Abstract
Background
Multidrug-resistant (MDR) bacterial infections have become an emerging health concern around the world. Antibiotics resistance among S. pneumoniae strains increased recently contributing to increase in incidence of pneumococcal infection. This necessitates the discovery of novel antipnemococcal such as compound C3-005 which target the interaction between RNA polymerase and σ factors. Chitosan nanoparticles (CNPs) exhibited antibacterial activity including S. pneumonia. Therefore, the aims of the current investigation were to formulate CNPs loaded with C3-005 and characteristic their antimicrobial properties against S. pneumonia.
Methods
The CNPs and C3-005 loaded CNPs were produced utilizing ionic gelation method, and their physicochemical characteristics including particle size, zeta potential, polydispersity index (PDI), encapsulation efficiency (EE%), and in vitro release profile were studied. Both differential scanning calorimetry (DSC) and fourier transform infrared spectroscopy (FTIR) were used for chemical characterization. The synthesized NPs' minimum inhibitory concentration (MIC) was determined using killing assay and broth dilution method, and their impact on bacteria induced hemolysis were also studied.
Results
The NPs encapsulating C3-005 were successfully prepared with particle size of 343.5 nm ± 1.3, zeta potential of 29.8 ± 0.37, and PDI of 0.20 ± 0.03. 70 % of C3-005 were encapsulated in CNPs and sustained release pattern of C3-005 from CNPs was revealed by an in vitro release study. CNPs containing C3-005 exhibited higher antipnomcoccal activity with MIC50 of 30 µg/ml when compared with C3-005 and empty CNPs alone. The prepared C3-CNPs showed a reduction of bacterial hemolysis in a concentration-related (dependent) manner and was higher than C3-005 alone.
Conclusions
The findings of this study showed the potential for using C3-005 loaded CNPs to treat pneumococcal infection.
Keywords: Streptococcus pneumonia, Chitosan, C3-005, Hemolysis, Nanoparticles, Toxin
1. Introduction
Infections from multidrug-resistant (MDR) bacteria (superbugs) have become an emerging health concern around the world (Tacconelli and Pezzani 2019). S. pneumoniae causes severe infection including pneumonia, septicemia, and meningitis (Cherazard et al., 2017). However, antibiotics resistance among S. pneumoniae strains increased recently contributing to increase in incidence of pneumococcal infection (Blasi et al., 2012). Pneumococci use multiple virulence factors to mediate infection including pneumolysin which enhance bacterial colonization, nutrition scavenging, and immunoevasion (Jedrzejas 2001). Undesirable elevation in S. pneumoniae toxin levels in host upon exposure to bacteriolytic antimicrobials such as β-lactams was observed and had impact on the treatment outcomes (Anderson et al., 2007). Resistance to penicillin was initially described in S. pneumoniae in 1967, followed by quinolone, fluoroquinolone, and macrolide resistance (Kang and Song 2013). In response to the worldwide antibiotics resistance threat, the World Health Organization (WHO) announced its initial list of 12 priority infections, involving S. pneumoniae, that are causing havoc on human health and thus require treatment with newer antibacterial drugs (WHO 2017). This necessitates the discovery of novel antimicrobial with new target (Hinsberger et al., 2013, Ma et al., 2013, Murakami, 2015, Ye et al., 2019), as recently reported by Ye et al (2019) study. In their study, anti-pnemococcal activity of compound C3-005 was shown and found to target the interaction between σ factors and RNA polymerase (Ye et al., 2019). It exhibited superior antimicrobial activity when compared with other compounds with MIC of 8 µg/ml (Ye et al., 2019). C3-005 reduced the ATP generation in S. pneumoniae, similar to how rifampicin inhibits transcription of bacteria, in which ATP production is one of the hallmarks of a successful antibiotic (Ye et al., 2019). The release of pneumolysin toxin into the extracellular milieu is a characteristic pneumococcal virulence factor (Jedrzejas 2001). C3-005 inhibited S. pneumoniae's virulence factor pneumolysin secretion (Ye et al., 2019).
Nanoparticles (NPs) have received a lot of attention in terms of infectious disease treatment and diagnosis, because of their unique physicochemical features that improve medical treatment with more effective, less toxic, and smart outcomes (Baptista et al., 2018, Shah et al., 2022, Sriharan et al., 2022). Nanomaterials are structures, materials, and functional systems made up of particles with sizes ranging from 1 to 500 nm (Baptista et al., 2018, Gao et al., 2018, Lee et al., 2019). The implication of NPs in infectious disease treatment depends on these nanostructures functioning as antimicrobial carriers, via either incorporation into the nanoformulation or integration, or adsorption to the surface, in order to enhance drug therapeutic efficiency, drug biodistribution and pharmacokinetics, and to regulate drug release (Maleki and Kamalzare, 2014, Baptista et al., 2018, Gao et al., 2018, Maleki, 2018, Maleki et al., 2018, Eivazzadeh-Keihan et al., 2019, Lee et al., 2019, Maleki et al., 2019, Ahghari et al., 2020, Soltaninejad et al., 2021). Chitosan (Cs) showed antibacterial properties toward both Gram-positive and Gram-negative microorganisms (Rabea et al., 2003, Kong et al., 2010). Chitin that has been partly or totally deacetylated (Rabea et al., 2003, Tikhonov et al., 2006, Kong et al., 2010). Although several theories have been presented to explain chitosan's antibacterial action, the precise mechanisms remain unclear (Rabea et al., 2003, Kong et al., 2010, Ke et al., 2021). Intracellular leakage is one of these possibilities, in which positively charged Cs binds to negatively charged bacterial surfaces like lipopolysaccharides (LPS) (Rabea et al., 2003, Kong et al., 2010, Ke et al., 2021). The integrity of the bacterial membrane has compromised as a result of this interaction, leading to the release of intracellular components and cell killing (Rabea et al., 2003, Kong et al., 2010, Ke et al., 2021). From previous study, CNPs exhibited greater antibacterial effect than chitosan against diverse range of fungal (Ing et al., 2012), Gram-positive including S. pneumonia (Chávez de Paz et al., 2011, Alqahtani et al., 2021), and Gram-negative bacteria (Qi et al., 2004, Aleanizy et al., 2018, Alqahtani et al., 2020). Given the superior anti-pnumococcal activity of C3-005 and CNPs, thus, this research purposed to develop CNPs loaded with C3-005 and characterize their antimicrobial properties against S. pneumonia.
