Abstract
Glutamine synthetase (GS) and superoxide dismutase (SOD), large multimeric enzymes that are thought to play important roles in the pathogenicity of Mycobacterium tuberculosis, are among the bacterium's major culture filtrate proteins in actively growing cultures. Although these proteins lack a leader peptide, their presence in the extracellular medium during early stages of growth suggested that they might be actively secreted. To understand their mechanism of export, we cloned the homologous genes (glnA1 and sodA) from the rapid-growing, nonpathogenic Mycobacterium smegmatis, generated glnA1 and sodA mutants of M. smegmatis by allelic exchange, and quantitated expression and export of both mycobacterial and nonmycobacterial GSs and SODs in these mutants. We also quantitated expression and export of homologous and heterologous SODs from M. tuberculosis. When each of the genes was expressed from a multicopy plasmid, M. smegmatis exported comparable proportions of both the M. tuberculosis and M. smegmatis GSs (in the glnA1 strain) or SODs (in the sodA strain), in contrast to previous observations in wild-type strains. Surprisingly, recombinant M. smegmatis and M. tuberculosis strains even exported nonmycobacterial SODs. To determine the extent to which export of these large, leaderless proteins is expression dependent, we constructed a recombinant M. tuberculosis strain expressing green fluorescent protein (GFP) at high levels and a recombinant M. smegmatis strain coexpressing the M. smegmatis GS, M. smegmatis SOD, and M. tuberculosis BfrB (bacterioferritin) at high levels. The recombinant M. tuberculosis strain exported GFP even in early stages of growth and at proportions very similar to those of the endogenous M. tuberculosis GS and SOD. Similarly, the recombinant M. smegmatis strain exported bacterioferritin, a large (∼500-kDa), leaderless, multimeric protein, in proportions comparable to GS and SOD. In contrast, high-level expression of the large, leaderless, multimeric protein malate dehydrogenase did not lead to extracellular accumulation because the protein was highly unstable extracellularly. These findings indicate that, contrary to expectations, export of M. tuberculosis GS and SOD in actively growing cultures is not due to a protein-specific export mechanism, but rather to bacterial leakage or autolysis, and that the extracellular abundance of these enzymes is simply due to their high level of expression and extracellular stability. The same determinants likely explain the presence of other leaderless proteins in the extracellular medium of actively growing M. tuberculosis cultures.
Mycobacterium tuberculosis, the primary etiologic agent of tuberculosis, is one of the world's leading causes of death, killing 2 million persons annually worldwide (23). New modalities to combat M. tuberculosis and a greater understanding of the biology and immunology of this pathogen are high priorities of tuberculosis research.
The extracellular proteins of M. tuberculosis have been the focus of many studies investigating their role in host immunity, their potential as vaccine candidates and diagnostic reagents, and more recently as drug targets (4, 5, 7, 31, 32, 46, 50). Although sensitive techniques have identified hundreds of proteins in culture filtrates of M. tuberculosis (36, 56, 62), approximately 12 proteins are released in relatively large amounts (accounting for ∼90% of total extracellular protein with a molecular mass of ≥10 kDa) and have been designated as the major extracellular proteins (reference 32 and unpublished data). Although most of the major extracellular proteins have typical leader peptides, the multimeric enzymes glutamine synthetase (GS) and superoxide dismutase (SOD) do not (29, 30, 71). GS and SOD are found in the extracellular medium of M. tuberculosis cultures in the early stages of growth, suggesting that they might be actively secreted by the bacterium. Both of these enzymes are considered to be strictly intracellular in most other bacteria.
GS is a dodecamer of identical 53-kDa subunits that has a central role in nitrogen metabolism, catalyzing the synthesis of l-glutamine from l-glutamate, ammonia, and ATP. Our group has identified GS as a major component of M. tuberculosis culture filtrates (28). We have proposed that GS may alter the ammonia level (and pH) of the host cell phagosome, possibly aiding the bacilli in preventing phagosome-lysosome fusion. We have also proposed that GS may be necessary for synthesis of poly-l-glutamate-glutamine present in the cell wall of pathogenic, but not nonpathogenic, mycobacteria.
SOD is ubiquitous in aerobes and is part of the mechanism to protect the cell from oxidative stress, catalyzing the dismutation of superoxide to oxygen and hydrogen peroxide (27). In bacteria, there are two families of SODs: the FeSOD/MnSOD family, whose members do not contain leader peptides, and the Cu,ZnSOD family, whose members do contain leader peptides (11). The two enzyme families do not share sequence similarities and are likely a result of convergent evolution. M. tuberculosis contains both a tetrameric FeSOD (SodA) and a less well characterized Cu,ZnSOD (SodC) (22, 39, 74, 76). Kusunose et al. first observed that M. tuberculosis exports large amounts of its FeSOD (SodA) (39). The Cu,ZnSOD is surface associated; however, SOD activity detectable in culture filtrates is mainly due to the presence of SodA. In several gram-negative pathogens, the periplasmic Cu,ZnSOD (SodC) has been associated with virulence (18, 25, 58, 72). However, a recent report by Dussurget et al. has shown that a SodC-deficient M. tuberculosis mutant is not attenuated in a guinea pig model (22). In humans, SOD may be important to M. tuberculosis survival in the phagocyte, where the bacterium may encounter host-generated superoxide and other reactive oxygen species.
Previous studies from this laboratory of recombinant M. tuberculosis GS and SOD in M. smegmatis found that a large percentage of the recombinant enzymes (>95% GS and 66% SOD) but a smaller percentage of the Mycobacterium smegmatis endogenous enzymes (1% GS and 21% SOD) were exported, suggesting that export of the two M. tuberculosis enzymes relied on information in the protein sequence and/or structure (29, 30). In this study we now provide evidence that strongly suggests that GS and SOD are not actively secreted but rather are released into the extracellular medium as a result of bacterial leakage or autolysis and that their extracellular abundance is simply dependent upon their high expression and extracellular stability.
MATERIALS AND METHODS
Materials.
Phenol, phenol-CHCl3-isoamyl alcohol (25:24:1), Superdex-75, and Sephacryl 300HR were purchased from Amersham Pharmacia Biotech. Reactive Red-120 Sepharose, xanthine oxidase, 2,3-bis[2-methoxy-4-nitro-5-sulfophenyl]-2H-tetrazolium-5-carboxanilide (XTT), and gamma-glutamic acid hydroxamate were purchased from Sigma. RNase (bovine pancreas) and proteinase K were purchased from Boehringer Mannheim.
Bacterial strains and plasmids.
The bacterial strains and plasmids used in this study are listed in Tables 1 and 2. M. smegmatis strains were grown on Middlebrook 7H11 agar (Difco) containing 10% (vol/vol) OADC (Becton Dickinson) and 0.5% (vol/vol) glycerol at 37°C or as shaken cultures in Middlebrook 7H9 broth (Difco) supplemented only with 0.2% (vol/vol) glycerol at 28 or 37°C. M. tuberculosis strains were grown on 7H11 agar containing 10% (vol/vol) OADC and 0.5% (vol/vol) glycerol or as unshaken cultures in 7H9 broth supplemented only with 0.2% (vol/vol) glycerol at 37°C in an atmosphere of 5% CO2–95% air. Hygromycin (50 μg ml−1) and/or kanamycin (20 μg ml−1) were included as appropriate. For the cultures shown in Fig. 12, glucose and/or additional (NH4)2SO4 was added to our standard 7H9 medium, which contains 0.2% (vol/vol) glycerol and 3.8 mM (NH4)2SO4. Other supplements are described in the text.
TABLE 1.
Bacterial strains and plasmids used in this study
| Strain or plasmid | Description | Promotera | Reference or source |
|---|---|---|---|
| Strains | |||
| E. coli DH5α | Gibco BRL | ||
| M. smegmatis 1-2c | Wild-type strain with increased electroporation efficiency | 76 | |
| M. smegmatis 1-2c glnA1 | Insertionally inactivated glnA1 locus, Kmr | This study | |
| M. smegmatis 1-2c sodA | Insertionally inactivated sodA locus, Kmr | This study | |
| M. tuberculosis Erdman | Wild-type strain | ATCC 35801 | |
| Plasmids | |||
| pUC18 | E. coli cloning vector | 48 | |
| pUC19 | E. coli cloning vector | 75 | |
| pUC18-Ms-glnA1 | M. smegmatis glnA1 genomic locus | This study | |
| pUC18-Ms-sodA | M. smegmatis sodA genomic locus | This study | |
| pPR27 | Mycobacterial allelic exchange vector, t.s. mycobacterial origin, sacB, Gmr | 51 | |
| pPR27-Ms-glnA1::Kmr | glnA1 allelic exchange construct | This study | |
| pPR27-Ms-sodA::Kmr | sodA allelic exchange construct | This study | |
| pUC19-Kmr | Kmr cassette | This study | |
| pNBV1 | E. coli-mycobacterial shuttle vector, Hygr | 33 | |
| pNBV1-MsGS | M. smegmatis glnA1 | Native | This study |
| pNBV1-MsSODA | M. smegmatis sodA | Native | This study |
| pNBV1-MtbGS | M. tuberculosis glnA1 | Native | This study |
| pNBV1-MtbSODA | M. tuberculosis sodA | Native | 29 |
| pNBV1-EcSODA | E. coli sodA | Mtb glnA1 | This study |
| pNBV1-EcSODB | E. coli sodB | Mtb glnA1 | This study |
| pNBV1-BsSODA | B. subtilis sodA | Mtb glnA1 | This study |
| pNBV1-StGS | S. enterica serovar Typhimurium glnA | Mtb glnA1 | This study |
| pNBV1-GFPuv | UV-optimized GFP | BCG hsp60 | This study |
| pNBV1-MDH | M. tuberculosis mdh | Mtb glnA1 | This study |
| pNBV1-BFRB | M. tuberculosis bfrB | Mtb glnA1 | This study |
| pNBV1-MsGS-MsSODA-MDH | M. smegmatis glnA1, M. smegmatis sodA, and M. tuberculosis mdh | Native, native, Mtb glnA1 | This study |
| pNBV1-MsSODA-MsGS-BFRB | M. smegmatis glnA1, M. smegmatis sodA, and M. tuberculosis bfrB | Native, native, Mtb glnA1 | This study |
| pSMT3 | Source of M. bovis BCG hsp60 promoter | 49 | |
| pGFPuv | Source of UV-optimized Aequorea victoria GFP gene | Clontech | |
| pCR2.1 | TA cloning vector; template DNA for Tn5 kanamycin resistance gene (aphA-2) | Invitrogen |
For expression constructs, the gene from which the promoter is derived is listed. The term “native” indicates that transcription is driven from the gene's own native promoter.
TABLE 2.
Recombinant mycobacterial strains used in this study
| Strain and plasmid |
|---|
| M. smegmatis 1-2c |
| pNBV1 |
| pNBV1-EcSODA |
| pNBV1-EcSODB |
| pNBV1-BsSODA |
| M. smegmatis 1-2c glnA1 |
| pNBV1 |
| pNBV1-MsGS |
| pNBV1-MtbGS |
| pNBV1-StGS |
| pNBV1-MsGS-MsSODA-MDH |
| pNBV1-MsSODA-MsGS-BFRB |
| M. smegmatis 1-2c sodA |
| pNBV1 |
| pNBV1-MsSODA |
| pNBV1-MtbSODA |
| pNBV1-EcSODA |
| pNBV1-EcSODB |
| pNBV1-BsSODA |
| M. tuberculosis Erdman |
| pNBV1 |
| pNBV1-MsSODA |
| pNBV1-EcSODB |
| pNBV1-GFPuv |
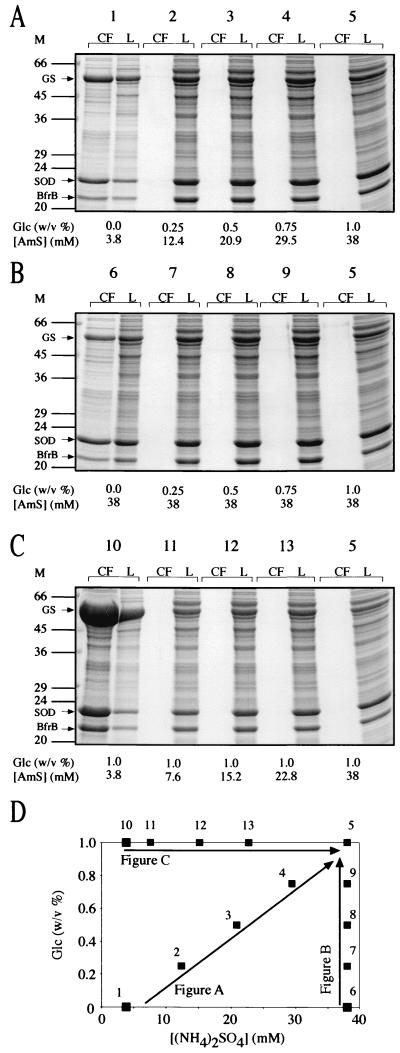
FIG. 12.
Expression and export of GS, SOD, and BfrB by M. smegmatis glnA1 pNBV1-MsSODA-MsGS-BFRB under various growth conditions. (A to C) M. smegmatis glnA1 pNBV1-MsGS-MsSODA-BFRB was grown for 4 days (cultures reached stationary phase by 2 to 3 days) in 7H9 medium containing 0.2% (vol/vol) glycerol with various amounts of glucose and/or additional (NH4)2SO4, and culture filtrates (CF) and lysates (L) were analyzed by SDS-PAGE. Tenfold more CF than L was loaded on the gels (the equivalent of 2 ml versus 0.2 ml of original culture volume). Arrows indicate the positions of the M. smegmatis GS, M. smegmatis SOD, and the M. tuberculosis BfrB (BfrB). Two to four independent cultures were grown and analyzed for each of the four extreme culture conditions (1, 5, 6, and 10) as well as for culture condition 2, and the results shown are representative. (D) Graph indicating the glucose and (NH4)2SO4 concentrations of the media that were tested in panels A to C. The standard 7H9 medium (culture 1) contains 82 mM carbon (as 0.2% [vol/vol] glycerol) and 11 mM nitrogen [as 3.8 mM (NH4)2SO4 and 3.4 mM l-glutamate]. The (NH4)2SO4 (AmS) concentration is stated as the total amount present in the medium for each culture condition. M, molecular mass markers in kilodaltons.
