Abstract
Background
Differences in the composition and diversity of the gut microbial communities among individuals are influenced by environmental factors. However, there is limited research on factors affecting microbiome variation in colorectal cancer patients, who display lower inter-individual variations than that of healthy individuals. In this study, we examined the association between modifiable factors and the microbiome variation in colorectal cancer patients.
Methods
A total of 331 colorectal cancer patients who underwent resection surgery at the Department of Surgery, Seoul National University Hospital between October 2017 and August 2019 were included. Fecal samples from colorectal cancer patients were collected prior to the surgery. Variations in the gut microbiome among patients with different lifestyles and metabolic diseases were examined through the network analysis of inter-connected microbial abundance, the assessment of the Anna Karenina principle effect for microbial stochasticity, and the identification of the enriched bacteria using linear discrimination analysis effect size. Associations of dietary diversity with microbiome variation were investigated using the Procrustes analysis.
Results
We found stronger network connectivity of microbial communities in non-smokers, non-drinkers, obese individuals, hypertensive subjects, and individuals without diabetes than in their counterparts. The Anna Karenina principle effect was found for history of smoking, alcohol consumption, and diabetes (with significantly greater intra-sample similarity index), whereas obesity and hypertension showed the anti-Anna Karenina principle effect (with significantly lower intra-sample similarity index). We found certain bacterial taxa to be significantly enriched in patients of different categories of lifestyles and metabolic diseases using linear discrimination analysis. Diversity of food and nutrient intake did not shape the microbial diversity between individuals (pProcrustes>0.05).
Conclusions
Our findings suggested an immune dysregulation and a reduced ability of the host and its microbiome in regulating the community composition. History of smoking, alcohol consumption, and diabetes were shown to affect partial individuals in shifting new microbial communities, whereas obesity and history of hypertension appeared to affect majority of individuals and shifted to drastic reductions in microbial compositions. Understanding the contribution of modifiable factors to microbial stochasticity may provide insights into how the microbiome regulates effects of these factors on the health outcomes of colorectal cancer patients.
Supplementary Information
The online version contains supplementary material available at 10.1186/s12866-023-02771-7.
Keywords: Anna Karenina principle, Dysbiosis, Colorectal cancer, Lifestyle, Metabolic disease, Diet
Introduction
Given the complex and dynamic nature of the microbial community in the gastrointestinal tract, the gut microbiome can actively interact with immune cells through the intestinal mucosal surface, and provide the host with colonization resistance against foreign microbes [1–3]. When the bacterial homeostasis is disrupted, there is an imbalance between the commensal and pathogenic bacteria in the gut that can lead to the formation of inflammatory biomarkers and stimulate the carcinogenesis process [1, 4]. This condition is called dysbiosis, which is determined to involve several physiological processes, such as inflammation, pathogenic bacteria, genotoxins, oxidative stress, metabolites, and biofilm [5].
To date, several researchers have proposed using an Anna Karenina principle (AKP) effect to describe the increase of stochastic transitions from stable to unstable states in the gut microbiome [6]. Regarding microbiome-associated factors, the AKP has been called “all healthy microbiomes are similar; each dysbiotic microbiome is dysbiotic in its own way” [6]. The AKP effect, therefore, indicates the increase of microbiome heterogeneity or stochasticity related to dysbiosis due to abnormal conditions and the high personalization of factor-associated microbial communities [7].
Differences in the composition and diversity of the gut microbial communities among individuals are contributed by environmental factors [8, 9]. Lifestyle, diet, and chronic disease have been generally described to affect specific components of the gut microbiome [10]. Under some conditions of immune system dysfunctions, there could be a rise of stochasticity and the dysbiotic communities much more veried from person to person [7]. However, the composition of the microbiome community in colorectal cancer (CRC) patients differs from the core microbiome and diversity levels of healthy individuals [11, 12]. More research is required to identify how modifiable factors affect the microbial instability in CRC patients and thus understand how the gut microbiome may regulate the effect of modifiable factors on the health outcomes of CRC patients.
In CRC patients, metabolic syndrome and lifestyle behaviors have been shown to contribute to the patient's prognosis [13, 14]. In addition, dietary factors can modify the gut microbial community via energy harvesting and several diet-derived metabolites, such as short-chain fatty acids (SCFAs) [15].11 Although several data-driven approaches are available for the determination of a person’s dietary patterns [16], individuals may differ not only in the specific food items they eat but also in their dietary diversity [17]. However, such a tree-based approach has not been applied to the habitual diets of the Korean population.
