parenchymal (n = 3), spinal/intradural lumbar (n = 2) and hepatic (n = 1) metastases.
3.2. Mutational analysis
A total of 17 out of 111 (15.3%) tumours presented a pathogenic mutation, including 6/64 (9.4%) non‐aggressive and 9/41 (21.9%) aggressive PAs/PitNETs, and 1/6 (16.6%) pituitary carcinoma.
Mutations mainly involved AIP (N = 6), NOTCH (N = 4) and TP53 (N = 3), that also presented the highest levels of mutation frequency. VAF was >70% in all the 3 cases with TP53 mutation, close to 100% in two, corresponding to a complete loss of function due to a missense mutation in one allele and probable deletion in the other. The same was true for 2 tumours with AIP mutation with a VAF close to 100%; the other 2 cases with pathogenic mutations displayed a VAF of 51%, corresponding to the loss of function of a single. Finally, the AIP mutation p.Q228K presented a VAF of 100%, and gave a PolyPhen‐2 score of 0.03, indicating a benign and germline mutation.
Three pathogenic mutations affected NOTHC1 with a VAF close to 30%, indicating the involvement of a single allele of the main clone of the tumour.
Mutated non‐aggressive PAs/PitNETs included 2 FSH‐/LH‐, 2 ACTH‐, 1 PRL‐secreting and 1 plurihormonal PIT‐1‐positive PAs/PitNETs; 2 (1 ACTH‐ and 1 PRL‐secreting) were functioning, while the other were non‐functioning; 5 were micro‐ while 1 was a macroadenoma. Four patients were females and 2 were males. Mean age at surgery was 49 years old (range: 36–66). Both USP8 mutated tumours were densely granulated ACTH‐secreting PAs/PitNETs.
Mutated aggressive PA/PitNETs were macroadenomas and included three sparsely granulated ACTH‐, 3 sparsely granulated GH‐, two FSH/LH‐ and two PRL‐secreting tumours. Five (two PRL‐secreting and three sparsely granulated GH‐secreting) were functioning, while five were non‐functioning (three sparsely granulated ACTH‐secreting and two FSH‐/LH‐secreting). Mean age at surgery was 42 years old (range: 1–62). The group included five females and five males.
Two pathogenic AIP and CDKN2A mutations were identified in the same lactotroph pituitary carcinoma, affecting a 59‐year‐old man.
Details on the type, VAF, genetic score, tumour histological type, tumour granulation (for ACTH‐ and GH‐secreting tumours), and distribution among non‐aggressive and aggressive PAs/PitNETs and carcinoma of each identified gene mutation are reported in Table 2.
Table 2.
Prevalence, VAF and pathogenic index of mutations identified in TP53, NOTCH, EGFR, AIP, USP8, KRAS and PIKI3 genes in non‐aggressive PAs/PitNETs, aggressive PAs/PitNETs and pituitary carcinomas
| Gene | Non‐aggressive PAs/PitNETs | Aggressive PAs/PitNETs | Pituitary carcinomas | Total (pathogenic) |
|---|---|---|---|---|
| TP53 | 0 | 3 | 0 | 3 |
| p.L114S$; VAF 74%; pathogenic (1.0); SG ACTH | ||||
| p.R196*; VAF 100%; pathogenic (0.96); SG GH | ||||
| p.P191L; VAF 100%; pathogenic (0.99); SG GH | ||||
| NOTCH1 | 3 | 1 | 0 | 3 |
| p.H1601Y; VAF 25%; pathogenic (0.99); FSH/LH | ||||
| p.R1608H; VAF 38%; pathogenic (0.99); Pit1+ | p.V1605I; VAF 25%; benign (0.036); SG ACTH | |||
| p.L1596F; VAF 24%; pathogenic (0.9); PRL | ||||
| FBXW7 | 0 | 1 | 0 | 1 |
| p.C453Y§; VAF 68%; pathogenic (1.0); FSH/LH | ||||
| PIK3CA | 0 | 2 | 0 | 1 |
| p.S535P; VAF 30%; pathogenic (0.99); SG ACTH | ||||
| p.Q1064§; VAF 28%; benign (0.021); FSH/LH | ||||
| CASP8 | 0 | 1 | 0 | 1 |
| p.R248V; VAF 28%; pathogenic (0.93); PRL | ||||
| GNAS | 0 | 1 | 0 | 1 |
| p.D858N; VAF 26%; pathogenic (0.99); SG GH | ||||
| USP8 | 2 | 0 | 0 | 2 |
| p.P690R; VAF 35% pathogenic (0.92); DG ACTH | ||||
| p.S718P; VAF 38% pathogenic (0.99); DG ACTH | ||||
| AIP | 1 | 4 | 1 | 4 |
| p.Q228K°; VAF 100%; benign, germline (0.03); PRL | ||||
| p.V301M$; VAF 25%; benign (0.01); SG ACTH | ||||
| p.R188W; VAF 51% pathogenic (0.98); FSH/LH | p.R307Q; VAF 72%; pathogenic (0.99); FSH/LH | |||
| p.E61K§; VAF 100%; pathogenic (0.99); FSH/LH | ||||
| p.S53N; VAF 38%; pathogenic (0.99); PRL | ||||
| CDKN2A | 0 | 0 | 1 | 1 |
| p.D74A°; VAF 30%; pathogenic (0.99); PRL |
Note: Genes were considered mutated for variant allele frequency (VAF) > 20%. §, $,° = mutations were detected in the same pituitary tumour.
