Abstract
Using chicken feathers as bait, Acremoniumglobosisporumsp. nov. and Acremoniumcurvumsp. nov. were collected from the soil of Yuncheng East Garden Wildlife Zoo and Zhengzhou Zoo in China. They were identified by combining the morphological characteristics and the two-locus DNA sequence (LSU and ITS) analyses. In the phylogenetic tree, both new species clustered into separate subclades, respectively. They were different from their allied species in their morphology. The description, illustrations, and phylogenetic tree of the two new species were provided.
Keywords: Acremonium , filamentous fungi, phylogeny, taxonomy
Introduction
The genus Acremonium Link, established in 1929, with A.alternatum Link as the type species, is one of the largest and most complex genera of asexually typified. The morphological characteristics consist of hyphae septate, mostly tapered and lateral phialides, produced singly or in small groups, and unicellular conidia produced in mucoid heads or unconnected chains (Summerbell et al. 2011). Nowadays, Acremonium has 217 records in the Index Fungorum (http://www.indexfungorum.org/Names/Names.asp, retrieval on 30 Jun. 2022). The traditionally circumscribed Acremonium is polyphyletic, which explains why many Acremonium species were transferred to other genera and families (Yang et al. 2019). Thus, there are still many unidentified, suspect or misidentified taxa that require taxonomic investigation.
Due to the poor differentiation of asexual forms of the genus Acremonium, it is difficult to identify species only by morphological differences. To address this issue, there are many unidentified and suspicious species that require further phylogenetic analysis. To date, many isolates of Acremonium spp. lack the gene loci such as SSU, TEF 1-α and RPB2 (Table 1), therefore, phylogenetic analyses of this genus are generally performed based on the single locus sequences, especially LSU (Hyde et al. 2020).
Table 1.
Strains included in the present study.
| Species | Strains | LSU | ITS | SSU | TEF 1-α | RPB2 |
|---|---|---|---|---|---|---|
| Acremoniumalcalophilum | CBS 114.92T | JX158443 | DQ825967 | JX158486 | JX158399 | JX158465 |
| Acremoniumalternatum | CBS 407.66T | HQ231988 | HE798150 | |||
| Acremoniumalternatum | CBS 831.97 | HQ231989 | ||||
| Acremoniumarthrinii | MFLU 18-1225T | MN036334 | MN036335 | MN038169 | ||
| Acremoniumbehniae | CBS 146824T | MW175400 | MW175360 | |||
| Acremoniumbiseptum | CBS 750.69T | HQ231998 | ||||
| Acremoniumblochii | CBS 993.69 | HQ232002 | HE608636 | |||
| Acremoniumborodinense | CBS 101148T | HQ232003 | HE608635 | |||
| Acremoniumbrachypenium | CBS 866.73T | HQ232004 | AB540570 | |||
| Acremoniumcamptosporum | CBS 756.69T | HQ232008 | HQ232186 | |||
| Acremoniumcavaraeanum | CBS 101149T | HF680202 | HF680220 | |||
| Acremoniumcavaraeanum | CBS 111656 | HF680203 | HF680221 | |||
| Acremoniumcavaraeanum | CBS 758.69 | HQ232012 | HF680222 | |||
| Acremoniumcerealis | CBS 207.65 | HQ232013 | ||||
| Acremoniumcerealis | CBS 215.69 | HQ232014 | ||||
| Acremoniumchiangraiense | MFLUCC 14-0397T | MN648329 | MN648324 | |||
| Acremoniumchrysogenum | CBS 144.62T | HQ232017 | HQ232187 | |||
| Acremoniumchrysogenum | CBS 401.65 | MH870276 | MH858636 | |||
| Acremoniumcitrinum | CBS 384.96T | HF680217 | HF680236 | |||
| Acremoniumdimorphosporum | CBS 139050T | LN810506 | LN810515 | |||
| Acremoniumexiguum | CBS 587.73T | HQ232035 | ||||
| Acremoniumexuviarum | UAMH 9995T | HQ232036 | AY882946 | |||
| Acremoniumfelinum | CBS 147.81T | AB540488 | AB540562 | |||
| Acremoniumflavum | CBS 596.70T | HQ232037 | HQ232191 | |||
| Acremoniumflavum | CBS 316.72 | MH872204 | MH860487 | |||
| Acremoniumfuci | CBS 112868T | AY632653 | ||||
| Acremoniumfuci | CBS 113889 | AY632652 | ||||
| Acremoniumfusidioides | CBS 109069 | HF680204 | HF680223 | |||
| Acremoniumfusidioides | CBS 991.69 | HF680211 | HF680230 | |||
| Acremoniumfusidioides | CBS 840.68T | HQ232039 | FN706542 | |||
| Acremoniumhansfordii | CBS 390.73 | HQ232043 | AB540578 | |||
| Acremoniumhennebertii | CBS 768.69T | HQ232044 | HF680238 | |||
| Acremoniuminflatum | CBS 212.69T | HQ232050 | ||||
