Abstract
Antiplatelet therapy is used in the treatment of patients with acute coronary syndromes, stroke, and those undergoing percutaneous coronary intervention. Clopidogrel is the most widely used antiplatelet P2Y12 inhibitor in clinical practice. Genetic variation in CYP2C19 may influence its enzymatic activity, resulting in individuals who are carriers of loss-of-function CYP2C19 alleles and thus have reduced active clopidogrel metabolites, have high on-treatment platelet reactivity and increased ischemic risk. Prospective studies have examined the utility of CYP2C19 genetic testing to guide antiplatelet therapy, and more recently published metanalyses suggest that pharmacogenetics represents a key treatment strategy to individualize antiplatelet therapy. Rapid genetic tests, including bedside genotyping platforms that are validated and have high reproducibility, are available to guide selection of P2Y12 inhibitors in clinical practice. The aim of this review is to provide an overview of the background and rationale for the role of a guided antiplatelet approach to enhance patient care.
Keywords: guided antiplatelet therapy, pharmacogenomics, pharmacogenetics, CYP2C19, precision medicine, dual antiplatelet therapy
INTRODUCTION
Dual antiplatelet therapy (DAPT), comprising aspirin and a P2Y12 inhibitor, is the mainstay of treatment in patients with acute coronary syndromes (ACSs) or undergoing percutaneous coronary intervention (PCI), including coronary stent placement. Although in recent years alternative P2Y12 inhibitors such as prasugrel and ticagrelor have been developed, clopidogrel continues to be most widely used because of its lower cost and rate of bleeding (1, 2). In fact, in an analysis of 64,600 patients who underwent PCI, the proportion of individuals receiving clopidogrel, prasugrel, and ticagrelor was respectively 72%, 20%, and 8% (3). Clopidogrel with aspirin is a well-established, commonly used DAPT for individuals with ischemic heart disease to reduce ischemic risk (4). However, variability in the clopidogrel response has been extensively described and is explained in part due to genetic variation, raising concerns of clopidogrel resistance or treatment failure (5).
GENETIC VARIATION AFFECTING ANTIPLATELET DRUG PATHWAYS
In particular, genetic variation in CYP2C19 may influence clopidogrel pharmacokinetics (PK) (6), providing the basis for the distinct PK phenotypes based on whether the patients carry variant CYP2C19 alleles, such as poor, normal, intermediate, extensive, or ultrarapid metabolizers (7). Clopidogrel is an oral second-generation thienopyridine with a half-life ranging from 7 to 8 h (8). Of the orally administered form of this prodrug, approximately 85% is hydrolyzed by endogenous esterases into an inactive metabolite. The remaining drug is bioactivated in the liver in a two-step oxidative process by cytochrome P450 enzymes, resulting in the production of 2-oxo-clopidogrel, which is eventually transformed to four isomeric thiol derivates, among which clopi-H4 is the only active metabolite (9). The only CYP450 enzyme that plays an important role in both steps of this oxidative process is CYP2C19. CYP2C19 is a 55.93-kDa protein expressed in hepatocytes and is encoded by a 9-exon highly polymorphic gene, of which the wild-type (WT) allele is usually indicated as *1 (e.g., homozygous WT *1/*1). Genetic variation of CYP2C19 results in variable clopidogrel active metabolite levels (7). Individuals who are carriers of loss-of-function (LOF) alleles of CYP2C19 have reduced active metabolite levels and persistent high platelet reactivity (HPR), which is associated with increased thrombotic or ischemic risk (10, 11). Currently, the Pharmacogene Variation Consortium (PharmVar) has defined over 35 star (*) allele haplotypes (12). These alleles are categorized into several functional groups: normal function (e.g., CYP2C19*1), reduced function (e.g., CYP2C19*9), loss of function (e.g., CYP2C19*2 and *3), and increased function (e.g., CYP2C19*17). The most extensively studied variants, CYP2C19*2 and CYP2C19*3, are characterized by the substitution of guanine with adenine at different nucleotide locations, effectively resulting in a truncated nonfunctional enzyme. Individuals who are homozygous for either CYP2C19*2 (i.e., *2/*2) or CYP2C19*3 (i.e., *3/*3) are poor metabolizers (PMs) of clopidogrel, resulting in significant variability in therapeutic response among patients (13, 14). Interestingly, the frequency of these alleles, and the distribution of the LOF CYP2C19 variants, significantly differs among various ancestral populations. Homozygosity for CYP2C19 LOF variants is present in 2% of Caucasians, 4% of African Americans, and 14% of East Asians individuals (15), while heterozygosity for CYP2C19*2 is noted in up to 30% of Caucasians, up to 40% of African Americans, and up to 50% of East Asians (6, 16). Moreover, CYP2C19*2 is the most prevalent haplotype in South Asians (32.5%) and East Asians (31%), while CYP2C19*3 is rare in individuals of European and African ancestry but is more common in East Asians (6.3%) and less common in South Asians (0.4%) (9). Although CYP2C19 is the most validated genetic determinant of clopidogrel response, the drug’s metabolism has also been reported to be influenced to a lesser extent by variants in other CYP450 genes such as CYP1A2 and CYP2B6 and by variants in other genes encoding the ATP-binding cassette subfamily B member 1 (ABCB1), paraoxonase-1 (PON1), carboxyl esterase 1 (CES1), and P2Y12 receptors (P2Y12) (17–21).
Pharmacogenetic determinants of other antiplatelet agents have also been studied. Unlike clopidogrel, ticagrelor is not a prodrug and does not require metabolic activation for its action. It acts directly on platelets by reversibly blocking platelet surface membrane P2Y12 receptors (22). After oral administration and absorption, ticagrelor is degraded to its primary active and inactive metabolites by CYP3A4/5 enzymes, and although it is suggested that variant alleles may potentially influence ticagrelor efficacy, this hypothesis has not been clinically validated (23, 24). Two other genetic variants have been described to be associated with plasma ticagrelor levels, including rs113681054 in the solute carrier organic anion transporter family member 1B1 (SLCO1B1) and rs61361928 in the UDP glucuronosyltransferase 2B7 (UGT2B7) (25). However, these variants have not been shown to have a significant effect on the PK and pharmacodynamics (PD) of ticagrelor (25–27).
Prasugrel, a third-generation thienopyridine, is hydrolyzed by carboxylesterases to generate a thiolactone (R-95913), which undergoes hepatic metabolism by CYP3A4, CYP2B6, CYP2C9, CYP2C19, and CYP2D6 to form its active metabolite (R-138727). However, no significant association has been detected for variation in these hepatic cytochrome P450 enzymes and either active metabolite exposure or PD response of prasugrel (28–30). Notably, ABCB1 was not significantly associated with platelet response or exposure to prasugrel’s active metabolite in the TRITON-TIMI 38 pharmacogenetic study (31).
As is well known, aspirin prevents the conversion of arachidonic acid to thromboxane A2 (TXA2) primarily by the irreversible acetylation of cyclooxygenase-1 (COX1), resulting in the inhibition of platelet aggregation. Interestingly, several studies have investigated many aspirin response candidate genes involving COX1, glycoprotein (GP)IIIa, GPIa/IIa, GPIba, GPVI, platelet endothelial aggregation receptor 1 (PEAR1), thromboxane A2 receptor (TBXA2R), ADP receptors (P2RY1 and P2RY12), coagulation factor XIII (F13A1), and UDP glucuronosyltransferase 1A6 (UGT1A6), with inconsistent results (32, 33).
IMPACT OF GENETIC VARIATION ON ANTIPLATELET DRUG AND/OR METABOLITE LEVELS
Because CYP2C19 genetic variants significantly influence variation in plasma levels of the active clopidogrel metabolite (34), with a resultant gene-dose effect, it is possible to classify individuals into several functional classes: normal metabolizers (NMs) carrying two normal function alleles (*1/*1), PMs carrying two nonfunctional alleles (*2/*2, *3/*3, or *2/*3), intermediate metabolizers (IMs) carrying one normal functional allele and one nonfunctional allele (*1/*2, *1/*3) or also one nonfunctional allele and one allele with increased function (*2/*17 or *3/17), extensive metabolizers (EMs) carrying one normal function allele and one so-called increased-function allele (*1/*17), and finally ultrarapid metabolizers (UMs)with two so-called increased-function alleles (*17/*17) (9). Furthermore, rare (<1% allelic frequency) and less studied alleles associated with CYP2C19 LOF status are CYP2C19*4, *5, *6, *7, and *8 (35, 36). Moreover, there are limited data regarding several other reduced-function alleles (e.g., CYP2C19*9 and *10); therefore, individuals who have two of these alleles, or one of these alleles combined with a normal or increased-function allele, are currently classified as likely IMs (33). Several studies have found that CYP2C19 IMs and PMs have reduced active clopidogrel metabolite concentration and HPR compared to NMs (37). In a cohort of 162 normal subjects from 6 separate studies, being a carrier of either CYP2C19*2 or *3 was associated with the highest reduction (32%, p < 0.001) in the area under the plasma concentration curve (AUC) for the active metabolite as compared to other CYP enzymes (38). In fact, CYP2C19 LOF carriers had significantly reduced active clopidogrel metabolite levels when treated with clopidogrel and were characterized by increased HPR rates and being at risk of thrombotic complications after undergoing PCI (39).
In a study by Brandt et al. (28) among CYP2C19 WT patients, the AUC over 0–24 h was 76 ± 17.9 ng·h/mL, and the maximum plasma concentration (Cmax) for the active metabolite of clopidogrel was 58.4 ± 9.2 ng/mL. Interestingly, considering CYP2C19*2 carriers, the mean AUC over 0–24 h was 41.5 ± 5.7 ng·h/mL and Cmax was 35.3 ± 4.3 ng/mL, levels that were 54% and 60%, respectively, of those of CYP2C19 WT individuals. Furthermore, considering CYP2C19*2 homozygous individuals, the AUC over 0–24 h and Cmax were 65% and 42%, respectively, of those of CYP2C19 WT individuals (28). A different study, including individuals with the CYP2C19*3 allele, as well as those with the CYP2C19*2 allele, showed that the AUC over 0–24 h and Cmax for the active metabolite of clopidogrel in heterozygote individuals were 71% and 67%, respectively, of those of WT individuals, while for homozygotes, the levels were 57% and 61%, respectively (40). Increasing clopidogrel dose may increase the bioavailability of the drug and may be useful in overcoming reduced active clopidogrel metabolite concentrations in CYP2C19 LOF carriers (41). A randomized pharmacogenetics study suggested that a clopidogrel loading dose of 600 mg with a maintenance dose of 150 mg led to improvement of active clopidogrel metabolite clopi-H4 levels in CYP2C19 LOF carriers comparable to levels observed with a lower loading dose of 300 mg followed by the typical 75 mg maintenance dose in CYP2C19 WT patients (6).