In this study, we first created the hierarchical tree of foods to reflect dietary diversity and investigated its associations with lifestyle factors and metabolic diseases. Then, we examined the association of lifestyle factors, metabolic disease, and dietary intake with the gut microbiome variation. We hypothesized that CRC participants with unhealthy lifestyles, such as smoking and alcohol consumption, and metabolic diseases may associate with a higher level of microbiome stochasticity.
Methods
Healthy lifestyles and non-metabolic diseases in the majority of study participants
The study was designed as a cross-sectional study and included study subjects who were diagnosed with CRC and underwent resection surgery between October 2017 and August 2019 in the Department of Surgery, Seoul National University Hospital, Seoul, Korea. Among the selected CRC patients, a total of 331 patients were included in this study after excluding those who could not be analyzed due to the absence or small amount of fecal sample prior to the surgery. Of these, 115 patients provided their dietary information.
Demographic characteristics, lifestyle behaviors, and metabolic diseases of CRC patients are shown in Table 1. The mean age of study participants was 61.9 years, with 14.8% of patients being early-onset (age ≤50 years old) cases (n=49). Of the included patients, 19.6% were ever smokers (n=65), 38.1% ever consumed alcohol (n=126), 40.5% were obese (BMI≥25.0 kg/m2) (n=134), 39.3% had hypertension (n=130), and and 21.1% had diabetes (n=70). Patient demographics, family history of CRC, neoadjuvant therapy, lifestyles, and metabolic diseases did not differ between those with and without dietary information (p>0.05).
Table 1.
Characteristics of study participants
| Factor | Total (N=331) | With dietary data (N=115) | Without dietary data (N=216) | P-value |
|---|---|---|---|---|
| Age (mean ± sd, years) | 61.9 ± 11.0 | 60.8 ± 11.8 | 62.4 ± 10.5 | 0.21 |
| ≤50 | 49 (14.8) | 23 (20.0) | 26 (12.0) | 0.23 |
| 50-≤60 | 94 (28.4) | 31 (27.0) | 63 (29.2) | |
| 60-≤70 | 118 (35.6) | 36 (31.3) | 82 (38.0) | |
| >70 | 70 (21.1) | 25 (21.7) | 45 (20.8) | |
| Sex | ||||
| Female | 122 (36.9) | 41 (35.7) | 81 (37.5) | 0.83 |
| Male | 209 (63.1) | 74 (64.3) | 135 (62.5) | |
| Family history of CRC | ||||
| No | 291 (87.9) | 105 (91.3) | 186 (86.1) | 0.23 |
| Yes | 40 (12.1) | 10 (8.7) | 30 (13.9) | |
| Neoadjuvant therapy | ||||
| No | 290 (87.6) | 104 (90.4) | 186 (86.1) | 0.34 |
| Chemotherapy and/or radiotherapy | 41 (12.4) | 11 (9.6) | 30 (13.9) | |
| Smoking status | ||||
| Never | 266 (80.4) | 92 (80.0) | 174 (80.6) | >0.99 |
| Ever | 65 (19.6) | 23 (20.0) | 42 (19.4) | |
| Alcohol consumption | ||||
| Never | 205 (61.9) | 74 (64.3) | 131 (60.6) | 0.59 |
| Ever | 126 (38.1) | 41 (35.7) | 85 (39.4) | |
| BMI (mean ± sd, kg/m2) | 24.4 ± 3.2 | 24.6 ± 3.3 | 24.3 ± 3.2 | 0.44 |
| Normal (<25.0) | 197 (59.5) | 38 (33.0) | 136 (63.0) | 0.10 |
| Obesity (≥25.0) | 134 (40.5) | 77 (67.0) | 80 (37.0) | |
| Hypertension | ||||
| No | 201 (60.7) | 75 (65.2) | 126 (58.3) | 0.27 |
| Yes | 130 (39.3) | 40 (34.8) | 90 (41.7) | |
| Diabetes | ||||
| No | 261 (78.9) | 94 (81.7) | 167 (77.3) | 0.43 |
| Yes | 70 (21.1) | 21 (18.3) | 49 (22.7) | |
Data are presented as mean ± standard deviation for continuous variables and counts (percentages) for categorical variables
P-values are calculated from a t-test for continuous variables and a Chi-square test for categorical variables
CRC Colorectal cancer, BMI Body mass index
Collection of lifestyles, dietary habits, and clinical data
On the study enrollment, we obtained patient information on age, sex, family history of CRC, neoadjuvant therapy, tobacco smoking and alcohol consumption experiences, and history of hypertension and diabetes. Additionally, the height and weight of the patients were measured and used to calculate the body mass index (BMI). After surgery, the disease stage was further assessed following the American Joint Committee on Cancer (AJCC) criteria. The average amount of diet consumption during the preceding year was assessed using validated a semi-quantitative food frequency questionnaire (SQFFQ), which was developed by the Korea Centers for Disease Control and Prevention [18]. By using the Computer-Aided Nutritional Analysis Program (CAN-Pro) 4.0 (Korean Nutrition Information Center, Seoul, Korea), we estimated the weight intake of macro- and micronutrients from 663 food subitems, which were generated from 106 food items in the SQFFQ. From this, 35 food groups were generated, which has been applied in previous studies [19]. The CAN-Pro software further identified the higher level with 17 food groups [20] and we determined the highest level with the plant-based foods, animal-based foods, beverages, and condiments. Details for the components of the tree-based diet are available in (Additional File 1: eTable1).