| Acremoniummali | ACCC 39305T | MF993114 | MF987658 | |||
| Acremoniummoniliforme | CBS 139051T | LN810507 | LN810516 | |||
| Acremoniummoniliforme | FMR 10363 | LN810508 | LN810517 | |||
| Acremoniumparvum | CBS 381.70A | HQ231986 | HF680219 | |||
| Acremoniumpersicinum | CBS 310.59T | HQ232077 | ||||
| Acremoniumpersicinum | CBS 101694 | HQ232085 | ||||
| Acremoniumpinkertoniae | CBS 157.70T | HQ232089 | HQ232202 | |||
| Acremoniumpolychroma | CBS 181.27T | HQ232091 | AB540567 | |||
| Acremoniumpotronii | CBS 189.70 | HQ232094 | ||||
| Acremoniumpseudozeylanicum | CBS 560.73T | HQ232101 | ||||
| Acremoniumpteridii | CBS 782.69T | HQ232102 | ||||
| Acremoniumpteridii | CBS 784.69 | HQ232103 | ||||
| Acremoniumsclerotigenum | CBS 124.42T | HQ232126 | FN706552 | HQ232209 | ||
| Acremoniumsclerotigenum | A101 | KC987215 | KC987139 | KC987177 | KC998961 | |
| Acremoniumsclerotigenum | A130 | KC987242 | KC987166 | KC987204 | KC998988 | |
| Acremonium sp. | E102 | KC987248 | KC987172 | KC987210 | KC998994 | KC999030 |
| Acremoniumspinosum | CBS 136.33T | HQ232137 | HE608637 | HQ232210 | ||
| Acremoniumstroudii | CBS 138820T | KM225291 | ||||
| Acremoniumtumulicola | CBS 127532T | AB540478 | AB540552 | |||
| Acremoniumvariecolor | CBS 130360T | HE608651 | HE608647 | |||
| Acremoniumvariecolor | CBS 130361 | HE608652 | HE608648 | |||
| Acremoniumverruculosum | CBS 989.69T | HQ232150 | ||||
| Acrophialophorahechuanensis | GZUIFR-H08-1T | MK926789 | DQ185070 | EU053286 | ||
| Brunneomycesbrunnescens | CBS 559.73T | HQ231966 | LN810520 | HQ232184 | LN810534 | |
| Brunneomyceshominis | UTHSC 06-415T | LN810509 | KP131517 | LN810535 | ||
| Bryocentriabrongniartii | M139 | EU940105 | EU940052 | |||
| Bryocentriabrongniartii | M190 | EU940125 | EU940052 | |||
| Bryocentriametzgeriae | M140 | EU940106 | ||||
| Bulbitheciumhyalosporum | CBS 318.91T | AF096187 | HE608634 | |||
| Cephalosporiumpurpurascens | CBS 149.62T | HQ232071 | ||||
| Cosmosporalavitskiae | CBS 530.68T | HQ231997 | ||||
| Emericellopsisalkalina | CBS 127350T | KC987247 | KC987171 | KC987209 | KC998993 | KC999029 |
| Emericellopsisterricola | CBS 120.40T | U57082 | U57676 | U44112 | ||
| Gliomastixroseogrisea | CBS 134.56T | HQ232121 | ||||
| Hapsidosporairregularis | ATCC 22087T | AF096192 | AF096177 | |||
| Kiflimoniumcurvulum | CBS 430.66T | HQ232026 | HE608638 | HQ232188 | ||
| Lanatonectriaflavolanata | CBS 230.31 | HQ232157 | ||||
| Leucosphaerinaarxii | CBS 737.84T | HE608662 | HE608640 | |||
| Nigrosabulumglobosum | ATCC 22102T | AF096195 | ||||
| Paracremoniumcontagium | CBS 110348T | HQ232118 | KM231831 | KM231966 | ||
| Parasarocladiumbreve | CBS 150.62T | HQ232005 | ||||
| Parasarocladiumradiatum | CBS 142.62T | HQ232104 | HQ232205 | |||
| Pestalotiopsishawaiiensis | CBS 114491T | KM116239 | KM199339 | KM199514 | ||
| Pestalotiopsisspathulata | CBS 356.86T | KM116236 | KM199338 | KM199513 | ||
| Phialemoniumatrogriseum | CBS 604.67T | HQ231981 | HE610367 | FJ176825 | ||
| Pseudoacremoniumsacchari | CBS 137990T | KJ869201 | KJ869144 | |||
| Sarcopodiumvanillae | CBS 100582 | HQ232174 | KM231780 | KM231911 | ||
| Sarocladiumbacillisporum | CBS 425.67T | HQ231992 | HE608639 | HQ232179 | ||
| Sarocladiumbactrocephalum | CBS 749.69T | HQ231994 | HG965006 | HQ232180 | ||
| Sarocladiumstrictum | CBS 346.70T | HQ232141 | AY214439 | HQ232211 | ||
| Sarocladiumterricola | CBS 243.59T | HQ232046 | HQ232196 | |||
| Seliniapulchra | AR 2812 | GQ505992 | HM484859 | HM484841 | ||
| Trichotheciumcrotocinigenum | CBS 129.64T | HQ232018 | AJ621773 | |||
| Trichotheciumindicum | CBS 123.78T | AF096194 | AF096179 | |||
| Trichotheciumroseum | DAOM 208997 | U69891 | U69892 | |||
| Trichotheciumsympodiale | ATCC 36477 | U69889 | U69890 | |||
| Acremoniumcurvum | CGMCC 3.20954 = GZUIFR 22.035T | ON041050 | ON041034 | ON876754 | ON494579 | ON494583 |
| Acremoniumglobosisporum | CGMCC 3.20955 = GZUIFR 22.036T | ON041051 | ON041035 | ON876755 | ON494580 | ON494584 |
| Acremoniumglobosisporum | GZUIFR 22.037 | ON041052 | ON041036 | ON876756 | ON494581 | ON494585 |
| Acremoniumglobosisporum | GZUIFR 22.038 | ON041053 | ON041037 | ON876757 | ON494582 | ON494586 |
Notes: “T” stands for Ex-type strains.