Escherichia coli DH5α, Salmonella enterica serovar Typhimurium (ATCC 13311), and Bacillus subtilis SB168 (ATCC 27689) were grown on Luria-Bertani agar, Luria-Bertani broth, or Terrific broth at 37°C. Ampicillin (100 μg ml−1), gentamicin (20 μg ml−1), hygromycin (250 μg ml−1), and kanamycin (50 μg ml−1) were included as appropriate.
Recombinant DNA methods.
Plasmid DNA was isolated using Quantum Prep (Bio-Rad) or Wizard Plus SV (Promega) miniprep kits. Genomic DNA was isolated from M. smegmatis, M. tuberculosis, E. coli, S. enterica serovar Typhimurium, and B. subtilis by extraction with hot phenol (65°C, Tris-equilibrated, pH ≥ 8) and ethanol precipitation. The DNA was further purified by sequential digestion with RNase and proteinase K, additional extractions with phenol and phenol-CHCl3-isoamyl alcohol (25:24:1), and a final ethanol precipitation.
Electroporation of mycobacteria.
Cells were washed two or three times with 10% (vol/vol) glycerol and resuspended in 10% (vol/vol) glycerol to a density of 109 to 1010 cells ml−1. M. smegmatis cells were kept at 0 to 4°C throughout the washing and electroporation procedures while M. tuberculosis cells were maintained at room temperature (68). DNA (1 to 5 μg) was mixed with 400 μl of cells, transferred to a 0.2-cm-gap electroporation cuvette (Bio-Rad), and a single electrical pulse was delivered (2.5 kV, 25 μF, 1,000 Ω). For M. smegmatis, the cells were immediately transferred to 2 ml of SOC medium (57) (with 5 mM l-glutamine for the glnA1 mutant) and incubated at 37°C with vigorous shaking for 4 to 6 h before being plated on selective media. For cells electroporated with temperature-sensitive pPR27-derived plasmids, the incubation was done at ≤32°C. For M. tuberculosis, 600 μl of SOC medium was added directly to the cells in the cuvette and the cells were incubated without shaking at 37°C for 18 to 24 h before being plated on selective media.
Southern and colony hybridizations.
Restriction fragments of genomic DNA or plasmids were electrophoresed in agarose gels, transferred to positively charged nylon membranes (Hybond-N+; Amersham Pharmacia Biotech) in 0.4 M NaOH, and hybridized to specific probes. Hybridizations were done at 60°C in 6× SSPE–2% skim milk for 15 to 20 h (1× SSPE is 150 mM NaCl, 10 mM NaH2PO4, 1 mM EDTA [pH 7.4]). Membranes were washed very stringently at 65°C for 15 to 20 min once with 1× SSPE–1% skim milk and twice with 0.1× SSPE–1% skim milk before being exposed to X-ray film for various times. Hybridization and washing conditions were identical for colony hybridizations. Probes were radiolabeled to a specific activity of 1 × 108 to 2 × 108 dpm μg−1 with [α-32P]dCTP by random priming using the Multiprime DNA Labeling Kit (Amersham Pharmacia Biotech).
Cloning and sequencing of the M. smegmatis glnA1 and sodA genes.
Primers were designed based on the M. tuberculosis glnA1 sequence (30) and used to amplify a 1-kb target region encoding ∼70% of the M. smegmatis glnA1 gene from M. smegmatis 1-2c genomic DNA. The amplification product was cloned into pCR2.1 (Invitrogen), sequenced, and found to be highly similar to the M. tuberculosis glnA1 gene. The M. smegmatis glnA1 gene fragment was released from pCR2.1 by EcoRI digestion, gel purified, and used as a probe for the identification and isolation of its genomic locus in Southern and colony hybridizations. Likewise, a 0.65-kb EcoRI fragment containing the entire coding region of the M. smegmatis 1-2c sodA gene from a PCR amplification product (29) was used as a probe for the genomic M. smegmatis 1-2c sodA locus.
M. smegmatis 1-2c genomic DNA was completely digested with both BamHI and EcoRI and subjected to Southern analysis. The glnA1 probe hybridized to a 2.2-kb fragment, and the sodA probe hybridized to a 1.1-kb fragment. Restriction fragments from 1.7 to 2.4 kb and 0.8 to 1.2 kb were isolated and ligated into pUC18 digested with BamHI and EcoRI. The ligation was transformed into E. coli DH5α, and clones were verified for both genes by colony hybridizations, restriction analysis, and Southern hybridizations. The plasmids were designated pUC18-Ms-glnA1 and pUC18-Ms-sodA. For both plasmids, the entire insert was sequenced in both directions by cycle sequencing with ABI PRISM dye terminators (Davis Sequencing). One region of M. smegmatis 1-2c sodA exhibited particularly strong secondary structure and the sequence was determined with the Fidelity DNA Sequencing System (Oncor) with reaction products resolved on a 40% formamide–6 M urea gel. The sequences were assembled using the programs Pregap4 (version 1.0) and Gap4 (version 4.4) from the Staden Package (13). The sequences were scanned against the GenBank database using the BLAST programs (3) at the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/).
Construction of M. smegmatis glnA1 and sodA mutants.
A promoterless, nonpolar kanamycin resistance (Kmr) cassette was constructed using an approach similar to the one described previously (43), using a different kanamycin resistance gene, replacing the gram-negative ribosomal binding site with the M. tuberculosis dnaK ribosomal binding site and including convenient restriction sites for exchanging resistance genes (Fig. 1). The cassette was constructed by amplification of the Tn5 kanamycin resistance gene (aphA-2) using pCR2.1 as the template DNA and contains stop codons in all three reading frames immediately upstream of the ATG start codon of the aphA-2 gene. Immediately downstream of the aphA-2 gene stop codon is the sequence for the ribosomal binding site and ATG start codon of the M. tuberculosis dnaK gene to allow for translation of the 3′ portion of the disrupted gene. The blunt-end PCR product was cloned into the SmaI site of pUC19. Clones resistant to ampicillin and kanamycin were selected and were confirmed by restriction analysis. The cassette was released from pUC19 as a 0.8-kb fragment by AvaI digestion, and the 5′ overhangs were filled with T4 DNA polymerase.
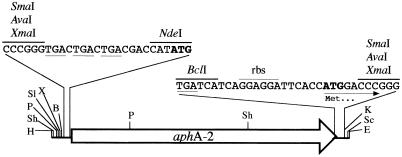
FIG. 1.
Nonpolar kanamycin resistance cassette cloned into the SmaI site of pUC19. The cassette contains the kanamycin resistance gene (aphA-2) coding region but lacks both a promoter and a transcriptional terminator. The stop codons in all three frames immediately upstream of aphA-2 are underlined, as is the stop codon of the gene. The start codon of aphA-2 and the start codon provided at the 3′ end of the cassette are shown in boldface type. The 15 nucleotides immediately downstream of the BclI site are identical to the sequence directly upstream of the M. tuberculosis dnaK gene (the ribosomal binding site [rbs] is indicated). NdeI and BclI sites were included to facilitate cloning of other resistance genes into the cassette. Abbreviations for restriction sites: H, HindIII; Sh, SphI; P, PstI; Sl, SalI; X, XbaI; B, BamHI; K, KpnI; Sc, SacI; E, EcoRI.
Unique sites in the coding regions of the M. smegmatis glnA1 and sodA genomic loci were identified (XhoI in pUC18-Ms-glnA1 and BstEII in pUC18-Ms-sodA). The plasmids were digested, and the 5′ overhangs were filled with T4 DNA polymerase. The Kmr cassette was blunt end ligated into the vectors such that the 3′ portion of glnA1 or sodA was in frame with the ATG start codon provided at the 3′ end of the Kmr cassette (Fig. 2B and D). Clones were identified by restriction analysis that had the Kmr cassette in the same orientation as glnA1 or sodA, allowing for transcription of the kanamycin resistance gene from the glnA1 or sodA promoter. The disrupted genes were released from pUC18 by digestion with BamHI and EcoRI. The EcoRI sites were modified by ligation to an adapter with multiple restriction sites (designated ECHXB) (Fig. 3) and the fragments were redigested with BamHI and cloned into the BamHI site of the temperature-sensitive allelic exchange vector, pPR27 (51). Constructions were verified by restriction analysis and designated pPR27-Ms-glnA1::Kmr and pPR27-Ms-sodA::Kmr.
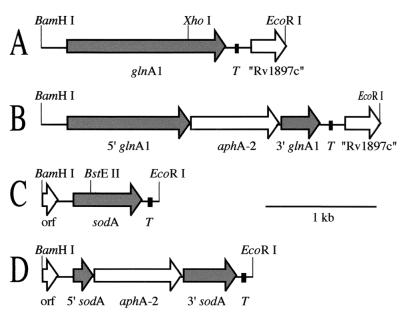
FIG. 2.
Maps of the M. smegmatis 1-2c glnA1 and sodA genomic loci. Both the glnA1 (A) and the sodA (C) loci were isolated as BamHI→EcoRI (5′-to-3′) genomic fragments. Both genes have putative transcriptional terminators (T) directly downstream of their coding regions. The glnA1 locus contains a truncated open reading frame (orf) with a high degree of similarity to the M. tuberculosis Rv1897c gene. The sodA locus contains a truncated open reading frame upstream of sodA with a predicted protein sequence that is highly similar (75% identity) to a truncated open reading frame in the same location in the M. fortuitum sodA locus (44). The glnA1 and the sodA loci were cloned into the multiple cloning site of pNBV1 for expression of GS and SOD in M. smegmatis and M. tuberculosis. The Kmr cassette (Fig. 1) was inserted into the unique XhoI site of glnA1 (B) and the BstEII site of sodA (D) to generate the disrupted loci used for allelic exchange.
FIG. 3.
ECHXB adapter. EcoRI adapter with several restriction sites useful for cloning into the multiple cloning site of pNBV1 and other vectors.
The two allelic exchange constructs were electroporated into M. smegmatis 1-2c, and transformants were selected on 7H11 with kanamycin (20 μg ml−1) at 32°C. Transformants were inoculated into 7H9 with kanamycin (20 μg ml−1) and 5 mM l-glutamine and incubated at 32°C for 8 h to allow allelic exchange to take place. Cells were then plated on 7H11 with kanamycin (20 μg ml−1) supplemented with 10% (wt/vol) sucrose and 5 mM l-glutamine and incubated at 39°C for 4 days to counterselect against those cells that retained the plasmid. Out of a total of 1,500 clones (each), six sodA clones and three glnA1 clones were selected for further analysis. Four of the six sodA clones no longer expressed SodA activity. All three glnA1 clones grew well on plates with added l-glutamine (5 mM) but did not grow at all on regular 7H11 plates. Genomic DNA was isolated from one sodA and one glnA1 clone, digested to completion with both BamHI and EcoRI, and subjected to Southern analysis (Fig. 4) using the following radiolabeled DNA probes: the M. smegmatis glnA1 genomic locus (Fig. 2A), the M. smegmatis sodA genomic locus (Fig. 2C), vector (pPR27), and the Kmr cassette. For the glnA1 mutant, the 2.2-kb band that is present in the wild-type strain was replaced by a 3-kb band indicating insertion of the 0.8 kb Kmr cassette. Likewise, for the sodA mutant, the 1.1-kb band that is present in the wild-type strain was replaced by an ∼2-kb band. When probed with just the Kmr cassette, only the 3- and 2-kb bands (glnA1 and sodA mutants, respectively) hybridized, and nothing hybridized in the wild-type DNA (data not shown). Also, the vector (pPR27) did not hybridize to DNA from the wild-type or mutant strains (data not shown).
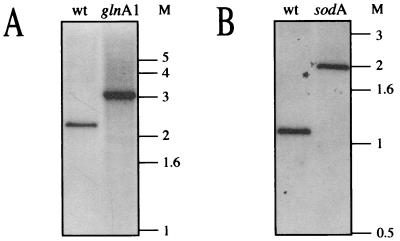
FIG. 4.
Southern analysis of M. smegmatis glnA1 and sodA mutants. Genomic DNA from M. smegmatis 1-2c (wild type [wt]), M. smegmatis glnA1, and M. smegmatis sodA strains was digested with BamHI and EcoRI and probed with the entire glnA1 locus (Fig. 2A) (A) or the entire sodA locus (Fig. 2C) (B). M, molecular mass markers in kilobases.
Construction of recombinant mycobacterial strains.
In previous studies, the M. tuberculosis glnA1 locus was cloned into pSMT3 for expression in the M. smegmatis 1-2c wild-type strain (30). This 1.8-kb locus was transferred as a BamHI→HindIII fragment (5′ to 3′) into the multiple cloning site of pNBV1. The M. tuberculosis sodA locus was previously cloned into pNBV1 as a ClaI→BamHI PCR product (5′ to 3′), and its expression was studied in M. smegmatis 1-2c (29). This plasmid was used without modification.