A genome-wide association study (GWAS) demonstrated that plasma levels of ticagrelor and its metabolite were associated with variation in SLCO1B1, (25) CYP3A4, and UGT2B7. However, there was no effect of genetic variation in CYP3A5, CYP2B6, CYP2C9, or CYP2C19 on prasugrel PK (28, 30).
IMPACT OF GENETIC VARIATION ON ON-TREATMENT PLATELET FUNCTION AND AGGREGATION
Reduced inhibition of platelet aggregation has been observed in CYP2C19 LOF carriers when treated with clopidogrel, leading to an increased thrombotic or ischemic risk (10, 11). HPR in response to clopidogrel therapy affected up to 40% of patients (42, 43), and in a recent meta-analysis of 4 studies and 4,341 subjects receiving a 600-mg loading dose of clopidogrel, there was significant residual HPR reflecting a gene-dose effect in CYP2C19*2 carriers as compared to noncarriers (44). Consequently, HPR has been recommended as a surrogate marker of clopidogrel resistance and adverse cardiovascular (CV) outcomes and has been used to individualize antiplatelet therapy (45). Although bioavailability of clopidogrel increased with an increased maintenance dose of 150 mg daily as compared to 75 mg, HPR did not significantly change in CYP2C19 LOF carriers (46). Notably, in the ELEVATE-TIMI 56 (Escalating Clopidogrel by Involving a Genetic Strategy—Thrombolysis in Myocardial Infarction 56) trial, a clopidogrel dose of 225 or 300 mg was required to significantly reduce the number of CYP2C19*2 LOF carriers who had HPR from 52% to 10% (p < 0.001), but homozygotes remained resistant with a dose as high as 300 mg (47). Platelet function tests (PFTs), which include bleeding time, platelet aggregation via light transmission, lumiaggregometry, blood impedance aggregometry, and flow cytometry, have been used to determine platelet function and reactivity (48).
Some studies have suggested that CYP2C19 genetic polymorphisms account only for 12% of the interindividual variability observed in platelet aggregation in individuals undergoing PCI, and that over 80% of the variability might be attributed to other genetic or nongenetic factors (34, 49, 50). Notably, it has been suggested that combining the PM phenotype with BMI and diabetes mellitus status could be a reliable predictor of resistance to platelet inhibition by clopidogrel (51). In addition, patients with chronic kidney disease demonstrated HPR on clopidogrel (52) and had increased adverse CV outcomes after PCI, in part related to a higher HPR at baseline prior to therapy (53). Therefore, a systems biology approach beyond pharmacogenomics encompassing clinical variables, multigenomic marker models, transcriptomics, epigenomics, exposomics, proteomics, metabolomics, and the microbiome has been proposed (9).
Despite its effectiveness, a considerable number of individuals show suboptimal aspirin response assessed by PFTs, and patients with HPR to aspirin show increased risk of ischemic events (54). Considering that aspirin leads to nearly complete inhibition of COX1 in approximately 95% of individuals (55), it is understandable how part of the variability in its response is mediated by factors outside of this pathway (54). In a study assessing the influence of CYP2C19 alleles on prasugrel response using the platelet reactivity index (PRI), as with clopidogrel, CYP2C19*2 carriers had a significantly higher PRI and risk of HPR than did noncarriers, which was not observed when platelet reactivity was assessed by adenosine diphosphate–induced platelet aggregation (56, 57). Genetic variation in CYP2C19 and other genes does not significantly affect the platelet response to ticagrelor (58).
The premise of using results of PFT as a reliable surrogate marker of adverse CV outcomes has been questioned given both variability in testing and that augmented antiplatelet therapeutic interventions to modify inhibition of platelet aggregation have not translated into improved clinical outcomes (59, 60). As an alternative to PFTs, performing a rapid genetic test to identify CYP2C19 LOF carriers who invariably have on-treatment HPR is another feasible and validated point-of-care assay that would facilitate guided selection of antiplatelet therapy in clinical practice with high reproducibility (61, 62). However, many such commercial genotyping platforms only detect common variants such as *2, *3, and *17 and could misclassify other rare CYP2C19 LOF genotypes such as *4–*7 and *20–*21 as normal. In fact, with the advent of sequencing, there are hundreds of other reported missense and predicted LOF variants, and current genotyping approaches could miss all of them (63). However, these rare genetic variants comprise less than 1% of actionable pharmacogenetic variants (9). A challenge will be establishing the functional significance, not only in the laboratory but also clinically, of this ever-increasing number of reported rarer variants for which some groups have begun establishing high-throughput techniques (63, 64).
IMPACT OF GENETIC VARIATION ON CLINICAL OUTCOMES OF ANTIPLATELET THERAPY
Clopidogrel for Coronary Artery Disease
Multiple variants in multiple genes have been investigated for an association with clinical outcomes in coronary artery disease (CAD) patients treated with clopidogrel (e.g., ABCB1, CYP2B6, CYP2C9, and PON1), but the strongest evidence is for CYP2C19 LOF variants and comes from several meta-analyses (65–67). Several organizations have determined that this body of evidence is sufficiently strong to make clinical recommendations for patients based on CYP2C19 LOF variants in clinical practice guidelines and drug labels, and these are summarized in Table 1. The evidence for CYP2C19 LOF variants and clinical outcomes in CAD patients has already been extensively reviewed elsewhere (7, 9). Thus, this section focuses on recently published studies that extend the evidence to other genetic variants such as the CYP2C19 so-called gain-of-function variant (*17; rs12248560), a polygenic score that includes multiple candidate variants, and a GWAS discovering novel variants.
Table 1.
Pharmacogenetic information for antiplatelet drugs in clinical practice guidelines and drug labels
| Antiplatelet drug | Clinical practice guidelines (134) | Regulatory agency drug labels (134) |
|---|---|---|
| Clopidogrel | CPIC,a DPWG,a RNPGx,a AHA/ACC (4), ESC (135) | EMA,a FDA,a HCSC,a PMDA,a Swissmedica |
| Aspirin | CPIC | Swissmedica |
| Ticagrelor | DPWG | EMA, FDA |
| Prasugrel | DPWG | EMA, FDA, Swissmedic |
These are defined as actionable clinical recommendations by the Pharmacogenomics Knowledgebase (PharmGKB) (134). Other entries are only informational (134).
Abbreviations: ACC, American College of Cardiology; AHA, American Heart Association; CPIC, Clinical Pharmacogenetics Implementation Consortium; DPWG, Dutch Pharmacogenetics Working Group; EMA, European Medicines Agency; ESC, European Society of Cardiology; FDA, US Food and Drug Administration; HCSC, Health Canada (Santé Canada); PMDA, Pharmaceuticals and Medical Devices Agency, Japan; RNPGx, French National Network of Pharmacogenetics.
Unlike for CYP2C19 LOF variants, the evidence supporting CYP2C19 gain-of-function variants, leading to CYP2C19 EM (e.g., *1/*17) and UM (e.g., *17/*17) phenotypes, was not as clear. Therefore, Lee et al. (68) recently published an analysis of atherothrombotic and bleeding outcomes in 3,342 CAD patients that underwent PCI at nine different sites implementing CYP2C19-guided antiplatelet therapy into clinical care [the Implementing Genomics in Practice (IGNITE) Network Pharmacogenetics Working Group] (69). Compared to CYP2C19 NMs (e.g., *1/*1) treated with clopidogrel, patients that were CYP2C19 EMs or UMs had a similar risk of atherothrombotic and bleeding events when treated with clopidogrel. These findings by Lee et al. are consistent with another recently published study by Claassens et al. (70), which was a prespecified subanalysis (n = 2,429) of the POPular Genetics trial. Claassens et al. also found that the CYP2C19*17 allele was not significantly associated with thrombotic or bleeding outcomes. In summary, this recent evidence shows that the so-called gain-of-function CYP2C19*17 variant does not play a significant role in the clinical outcomes of CAD patients treated with clopidogrel.
Some evidence suggests that variants in other metabolic genes may also play a role in clopidogrel response, such as those in carboxylesterase 1 (CES1) and CYP2B6. Therefore, Lewis et al. (18) calculated a polygenic clopidogrel response score with variants from six different candidate genes: CYP2C19*2 (rs4244285), CES1 G143E (rs71647871), CYP2B6 1294 + 53 C>T (rs8192719), CYP2B6 Q172 (rs3745274), CYP2C9*3 (rs1057910), and PEAR1 rs12041331. They calculated the score by adding together the number of risk alleles (0, 1, 2) in each CAD patient that significantly increased on-clopidogrel platelet reactivity. Then they tested the association of the polygenic response score with major adverse CV events (MACEs), including myocardial infarction, ischemic stroke, stent thrombosis, and CV death, in 2,134 Caucasian patients with CAD in the International Clopidogrel Pharmacogenomics Consortium (ICPC) (71). An increase in polygenic response score was significantly associated with an increased risk of MACEs [odds ratio (OR) = 1.19, p = 0.01] and CV death specifically (OR = 1.54, p = 0.007). The clinical utility of the polygenic clopidogrel response score will need to be evaluated in the future, especially compared to the use of CYP2C19 LOF variants alone, but this study by Lewis et al. lays the groundwork for the use of additional variants beyond CYP2C19 variants.
Finally, another recent study of interest in CAD patients is a GWAS of clinical outcomes on clopidogrel by Verma et al. (17). A previous GWAS of clopidogrel response was published by Shuldiner et al. (72) in 2009, but it was a small and specific sample (i.e., 429 healthy Amish participants). The Shuldiner et al. 2009 GWAS estimated that approximately 70% of the variability in on-clopidogrel platelet reactivity can be explained by genetics, but only approximately 12% of the variability is explained by the CYP2C19*2 variant. This means that other, yet-undiscovered genetic variants play a role in clopidogrel response. Therefore, in 2020, Verma et al. (17) published a much larger GWAS (n = 2,750) of participants of European ancestry treated for different indications with clopidogrel (mostly CAD). They analyzed multiple different clinical outcomes, but only the results for a combined endpoint of CV death and myocardial infarction are summarized herein. No individual genetic variants reached the level of genome-wide statistical significance in the overall analysis, but genome-wide significant results were detected in every analysis when the patients were stratified into specific clinical subgroups (i.e., patients with CAD, n = 2,509; those who underwent PCI, n = 2,065; and those with ACSs, n = 1,188). Variants mapping to SOCS5P1 (pseudogene 1 of the suppressor of cytokine signaling 5) were statistically significant in all the clinical subgroup analyses. The mechanism of SOCS5P1 in clopidogrel response is unknown and thus needs to be understood, and these findings will need to be replicated in an independent sample to confirm their clinical validity. These GWAS findings support a potential role for genetic variants outside of previously studied candidate genes, which is consistent with a recent study showing that most pharmacogenomic GWASs discover novel genes outside of previously studied candidate genes (73).
