Abstract
Aromatase inhibitors (AIs) are a class of drugs commonly given to patients with estrogen receptor (ER)-dependent breast cancers to reduce estrogenic stimulation. However, AIs like Letrozole are associated with negative side effects such as cognitive deficits, sleep disturbances and hot flashes. We have previously shown that these negative effects can be recapitulated in common marmosets (Callithrix jacchus) treated with Letrozole (20 μg daily) for 4 weeks and that marmosets treated with Letrozole show increased levels of estradiol in the hippocampus (Gervais et al., 2019). In order to better understand the mechanisms through which AIs affect cognitive function and increase steroid levels in the hippocampus, we used bulk, paired-end RNA-sequencing to examine differentially expressed genes among Letrozole-treated (LET; n=8) and vehicle-treated (VEH; n=8) male and female animals. Gene ontology results show significant reduction across hundreds of categories (P < 0.05), some of the most significant being inflammatory response, stress response, MHC Class II protein complex binding, T-cell activation, carbohydrate binding and signaling receptor binding in LET animals (P < 5.0E-5). GSEA results indicate that LET females, but not LET males (FDR = 0.67), show enrichment for hormonal gene sets (FDR = 0.1). Based on the transcriptional changes observed, we conclude that AIs may differentially affect the sexes in part due to processes mediated by the CYP-450 superfamily. Ongoing studies will further investigate the longitudinal effects of AIs on behavior and whether AIs increase the risk of stress-induced neurodegeneration.
Keywords: Aromatase inhibitor, Letrozole, estradiol, hippocampus, RNA-sequencing
Introduction
Estrogen-receptor (ER) positive breast cancers represent one of the most common forms of breast cancer in cisgender women. In ER-positive breast cancers, estrogens are derived from the aromatization of testosterone and other steroid hormones and facilitate cancerous growth via ER signaling pathways and estrogen metabolites (Bhatnagar, 2007; Fabian 2007). One method to disrupt estrogen-induced cancerous growth is through the administration of aromatase inhibitors (AIs) which function by preventing the synthesis of estrogens or blocking estrogen receptors. While AIs are the standard for hormone based adjuvant therapies, they are accompanied by several manifestations of central nervous system (CNS) deficits such as cognitive impairment, insomnia, and hot flashes. These side effects can be severe enough to deter patients from continuing this life-saving treatment (Garreau et al., 2006).
Understanding the mechanisms by which AIs induce CNS deficits is critical to finding improved alternatives. In vitro studies in rat hippocampal slices have provided convincing evidence that Letrozole treatment decreases estradiol levels, synaptic density, and levels of spinophilin and synaptophysin (Kretz et al., 2004), as well as reduced LTP (Di Mauro et al., 2017; Vierk et al., 2012). Some of these findings have been replicated in vivo; Letrozole and other AIs applied directly to the brain via cannulation or microdialysis suppress local estradiol synthesis (Marbouti et al., 2020; Tuscher et al., 2016) and lead to decreases in hippocampal spine density (Prange-Kiel et al., 2006) as well as a decrease the expression of genes involved in steroidogenesis and synapse formation (Marbouti et al., 2020). Central aromatase inhibition also induces cognitive deficits in a variety of species, including song birds (Bailey et al., 2013; Macedo-Lima & Remage-Healey, 2020), rats (Marbouti et al., 2020; Tuscher et al., 2016) and mice (Liu et al., 2019; Zhao et al., 2018).
However, central administration of Letrozole does not model the systemic administration used in patients and may miss important cross-talk between the periphery and CNS or interactions within the blood brain barrier. Letrozole administered intravenously crosses the blood brain barrier in a dose-dependent manner in rats (Dave et al., 2013) and is reported to have CNS effects in cisgender women (Vierk et al., 2012), but its effect on estradiol synthesis and hippocampal function remains unclear. Zhou et al. (2010) reported that systemic administration of Letrozole for 4 weeks decreased hippocampal synapses in intact and ovariectomized female mice, while Fester et al. (2012) found similar effects in intact female rats, but no effect in male rats. Additionally, sex and brain region-specific differences have been found in intact and gonadectomized rodents. Recent studies have shown that Letrozole decreases aromatase activity in the hypothalamus of intact and gonadectomized Wistar rats (Kokras et al., 2018) and decreases estradiol levels in the hypothalamus of common marmosets (Kraynak et al., 2022). However, the hypothalamic blood brain barrier is unique in its permeability and sensitivity to systemically circulating substances; it is possible that this region’s sensitivity to aromatase inhibition is dissimilar to other brain regions with tighter junctions between endothelial cells within the blood brain barrier (Haddad-Tóvolli et al., 2017).
Interestingly, conflicting results have been found using non-rodent models. A study by Villeneuve et al. (2009) previously reported a dose- and time-dependent increase of aromatase in the brain and in peripheral tissues of zebrafish exposed to the second-generation AI Fadrozole. Similarly, Gervais et al. (2019) reported an increase in estradiol in the hippocampus of marmosets treated with oral Letrozole for 4 weeks. While these outcomes may reflect compensatory mechanisms for the loss of peripheral estradiol, there is a need to elucidate these mechanisms in a nonhuman primate that most closely models the dose, mode of administration and consequences of Letrozole use in patients.
The aim of this study was to assess gene expression in the hippocampus of male and female marmosets treated with an oral dose of Letrozole or vehicle for 4 weeks. We previously reported that Letrozole-treated monkeys exhibited adverse changes in cognition, thermoregulation and anxiety as well as compromised hippocampal intrinsic excitability (Gervais et al., 2019). Here, we used RNA-sequencing to identify differentially expressed genes in the hippocampus of these animals according to treatment and sex. We hypothesized that genes involved in steroidogenesis would be upregulated in Letrozole-treated (LET) animals, reflecting compensatory mechanisms due to peripherally low estrogen levels. Our findings highlight differentially expressed genes involved overwhelmingly in immune response in LET animals while genes involved in steroidogenesis and cognitive impairment were enriched only in LET females. Collectively, these results may improve our understanding of the impact AIs have on the human brain and behavior.
Materials and Methods
Experimental Design and Statistical Analysis.
Brain tissue from sixteen (males: n = 9; females: n = 7) common marmosets (Callithrix jacchus) previous studied in Gervais et al. (2019) was used to assess whether Letrozole affected hippocampal gene expression and evaluate sex differences in differential expression (Figure 1). Treatment groups were matched based on age and sex, with animals receiving 20 μg of either Letrozole mixed in pudding or pudding without the drug. These animals were part of a previous study (Gervais et al., 2019) that measured behavior, memory and thermoregulation as well as CA1 hippocampal physiology to evaluate if Letrozole’s negative side effects reported in cisgender women could be recapitulated in common marmosets. All animals were gonadectomized approximately 3 years before the experiment began which prevented socially housed animals from breeding or undergoing ovulatory cycles. Our previous study measured peripheral levels of estradiol in both control and Letrozole-treated animals and found low circulating levels of estradiol in the urine of control animals and nearly no estradiol in the urine of Letrozole-treated animals (Gervais et al., 2019). The animals were cared for and humanely euthanized in accordance with the Guide for the Care and Use of Laboratory Animals and University of Massachusetts Institutional Animal Care and Use Committee.
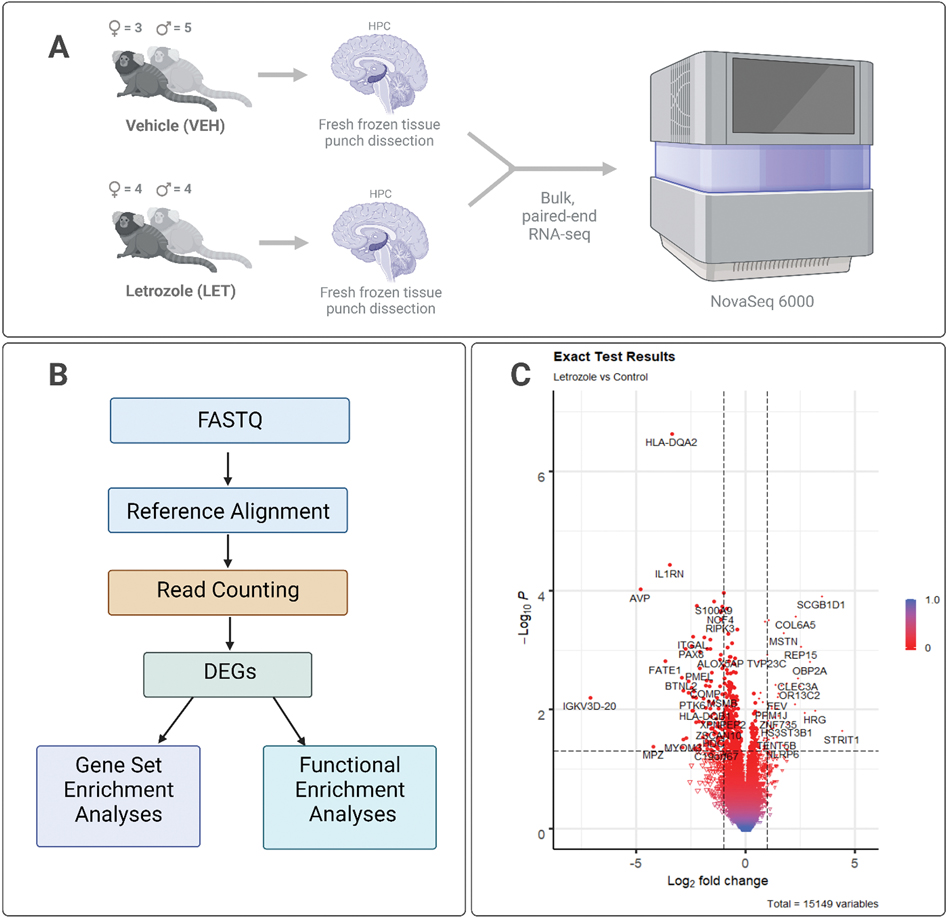
Figure 1. Experimental Design and Differentially Expressed Genes among LET and VEH animals.
(A) Common marmosets were assigned to vehicle (VEH; n = 3 females, n = 5 males) or Letrozole (LET; n = 4 females, n = 4 males). The right hippocampus was dissected from each sample and processed for bulk, paired-end RNA-sequencing.
(B) The data processing workflow involved downloading forward and reverse FASTQ files and mapping them to a reference genome. Reads were counted before assessing differentially expressed genes (DEGs) via gene set enrichment analyses (GSEAs) or functional enrichment analyses.