Pathogenic scores were calculated by Poly‐Phen‐2 web tool (http://genetics.bwh.harvard.edu/pph2/) and verified on COSMIC database. 16
Abbreviations: DG, densely granulated; PA, pituitary adenoma; PitNET, pituitary neuroendocrine tumour; SG, sparsely granulated.
3.3. DNA methylation analysis
Methylation levels detected in normal pituitaries, non‐aggressive and aggressive PAs/PitNETs, and carcinomas are summarised in Figure 1.
Figure 1.
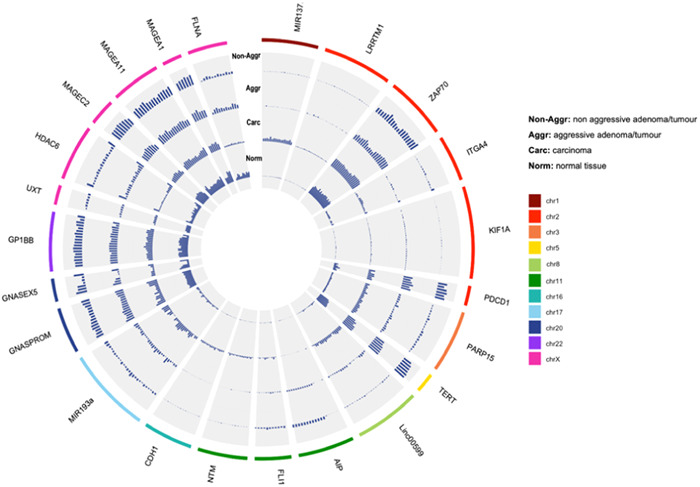
Circle plot depicting methylation levels of autosomes and X‐linked genes in normal pituitary, aggressive and non‐aggressive PA/PitNETs and carcinomas. PA, pituitary adenoma; PitNET, pituitary neuroendocrine tumour
Methylation data of CpGs of all investigated genes, including mean levels, standard deviation and Kruskal–Wallis p values are reported in Table S5.
The methylation level of PARP15, LINC00599, ZAP70 was higher in aggressive PAs/PITNETs (mean values for informative CpGs: 0.21, 0.08 and 0.48, respectively) compared to non‐aggressive ones (mean values for informative CpGs: 0.11, 0.03 and 0.40, respectively; according to Kruskal–Wallis test with p < .05).
Levels of methylation of AIP, GNAS and PDCD1 were significantly lower in aggressive PAs/PITNETs (mean values for informative CpGs: 0.01, 0.31 and 0.64, respectively) than in non‐aggressive ones (mean values for informative CpGs: 0.18, 0.48 and 0.70, respectively; according to Kruskal–Wallis test with p < .05). Informative CpGs are reported in Figure 2 and highlighted with an asterisk.
Figure 2.