Clopidogrel for Stroke
The body of literature investigating the association of genetic variants with clinical outcomes in patients treated with clopidogrel for stroke is not as extensive as that for CAD, but the evidence also strongly supports a role for CYP2C19 LOF variants in large studies and meta-analyses (74–76). Some organizations have published clinical recommendations for clopidogrel pharmacogenetics specifically for stroke as well as for CAD (77) (Table 1), and some of the pharmacogenetic clinical recommendations are broad and not limited to specific indications (78). Others have recently reviewed the body of literature on stroke in detail (79, 80); thus, this section focuses on highlighting three of the most recently published studies, which extend the evidence for CYP2C19 LOF variants to non-European patients.
Li et al. (81) recently published a retrospective analysis of 201 elderly Chinese patients with noncardioembolic ischemic stroke treated with clopidogrel for a composite primary outcome of stroke or myocardial infarction at 2 years. They compared the primary outcome in a CYP2C19 genotype–guided group (n = 99) and a non-genotype-guided group (n = 102). The rate of the primary outcome was significantly lower in the CYP2C19 genotype–guided group than in the non-genotype-guided group. Lv et al. (82) had similar findings in a slightly larger study of Chinese patients with much longer follow-up. Lv et al. prospectively studied 314 Chinese patients with ischemic stroke who were treated with clopidogrel for the primary composite outcome of recurrent ischemic stroke, transient ischemic attack (TIA), myocardial infarction, or ischemic vascular death for approximately 4.5 years. Patients carrying a CYP2C19 LOF allele had significantly increased risk of the primary outcome. Finally, Al-Rubaish et al. (83) had similar findings in their small, prospective study of 42 Saudi Arabian patients with ischemic stroke who were treated with clopidogrel for the outcome of recurrent stroke at 6 months. Even in this very small sample with much shorter follow-up, patients carrying CYP2C19 LOF alleles had a significantly increased risk of recurrent stroke. Therefore, these most recent studies provide further evidence in non-European stroke patients that CYP2C19 LOF variants significantly increase the risk of adverse clinical outcomes in patients treated with clopidogrel.
Aspirin
Early evidence indicated that variants in cyclooxygenase, thromboxane, and platelet receptor genes were associated with the antiplatelet effects of aspirin, but the evidence translating those effects to clinical outcomes is in much earlier stages than the evidence for clopidogrel. To our knowledge, only a single meta-analysis that evaluated the association of a genetic variant with clinical outcomes in patients treated with aspirin has been published (84), and no significant association was found for the rs5918, a single-nucleotide substitution in the gene for GPIIIa, with MACEs. Therefore, it is not surprising that there are no pharmacogenetic clinical recommendations for aspirin in clinical practice guidelines or drug labels yet (Table 1). Others have recently reviewed the literature for aspirin pharmacogenetics in detail (85, 86); thus, this section focuses on highlighting the most recently published aspirin pharmacogenetic studies that assessed clinical outcomes.
Genetic variants in PEAR1 had been previously associated with platelet aggregation response to aspirin (87), but the association with CV clinical outcomes was less clear. Therefore, Lewis et al. (88) recently published a clinical outcomes analysis of aspirin and the rs12041331 G>A variant in PEAR1. They performed a post hoc pharmacogenetic analysis of 13,547 healthy older individuals in the Aspirin in Reducing Events in the Elderly (ASPREE) trial, which was a double-blind, randomized (1:1), placebo-controlled trial to examine the effect of low-dose aspirin (100 mg daily) on disability-free survival. They analyzed incident CV, cerebrovascular, and bleeding outcomes, but the PEAR1 variant was not significantly associated with any of these clinical outcomes. However, the effects of the PEAR1 rs12041331 may depend on whether aspirin is being used for primary or secondary stroke prevention and the subtype of stroke. In a recently published pharmacogenetic analysis of 868 ischemic stroke patients treated with aspirin alone, Li et al. (89) found that PEAR1 rs12041331 was only significantly associated with functional outcomes in patients with the small-artery occlusion subtype of stroke. Therefore, the difference in findings for the PEAR1 rs12041331 variant between the studies by Lewis et al. and Li et al. may be because the Lewis et al. patient sample was healthier than the patient sample analyzed by Li et al., and perhaps PEAR1 rs12041331 only interacts with aspirin benefit for a particular subtype of stroke. These potential explanations need to be confirmed in future research.
In addition to those recent studies of efficacy, multiple recent studies have evaluated the association of genetic variants with the adverse gastrointestinal effects of aspirin (90–94). Bourgeois et al. (90) recently published a GWAS of 247 aspirin-induced cases of endoscopically confirmed peptic ulcer disease compared to 476 controls. They found one statistically significant variant that replicated in an additional 84 cases and 162 controls, rs12678747, which is an intronic variant in EYA1 (eyes absent transcriptional coactivator and phosphatase 1). In the meta-analysis of the discovery and replication data sets, rs12678747 reached the genome-wide level of statistical significance. The mechanism of this intronic EYA1 variant in aspirin-induced peptic ulcer disease is unclear and requires further replication and mechanistic investigation to confirm biologic plausibility. Recently published candidate gene association studies found variants in PTGS1, NOS3, CHST2, and GSTP1 to be associated with gastrointestinal bleeding in patients treated with aspirin, but the findings for these variants need to be independently replicated to confirm their clinical validity (91–94). In summary, although there is not as much pharmacogenetic evidence for clinical outcomes with aspirin as there is for clopidogrel, there appears to be some evidence supporting a role for pharmacogenetics in identifying the adverse gastrointestinal effects of aspirin.
Ticagrelor
There are fewer pharmacogenetic studies of clinical outcomes in patients treated with ticagrelor. Most of the recently published studies (within the past 5 years) investigating the pharmacogenetics of ticagrelor had platelet reactivity, and not clinical outcomes, as the primary outcome. A few of those studies assessed clinical outcomes as well as platelet reactivity (95–97), but they did not detect significant differences in clinical outcomes, which is not surprising given that these studies were underpowered to detect clinical outcomes. This section focuses on the pivotal, albeit older, studies assessing the association of genetic variants with clinical outcomes in patients treated with ticagrelor [published by Wallentin et al. (98) in 2010 and Varenhorst et al. (25) in 2015]. Readers are also referred to recently published detailed reviews of the ticagrelor pharmacogenetics literature (79, 99).
Wallentin et al. performed a genetic substudy of 10,285 patients from the landmark PLATO (Platelet Inhibition and Patient Outcomes) trial (98, 100) and compared the efficacy of ticagrelor versus clopidogrel in patients with ACSs. The primary efficacy endpoint was the composite of CV death, myocardial infarction, or stroke up to 12 months, and they also investigated bleeding as a safety outcome. Patients were genotyped for CYP2C19 LOF and gain-of-function alleles and the ABCB1 3435 C>T variant. Regardless of CYP2C19 or ABCB1 genotypes, ticagrelor was consistently associated with better clinical outcomes than was clopidogrel, and there were no significant differences in bleeding events by genotypes. Therefore, the large study by Wallentin et al. (98) demonstrates that genetic variants in CYP2C19 and ABCB1 do not significantly affect clinical outcomes in ACS patients treated with ticagrelor. Five years later, Varenhorst et al. (25) published a GWAS of the PLATO trial to discover whether other genetic variants significantly predicted clinical outcomes in ACS patients treated with ticagrelor (clinical outcomes assessed in 4,990 patients). Even though they found significant effects of variants in SLCO1B1, CYP3A4, and UGT2B7 on ticagrelor parent and active metabolite plasma levels, those effects did not translate into significant differences in the primary composite ischemic outcome or bleeding. In summary, although there are not as many pharmacogenetic studies published on the clinical outcomes from ticagrelor, the available evidence is strong and shows that genetic variants do not play a large role in the clinical outcomes of patients treated with ticagrelor. This has led the pharmacogenetic recommendations for ticagrelor in clinical practice guidelines and drug labels to be mostly informational (Table 1).
Prasugrel
Similar to ticagrelor, most of the recently published pharmacogenetic literature (within the past 5 years) for prasugrel has focused on platelet reactivity as opposed to clinical outcomes, and the strongest evidence for clinical outcomes is from older subanalyses of a landmark clinical trial: TRITON-TIMI 38 (Trial to Assess Improvement in Therapeutic Outcomes by Optimizing Platelet Inhibition with Prasugrel–Thrombolysis in Myocardial Infarction) (2). Therefore, this section focuses on the pivotal, albeit older, studies assessing the association of genetic variants with clinical outcomes in patients treated with prasugrel [published by Mega et al. (31, 101, 102)] and the recently published study by Sawayama et al. (103) that assessed clinical outcomes.
The TRITON-TIMI 38 trial compared clinical outcomes of patients with ACSs and planned PCI who were randomly assigned to treatment with prasugrel or clopidogrel for up to 15 months (2). The primary efficacy end point was a composite of CV death, myocardial infarction, or stroke. A prespecified secondary end point was definite or probable stent thrombosis, and safety end points included non–coronary artery bypass graft–related thrombolysis in myocardial infarction major or minor bleeding. A total of 1,466 patients treated with prasugrel were available for the pharmacogenetic substudies. Mega et al. (31, 101, 102) assessed the association of genetic variants in ABCB1, CYP2C19, CYP2C9, CYP2B6, CYP3A5, CYP3A4, CYP1A2, and PON1 with clinical outcomes, but none were statistically significant.
To our knowledge, only one study assessing the association of genetic variants with clinical outcomes in patients treated with prasugrel has been recently published (103), and the findings by Sawayama et al. are consistent with the pharmacogenetic sub-analyses of the TRITON-TIMI 38 trial by Mega et al. Sawayama et al. performed a retrospective, observational analysis of 193 clopidogrel-treated and 217 prasugrel-treated patients that underwent PCI at a single center in Japan. They genotyped the patients for CYP2C19*2, *3, and *17, and the primary outcome was a MACE or major bleeding at 1 year. They found a significant difference in the primary outcome between patients treated with clopidogrel versus prasugrel, but only if the patients were carrying CYP2C19 LOF alleles. In patients without CYP2C19 LOF alleles, there was no significant difference between clopidogrel and prasugrel. This recent study by Sawayama et al. and the larger studies published by Mega et al. consistently demonstrate that, unlike clopidogrel, clinical outcomes in patients treated with prasugrel are not affected by variants in several candidate genes. As with ticagrelor, this finding supports the lack of clinical pharmacogenetic recommendations for prasugrel in clinical practice guidelines and drug labels (Table 1).