2. Materials and methods
2.1. Materials
Chitosan with a low molecular weight (LMW) ranged from 50 to190 kDa and a high degree of deacetylation (DD) ranged from 75 to 85 %, acetic acid, and tripolyphosphate (TPP) were acquired from Sigma Aldrich (St. Louis, MO, USA). Bacterial media including blood agar plates (BA), and Todd Hewitt broth containing 0.5 % yeast extract (THYB) were purchased from Merck (Darmstadt, Germany). C3-005 compound was synthesized locally in the Department of Pharmaceutical Chemistry, College of Pharmacy, King Saud University, Riyadh, Saudi Arabia. All solvents, and chemicals were of analytical grades.
2.2. Preparation of CNPs
Chitosan nanoparticles (CNPs) and chitosan nanoparticles containing C3-005 (C3-CNPs) were generated via ionic gelation technique as performed by Calvo et al. with minor variations (Calvo et al., 1997). In 1 % acetic acid, a Cs stock solution (1 mg/ml) was prepared. The pH of Cs solution was determined and set at 5. TPP solution (1 mg/ml) was added dropwise into 5 mL of Cs solution while being stirred magnetically to produce ECNPs at 4 to 1 ratio of Cs to TPP. For C3-005 loaded CNPs, drug was added to stock Cs solution at 1 to 1 ratio under continuous stirring for 10 min and TPP was added as mentioned before in the preparation of ECNP. Lastly, NPs were harvested by centrifugation for 30 min at 21,700 × g. The NPs reconstituted in sterile distil water (DW) for further experiment.
2.3. Determination of PS, PDI and ZP
The PS, PDI, and zeta potential of the formed NPs were evaluated employing Zetasizer Nano ZS90 (Malvern Instruments ltd, UK) through detection angle of 90° at 25 °C in triplicate.
2.4. Scanning electron microscope (SEM)
The shape of synthesized NPs was examined utilizing scanning electron microscope (JEM-1230EX; Tokyo, Japan).
2.5. Fourier transform infrared spectroscopy (FTIR)
The FTIR spectroscopy was carried out by Shimadzu IRAffinity-1 Fourier Transform Infrared Spectrophotometer utilizing potassium bromide (KBr) disk method after adsorption of a smaller amount of chitosan, C3-005, ECNPs, and C3-CNPs on KBr. The wavelength range measured from 4000 to 400 cm−1, and at a resolution of 2 cm−2.
2.6. Differential scanning calorimetry (DSC) analysis
The differential scanning calorimeter (Perkin-Elmerm, DSC4000, Shelton, CT 0648–4794, USA) was used to perform the DSC measurements. Samples of approximately 1.1 to 4.8 mg (chitosan powder, freeze-dried empty chitosan nanoparticles powder, freeze-dried loaded chitosan nanoparticles powder, and the drug powder) were weighed, transferred into a typical aluminum pan, and covered with a lid. In the range of 30 to 360 °C under nitrogen gas (flow rate 20 mL/min), a heating rate of 10 °C min1 was used. With empty pans, the DSC was previously calibrated.
2.7. HPLC analysis
The Water HPLC system was used to analyze C3-005 at 25 ± 1 °C. The 1515 isocratic (Waters, Milford, MA, USA) pump, 717 autosampler, quaternary LC-10A VP pumps, a programmable UV–visible variable-wavelength detector, a column oven, a SCL 10AVP system controller and an inline vacuum degasser were utilized. The HPLC system utilized the use of the Millennium software (version 32) for data processing and analysis. The column used for this estimation was a Nucleodur (150 mm × 4.6 mm) RP C18 with particle size of 5 μm. The eluent/solvent system was constituted of ethanol: ethyl acetate (50:50 % v/v). The solvent system was flowed with a flow rate of 1.0 mL/min. The detection of target drug was carried out at 350 nm. The samples (20 μL) were inserted into the system via a water auto sampler. The proposed method's linearity, accuracy, precision, robustness, and sensitivity were all verified.
2.8. Encapsulation efficiency (EE%)
For the purpose of calculating the percentage of EE, an indirect method was utilized via determination of drug amount in the supernatant through HPLC. The equation shown below was applied to quantify amount of drug entrapped into CNPs:
2.9. In vitro study
For the in vitro study, the PBS was mixed with a predetermined quantity of C3-CNPs in an eppendorf centrifuge tube. The water bath used for the in vitro release investigation was set at 37 ± 1 °C with constant stirring. The samples were withdrawn of the water bath and centrifuged at 18.000 g for 20 min at specified intervals. One millilitre of the sample was withdrawn and immediately replaced with 1 mL of fresh buffer. The quantity of released C3-005 was estimated by HPLC. At the set time intervals, the cumulative drug releases for each sample were determined.
2.10. Bacterial strains
The Streptococcus pneumonia strain (TIGR4)) (Tettelin et al., 2001) was utilized in the antimicrobial testing after being grown on blood agar plates (BA) overnight or in THYB at 37 °C in 5 % CO2.
2.11. Killing assay
The bacteria were diluted to 1.5 × 106 CFU/mL in THYB at log phase and incubated at 37 °C for 5 h with C3-005, ECNPs, and C3-CNPs at different concentrations ranging from 2.5 to 0.08 mg/ml at 200 rpm. Bacteria cultured in THYB medium alone was included as an untreated control. Bacteria after incubation were serially diluted and plated on BA, where they were cultured overnight at 37 °C and 5 % CO2, and viable colonies were counted. CFUs/mL was used to represent the concentrations. The MIC was described as the lowest concentration of antimicrobial with no observable growth. Experiments were carried out in triplicate.