(C) Post-data normalization and filtering for lowly expressed genes, 15,149 genes were assessed for differential expression between Letrozole- and Vehicle-treated groups. Horizontal dotted lines indicate the threshold at which individual genes’ p-value < 0.05, whereas vertical lines represent a log2 fold change greater than 0.5 (a fold change of 1). The triangle shapes for individual genes represent genes which have a significant fold change but non-significant p-value.
Brain Matrix and Tissue Dissection.
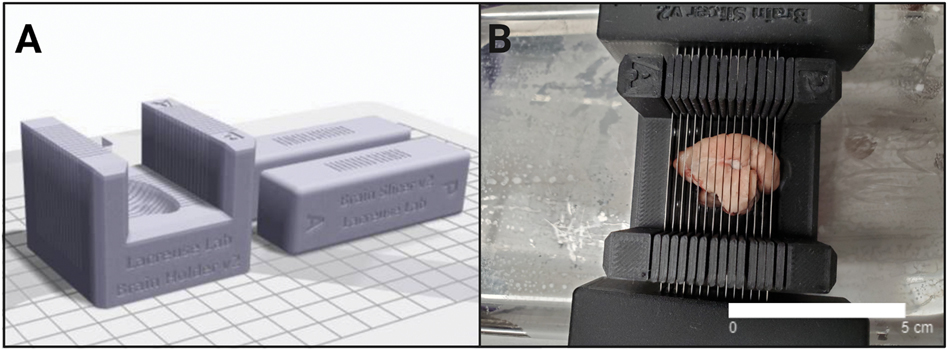
This study used fresh frozen brain tissue (right hemisphere) which was stored in a −80°C freezer. In order to carefully dissect the right hippocampus, a custom made, 3D-printed brain slicer and blade holders were adapted from Guy et al. (2016) whose design methods have previously been described (Figure 2A). The brain slicer and accompanying blade holders were customized in SolidWorks® and 3D-printed in onyx at the Advanced Digital Design and Fabrication (ADDFab) Core Facility at the University of Massachusetts Amherst. An MRI of a marmoset brain from the Silva lab (see Guy et al., 2016) was superimposed onto the brain matrix to allow for a sagittal mold. Onyx was chosen as an ideal material due to its durability and ability to withstand −20°C freezer incubation prior to dissection. Low-profile microtome blades (0.24×80mm) (Fisher Scientific) and their measurements were incorporated into the SolidWorks design of the custom blade holders.
Figure 2. Brain Matrix Design.
(A) 3D-printed brain matrix and accompanying blade holders were designed using SOLIDWORKS® software and a matrix template provided by the Silva lab (Guy et al., 2016). An MRI of a marmoset brain was superimposed in a sagittal position on the matrix. (B) The brain matrix and blade holders were printed in onyx and dissected over dry ice to preserve the brain while obtaining ~ 2mm sections before tissue punch and homogenization.
Prior to tissue dissection, the right hemisphere from each animal was briefly removed from a −80°C freezer and immediately placed in a −20°C freezer overnight to slowly thaw the tissue. Several hours before dissection, the brain slicer was also placed in the −20°C freezer and all materials were autoclaved, wiped down with RNAse wipes and allowed to dry completely. The tissue dissection took place over a bed of dry ice and 2mm slices were dissected, then immediately transferred to polystyrene petri dishes directly atop dry ice (Figure 2B). The right hippocampus was identified with a common marmoset brain atlas (Palazzi & Bordier, 2009) and dissected via a 2000-micron biopsy punch (Harvard Apparatus). Once the hippocampus tissue punches were removed, the tissue was transferred to 1.5 mL centrifuge tubes and weighed before storing on dry ice. Due to the high lipid content of the marmoset brain, tissue punches performed best for downstream analysis if kept under 10 mg total weight (average = 6.61 mg). All samples were dissected by the same researcher in order to maintain consistency.
RNA extraction.
Qiazol (Qiagen) was added to each vial before homogenization with the FastPrep-24 5G Sample Homogenizer (MP Biomedicals) courtesy of the Genomics Resource Laboratory at the University of Massachusetts Amherst. Total RNA was isolated using an RNeasy Mini Kit (Qiagen). RNA concentration and quality were assessed using a Qubit Fluorometer (RNA HS Assay, ThermoFisher Scientific) and Agilent 2100 Bioanalyzer (RNA 6000 Pico Kit, Agilent) respectively. Samples with an RNA integrity score less than seven were not used for library construction.
Library Preparation.
Illumina-compatible libraries were prepared using the QIAseq Stranded mRNA Select Kit (Qiagen). Poly(A)+ mRNA was isolated by binding to the surface of magnetized Qiagen Pure mRNA beads in the presence of buffer. The mRNA was then washed and eluted with nuclease-free water followed by fragmentation into ~350 bp fragments. First strand cDNA synthesis was performed per protocol by using an RNase H-Reverse Transcriptase in tandem with random primers. Second strand synthesis was performed using 5’ phosphorylated random primers and QIAseq Unique Dual-Index (UDI) adapters (two unique barcode motifs per adapter molecular) are ligated to the to 5’ phosphorylated strand for sample identification. Libraries were amplified using the Mastercycler Pro thermal cycler (Eppendorf) and quality control checks were performed using the Agilent Bioanalyzer for correct size of library fragments and absence of adapter-dimers. The libraries were sequenced at a depth of 20 million paired-end reads per sample using the Novaseq 6000 at Medgenome, Foster City, CA, USA.
Data Preprocessing.
After sequencing, data quality was assessed with FASTQC analysis. Sequences were mapped to the marmoset reference genome (GCA_011100555.1) using Bowtie (Langmead et al., 2009) and counts per gene were obtained using HTSeq (Anders et al., 2014). Raw counts were imported into RStudio (v4.0.4) and human one-to-one ortholog names for each gene were obtained with the RStudio package gProfiler2 (Kolberg et al., 2020). We then performed library normalization for composition bias using edgeR and transformed the data into counts per million (CPM). Lowly expressed genes were filtered using a CPM threshold of 0.3 (a minimum of four counts per gene) in at least two samples and only one-to-one human-marmoset orthologs were used. Thus, 19,359 unique orthologous genes were assayed after filtering. The FASTQ files and raw counts for each animal can be found on the NCBI Sequence Read Archive (SRA) and extended data.
Differential Expression Analyses.
We used quantile-adjusted conditional maximum likelihood (qCML) methods in edgeR to normalize and compare the data (Chen et al., 2016; Robinson et al., 2009). In order to visualize dispersion estimates, biological coefficient of variation (BCV) plots were made and common dispersion was chosen for Fisher’s exact tests. The exact test functionality in edgeR was used to examine differences in the means between two groups before extracting significant tags (Robinson & Smyth, 2007). This function defaults to using the Benjamin and Hochberg’s algorithm to control the false discovery rate (FDR) at level alpha (α = 0.05) and is appropriate for small sample sizes. Because the exact test in edgeR is suitable for experiments with a single factor, female and male subjects were analyzed in separate dataframes to evaluate sex-specific outcomes across LET and vehicle-treated (VEH) groups.
Gene Ontology and Gene Set Enrichment Analyses.
Functional enrichment analyses were used to analyze relevant biological processes, biological pathways, cellular components and molecular function using Panther (http://pantherdb.org), gProfiler (g:GOst; https://biit.cs.ut.ee/gprofiler) and REVIGO (http://revigo.irb.hr/) using up- and down-regulated genes with an FDR < 0.05. Five types of gene set enrichment analyses (GSEAs) were performed using the GSEA Software version 4.2.1 (Subramanian et al., 2005) to evaluate gene sets and network models. Using curated gene sets from MSigDB based on published literature (Subramanian et al., 2005), we focused on three curated hormonal gene sets: regulation of hormone levels, cellular hormone metabolic processes and steroid hormone biosynthesis. Since the Cytochrome P450 family is essential for metabolizing 70–80% of all clinical drugs (Meyer et al., 2007; Miksys & Tyndale, 2013; Zanger & Schwab, 2013), we also included a drug metabolism gene set. Lastly, due to the cognitive deficits Gervais et al. (2018) observed for LET animals and our findings of downregulated amyloid beta clearance in LET animals (Figure 3-1), we included a gene set for Alzheimer’s disease to evaluate whether Letrozole increases transcripts involved in neurodegeneration. The GSEA report generated from each analysis highlights enrichment gene sets with an FDR less than 25% as biologically relevant and likely to drive further research (Subramanian et al., 2005). When appropriate, 1,000 phenotype-based permutations were selected for to generate a null distribution for calculating the enrichment score and significance (Subramanian et al., 2005). When making comparisons with too few samples for each group (e.g. sex and treatment), gene set permutations were used instead with a more stringent threshold for significance (FDR < 0.10).
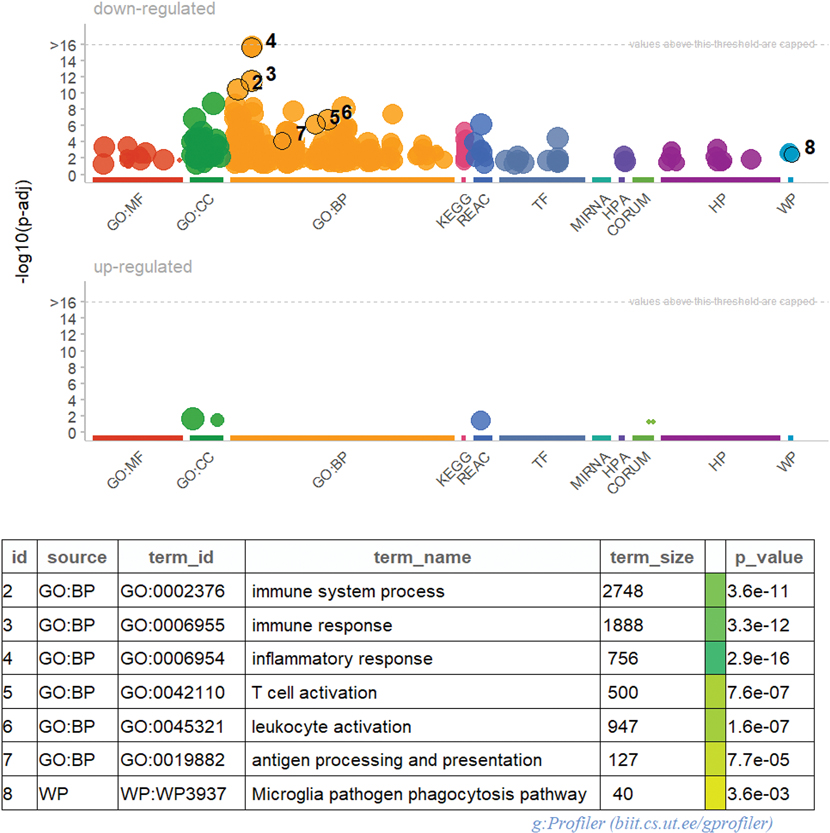
Figure 3. Functional profiling of upregulated and downregulated genes (FDR<0.05) in LET animals.