DOES ALTERING ANTIPLATELET THERAPY BASED ON GENETIC TEST RESULTS CHANGE OUTCOMES?
Clinical utility of pharmacogenetic testing can be established if prospective data demonstrate that altering antiplatelet therapy based on the pharmacogenetic test can result in a change in clinical outcomes. There have been several prospective studies and large meta-analyses that have been performed that primarily support the use of CYP2C19 pharmacogenetic testing in clinical practice. A prospective study of 1,815 stable CAD and ACS post-PCI patients, in which genotyping results were made available and the decision to choose a P2Y12 inhibitor was left to the medical practitioner, demonstrated that patients with CYP2C19 LOF alleles receiving ticagrelor or prasugrel as compared to clopidogrel had significantly reduced MACEs (104). Lack of randomization with assigned therapy may have confounded the results, although propensity matching was performed to adjust for group differences (104). This study demonstrated the beneficial effect on real-world clinical outcomes of providing CYP2C19 pharmacogenetic test results to the medical practitioner.
Multiple other studies have examined the utility of CYP2C19 genetic testing in guiding P2Y12 inhibitor therapy (105). A prospective randomized clinical trial (RCT) comprising 600 Chinese PCI patients reported that CYP2C19 genotype–directed antiplatelet therapy significantly decreased the cumulative incidence of MACEs (106). In the POPular Genetics noninferiority RCT, 2,488 ST elevation myocardial infarction patients undergoing primary PCI were randomized to CYP2C19 genotype–guided P2Y12 inhibitor therapy or routine treatment with ticagrelor or prasugrel (107). This trial demonstrated that a genotype-guided strategy was noninferior to standard treatment with ticagrelor or prasugrel at 12 months in reducing ischemic events and actually resulted in a lower incidence of bleeding (107).
In the TAILOR PCI trial, CAD patients were randomized to CYP2C19 genotype–guided P2Y12 inhibitors or conventional therapy with clopidogrel after PCI (108). The trial included 5,302 patients, and follow-up was for 12 months, with the primary composite ischemic endpoint occurring in 4.0% of the genotype-guided group and 5.9% of the conventional group of CYP2C19 LOF patients (hazard ratio 0.66, p = 0.06). The 34% risk reduction with genetic-guided therapy did not meet the predetermined 50% reduction that was specified in the statistical analysis plan (108). A post hoc Bayesian reanalysis of the trial, using informative priors from four previous clinical trials (POPular Genetics, PHARMCLO, IAC-PCI, and ADAPT-PCI), demonstrated that the probability of genotype-guided P2Y12 inhibitor therapy in reducing MACEs was 99% (109), clearly demonstrating the potential benefit of genetic testing in this clinical scenario.
A recent meta-analysis that included 15,949 patients (77% PCI, 98% ACS) from 7 RCTs reported that treatment with prasugrel or ticagrelor reduced major ischemic events compared to clopidogrel in CYP2C19 LOF carriers, whereas no difference was observed in noncarriers (110). Notably, a significant genotype-treatment interaction was reported (p = 0.013), leading to the conclusion that the benefit of prasugrel or ticagrelor over clopidogrel was primarily due to CYP2C19 genotype status (110). This study therefore establishes the basis for CYP2C19 pharmacogenetic testing to identify and treat LOF carriers with ticagrelor or prasugrel and noncarriers who could be prescribed clopidogrel (Figure 1). In addition, another meta-analysis of 20,743 patients, including 11 randomized controlled trials and 3 observational studies, demonstrated that guided selection of antiplatelet therapy using CYP2C19 genetic testing and PFTs significantly improves the composite of MACEs and reduces individual ischemic outcomes, including CV death, myocardial infarction, stent thrombosis, and stroke, with a significant reduction in minor bleeding. This study, along with other studies described above, supports the practice of a precision medicine approach for the selection of antiplatelet therapy in patients with CAD (111).
Figure 1.
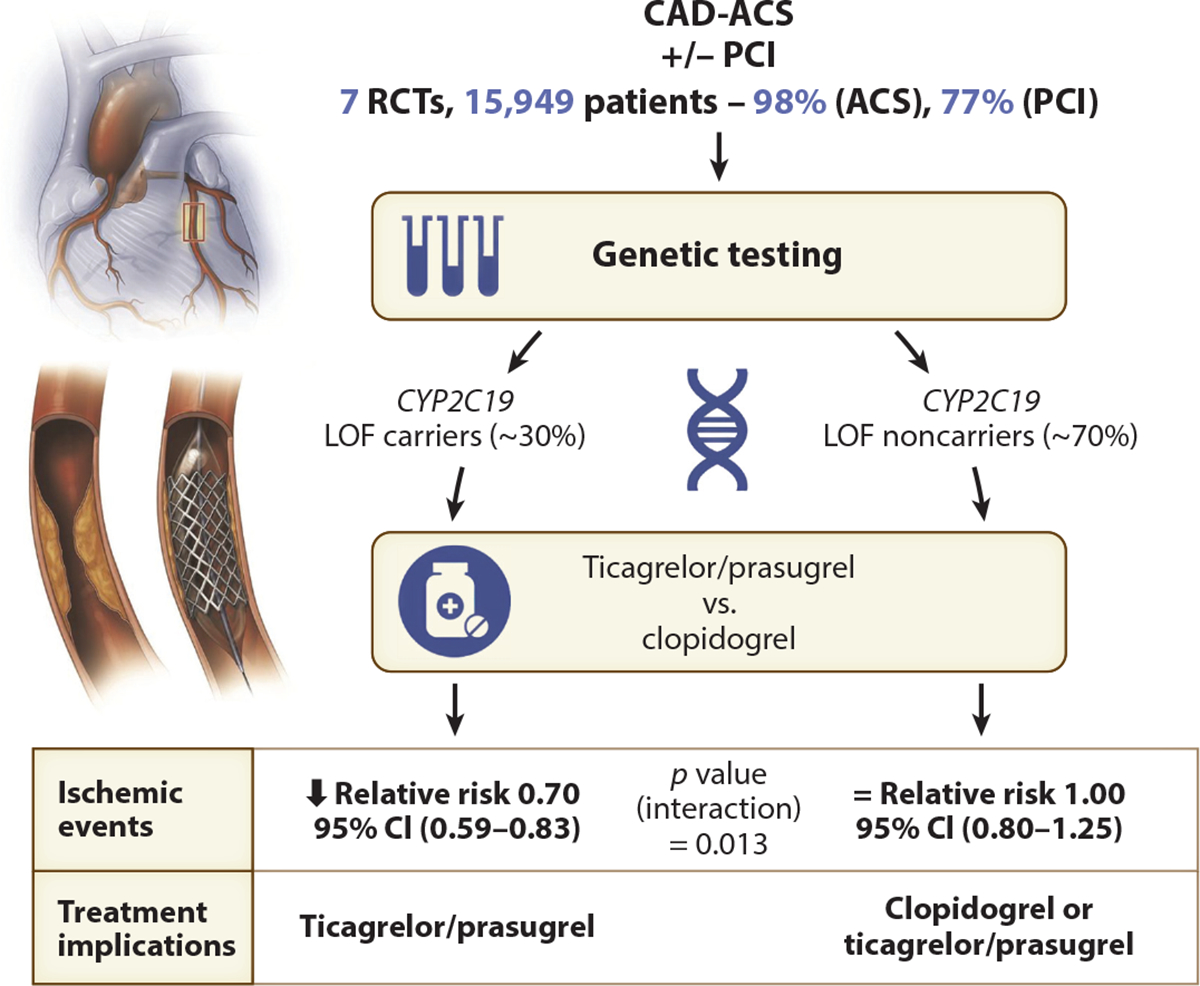
A proposed algorithm using CYP2C19 pharmacogenetic testing to individualize oral P2Y12 inhibitor therapy in patients with coronary artery disease on the basis of the meta-analysis results. The overall prevalence of 30% CYP2C19 LOF carriers does not reflect the prevalence of up to 50% observed in Asian populations. Abbreviations: ACS, acute coronary syndrome; CAD, coronary artery disease; CI, confidence interval; LOF, loss-of-function; PCI, percutaneous coronary intervention; RCT, randomized clinical trial. Figure adapted from Reference 110. Illustrations copyright Mayo Clinic.
The benefit of pharmacogenetic testing has also been observed in the treatment of stroke and TIAs. In the CHANCE-2 trial, DAPT with ticagrelor and aspirin was superior to clopidogrel and aspirin in reducing the risk of stroke in Chinese patients who were CYP2C19 LOF carriers (112). Therefore, identifying LOF patients by pharmacogenetic testing prior to treatment with P2Y12 inhibitors would be important for stroke patients as well. The risk of severe or moderate bleeding did not differ between the two groups, but ticagrelor was associated with more minor bleeding events than was clopidogrel (112).
COST-EFFECTIVENESS AND INSURANCE REIMBURSEMENT FOR GENETIC TESTING
Current guidelines that have yet to be updated support the universal use of ticagrelor or prasugrel regardless of CYP2C19 genotype status (113, 114). However, the decreased bleeding risk with the equivalent ischemic outcomes of a genetics-guided approach may be cost effective (113, 115, 116). The implementation of such an approach supports the use of the less expensive P2Y12 inhibitor clopidogrel in noncarriers, which in most populations comprise the majority as compared to LOF carriers (111). Insurance coverage for CYP2C19 genetic testing is available from selected providers; moreover, the cost of genetic testing has dramatically decreased over the past decade. Notably, the University of Florida Health Personalized Medicine Program reported that approximately 85% of third-party payers (including Medicare) reimbursed CYP2C19 genotyping to select P2Y12 inhibitors for PCI patients in 2014 (117), and reimbursement for pharmacogenetic testing has further improved since then (118).