2.12. S. pneumoniae toxin secretion
Bacterial toxin was secreted as previously described (Ye et al., 2019). First, S. pneumoniae cells were sub-cultured from overnight culture to reach log phase then diluted to OD 0f 0.09 and incubated with C3-005, C3-CNPs, and ECNPs at concentration ranged from 1 to 0.015 mg/ml for 5 hrs at 37 ◦C. Then, bacterial cultures were centrifuged at 3000 g for 3 min. Afterward, the supernatants were utilized for hemolytic assay.
2.13. Hemolytic assay
As previously reported, hemolytic activity was carried out with some modifications (Alqahtani et al., 2021). In THYB, TIGR4 strain were sub-cultured from overnight culture till reaching an OD600 of 0.5. After that, the culture was co-incubated for 5 h with serially diluted C3-005, ECNPs, and C3-CNPs after being diluted to an OD600 of 0.09. The bacterial cells were pelleted after incubation via centrifugation for 5 min (min) at 12000 rpm, and subsequently the supernatant obtained from every sample was then collected and filtered through a 0.22 m syringe filter (Merck Millipore, Burlington, MA, USA). A healthy volunteer's blood was drawn and immediately transferred to an anticoagulant citrate tube, centrifuged for 5 min at 500 g, then the supernatant was discarded, and platelets was removed with PBS. This process was performed twice, and the pellet was then resuspended in 1.5 mL PBS. Then hemolytic activity was carried out by mixing 675 µL of THYB + 300 µL of bacterial supernatant + 25 µL of purified erythrocyte suspension together and incubated at 37 °C for 30 min. PBS was utilized as a negative control, and freshly made 0.1 % Triton X-100 was utilizied as a positive control (100 % hemolysis). Then, the mixture was centrifuges for 1 min at 10,000 g, and collected supernatants were transferred into 96 well plates, where hemoglobin content was quantified by recording absorbance at 590 nm in triplicate using an ELISA plate reader.
2.14. Statistical analysis
Data were displayed as the mean ± standard deviation (SD). Analysis of the data was conducted utilizing GraphPad Prism version 8.0.0 for Mac OS (GraphPad Software, Inc, CA, USA). Student’s t-test and ANOVA tests were utilized as appropriate. A p-value of < 0.05 was counted significant.
3. Results
3.1. Characterization of formulated nanoparticles
In the current study, CNPs were successfully generated in this research using the ion gelation method, which is depends on crosslinking between cationic Cs and anionic TPP and has been reported in previous reports (Calvo et al., 1997, Aleanizy et al., 2018, Alqahtani et al., 2021). The PS, PDI, ZP and EE% of prepared empty CNPs and C3-005 loaded CNPs were presented in Table 1. The diameter of ECNPs and C3-CNPs found to be 255.3 ± 1.8 and 343.5 ± 1.3, respectively.
Table 1.
Physicochemical properties of synthesized CNPs.
| Formula | PZ (nm) ± SD | PDI ± SD | ZP (mV) ± SD | Encapsulation efficiency |
|---|---|---|---|---|
| Empty CNPs | 255.3 ± 1.8 | 0.23 ± 0.01 | 32.1 ± 0.08 | —— |
| C3-CNPs | 343.5 ± 1.3 | 0.20 ± 0.03 | 29.8 ± 0.37 | 70 % |
All formulated nanochitosans generated in this study are monodisperse, with a PDI of<0.23. The zeta potential values of ECNPs were 32.1 mV and 29.8 mV for C3-CNPs. The EE% of C3-005 into CNPs was found to be 70 %. The SEM images revealed spherical morphology of ECNPs and C3-CNPs (Fig. 1).
Fig. 1.
SEM images of ECNPs and C3-CNPs.
3.2. DSC analysis
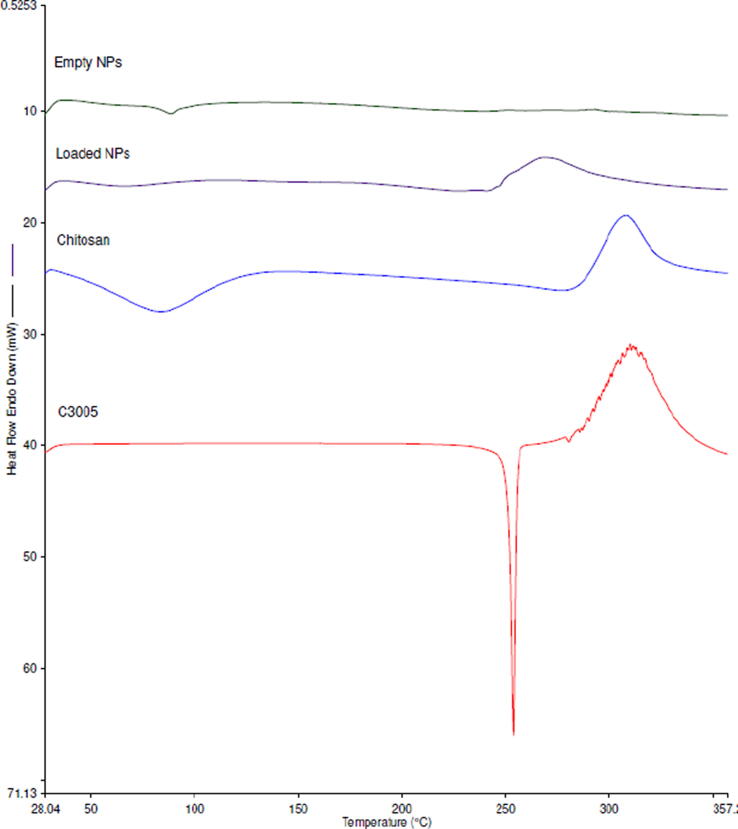
Fig. 2 showed the DSC thermograms of pure C3-005, pure chitosan, ECNPs, and loaded CNPs. Pure C3-005 showed a strong endothermic peak at 253.81 °C and exothermic peak at 310.35 °C. For chitosan, broad endothermic peak at 83.4 °C and exothermic peak at 307.95 °C. The endothermic and exothermic peaks of chitosan of 83.4 °C and 307.95 °C were shifted to 88.82 °C and 293.36 °C, respectively in the empty CNPs. Similarly, the endothermic peak of C3-005 at 253.81 °C was shifted to 67.80 °C, and the exothermic peak at 310.35 °C was shifted to 269.20 °C in C3-005 loaded CNPs.