In the present study, two new species of Acremonium were identified in a survey of keratinolytic fungi from China, which were enriched by the baiting technique. We provided a description, illustrations, and phylogenetic tree for the two new species.
Materials and methods
Fungal isolation and morphology
Soil samples were collected from Yuncheng East Garden Wildlife Zoo (35°6'26"N, 111°4'24"E) (three isolates), Yuncheng City, Shanxi Province and Zhengzhou Zoo (34°47'20"N, 113°40'41"E) (one isolate), Zhengzhou City, Henan Province, China by Yu-Lian Ren on July 2021. We collected 3–10 cm below the soil surface, placed the samples in sterile Ziploc plastic bags (Kaixin Biotechnology, Guizhou, China), and transported them to the laboratory (Zhang et al. 2019a, b). Then, they were treated and isolated according to the baiting method (using chicken feathers as bait: a method specifically designed for isolating keratinophilic microbes) of Zhang et al. (2020a, b; 2021). We washed the chicken feathers, sterilized them in an autoclave for 30 minutes at 121 °C, and dried them in an oven at 50 °C. The sterile and dried chicken feathers were mixed with soil samples and then wet with sterile distilled water and cultured at darkroom temperature for 1 month (Li et al. 2022).
Then, the 2 g samples were weighed in a conical flask with glass beads containing 20 mL sterile water and mixed evenly by eddy shock for 10 min. Next, 1 mL samples were mixed evenly in 9 mL sterile water in a sterile environment and diluted to 10-3. Then, 1 mL 10-3 samples were put into a sterile petri dish, and SDA medium containing 50 mg/L penicillin and 50 mg/L streptomycin was added and mixed. The target strains were isolated. The purified strains were transferred to PDA, OA, and MEA plates for dark culture at 25 °C for 7 days. Microscopic features were examined by making direct wet mounts with 25% lactic acid on PDA, with a light microscope.
The cultures were placed to slowly dry at 50 °C to produce the dried holotype. The dried holotype was deposited in the Mycological Herbarium of the Institute of Microbiology, Chinese Academy of Sciences, Beijing, China (HMAS), while ex-type living culture was stored in PDA test tubes which were deposited in the China General Microbiological Culture Collection Center (CGMCC), and the Institute of Fungus Resources, Guizhou University, Guiyang City, Guizhou, China (GZUIFR).
DNA extraction, PCR amplification, and sequencing
We used a 5% chelex-100 solution for total genomic DNA extraction. ITS1/ITS4 (White et al. 1990), LROR/LR7 (Vilgalys and Hester 1990), EF1-983F/ EF1-2218R (Rehner and Buckley 2005), fRPB2-5f/ fRPB2-7cR (Liu et al. 1999), and NS1 and NS4 (White et al. 1990) primers were used for amplification of the internal transcribed spacers (ITS), the 28S nrRNA locus (LSU), translation elongation factor 1-alpha gene region (TEF 1-α), RNA polymerase II second largest subunit gene (RPB2), and small subunit rDNA (SSU), respectively. Purification and sequencing were performed by Quintarabio (Wuhan, China). The new sequences were submitted to GenBank (Table 1).
Phylogenetic analyses
The ITS and LSU sequences of Acremonium were downloaded from GenBank (Table 1). Two strains of Pestalotiopsisspathulata (CBS 356.86) and P.hawaiiensis (CBS 114491) were chosen as the outgroup taxa. The TBtools were used for name simplification and renaming (Chen et al. 2020). Sequences were aligned by MAFFT v7.037 (Katoh and Standley 2013). Multi-locus was concatenated by PhyloSuite v1.16 (Zhang et al. 2020a).