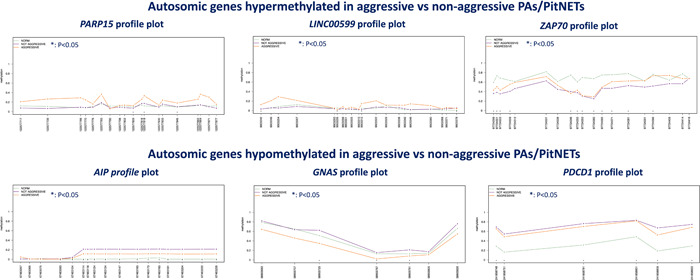
Methylation plots of autosomes in normal pituitary tissue, non‐aggressive and aggressive PAs/PitNETs. The asterisk (*) indicates CpGs that show statistical differences according to the non‐parametric Kruskal–Wallis test. PA, pituitary adenoma; PitNET, pituitary neuroendocrine tumour
No significant differences among groups were detected for other investigated autosomal genes.
X‐linked genes were analyzed separately according to patient gender.
In males, levels of methylation of FLNA, UXT and MAGE family (MAGEA11, MAGEA1, MAGEC2) genes were significantly higher in aggressive PAs/PITNETs (mean values for informative CpGs: 0.15, 0.096, 0.81, 0.88 and 0.95, respectively) than in non‐aggressive ones (mean values for informative CpGs: 0.007, 0.025, 0.77, 0.69 and 0.88, respectively; according to Kruskal–Wallis test with p < .05). Informative CpGs are reported in Figure 3 and highlighted with an asterisk.
Figure 3.
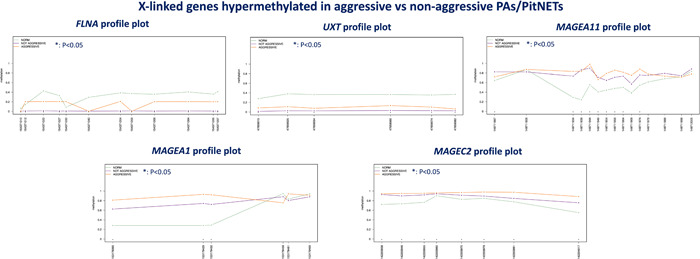
Methylation plots of X‐linked genes in males in normal pituitary tissue, non‐aggressive and aggressive PAs/PitNETs. The asterisk (*) indicates CpGs that show statistical differences according to the non‐parametric Kruskal–Wallis test. PA, pituitary adenoma; PitNET, pituitary neuroendocrine tumour
In females, no significant differences were detected between aggressive and non‐aggressive PAs/PITNETs.
In the low number of pituitary carcinomas examined, the levels of methylation of autosomal genes (PARP15, LINC00599, MIR193 and ZAP70) was higher, while for X‐linked genes methylation level was lower than in PAs/PITNETs.
3.4. Patient outcome
All patients underwent selective and radical adenoma resection, defined by the absence of visible remnants at the end of surgery (confirmed by the surgeon) and the MRI with contrast medium performed 3 months after surgery. Tumour recurrence was observed in 38 patients (34.2%), that received additional post‐surgical therapy, often in combination. Specifically, 25 (22.5%) patients, including 15 (36.6%) with aggressive and 4 (6.3%) with non‐aggressive PAs/PitNETs, and 6 (100%) with carcinomas—underwent second surgery; 27 (24.3%) patients including 23 (56.1%) with aggressive and 3 (4.7%) with non‐aggressive PAs/PitNETs, and 1 (16.7%) with carcinoma received medical therapy; and 9 (8.1%) radiotherapy including 7 (17.1%) with aggressive PAs/PitNETs and 2 (33.3%) with carcinomas. Two patients with aggressive ACTH‐secreting PAs/PitNETs underwent adrenalectomy (Table 1). The rate of patients requiring second surgery was significantly higher among carcinomas than PAs/PitNETs (p = .0001). No difference was found for medical and radiation therapy.
Patients with aggressive PAs/PitNETs required second surgery, medical and radiation therapy more frequently than those with non‐aggressive tumours (p = .0001 for all treatments).
4. DISCUSSION
Epigenetic alterations have been increasingly investigated in association with somatic gene mutations in pituitary tumours to understand the mechanisms underlying tumorigenesis, the progression to carcinomas, and to identify predictors of clinical behaviour and response to treatment.