Fig. 2.
DSC thermograms of C3-005, Cs, ECNPs and C3-CNPs.
3.3. FTIR analysis
The FTIR spectroscopy was used to evaluate encapsulation of compound C3-005 by CNPs. The stretching of the carboxylic and ketonic carbonyl groups is shown at 1686 cm−1 and 1676 cm−1, respectively. In addition, another characteristic absorption was observed around 1527 cm−1 which is due to aromatic nitro group stretching. All of these characteristic stretchings were observed in the FT-IR spectrum of the C3-005 loaded CNPs (Fig. 3) which confirms a successful encapsulation of compound C3-005 into CNPs.
Fig. 3.
FTIR spectra of C3-CNPs, ECNPs and C3-005 compound.
3.4. In vitro release of C3-005 from CNPs
The in vitro release study was performed at pH of 7.4 and showed that C3-005 released from CNPs first at burst release pattern in which 23.5 % release within 6 hrs followed by sustained release (Fig. 4). After 48 hrs, 45.3 % of C3-005 released from CNPs.
Fig. 4.
In vitro release of C3-005 from CNPs.
3.5. Antipneumococcal activity
The results of killing assay are presented in Fig. 5, in which TIGR4 were incubated for five hours with different concentrations of C3-005, C3-CNPs, and ECNPs, then serially diluted, plated and the CFU was determined. The MIC50 of C3-005 was 8 µg/ml when the main stock of the compound dissolved in 100 % DMSO (data not shown). However, when we use the compound dissolved at 5 % DMSO as the same % used in the formula preparation, the MIC50 of C3-005 was increased to 60 µg/ml (Fig. 5). As demonstrated in Fig. 5, the MIC50 of C3-CNPs was decreased to 30 µg/ml. The ECNPs produced anti-pneumococcal activities in dose dependent manner to less extent than both C3-005 and C3-CNPs (Fig. 5).
Fig. 5.
C3-005 loaded CNPs kills S. pneumoniae strain T4 in a concentration dependent manner. After 5 hrs of co-incubation of bacteria with several concentrations of C3-005, C3-CNPs, and ECNPs, T4 were serially diluted and plated. After overnight incubation, the colonies (CFU) were calculated. *p < 0.05 in ANOVA test, in which the viability of C3-CNPs treated bacteria compared to the one treated with C3-005.
In addition, bacterial growth in THYB in the presence of C3-005, C3-CNPs, and ECNPs for 5 h were studied at MIC50, 2x and 3x MIC50 concentration, and results illustrated in Fig. 6. At 30 µg/ml the C3-CNPs reduced the growth of TIGR 4 by approximately 50 %. While C3-005 alone reduced bacterial growth by 40 % at the same concentration (30 µg/ml). These findings support the killing assay results in which C3-CNPs exhibited higher antibacterial activity than C3-005 alone.
Fig. 6.
Growth of S. pneumoniae strain T4 in the presence of prepared NPs. In THYB medium with varying concentrations of C3-CNPs, C3-005, and ECNPs bacteria were grown. In the presence of CNPs, Further reduction in bacterial growth was observed with C3-CNPs treatment. The experiment was carried out in triplicate. Data illustrate a representative experiment.
3.6. The C3-CNPs enhanced the reduction in pneumococcal hemolysis activity
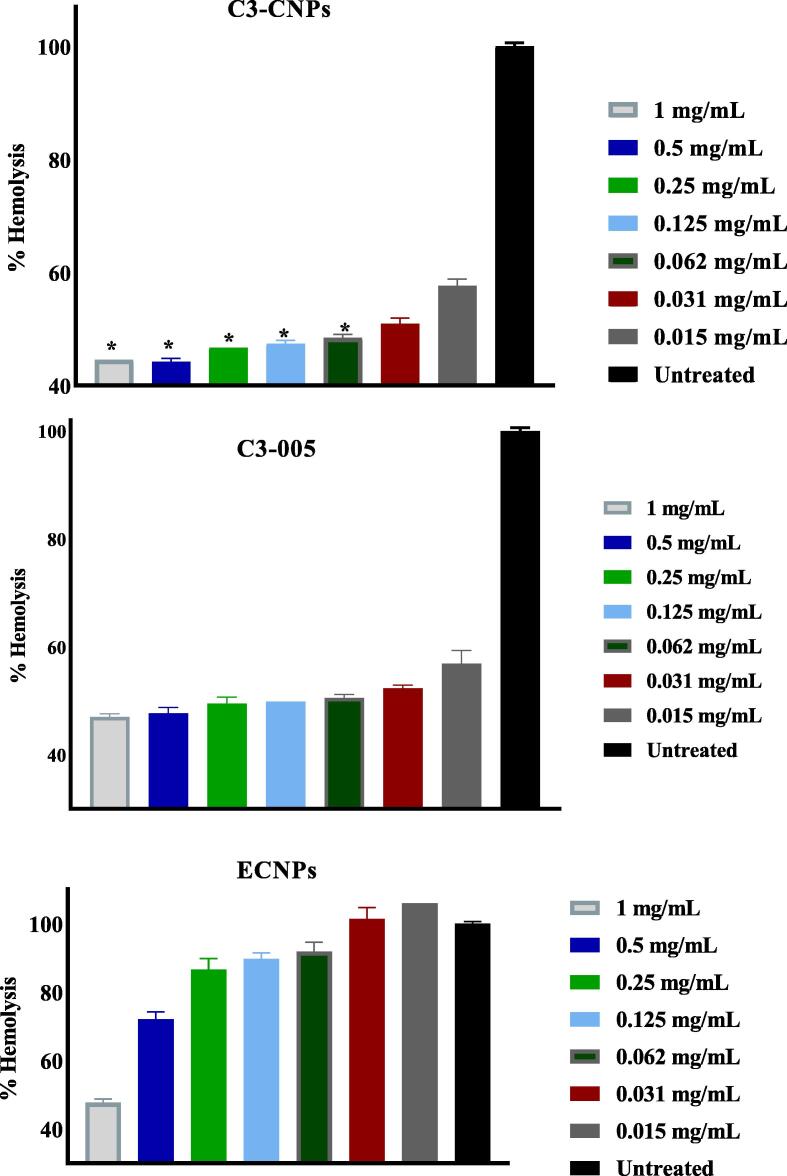
The effect of empty CNPs on the hemolysis activity of pneumococcal culture supernatants was reported previously (Alqahtani et al., 2021). Therefore, the influence of C3-005 and C3-CNPs on the hemolytic activity of S. pneumonia were evaluated utilizing a hemolysis assay. As shown in Fig. 7, to different extent C3-005, ECNPs and C3-CNPs reduced hemolytic activity of supernatant from TIGR4 in dose dependent manner. In comparison with C3-005 alone, significant further decease in bacterial hemolytic activity with C3-CNPs in concentrations ranged from 1 to 0.62 mg/ml was observed (Fig. 7). This shows that CNPs prevent pneumococcal supernatant hemolysis caused by bacteria. Pneumolysin is probably the reason, however this cannot be concluded merely from the study's findings.