The open source version of gProfiler (g:GOSt) (Raudvere et al., 2019) enables functional profiling of genes significantly downregulated in LET animals. Each circle dot reflects a specific gene ontology (GO) term within eleven distinct categories: molecular function (MF), cellular component (CC), biological processes (BP), KEGG pathways (KEGG), reactome (REAC), transcription binding sites (TF), genes targeted by miRNAs (MIRNA), and their respective adjusted p-values. The BP category returned the most categories with 1,174 GO terms.
Results
Differential gene expression and enrichment analysis
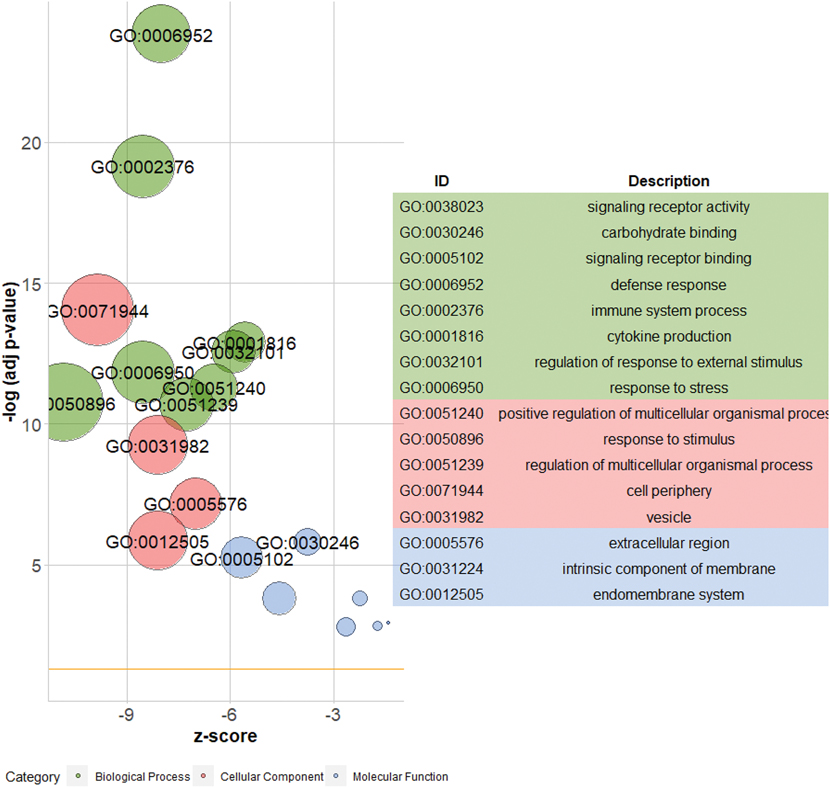
To better understand the effect of AIs on hippocampal gene expression, we used bulk RNA-seq to examine mRNA expression in the hippocampus of 16 marmosets treated with Letrozole or vehicle. We found 556 genes downregulated in LET animals with 125 of these genes significantly downregulated (FDR < 0.05) (Figure 1). Next, we sought to identify the proportion of differentially expressed genes belonging to established gene ontology (GO) categories to determine if Letrozole treatment increased the prevalence of certain genes relative to what might occur by chance. Genes with an FDR < 0.05 were submitted to gProfiler for functional enrichment analyses to focus on four GO categories: biological process (BP), molecular function (MF), cellular components (CC) and reactome (REAC). For downregulated genes in LET animals, the BP category returned the most GO terms (1,174) (Figure 3). Of this downregulated gene list, the most significantly enriched processes were carbohydrate binding, signaling receptor binding, defense response, immune response and MHC Class II protein complex binding as part of the BP, CC and MF categories (Figure 3; Table 1-1). To reduce the redundancy of returned GO terms, we used REVIGO to cluster GO terms which share a significant number of overlapping genes and graphed the most significant terms (Figure 4). The upregulated transcripts in LET animals returned few GO terms (28) across three categories: CORUM database (CORUM), MF and CC (Table 1). To visualize the overlap between highly differentially expressed genes and their corresponding GO categories, we plotted transcripts with a logFC < −1 from six different highly downregulated processes (Figure 5).
Table 1.
Functional profiling results for upregulated genes in LET animals.
| id | source | term_id | term_name | term_size | intersection_size | p_value | |
|---|---|---|---|---|---|---|---|
| 1 | CORUM | CORUM:1140 | Complexin complex (STX3, CPLX1, CPLX3) | 3 | 1 | 1.5e-02 | |
| 2 | CORUM | CORUM:1138 | SNARE complex (VAMP2, SNAP25, STX1a, CPLX3, CPLX4) | 5 | 1 | 1.5e-02 | |
| 3 | CORUM | CORUM:1137 | SNARE complex (VAMP2, SNAP25, STX1a, CPLX1, CPLX3) | 5 | 1 | 1.5e-02 | |
| 4 | CORUM | CORUM:6571 | KCNE1-KCNH2 complex | 2 | 1 | 1.5e-02 | |
| 5 | CORUM | CORUM:7507 | Potassium channel complex (KCNB1, KCNE1) | 2 | 1 | 1.5e-02 | |
| 6 | CORUM | CORUM:1139 | SNARE complex (VAMP2, SNAP25, STX1a, STX3, CPLX1, CPLX3, CPLX4) | 7 | 1 | 1.6e-02 | |
| 7 | GO:CC | GO:0005576 | extracellular region | 4303 | 11 | 6.7e-04 | |
| 8 | GO:CC | GO:0098093 | anchored component of synaptic vesicle membrane | 12 | 2 | 3.8e-03 | |
| 9 | GO:CC | GO:0031225 | anchored component of membrane | 171 | 3 | 2.2e-02 | |
| 10 | GO:CC | GO:0098563 | intrinsic component of synaptic vesicle membrane | 42 | 2 | 2.5e-02 | |
| 11 | GO:CC | G0:0098590 | plasma membrane region | 1237 | 6 | 3.4e-02 | |
| 12 | GO:CC | GO:0099029 | anchored component of presynaptic active zone membrane | 3 | 1 | 3.4e-02 | |
| 13 | GO:CC | GO:0005615 | extracellular space | 3353 | 13 | 3.7e-02 | |
| 14 | GO:CC | GO:0098684 | photoreceptor ribbon synapse | 7 | 1 | 4.5e-02 | |
| 15 | GO:CC | GO:0099026 | anchored component of presynaptic membrane | 7 | 1 | 4.5e-02 | |
| 16 | GO:MF | GO:0005109 | frizzled binding | 38 | 2 | 4.0e-02 | |
| 17 | GO:MF | GO:0005125 | cytokine activity | 235 | 4 | 4.0e-02 | |
| 18 | GO:MF | GO:0030546 | signaling receptor activator activity | 502 | 5 | 4.0e-02 | |
| 19 | GO:MF | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs | 2 | 1 | 4.0e-02 | |
| 20 | GO:MF | GO:0062183 | all-trans retinoic acid 18-hydroxylase activity | 2 | 1 | 4.0e-02 | |
| 21 | GO:MF | GO:0062182 | all-trans retinoic acid 4-hydrolase activity | 1 | 1 | 4.0e-02 | |
| 22 | GO:MF | GO:0048018 | receptor ligand activity | 495 | 5 | 4.0e-02 | |
| 23 | GO:MF | GO:0043035 | chromatin insulator sequence binding | 5 | 1 | 4.0e-02 | |
| 24 | GO:MF | GO:0008188 | neuropeptide receptor activity | 46 | 2 | 5.0e-02 | |
| 25 | GO:MF | GO:0038062 | protein tyrosine kinase collagen receptor activity | 3 | 1 | 5.0e-02 | |
| 26 | GO:MF | GO:0004952 | dopamine neurotransmitter receptor activity | 5 | 1 | 5.0e-02 | |
| 27 | GO:MF | GO:0030545 | signaling receptor regulator activity | 548 | 5 | 5.0e-02 | |
| 28 | GO:MF | GO:0031729 | CCR4 chemokine receptor binding | 2 | 1 | 5.0e-02 |
The open source web version of gProfiler (Raudvere et al., 2019) was used to profile upregulated genes in LET animals. Our results highlight 28 terms which were significant (adjusted p-value < 0.05) across three distinct source categories: CORUM database (CORUM), cellular component (CC), and molecular function (MF). Term id refers to the Journal Pre-proof unique seven-digit identifier of the gene ontology term name while the term size refers to the number of genes listed within each gene ontology term. Intersection size refers to the number of significantly enriched transcripts within LET animals.
Figure 4. Top GO terms for downregulated genes in three GO categories: biological processes (BP); cellular component (CC); molecular function (MF).
GOplot (Walter et al., 2015) was used in R to reduced redundant or significantly overlapping GO IDs in each category. Size of the GO term bubble indicates number of significantly downregulated genes within GO term ID.
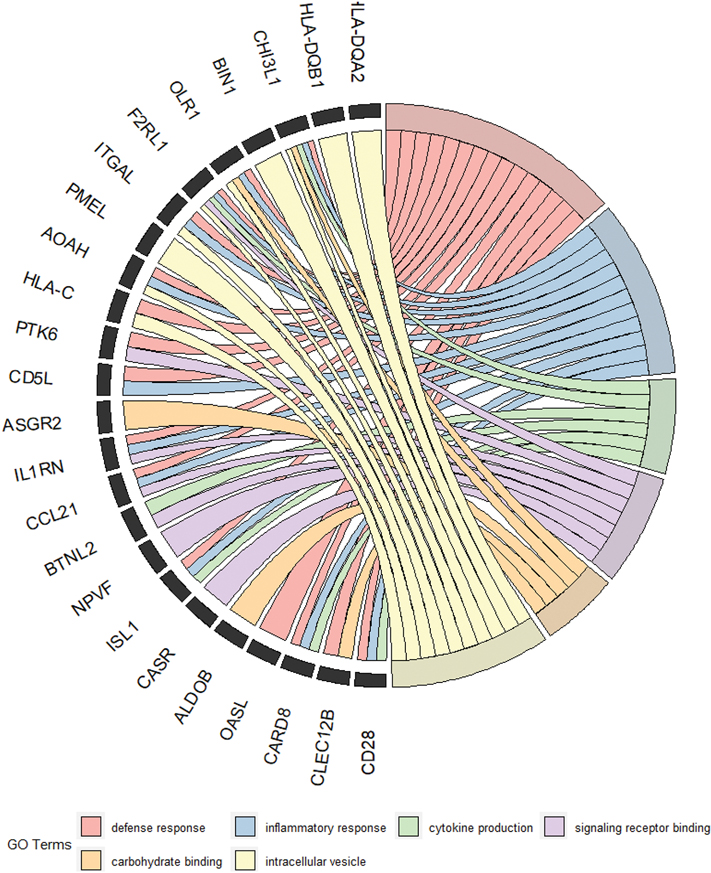
Figure 5. Visualization of interactions between significant GO terms and overlapping genes with a logFC ≤ −1.