The M. smegmatis glnA1 and sodA genomic loci cloned into pUC18 likely contained sufficient upstream DNA sequence to encode all of the necessary promoter elements for expression in a mycobacterial host. Therefore, the loci were released by digestion with BamHI and EcoRI and the EcoRI site was modified by ligation to the ECHXB adapter (Fig. 3). The DNA was digested with XbaI or HindIII and cloned into the mycobacterial shuttle vector pNBV1 cut with BamHI and HindIII or BamHI and XbaI so as to obtain the genes in both orientations on the plasmid. Prior to electroporation into mycobacteria as described above, all constructs were confirmed by restriction analysis. The glnA1 plasmids were electroporated into the glnA1 mutant, and the sodA plasmids were electroporated into the sodA mutant. Transformants expressed high levels of GS and SOD for both orientations of the glnA1 and sodA loci, respectively. One clone of each with the gene cloned (5′ to 3′) in the BamHI and HindIII sites was chosen for all further studies (designated pNBV1-MsGS and pNBV1-MsSODA).
The gene encoding a UV-optimized green fluorescent protein (GFPuv) was amplified by PCR from pGFPuv (Clontech), modifying the upstream region to contain an NdeI site at the ATG start codon and adding a HindIII site downstream of the stop codon. The PCR product was digested with NdeI and HindIII and cloned into a pNBV1 construct that contains a modified M. bovis BCG hsp60 promoter (amplified from pSMT3) in which the sequence upstream of the ATG start codon was modified to contain an NdeI site. The gene contains an internal NdeI site which required first cloning the 3′ portion of the gene into the vector as an NdeI→HindIII (5′ to 3′) fragment. A clone was isolated and digested with NdeI, and the 5′ portion of the gene was cloned as an NdeI fragment.
The remaining genes (M. tuberculosis mdh, M. tuberculosis bfrB, E. coli sodA, E. coli sodB, B. subtilis sodA, and S. enterica serovar Typhimurium glnA) were amplified from genomic DNA using the Advantage-GC2 PCR kit (Clontech) and cloned downstream of the M. tuberculosis glnA1 promoter. For all of the genes except for bfrB the same strategy was employed, cloning into the BsmI site at the 5′ end (immediately upstream of the M. tuberculosis glnA1 GTG start codon) and the HindIII site at the 3′ end. The 5′ forward primers started with the sequence 5′-GCTAGCATTCTGTG with 18 to 20 additional nucleotides corresponding to codons 2 through 8 of the gene being amplified. The BsmI site is underlined, and the GTG start codon is shown in italics. The 3′ reverse primers incorporated a HindIII site (underlined) immediately downstream of the gene's stop codon, 5′-CCCAAGCTT, with 21 or 22 additional nucleotides complementary to the 3′ end of the gene. Because a BsmI site is present in pNBV1, the 1.8-kb M. tuberculosis glnA1 locus was first transferred into pUC19. The PCR-amplified genes were cloned downstream of the glnA1 promoter, replacing the glnA1 coding region, and then the gene plus promoter was transferred to pNBV1 as a BamHI→HindIII fragment. The B. subtilis sodA gene has an internal HindIII site that was first removed by PCR mutagenesis.
One of the genes (bfrB) was cloned downstream of an M. tuberculosis glnA1 promoter modified to contain an NdeI site at the start codon. There are no NdeI sites in pNBV1, which allowed for the PCR product to be cloned downstream of the promoter directly in pNBV1, avoiding the initial cloning into pUC19 required for the earlier constructs described above. The sequence upstream of and including the GTG start codon, TTCTGTG, was modified to TTCaTaTG in otherwise identical promoters (start codons are underlined and lowercase letters indicate added or modified nucleotides). The forward bfrB primer incorporated an NdeI site at the ATG start codon and the reverse bfrB primer incorporated a HindIII site as described above.
Recombinant plasmids carrying both sodA and glnA1 as well as a third gene were constructed by first cloning the M. smegmatis sodA locus into the BamHI and XbaI (5′-to-3′) sites of pNBV1-MsGS to yield a plasmid (designated pNBV1-MsGS-MsSODA) containing both M. smegmatis glnA1 and sodA in opposite orientations. A 3.3-kb HindIII fragment from pNBV1-MsGS-MsSODA, containing the entire sequence of both loci, was cloned into the HindIII site of pNBV1-MDH and pNBV1-BFRB (Fig. 5).
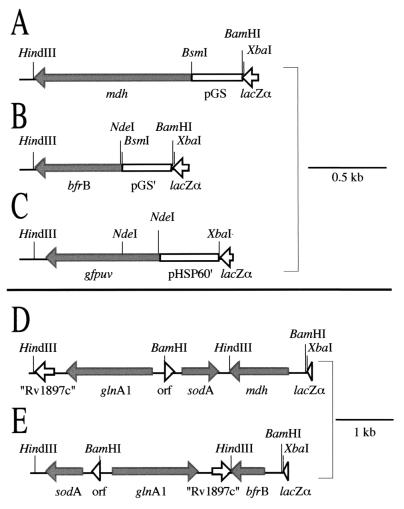
FIG. 5.
Maps of the mycobacterial expression constructs. All of the constructs were cloned into the HindIII and BamHI or XbaI sites of the multiple cloning site of pNBV1. (A) The M. tuberculosis mdh gene was cloned downstream of the M. tuberculosis glnA1 promoter (pGS). The following genes were cloned in the same way as mdh: E. coli sodA, E. coli sodB, B. subtilis sodA, and S. enterica serovar Typhimurium glnA. (B) The M. tuberculosis bfrB gene was cloned downstream of an M. tuberculosis glnA1 promoter (pGS′) modified to have an NdeI site at the start codon. (C) The UV-optimized GFP gene (gfpuv) was cloned downstream of an M. bovis BCG hsp60 promoter (pHSP60′) modified to contain an NdeI site at the start codon. (D and E) A 3.3-kb HindIII fragment containing the M. smegmatis glnA1 and sodA loci was cloned into the HindIII site of pNBV1-MDH (A) or pNBV1-BFRB (B). The M. smegmatis glnA1 and sodA genes were transcribed from their own promoters.
Plasmids were electroporated into mycobacteria as described above. Recombinant M. smegmatis and M. tuberculosis strains are listed in Table 2.
Analysis of M. smegmatis and M. tuberculosis recombinant strains.
Cultures were typically inoculated with cells from 7H9 cultures in late log phase or early stationary phase to give a calculated A550 of 0.01 to 0.02 (100-fold dilution). When the cultures were intended for analysis of early-log-phase extracellular proteins, the cells were washed in fresh media before inoculation to avoid carryover of extracellular proteins. Because some proteins were expressed at higher levels at 28 than 37°C in M. smegmatis, particularly malate dehydrogenase (MDH), all M. smegmatis cultures were grown at 28°C except where indicated. When single time points of a culture were analyzed, the cultures were grown to late log phase or early stationary phase prior to harvest. Aliquots (10, 20, 40, or 100 ml) were removed for analysis and centrifuged. The supernate was filtered (Acrodisc PF 0.8/0.2 μm filter; Pall Corporation) and concentrated to 100 to 200 μl with a Centricon Plus-20 concentrator (5000 or 8000 MWCO; Millipore), and the concentrate was then diluted to 0.5 or 1.0 ml with phosphate-buffered saline (PBS). The cell pellet was washed by resuspending in 5 ml of PBS, centrifuging, and decanting the supernatant fluid. The cells were stored frozen at −80°C or used immediately. Cells were lysed by resuspending in 5 ml of PBS and sonicating twice for 2 min each (M. smegmatis) or once for 4 min (M. tuberculosis) on ice with a Heat Systems Ultrasonics W-375 sonicator (50% pulse; maximum setting with a microtip). Cellular debris was removed by centrifugation and the cleared lysate was filtered (pore size, 0.2 μm).
Enzyme assays.
GS was assayed by the transfer reaction (73). Enzyme (100 μl) was added to 900 μl of assay mix (prewarmed to 37°C) to give the following concentrations of components: 30 mM l-glutamine, 60 mM NH2OH, 20 mM arsenate, 20 mM imidazole, 1 mM MnCl2, and 0.4 mM ADP at pH 7.0. The reaction was incubated at 37°C for 10 to 120 min (depending on activity) and stopped by the addition of 250 μl of 8% (wt/vol) trichloroacetic acid, 2 N HCl, and 3.3% (wt/vol) FeCl3. The samples were centrifuged for 1 min to remove precipitated protein, and the absorbance was read at 540 nm. Blank reactions, which substituted PBS for enzyme, were subtracted. A standard curve was prepared from gamma-glutamic acid hydroxamate. A unit is defined as the amount of enzyme that catalyzes the formation of 1 μmol of gamma-glutamic acid hydroxamate per min under the assay conditions.
SOD was assayed using a modified version of the method described by Ukeda et al. (66). Enzyme (100 μl) was added to 800 μl of assay mix, and the reaction was initiated by the addition of xanthine oxidase (100 μl; 19 mU ml−1 in PBS containing 1 mg of bovine serum albumin ml−1) to give the following concentration of components: 50 mM NaH2PO4 (pH 8.0), 200 mM NaCl, 0.1 mM EDTA, 0.1 mM xanthine, 25 μM XTT, and 1.9 mU of xanthine oxidase per ml. The reaction was monitored continuously at 470 nm at room temperature (25°C) and was linear for 12 min. One unit is defined as the amount of SOD that inhibits by 50% the reduction of XTT by superoxide.
MDH was assayed by adding 100 μl of enzyme to 900 μl of assay mix to give the following concentrations of components: 50 mM NaH2PO4 (pH 7.5), 200 mM NaCl, 1 mM KCN, 0.2 mM NADH, and 2 mM oxaloacetate. The reaction was monitored continuously at 340 nm for 2 to 3 min during which time the reaction was linear. Blank reactions in which oxaloacetate was omitted were subtracted. KCN at 1 mM was useful in inhibiting respiratory chain oxidation of NADH (17). For M. smegmatis strains overexpressing the M. tuberculosis MDH, lysates were diluted 100-fold in PBS immediately before assay. A unit is defined as the amount of enzyme that catalyzes the consumption of 1 μmol of NADH per min under the assay conditions (ɛ = 6,300 M−1 cm−1).
SDS-polyacrylamide gel electrophoresis (PAGE) analysis.
Culture filtrates and cell lysates were analyzed on sodium dodecyl sulfate (SDS)–12.5% polyacrylamide gels. The gels were stained with colloidal Coomassie brilliant blue G-250 (47).
SOD activity gels.
Culture filtrates and cell lysates were electrophoresed on 15% nondenaturing polyacrylamide gels at 80 V for 16 to 20 h at room temperature. Gels were negatively stained for SOD activity as described in reference 10.
Immunoblotting.
Culture filtrates and cell lysates of M. tuberculosis pNBV1-GFPuv were electrophoresed on an SDS–12.5% polyacrylamide gel and transferred to a nitrocellulose membrane. Strips containing GS, SOD, and GFPuv antigens were probed separately with rabbit polyclonal antibodies specific for the M. tuberculosis GlnA1 (diluted 1:2,500) (28), the M. tuberculosis SodA (diluted 1:2,500) (29), and GFP (diluted 1:1,000) (Molecular Probes). The membranes were subsequently incubated with horseradish peroxidase-conjugated goat anti-rabbit antibodies (diluted 1:10,000; Bio-Rad), a chemiluminescent substrate (SuperSignal West Pico; Pierce) was added, and the proteins were visualized on X-ray film.
Gel and immunoblot analysis.
Gels were photographed wet and the negatives were scanned to obtain an 8-bit gray-scale image. Developed X-ray film from chemiluminescent immunoblotting was scanned directly. Protein bands were quantitated using the public domain NIH Image (version 1.62) program (developed at the U.S. National Institutes of Health and available on the Internet at http://rsb.info.nih.gov/nih-image/).
Metabolic labeling of M. tuberculosis pNBV1-GFPuv.
An actively growing culture of M. tuberculosis pNBV1-GFPuv (10 ml) was diluted into 40 ml of fresh medium (7H9 with hygromycin at 50 μg ml−1) to give an initial A550 of ∼0.06 and grown for 4 days at 37°C to an A550 of 0.18. Then, a mixture of [35S]l-methionine and [35S]l-cysteine (Promix; Amersham Pharmacia Biotech) was sterile filtered and added to the culture at a final concentration of 50 μCi ml−1. The culture was incubated at 37°C for 3 more days, reaching a final A550 of 0.28, during which time 10-ml aliquots were removed at 7, 23, 48, and 72 h. Culture filtrates and cell lysates were prepared as described above and analyzed by SDS-PAGE. After staining with Coomassie brilliant blue, the gel was dried and exposed to X-ray film for various times (16 to 160 h).
Purification and stability of MDH.
M. smegmatis cells overexpressing the M. tuberculosis MDH (M. smegmatis glnA1 pNBV1-MsGS-MsSODA-MDH) were grown to stationary phase in 7H9 and harvested by centrifugation. The supernate was sterilized by filtration (pore size, 0.2 μm) and saved at 4°C. The cells were resuspended in PBS, lysed by sonication, and centrifuged, and the cleared lysate was filtered (pore size, 0.2 μm). The lysate (15 ml) was loaded on a Reactive Red-120 Sepharose column (20 ml, 2.6 by 3.5 cm) and eluted by gravity flow. The column was eluted with 90 ml of PBS, 60 ml of PBS–1 mM AMP, 30 ml of 0.1× malate buffer, 90 ml of 0.1× malate buffer–1 mM AMP, and finally with 90 ml of 1× malate buffer–1 mM AMP (1× malate buffer is 50 mM NaH2PO4, 100 mM l-malic acid, and 500 mM NaCl [pH 7.5]). Fractions were assayed for MDH activity, which was found only in the final wash with 1× malate buffer. The active fractions were concentrated with Centricon Plus-20 concentrators (MWCO 8000) and loaded on a Superdex-75 column (1.6 by 55 cm) equilibrated in PBS. The column was eluted at 1 ml min−1 with PBS and MDH was collected as a single, symmetrical peak that eluted very close to the void volume.