Based on our data, mutation rate was low, although significantly higher in aggressive versus non‐aggressive tumours, in agreement with previous studies performed in sporadic PAs/PitNETs. 7 , 8 AIP, and TP53 were the most frequently mutated genes in aggressive PAs/PitNETs, NOTCH1 and USP8 in non‐aggressive PAs/PitNETs while pituitary carcinoma showed mutation in both AIP and CDKN2A. 7 , 8 , 20 , 21 AIP gene showed pathogenic mutations in only 3.6% of cases, a rate similar to previous studies. 22 The identification of TP53 mutation in PAs/PitNETs suggests its implication in the neoplastic transformation of pituitary cells at least in a small fraction of cases, although its role remains debated. 8 , 23 , 24 , 25 We also detected USP8 mutation in two densely granulated functioning corticotroph tumours, confirming literature data on the specificity of USP8 mutation for corticotroph tumours, reported in 25%–60% of the examined cases, with a significantly higher prevalence in functioning than silent forms. 26 , 27 These two cases presented wildtype TP53, supporting the hypothesis that, in corticotroph tumours, TP53 and USP8 mutations are mutually exclusive. 24 , 25
Alterations in methylation levels in PAs/PitNETs, as well as carcinomas, were more common than gene mutations, and were identified in the majority of the genes analyzed in the study, supporting previous literature data. 28 Differences were found in methylation levels among non‐aggressive, aggressive PAs/PitNETs and carcinomas in genes involved in pituitary tumorigenesis (GNAS and AIP), microRNA (miRNA)/long noncoding RNA (LncRNA)‐post‐transcriptional regulation (i.e., LINC00599), targets of new therapies for aggressive tumours/carcinomas (PDCD1, PARP15 and ZAP70), and cell proliferation/differentiation (MAGEA family, UXT and FLNA). 7 , 29 , 30
We also noticed that levels of methylation of PARP15, LINC00599, ZAP70 were significantly higher in aggressive PA/PITNET than in non‐aggressive PA/PITNET. Since hypermethylation often results in gene silencing and reduced protein expression, our data is in keeping with previous observations of the lower expression of PARP15 and LINC00599 in aggressive extra‐pituitary tumours, and on the association between hypermethylation of ZAP70 and worse prognosis. 29
The demonstration of altered expression (secondary to different levels of methylation) of factors modulating oncogenes/oncosuppressors in non‐aggressive and aggressive PAs/PitNETs, while oncogenes/oncosuppressor mutations occur only in a minority of aggressive tumours, suggest the alteration of gene methylation may be an early event, potentially promoting aggressive tumour behaviour.
In addition, hypermethylation of PDCD1 in PAs/PitNETs seem to support the use of anti‐PD1 and PDL‐1 inhibitors in PAs/PitNETs. 30 , 31 At the same time, reduced methylation levels in carcinomas suggest a potential mechanism of escape of non‐hypermutated pituitary carcinomas from immunological control.
An important finding of this study is the identification of different methylation patterns of X‐linked genes, i.e., androgen receptor regulator genes, including MAGE family, UXT and FLNA, 32 in PAs/PitNETs according to patient sex.
In 2008 Zhu et al., 33 examining gene expression in normal and neoplastic human and mouse pituitary cells, first reported higher levels of MAGEA3/CTAG in females as compared with males and the correlation between MAGEA3/CTAG expression and the extent of DNA promoter methylation. In addition, MAGEA3/CTAG was suggested as a candidate gene responsible for p53 dysregulation in pituitary tumours.
A variation in methylation levels of MAGE family members according to gender was found in our study, supporting the role of AR expression in pituitary tumorigenesis, suggested by Scheithauer et al. 34 Furthermore, variation in methylation levels of MAGE family members and UXT could contribute to PAs/PitNETs cell proliferation indirectly by the regulation of p53 expression/activity, 35 as reported above for AIP, reinforcing the hypothesis of the synergy of epigenetics and somatic gene mutations in pituitary tumorigenesis.
Finally, differences in the levels of methylation of X‐linked genes could be associated with gender differences in tumour behaviour. 36
The restricted number of genes screened for mutation and methylation analysis represents the main study limitation. Further researches, performed on larger tumour series and assessing a wider panel of genes, ideally using fresh/frozen specimens (enabling the extraction of high quality and quantity of DNA) and with a prospective design, are strongly recommended.