PATIENT-PHYSICIAN PERCEPTIONS AND FACILITATION OF CLINICAL IMPLEMENTATION
Physician adoption of clinical CYP2C19 testing has not been widespread, which is likely due to a number of barriers, including the need for rapid results, testing logistics, practice guidelines, inclusion of genotype data in electronic medical records, intuitive decision support software and tools, insurance coverage, physician and patient education, and perception (119–122). Education in pharmacogenetics at all levels (physicians, assistants, pharmacists, nurses, patients) is going to be important for effective clinical implementation (123). Pharmacogenetic education significantly improved physicians’ attitudes toward pharmacogenetic testing, particularly for clopidogrel (122). A recent survey of patients focused on evaluating perceptions regarding pharmacogenetic testing and exploring geographical differences (124). Fewer Korean (44%) as compared to Canadian (91%) and US (89%) patients identified pharmacogenetics as important in their care (p < 0.001), which was primarily based on patients’ confidence in their capacity to understand genetic information emphasizing the importance of genetic education to increase clinical uptake (124). Failure to clinically implement a genotyping approach to guiding P2Y12 inhibitor therapy, especially in populations that have a high prevalence of CYP2C19 LOF alleles, may also have legal implications (125). A recent judicial verdict resulted in Bristol Myers Squibb and Sanofi S.A. being fined $834 million for failing to warn healthcare providers and patients regarding clopidogrel ineffectiveness in PMs in the drug labeling information (126). Whether the medical practitioner could be liable for not acting upon this drug labeling information is not known.
FUTURE DIRECTIONS (PREEMPTIVE GENETIC TESTING AND ARTIFICIAL INTELLIGENCE APPROACHES)
Preemptive pharmacogenetic testing may be an approach that could overcome the issue of having genetic test results rapidly available, which is especially important in the CAD, stroke, and PCI setting (127). In fact, this strategy allows incorporation of CYP2C19 genotype data into patient electronic medical records that allows the creation of alerts when clopidogrel is prescribed for a patient with a CYP2C19 LOF allele (128–130). It is also important to recognize that patients are increasingly undergoing direct-to-consumer genetic testing, the results of which are being made available to their health-care providers (131). Given the overall low adverse event rate in the population of patients undergoing PCI, especially with advances in coronary stent technology, it becomes increasingly important to identify the high- or low-risk patient for whom genetic testing could potentially guide escalation or de-escalation of antiplatelet therapy Scores that combine CYP2C19 genetic testing results with clinical factors have been successfully developed for this purpose (132). Unbiased or agnostic machine learning approaches are also being used to integrate pharmacogenomic and clinical data to identify patients’ risk profiles, which eventually may utilize other high-resolution omic profiling to realize the goal of individualizing the selection of antiplatelet therapy for patients with stroke and CAD (133).
Footnotes
DISCLOSURE STATEMENT
The authors are not aware of any affiliations, memberships, funding, or financial holdings that might be perceived as affecting the objectivity of this review.
LITERATURE CITED
- 1.Wallentin L, Becker RC, Budaj A, Cannon CP, Emanuelsson H, et al. 2009. Ticagrelor versus clopidogrel in patients with acute coronary syndromes. N. Engl. J. Med. 361(11):1045–57 [DOI] [PubMed] [Google Scholar]
- 2.Wiviott SD, Braunwald E, McCabe CH, Montalescot G, Ruzyllo W, et al. 2007. Prasugrel versus clopidogrel in patients with acute coronary syndromes. N. Engl. J. Med. 357(20):2001–15 [DOI] [PubMed] [Google Scholar]
- 3.Karve AM, Seth M, Sharma M, LaLonde T, Dixon S, et al. 2015. Contemporary use of ticagrelor in interventional practice (from Blue Cross Blue Shield of Michigan Cardiovascular Consortium). Am. J. Cardiol. 115(11):1502–6 [DOI] [PubMed] [Google Scholar]
- 4.Levine GN, Bates ER, Bittl JA, Brindis RG, Fihn SD, et al. 2016. 2016 ACC/AHA guideline focused update on duration of dual antiplatelet therapy in patients with coronary artery disease: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines: an update of the 2011 ACCF/AHA/SCAI guideline for percutaneous coronary intervention, 2011 ACCF/AHA guideline for coronary artery bypass graft surgery, 2012 ACC/AHA/ACP/AATS/PCNA/SCAI/STS guideline for the diagnosis and management of patients with stable ischemic heart disease, 2013 ACCF/AHA guideline for the management of ST-elevation myocardial infarction, 2014 AHA/ACC guideline for the management of patients with non-ST-elevation acute coronary syndromes, and 2014 ACC/AHA guideline on perioperative cardiovascular evaluation and management of patients undergoing noncardiac surgery. Circulation 134(10):e123–55. Erratum. 2016. Circulation 134(10):e192–94 [DOI] [PubMed] [Google Scholar]
- 5.Gurbel PA, Bliden KP, Hiatt BL, O’Connor CM. 2003. Clopidogrel for coronary stenting: response variability, drug resistance, and the effect of pretreatment platelet reactivity. Circulation 107(23):2908–13 [DOI] [PubMed] [Google Scholar]
- 6.Simon T, Bhatt DL, Bergougnan L, Farenc C, Pearson K, et al. 2011. Genetic polymorphisms and the impact of a higher clopidogrel dose regimen on active metabolite exposure and antiplatelet response in healthy subjects. Clin. Pharmacol. Ther. 90(2):287–95 [DOI] [PubMed] [Google Scholar]
- 7.Pereira NL, Rihal CS, So DYF, Rosenberg Y, Lennon RJ, et al. 2019. Clopidogrel pharmacogenetics. Circ. Cardiovasc. Interv. 12(4):e007811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Schrör K. 1998. Clinical pharmacology of the adenosine diphosphate (ADP) receptor antagonist, clopidogrel. Vasc. Med. 3(3):247–51 [DOI] [PubMed] [Google Scholar]
- 9.Brown SA, Pereira N. 2018. Pharmacogenomic impact of CYP2C19 variation on clopidogrel therapy in precision cardiovascular medicine. J. Pers. Med. 8(1):8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sibbing D, Aradi D, Alexopoulos D, Ten Berg J, Bhatt DL, et al. 2019. Updated expert consensus statement on platelet function and genetic testing for guiding P2Y(12) receptor inhibitor treatment in percutaneous coronary intervention. JACC Cardiovasc. Interv. 12(16):1521–37 [DOI] [PubMed] [Google Scholar]
- 11.Aradi D, Kirtane A, Bonello L, Gurbel PA, Tantry US, et al. 2015. Bleeding and stent thrombosis on P2Y12-inhibitors: collaborative analysis on the role of platelet reactivity for risk stratification after percutaneous coronary intervention. Eur. Heart J. 36(27):1762–71 [DOI] [PubMed] [Google Scholar]
- 12.Botton MR, Whirl-Carrillo M, Del Tredici AL, Sangkuhl K, Cavallari LH, et al. 2021. PharmVar Gene-Focus: CYP2C19. Clin. Pharmacol. Ther. 109(2):352–66 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Helsby NA, Burns KE. 2012. Molecular mechanisms of genetic variation and transcriptional regulation of CYP2C19. Front. Genet. 3:206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wei YQ, Wang DG, Yang H, Cao H. 2015. Cytochrome P450 CYP 2C19*2 Associated with adverse 1-year cardiovascular events in patients with acute coronary syndrome. PLOS ONE 10(7):e0132561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Desta Z, Zhao X, Shin JG, Flockhart DA. 2002. Clinical significance of the cytochrome P450 2C19 genetic polymorphism. Clin. Pharmacokinet. 41(12):913–58 [DOI] [PubMed] [Google Scholar]
- 16.Luo HR, Poland RE, Lin KM, Wan YJ. 2006. Genetic polymorphism of cytochrome P450 2C19 in Mexican Americans: a cross-ethnic comparative study. Clin. Pharmacol. Ther. 80(1):33–40 [DOI] [PubMed] [Google Scholar]
- 17.Verma SS, Bergmeijer TO, Gong L, Reny JL, Lewis JP, et al. 2020. Genomewide association study of platelet reactivity and cardiovascular response in patients treated with clopidogrel: a study by the International Clopidogrel Pharmacogenomics Consortium. Clin. Pharmacol. Ther. 108(5):1067–77 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Lewis JP, Backman JD, Reny JL, Bergmeijer TO, Mitchell BD, et al. 2020. Pharmacogenomic polygenic response score predicts ischaemic events and cardiovascular mortality in clopidogrel-treated patients. Eur. Heart J. Cardiovasc. Pharmacother. 6(4):203–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Scott SA, Collet JP, Baber U, Yang Y, Peter I, et al. 2016. Exome sequencing of extreme clopidogrel response phenotypes identifies B4GALT2 as a determinant of on-treatment platelet reactivity. Clin. Pharmacol. Ther. 100(3):287–94 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lewis JP, Horenstein RB, Ryan K, O’Connell JR, Gibson Q, et al. 2013. The functional G143E variant of carboxylesterase 1 is associated with increased clopidogrel active metabolite levels and greater clopidogrel response. Pharmacogenet. Genom. 23(1):1–8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Staritz P, Kurz K, Stoll M, Giannitsis E, Katus HA, Ivandic BT. 2009. Platelet reactivity and clopidogrel resistance are associated with the H2 haplotype of the P2Y12-ADP receptor gene. Int. J. Cardiol. 133(3):341–45 [DOI] [PubMed] [Google Scholar]
- 22.Htun WW, Steinhubl SR. 2013. Ticagrelor: the first novel reversible P2Y12 inhibitor. Expert Opin. Pharmacother. 14(2):237–45 [DOI] [PubMed] [Google Scholar]
- 23.Teng R 2012. Pharmacokinetic, pharmacodynamic and pharmacogenetic profile of the oral antiplatelet agent ticagrelor. Clin. Pharmacokinet. 51(5):305–18 [DOI] [PubMed] [Google Scholar]