Purified MDH (∼500 μg in 5 ml) was sterile filtered (pore size, 0.2 μm), added to 195 ml of the sterile culture filtrate (see above), and incubated at 28°C with shaking (same conditions as for the M. smegmatis cultures). Aliquots of 20 ml were removed immediately after adding MDH and 1, 2, 3, and 7 days later. The samples were concentrated to 100 to 200 μl with Centricon Plus-20 concentrators (8000 MWCO) and then diluted to 1.0 ml with PBS. The concentrated samples were stored at 4°C until day 3, at which point all of the samples were assayed for MDH and GS activity. On day 7, all of the samples were analyzed by SDS-PAGE.
Nucleotide sequence accession numbers.
The nucleotide sequences have been deposited in the GenBank database (glnA1, GenBank accession number AY008693; sodA, GenBank accession number AF061031).
RESULTS
The proportion of GS and SOD exported from M. smegmatis and M. tuberculosis is not dependent upon the species from which the protein is derived.
Previously, we found a high level of expression and export of recombinant M. tuberculosis GS and SOD compared with the endogenous M. smegmatis homologues in a wild-type M. smegmatis host (29, 30). To explore export of these large, leaderless, multimeric proteins further, we cloned the M. smegmatis glnA1 and sodA genes and constructed glnA1 and sodA mutants (see Materials and Methods), allowing for high-level expression of the M. tuberculosis and M. smegmatis proteins in a null background. The M. smegmatis SodA was found to be highly similar (>80% identity) to many mycobacterial SodA proteins as well as the SodA from Nocardia asteroides (Mycobacterium fortuitum, accession no. X70914; Mycobacterium avium, accession no. U11550; Mycobacterium lepraemurium, accession no. D13288; M. leprae, accession no. X16453; N. asteroides, accession no. U02341; M. tuberculosis, accession no. AF061030). The M. smegmatis GlnA1 was most similar (83 to 84% identity) to the M. tuberculosis GlnA1 (accession no. U87280) and M. leprae GlnA1 (accession no. CAC31306), with the next most similar GlnA proteins (70 to 73% identity) being from Streptomyces coelicolor (accession no. M23172), Corynebacterium glutamicum (accession no. Y13221), and Amycolatopsis mediterranei (accession no. AF050112). In contrast, the mycobacterial SodA proteins share only 39 to 45% identity with E. coli and B. subtilis SODs (Table 3), and the GlnA1 proteins share ∼50% identity with S. enterica serovar Typhimurium GlnA (accession no. M14536).
TABLE 3.
Identity matrix of bacterial superoxide dismutases
| Proteina | %
Identityb with:
|
Accession no. | ||||
|---|---|---|---|---|---|---|
| M. fortuitum SodA | M. tuberculosis SodA | E. coli SodA | E. coli SodB | B. subtilis SodA | ||
| M. smegmatis SodA (Mn)a | 96 | 81 | 43 | 39 | 44 | AF061031 |
| M. fortuitum SodA (Mn) | 81 | 42 | 38 | 44 | X70914 | |
| M. tuberculosis SodA (Fe) | 41 | 40 | 45 | AF061030 | ||
| E. coli SodA (Mn) | 45 | 60 | X03951 | |||
| E. coli SodB (Fe) | 53 | J03511 | ||||
| B. subtilis SodA (Mn) | D86856 | |||||
The metal cofactor used by the SOD is given in parentheses.
The percentage of identical residues is given for each pair of proteins based on a multiple alignment of all the proteins using the program Clustal X (version 1.8) (64).
Expression and export of homologous and heterologous GSs from the M. smegmatis wild-type and glnA1 strains.
The M. smegmatis glnA1 strain has an absolute requirement for l-glutamine. Complementation of the glnA1 strain was achieved by supplying either a copy of the M. smegmatis glnA1 or M. tuberculosis glnA1 in trans on pNBV1 with expression driven from their own promoters. Because genes are often poorly expressed from E. coli promoters in M. smegmatis (8), expression of S. enterica serovar Typhimurium glnA was driven from the strong M. tuberculosis glnA1 promoter (Fig. 5A). This construct was also capable of complementing the glnA1 mutant. The M. smegmatis wild type, glnA1 mutant, and complemented strains were analyzed for expression and export (extracellular accumulation) of GS by measuring enzyme activity (Table 4) and by SDS-PAGE analysis (Fig. 6). The M. smegmatis wild-type strain expressed GS activity at 94 mU per ml of original culture volume, while virtually no GS activity (0.1 mU ml−1) was detected for the glnA1 mutant (grown with 10 mM l-glutamine). Growth of the wild-type strain with 10 mM l-glutamine did decrease the GS activity to approximately 20% compared to growth in regular 7H9 but still gave clearly detectable activity. The level of GS activity in the mutant strain complemented with the M. smegmatis glnA1 (Ms glnA1 pNBV1-MsGS) or M. tuberculosis glnA1 (Ms glnA1 pNBV1-MtbGS) was 6- to 13-fold higher than in the wild-type strain. Consistent with this, bands of much greater intensity (compared with the wild-type strain) were observed on SDS-PAGE analysis. Even though the S. enterica serovar Typhimurium glnA was expressed from a strong mycobacterial promoter, the mutant strain complemented with this gene (Ms glnA1 pNBV1-StGS) had very low levels of GS activity (4 mU ml−1), but the levels were clearly detectable above background and no protein band for GS could be detected by SDS-PAGE analysis. However, this strain grew just as well as the strains complemented with the mycobacterial glnA1 genes.
TABLE 4.
Expression and export of GS from M. smegmatis
| M. smegmatis strain | Activity (mU
ml−1)a
|
Extracellular activity (%) | Relative activityb | |
|---|---|---|---|---|
| Culture filtrate | Cell lysate | |||
| pNBV1 | 6.3 | 87.9 | 6.7 | 1.00 |
| pNBV1 (10 mM L-Gln) | 0.4 | 17.9 | 2.3 | 0.19 |
| glnA1 pNBV1 (10 mM L-Gln) | 0.0 | 0.1 | 0.0 | 0.00 |
| glnA1 pNBV1-MsGS | 62.3 | 1,238.4 | 4.8 | 13.81 |
| glnA1 pNBV1-MtbGS | 13.1 | 520.6 | 2.5 | 5.67 |
| glnA1 pNBV1-StGS | 0.1 | 3.9 | 2.7 | 0.04 |
Activity is expressed in milliunits per milliliter of the original culture volume.
Based on the total GS activity in the culture filtrate and lysate. Values are relative to the activity of the wild-type strain (pNBV1).
FIG. 6.
GS expression and export in M. smegmatis wild-type and M. smegmatis glnA1 strains. SDS-PAGE analysis of culture filtrates (CF) and lysates (L). Tenfold more CF than L was loaded on the gel (the equivalent of 2 ml versus 0.2 ml of the original culture volume). The results are representative of three independent cultures. Arrows indicate the positions of the M. smegmatis (Ms) and M. tuberculosis (Mtb) GSs. M, molecular mass markers in kilodaltons.
For all of the strains, the proportion of total GS activity detected in culture filtrates was quite low (2 to 7%), and no significant difference in the proportion of the M. smegmatis and M. tuberculosis GSs exported was observed.
Expression and export of homologous and heterologous SODs from M. smegmatis, M. smegmatis sodA, and M. tuberculosis.
The sodA mutation in M. smegmatis did not appear to result in any growth defect (i.e., the mutant grew well in 7H9 medium). Strains were constructed from the sodA mutant that expressed the M. smegmatis SodA, M. tuberculosis SodA, E. coli SodA, E. coli SodB, or B. subtilis SodA. Transcription of the mycobacterial sodA genes was from their own promoters. Nonmycobacterial SOD genes were all transcribed from the M. tuberculosis glnA1 promoter (Fig. 5A), as was done for the S. enterica serovar Typhimurium glnA. The plasmids containing the nonmycobacterial SOD genes were also transformed into the wild-type M. smegmatis to investigate whether the nonmycobacterial SOD subunits could mix with the M. smegmatis SodA subunits.
The strains were grown in 7H9 medium, and the culture filtrates and lysates were analyzed by SDS-PAGE and SOD activity gels (Fig. 7A to C and Table 5). By SDS-PAGE analysis, the 23-kDa band for the M. smegmatis SodA was absent from the sodA strain, as was the major band on the activity gel. However, there was a small amount of SOD activity in the sodA mutant that ran slightly higher than the M. smegmatis SodA on the activity gel. This band was not evident in the wild-type strain or the sodA strain expressing the M. smegmatis SodA from a plasmid (Ms sodA pNBV1-MsSODA). However, it may have run differently in the presence of the M. smegmatis SodA, perhaps forming hybrids, and been hidden by the larger band. A nearly identical phenotype was previously observed for an M. smegmatis sodA mutant resulting from a single crossover event (21). All of the recombinant SODs were expressed at levels comparable to, or greater than, the endogenous M. smegmatis SodA as judged by SDS-PAGE and SOD activity (Table 5).
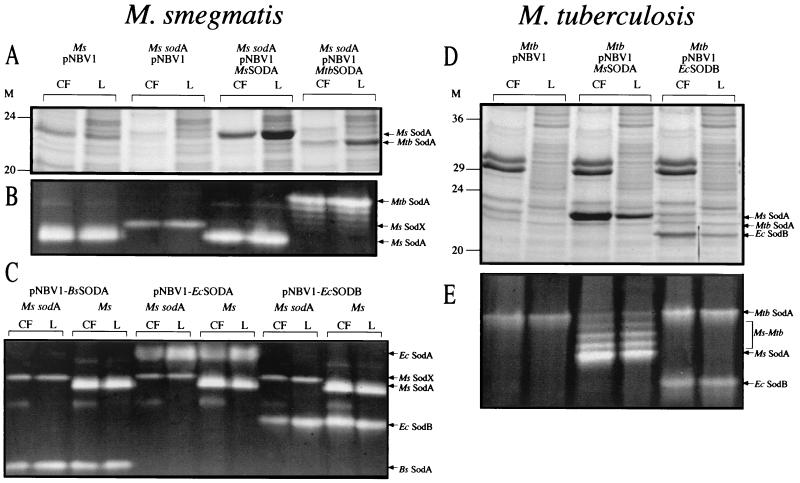
FIG. 7.
SOD expression and export in M. smegmatis wild type, M. smegmatis sodA, and M. tuberculosis wild-type strains. M. smegmatis strains were analyzed for SOD expression by SDS-PAGE (A) and SOD activity gel (B and C). M. tuberculosis strains were also analyzed for SOD expression by SDS-PAGE (D) and SOD activity gel (E). Tenfold more culture filtrate (CF) than lysate (L), based on the original culture volume, was loaded on the gels for each analysis (4 ml of CF and 0.4 ml of L [A and D], 6 ml of CF and 0.6 ml of L [B and C], and 2 ml of CF and 0.2 ml of L [E]). The results are representative of two or three independent cultures. Arrows indicate the positions of the M. smegmatis (Ms) SodA, M. tuberculosis (Mtb) SodA, M. smegmatis-M. tuberculosis hybrid SODs (Ms-Mtb), E. coli (Ec) SodA, E. coli SodB, B. subtilis (Bs) SodA, and the M. smegmatis SOD activity in the sodA mutant (Ms SodX). M, molecular mass markers in kilodaltons.
TABLE 5.
Expression and export of SOD from M. smegmatis and M. tuberculosis
| Species and strain | SODa | %
Extracellularb
|
Relative
amountc
|
||
|---|---|---|---|---|---|
| Activity gel | SDS-PAGE | Activity gel | SDS-PAGE | ||
| M. smegmatis | |||||
| pNBV1 | M. smegmatis SodA | 10 | 12 | 1.0 | 1.0 |
| sodA pNBV1 | M. smegmatis SodX | 6 | —d | 0.3 | — |
| sodA pNBV1-MsSODA | M. smegmatisSodA | 8 | 4 | 1.1 | 5.6 |
| sodA pNBV1-MtbSodA | M. tuberculosis SodA | 7 | 4 | 1.4 | 1.7 |
| sodA pNBV1-BsSODA | M. smegmatis SodX | 7 | — | 0.3 | — |
| B. subtilis SodA | 6 | 5 | 0.6 | 0.7 | |
| pNBV1-BsSODA | M. smegmatisSodA | 9 | |||
| B. subtilis SodA | 9 | 11 | 0.5 | — | |
| sodA pNBV1-EcSODA | M. smegmatis SodX | 7 | — | 0.2 | — |
| E. coli SodA | 4 | 7 | 1.1 | 1.6 | |
| pNBV1-EcSODA | M. smegmatis SodA | 12 | 14 | ||
| E. coli SodA | 7 | 14 | 1.0 | 2.4 | |
| sodA pNBV1-EcSODX | M. smegmatis SodX | 6 | — | 0.3 | — |
| E. coli SodB | 5 | 3 | 0.9 | 0.8 | |
| pNBV1-EcSODB | M. smegmatis SodA | 10 | 13 | ||
| E. coli SodB | 12 | 15 | 0.8 | 1.0 | |
| M. tuberculosis | |||||
| pNBV1 | M. tuberculosis SodA | 15 | 20 | 1.0 | 1.0 |
| pNBV1-MsSODA | M. tuberculosis SodA | 15 | 23 | ||
| M. smegmatis SodA | 20 | 20 | 3.2 | 7.9 | |
| pNBV1-EcSODB | M. tuberculosis SodA | 14 | 20 | ||
| E. coli SodB | 16 | 19 | 0.5 | 2.2 | |
Many of the strains have two SOD activities which often could be analyzed separately. The SOD activity present in the M. smegmatis sodA mutant, which was not evident in the wild-type strain, is designated SodX.