A circular ribbon chat depicts interaction between top significant GO terms and genes with a logFC < −1. Ribbons connect individual genes on the left to their appropriate, color-coded GO category on the right. These gene names reflect some of the most significant transcripts that were affected within each GO category.
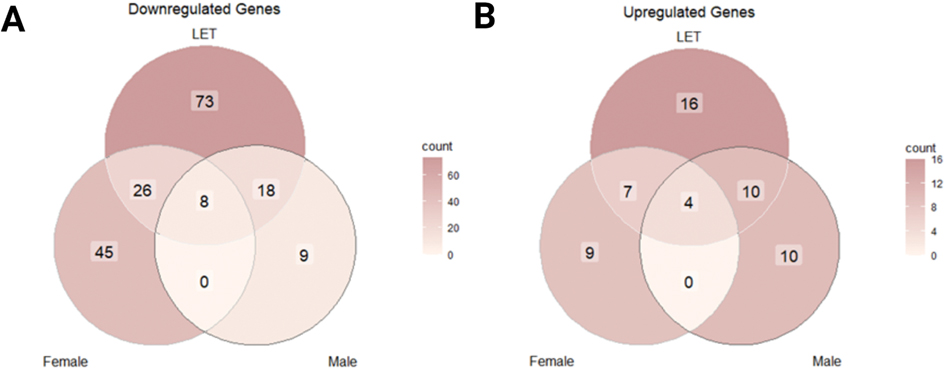
Next, we examined whether treatment effects on gene expression varied by sex. We found a fivefold difference in the number of unique downregulated genes in LET females (45) relative to LET males (9) (Figure 6A). Only eight downregulated genes were shared among LET females, LET males and the aggregation of both groups. We functionally profiled the 45 genes uniquely downregulated in LET females using the g:GOst functionality of gprofiler2 and found 423 significant GO terms associated with these genes with many involved in cell secretion and exocytosis (Figure 7; Figure 7-1). These results highlight significant sex differences in the brain in response to systemic Letrozole treatment, suggesting that different mechanisms for drug responsivity exist in males and females.
Figure 6. Venn Diagram of downregulated transcripts.
(A) Downregulated transcripts are shown for LET (Letrozole-treated animals relative to vehicle), Male (Letrozole-treated males relative to vehicle males), and Female (Letrozole-treated females relative to vehicle females) (FDR < 0.05). Only 8 transcripts were found to be shared between all contrasted groups while females had a five-fold difference in uniquely downregulated transcripts (45) than males (9). (B) Significantly fewer genes were upregulated across all groups.
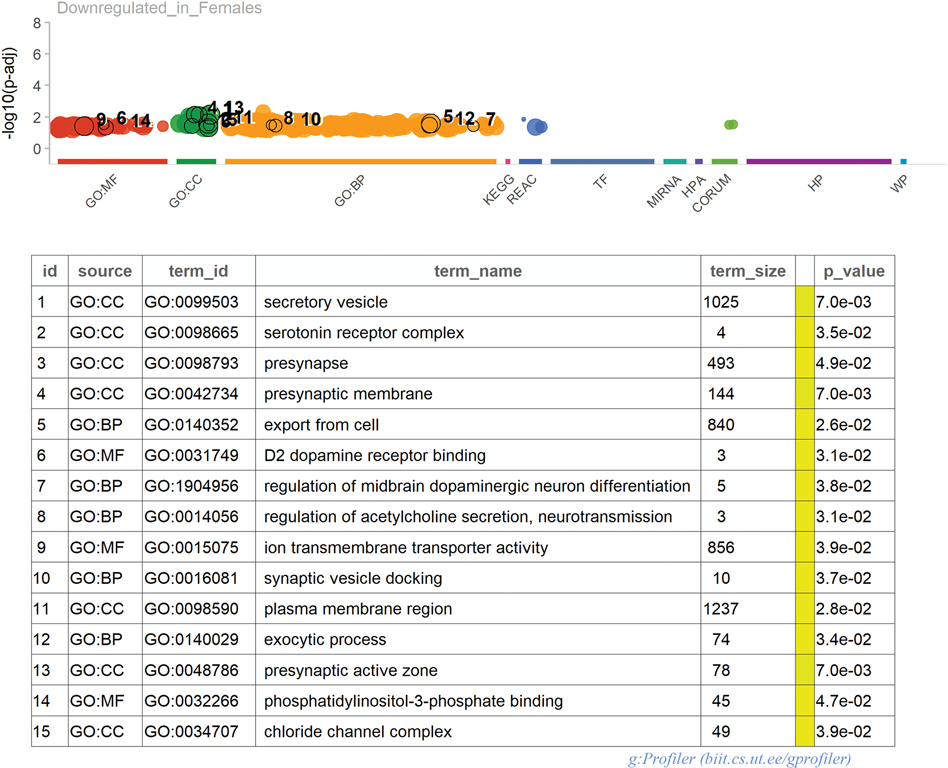
Figure 7. Functional profiling of uniquely downregulated genes in LET Females.
Using gOSt functionality as part of the gProfiler2 package (Raudvere et al., 2019), 45 downregulated genes unique to the LET females were submitted for functional profiling.
Gene set testing
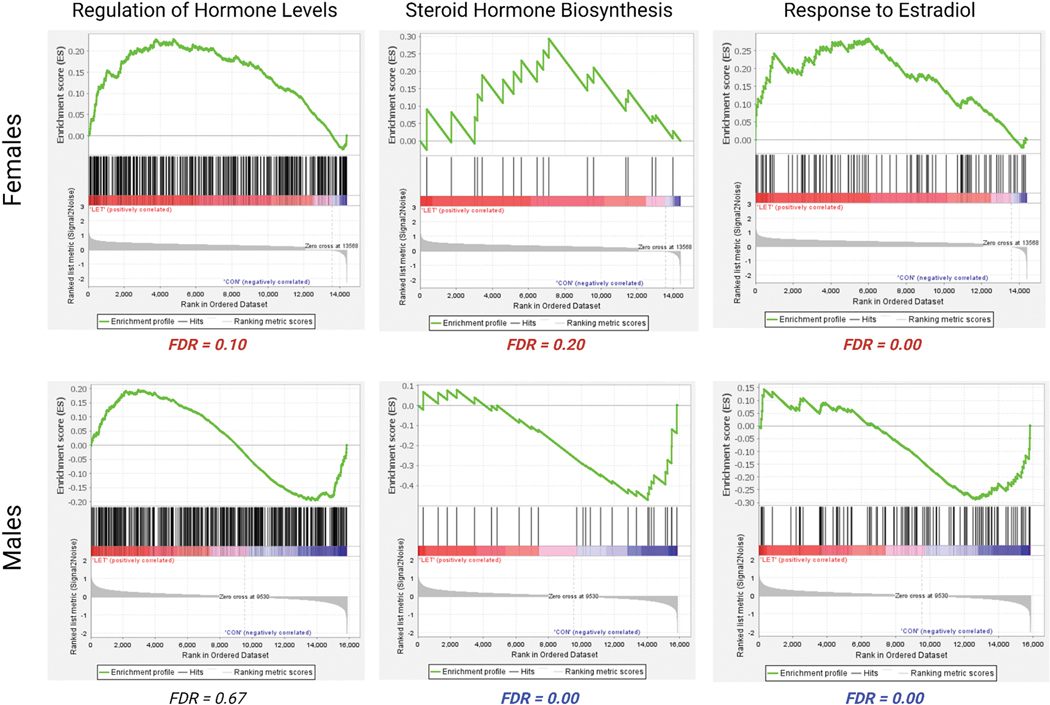
Previously, Gervais et al. (2019) found that LET animals, particularly females, had increased levels of 17β-estradiol in the hippocampus relative to vehicle-treated animals. Therefore, we used GSEAs to evaluate the effect of sex and treatment on genes involved in hormone metabolism and synthesis. We used curated gene sets from the Molecular Signatures Database (MSigDB) (Subramanian et al., 2005) involved in the regulation of hormone levels (M11178), response to estradiol (M105090 and steroid hormone biosynthesis (M14933). These gene sets were all enriched in LET females compared to VEH females (Figure 8) In males, no hormonal gene sets were enriched in the LET group, however one gene set was enriched in vehicle-treated males: steroid hormone biosynthesis (FDR = 0.00) (Figure 8). These results suggest that compensatory mechanisms to synthesize estradiol may be uniquely recruited or enriched in the brains of females.
Figure 8. Gene set enrichment analyses reveal sex differences.
LET females show enrichment for genes involved in the regulation of hormone levels (M11178), steroid hormone biosynthesis (M14933) and response to estradiol (M10509) compared to vehicle-treated females. LET males do not show enrichment for genes involved in the regulation of hormone levels compared to control males (FDR = 0.67) and only vehicle-treated males show enrichment for genes involved in steroid hormone biosynthesis and response to estradiol.
Due to the significant species differences in Cytochrome P450 distribution and drug metabolism (Shi et al., 2016; Stamou et al., 2013) we chose to look at a Cytochrome P450 drug metabolism gene set (M9257) which included members of the UDP glucuronosyltransferase family as well as Glutathione S-Transferase family and others involved in pathways responsible for processing pharmacologically active compounds. Interestingly, when we ran a GSEA for Cytochrome P450 drug metabolism, we found an enrichment in LET males that was not observed in LET females (Figure 9). These results are consistent with studies which have shown that Letrozole has a longer half-life in female Sprague-Dawley rats (~ 34 hours) than male rats (~ 9 hours), both in the plasma and brain extracellular fluid (Arora et al., 2021), suggesting a more efficient clearance or metabolism of Letrozole by males. Notably, mRNA expression in the human liver differs significantly by sex with 70% of differentially expressed genes showing a female bias (Zhang et al., 2011). It is possible that different outcomes in hippocampal gene expression are related to the processing of Letrozole by the liver and disparate metabolic rates in other peripheral organs and the brain.