5. CONCLUSIONS
A different profile of somatic mutations and methylation of genes involved in cell proliferation/differentiation, miRNA/LncRNA‐post‐transcriptional regulators, and targets of antineoplastic therapies has been identified in non‐aggressive and aggressive PAs/PitNETs, and pituitary carcinomas. Alteration of methylation levels appears to be an early event that could contribute to determine tumour aggressiveness, response to treatment and progression to carcinoma, directly or indirectly by inducing mutation of oncogenes and oncosuppressors. The involvement of X‐linked genes could explain differences in tumour behaviour according to gender. These data, although preliminary, seem to suggest the importance of identifying molecular markers to be integrated with clinico‐radiological and pathological data to predict pituitary tumour behaviour. Future studies should include larger patient cohorts and expand the analysis to other emerging genes in the molecular landscape of pituitary tumours (e.g., DNA mismatch repair genes, ATRX, RB1, NF2, CDKN2B and CTAG 2).
CONFLICTS OF INTEREST
The authors declare no conflicts of interest.
ETHICS STATEMENT
The study was performed according to the principles of the Declaration of Helsinki. The study was approved by the local ethics committee (Protocol Number: 269‐2020‐OSS‐AUSLBO evaluated on 25/03/2020). Sensitive data information was anonymized by replacing patient names by numerical codes. Informed consent was obtained from all participants to the study.
Supporting information
Supplementary information.
Supplementary information.
Supplementary information.
Supplementary information.
Supplementary Table 5: Methylation data of CpGs of all investigated genes, including mean levels, standard deviation and Kruskal‐Wallis p values.
ACKNOWLEDGEMENTS
The study was funded by the research grant ‘Aggressive and recurrent Pituitary Tumours: study of risk factors and involved genes (RECurrence in Aggressive PItuitary Tumours: RECAPIT; P.I.: Prof. Diego Mazzatenta)’, winner of the Tender for Translational Medical Research 2019, by the private non‐profit organisation ‘Fondazione Cassa di Risparmio di Bologna’, Bologna, Italy; and by the ‘Ricerca Corrente’ grant (P.I.: Prof. Caterina Tonon) by The Italian Ministry of Health to the IRCCS Istituto delle Scienze Neurologiche di Bologna. Open access funding provided by BIBLIOSAN.
Guaraldi F, Morandi L, Zoli M, et al. Epigenomic and somatic mutations of pituitary tumors with clinical and pathological correlations in 111 patients. Clin Endocrinol (Oxf). 2022;97:763‐772. 10.1111/cen.14827
Federica Guaraldi and Luca Morandi equally contributed to the manuscript.
This work was partially presented as platform presentation at the 109th Meeting of the United States and Canadian Academy of Pathology in Los Angeles, CA, USA, February 28‐March 5, 2020.
DATA AVAILABILITY STATEMENT
The authors declare transparency and availability of data, material, and code (all data that support the findings of this research is deposited in Supporting Information Files).
REFERENCES
- 1. Trouillas J, Jaffrain‐Rea M‐L, Vasiljevic A, Raverot G, Roncaroli F, Villa C. How to classify the pituitary neuroendocrine tumors (PitNET)s in 2020. Cancers. 2020;12(2):514. 10.3390/cancers12020514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Laws ER, Penn DL, Repetti CS. Advances and controversies in the classification and grading of pituitary tumors. J Endocrinol Invest. 2019;42(2):129‐135. 10.1007/s40618-018-0901-5 [DOI] [PubMed] [Google Scholar]
- 3. Raverot G, Burman P, McCormack A, et al. European Society of Endocrinology Clinical Practice Guidelines for the management of aggressive pituitary tumours and carcinomas. Eur J Endocrinol. 2018;178(1):G1‐G24. 10.1530/EJE-17-0796 [DOI] [PubMed] [Google Scholar]
- 4. Asa SL, Asioli S, Bozkurt S, et al. Pituitary neuroendocrine tumors (PitNETs): nomenclature evolution, not clinical revolution. Pituitary. 2020;23(3):322‐325. 10.1007/s11102-019-01015-0 [DOI] [PubMed] [Google Scholar]