Bayesian inference (BI) and maximum likelihood (ML) methods were used in the analysis. For BI analysis was conducted with MrBayes v3.2 (Ronquist et al. 2012) and Markov chain Monte Carlo (MCMC) simulations; ML analysis was performed using IQ-TREE v1.6.11 (Nguyen et al. 2015), as outlined in Li et al (2022). All analyses were performed in PhyloSuite V1.16 (Zhang et al. 2020b).
Results
Phylogeny
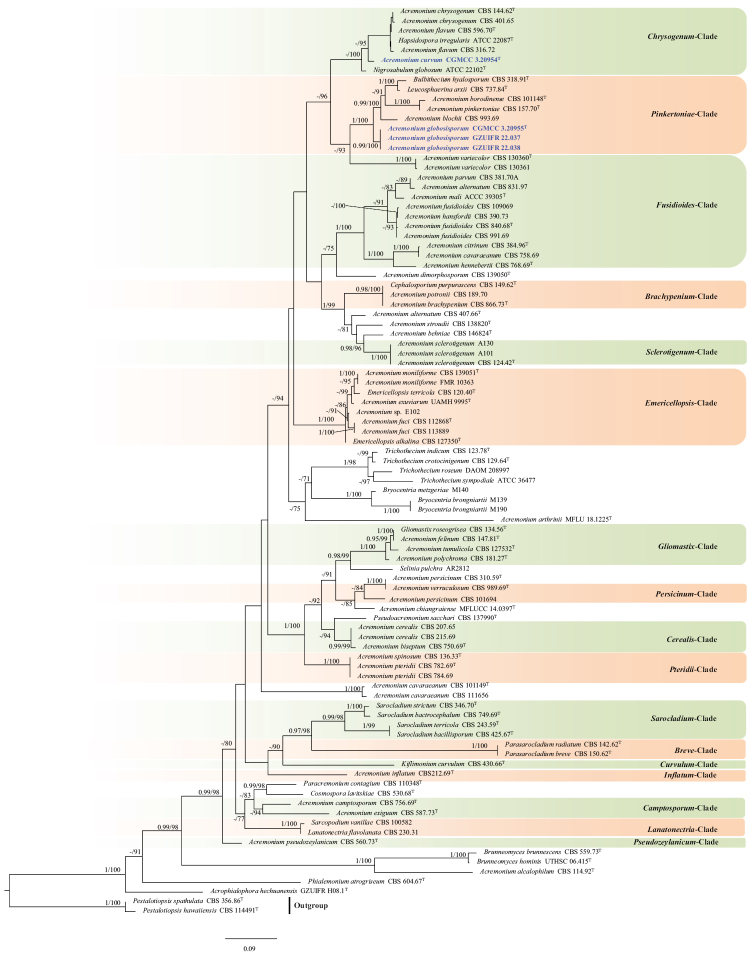
Based on a BLAST search (https://blast.ncbi.nlm.nih.gov/Blast.cgi) using the LSU sequences, our isolates were identified as belonging to the genus Acremonium. To further determine the phylogenetic position of these strains, we performed a multi-locus phylogenetic analysis. The dataset was composed of LSU (1–430 bp) and ITS (431–1005 bp) gene, comprising a total of 1005 characters (including gaps). The best-fit partition model for ML analysis and BI analysis is shown in Table 2. The results showed that the CGMCC 3.20955, GZUIFR 22.037, and GZUIFR 22.038 are still grouped in the Pinkertoniae-clade (Fig. 1). The CGMCC 3.20954 is still grouped in the Chrysogenum-clade (Fig. 1).
Table 2.
The best-fit substitution models are used in multi-locus phylogenetic construction.
| LSU | ITS | |
|---|---|---|
| ML analysis | TN+F+R5 | GTR+F+R4 |
| BI analysis | GTR+F+I+G4 | GTR+F+I+G4 |
Figure 1.
Phylogenetic tree of the genus Acremonium constructed from LSU and ITS. Bayesian posterior probability (≥ 0.95) and ML bootstrap values (≥ 70%) are indicated along branches (BPP/ML).
Taxonomy
. Acremonium globosisporum
Xin Li, Y.F. Han & Z.Q. Liang sp. nov.
35BF27DA-FC43-57D8-989A-49581DCE8C51
843765
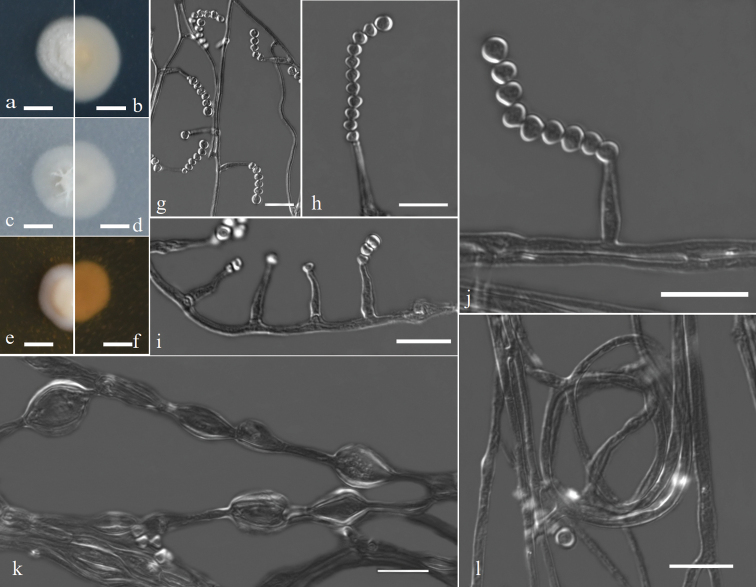
Figure 2.