Fig. 7.
Effect of CNPs encapsulating C3-005 on the hemolytic activity of S. pneumonia TIGR4. Supernatants of bacteria treated with C3-005, C3-CNPs, and ECNPs for 5 hr were incubated with blood for 30 min and the absorbance recorded after centrifugation. The results are shown as the mean ± SD. Student t test (*p < 0.05) revealed significant reduction in hemolysis compared to bacterial supernatant treated with C3-005 alone.
4. Discussions
Infection with Streptococcus pneumonia associates with invasive life-threatening pneumonia in children and adults, sepsis, and other conditions. Despite the fact that the pneumococcal conjugate vaccine (PCV) is widely used, it is still difficult to treat invasive pneumococcal disease because of the complex serotype and resistance patterns of the bacteria (Kang and Song, 2013, WHO, 2017). Thus, alternative antimicrobial approaches must be developed, and design of novel antimicrobials with new targets, such as compound C3-005 (Hinsberger et al., 2013, Ma et al., 2013, Murakami, 2015, Ye et al., 2019), and use of nanoparticles (NPs) are among these strategies (Czaplewski et al., 2016). In our previous study, NPs formulated from Cs found to exhibited antipneumococcal activity and reduced bacterial hemolysis (Alqahtani et al., 2021). Therefore, our study aims to formulate C3-005 loaded CNPs and evaluate their antibacterial activity and effect on bacterial hemolysis.
In the current study, C3-005 loaded CNPs and ECNPs were successfully prepared using ion gelation method and produced NPs in the nano size range and with low PDI as reported previously (Aleanizy et al., 2018, Alqahtani et al., 2021). The surface charge of the formulated C3-CNPs was cationic, facilitating their uptake by bacteria which covered by negative charge. Loading of C3-005 into CNPs did not significantly change their zeta potential but increased the particle size by approximately 87 nm, which revealed successful encapsulation of C3-005 into CNPs as observed in previous study (Othman et al., 2018). The encapsulation efficiency of C3-005 into CNPs was 70 %, which within the range of other antibiotics and antimicrobials loaded CNPs that previously reported (Jamil et al., 2016, Rawal et al., 2017).
SEM images revealed spherical morphology. The chemical characterization of prepared NPs. The in vitro release results of this study showed burst release of C3-005 from CNPs followed by sustained release pattern in which 45.3 % of C3-005 released from CNPs after 48 hrs. consistent with our study, such type of dual release behavior is described for other drugs by previous research (Tığlı Aydın and Pulat, 2012, Sun et al., 2017).
Cs was used as polymer for synthesis of various nanoparticles loaded with antimicrobials including antibiotics, antimicrobial peptides, natural compound and proteins to enhance their antimicrobial activity (Kong et al., 2010, Chandrasekaran et al., 2020). The antipneumococcal activity of CNPs and C3-005 against S. pneumonia have been previously shown (Ye et al., 2019, Alqahtani et al., 2021). Of note, in the current study, C3-005 loaded CNPs showed enhanced antipneumococcal activity more than C3-005 compound and ECNPs individually. Data obtained from this antimicrobial study are consistent with findings of previous studies (Maya et al., 2012, Zaki and Hafez, 2012, Safhi et al., 2014, Jamil et al., 2016, Sobhani et al., 2017) that showed increase in antibacterial efficacy of different antimicrobial against different pathogens when loaded in CNPs. For example, study conducted by Zaki et al. showed that ceftriaxone sodium loaded CNPs was taken up by macrophages and had a stronger antibacterial action against S. typhimurium than the drug in solution (Zaki and Hafez 2012). In a related study, tetracycline-loaded O-carboxymethyl chitosan nanoparticles were utilized, and it was demonstrated that the tetracycline loaded chitosan NPs improved the effectiveness of the antibiotics in combating S. aureus intracellular infections (Maya et al., 2012).
In addition, the effectiveness of cefazolin containing CNPs against multi-resistant Gram-negative bacteria such E. coli, K. pneumoniae, and P. aeruginosa was assessed by Jamil et al. Their findings revealed that cefazolin loaded CNPs exhibited higher antibacterial activity than drug alone against tested bacteria as proven by agar diffusion and broth dilution methods (Jamil et al., 2016). Similar study demonstrated enhanced antibacterial efficacy of ampicillin combined with chitosan-capped Au NPs against multidrug-resistant clinical isolates of Pseudomonas aeruginosa and Escherichia coli when compared to ampicillin used alone (Chamundeeswari et al., 2010). In another study, penicillin G-loaded CNPs inhibited Streptococcus pyogenes, B. subtilis, and S. aureus higher than penicillin alone (Safhi et al., 2014). Moreover, MIC90 values for CNPs that contained amoxicillin or ciprofloxacin against E. coli and S. aureus were reported to be lower than two drugs used alone (Sobhani et al., 2017).