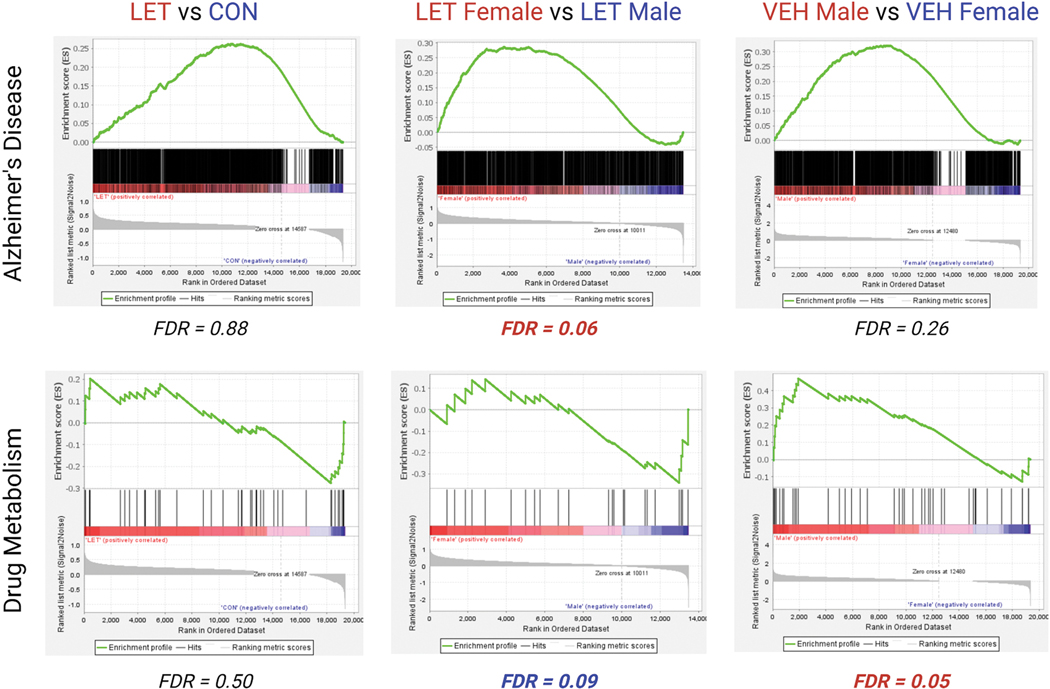
Figure 9. Gene set testing results reveal sex differences not observed in aggregated group.
Gene set for Alzheimer’s disease (M12921) shows no effect when LET animals are contrasted against control animals. (FDR = 0.88). Gene set testing for Alzheimer’s disease (AD) and drug metabolism show no effect on LET animals relative to VEH animals (FDR > 0.25). Interestingly, an enrichment for Alzheimer’s disease was found for LET females, while genes involved in drug metabolism (M9257) were enriched in LET males (FDR = 0.06; FDR = 0.09). Finally, there was no effect when comparing VEH males and VEH females for genes enriched in Alzheimer’s disease, however the drug metabolism gene set was also enriched in VEH males (FDR = 0.05).
Lastly, we used a gene set for genes involved in Alzheimer’s disease (1600 genes) to evaluate whether transcripts involved in Alzheimer’s disease would be enriched in all LET animals relative to vehicle-treated animals. We found no effect, however this gene set was enriched in LET females when compared to vehicle-treated females (Figure 9). This effect was not found when comparing LET males to vehicle-treated males. These results highlight unique mRNA enrichment in females that may predispose women to neurodegeneration if given prolonged Letrozole treatment (Rosenfeld et al., 2018).
Discussion
Our results show that long-term Letrozole administration significantly reduced transcripts involved in many biological, cellular and molecular processes such as immune signaling, carbohydrate binding, cellular homeostasis, phagocytosis, MHC protein complex binding and defense response. The observed decrease in immune response and previously reported increase of estradiol in the brains of LET animals are consistent with studies which have highlighted estradiol’s role in dampening immune function, particularly in microglia (Vegeto et al., 2003; Wu et al., 2016; for review see Villa et al., 2016). Activated microglia are often characterized by increased expression of major histocompatibility complex (MHC) class II proteins, pro-inflammatory cytokines, carbohydrate binding and induce effector T-cell function (Dimayuga et al., 2005; Linnartz et al., 2012; Schetters et al., 2018), all of which were significantly enriched GO terms revealed by functional profiling of downregulated genes in LET marmosets (Table 1-1). This suppression of immune response may be particularly harmful with aging, as intact immune responses may be crucial to protect against age-related accumulation of amyloid-beta in the brain (Palazzi et al., 2006; Villemagne et al., 2013).
Evidence for Letrozole-induced estrogenic regulation of cellular responses
Our analyses showed 28 enriched GO terms for upregulated genes in LET animals with many significant GO terms relating to cell signaling (Table 1). Genes listed within these GO terms are primarily involved in regulating neurotransmitter release, SNARE complex, cell binding and xenobiotic recognition. Interestingly, several of the upregulated genes in LET animals, including WNT genes and SCGB1D1, have also been shown to have estrogen responsive elements (EREs) or are transcriptionally regulated by steroid hormones (Bourdeau et al., 2004). These results suggest a concurrent depression of immune response and increase in cell signaling, which may be tied to increased estradiol levels in the hippocampus (Gervais et al., 2019). Much of the literature on the neuromodulatory properties of estradiol have focused on its facilitation of excitatory neurotransmission in brain regions important for cognition and memory (for review, see Brandt & Rune, 2019; Frick et al., 2018). However, one limitation of bulk RNA-sequencing is the masking of effects for specific cell types; it is unclear which cell types may have increased signaling from the functional enrichment analyses.
A recent study in common marmosets showed that aromatase inhibition induced hypothalamic hypoestrogenemia (or a reduction in estradiol levels) and disrupted metabolic homeostasis; these effects were rescued by systemically administering capsules of estradiol (Kraynak et al., 2022). This ablation of hypothalamic estradiol may be due to the higher dose used (400 μg) by Kraynak et al. (2022) whereas, in the current study, we use a dose that more closely resembles the recommended dosing in cisgender women (20 μg). Our results suggest that a sub-maximal dose of Letrozole may not be sufficient to cross the blood brain barrier and/or impact de novo neuroestrogen synthesis.
Letrozole treatment reveals sex differences in DEGs
Sex-specific DEGs were also observed within LET animals. 79 genes were significantly downregulated (FDR < 0.05) in LET females relative to vehicle-treated females. Of these 79 genes, only 10% shared overlap with genes significantly downregulated in LET males relative to vehicle-treated males. Additionally, 45 genes were uniquely downregulated in females which did not share overlap with the aggregation of males and females (LET) or LET males. We functionally profiled these 45 genes and found the top GO terms reflected processes involved in inflammation, cell secretion and release of intracellular molecules (Figure 7; Figure 7–1). For the 9 genes uniquely downregulated in males, GO terms only returned 16 GO terms primarily involved in immune response, IL12A complexes and interleukin-17 binding. These important findings highlight sex-specific responsivity to aromatase inhibition, which other studies have also shown in rodents (Borbélyová et al., 2017; Kokras et al., 2018), fish (Villeneuve et al., 2009) and primates (Gervais et al., 2019).
Cytochrome P450, steroidogenesis and drug metabolism in the brain
Our results also highlight sex differences in GSEAs. LET females showed unique enrichment of genes involved in the regulation of hormone levels, cellular hormone metabolic processes and steroid hormone biosynthesis relative to vehicle-treated females—an effect that was not observed in LET males. Interestingly, the oxidation by Cytochrome P450 family gene set was enriched in LET males when compared to LET females, which suggests that sex differences could arise in AI metabolism especially if Letrozole metabolites crossed the blood brain barrier. Sex differences in the expression of Cytochrome P450 enzymes have been reported in the brains of primates (Scallet et al., 2005), rodents (Roselli & Resko, 1997; Stamou et al., 2013) and humans (Sakuma et al., 2009). Sex differences have also been found in the metabolism of clinical drugs (Sakuma et al., 2009; Stamou et al., 2013; Fuscoe et al., 2020). Approximately 70% of all clinical drugs are metabolized by the Cytochrome P450 superfamily, but in vitro studies have shown that Letrozole is primarily metabolized by the CYP2A6 enzyme (Bhatnagar, 2007; Hertz et al., 2017). It is unclear whether this isoform is also responsible for metabolizing Letrozole in the marmoset brain.
Aromatase inhibition and susceptibility for neurodegenerative disease
An enrichment for Alzheimer’s disease-associated genes was found only in LET females when compared to LET males (Figure 9). Consistent with the cognitive impairments observed in women, Gervais et al. (2019) found cognitive impairments in marmosets treated with Letrozole and there are mounting concerns that AIs may exacerbate cognitive decline, particularly in postmenopausal people (Rosenfeld et al., 2018). Studies have shown that Letrozole treatment decreases mitochondrial volume, dendritic spine density, and synaptic proteins (Chang et al., 2013; Kretz et al., 2004). When Letrozole was combined with conformationally altered amyloid beta treatment in hippocampal neurons, these effects were further exacerbated (Chang et al., 2013). From our results indicating a decreased immune response, one hypothesis may be that decreased immune function via Letrozole treatment allows for greater aggregation of amyloid-beta plaques and other debris which would normally be cleared by proinflammatory glia (reviewed in Ries & Sastre, 2016).