Morphology of Acremoniumglobosisporum sp. nov. a–f colony on PDA, OA and MEA after 7 d at 25 °C (upper surface and lower surface) g–j conidia are borne on the phialides k–l winding hyphae and inflate hyphae. Scale bars: 4 mm (a–f); 10 μm (g–l).
Type.
Yuncheng East Garden Wildlife Zoo, Yuncheng City, Shanxi Province, China N35°6'26", E111°4'24", isolated from green belt soil, July 2021, Yu-Lian Ren (dried holotype culture HMAS 351939, ex-holotype culture CGMCC 3.20955 = GZUIFR 22.036). ITS sequences, GenBank ON041035; LSU sequences, GenBank ON041051; SSU sequences, GenBank ON876755; TEF 1-α sequences, GenBank ON494580; RPB2 sequences, GenBank ON494584.
Description.
Colonies on PDA and OA at 25 °C attaining 11–13 mm and 9–11 mm diam respectively after 7 d, white, flat or raised, velvety to slightly cottony. On MEA at 25 °C, reaching 8–10 mm after 7 d, white to yellowish white, raised, slimy. Hyphae hyaline, septate, sometimes winding and inflate, 1.5–11.0 µm wide. Sporulation abundant. Phialides are mostly borne singly, hyaline, erect to slightly curved, sometimes forming a collarette, 9.0–22.0 µm long, tapering from 1.5–3.5 µm near the base to 0.5–1.5 µm. Conidia cohering in long chains, with minutely truncate ends, up to 27.5 µm long, globose or subglobose, 2.5–4.5 × 2.5–4.5 µm (x– ± SD = 3.4 ± 0.77 × 3.6 ± 0.52, n = 50) diam. Chlamydospores and teleomorph stage were not observed.
Etymology.
globosisporum. A reference to the global conidia.
Additional specimens examined.
Yuncheng East Garden Wildlife Zoo, Yuncheng City, Shanxi Province, China N35°6'26", E111°4'24", isolated from green belt soil, July 2021, Yu-Lian Ren, GZUIFR 22.037, ITS, LSU, SSU, TEF 1-α, RPB2 sequences GenBank ON041036, ON041052, ON876756, ON494581, ON494585; GZUIFR 22.038, ITS, LSU, SSU, TEF 1-α, RPB2 sequences GenBank ON041037, ON041053, ON876757, ON494582, ON494586.
Known distribution.
Yuncheng City, Shanxi Province, China.
Notes.
The phylogeny results showed that the CGMCC 3.20955, GZUIFR 22.037 and GZUIFR 22.038 still nested in the Pinkertoniae-clade. The morphological characteristics of Acremoniumglobosisporum were similar to other species of the Pinkertoniae-clade in that phialides were erect on the hyphae; sporulation was abundant, and conidia were subglobose (Ito et al. 2000). However, Acremoniumglobosisporum hyphae were sometimes winding and inflated, with conidia cohering in long chains, unlike other species.
. Acremonium curvum
Xin Li, Y.F. Han & Z.Q. Liang sp. nov.
7DC71370-B32D-59C2-B927-D407C2A66239
843766
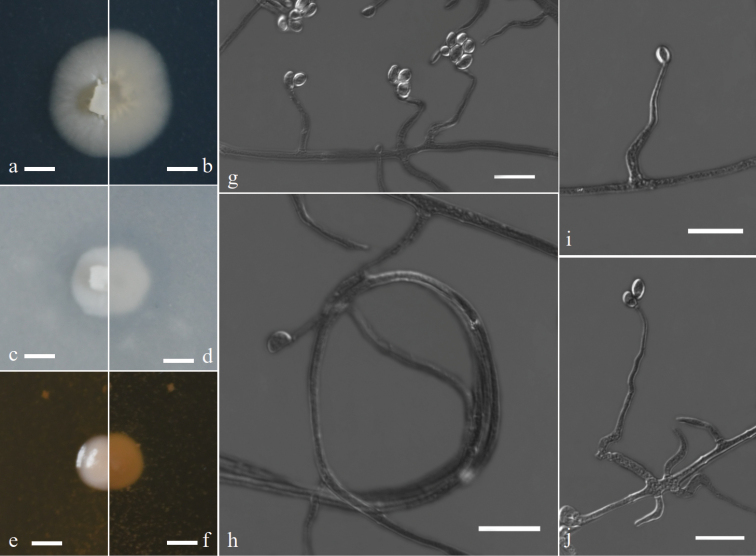
Figure 3.