In addition to the enhanced antibacterial effects, the C3-CNPs formulated in this study also showed enhanced inhibition of hemolysis induced by S. pneumonia. In our previous study (Alqahtani et al., 2021), the CNPs produced concentration-dependent reduction of pneumococcal hemolysis which consistent with the results of ECNPs obtained in this study. This suggests that CNPs may have an antivirulence effect on S. pneumoniae, but the precise mechanism underlying the attenuated hemolysis reported is unknown and require further future evaluation. One of the virulence factors of S. pneumoniae is the release of toxin pneumolysin extracellularly. Earlier study showed that the C3-005 compound significantly reduced post-culture pneumolysin levels when compared to the untreated control (Ye et al., 2019). Given that the obtained findings from the current study regarding enhanced reduction in the hemolysis induced by S. pneumonia in response to C3-CNPs is due to the synergistic effect between Cs and C3-005. Whether Cs exerted this effect through binding or interfering with pneumolysin cannot be concluded from the current study.
5. Conclusions
In the current study, CNPs containing C3-005 were produced and characterized in term of their physicochemical characteristics, antibacterial activity, and effect on pneumococcal induced hemolysis. C3-005 was successfully loaded into CNPs with particle size in nano range, lower PDI, and showed sustained release pattern. The chemical and physical characteristics of prepared nanoparticle were revealed. In comparison with C3-005 and ECNPs, C3-CNP showed enhanced antibacterial activity against S. pneumoniae and further reduction in bacterial induced hemolysis. Further research into the therapeutic potential of C3-005 loaded CNPs in the management of S. pneumoniae infections in animal models can provide evidence implying their clinical use.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgement
This research project was supported by Researchers Supporting Project number (RSP-2022R423), King Saud University, Riyadh, Saudi Arabia.
Footnotes
Peer review under responsibility of King Saud University.
References
- Ahghari M.R., Soltaninejad V., Maleki A. Synthesis of nickel nanoparticles by a green and convenient method as a magnetic mirror with antibacterial activities. Sci. Rep. 2020;10:12627. doi: 10.1038/s41598-020-69679-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aleanizy F.S., Alqahtani F.Y., Shazly G., Alfaraj R., Alsarra I., Alshamsan A., Gareeb Abdulhady H. Measurement and evaluation of the effects of pH gradients on the antimicrobial and antivirulence activities of chitosan nanoparticles in Pseudomonas aeruginosa. Saudi Pharm. J. 2018;26(1):79–83. doi: 10.1016/j.jsps.2017.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alqahtani F., Aleanizy F., El Tahir E., et al. Antibacterial activity of chitosan nanoparticles against pathogenic N. gonorrhoea. Int. J. Nanomed. 2020;15:7877–7887. doi: 10.2147/IJN.S272736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alqahtani F.Y., Aleanizy F.S., El Tahir E., Alowais H., Binkelaib A., Alwathlan B., Al-Bdrawy A., Håkansson A.P., Alsarra I. Capsule independent antimicrobial activity induced by nanochitosan against Streptococcus pneumoniae. Polymers (Basel) 2021;13(17):2924. doi: 10.3390/polym13172924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anderson R., Steel H.C., Cockeran R., Smith A.M., von Gottberg A., de Gouveia L., Brink A., Klugman K.P., Mitchell T.J., Feldman C. Clarithromycin alone and in combination with ceftriaxone inhibits the production of pneumolysin by both macrolide-susceptible and macrolide-resistant strains of Streptococcus pneumoniae. J. Antimicrob. Chemother. 2007;59(2):224–229. doi: 10.1093/jac/dkl479. [DOI] [PubMed] [Google Scholar]
- Baptista P.V., McCusker M.P., Carvalho A., Ferreira D.A., Mohan N.M., Martins M., Fernandes A.R. Nano-Strategies to fight multidrug resistant bacteria-“a battle of the titans”. Front. Microbiol. 2018;9 doi: 10.3389/fmicb.2018.01441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blasi F., Mantero M., Santus P., et al. Understanding the burden of pneumococcal disease in adults. Clin. Microbiol. Infect. 2012;18(Suppl 5):7–14. doi: 10.1111/j.1469-0691.2012.03937.x. [DOI] [PubMed] [Google Scholar]
- Calvo, P., C. Remuñán-López, J. L. Vila-Jato, et al., 1997. Novel hydrophilic chitosan-polyethylene oxide nanoparticles as protein carriers. Journal of Applied Polymer Science. 63, 125-132. https://doi.org/https://doi.org/10.1002/(SICI)1097-4628(19970103)63:1<125::AID-APP13>3.0.CO;2-4.