Potential mechanisms underlying increased estradiol in the hippocampus
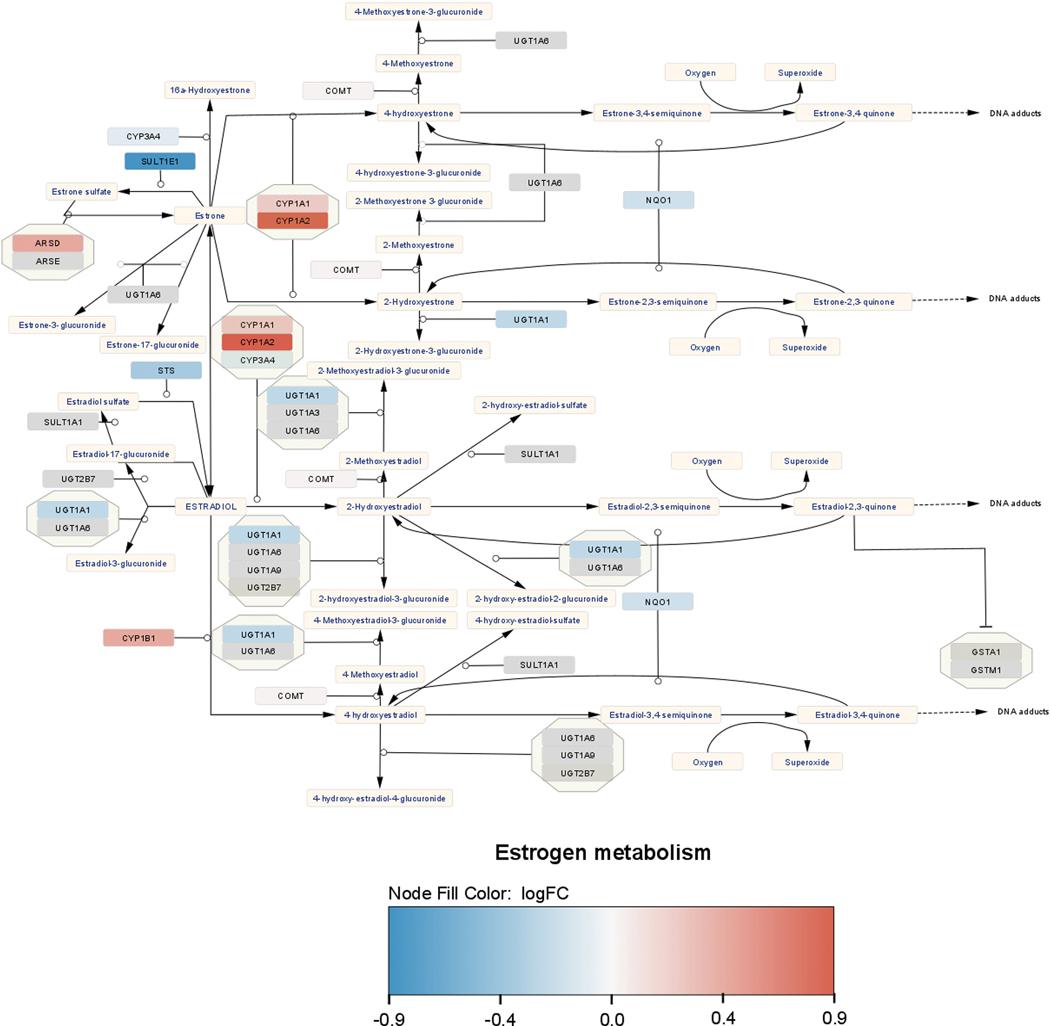
The mechanism by which Letrozole increased estradiol in the hippocampus of treated animals remains unknown. It is unclear whether the Letrozole dose used in this study was sufficient to cross the blood brain barrier and act directly upon cells in the hippocampus. It is possible that a compensatory mechanism involving sulfated estrogens in the “sulfatase pathway” (STS) may be utilized if the “aromatase pathway” is affected by Letrozole (Figure 10) (Kříž et al., 2008; Secky et al., 2013). In this pathway, biologically inactive steroid sulfates are precursors for estradiol synthesis whereas testosterone and other steroid hormones are converted into estradiol in the classic aromatase pathway (Kříž et al, 2008; Secky et al., 2013). These conjugated forms of steroids are often much more abundant than free steroids and differ in their non-genomic and genomic effects on cellular subtypes (Kim et al., 2015; for review see Vitku et al., 2022). Previous work has shown that tumor cells treated with Letrozole respond by upregulating steroid sulphatase expression and activity (Foster, 2021). In the current study, it is possible that circulating estrone sulfates (E1S) and estrogen sulfates (E2S) were converted into forms of estradiol with varying affinities for estrogen targets (Figure 10). Although Letrozole has been shown to affect serum E2 and E1S in the periphery (Dixon et al., 2008; Folkerd et al., 2012) there is little evidence that this mechanism exists in the primate brain, particularly in non-cancerous tissue microenvironments in pre-menopausal, post-menopausal or gonadectomized animal subjects.
Figure 10. Estrogen metabolism pathway for LET animals.
Using Cytoscape, the enrichment of genes involved in the estrogen metabolism pathway are shown with up- and downregulated transcripts. Metabolic byproducts (e.g. estradiol) are shown in the light tan, rectangular boxes. Black arrows denote the metabolic byproducts while the enzymes responsible for these products are shown in octagonal shapes colored in red (upregulated), blue (downregulated) or gray (no effect).
The observed increase in hippocampal E2 (Gervais et al., 2019) may also reflect de novo synthesis in order to compensate for low peripheral levels of estradiol. Due to the unique blood brain barrier properties of circumventricular organs (i.e. the hypothalamus and pituitary), it is also possible that these areas are more sensitive to aromatase inhibition and could respond in a way that affects gene expression in neighboring or distal brain regions. Aromatase inhibitors are frequently used to stimulate ovulation (for review, see Yang et al., 2021) and their usage has been shown to increase levels of luteinizing hormone (LH) and follicle stimulating hormone (FSH) (Kucherov et al., 2011). The rodent hippocampus harbors LH and FSH receptors (Chu et al., 2007) and blocking FSH receptors in the hippocampus has been shown to improve cognition in mice (Xiong et al., 2022). Thus, aromatase inhibition could indirectly affect de novo E2 synthesis the hippocampus via hypothalamic–pituitary–gonadal (HPG) axis modulation. Future studies will need to establish the precise feedback mechanism(s) in which the hippocampus and other brain regions detect and respond to aromatase inhibitors and adjust de novo synthesis of hormones.
Lastly, it is unknown which cell types may be responsible for the increase in brain estradiol in LET animals—a response that seems to be driven by females. One hypothesis is that neuroglia and other immune cells in the brain respond to xenobiotics and drugs by releasing estrogens to maintain blood brain barrier integrity. Studies have shown that ovariectomy and age increase the permeability of the blood brain barrier (Wilson et al., 2008) and that Letrozole can cross the blood brain barrier in a dose dependent manner in rats (Dave et al., 2013). It is possible that immune cells—namely astrocytes—synthesize estradiol in response to gonadectomy and/or drug treatment without entering a hyperactive, damaging state (Prat et al., 2011). Estrogens can exert pro- or anti-inflammatory effects depending on sex, cell type, estrogen levels and the status of the surrounding microenvironment (Dragin et al., 2017). Collectively, the actions of immune and glial cells may help to explain sex differences observed in our gene set testing as well as sex differences in aging trajectories (for review see Li et al., 2014; Villa et al., 2016.)
There are a few key limitations and considerations for the current study. While bulk RNA-sequencing is a powerful tool for gene expression quantification and analysis, transcripts are also not always predictive of protein levels or enzymatic activity (Koussounadis et al., 2015). Additionally, bulk RNA-sequencing may mask unique effects on cellular subtypes that respond differently to xenobiotics and steroid hormones. Our small sample size, especially for sex and treatment comparisons, in tandem with large inter-individual variability leads to low statistical power and may affect our inferences. Lastly, the importance of understanding peripheral influence on the brain cannot be overstated; future studies would greatly benefit from analyzing gene and protein expression in extra-gonadal tissues and additional brain areas to better understand how AIs are metabolized and their unique localized effects, which have implications on the brain and behavior.
In summary, our findings highlight treatment and sex differences in the hippocampus of animals treated with Letrozole. To our knowledge, no studies exist which have used nonhuman primates and long-term, oral Letrozole administration to look at gene expression changes in the brain. Our results are not consistent with studies on AIs which use in vitro and in vivo methods to assess the effects of Letrozole treatment on the brain (Bian et al., 2014; Kretz et al., 2004; Prange-Kiel et al., 2006). This may be due to the usage of isolated slice cultures—which remove the interaction between peripheral and central nervous systems—and species-level differences in cytochrome P450 enzyme distribution (Stamou et al., 2013). There are many additional factors which may explain the conflicting results in study outcomes, such as species used, drug dose, length of treatment, gonadal status, body mass index (Folkerd et al., 2012), age, type of AI used, sex and route of drug administration. Regional brain differences have also been reported with synaptic proteins more robustly downregulated in CA1 neurons than CA3 neurons (Prange-Kiel et al., 2006).
Studies using oral Letrozole administration in nonhuman primates—which are behaviorally and neuroanatomically more similar to humans—are translationally advantageous for understanding how AIs impact human health. Future studies which leverage nonhuman primates are essential for understanding how aromatase inhibitors affect the brain and how crosstalk between the brain and periphery may affect patient behavior and biology.
Edwards et al. Highlights.
Transcripts involved in immunity were downregulated in LET animals’ hippocampus
Letrozole upregulated transcripts in hippocampus involved in cell signaling
Gene set testing revealed sex differences in Letrozole treatment
Systemic aromatase inhibition may predispose patients to CNS impairment
Acknowledgements:
This study was supported in part by NIH R01 AG053841. Mélise Edwards was supported by the Ford Fellowship. We would like to acknowledge Dr. Dave Follette and the Advanced Digital Design & Fabrication Core Facility (ADDFab) at the Institute for Applied Life Sciences at UMass Amherst for help with 3D-printing brain matrices, the Genomics Resource Laboratory at UMass Amherst and Dr. Alfonso Silva and colleagues for generously sharing their original marmoset brain matrix design (Guy et 17 al., 2016).
Funding:
This work was funded in part by NIH R01 AG046266.