Morphology of Acremoniumcurvum sp. nov. a–f colony on PDA, OA and MEA after 7 d at 25 °C (upper surface and lower surface) g, I, j conidia are borne on the phialides h Winding hyphae. Scale bars: 4 mm (a–f); 10 μm (g–j).
Type.
Zhengzhou Zoo, Zhengzhou City, Henan Province, China N34°47'20", E113°40'41", isolated from green belt soil, July 2021, Yu-Lian Ren (dried holotype culture HMAS 351938, ex-holotype culture CGMCC 3.20954 = GZUIFR 22.035). ITS sequences, GenBank ON041034, LSU sequences, GenBank ON041050; SSU sequences, GenBank ON876754; TEF 1-α sequences, GenBank ON494579; RPB2 sequences, GenBank ON494583.
Description.
Colonies on PDA and OA at 25 °C attaining 11–14 mm and 7–9 mm diam respectively after 7 d, white, flat, radially folded or rugose. On MEA at 25 °C, reaching 6–8 mm after 7 d, white to yellowish-white, slimy. Hyphae hyaline, septate, sometimes winding, 1.5–2.5 µm wide. Sporulation abundant. Phialides are mostly borne singly, curved, slightly inflated at the base, tapered at the tip, up to 38.0 µm long. tapering from 1.5–3.5 µm near the base to 0.5–1.5 µm. Conidia cohering together on the top of phialides, one-celled, solitary, or several fascicled, ovoid or subglobose, 3.0–7.0 × 2.5–3.5 µm (x– ± SD = 4.1 ± 1.18 ×3.2 ± 0.77, n = 50) diam. Chlamydospores and teleomorph stage were not observed.
Etymology.
curvum. Referring to the curved Phialides.
Known distribution.
Henan Province, China.
Notes.
Based on the multi-locus analysis we found that Acremoniumcurvum had close phylogenetic affinities to other taxa of the Chrysogenum-clade. Morphologically, A.curvum was similar to other taxa of the Chrysogenum-clade in having simple or rarely branched conidiophores, slightly inflated at the base and tapered at tip phialides, and ovoid to subglobose conidia (Yang et al. 2019). Conidia of Hapsidosporairregularis and A.curvum had several fascicled at the tips of the conidiophores (Malloch and Cain 1970). However, A.curvum was differentiated by having mostly curved phialides and the conidia were several fascicled at the tips of the phialides.
Discussion
In the present study, four strains of Acremonium fungi were isolated from soil in the Shanxi and Henan Province, China. Two-locus (LSU and ITS) phylogenetic analyses in combination with morphological characteristics were used for identification. As a result, two new species of A.curvum (one isolate) and A.globosisporum (three isolates) were proposed.
With the development of biotechnology, a growing number of studies have combined morphological and phylogenetic features to distinguish between species. This provides the basis for more precise species naming. Generally, the fungal ITS marker includes considerably more sequence variability, and consequently provides high interspecific resolution, and also some degree of intraspecific variability (Nilsson et al. 2008). Therefore, ITS has been widely used in studies of fungal inter- and intraspecific relationships (Dai et al. 2020; Szczepańska et al. 2021). There are numerous ITS sequences stored in public databases, which are incomparable to other molecular markers (Zhang et al. 2022). In addition, according to Vu et al. (2019), combining ITS and LSU can improve the accuracy of fungal species discrimination with high generality. They think that fungi commonly present in clinical, environmental, or economically relevant communities can often be identified to species level by their ITS and LSU barcodes (Vu et al. 2019).
Although, Yang et al. (2019) used a multi-locus phylogenetic analysis in introducing the new species Acremoniumarthrinii, lacking loci such as SSU, TEF 1-α and RPB2 (Table 1) in the isolates of Acremonium spp. were relatively serious, so it is not difficult to find that the strains with SSU and TEF 1-α in this analysis were not yet 50% or even 30% of the total number of strains. Therefore, although we sequenced these above loci in the new isolates, they were not included in the phylogenetic analysis. In the future, the phylogeny relationships of Acremonium members will undoubtedly vary and become clearer with the increase of the number and type of molecular used.
In recent years, Acremonium spp. has been reported to cause immunocompetent and immunocompromised individual diseases, such as brain abscess (Anis et al. 2021), fungal keratitis (Liu et al. 2021), fungal osteomyelitis (Jalan et al. 2021), and fungal maxillary sinusitis (Durbec et al. 2011). In the present study, all strains were isolated by a method specifically designed for the isolation of keratinophilic microbes. Therefore, more studies are necessary to confirm whether A.curvum and A.globosisporum are opportunistic infectious pathogens that infect the skin and cause skin infection, as well as their potential application in the degradation of keratin-rich matrices.