- 5. Louis DN, Perry A, Wesseling P, et al. WHO classification of tumors of the central nervous system: a summary. Neuro Oncol. 2021. The 2021;23(8):1231‐1251. 10.1093/neuonc/noab106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Osamura RY, Grossman A, Korbonits M, Kovacs K, Lopes MBS, Matsuno A. Pituitary adenoma. In: Llyod RV, Osamura RY, Kloppel G, Rosai J, (eds), WHO Classification of Tumours of Endocrine Organs. 4th edn, IARC, Lyon 2017:14‐18. [Google Scholar]
- 7. Asa SL, Mete O. Ezzat S. Genomicsgenomics and epigenomics of pituitary tumors: what do pathologists need to know? Endocr Pathol. 2021;32:3‐16. 10.1007/s12022-021-09663-4 [DOI] [PubMed] [Google Scholar]
- 8. Spada A, Mantovani G, Lania AG, et al. Pitu itary tumors: genetic and molecular factors underlying pathogenesis and clinical behavior. Neuroendocrinology. 2022;112(1):15‐33. 10.1159/000514862 [DOI] [PubMed] [Google Scholar]
- 9. Neou M, Villa C, Armignacco R, et al. Pangenomic classification of pituitary neuroendocrine tumors. Cancer Cell. 2020;37:123‐134.e5. 10.1016/j.ccell.2019.11.002 [DOI] [PubMed] [Google Scholar]
- 10. Castellnou S, Vasiljevic A, Lapras V, et al. SST5 expression and USP8 mutation in functioning and silent corticotroph pituitary tumors. Endocr Connect. 2020;9:243‐253. 10.1530/EC-20-0035 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Asioli S, Righi A, Iommi M, et al. Validation of a clinicopathological score for the prediction of post‐surgical evolution of pituitary adenoma: retrospective analysis on 566 patients from a tertiary care centre. Eur J Endocrinol. 2019;180(2):127‐134. 10.1530/EJE-18-0749 [DOI] [PubMed] [Google Scholar]
- 12. Mete O, Ezzat S, Perry A, et al. The pangenomic classification of pituitary neuroendocrine tumors: quality histopathology is required for accurate translational research. Endocr Pathol. 2021. 2021;32(3):415‐417. 10.1007/s12022-021-09671-4 [DOI] [PubMed] [Google Scholar]
- 13. Caimari F, Korbonits M. Novel genetic causes of pituitary adenomas. Clin Cancer Res. 2016;22:5030‐5042. 10.1158/1078-0432.CCR-16-0452 [DOI] [PubMed] [Google Scholar]
- 14. Gabusi A, Gissi DB, Tarsitano A, et al. Intratumoral heterogeneity in recurrent metastatic squamous cell carcinoma of the oral cavity: new perspectives afforded by multiregion DNA sequencing and mtDNA analysis. J Oral Maxillofac Surg. 2019;77(2):440‐455. 10.1016/j.joms.2018.09.014 [DOI] [PubMed] [Google Scholar]
- 15. Afgan E, Baker D, Batut B, et al. The galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018;46(W1):W537‐W544. 10.1093/nar/gky379 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Tate JG, Bamford S, Jubb HC, et al. COSMIC: the catalogue of somatic mutations in cancer. Nucleic Acids Res. 2019;47:D941‐D947. 10.1093/nar/gky1015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Li L‐C, Dahiya R. MethPrimer: designing primers for methylation PCRs. Bioinformatics. 2002;18:1427‐1431. 10.1093/bioinformatics/18.11.1427 [DOI] [PubMed] [Google Scholar]
- 18. Gissi DB, Tarsitano A, Gabusi A, et al. 13‐Gene DNA methylation analysis from oral brushing: a promising non invasive tool in the follow‐up of oral cancer patients. J Clin Med. 2019;8(12):2107. 10.3390/jcm8122107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Krainer J, Weinhäusel A, Hanak K, et al. EPIC‐TABSAT: analysis tool for targeted bisulfite sequencing experiments and array‐based methylation studies. Nucleic Acids Res. 2019;47:W166‐W170. 10.1093/nar/gkz398 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Guaraldi F, Salvatori R. Familial isolated pituitary adenomas: from genetics to therapy. Clin Transl Sci. 2011;4(1):55‐62. 10.1111/j.1752-8062.2010.00254.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Peculis R, Niedra H, Rovite V. Large scale molecular studies of pituitary neuroendocrine tumors: novel markers, mechanisms and translational perspectives. Cancers. 2021;13:1395. 10.3390/cancers13061395 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Coopmans EC, Márta Korbonits M. Molecular genetic testing in the management of pituitary disease. Clin Endocrinol (Oxf). 2022;97:424‐435. 10.1111/cen.14706 [DOI] [PubMed] [Google Scholar]