- Chamundeeswari M., Sobhana S.S.L., Jacob J., Kumar M.G., Devi M.P., Sastry T., Mandal A. Preparation, characterization and evaluation of a biopolymeric gold nanocomposite with antimicrobial activity. Biotechnol. Appl. Biochem. 2010;55(1):29–35. doi: 10.1042/BA20090198. [DOI] [PubMed] [Google Scholar]
- Chandrasekaran M., Kim K.D., Chun S.C. Antibacterial activity of chitosan nanoparticles: a review. Processes. 2020;8:1173. [Google Scholar]
- Chávez de Paz L.E., Resin A., Howard K.A., Sutherland D.S., Wejse P.L. Antimicrobial effect of chitosan nanoparticles on streptococcus mutans biofilms. Appl. Environ. Microbiol. 2011;77(11):3892–3895. doi: 10.1128/AEM.02941-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cherazard R., Epstein M., Doan T.-L., Salim T., Bharti S., Smith M.A. Antimicrobial resistant streptococcus pneumoniae: prevalence, mechanisms, and clinical implications. Am. J. Ther. 2017;24(3):e361–e369. doi: 10.1097/MJT.0000000000000551. [DOI] [PubMed] [Google Scholar]
- Czaplewski L., Bax R., Clokie M., Dawson M., Fairhead H., Fischetti V.A., Foster S., Gilmore B.F., Hancock R.E.W., Harper D., Henderson I.R., Hilpert K., Jones B.V., Kadioglu A., Knowles D., Ólafsdóttir S., Payne D., Projan S., Shaunak S., Silverman J., Thomas C.M., Trust T.J., Warn P., Rex J.H. Alternatives to antibiotics-a pipeline portfolio review. Lancet Infect. Dis. 2016;16(2):239–251. doi: 10.1016/S1473-3099(15)00466-1. [DOI] [PubMed] [Google Scholar]
- Eivazzadeh-Keihan R., Radinekiyan F., Maleki A., Salimi Bani M., Hajizadeh Z., Asgharnasl S. A novel biocompatible core-shell magnetic nanocomposite based on cross-linked chitosan hydrogels for in vitro hyperthermia of cancer therapy. Int. J. Biol. Macromol. 2019;140:407–414. doi: 10.1016/j.ijbiomac.2019.08.031. [DOI] [PubMed] [Google Scholar]
- Gao W., Chen Y., Zhang Y., Zhang Q., Zhang L. Nanoparticle-based local antimicrobial drug delivery. Adv. Drug Deliv. Rev. 2018;127:46–57. doi: 10.1016/j.addr.2017.09.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinsberger S., Hüsecken K., Groh M., Negri M., Haupenthal J., Hartmann R.W. Discovery of novel bacterial RNA polymerase inhibitors: pharmacophore-based virtual screening and hit optimization. J. Med. Chem. 2013;56(21):8332–8338. doi: 10.1021/jm400485e. [DOI] [PubMed] [Google Scholar]
- Ing, L. Y., N. M. Zin, A. Sarwar, et al., 2012. Antifungal activity of chitosan nanoparticles and correlation with their physical properties. Int J Biomater. 2012, 632698. https://doi.org/10.1155/2012/632698. [DOI] [PMC free article] [PubMed]
- Jamil B., Habib H., Abbasi S., Nasir H., Rahman A., Rehman A., Bokhari H., Imran M. Cefazolin loaded chitosan nanoparticles to cure multi drug resistant Gram-negative pathogens. Carbohydr. Polym. 2016;136:682–691. doi: 10.1016/j.carbpol.2015.09.078. [DOI] [PubMed] [Google Scholar]
- Jedrzejas M.J. Pneumococcal virulence factors: structure and function. Microbiol. Mol. Biol. Rev. 2001;65(2):187–207. doi: 10.1128/MMBR.65.2.187-207.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang C.I., Song J.H. Antimicrobial resistance in Asia: current epidemiology and clinical implications. Infect. Chemother. 2013;45:22–31. doi: 10.3947/ic.2013.45.1.22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ke C.-L., Deng F.-S., Chuang C.-Y., Lin C.-H. Antimicrobial actions and applications of chitosan. Polymers (Basel) 2021;13(6):904. doi: 10.3390/polym13060904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kong M., Chen X.G., Xing K.e., Park H.J. Antimicrobial properties of chitosan and mode of action: a state of the art review. Int. J. Food Microbiol. 2010;144(1):51–63. doi: 10.1016/j.ijfoodmicro.2010.09.012. [DOI] [PubMed] [Google Scholar]
- Lee N.Y., Ko W.C., Hsueh P.R. Nanoparticles in the treatment of infections caused by multidrug-resistant organisms. Front. Pharmacol. 2019;10:1153. doi: 10.3389/fphar.2019.01153. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma C., Yang X., Kandemir H., Mielczarek M., Johnston E.B., Griffith R., Kumar N., Lewis P.J. Inhibitors of bacterial transcription initiation complex formation. ACS Chem. Biol. 2013;8(9):1972–1980. doi: 10.1021/cb400231p. [DOI] [PubMed] [Google Scholar]
- Maleki A. Green oxidation protocol: Selective conversions of alcohols and alkenes to aldehydes, ketones and epoxides by using a new multiwall carbon nanotube-based hybrid nanocatalyst via ultrasound irradiation. Ultrason. Sonochem. 2018;40:460–464. doi: 10.1016/j.ultsonch.2017.07.020. [DOI] [PubMed] [Google Scholar]
- Maleki A., Ghassemi M., Firouzi-Haji R. Green multicomponent synthesis of four different classes of six-membered N-containing and O-containing heterocycles catalyzed by an efficient chitosan-based magnetic bionanocomposite. Pure Appl. Chem. 2018;90:387–394. doi: 10.1515/pac-2017-0702. [DOI] [Google Scholar]
- Maleki A., Hajizadeh Z., Salehi P. Mesoporous halloysite nanotubes modified by CuFe2O4 spinel ferrite nanoparticles and study of its application as a novel and efficient heterogeneous catalyst in the synthesis of pyrazolopyridine derivatives. Sci. Rep. 2019;9:5552. doi: 10.1038/s41598-019-42126-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maleki A., Kamalzare M. An efficient synthesis of benzodiazepine derivatives via a one-pot, three-component reaction accelerated by a chitosan-supported superparamagnetic iron oxide nanocomposite. Tetrahedron Lett. 2014;55:6931–6934. doi: 10.1016/j.tetlet.2014.10.120. [DOI] [Google Scholar]