Supplementary Material
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (no. 32060011, 32160007, 31860002), “Hundred” Talent Projects of Guizhou Province (Qian Ke He [2020] 6005), and the Key Areas of Research and Development Program of Guangdong Province (no. 2018B020205003), and Construction Program of Biology First-class Discipline in Guizhou (GNYL [2017] 009). We appreciate MDPI for the English-language editing of the whole manuscript.
Citation
Li X, Zhang Z-Y, Ren Y-L, Chen W-H, Liang J-D, Pan J-M, Huang J-Z, Liang Z-Q, Han Y-F (2022) Morphological characteristics and phylogenetic evidence reveal two new species of Acremonium (Hypocreales, Sordariomycetes). MycoKeys 91: 85–96. https://doi.org/10.3897/mycokeys.91.86257
References
- Anis A, Sameeullah F, Bhatti JM. (2021) A rare case of brain abscesses caused by Acremonium species. Cureus 13(4): e14396. 10.7759/cureus.14396 [DOI] [PMC free article] [PubMed]
- Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R. (2020) TBtools: An integrative toolkit developed for interactive analyses of big biological data. Molecular Plant 13(8): 1194–1202. 10.1016/j.molp.2020.06.009 [DOI] [PubMed] [Google Scholar]
- Dai YD, Wu CK, Yuan F, Wang YB, Huang LD, Chen ZH, Zeng WB, Wang Y, Yang ZL, Zeng PS, Lemetti P, Mo XX, Yu H. (2020) Evolutionary biogeography on Ophiocordycepssinensis: An indicator of molecular phylogeny to geochronological and ecological exchanges. Geoscience Frontiers 11(3): 807–820. 10.1016/j.gsf.2019.09.001 [DOI] [Google Scholar]
- Durbec M, Bienvenu AL, Picot S, Dubreuil C, Cosmidis A, Tringali S. (2011) Maxillary sinus fungal infection by Acremonium. European Annals of Otorhinolaryngology, Head and Neck Diseases 128(1): 41–43. 10.1016/j.anorl.2010.10.004 [DOI] [PubMed] [Google Scholar]
- Hyde KD, Dong Y, Phookamsak R, Jeewon R, Bhat DJ, Jones EBG, Liu NG, Abeywickrama PD, Mapook A, Wei D, Perera RH, Manawasinghe IS, Pem D, Bundhun D, Karunarathna A, Ekanayaka AH, Bao DF, Li JF, Samarakoon MC, Chaiwan N, Lin CG, Phutthacharoen K, Zhang SN, Senanayake IC, Goonasekara ID, Thambugala KM, Phukhamsakda C, Tennakoon DS, Jiang HB, Yang J, Zeng M, Huanraluek N, Liu JK, Wijesinghe SN, Tian Q, Tibpromma S, Brahmanage RS, Boonmee S, Huang SK, Thiyagaraja V, Lu YZ, Jayawardena RS, Dong W, Yang EF, Singh SK, Singh SM, Rana S, Lad SS, Anand G, Devadatha B, Niranjan M, Sarma VV, Liimatainen K, Aguirre-Hudson B, Niskanen T, Overall A, Alvarenga RLM, Gibertoni TB, Pfliegler WP, Horváth E, Imre A, Alves AL, Bahkali AH, Doilom M, Elgorban AM, Maharachchikumbura SSN, Rajeshkumar KC, Haelewaters D, Mortimer PE, Zhao Q, Lumyong S, Xu JC, Sheng J. (2020) Fungal diversity notes 1151–1276: Taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Diversity 100(1): 5–277. 10.1007/s13225-020-00439-5 [DOI] [Google Scholar]
- Ito T, Okane I, Nakagiri A, Gams W. (2000) Two species of AcremoniumsectionAcremonium: A.borodinense sp. nov. and A.cavaraeanum rediscovered. Mycological Research 104(1): 77–80. 10.1017/S0953756299008977 [DOI] [Google Scholar]
- Jalan D, Saini MK, Elhence P, Elhence A, Jain P. (2021) Calcaneal osteomyelitis caused by Acremonium sp. in an immunocompetent adult – A case report. The Foot 47: е101781. 10.1016/j.foot.2021.101781 [DOI] [PubMed]
- Katoh K, Standley DM. (2013) MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution 30(4): 772–780. 10.1093/molbev/mst010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X, Zhang ZY, Chen WH, Liang JD, Huang JZ, Han YF, Liang ZQ. (2022) A new species of Arthrographis (Eremomycetaceae, Dothideomycetes), from the soil in Guizhou, China. Phytotaxa 538(3): 175–181. 10.11646/phytotaxa.538.3.1 [DOI] [Google Scholar]
- Liu YJ, Whelen S, Hall BD. (1999) Phylogenetic relationships among Ascomycetes: Evidence from an RNA polymerse II subunit. Molecular Biology and Evolution 16(12): 1799–1808. 10.1093/oxfordjournals.molbev.a026092 [DOI] [PubMed] [Google Scholar]
- Liu J, Freiberg FJ, Yeung SN, Iovieno A. (2021) Late-onset recurrent Acremonium fungal keratitis after therapeutic penetrating keratoplasty. Canadian Journal of Ophthalmology 56(4): e135–e137. 10.1016/j.jcjo.2021.02.006 [DOI] [PubMed]