Footnotes
Code Accessibility
All code available upon request. FASTQ files available in extended data and NCBI SRA.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Anders S, Pyl PT, & Huber W (2014). HTSeq--a Python framework to work with high-throughput sequencing data. Bioinformatics, 31(2), 166–169. 10.1093/bioinformatics/btu638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arora P, Gudelsky G, & Desai PB (2021). Gender-based differences in brain and plasma pharmacokinetics of letrozole in sprague-dawley rats: Application of physiologically-based pharmacokinetic modeling to gain quantitative insights. PLOS ONE, 16(4), e0248579. 10.1371/journal.pone.0248579 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bailey DJ, Ma C, Soma KK, & Saldanha CJ (2013). Inhibition of hippocampal aromatization impairs spatial memory performance in a male songbird. Endocrinology, 154(12), 4707–4714. 10.1210/en.2013-1684 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhatnagar AS (2007). The discovery and mechanism of action of letrozole. Breast Cancer Research and Treatment, 105(S1), 7–17. 10.1007/s10549-007-9696-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bian C, Zhao Y, Guo Q, Xiong Y, Cai W, & Zhang J (2014). Aromatase inhibitor letrozole downregulates steroid receptor coactivator-1 in specific brain regions that primarily related to memory, neuroendocrine and integration. The Journal of Steroid Biochemistry and Molecular Biology, 141, 37–43. 10.1016/j.jsbmb.2013.12.020 [DOI] [PubMed] [Google Scholar]
- Borbélyová V, Domonkos E, Csongová M, Kačmárová M, Ostatníková D, Celec P, & Hodosy J (2017). Sex-dependent effects of letrozole on anxiety in middle-aged rats. Clinical and Experimental Pharmacology and Physiology, 44, 93–98. 10.1111/1440-1681.12731 [DOI] [PubMed] [Google Scholar]
- Bourdeau V, Deschênes J, Métivier R, Nagai Y, Nguyen D, Bretschneider N, Gannon F, White JH, & Mader S (2004). Genome-Wide Identification of High-Affinity Estrogen Response Elements in Human and Mouse. Molecular Endocrinology, 18(6), 1411–1427. 10.1210/me.2003-0441 [DOI] [PubMed] [Google Scholar]
- Brandt N, & Rune GM (2019). Sex-dependency of oestrogen-induced structural synaptic plasticity: Inhibition of aromatase versus application of estradiol in rodents. European Journal of Neuroscience, 52(1), 2548–2559. 10.1111/ejn.14541 [DOI] [PubMed] [Google Scholar]
- Chang PKY, Boridy S, McKinney RA, & Maysinger D (2013). Letrozole Potentiates Mitochondrial and Dendritic Spine Impairments Induced byβAmyloid. Journal of Aging Research, 2013, 1–11. 10.1155/2013/538979 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Lun ATL, & Smyth GK (2016). From reads to genes to pathways: differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Research, 5, 1438. 10.12688/f1000research.8987.2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu C, Gao G & Huang W (2008). A study on co-localization of FSH and its receptor in rat hippocampus. J Mol Hist, 39, 49–55. 10.1007/s10735-007-9125-2 [DOI] [PubMed] [Google Scholar]
- Cui J, Shen Y, & Li R (2013). Estrogen synthesis and signaling pathways during aging: from periphery to brain. Trends in Molecular Medicine, 19(3), 197–209. 10.1016/j.molmed.2012.12.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dave N, Gudelsky GA, & Desai PB (2013). The pharmacokinetics of letrozole in brain and brain tumor in rats with orthotopically implanted C6 glioma, assessed using intracerebral microdialysis. Cancer Chemotherapy and Pharmacology, 72(2), 349–357. 10.1007/s00280-013-2205-y [DOI] [PubMed] [Google Scholar]
- Di Mauro M, Tozzi A, Calabresi P, Pettorossi VE, & Grassi S (2017). Different synaptic stimulation patterns influence the local androgenic and estrogenic neurosteroid availability triggering hippocampal synaptic plasticity in the male rat. European Journal of Neuroscience, 45(4), 499–509. 10.1111/ejn.13455 [DOI] [PubMed] [Google Scholar]
- Dixon JM, Renshaw L, Young O, Murray J, Macaskill EJ, McHugh M, Folkerd E, Cameron DA, A’Hern RP, & Dowsett M (2008). Letrozole Suppresses Plasma Estradiol and Estrone Sulphate More Completely Than Anastrozole in Postmenopausal Women with Breast Cancer. Journal of Clinical Oncology, 26(10), 1671–1676. 10.1200/jco.2007.13.9279 [DOI] [PubMed] [Google Scholar]
- Dimayuga FO, Reed JL, Carnero GA, Wang C, Dimayuga ER, Dimayuga VM, Perger A, Wilson ME, Keller JN, & Bruce-Keller AJ (2005). Estrogen and brain inflammation: Effects on microglial expression of MHC, costimulatory molecules and cytokines. Journal of Neuroimmunology, 161(1–2), 123–136. 10.1016/j.jneuroim.2004.12.016 [DOI] [PubMed] [Google Scholar]
- Dragin N, Nancy P, Villegas J, Roussin R, Le Panse R, & Berrih-Aknin S (2017). Balance between Estrogens and Proinflammatory Cytokines Regulates Chemokine Production Involved in Thymic Germinal Center Formation. Scientific Reports, 7(1). 10.1038/s41598-017-08631-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fabian CJ (2007). The what, why and how of aromatase inhibitors: hormonal agents for treatment and prevention of breast cancer. International Journal of Clinical Practice, 61(12), 2051–2063. 10.1111/j.1742-1241.2007.01587.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fester L, Prange-Kiel J, Zhou L, Blittersdorf BV, Böhm J, Jarry H, Schumacher M, & Rune GM (2012). Estrogen-regulated synaptogenesis in the hippocampus: Sexual dimorphism in vivo but not in vitro. The Journal of Steroid Biochemistry and Molecular Biology, 131(1–2), 24–29. 10.1016/j.jsbmb.2011.11.010 [DOI] [PubMed] [Google Scholar]
- Folkerd EJ, Dixon JM, Renshaw L, A’Hern RP, & Dowsett M (2012). Suppression of Plasma Estrogen Levels by Letrozole and Anastrozole Is Related to Body Mass Index in Patients With Breast Cancer. Journal of Clinical Oncology, 30(24), 2977–2980. 10.1200/jco.2012.42.0273 [DOI] [PubMed] [Google Scholar]
- Foster PA (2021). Steroid Sulphatase and Its Inhibitors: Past, Present and Future. Molecules, 26(10), 2852. 10.3390/molecules26102852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frick KM, Kim J, & Koss WA (2018). Estradiol and hippocampal memory in female and male rodents. Current Opinion in Behavioral Sciences, 23, 65–74. 10.1016/j.cobeha.2018.03.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuscoe JC, Vijay V, Hanig JP, Han T, Ren L, Greenhaw JJ, Beger RD, Pence LM, & Shi Q (2020). Hepatic Transcript Profiles of Cytochrome P450 Genes Predict Sex Differences in Drug Metabolism. Drug Metabolism and Disposition, 48(6), 447–458. 10.1124/dmd.119.089367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garreau JR, DeLaMelena T, Walts D, Karamlou K, & Johnson N (2006). Side effects of aromatase inhibitors versus tamoxifen: the patients’ perspective. The American Journal of Surgery, 192(4), 496–498. 10.1016/j.amjsurg.2006.06.018 [DOI] [PubMed] [Google Scholar]
- Gervais NJ, Remage-Healey L, Starrett JR, Pollak DJ, Mong JA, & Lacreuse A (2019). Adverse Effects of Aromatase Inhibition on the Brain and Behavior in a Nonhuman Primate. The Journal of Neuroscience, 39(5), 918–928. 10.1523/jneurosci.0353-18.2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guy JR, Sati P, Leibovitch E, Jacobson S, Silva AC, & Reich DS (2016). Custom fit 3D-printed brain holders for comparison of histology with MRI in marmosets. Journal of Neuroscience Methods, 257, 55–63. 10.1016/j.jneumeth.2015.09.002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haddad-Tóvolli R, Dragano NRV, Ramalho AFS, & Velloso LA (2017). Development and function of the blood-brain barrier in the context of metabolic control. Frontiers. https://www.frontiersin.org/articles/10.3389/fnins.2017.00224/full [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hertz DL, Henry NL, & Rae JM (2017). Germline genetic predictors of aromatase inhibitor concentrations, estrogen suppression and drug efficacy and toxicity in breast cancer patients. Pharmacogenomics, 18(5), 481–499. 10.2217/pgs-2016-0205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim H, Cam-Etoz B, Zhai G, Hubbard WJ, Zinn KR, & Chaudry IH (2015). Salutary Effects of Estrogen Sulfate for Traumatic Brain Injury. Journal of Neurotrauma, 32(16), 1210–1216. 10.1089/neu.2014.3771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kokras N, Pastromas N, Papasava D, de Bournonville C, Cornil C, & Dalla C (2018). Sex differences in behavioral and neurochemical effects of gonadectomy and aromatase inhibition in rats. Psychoneuroendocrinology, 87, 93–107. 10.1016/j.psyneuen.2017.10.007 [DOI] [PubMed] [Google Scholar]
- Kolberg L, Raudvere U, Kuzmin I, Vilo J, & Peterson H (2020). gprofiler2 -- an R package for gene list functional enrichment analysis and namespace conversion toolset g:Profiler. F1000Research, 9, 709. 10.12688/f1000research.24956.2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koussounadis A, Langdon SP, Um IH, Harrison DJ, & Smith VA (2015). Relationship between differentially expressed mRNA and mRNA-protein correlations in a xenograft model system. Scientific Reports, 5(1). 10.1038/srep10775 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kraynak M, Willging MM, Kuehlmann AL, Kapoor AA, Flowers MT, Colman RJ, Levine JE, & Abbott DH (2022). Aromatase Inhibition Eliminates Sexual Receptivity Without Enhancing Weight Gain in Ovariectomized Marmoset Monkeys. Journal of the Endocrine Society, 6(6), bvac063. 10.1210/jendso/bvac063 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kretz O (2004). Hippocampal Synapses Depend on Hippocampal Estrogen Synthesis. Journal of Neuroscience, 24(26), 5913–5921. 10.1523/jneurosci.5186-03.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kříž L, Bičíková M, & Hampl R (2008). Roles of steroid sulfatase in brain and other tissues. Physiological Research, 657–668. 10.33549/physiolres.931207 [DOI] [PubMed] [Google Scholar]
- Langmead B, Trapnell C, Pop M, & Salzberg SL (2009). Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biology, 10(3), R25. 10.1186/gb-2009-10-3-r25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li R, Cui J, & Shen Y (2014). Brain sex matters: Estrogen in cognition and Alzheimer’s disease. Molecular and Cellular Endocrinology, 389(1–2), 13–21. 10.1016/j.mce.2013.12.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linnartz B, Bodea LG, & Neumann H (2012). Microglial carbohydrate-binding receptors for neural repair. Cell and Tissue Research, 349(1), 215–227. 10.1007/s00441-012-1342-7 [DOI] [PubMed] [Google Scholar]