The percentages of extracellular SOD, from both SOD activity gels and SDS-polyacrylamide gels (Fig. 7), were determined with NIH Image software (version 1.62). Because B. subtilis SodA and M. smegmatis SodA have nearly identical mobilities on SDS-polyacrylamide gels, the extracellular percentage is based on the combination of the two SODs.
For M. tuberculosis and M. smegmatis wild-type strains, relative amount refers to the amount of recombinant SOD compared with the endogenous M. tuberculosis or M. smegmatis SodA, respectively (i.e., an internal comparison). For the M. smegmatis sodA strains, “relative amount” refers to the amount of recombinant SOD compared with the amount of M. smegmatis SodA produced by a culture of the M. smegmatis pNBV1 wild-type strain grown under the same conditions.
—, unable to analyze because a band could not be identified or was not clearly resolved.
All SODs, recombinant and endogenous, were present in the culture filtrates of M. smegmatis strains at relatively low percentages (3 to 15%) of the total expressed. M. tuberculosis and M. smegmatis SodA subunits are known to mix, forming hybrid enzymes (29, 76). Also, the E. coli SodA and SodB subunits form a hybrid SOD and the B. subtilis SodA subunits are capable of forming hybrid SODs with both the E. coli SodA and SodB subunits (11, 35). However, no mixing of the E. coli and B. subtilis SOD subunits with M. smegmatis SodA subunits was detected (Fig. 7C).
The recombinant M. tuberculosis strain expressing the M. smegmatis SodA (Mtb pNBV1-MsSODA) produced the recombinant enzyme at levels even greater than its own SodA (Fig. 7D and E and Table 5). The M. smegmatis SodA subunits formed hybrids with the endogenous M. tuberculosis SodA subunits as observed previously for M. smegmatis expressing the recombinant M. tuberculosis SodA (29, 76). The recombinant M. tuberculosis strain expressing the E. coli SodB (Mtb pNBV1-EcSODB) produced the recombinant SOD at levels comparable to its endogenous SodA. Mixing of the endogenous M. tuberculosis SodA subunits with the E. coli SodB subunits was not detected (Fig. 7E), as was previously observed for the M. smegmatis SodA and E. coli SodB subunits in an M. smegmatis host (Fig. 7C).
All three SODs (M. tuberculosis SodA, M. smegmatis SodA, and E. coli SodB) were present in the culture filtrate of M. tuberculosis at similar proportions (14 to 23%) of the total expressed.
The results presented thus far showed that while the absolute amounts of GS and SOD exported were high relative to other culture filtrate proteins, the proportion exported represented only a minority of the total expressed. Moreover, there was no significant difference in the proportion of GS and SOD exported depending upon whether they were native to pathogenic mycobacteria, nonpathogenic mycobacteria, or nonmycobacteria. This suggested that export was largely expression dependent rather than protein specific. To explore this possibility further, we prepared recombinant strains of M. smegmatis and M. tuberculosis expressing large amounts of three other leaderless proteins, all typically intracellular, and studied their expression and export.
The presence of large amounts of GS, SOD, and other leaderless proteins in the culture filtrate of mycobacteria reflects their high expression and extracellular stability. (i) Expression of MDH in M. smegmatis.
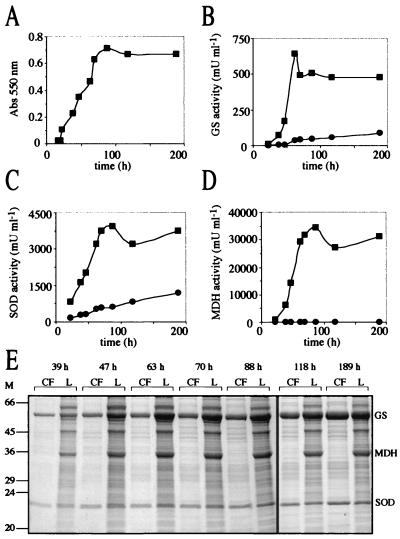
The cytoplasmic enzyme lactate dehydrogenase is frequently assayed as a marker for lysis of bacterial and eukaryotic cells (41). However, we chose MDH, a Krebs cycle enzyme that is also expected to be strictly intracellular, as a marker for autolysis in M. smegmatis primarily because M. smegmatis does not express an MDH activity (54) while M. tuberculosis expresses a high level of activity. This suggested that overexpression of the M. tuberculosis MDH in M. smegmatis would be successful and analysis would not be complicated by the presence of endogenous enzyme. MDH forms either dimers or tetramers of identical 30- to 35-kDa subunits and has a three-dimensional structure similar to LDH (45). The M. tuberculosis mdh gene was cloned downstream of the M. tuberculosis glnA1 promoter in pNBV1 (Fig. 5A), and a plasmid was constructed containing the M. smegmatis glnA1, M. smegmatis sodA, and mdh as separate transcriptional units (Fig. 5D) in order to follow MDH activity in recombinant bacteria overexpressing GS and SOD. This plasmid was transformed into the M. smegmatis glnA1 mutant, and expression and export of GS, SOD, and MDH were analyzed by SDS-PAGE and enzyme activity as a function of growth (Fig. 8). The strain expressed large amounts of all three enzymes. The percentage of GS and SOD activity in the culture filtrate increased during growth and reached a maximum of 15 and 25% after the cells were well into stationary phase. During the earlier stages of growth (39 to 88 h), the percentages (means ± standard deviations) of extracellular GS protein (5.6% ± 1.4%; n = 5) and GS activity (6.2% ± 1.9%; n = 5) were similar to previous results (Fig. 6 and Table 4). The percentages of extracellular SOD protein (8.5% ± 0.6%; n = 5) and SOD activity (13.6% ± 0.6%; n = 5) were also similar to previous results (Fig. 7 and Table 5). In contrast, although there was a readily detectable quantity of MDH in the culture filtrate at all time points (10 to 70 mU per ml of original culture volume), this extracellular MDH activity represented less than 0.5% of the total MDH activity. The SDS-PAGE results were consistent, showing no accumulation of MDH in the culture filtrate.
FIG. 8.
Expression and export of GS, SOD, and MDH by M. smegmatis glnA1 pNBV1-MsGS-MsSODA-MDH. (A) Growth of M. smegmatis glnA1 pNBV1-MsGS-MsSODA-MDH. (B, C, and D) Enzyme activity of GS, SOD, and MDH in the culture filtrate (●) and lysate (▪) over time. (E) SDS-PAGE analysis of culture filtrates (CF) and lysates (L). Tenfold more CF than L, based on the original culture volume, was loaded on the gels (at 39 and 47 h, 4 ml of CF and 0.4 ml of L; at 63 to 189 h, 2 ml of CF and 0.2 ml of L). The positions of the M. smegmatis GS, M. smegmatis SOD, and the M. tuberculosis MDH are indicated. M, molecular mass markers in kilodaltons.
However, after 3 weeks in stationary phase, by which time the enzyme would have been expected to have been released extracellularly by autolysis, only 0.2% of the total MDH activity was present in the culture filtrate, while intracellular MDH activity was at 96% of its maximum value (data not shown). This suggested that MDH was not stable in the extracellular medium. Therefore, we examined the stability of the recombinant M. tuberculosis MDH in culture filtrate. We partially purified MDH from a cell lysate of M. smegmatis glnA1 pNBV1-MsGS-MsSODA-MDH, added the purified MDH to the sterile culture filtrate from the same strain, and incubated the mixture under the same conditions as a growing culture. Aliquots were removed at various times, analyzed by SDS-PAGE, and assayed for MDH and GS activity (Fig. 9). MDH was clearly unstable in 7H9 culture filtrate, while the amounts of other proteins such as GS and SOD remained constant over the course of 1 week at 28°C. Further experiments showed that MDH had a half-life of 2.3 to 3 h in culture filtrate or fresh 7H9 medium at 28°C, and the addition of a combination of protease inhibitors (2 mM EDTA, 1 mM phenylmethylsulfonyl fluoride, and 1 μM pepstatin A) or bovine serum albumin (10 mg ml−1) did little to improve stability (data not shown).
FIG. 9.
Stability of MDH in 7H9 culture filtrate. (A) SDS-PAGE analysis of purified MDH (lane 1) and its stability after addition to culture filtrate. An amount equivalent to 2 ml of the original culture volume was loaded for each culture filtrate time point. The positions of the M. smegmatis GS, M. smegmatis SOD, and the M. tuberculosis MDH are indicated. The purified MDH was judged to be 85% pure by analysis with NIH Image software (version 1.62). (B) The percentage of initial activity of MDH and GS is listed beneath each lane, with the activity at day zero defined to be 100%. Abbreviations: ND, not determined; M, molecular mass markers in kilodaltons.
(ii) Expression of bacterioferritin in M. smegmatis.
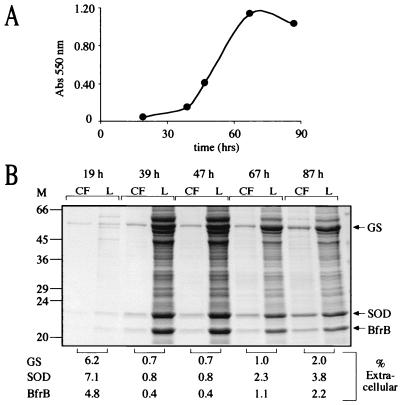
Because of the significant and unexpected instability of MDH in culture filtrates, we sought a more stable protein for use as an intracellular marker for M. smegmatis. Ferritins and bacterioferritins are large (∼450-kDa), highly stable, multimeric, iron storage proteins composed of 24 subunits of ∼19 kDa (6). M. tuberculosis has two bacterioferritin genes (bfrA and bfrB) that do not encode leader peptides. One of the bacterioferritin genes, bfrB, was cloned downstream of the M. tuberculosis glnA1 promoter (Fig. 5B), and a plasmid was constructed containing the M. smegmatis glnA1, M. smegmatis sodA, and M. tuberculosis bfrB as separate transcriptional units (Fig. 5E), as was done for mdh. This plasmid was transformed into the glnA1 mutant, and expression and export of GS, SOD, and BfrB were analyzed by SDS-PAGE as a function of growth (Fig. 10). In contrast to MDH, BfrB was detected in the culture filtrate at levels comparable to GS and SOD at all time points analyzed, even in early log phase. Furthermore, BfrB in the culture filtrate likely exists as a functional 24-subunit multimer based on analysis of culture filtrate proteins in which BfrB largely coeluted with GS (molecular mass, ∼640 kDa) on a Sephacryl-300HR size exclusion column (data not shown). Thus, in addition to GS, another very large and typically intracellular protein was released by M. smegmatis cultures into the growth medium.
FIG. 10.
Expression and export of GS, SOD, and BfrB by M. smegmatis glnA1 pNBV1-MsSODA-MsGS-BFRB. (A) Growth of M. smegmatis glnA1 pNBV1-MsGS-MsSODA-BFRB. (B) SDS-PAGE analysis of culture filtrates (CF) and lysates (L). Tenfold more CF than L, based on the original culture volume, was loaded on the gels (at 19 and 39 h, 8 ml of CF and 0.8 ml of L; at 47 h, 4 ml of CF and 0.4 ml of L; at 67 and 87 h, 2 ml of CF and 0.2 ml of L). The percentages of extracellular GS, SOD, and BfrB (listed below the gel) were determined with NIH Image software (version 1.62). Arrows indicate the positions of the M. smegmatis GS, M. smegmatis SOD, and the M. tuberculosis BfrB (BfrB). M, molecular mass markers in kilodaltons.
(iii) Expression of GFP in M. tuberculosis.
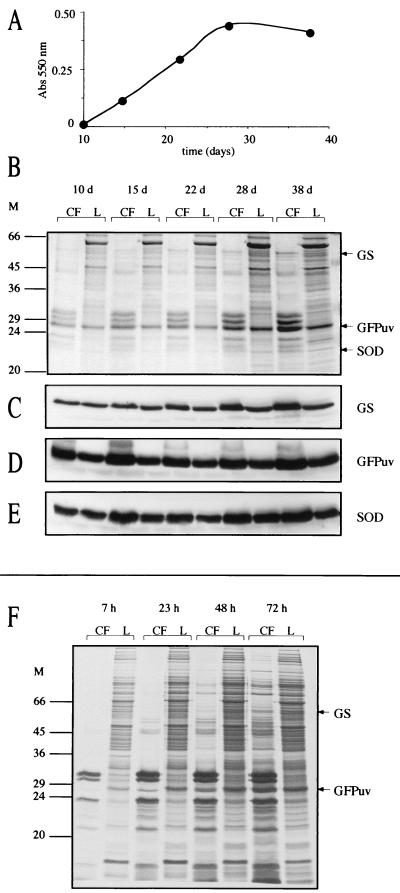
We chose the nonbacterial GFP as an intracellular marker for M. tuberculosis because we anticipated that (i) it would be expected to remain strictly intracellular since it lacks a leader peptide, (ii) it would not be exported by M. tuberculosis based on similarities to endogenous proteins since it is not a bacterial protein, and (iii) it would persist in the culture medium if released since it is highly stable (14). M. tuberculosis was transformed with a plasmid containing the gene for GFPuv downstream of the M. bovis BCG hsp60 promoter (Fig. 5C), a strong mycobacterial promoter that has been successfully used to express GFP in mycobacteria (20, 37). High level expression of GFPuv was obtained, and the cells were bright green when examined under long-wavelength UV light. Expression and export of the recombinant GFPuv along with the endogenous M. tuberculosis GlnA1 and M. tuberculosis SodA was analyzed by SDS-PAGE and immunoblotting as a function of growth (Fig. 11). All three proteins were clearly present in the M. tuberculosis culture filtrate during all stages of growth and at similar percentages (GS, 8 to 14%; GFPuv, 11 to 18%; SOD, 8 to 15%). Expression of GFPuv had no noticeable toxicity on the cells, and the export of SodA by the GFPuv-expressing strain was consistent with previous results obtained with three different M. tuberculosis strains not expressing GFPuv (Fig. 7 and Table 5) (SodA protein, 21.0% ± 1.7%; SodA activity, 14.7% ± 0.6%).