- Maya S., Indulekha S., Sukhithasri V., Smitha K.T., Nair S.V., Jayakumar R., Biswas R. Efficacy of tetracycline encapsulated O-carboxymethyl chitosan nanoparticles against intracellular infections of Staphylococcus aureus. Int. J. Biol. Macromol. 2012;51(4):392–399. doi: 10.1016/j.ijbiomac.2012.06.009. [DOI] [PubMed] [Google Scholar]
- Murakami K.S. Structural biology of bacterial RNA polymerase. Biomolecules. 2015;5:848–864. doi: 10.3390/biom5020848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Othman N., Masarudin M., Kuen C., Dasuan N., Abdullah L., Md. Jamil S. Synthesis and optimization of chitosan nanoparticles loaded with L-ascorbic acid and thymoquinone. Nanomaterials (Basel) 2018;8(11):920. doi: 10.3390/nano8110920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qi L., Xu Z., Jiang X., Hu C., Zou X. Preparation and antibacterial activity of chitosan nanoparticles. Carbohydr. Res. 2004;339(16):2693–2700. doi: 10.1016/j.carres.2004.09.007. [DOI] [PubMed] [Google Scholar]
- Rabea E.I., Badawy M.E., Stevens C.V., et al. Chitosan as antimicrobial agent: applications and mode of action. Biomacromolecules. 2003;4:1457–1465. doi: 10.1021/bm034130m. [DOI] [PubMed] [Google Scholar]
- Rawal T., Parmar R., Tyagi R.K., Butani S. Rifampicin loaded chitosan nanoparticle dry powder presents an improved therapeutic approach for alveolar tuberculosis. Colloids Surf. B Biointerfaces. 2017;154:321–330. doi: 10.1016/j.colsurfb.2017.03.044. [DOI] [PubMed] [Google Scholar]
- Safhi M.M., Sivakumar S.M., Jabeen A., et al. Chitosan nanoparticles as a sustained delivery of penicillin G prepared by ionic gelation technique. J. Pharm. Res. 2014;88:1352–1354. [Google Scholar]
- Shah S.M.K., Rasheed T.U., Ali H.M., Umar A. Smart integrated decentralization strategies of solar power system in buildings. Int. J. Photoenergy. 2022;2022:1–14. doi: 10.1155/2022/9311686. [DOI] [Google Scholar]
- Sobhani Z., Mohammadi Samani S., Montaseri H., et al. Nanoparticles of chitosan loaded ciprofloxacin: fabrication and antimicrobial activity. Adv. Pharma. Bull. 2017;7:427–432. doi: 10.15171/apb.2017.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soltaninejad V., Ahghari M.R., Taheri-Ledari R., Maleki A. Bifunctional PVA/ZnO/AgI/chlorophyll nanocomposite film: enhanced photocatalytic activity for degradation of pollutants and antimicrobial property under visible-light irradiation. Langmuir : ACS J. Surf. Colloids. 2021;37(15):4700–4713. doi: 10.1021/acs.langmuir.1c00501. [DOI] [PubMed] [Google Scholar]
- Sriharan G., Harikrishnan S., Ali H.M. Enhanced heat transfer characteristics of the mini hexagonal tube heat sink using hybrid nanofluids. Nanotechnology. 2022;33(47):475403. doi: 10.1088/1361-6528/ac8995. [DOI] [PubMed] [Google Scholar]
- Sun L.i., Chen Y., Zhou Y., Guo D., Fan Y., Guo F., Zheng Y., Chen W. Preparation of 5-fluorouracil-loaded chitosan nanoparticles and study of the sustained release in vitro and in vivo. Asian J. Pharm. Sci. 2017;12(5):418–423. doi: 10.1016/j.ajps.2017.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tacconelli E., Pezzani M.D. Public health burden of antimicrobial resistance in Europe. Lancet Infect. Dis. 2019;19:4–6. doi: 10.1016/S1473-3099(18)30648-0. [DOI] [PubMed] [Google Scholar]
- Tettelin H., Nelson K.E., Paulsen I.T., Eisen J.A., Read T.D., Peterson S., Heidelberg J., DeBoy R.T., Haft D.H., Dodson R.J., Durkin A.S., Gwinn M., Kolonay J.F., Nelson W.C., Peterson J.D., Umayam L.A., White O., Salzberg S.L., Lewis M.R., Radune D., Holtzapple E., Khouri H., Wolf A.M., Utterback T.R., Hansen C.L., McDonald L.A., Feldblyum T.V., Angiuoli S., Dickinson T., Hickey E.K., Holt I.E., Loftus B.J., Yang F., Smith H.O., Venter J.C., Dougherty B.A., Morrison D.A., Hollingshead S.K., Fraser C.M. Complete genome sequence of a virulent isolate of Streptococcus pneumoniae. Science. 2001;293(5529):498–506. doi: 10.1126/science.1061217. [DOI] [PubMed] [Google Scholar]
- Tığlı Aydın, R. S. and M. Pulat, 2012. 5-Fluorouracil Encapsulated Chitosan Nanoparticles for pH-Stimulated Drug Delivery: Evaluation of Controlled Release Kinetics. Journal of Nanomaterials. 2012, 313961. https://doi.org/10.1155/2012/313961.
- Tikhonov V.E., Stepnova E.A., Babak V.G., Yamskov I.A., Palma-Guerrero J., Jansson H.-B., Lopez-Llorca L.V., Salinas J., Gerasimenko D.V., Avdienko I.D., Varlamov V.P. Bactericidal and antifungal activities of a low molecular weight chitosan and its N-/2(3)-(dodec-2-enyl)succinoyl/-derivatives. Carbohydr. Polym. 2006;64(1):66–72. doi: 10.1016/j.carbpol.2005.10.021. [DOI] [Google Scholar]
- WHO. 2017. Retrieved 20th August, 2022, from https://www.who.int/medicines/publications/global-priority-list-antibioticresistant-bacteria/en/.
- Ye, J., A. J. Chu, L. Lin, et al., 2019. First-In-Class Inhibitors Targeting the Interaction between Bacterial RNA Polymerase and Sigma Initiation Factor Affect the Viability and Toxin Release of Streptococcus pneumoniae. Molecules. 24, https://doi.org/10.3390/molecules24162902 [DOI] [PMC free article] [PubMed]
- Zaki N.M., Hafez M.M. Enhanced antibacterial effect of ceftriaxone sodium-loaded chitosan nanoparticles against intracellular Salmonella typhimurium. AAPS PharmSciTech. 2012;13:411–421. doi: 10.1208/s12249-012-9758-7. [DOI] [PMC free article] [PubMed] [Google Scholar]