- 24.Dinicolantonio JJ, Serebruany VL. 2013. Exploring the ticagrelor-statin interplay in the PLATO trial. Cardiology 124(2):105–7 [DOI] [PubMed] [Google Scholar]
- 25.Varenhorst C, Eriksson N, Johansson Å, Barratt BJ, Hagström E, et al. 2015. Effect of genetic variations on ticagrelor plasma levels and clinical outcomes. Eur. Heart J. 36(29):1901–12 [DOI] [PubMed] [Google Scholar]
- 26.Pourdjabbar A, Hibbert B, Chong AY, Abunassar J, Malhotra N, et al. 2016. A pharmacodynamic analysis for the co-administration of inducers of CYP3A with ticagrelor: a cautionary tale in managing patients with acute coronary syndromes. Int. J. Cardiol. 214:423–25 [DOI] [PubMed] [Google Scholar]
- 27.Li M, Hu Y, Li H, Wen Z, Hu X, et al. 2017. No effect of SLCO1B1 and CYP3A4/5 polymorphisms on the pharmacokinetics and pharmacodynamics of ticagrelor in healthy Chinese male subjects. Biol. Pharm. Bull. 40(1):88–96 [DOI] [PubMed] [Google Scholar]
- 28.Brandt JT, Close SL, Iturria SJ, Payne CD, Farid NA, et al. 2007. Common polymorphisms of CYP2C19 and CYP2C9 affect the pharmacokinetic and pharmacodynamic response to clopidogrel but not prasugrel. J. Thromb. Haemost. 5(12):2429–36 [DOI] [PubMed] [Google Scholar]
- 29.Kelly RP, Close SL, Farid NA, Winters KJ, Shen L, et al. 2012. Pharmacokinetics and pharmacodynamics following maintenance doses of prasugrel and clopidogrel in Chinese carriers of CYP2C19 variants. Br. J. Clin. Pharmacol. 73(1):93–105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Varenhorst C, James S, Erlinge D, Brandt JT, Braun OO, et al. 2009. Genetic variation of CYP2C19 affects both pharmacokinetic and pharmacodynamic responses to clopidogrel but not prasugrel in aspirin-treated patients with coronary artery disease. Eur. Heart J. 30(14):1744–52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mega JL, Close SL, Wiviott SD, Shen L, Walker JR, et al. 2010. Genetic variants in ABCB1 and CYP2C19 and cardiovascular outcomes after treatment with clopidogrel and prasugrel in the TRITON-TIMI 38 trial: a pharmacogenetic analysis. Lancet 376(9749):1312–19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Würtz M, Kristensen SD, Hvas AM, Grove EL. 2012. Pharmacogenetics of the antiplatelet effect of aspirin. Curr. Pharm. Des. 18(33):5294–308 [DOI] [PubMed] [Google Scholar]
- 33.Yang Y, Lewis JP, Hulot JS, Scott SA. 2015. The pharmacogenetic control of antiplatelet response: candidate genes and CYP2C19. Expert Opin. Drug Metab. Toxicol. 11(10):1599–617 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Shuldiner AR, O’Connell JR, Bliden KP, Gandhi A, Ryan K, et al.2009.Association of cytochrome P450 2C19 genotype with the antiplatelet effect and clinical efficacy of clopidogrel therapy. JAMA 302(8):849–57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Luzum JA, Pakyz RE, Elsey AR, Haidar CE, Peterson JF, et al. 2017. The Pharmacogenomics Research Network Translational Pharmacogenetics Program: outcomes and metrics of pharmacogenetic implementations across diverse healthcare systems. Clin. Pharmacol. Ther. 102(3):502–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sistonen J, Fuselli S, Palo JU, Chauhan N, Padh H, Sajantila A. 2009. Pharmacogenetic variation at CYP2C9, CYP2C19, and CYP2D6 at global and microgeographic scales. Pharmacogenet. Genom. 19(2):170–79 [DOI] [PubMed] [Google Scholar]
- 37.Scott SA, Sangkuhl K, Stein CM, Hulot JS, Mega JL, et al. 2013. Clinical Pharmacogenetics Implementation Consortium guidelines for CYP2C19 genotype and clopidogrel therapy: 2013 update. Clin. Pharmacol. Ther. 94(3):317–23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mega JL, Close SL, Wiviott SD, Shen L, Hockett RD, et al. 2009. Cytochrome P-450 polymorphisms and response to clopidogrel. N. Engl. J. Med. 360(4):354–62 [DOI] [PubMed] [Google Scholar]
- 39.Mega JL, Simon T, Collet JP, Anderson JL, Antman EM, et al. 2010. Reduced-function CYP2C19 genotype and risk of adverse clinical outcomes among patients treated with clopidogrel predominantly for PCI: a meta-analysis. JAMA 304(16):1821–30 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Umemura K, Furuta T, Kondo K. 2008. The common gene variants of CYP2C19 affect pharmacokinetics and pharmacodynamics in an active metabolite of clopidogrel in healthy subjects. J. Thromb. Haemost. 6(8):1439–41 [DOI] [PubMed] [Google Scholar]
- 41.Ernest CS II, Small DS, Rohatagi S, Salazar DE, Wallentin L, et al. 2008. Population pharmacokinetics and pharmacodynamics of prasugrel and clopidogrel in aspirin-treated patients with stable coronary artery disease. J. Pharmacokinet. Pharmacodyn. 35(6):593–618 [DOI] [PubMed] [Google Scholar]
- 42.Gurbel PA, Tantry US. 2006. Drug insight: clopidogrel nonresponsiveness. Nat. Clin. Pract. Cardiovasc. Med. 3(7):387–95 [DOI] [PubMed] [Google Scholar]
- 43.Gurbel PA, Tantry US. 2007. Clopidogrel resistance? Thromb. Res. 120(3):311–21 [DOI] [PubMed] [Google Scholar]
- 44.Holmes MV, Perel P, Shah T, Hingorani AD, Casas JP. 2011. CYP2C19 genotype, clopidogrel metabolism, platelet function, and cardiovascular events: a systematic review and meta-analysis. JAMA 306(24):2704–14 [DOI] [PubMed] [Google Scholar]
- 45.Gurbel PA, Tantry US. 2012. Do platelet function testing and genotyping improve outcome in patients treated with antithrombotic agents? Platelet function testing and genotyping improve outcome in patients treated with antithrombotic agents. Circulation 125(10):1276–87 [DOI] [PubMed] [Google Scholar]
- 46.Price MJ, Murray SS, Angiolillo DJ, Lillie E, Smith EN, et al. 2012. Influence of genetic polymorphisms on the effect of high- and standard-dose clopidogrel after percutaneous coronary intervention: the GIFT (Genotype Information and Functional Testing) study. J. Am. Coll. Cardiol. 59(22):1928–37 [DOI] [PubMed] [Google Scholar]
- 47.Mega JL, Hochholzer W, Frelinger AL 3rd, Kluk MJ, Angiolillo DJ, et al. 2011. Dosing clopidogrel based on CYP2C19 genotype and the effect on platelet reactivity in patients with stable cardiovascular disease. JAMA 306(20):2221–28 [DOI] [PubMed] [Google Scholar]
- 48.Jeong YH, Bliden KP, Antonino MJ, Park KS, US Tantry, Gurbel PA. 2012. Usefulness of the VerifyNow P2Y12 assay to evaluate the antiplatelet effects of ticagrelor and clopidogrel therapies. Am. Heart J. 164(1):35–42 [DOI] [PubMed] [Google Scholar]
- 49.Hochholzer W, Trenk D, Fromm MF, Valina CM, Stratz C, et al. 2010. Impact of cytochrome P450 2C19 loss-of-function polymorphism and of major demographic characteristics on residual platelet function after loading and maintenance treatment with clopidogrel in patients undergoing elective coronary stent placement. J. Am. Coll. Cardiol. 55(22):2427–34 [DOI] [PubMed] [Google Scholar]
- 50.Geisler T, Schaeffeler E, Dippon J, Winter S, Buse V, et al. 2008. CYP2C19 and nongenetic factors predict poor responsiveness to clopidogrel loading dose after coronary stent implantation. Pharmacogenomics 9(9):1251–59 [DOI] [PubMed] [Google Scholar]
- 51.Liu T, Yin T, Li Y, Song LQ, Yu J, et al. 2014. CYP2C19 polymorphisms and coronary heart disease risk factors synergistically impact clopidogrel response variety after percutaneous coronary intervention. Coron. Artery Dis. 25(5):412–20 [DOI] [PubMed] [Google Scholar]
- 52.Muller C, Caillard S, Jesel L, El Ghannudi S, Ohlmann P, et al. 2012. Association of estimated GFR with platelet inhibition in patients treated with clopidogrel. Am. J. Kidney Dis. 59(6):777–85 [DOI] [PubMed] [Google Scholar]
- 53.Gremmel T, Müller M, Steiner S, Seidinger D, Koppensteiner R, et al. 2013. Chronic kidney disease is associated with increased platelet activation and poor response to antiplatelet therapy. Nephrol. Dial. Transplant. 28(8):2116–22 [DOI] [PubMed] [Google Scholar]
- 54.Faraday N, Becker DM, Becker LC. 2007. Pharmacogenomics of platelet responsiveness to aspirin. Pharmacogenomics 8(10):1413–25 [DOI] [PubMed] [Google Scholar]
- 55.Patrono C 1994. Aspirin as an antiplatelet drug. N. Engl. J. Med. 330(18):1287–94 [DOI] [PubMed] [Google Scholar]
- 56.Cuisset T, Loosveld M, Morange PE, Quilici J, Moro PJ, et al. 2012. CYP2C19*2 and *17 alleles have a significant impact on platelet response and bleeding risk in patients treated with prasugrel after acute coronary syndrome. JACC Cardiovasc. Interv. 5(12):1280–87 [DOI] [PubMed] [Google Scholar]
- 57.Grosdidier C, Quilici J, Loosveld M, Camoin L, Moro PJ, et al. 2013. Effect of CYP2C19*2 and *17 genetic variants on platelet response to clopidogrel and prasugrel maintenance dose and relation to bleeding complications. Am. J. Cardiol. 111(7):985–90 [DOI] [PubMed] [Google Scholar]
- 58.Storey RF, Thornton SM, Lawrance R, Husted S, Wickens M, et al. 2009. Ticagrelor yields consistent dose-dependent inhibition of ADP-induced platelet aggregation in patients with atherosclerotic disease regardless of genotypic variations in P2RY12, P2RY1, and ITGB3. Platelets 20(5):341–48 [DOI] [PubMed] [Google Scholar]
- 59.Collet J-P, Cuisset T, Rangé G, Cayla G, Elhadad S, et al. 2012. Bedside monitoring to adjust antiplatelet therapy for coronary stenting. N. Engl. J. Med. 367(22):2100–9 [DOI] [PubMed] [Google Scholar]
- 60.Price MJ, Berger PB, Teirstein PS, Tanguay JF, Angiolillo DJ, et al. 2011. Standard- versus high-dose clopidogrel based on platelet function testing after percutaneous coronary intervention: the GRAVITAS randomized trial. JAMA 305(11):1097–105 [DOI] [PubMed] [Google Scholar]
- 61.Roberts JD, Wells GA, Le May MR, Labinaz M, Glover C, et al. 2012. Point-of-care genetic testing for personalisation of antiplatelet treatment (RAPID GENE): a prospective, randomised, proof-of-concept trial. Lancet 379(9827):1705–11 [DOI] [PubMed] [Google Scholar]