FIG. 11.
Expression and export of GS, SOD, and GFPuv by M. tuberculosis pNBV1-GFPuv. (A) Growth of M. tuberculosis pNBV1-GFPuv. (B) SDS-PAGE analysis of culture filtrates (CF) and lysates (L). Tenfold more CF than L, based on the original culture volume, was loaded on the gels (at 10 days; 15 ml of CF and 1.5 ml of L; at 15 days, 6 ml of CF and 0.6 ml of L; at 22 to 38 days, 2 ml of CF and 0.2 ml of L). (C to E) Immunoblot analysis of GS, GFPuv, and SOD. (F) Autoradiogram of CF and L from M. tuberculosis pNBV1-GFPuv metabolically labeled with [35S]l-methionine and [35S]l-cysteine. The bacteria were grown for 4 days, radiolabel was added, and aliquots were removed for analysis 7, 23, 48, and 72 h later. The culture was actively growing over the 72-h period, increasing in turbidity (A550) from 0.18 to 0.28. Twentyfold more CF than L was loaded on the gel (the equivalent of 2 ml versus 0.1 ml of original culture volume). Arrows indicate the positions of the M. tuberculosis GS, M. tuberculosis SOD, and GFPuv. M, molecular mass markers in kilodaltons.
Because all three proteins appeared in the culture filtrate even during the early stages of growth, the kinetics of export were investigated by metabolic labeling of M. tuberculosis pNBV1-GFPuv with [35S]l-methionine and [35S]l-cysteine. Export of GFPuv and the endogenous M. tuberculosis GlnA1 was analyzed over a 3-day period (Fig. 11F). By 7 h, the cell lysate proteins were fairly heavily labeled, whereas the culture filtrate was nearly devoid of labeled proteins with the exception of the 30/32-kDa complex (antigen 85 complex) and 23.5-kDa (MPT64) major secretory antigens. These antigens all contain N-terminal signal peptides and have high localization indexes (70). Radiolabeled GFPuv and GlnA1 in the 7-h culture filtrate were close to, or below, detection limits and accounted for ≤0.5% of the total metabolically labeled GFPuv and GlnA1. By 72 h, only 5% of GFPuv and 3% of GlnA1 were present in the culture filtrate. The 30/32-kDa and 23.5-kDa antigens were very heavily labeled at this time, and many other proteins were evident in the culture filtrate. SodA could not be analyzed by this technique because it is poorly labeled, possessing only two methionine residues (one of which is cleaved off [29]) and no cysteine residues. GlnA1 and GFPuv have 13 and 7 total methionine and cysteine residues, respectively.
These experiments on expression and export of MDH and bacterioferritin by M. smegmatis and GFPuv by M. tuberculosis demonstrate that proteins accumulate extracellularly in large amounts if they are abundantly expressed and resistant to degradation in the extracellular milieu. Also, the metabolic labeling showed that the export is very likely not an active process because of the very slow release of GS and GFPuv compared to actively secreted mycobacterial proteins.
The role of carbon and nitrogen on release of proteins from M. smegmatis and evidence that autolysis plays a role in the accumulation of extracellular proteins.
To investigate the potential role of carbon and nitrogen in protein export, we cultured M. smegmatis glnA1 pNBV1-MsSODA-MsGS-BFRB for 4 days in 7H9 medium in the presence of various amounts of added glucose and (NH4)2SO4. The addition of both (NH4)2SO4 and glucose to 7H9 medium resulted in a dramatic decrease in extracellular proteins (Fig. 12A). In fact, rather modest absolute amounts of the two substances [12.4 mM (NH4)2SO4 and 0.25% (wt/vol) Glc] were capable of preventing the accumulation of extracellular proteins (Fig. 12A, culture 2). Growth in 7H9 medium with additional (NH4)2SO4 but without glucose did not affect the profile of extracellular proteins (Fig. 12B, culture 6). However, growth in 7H9 medium with 1% (wt/vol) glucose and no additional (NH4)2SO4 resulted in a much greater level of extracellular proteins than growth in standard 7H9 (Fig. 12C, culture 10). The increase in extracellular proteins was most likely due to autolysis, because cultures analyzed after 2 days rather than after 4 days of growth showed no difference in culture filtrate proteins compared with growth in standard 7H9 medium (data not shown). Glycerol (1%, vol/vol) could substitute for glucose (1% [wt/vol]), and NH4Cl could substitute for (NH4)2SO4. MgCl2, MgSO4, and NaCl could not substitute for a source of NH4+ (data not shown).
After prolonged incubation of the M. smegmatis strain in stationary phase (10 days) in 7H9 medium containing 12.4 mM (NH4)2SO4 and 0.25% (wt/vol) glucose, extracellular proteins again accumulated, resulting in a culture filtrate protein profile similar to growth for 4 days in standard 7H9 medium. This suggested that the added nutrients were eventually consumed after which a portion of the culture lysed (data not shown). However, after prolonged incubation of the M. smegmatis strain in stationary phase (10 days) in 7H9 containing 38 mM (NH4)2SO4 and 1% (wt/vol) glucose, almost no extracellular proteins were present, presumably because an excess of carbon and nitrogen was maintained. It was also observed that high-molecular-weight DNA (≥12 kb) was present in culture filtrates and was highly correlated with the presence of extracellular proteins (i.e., DNA was readily detected in culture filtrates when proteins were present but not when proteins were absent [data not shown]), providing further evidence that autolysis was occurring and that increased levels of glucose and nitrogen could prevent it.
When M. smegmatis glnA1 pNBV1-MsSODA-MsGS-BFRB was grown under the same conditions as M. tuberculosis (37°C unshaken), additional glucose and NH4+ still prevented the release of GS, SOD, and BfrB into the growth medium (data not shown). However, when M. tuberculosis was grown under the four extreme culture conditions (cultures 1, 5, 6, and 10 in Fig. 12D) the extracellular protein profiles of the cultures were very similar; i.e., additional carbon and nitrogen in the medium did not prevent the release of proteins as it did for M. smegmatis. DNA was readily detected in culture filtrates under both standard conditions and with additional glucose and NH4+ in the medium (data not shown), suggesting that autolysis was also occurring in M. tuberculosis cultures but that it could not be prevented by the addition of glucose and NH4+.
DISCUSSION
The culture filtrate proteins of M. tuberculosis have been the subject of numerous studies (see, for example, references 4, 5, 7, 31, 32, 46, and 50). One often-discussed topic is to what degree autolysis contributes to the presence of antigens in the culture filtrate (for a recent discussion, see reference 69). Culture filtrate proteins have largely been defined as secreted based on their presence in early culture filtrates and/or the lack of an intracellular marker (enzyme activity or antigen) in the culture filtrate, but only a few studies have determined a localization index and/or an extracellular percentage for selected culture filtrate proteins (2, 5, 28–30, 46, 55, 62, 70, 71). Occasionally, when Hsp65 (GroEL, antigen 82) was not present in the culture filtrate, it was assumed that minimal autolysis had occurred (2, 62, 76). However, Wiker et al. noted that this antigen is not very stable (70). In the present study, under conditions in which substantial amounts of recombinant GFPuv were released by M. tuberculosis, Hsp65 was virtually undetectable in culture filtrates despite a high intracellular level (Fig. 11). This suggests that Hsp65 should not be used as an intracellular marker to measure autolysis. There is consensus, however, that most of the major extracellular proteins of M. tuberculosis are encoded by genes containing leader peptide sequences, and they have been shown to be greatly enriched in the culture filtrate (70, 71).
With respect to export of leaderless proteins by M. tuberculosis, we previously showed that M. tuberculosis and other pathogenic mycobacteria export abundant amounts of GS, whereas M. smegmatis and Mycobacterium phlei export small amounts (28). Moreover, recombinant M. tuberculosis GS was abundantly exported by M. smegmatis while very little of the endogenous M. smegmatis GS was released (30). Others have also identified M. tuberculosis GS in culture filtrates (55, 56, 62, 69). In this study, we found a somewhat lower percentage of total GS (8 to 14%) in M. tuberculosis culture filtrates compared with our previous work (33%) and that of Raynaud et al. (17%) (55). We also found, in contrast to our earlier work, that abundantly expressed and active M. tuberculosis GS is not released in large proportions by M. smegmatis. In fact, the M. smegmatis wild-type strain as well as glnA1 strains expressing high levels of the M. smegmatis or M. tuberculosis GS, all exported a similar, relatively low, percentage of total GS (3 to 7%). The large, quantitative differences between these and previous results may be due to the different growth conditions used, as previously strains were grown with added glucose (2%, wt/vol). In this study, we have shown that inclusion of high levels of glucose in 7H9 medium may well result in increased autolysis of M. smegmatis (Fig. 12C, culture 10). It is known that M. tuberculosis undergoes autolysis under low-nitrogen conditions when glucose is present in excess (59), and M. smegmatis may suffer a similar fate. For all conditions and strains examined, the origin of GS was not a major determinant of export.
There are conflicting reports on the export of the M. tuberculosis SodA. Kusunose et al. first reported that M. tuberculosis SodA could be found in substantial amounts in the culture filtrate (39). M. tuberculosis SodA was later identified as a major component of an early culture filtrate by Andersen et al. (5). Also, we previously found that 76% of SodA was extracellular, and Raynaud et al. found 92% of SOD activity to be extracellular (29, 55). However, Nagai et al. noted that SOD was released during a late stage of growth, which they attributed to autolysis, and Alito et al. found low levels of SOD in culture filtrates compared to lysates (2, 46). In the present study, we found a substantially lower amount of SodA (∼20%) in M. tuberculosis culture filtrates compared with our previous work (76%). Several other bacterial pathogens have also been reported to release leaderless SOD into the extracellular environment. N. asteroides, a phylogenetically close relative of the mycobacteria, produces abundant amounts of SodA (2 to 5% of total extractable protein) and releases SodA as a major component of the extracellular medium (1, 9). The M. avium SodA has also been reported to be surface associated and extracellular (19, 24, 42). The Helicobacter pylori SodB is abundantly expressed and has been found to be surface associated and extracellular as well (60, 63).
Several investigators have studied export of recombinant SodAs in M. smegmatis hosts and assessed the proportion of endogenous and recombinant SOD exported. Here too, there have been conflicting reports on export. In our previous study, we found that recombinant M. smegmatis exported a larger proportion of M. tuberculosis SodA than the endogenous enzyme (66 versus 21%) (29). Although Escuyer et al. observed release of recombinant M. avium SodA from M. smegmatis, they did not detect any of the endogenous M. smegmatis enzyme in the culture filtrate (24). Also, Zhang et al. did not detect release of either the endogenous M. smegmatis enzyme or the recombinant M. tuberculosis SodA (76).
Thus, while it was clear from previous studies that the M. tuberculosis SodA was abundantly expressed and sometimes found at high levels in the extracellular medium, it was not clear what the mechanism of export was and whether or not the origin of SOD itself was a major determinant of export, i.e., whether SOD native to a pathogenic species had a greater inherent propensity for export than SOD native to a nonpathogenic species. To explore these issues more definitively, we expressed M. tuberculosis and M. smegmatis SodA, as well as three nonmycobacterial SODs, in an M. smegmatis sodA mutant, and we expressed M. smegmatis SodA and E. coli SodB in wild-type M. tuberculosis. We found that a similar proportion of both recombinant and endogenous mycobacterial SODs were exported in both M. smegmatis and M. tuberculosis. Moreover, nonmycobacterial SODs were released into the medium in proportions similar to those of mycobacterial SODs. This was especially surprising considering that the nonmycobacterial SODs are only 39 to 45% identical to the mycobacterial SODs and their structure is different enough that is precludes mixing of nonmycobacterial and mycobacterial subunits. As for GS, the origin of SOD was not a major determinant of export.
Our finding that the proportion of GS and SOD exported was independent of the bacterial species from which the protein was derived indicated that export was not protein specific. To explore this issue further, we studied three additional leaderless proteins as intracellular markers. We first examined expression and export in M. smegmatis, choosing the M. tuberculosis MDH and coexpressing it at a high level with the recombinant M. smegmatis GlnA1 and SodA. SDS-PAGE and enzymatic analysis clearly showed preferential export of GS and SOD compared with MDH. However, this was due to the unexpected instability of MDH in the culture medium. Although MDH activity was present in the culture filtrate along with GS and SOD at all time points, it was always less than 0.5% of the total MDH activity. Given the very short half-life of MDH in the culture medium, much greater amounts must have leaked out but were degraded. Therefore, a protein noted for its stability (bacterioferritin) was used as an intracellular marker, and in contrast to MDH, it was present in the culture filtrate at levels comparable to those of GS and SOD at all stages of growth, even in early log phase. The amount of extracellular GS and SOD in this experiment was somewhat less than that in the experiment where GS and SOD were coexpressed with MDH. This was typical of the variability observed between cultures and highlights the importance of examining multiple proteins in the same culture when determining intracellular and extracellular location as previously described (70). For the analysis of expression and export in M. tuberculosis, we chose GFP, a nonbacterial protein that lacks a leader peptide, so that there should be no mechanism for its release. GFP, being quite stable, was expected to accumulate in the culture medium if it was released (14). Indeed, GFP was found in the culture filtrate, even in the early stages of growth, and at levels very similar to the endogenous GS and SOD.