- Malloch D, Cain RF. (1970) Five new genera in the new family Pseudeurotiaceae. Canadian Journal of Botany 48(10): 1815–1825. 10.1139/b70-267 [DOI] [Google Scholar]
- Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ. (2015) IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32(1): 268–274. 10.1093/molbev/msu300 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson KH. (2008) Intraspecific ITS variability in the kingdom fungi as ex-pressed in the international sequence databases and its implications for molecular species identification. Evolutionary Bioinformatics Online 4: 193–201. 10.4137/EBO.S653 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rehner SA, Buckley E. (2005) A Beauveria phylogeny inferred from nuclear ITS and EF1-α sequences: Evidence for cryptic diversification and links to Cordycepsteleomorphs. Mycologia 97(1): 84–98. 10.3852/mycologia.97.1.84 [DOI] [PubMed] [Google Scholar]
- Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP. (2012) MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic Biology 61(3): 539–542. 10.1093/sysbio/sys029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Summerbell RC, Gueidan C, Schroers HJ, de Hoog GS, Starink M, Arocha Rosete Y, Guarro J, Scott JA. (2011) Acremonium phylogenetic overview and revision of Gliomastix, Sarocladium, and Trichothecium. Studies in Mycology 68(1): 139–162. 10.3114/sim.2011.68.06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szczepańska K, Guzow-Krzemińska B, Urbaniak J. (2021) Infraspecific variation of some brown Parmeliae (in Poland) – a comparison of ITS rDNA and non-molecular characters. MycoKeys 85: 127–160. 10.3897/mycokeys.85.70552 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vilgalys R, Hester M. (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. Journal of Bacteriology 172(8): 4238–4246. 10.1128/jb.172.8.4238-4246.1990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vu D, Groenewald M, de Vries M, Gehrmann T, Stielow B, Eberhardt U, Al-Hatmi A, Groenewald JZ, Cardinali G, Houbraken J, Boekhout T, Crous PW, Robert V, Verkley GJM. (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Studies in Mycology 92(1): 135–154. 10.1016/j.simyco.2018.05.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- White TJ, Bruns T, Lee S, Taylor J. (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (Eds) PCR protocols: a guide to methods and applications, Academic Press, San Diego, California, 315–322. 10.1016/B978-0-12-372180-8.50042-1 [DOI]
- Yang CL, Xu XL, Jeewon R, Boonmee S, Liu YG, Hyde KD. (2019) Acremoniumarthrinii sp. nov., a mycopathogenic fungus on Arthriniumyunnanum. Phytotaxa 420(4): 283–299. 10.11646/phytotaxa.420.4.4 [DOI] [Google Scholar]
- Zhang ZY, Chen WH, Zou X, Han YF, Huang JZ, Liang ZQ, Deshmukh SK. (2019a) Phylogeny and taxonomy of two new Plectosphaerella (Plectosphaerellaceae, Glomerellales) species from China. MycoKeys 57: 47–60. 10.3897/mycokeys.57.36628 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang ZY, Han YF, Chen WH, Liang ZQ. (2019b) Gongronellasichuanensis (Cunninghamellaceae, Mucorales), a new species isolated from soil in China. Phytotaxa 416(2): 167–174. 10.11646/phytotaxa.416.2.4 [DOI] [Google Scholar]
- Zhang ZY, Dong CB, Chen WH, Mou QR, Lu XX, Han YF, Huang JZ, Liang ZQ. (2020a) The enigmatic Thelebolaceae (Thelebolales, Leotiomycetes): One new genus Solomyces and five new species. Frontiers in Microbiology 11: е572596. 10.3389/fmicb.2020.572596 [DOI] [PMC free article] [PubMed]
- Zhang ZY, Zhao YX, Shen X, Chen WH, Han YF, Huang JZ, Liang ZQ. (2020b) Molecular phylogeny and morphology of Cunninghamellaguizhouensis sp. nov. (Cunninghamellaceae, Mucorales), from soil in Guizhou, China. Phytotaxa 455(1): 31–39. 10.11646/phytotaxa.455.1.4 [DOI] [Google Scholar]
- Zhang ZY, Shao QY, Li X, Chen WH, Liang JD, Han YF, Huang JZ, Liang ZQ. (2021) Culturable fungi from urban soils in China I: Description of 10 new taxa. Microbiology Spectrum 9(2): e00867–e21. 10.1128/Spectrum.00867-21 [DOI] [PMC free article] [PubMed]
- Zhang Z, Chen W, Liang Z, Zhang L, Han Y, Huang J, Liang Z. (2022) Revealing the non-overlapping characteristics between original centers and genetic diversity of Purpureocilliumlilacinum. Fungal Ecology [accepted].
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.