- 62.Erlinge D, James S, Duvvuru S, Jakubowski JA, Wagner H, et al. 2014. Clopidogrel metaboliser status based on point-of-care CYP2C19 genetic testing in patients with coronary artery disease. Thromb. Haemost. 111(5):943–50 [DOI] [PubMed] [Google Scholar]
- 63.Morales-Rosado JA, Goel K, Zhang L, Åkerblom A, Baheti S, et al.2021.Next-generation sequencing of CYP2C19 in stent thrombosis: implications for clopidogrel pharmacogenomics. Cardiovasc. Drugs Ther. 35(3):549–59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Zhang L, Sarangi V, Moon I, Yu J, Liu D, et al. 2020. CYP2C9 and CYP2C19: deep mutational scanning and functional characterization of genomic missense variants. Clin. Transl. Sci. 13(4):727–42 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sorich MJ, Rowland A, McKinnon RA, Wiese MD. 2014. CYP2C19 genotype has a greater effect on adverse cardiovascular outcomes following percutaneous coronary intervention and in Asian populations treated with clopidogrel: a meta-analysis. Circ. Cardiovasc. Genet. 7(6):895–902 [DOI] [PubMed] [Google Scholar]
- 66.Niu X, Mao L, Huang Y, Baral S, Li JY, et al. 2015. CYP2C19 polymorphism and clinical outcomes among patients of different races treated with clopidogrel: a systematic review and meta-analysis. J. Huazhong. Univ. Sci. Technol. Med. Sci. 35(2):147–56 [DOI] [PubMed] [Google Scholar]
- 67.Xi Z, Fang F, Wang J, AlHelal J, Zhou Y, Liu W. 2019. CYP2C19 genotype and adverse cardiovascular outcomes after stent implantation in clopidogrel-treated Asian populations: a systematic review and meta-analysis. Platelets 30(2):229–40 [DOI] [PubMed] [Google Scholar]
- 68.Lee CR, Thomas CD, Beitelshees AL, Tuteja S, Empey PE, et al. 2021. Impact of the CYP2C19*17 allele on outcomes in patients receiving genotype-guided antiplatelet therapy after percutaneous coronary intervention. Clin. Pharmacol. Ther. 109(3):705–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Cavallari LH, Beitelshees AL, Blake KV, Dressler LG, Duarte JD, et al. 2017.The IGNITE Pharmacogenetics Working Group: an opportunity for building evidence with pharmacogenetic implementation in a real-world setting. Clin. Transl. Sci. 10(3):143–46 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Claassens DMF, Bergmeijer TO, Vos GJA, Hermanides RS, van ‘t Hof AWJ, et al. 2021. Clopidogrel versus ticagrelor or prasugrel after primary percutaneous coronary intervention according to CYP2C19 genotype: a POPular genetics subanalysis. Circ. Cardiovasc. Interv. 14(4):e009434. [DOI] [PubMed] [Google Scholar]
- 71.Bergmeijer TO, Reny JL, Pakyz RE, Gong L, Lewis JP, et al. 2018. Genome-wide and candidate gene approaches of clopidogrel efficacy using pharmacodynamic and clinical end points—rationale and design of the International Clopidogrel Pharmacogenomics Consortium (ICPC). Am. Heart J. 198:152–59 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Shuldiner AR, O’Connell JR, Bliden KP, Gandhi A, Ryan K, et al. 2009. Association of cytochrome P450 2C19 genotype with the antiplatelet effect and clinical efficacy of clopidogrel therapy. JAMA 302(8):849–57 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Linskey DW, Linskey DC, McLeod HL, Luzum JA. 2021. The need to shift pharmacogenetic research from candidate gene to genome-wide association studies. Pharmacogenomics 22(17):1143–50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Pan Y, Chen W, Xu Y, Yi X, Han Y, et al. 2017. Genetic polymorphisms and clopidogrel efficacy for acute ischemic stroke or transient ischemic attack: a systematic review and meta-analysis. Circulation 135(1):21–33 [DOI] [PubMed] [Google Scholar]
- 75.Tornio A, Flynn R, Morant S, Velten E, Palmer CNA, et al. 2018. Investigating real-world clopidogrel pharmacogenetics in stroke using a bioresource linked to electronic medical records. Clin. Pharmacol. Ther. 103(2):281–86 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Wang Y, Zhao X, Lin J, Li H, Johnston SC, et al. 2016. Association between CYP2C19 loss-of-function allele status and efficacy of clopidogrel for risk reduction among patients with minor stroke or transient ischemic attack. JAMA 316(1):70–78 [DOI] [PubMed] [Google Scholar]
- 77.Lee CR, Luzum JA, Sangkuhl K, Gammal RS, Sabatine MS, et al. 2022. Clinical Pharmacogenetics Implementation Consortium guideline for CYP2C19 genotype and clopidogrel therapy: 2022 update. Clin. Pharmacol. Ther. In press [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.FDA (US Food Drug Admin.). 2021. Clopidogrel prescribing information 2021. FDA, Silver Spring, MD. https://www.accessdata.fda.gov/drugsatfda_docs/label/2021/020839s074lbl.pdf [Google Scholar]
- 79.Alhazzani A, Venkatachalapathy P, Padhilahouse S, Sellappan M, Munisamy M, et al. 2021. Biomarkers for antiplatelet therapies in acute ischemic stroke: a clinical review. Front. Neurol. 12:667234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Gower MN, Ratner LR, Williams AK, Rossi JS, Stouffer GA, Lee CR. 2020. Clinical utility of CYP2C19 genotype-guided antiplatelet therapy in patients at risk of adverse cardiovascular and cerebrovascular events: a review of emerging evidence. Pharmacogenom. Pers. Med. 13:239–52 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Li C, Jia W, Li J, Li F, Ma J, Zhou L. 2021. Association with CYP2C19 polymorphisms and clopidogrel in treatment of elderly stroke patients. BMC Neurol. 21(1):104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Lv H, Yang Z, Wu H, Liu M, Mao X, et al. 2021. High on-treatment platelet reactivity as predictor of long-term clinical outcomes in stroke patients with antiplatelet agents. Transl. Stroke Res. 13(3):391–98 [DOI] [PubMed] [Google Scholar]
- 83.Al-Rubaish AM, Al-Muhanna FA, Alshehri AM, Alsulaiman AA, Alabdulali MM, et al. 2021. Prevalence of CYP2C19*2 carriers in Saudi ischemic stroke patients and the suitability of using genotyping to guide antiplatelet therapy in a university hospital setup. Drug. Metab. Pers. Ther In press [DOI] [PubMed] [Google Scholar]
- 84.Wang J, Liu J, Zhou Y, Wang F, Xu K, et al. 2019. Association among PlA1/A2 gene polymorphism, laboratory aspirin resistance and clinical outcomes in patients with coronary artery disease: an updated meta-analysis. Sci. Rep. 9(1):13177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Ferreira M, Freitas-Silva M, Assis J, Pinto R, Nunes JP, Medeiros R. 2020. The emergent phenomenon of aspirin resistance: insights from genetic association studies. Pharmacogenomics 21(2):125–40 [DOI] [PubMed] [Google Scholar]
- 86.Ross S, Paré G. 2018. Pharmacogenetics of stroke. Stroke 49(10):2541–48 [DOI] [PubMed] [Google Scholar]
- 87.Herrera-Galeano JE, Becker DM, Wilson AF, Yanek LR, Bray P, et al. 2008. A novel variant in the platelet endothelial aggregation receptor-1 gene is associated with increased platelet aggregability. Arterioscler. Thromb. Vasc. Biol. 28(8):1484–90 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Lewis JP, Riaz M, Xie S, Polekhina G, Wolfe R, et al. 2020. Genetic variation in PEAR1, cardiovascular outcomes and effects of aspirin in a healthy elderly population. Clin. Pharmacol. Ther. 108(6):1289–98 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Li Z, Jiang H, Ding Y, Zhang D, Zhang X, et al. 2021. Platelet endothelial aggregation receptor 1 polymorphism is associated with functional outcome in small-artery occlusion stroke patients treated with aspirin. Front. Cardiovasc. Med. 8:664012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Bourgeois S, Carr DF, Musumba CO, Penrose A, Esume C, et al. 2021. Genome-wide association between EYA1 and aspirin-induced peptic ulceration. EBioMedicine 74:103728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Mallah N, Zapata-Cachafeiro M, Aguirre C, Ibarra-García E, Palacios-Zabalza I, et al. 2021. A multicenter case-control study of the effect of e-nos VNTR polymorphism on upper gastrointestinal hemorrhage in NSAID users. Sci. Rep. 11(1):19923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Mallah N, Zapata-Cachafeiro M, Aguirre C, Ibarra-García E, Palacios-Zabalza I, et al. 2022. Synergism interaction between genetic polymorphisms in drug metabolizing enzymes and NSAIDs on upper gastrointestinal haemorrhage: a multicenter case-control study. Ann. Med. 54(1):379–92 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Forgerini M, Urbano G, de Nadai TR, Batah SS, Fabro AT, Mastroianni PC. 2021. Genetic variants in PTGS1 and NOS3 genes increase the risk of upper gastrointestinal bleeding: a case-control study. Front. Pharmacol. 12:671835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Handa Y, Fukushima S, Yo S, Osawa M, Murao T, et al. 2021. A novel gene associated with small bowel bleeding in patients taking low-dose aspirin. Dig. Liver Dis. 53(7):841–45 [DOI] [PubMed] [Google Scholar]
- 95.Wang Y, Chen W, Lin Y, Meng X, Chen G, et al. 2019. Ticagrelor plus aspirin versus clopidogrel plus aspirin for platelet reactivity in patients with minor stroke or transient ischaemic attack: open label, blinded endpoint, randomised controlled phase II trial. BMJ 365:l2211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Verdoia M, Tonon F, Gioscia R, Nardin M, Fierro N, et al. 2020. Impact of the rs73598374 polymorphism of the adenosine deaminase gene on platelet reactivity and long-term outcomes among patients with acute coronary syndrome treated with ticagrelor. Thromb. Res. 196:231–37 [DOI] [PubMed] [Google Scholar]