Although several export and secretion pathways have been identified in bacteria, including mechanisms to secrete large, folded proteins, export of GS and SOD by M. tuberculosis does not seem to fit any of the models (12, 15, 34, 53, 61, 65). Type I secretion is sec independent (no N-terminal signal peptide) and involves direct secretion of a protein to the extracellular space by an ABC transporter (12). Three proteins are required, and their genes are usually clustered with the gene for the secreted protein. The secretion signal is believed to reside in the C-terminal ∼60 amino acids of the protein, which is preceded by a glycine-rich domain. M. tuberculosis does have many ABC transporters, but sodA and glnA1 are not clustered with any, and SodA and GlnA1 do not contain glycine-rich domains. The general export pathway (type II) is sec dependent and requires an N-terminal signal peptide that is cleaved off as the unfolded protein is exported across the inner membrane (53). In gram-negative bacteria, some specific proteins exported to the periplasm in this manner are then secreted in a folded state across the outer membrane, requiring a large cluster of genes for their secretion. As stated before, GS and SOD do not have N-terminal signal peptides. Autosecreted proteins such as the Neisseria gonorrhoeae immunoglobulin A protease (sometimes referred to as type IV secretion, but different from the type IV secretion discussed below) are exported to the periplasm by the type II pathway and then pass through the outer membrane with no assistance from other proteins (26). In contrast to GS and SOD, autosecreted proteins have N-terminal signal peptides, and only the N-terminal portions of the proteins are translocated across the gram-negative outer membrane; their C-terminal portions contain transmembrane domains that form a pore for translocation (26). Type III secretion has also been identified in gram-negative bacteria and involves the direct transfer of a protein from the bacterial cytoplasm to the cytoplasm of a host cell, although these proteins may accumulate in culture medium as well (34). Secretion is sec independent and requires a cluster of approximately 20 genes. Finally, type IV secretion is similar to type III secretion in that it transports macromolecules (DNA and/or proteins) from the bacterial cytoplasm to a target cell by a contact-dependent mechanism which requires a large cluster of genes (15). No clusters of genes necessary for type III or type IV secretion are present in the M. tuberculosis genome (16).
While the presence of GS and SOD in early culture filtrates from actively growing M. tuberculosis cultures initially suggested a protein-specific export mechanism, the following observations strongly suggest that GS and SOD are released into the culture medium as a result of bacterial leakage or autolysis. (i) The percentage of GS and SOD found in M. tuberculosis and M. smegmatis culture filtrates was quite low, typically less than 20%. (ii) Heterologous SODs, bacterioferritin, and GFP all accumulate in culture filtrates essentially in parallel with the endogenous mycobacterial GSs and SODs. (iii) Metabolic labeling of M. tuberculosis pNBV1-GFPuv showed that GFPuv and GS were slowly released after synthesis, in stark contrast to actively secreted proteins with N-terminal signal peptides. (iv) The presence of high-molecular-weight DNA in culture filtrates was highly correlated with the presence of extracellular proteins. (v) In the case of M. smegmatis, a richer growth medium prevented the release of both proteins and DNA. Recently, studies utilizing highly sensitive two-dimensional PAGE techniques have detected hundreds of proteins in M. tuberculosis culture filtrates, many of them regarded as cytoplasmic (e.g., ribosomal proteins, transcriptional regulators, and elongation factors [36, 56, 62]). It is likely that many of these proteins are also present due to bacterial leakage or limited autolysis of the cultures.
Our study cannot differentiate between autolysis and leakage as a mechanism by which GS and SOD are released. Thus, as an alternative to lysis it is possible that the proteins leak out of viable bacteria as they remodel their cell wall and divide. One observation favoring this is that, in human macrophages infected with M. tuberculosis and stained for GS by the cryosection immunogold technique, extracellular GS is observed in the phagosome of intact healthy-appearing mycobacteria (28). However, it is also a possibility that the extracellular GS came from autolyzed bacteria and was absorbed onto the cell wall of the healthy bacterium before infection of the macrophage. Whether released by autolysis or leakage from intact, viable cells, the extracellular abundance of M. tuberculosis GS and SOD is readily explained by their high expression and extracellular stability.
The presence of leaderless proteins in the culture supernatant fluids of the gram-negative pathogen H. pylori has also been the subject of several papers recently, with some papers claiming that the proteins are specifically secreted and others claiming that the proteins are released by autolysis (38, 52, 60, 67). Interestingly, SodA was one of the H. pylori culture supernatant proteins identified. Also, SodA, as well as some of the other leaderless, extracellular proteins, is expressed in high amounts by H. pylori. Phadnis et al. have proposed that cytoplasmic proteins from H. pylori are released by autolysis, some of which become adsorbed to the outer membrane of intact bacteria (52). They suggest that the amount of protein is variable and depends upon the culture age and subculture status, accumulating on repeatedly subcultured cells. It is possible that a similar phenomenon also occurs in M. tuberculosis cultures.
While autolysis no doubt occurs during the late stages of bacterial growth, the issue here is to what extent it occurs in the early growth phase and under physiologic conditions. Clearly, autolysis can play physiologically important roles in development, genetic exchange, and elimination of defective cells in bacterial cultures and is not necessarily simply the result of a prolonged period in stationary phase (40). Interestingly, Krishnamurthy et al. have proposed that H. pylori undergoes altruistic suicide to release its abundantly expressed urease, which then coats neighboring cells and is protective against an acidic environment (38). While autolysis of a limited proportion of M. tuberculosis bacilli in an otherwise multiplying culture may reflect suboptimal in vitro growth conditions, it is also possible that limited autolysis occurs under physiologic conditions. Hence, we should allow for the possibility that release of proteins by autolysis from a small proportion of the bacteria at a site of bacterial invasion in the tissues may contribute to the overall success of the infection. Viewed in this way, limited autolysis may result in the release of mycobacterial proteins which function as biologically important virulence factors.
ACKNOWLEDGMENTS
This work was supported by grant AI 42925 from the National Institutes of Health.
We thank Chalermchai Chaloyphian for technical assistance and Bai-yu Lee for helpful discussions. We are grateful to Christophe Guilhot and Brigitte Gicquel for supplying the allelic exchange vector, pPR27.
REFERENCES
- 1.Alcendor D J, Chapman G D, Beaman B L. Isolation, sequencing and expression of the superoxide dismutase-encoding gene (sod) of Nocardia asteroides strain GUH-2. Gene. 1995;164:143–147. doi: 10.1016/0378-1119(95)00476-m. [DOI] [PubMed] [Google Scholar]
- 2.Alito A, Romano M I, Bigi F, Zumarraga M, Cataldi A. Antigenic characterization of mycobacteria from South American wild seals. Vet Microbiol. 1999;68:293–299. doi: 10.1016/s0378-1135(99)00107-8. [DOI] [PubMed] [Google Scholar]
- 3.Altschul S F, Madden T L, Schaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Andersen P. Effective vaccination of mice against Mycobacterium tuberculosisinfection with a soluble mixture of secreted mycobacterial proteins. Infect Immun. 1994;62:2536–2544. doi: 10.1128/iai.62.6.2536-2544.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Andersen P, Askgaard D, Ljungqvist L, Bennedsen J, Heron I. Proteins released from Mycobacterium tuberculosisduring growth. Infect Immun. 1991;59:1905–1910. doi: 10.1128/iai.59.6.1905-1910.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Andrews S C. Iron storage in bacteria. Adv Microb Physiol. 1998;40:281–351. doi: 10.1016/s0065-2911(08)60134-4. [DOI] [PubMed] [Google Scholar]
- 7.Arend S M, Geluk A, van Meijgaarden K E, van Dissel J T, Theisen M, Andersen P, Ottenhoff T H. Antigenic equivalence of human T-cell responses to Mycobacterium tuberculosis-specific RD1-encoded protein antigens ESAT-6 and culture filtrate protein 10 and to mixtures of synthetic peptides. Infect Immun. 2000;68:3314–3321. doi: 10.1128/iai.68.6.3314-3321.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bashyam M D, Kaushal D, Dasgupta S K, Tyagi A K. A study of mycobacterial transcriptional apparatus: identification of novel features in promoter elements. J Bacteriol. 1996;178:4847–4853. doi: 10.1128/jb.178.16.4847-4853.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Beaman B L, Scates S M, Moring S E, Deem R, Misra H P. Purification and properties of a unique superoxide dismutase from Nocardia asteroides. J Biol Chem. 1983;258:91–96. [PubMed] [Google Scholar]
- 10.Beauchamp C, Fridovich I. Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem. 1971;44:276–287. doi: 10.1016/0003-2697(71)90370-8. [DOI] [PubMed] [Google Scholar]
- 11.Beyer W, Imlay J, Fridovich I. Superoxide dismutases. Prog Nucleic Acid Res Mol Biol. 1991;40:221–253. doi: 10.1016/s0079-6603(08)60843-0. [DOI] [PubMed] [Google Scholar]
- 12.Binet R, Letoffe S, Ghigo J M, Delepelaire P, Wandersman C. Protein secretion by Gram-negative bacterial ABC exporters–a review. Gene. 1997;192:7–11. doi: 10.1016/s0378-1119(96)00829-3. [DOI] [PubMed] [Google Scholar]
- 13.Bonfield J K, Smith K, Staden R. A new DNA sequence assembly program. Nucleic Acids Res. 1995;23:4992–4999. doi: 10.1093/nar/23.24.4992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chalfie M. Green fluorescent protein. Photochem Photobiol. 1995;62:651–656. doi: 10.1111/j.1751-1097.1995.tb08712.x. [DOI] [PubMed] [Google Scholar]
- 15.Christie P J, Vogel J P. Bacterial type IV secretion: conjugation systems adapted to deliver effector molecules to host cells. Trends Microbiol. 2000;8:354–360. doi: 10.1016/s0966-842x(00)01792-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Cole S T, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Gordon S V, Eiglmeier K, Gas S, Barry III C E, Tekaia F, Badcock K, Basham D, Brown D, Chillingworth T, Connor R, Davies R, Devlin K, Feltwell T, Gentles S, Hamlin N, Holroyd S, Hornsby T, Jagels K, Barrell B G, et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–544. doi: 10.1038/31159. [DOI] [PubMed] [Google Scholar]
- 17.Crabtree B, Leech A R, Newsholme E A. Techniques in metabolic research. B2. Limerick, Ireland: Elsevier/North-Holland Biomedical Press; 1979. Measurement of enzyme activities in crude extracts of tissues; pp. 1–37. [Google Scholar]
- 18.De Groote M A, Ochsner U A, Shiloh M U, Nathan C, McCord J M, Dinauer M C, Libby S J, Vazquez-Torres A, Xu Y, Fang F C. Periplasmic superoxide dismutase protects Salmonella from products of phagocyte NADPH-oxidase and nitric oxide synthase. Proc Natl Acad Sci USA. 1997;94:13997–14001. doi: 10.1073/pnas.94.25.13997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Deshpande R G, Khan M B, Savariar L S, Windham Y Z, Navalkar R G. Superoxide dismutase activity of Mycobacterium avium complex (MAC) strains isolated from AIDS patients. Tuber Lung Dis. 1993;74:305–309. doi: 10.1016/0962-8479(93)90104-6. [DOI] [PubMed] [Google Scholar]
- 20.Dhandayuthapani S, Via L E, Thomas C A, Horowitz P M, Deretic D, Deretic V. Green fluorescent protein as a marker for gene expression and cell biology of mycobacterial interactions with macrophages. Mol Microbiol. 1995;17:901–912. doi: 10.1111/j.1365-2958.1995.mmi_17050901.x. [DOI] [PubMed] [Google Scholar]
- 21.Dussurget O, Rodriguez M, Smith I. Protective role of the Mycobacterium smegmatis IdeR against reactive oxygen species and isoniazid toxicity. Tuber Lung Dis. 1998;79:99–106. doi: 10.1054/tuld.1998.0011. [DOI] [PubMed] [Google Scholar]
- 22.Dussurget O, Stewart G, Neyrolles O, Pescher P, Young D, Marchal G. Role of Mycobacterium tuberculosiscopper-zinc superoxide dismutase. Infect Immun. 2001;69:529–533. doi: 10.1128/IAI.69.1.529-533.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Dye C, Scheele S, Dolin P, Pathania V, Raviglione M C. Consensus statement. Global burden of tuberculosis: estimated incidence, prevalence, and mortality by country. WHO Global Surveillance and Monitoring Project. JAMA. 1999;282:677–686. doi: 10.1001/jama.282.7.677. [DOI] [PubMed] [Google Scholar]
- 24.Escuyer V, Haddad N, Frehel C, Berche P. Molecular characterization of a surface-exposed superoxide dismutase of Mycobacterium avium. Microb Pathog. 1996;20:41–55. doi: 10.1006/mpat.1996.0004. [DOI] [PubMed] [Google Scholar]