- 23. Sbiera S, Perez‐Rivas LG, Taranets L, et al. Driver mutations in USP8 wild‐type cushing's disease. Neuro Oncol. 2019;21(10):1273‐1283. 10.1093/neuonc/noz109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Uzilov AV, Taik P, Cheesman KC, et al. USP8 and TP53 drivers are associated with CNV in a corticotroph adenoma cohort enriched for aggressive tumors. J Clin Endocrinol Metab. 2021;106(3):826‐842. 10.1210/clinem/dgaa853 [DOI] [PubMed] [Google Scholar]
- 25. Hasanov R, Bİ A, Kiremitçi S, Erden E, Güllü S. The prognostic roles of the Ki‐67 proliferation index, P53 expression, mitotic index, and radiological tumor invasion in pituitary adenomas. Endocr Pathol. 2019;30:49‐55. 10.1007/s12022-018-9563-2 [DOI] [PubMed] [Google Scholar]
- 26. Ma Z‐Y, Song Z‐J, Chen J‐H, et al. Recurrent gain‐of‐function USP8 mutations in cushing's disease. Cell Res. 2015;25(3):306‐17. 10.1038/cr.2015.20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Perez‐Rivas LG, Theodoropoulou M, Ferraù F, et al. The gene of the Ubiquitin‐Specific protease 8 is frequently mutated in adenomas causing cushing's disease. J Clin Endocrinol Metab. 2015;100(7):E997‐E1004. 10.1210/jc.2015-1453 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Ezzat S, Cheng S, Asa SL. Epigenetics of pituitary tumors: pathogenetic and therapeutic implications. Mol Cell Endocrinol. 2018;469:70‐76. 10.1016/j.mce.2017.07.011 [DOI] [PubMed] [Google Scholar]
- 29. Khordadmehr M, Shahbazi R, Baradaran B, Sadreddini S, Shanebandi D, Hajiasgharzadeh K. Restoring of miR‐193a‐5p sensitizes breast cancer cells to paclitaxel through P53 pathway. Adv Pharm Bull. 2020;10:595‐601. 10.34172/apb.2020.071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Dai C, Liang S, Sun B, Kang J. The progress of immunotherapy in refractory pituitary adenomas and pituitary carcinomas. Front Endocrinol. 2020;11:608422. 10.3389/fendo.2020.608422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Ricci C, Morandi L, Righi A, et al. PD‐1 (PDCD1) promoter methylation in Merkel cell carcinoma: prognostic relevance and relationship with clinico‐pathological parameters. Mod Pathol. 2019;32:1359‐1372. 10.1038/s41379-019-0261-5 [DOI] [PubMed] [Google Scholar]
- 32. Foschini MP, Morandi L, Sanchez AM, et al. Methylation profile of X‐chromosome‐related genes in male breast cancer. Front Oncol. 2020;10:784. 10.3389/fonc.2020.00784 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Zhu X, Asa Sl, Ezzat S. Fibroblast growth factor 2 and estrogen control the balance of histone 3 modifications targeting MAGE‐A3 in pituitary neoplasia. Clin Cancer Res. 2008;14:1984‐1996. 10.1158/1078-0432.CCR-07-2003 [DOI] [PubMed] [Google Scholar]
- 34. Scheithauer BW, Kovacs K, Zorludemir S, Lloyd RV, Erdogan S, Slezak J. Immunoexpression of androgen receptor in the nontumorous pituitary and in adenomas. Endocr Pathol. 2008;19:27‐33. 10.1007/s12022-007-9012-0 [DOI] [PubMed] [Google Scholar]
- 35. Qi M, Ganapathy S, Zeng W, Zhang J, Little JB, Yuan Z. A novel MDMX‐binding protein, promotes glycolysis by mitigating p53‐mediated restriction of NF‐κB activity. Oncotarget. 2015;6:17584‐17593. 10.18632/oncotarget.3770 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Delgrange E, Vasiljevic A, Wierinckx A, et al. Expression of estrogen receptor alpha is associated with prolactin pituitary tumor prognosis and supports the sex‐related difference in tumor growth. Eur J Endocrinol. 2015;172:791‐801. 10.1530/EJE-14-0990 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary information.
Supplementary information.
Supplementary information.
Supplementary information.
Supplementary Table 5: Methylation data of CpGs of all investigated genes, including mean levels, standard deviation and Kruskal‐Wallis p values.
Data Availability Statement
The authors declare transparency and availability of data, material, and code (all data that support the findings of this research is deposited in Supporting Information Files).