- Liu M, Xing F, Bian C, Zhao Y, Zhao J, Liu Y, & Zhang J (2019). Letrozole induces worse hippocampal synaptic and dendritic changes and spatial memory impairment than ovariectomy in adult female mice. Neuroscience Letters, 706, 61–67. 10.1016/j.neulet.2019.05.006 [DOI] [PubMed] [Google Scholar]
- Macedo-Lima M, & Remage-Healey L (2020). Auditory learning in an operant task with social reinforcement is dependent on neuroestrogen synthesis in the male songbird auditory cortex. Hormones and Behavior, 121, 104713. 10.1016/j.yhbeh.2020.104713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marbouti L, Zahmatkesh M, Riahi E, & Sadr SS (2020). Inhibition of brain 17β-estradiol synthesis by letrozole induces cognitive decline in male and female rats. Neurobiology of Learning and Memory, 175, 107300. 10.1016/j.nlm.2020.107300 [DOI] [PubMed] [Google Scholar]
- Meyer R, Gehlhaus M, Knoth R, & Volk B (2007). Expression and Function of Cytochrome P450 in Brain Drug Metabolism. Current Drug Metabolism, 8(4), 297–306. 10.2174/138920007780655478 [DOI] [PubMed] [Google Scholar]
- Miksys S, & Tyndale R (2013). Cytochrome P450–mediated drug metabolism in the brain. Journal of Psychiatry & Neuroscience, 38(3), 152–163. 10.1503/jpn.120133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palazzi X, & Bordier N (2009). The Marmoset Brain in Stereotaxic Coordinates (2008th ed.). Springer. [Google Scholar]
- Palazzi X, Switzer R, & George C (2006). Natural Occurrence of Amyloid-Aβ Deposits in the Brain of Young Common Marmosets (Callithrix jacchus): A Morphological and Immunohistochemical Evaluation. Veterinary Pathology, 43(5), 777–779. 10.1354/vp.43-5-777 [DOI] [PubMed] [Google Scholar]
- Prange-Kiel J, Fester L, Zhou L, Lauke H, Carrétero J, & Rune GM (2006). Inhibition of hippocampal estrogen synthesis causes region-specific downregulation of synaptic protein expression in hippocampal neurons. Hippocampus, 16(5), 464–471. 10.1002/hipo.20173 [DOI] [PubMed] [Google Scholar]
- Prat A, Behrendt M, Marcinkiewicz E, Boridy S, Sairam RM, Seidah NG, & Maysinger D (2011). A Novel Mouse Model of Alzheimer’s Disease with Chronic Estrogen Deficiency Leads to Glial Cell Activation and Hypertrophy. Journal of Aging Research, 2011, 1–12. 10.4061/2011/251517 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raudvere U, Kolberg L, Kuzmin I, Arak T, Adler P, Peterson H, & Vilo J (2019). g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Research, 47(W1), W191–W198. 10.1093/nar/gkz369 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ries M, & Sastre M (2016). Mechanisms of Aβ Clearance and Degradation by Glial Cells. Frontiers in Aging Neuroscience, 8. 10.3389/fnagi.2016.00160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MD, & Smyth GK (2007). Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics, 9(2), 321–332. 10.1093/biostatistics/kxm030 [DOI] [PubMed] [Google Scholar]
- Robinson MD, McCarthy DJ, & Smyth GK (2009). edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26(1), 139–140. 10.1093/bioinformatics/btp616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roselli CE, & Resko JA (1997). Sex differences in androgen-regulated expression of cytochrome P450 aromatase in the rat brain. The Journal of Steroid Biochemistry and Molecular Biology, 61(3–6), 365–374. 10.1016/s0960-0760(97)80034-9 [DOI] [PubMed] [Google Scholar]
- Rosenfeld CS, Shay DA, & Vieira-Potter VJ (2018). Cognitive Effects of Aromatase and Possible Role in Memory Disorders. Frontiers in Endocrinology, 9. 10.3389/fendo.2018.00610 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakuma T, Kawasaki Y, Jarukamjorn K, & Nemoto N (2009). Sex Differences of Drug-metabolizing Enzyme: Female Predominant Expression of Human and Mouse Cytochrome P450 3A Isoforms. Journal of Health Science, 55(3), 325–337. 10.1248/jhs.55.325 [DOI] [Google Scholar]
- Scallet AC, Muskhelishvili L, Slikker W, & Kadlubar FF (2005). Sex differences in cytochrome P450 1B1, an estrogen-metabolizing enzyme, in the rhesus monkey telencephalon. Journal of Chemical Neuroanatomy, 29(1), 71–80. 10.1016/j.jchemneu.2004.09.003 [DOI] [PubMed] [Google Scholar]
- Schetters STT, Gomez-Nicola D, Garcia-Vallejo JJ, & van Kooyk Y (2018). Neuroinflammation: Microglia and T Cells Get Ready to Tango. Frontiers in Immunology, 8. 10.3389/fimmu.2017.01905 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Secky L, Svoboda M, Klameth L, Bajna E, Hamilton G, Zeillinger R, Jäger W, & Thalhammer T (2013). The Sulfatase Pathway for Estrogen Formation: Targets for the Treatment and Diagnosis of Hormone-Associated Tumors. Journal of Drug Delivery, 2013, 1–13. 10.1155/2013/957605 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi X, Zhang G, Mackie B, Yang S, Wang J, & Shan L (2016). Comparison of the in vitro metabolism of psoralidin among different species and characterization of its inhibitory effect against UDP- glucuronosyltransferase (UGT) or cytochrome p450 (CYP450) enzymes. Journal of Chromatography B, 1029–1030, 145–156. 10.1016/j.jchromb.2016.06.031 [DOI] [PubMed] [Google Scholar]
- Stamou M, Wu X, Kania-Korwel I, Lehmler HJ, & Lein PJ (2013). Cytochrome P450 mRNA Expression in the Rodent Brain: Species-, Sex-, and Region-Dependent Differences. Drug Metabolism and Disposition, 42(2), 239–244. 10.1124/dmd.113.054239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, & Mesirov JP (2005). Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences, 102(43), 15545–15550. 10.1073/pnas.0506580102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuscher JJ, Szinte JS, Starrett JR, Krentzel AA, Fortress AM, Remage-Healey L, & Frick KM (2016). Inhibition of local estrogen synthesis in the hippocampus impairs hippocampal memory consolidation in ovariectomized female mice. Hormones and Behavior, 83, 60–67. 10.1016/j.yhbeh.2016.05.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vegeto E, Belcredito S, Etteri S, Ghisletti S, Brusadelli A, Meda C, Krust A, Dupont S, Ciana P, Chambon P, & Maggi A (2003). Estrogen receptor-α mediates the brain antiinflammatory activity of estradiol. Proceedings of the National Academy of Sciences, 100(16), 9614–9619. 10.1073/pnas.1531957100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villa A, Vegeto E, Poletti A, & Maggi A (2016). Estrogens, Neuroinflammation, and Neurodegeneration. Endocrine Reviews, 37(4), 372–402. 10.1210/er.2016-1007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villemagne VL, Burnham S, Bourgeat P, Brown B, Ellis KA, Salvado O, Szoeke C, Macaulay SL, Martins R, Maruff P, Ames D, Rowe CC, & Masters CL (2013). Amyloid β deposition, neurodegeneration, and cognitive decline in sporadic Alzheimer’s disease: a prospective cohort study. The Lancet Neurology, 12(4), 357–367. 10.1016/s1474-4422(13)70044-9 [DOI] [PubMed] [Google Scholar]
- Villeneuve DL, Wang RL, Bencic DC, Biales AD, Martinović D, Lazorchak JM, Toth G, & Ankley GT (2009). ALTERED GENE EXPRESSION IN THE BRAIN AND OVARIES OF ZEBRAFISH (DANIO RERIO) EXPOSED TO THE AROMATASE INHIBITOR FADROZOLE: MICROARRAY ANALYSIS AND HYPOTHESIS GENERATION. Environmental Toxicology and Chemistry, 28(8), 1767. 10.1897/08-653.1 [DOI] [PubMed] [Google Scholar]
- Vierk R, Glassmeier G, Zhou L, Brandt N, Fester L, Dudzinski D, Wilkars W, Bender RA, Lewerenz M, Gloger S, Graser L, Schwarz J, & Rune GM (2012). Aromatase inhibition abolishes LTP generation in female but not in male mice. Journal of Neuroscience, 32(24), 8116–8126. 10.1523/jneurosci.5319-11.2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vitku J, Hill M, Kolatorova L, Kubala Havrdova E, & Kancheva R (2022). Steroid Sulfation in Neurodegenerative Diseases. Frontiers in Molecular Biosciences, 9. 10.3389/fmolb.2022.839887 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walter W, Sánchez-Cabo F, & Ricote M (2015). GOplot: an R package for visually combining expression data with functional analysis: Fig. 1. Bioinformatics, 31(17), 2912–2914. 10.1093/bioinformatics/btv300 [DOI] [PubMed] [Google Scholar]
- Wilson AC, Clemente L, Liu T, Bowen RL, Meethal SV, & Atwood CS (2008). Reproductive hormones regulate the selective permeability of the blood-brain barrier. Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, 1782(6), 401–407. 10.1016/j.bbadis.2008.02.011 [DOI] [PubMed] [Google Scholar]
- Wu SY, Chen YW, Tsai SF, Wu SN, Shih YH, Jiang-Shieh YF, Yang TT, & Kuo YM (2016). Estrogen ameliorates microglial activation by inhibiting the Kir2.1 inward-rectifier K+ channel. Scientific Reports, 6(1). 10.1038/srep22864 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiong J, Kang SS, Wang Z et al. (2022). FSH blockade improves cognition in mice with Alzheimer’s disease. Nature, 603, 470–476. 10.1038/s41586-022-04463-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Ai-Min, et al. (2021). “Letrozole for Female Infertility.” Frontiers. https://www.frontiersin.org/articles/10.3389/fendo.2021.676133/full. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zanger UM, & Schwab M (2013). Cytochrome P450 enzymes in drug metabolism: Regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacology & Therapeutics, 138(1), 103–141. 10.1016/j.pharmthera.2012.12.007 [DOI] [PubMed] [Google Scholar]
- Zhang Y, Klein K, Sugathan A, Nassery N, Dombkowski A, Zanger UM, & Waxman DJ (2011). Transcriptional Profiling of Human Liver Identifies Sex-Biased Genes Associated with Polygenic Dyslipidemia and Coronary Artery Disease. PLoS ONE, 6(8), e23506. 10.1371/journal.pone.0023506 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao J, Bian C, Liu M, Zhao Y, Sun T, Xing F, & Zhang J (2018). Orchiectomy and letrozole differentially regulate synaptic plasticity and spatial memory in a manner that is mediated by SRC-1 in the hippocampus of male mice. The Journal of Steroid Biochemistry and Molecular Biology, 178, 354–368. 10.1016/j.jsbmb.2018.02.007 [DOI] [PubMed] [Google Scholar]
- Zhou L, Fester L, von Blittersdorff B, Hassu B, Nogens H, Prange-Kiel J, Jarry H, Wegscheider K, & Rune GM (2010). Aromatase Inhibitors Induce Spine Synapse Loss in the Hippocampus of Ovariectomized Mice. Endocrinology, 151(3), 1153–1160. 10.1210/en.2009-0254 [DOI] [PubMed] [Google Scholar]