- 25.Farrant J L, Sansone A, Canvin J R, Pallen M J, Langford P R, Wallis T S, Dougan G, Kroll J S. Bacterial copper- and zinc-cofactored superoxide dismutase contributes to the pathogenesis of systemic salmonellosis. Mol Microbiol. 1997;25:785–796. doi: 10.1046/j.1365-2958.1997.5151877.x. [DOI] [PubMed] [Google Scholar]
- 26.Finlay B B, Falkow S. Common themes in microbial pathogenicity revisited. Microbiol Mol Biol Rev. 1997;61:136–169. doi: 10.1128/mmbr.61.2.136-169.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Fridovich I. Fundamental aspects of reactive oxygen species, or what's the matter with oxygen? Ann N Y Acad Sci. 1999;893:13–18. doi: 10.1111/j.1749-6632.1999.tb07814.x. [DOI] [PubMed] [Google Scholar]
- 28.Harth G, Clemens D L, Horwitz M A. Glutamine synthetase of Mycobacterium tuberculosis: extracellular release and characterization of its enzymatic activity. Proc Natl Acad Sci USA. 1994;91:9342–9346. doi: 10.1073/pnas.91.20.9342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Harth G, Horwitz M A. Export of recombinant Mycobacterium tuberculosis superoxide dismutase is dependent upon both information in the protein and mycobacterial export machinery. A model for studying export of leaderless proteins by pathogenic mycobacteria. J Biol Chem. 1999;274:4281–4292. doi: 10.1074/jbc.274.7.4281. [DOI] [PubMed] [Google Scholar]
- 30.Harth G, Horwitz M A. Expression and efficient export of enzymatically active Mycobacterium tuberculosis glutamine synthetase in Mycobacterium smegmatis and evidence that the information for export is contained within the protein. J Biol Chem. 1997;272:22728–22735. doi: 10.1074/jbc.272.36.22728. [DOI] [PubMed] [Google Scholar]
- 31.Harth G, Horwitz M A. An inhibitor of exported Mycobacterium tuberculosis glutamine synthetase selectively blocks the growth of pathogenic mycobacteria in axenic culture and in human monocytes: extracellular proteins as potential novel drug targets. J Exp Med. 1999;189:1425–1436. doi: 10.1084/jem.189.9.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Horwitz M A, Lee B W, Dillon B J, Harth G. Protective immunity against tuberculosis induced by vaccination with major extracellular proteins of Mycobacterium tuberculosis. Proc Natl Acad Sci USA. 1995;92:1530–1534. doi: 10.1073/pnas.92.5.1530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Howard N S, Gomez J E, Ko C, Bishai W R. Color selection with a hygromycin-resistance-based Escherichia coli-mycobacterial shuttle vector. Gene. 1995;166:181–182. doi: 10.1016/0378-1119(95)00597-x. [DOI] [PubMed] [Google Scholar]
- 34.Hueck C J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol Mol Biol Rev. 1998;62:379–433. doi: 10.1128/mmbr.62.2.379-433.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Inaoka T, Matsumura Y, Tsuchido T. Molecular cloning and nucleotide sequence of the superoxide dismutase gene and characterization of its product from Bacillus subtilis. J Bacteriol. 1998;180:3697–3703. doi: 10.1128/jb.180.14.3697-3703.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Jungblut P R, Schaible U E, Mollenkopf H J, Zimny-Arndt U, Raupach B, Mattow J, Halada P, Lamer S, Hagens K, Kaufmann S H. Comparative proteome analysis of Mycobacterium tuberculosis and Mycobacterium bovis BCG strains: towards functional genomics of microbial pathogens. Mol Microbiol. 1999;33:1103–1117. doi: 10.1046/j.1365-2958.1999.01549.x. [DOI] [PubMed] [Google Scholar]
- 37.Kremer L, Baulard A, Estaquier J, Poulain-Godefroy O, Locht C. Green fluorescent protein as a new expression marker in mycobacteria. Mol Microbiol. 1995;17:913–922. doi: 10.1111/j.1365-2958.1995.mmi_17050913.x. [DOI] [PubMed] [Google Scholar]
- 38.Krishnamurthy P, Parlow M, Zitzer J B, Vakil N B, Mobley H L, Levy M, Phadnis S H, Dunn B E. Helicobacter pyloricontaining only cytoplasmic urease is susceptible to acid. Infect Immun. 1998;66:5060–5066. doi: 10.1128/iai.66.11.5060-5066.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kusunose E, Ichihara K, Noda Y, Kusunose M. Superoxide dismutase from Mycobacterium tuberculosis. J Biochem (Tokyo) 1976;80:1343–1352. doi: 10.1093/oxfordjournals.jbchem.a131407. [DOI] [PubMed] [Google Scholar]
- 40.Lewis K. Programmed death in bacteria. Microbiol Mol Biol Rev. 2000;64:503–514. doi: 10.1128/mmbr.64.3.503-514.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Loo D T, Rillema J R. Measurement of cell death. Methods Cell Biol. 1998;57:251–264. doi: 10.1016/s0091-679x(08)61583-6. [DOI] [PubMed] [Google Scholar]
- 42.Mayer B K, Falkinham J O I. Superoxide dismutase activity of Mycobacterium avium, M. intracellulare, and M. scrofulaceum. Infect Immun. 1986;53:631–635. doi: 10.1128/iai.53.3.631-635.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Menard R, Sansonetti P J, Parsot C. Nonpolar mutagenesis of the ipa genes defines IpaB, IpaC, and IpaD as effectors of Shigella flexnerientry into epithelial cells. J Bacteriol. 1993;175:5899–5906. doi: 10.1128/jb.175.18.5899-5906.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Menendez M C, Domenech P, Prieto J, Garcia M J. Cloning and expression of the Mycobacterium fortuitum superoxide dismutase gene. FEMS Microbiol Lett. 1995;134:273–278. doi: 10.1111/j.1574-6968.1995.tb07950.x. [DOI] [PubMed] [Google Scholar]
- 45.Musrati R A, Kollarova M, Mernik N, Mikulasova D. Malate dehydrogenase: distribution, function and properties. Gen Physiol Biophys. 1998;17:193–210. [PubMed] [Google Scholar]
- 46.Nagai S, Wiker H G, Harboe M, Kinomoto M. Isolation and partial characterization of major protein antigens in the culture fluid of Mycobacterium tuberculosis. Infect Immun. 1991;59:372–382. doi: 10.1128/iai.59.1.372-382.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Neuhoff V, Arold N, Taube D, Ehrhardt W. Improved staining of proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis. 1988;9:255–262. doi: 10.1002/elps.1150090603. [DOI] [PubMed] [Google Scholar]
- 48.Norrander J, Kempe T, Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983;26:101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- 49.O'Gaora P, Barnini S, Hayward C, Filley E, Rook G, Young D, Thole J. Mycobacteria as immunogens: development of expression vectors for use in multiple mycobacterial species. Med Principles Pract. 1997;6:91–96. [Google Scholar]
- 50.Pal P G, Horwitz M A. Immunization with extracellular proteins of Mycobacterium tuberculosisinduces cell-mediated immune responses and substantial protective immunity in a guinea pig model of pulmonary tuberculosis. Infect Immun. 1992;60:4781–4792. doi: 10.1128/iai.60.11.4781-4792.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Pelicic V, Jackson M, Reyrat J M, Jacobs W R, Jr, Gicquel B, Guilhot C. Efficient allelic exchange and transposon mutagenesis in Mycobacterium tuberculosis. Proc Natl Acad Sci USA. 1997;94:10955–10960. doi: 10.1073/pnas.94.20.10955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Phadnis S H, Parlow M H, Levy M, Ilver D, Caulkins C M, Connors J B, Dunn B E. Surface localization of Helicobacter pyloriurease and a heat shock protein homolog requires bacterial autolysis. Infect Immun. 1996;64:905–912. doi: 10.1128/iai.64.3.905-912.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Pugsley A P. The complete general secretory pathway in gram-negative bacteria. Microbiol Rev. 1993;57:50–108. doi: 10.1128/mr.57.1.50-108.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ratledge C. Nutrition, growth and metabolism. In: Ratledge C, Stanford J, editors. The biology of the mycobacteria. London, United Kingdom: Academic Press; 1982. pp. 185–271. [Google Scholar]
- 55.Raynaud C, Etienne G, Peyron P, Laneelle M A, Daffe M. Extracellular enzyme activities potentially involved in the pathogenicity of Mycobacterium tuberculosis. Microbiology. 1998;144:577–587. doi: 10.1099/00221287-144-2-577. [DOI] [PubMed] [Google Scholar]
- 56.Rosenkrands I, Weldingh K, Jacobsen S, Hansen C V, Florio W, Gianetri I, Andersen P. Mapping and identification of Mycobacterium tuberculosis proteins by two-dimensional gel electrophoresis, microsequencing and immunodetection. Electrophoresis. 2000;21:935–948. doi: 10.1002/(SICI)1522-2683(20000301)21:5<935::AID-ELPS935>3.0.CO;2-P. [DOI] [PubMed] [Google Scholar]
- 57.Sambrook J, Maniatis T, Fritsch E F. Molecular cloning: a laboratory manual. 2nd ed. Vol. 3. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory; 1989. [Google Scholar]
- 58.San Mateo L R, Toffer K L, Orndorff P E, Kawula T H. Neutropenia restores virulence to an attenuated Cu,Zn superoxide dismutase-deficient Haemophilus ducreyistrain in the swine model of chancroid. Infect Immun. 1999;67:5345–5351. doi: 10.1128/iai.67.10.5345-5351.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Schaefer W B, Marshak A, Burkhart B. The growth of Mycobacterium tuberculosisas a function of its nutrients. J Bacteriol. 1949;58:549–563. doi: 10.1128/jb.58.5.549-563.1949. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Schraw W, McClain M S, Cover T L. Kinetics and mechanisms of extracellular protein release by Helicobacter pylori. Infect Immun. 1999;67:5247–5252. doi: 10.1128/iai.67.10.5247-5252.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Simonen M, Palva I. Protein secretion in Bacillusspecies. Microbiol Rev. 1993;57:109–137. doi: 10.1128/mr.57.1.109-137.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Sonnenberg M G, Belisle J T. Definition of Mycobacterium tuberculosisculture filtrate proteins by two-dimensional polyacrylamide gel electrophoresis, N-terminal amino acid sequencing, and electrospray mass spectrometry. Infect Immun. 1997;65:4515–4524. doi: 10.1128/iai.65.11.4515-4524.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Spiegelhalder C, Gerstenecker B, Kersten A, Schiltz E, Kist M. Purification of Helicobacter pylorisuperoxide dismutase and cloning and sequencing of the gene. Infect Immun. 1993;61:5315–5325. doi: 10.1128/iai.61.12.5315-5325.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Thompson J D, Gibson T J, Plewniak F, Jeanmougin F, Higgins D G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25:4876–4882. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Tjalsma H, Bolhuis A, Jongbloed J D H, Bron S, van Dijl J M. Signal peptide-dependent protein transport in Bacillus subtilis: a genome-based survey of the secretome. Microbiol Mol Biol Rev. 2000;64:515–547. doi: 10.1128/mmbr.64.3.515-547.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Ukeda H, Maeda S, Ishii T, Sawamura M. Spectrophotometric assay for superoxide dismutase based on tetrazolium salt 3′-1-(phenylamino)-carbonyl-3,4-tetrazolium]-bis(4-methoxy-6-nitro)benzenesulfonic acid hydrate reduction by xanthine-xanthine oxidase. Anal Biochem. 1997;251:206–209. doi: 10.1006/abio.1997.2273. [DOI] [PubMed] [Google Scholar]
- 67.Vanet A, Labigne A. Evidence for specific secretion rather than autolysis in the release of some Helicobacter pyloriproteins. Infect Immun. 1998;66:1023–1027. doi: 10.1128/iai.66.3.1023-1027.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Wards B J, Collins D M. Electroporation at elevated temperatures substantially improves transformation efficiency of slow-growing mycobacteria. FEMS Microbiol Lett. 1996;145:101–105. doi: 10.1111/j.1574-6968.1996.tb08563.x. [DOI] [PubMed] [Google Scholar]
- 69.Weldingh K, Hansen A, Jacobsen S, Andersen P. High resolution electroelution of polyacrylamide gels for the purification of single proteins from Mycobacterium tuberculosis culture filtrate. Scand J Immunol. 2000;51:79–86. doi: 10.1046/j.1365-3083.2000.00655.x. [DOI] [PubMed] [Google Scholar]
- 70.Wiker H G, Harboe M, Nagai S. A localization index for distinction between extracellular and intracellular antigens of Mycobacterium tuberculosis. J Gen Microbiol. 1991;137:875–884. doi: 10.1099/00221287-137-4-875. [DOI] [PubMed] [Google Scholar]
- 71.Wiker H G, Michell S L, Hewinson R G, Spierings E, Nagai S, Harboe M. Cloning, expression and significance of MPT53 for identification of secreted proteins of Mycobacterium tuberculosis. Microb Pathog. 1999;26:207–219. doi: 10.1006/mpat.1998.0267. [DOI] [PubMed] [Google Scholar]
- 72.Wilks K E, Dunn K L, Farrant J L, Reddin K M, Gorringe A R, Langford P R, Kroll J S. Periplasmic superoxide dismutase in meningococcal pathogenicity. Infect Immun. 1998;66:213–217. doi: 10.1128/iai.66.1.213-217.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Woolfolk C A, Shapiro B, Stadtman E R. Regulation of glutamine synthetase. I. Purification and properties of glutamine synthetase from Escherichia coli. Arch Biochem Biophys. 1966;116:177–192. doi: 10.1016/0003-9861(66)90026-9. [DOI] [PubMed] [Google Scholar]
- 74.Wu C H, Tsai-Wu J J, Huang Y T, Lin C Y, Lioua G G, Lee F J. Identification and subcellular localization of a novel Cu,Zn superoxide dismutase of Mycobacterium tuberculosis. FEBS Lett. 1998;439:192–196. doi: 10.1016/s0014-5793(98)01373-8. [DOI] [PubMed] [Google Scholar]
- 75.Yanisch-Perron C, Vieira J, Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33:103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- 76.Zhang Y, Lathigra R, Garbe T, Catty D, Young D. Genetic analysis of superoxide dismutase, the 23 kilodalton antigen of Mycobacterium tuberculosis. Mol Microbiol. 1991;5:381–391. doi: 10.1111/j.1365-2958.1991.tb02120.x. [DOI] [PubMed] [Google Scholar]