- 97.Yuan D, Shi X, Gao L, Wan G, Zhang H, et al. 2022. Identification of potential biological factors affecting the treatment of ticagrelor after percutaneous coronary intervention in the Chinese population. Pharmacogenom. Pers. Med. 15:29–43 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Wallentin L, James S, Storey RF, Armstrong M, Barratt BJ, et al. 2010. Effect of CYP2C19 and ABCB1 single nucleotide polymorphisms on outcomes of treatment with ticagrelor versus clopidogrel for acute coronary syndromes: a genetic substudy of the PLATO trial. Lancet 376(9749):1320–28 [DOI] [PubMed] [Google Scholar]
- 99.Danielak D, Karaźniewicz-Łada M, Główka F. 2018. Ticagrelor in modern cardiology—an up-to-date review of most important aspects of ticagrelor pharmacotherapy. Expert Opin. Pharmacother. 19(2):103–12 [DOI] [PubMed] [Google Scholar]
- 100.Wallentin L, Becker RC, Budaj A, Cannon CP, Emanuelsson H, Held C, et al. 2009. Ticagrelor versus clopidogrel in patients with acute coronary syndromes. N. Engl. J. Med. 361(11):1045–57 [DOI] [PubMed] [Google Scholar]
- 101.Mega JL, Close SL, Wiviott SD, Man M, Duvvuru S, et al. 2016. PON1 Q192R genetic variant and response to clopidogrel and prasugrel: pharmacokinetics, pharmacodynamics, and a meta-analysis of clinical outcomes. J. Thromb. Thrombolys. 41(3):374–83 [DOI] [PubMed] [Google Scholar]
- 102.Mega JL, Close SL, Wiviott SD, Shen L, Hockett RD, et al. 2009. Cytochrome P450 genetic polymorphisms and the response to prasugrel: relationship to pharmacokinetic, pharmacodynamic, and clinical outcomes. Circulation 119(19):2553–60 [DOI] [PubMed] [Google Scholar]
- 103.Sawayama Y, Yamamoto T, Tomita Y, Asada K, Yagi N, et al. 2020. Comparison between clopidogrel and prasugrel associated with CYP2C19 genotypes in patients receiving percutaneous coronary intervention in a Japanese population. Circ. J. 84(9):1575–81 [DOI] [PubMed] [Google Scholar]
- 104.Cavallari LH, Lee CR, Beitelshees AL, Cooper-DeHoff RM, Duarte JD, et al. 2018. Multisite investigation of outcomes with implementation of CYP2C19 genotype-guided antiplatelet therapy after percutaneous coronary intervention. JACC Cardiovasc. Interv. 11(2):181–91 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Galli M, Franchi F, Rollini F, Angiolillo DJ. 2021. Role of platelet function and genetic testing in patients undergoing percutaneous coronary intervention. Trends Cardiovasc. Med In press [DOI] [PubMed] [Google Scholar]
- 106.Xie X, Ma YT, Yang YN, Li XM, Zheng YY, et al. 2013. Personalized antiplatelet therapy according to CYP2C19 genotype after percutaneous coronary intervention: a randomized control trial. Int. J. Cardiol. 168(4):3736–40 [DOI] [PubMed] [Google Scholar]
- 107.Claassens DMF, Vos GJA, Bergmeijer TO, Hermanides RS, van ‘t Hof AWJ, et al. 2019. A genotype-guided strategy for oral P2Y12 inhibitors in primary PCI. N. Engl. J. Med. 381(17):1621–31 [DOI] [PubMed] [Google Scholar]
- 108.Pereira NL, Farkouh ME, So D, Lennon R, Geller N, et al. 2020. Effect of genotype-guided oral P2Y12 inhibitor selection versus conventional clopidogrel therapy on ischemic outcomes after percutaneous coronary intervention: the TAILOR-PCI randomized clinical trial. JAMA 324(8):761–71 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Parcha V, Heindl BF, Li P, Kalra R, Limdi NA, et al. 2021. Genotype-guided P2Y12 inhibitor therapy after percutaneous coronary intervention: a Bayesian analysis. Circ. Genom. Precis Med. 14(6):e003353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Pereira NL, Rihal C, Lennon R, Marcus G, Shrivastava S, et al. 2021. Effect of CYP2C19 genotype on ischemic outcomes during oral P2Y12 inhibitor therapy: a meta-analysis. JACC Cardiovasc. Interv. 14(7):739–50 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Galli M, Benenati S, Capodanno D, Franchi F, Rollini F, et al. 2021. Guided versus standard antiplatelet therapy in patients undergoing percutaneous coronary intervention: a systematic review and meta-analysis. Lancet 397(10283):1470–83 [DOI] [PubMed] [Google Scholar]
- 112.Wang Y, Meng X, Wang A, Xie X, Pan Y, et al. 2021. Ticagrelor versus clopidogrel in CYP2C19 loss-of-function carriers with stroke or TIA. N. Engl. J. Med. 385(27):2520–30 [DOI] [PubMed] [Google Scholar]
- 113.Kazi DS, Garber AM, Shah RU, Dudley RA, Mell MW, et al. 2014. Cost-effectiveness of genotype-guided and dual antiplatelet therapies in acute coronary syndrome. Ann. Intern. Med. 160(4):221–32 [DOI] [PubMed] [Google Scholar]
- 114.Sorich MJ, Horowitz JD, Sorich W, Wiese MD, Pekarsky B, Karnon JD. 2013. Cost-effectiveness of using CYP2C19 genotype to guide selection of clopidogrel or ticagrelor in Australia. Pharmacogenomics 14(16):2013–21 [DOI] [PubMed] [Google Scholar]
- 115.Lala A, Berger JS, Sharma G, Hochman JS, Braithwaite RS, Ladapo JA.2013.Genetic testing in patients with acute coronary syndrome undergoing percutaneous coronary intervention: a cost-effectiveness analysis. J. Thromb. Haemost. 11(1):81–91 [DOI] [PubMed] [Google Scholar]
- 116.Deiman BA, Tonino PA, Kouhestani K, Schrover CE, Scharnhorst V, et al. 2016. Reduced number of cardiovascular events and increased cost-effectiveness by genotype-guided antiplatelet therapy in patients undergoing percutaneous coronary interventions in the Netherlands. Neth. Heart J. 24(10):589–99 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Weitzel KW, Elsey AR, Langaee TY, Burkley B, Nessl DR, et al. 2014. Clinical pharmacogenetics implementation: approaches, successes, and challenges. Am. J. Med. Genet. C Semin. Med. Genet. 166c(1):56–67 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Empey PE, Pratt VM, Hoffman JM, Caudle KE, Klein TE. 2021. Expanding evidence leads to new pharmacogenomics payer coverage. Genet. Med. 23(5):830–32 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Scott SA. 2011. Personalizing medicine with clinical pharmacogenetics. Genet. Med. 13(12):987–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Abul-Husn NS, Owusu Obeng A, Sanderson SC, Gottesman O, Scott SA. 2014. Implementation and utilization of genetic testing in personalized medicine. Pharmacogenom. Pers. Med. 7:227–40 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Pereira NL, Stewart AK. 2015. Clinical implementation of cardiovascular pharmacogenomics. Mayo Clin. Proc. 90(6):701–4 [DOI] [PubMed] [Google Scholar]
- 122.Luzum JA, Luzum MJ. 2016. Physicians’ attitudes toward pharmacogenetic testing before and after pharmacogenetic education. Pers. Med. 13(2):119–27 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Nickola TJ, Green JS, Harralson AF, O’Brien TJ. 2012. The current and future state of pharmacogenomics medical education in the USA. Pharmacogenomics 13(12):1419–25 [DOI] [PubMed] [Google Scholar]
- 124.Pereira NL, So D, Bae JH, Chavez I, Jeong MH, et al. 2019. International survey of patients undergoing percutaneous coronary intervention and their attitudes toward pharmacogenetic testing. Pharmacogenet. Genom. 29(4):76–83 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125.Wu AH, White MJ, Oh S, Burchard E. 2015. The Hawaii clopidogrel lawsuit: the possible effect on clinical laboratory testing. Pers. Med. 12(3):179–81 [DOI] [PubMed] [Google Scholar]
- 126.Dep. Atty. Gen. 2021. $834 Million order entered in Hawai’i state court against Bristol-Myers Squibb and Sanofi for failing to investigate and disclose ineffectiveness of Plavix®. News Release, Febr. 15. https://ag.hawaii.gov/wp-content/uploads/2021/02/News-Release-2021-13.pdf [Google Scholar]
- 127.Haga SB, Moaddeb J. 2014. Comparison of delivery strategies for pharmacogenetic testing services. Pharmacogenet. Genom. 24(3):139–45 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128.Gottesman O, Scott SA, Ellis SB, Overby CL, Ludtke A, et al. 2013. The CLIPMERGE PGx program: clinical implementation of personalized medicine through electronic health records and genomics-pharmacogenomics. Clin. Pharmacol. Ther. 94(2):214–17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129.Pulley JM, Denny JC, Peterson JF, Bernard GR, Vnencak-Jones CL, et al. 2012. Operational implementation of prospective genotyping for personalized medicine: the design of the Vanderbilt PREDICT project. Clin. Pharmacol. Ther. 92(1):87–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130.Rasmussen-Torvik LJ, Stallings SC, Gordon AS, Almoguera B, Basford MA, et al. 2014. Design and anticipated outcomes of the eMERGE-PGx project: a multicenter pilot for preemptive pharmacogenomics in electronic health record systems. Clin. Pharmacol. Ther. 96(4):482–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131.AACC (Am. Assoc. Clin. Chem.). 2021. FDA OKs 23andMe to report pharmacogenomic variants without confirmatory testing. AACC, July 1. https://www.aacc.org/cln/articles/2021/july/fda-oks-23andme-to-report-pharmacogenomic-variants-without-confirmatory-testing [Google Scholar]
- 132.Capodanno D, Angiolillo DJ, Lennon RJ, Goodman SG, Kim SW, et al. 2022. ABCD-GENE score and clinical outcomes following percutaneous coronary intervention: insights from the TAILOR-PCI trial. J. Am. Heart Assoc. 11(4):e024156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133.Grant C, Raina A, Lennon R, Goodman SG, Gulati R, et al. 2021. Baseline clinical and genetic data based machine learning predictions of 1-year ischemic outcomes following percutaneous coronary intervention. Circulation 144(Suppl. 1):A10602 [Google Scholar]
- 134.Whirl-Carrillo M, Huddart R, Gong L, Sangkuhl K, Thorn CF, et al. 2021. An evidence-based framework for evaluating pharmacogenomics knowledge for personalized medicine. Clin. Pharmacol. Ther. 110(3):563–72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 135.Collet JP, Thiele H, Barbato E, Barthélémy O, Bauersachs J, et al. 2021. 2020 ESC guidelines for the management of acute coronary syndromes in patients presenting without persistent ST-segment elevation. Eur. Heart J. 42(14):1289–367 [DOI] [PubMed] [Google Scholar]