Fecal sample collection and 16S rRNA sequencing process
In this study, participants received a kit (DNeasyPowerSoil Kit, Qiagen, Hilden, Germany) and collected a single stool sample according to the kit instructions. The sample was collected before the operation date, and fecal microbiota was analyzed using 16S rRNA gene amplicon sequencing with V3-V4 primers. The first polymerase chain reaction (PCR) product was purified with AMPure beads (Agencourt Bioscience, Beverly, MA). Following purification, 2μl of the first PCR product was PCR amplified for final library construction containing the index using NexteraXT Indexed Primer. The cycle conditions for the second PCR were the same as the first PCR except for 10 cycles. The final PCR product was purified with AMPure beads and then quantified using qPCR according to the qPCR Quantification Protocol Guide (KAPA Library Quantificatoin kits for IlluminaSequecing platforms) and qualified using the TapeStation D1000 ScreenTape (Agilent Technologies, Waldbronn, Germany). The paired-end (2×300 bp) sequencing was performed by the Macrogen using the MiSeq™ platform (Illumina, San Diego, USA). The number of operational taxonomic units was identified by utilizing the preprocessed reads of samples and clustering the sequences from samples using a 97% sequence identity cut-off.
Statistical analysis
Lifestyle factors, metabolic diseases, and microbial variation
The interaction of microbiome relative abundance was visualized using a network analysis approach. Given the compositional and zero-inflated properties of the microbiome data, numerous correlation-focused approaches have been developed to overcome the difficulty of inferring dependencies in microbial data, such as CCREPE, SparCC, CCLasso, and REBACCA [21, 22]. However, these methods may be limited in reflecting indirect relationships and causing spurious associations from the creation of pseudo-counts [21, 22]. Ha et al. introduced a COpositional Zero-Inflated Network Estimation (COZINE) to address this challenge by generating a binary incidence matrix and a compositional abundance matrix in which the centered log-ratio transformation can be applied for non-zero counts only [21, 22]. In this study, the network structure was constructed for each group of smoking status, alcohol consumption, BMI, and underlying diseases using the COZINE method (R packages ‘COZINE’ and ‘HurdleNormal’) [21, 22].
For the estimation of the AKP effect on the increase of microbial stochasticity associated with dysbiosis due to the external factor, Ning et al. recently developed a framework to assess and present the ecological stochasticity as a single index, which is called normalized stochasticity ratio [23]. Borrowing the Ružička similarity concept from this framework, we calculated the intra-sample similarity (C) index to reflect the similarity in microbiome composition of individuals in each group according to exposure status [7]. In general, if the C-index of the exposed group was higher than that of the non-exposed group, it implied presence of the AKP effect according to that exposed factor and the higher stochasticity or heterogeneity in the exposed group [7]. The C-indexes of groups were then compared using a Wilcoxon test, with the level of significance defined as p<0.05.
Furthermore, we conducted the linear discrimination analysis effect size (LEfSe) to identify bacteria that are phylogenetically abundant in each group of lifestyle factors and metabolic diseases.
Dietary diversity and microbial variation
Our previous study identified several dietary factors that were correlated with the relative abundance of several taxon [24], however, whether the overall diversity of source food intake reflecting differences of microbiome composition among CRC patients remained unclear. Given an estimate of almost 10% of dietary energy was obtained due to microbial fermentation and volatile fatty acid production [25], we used a tree-based approach to assess the dietary diversity of energy intake and its components in associations with microbiome variations. The assessment of dietary diversity included considerations of the consumption of energy intake (kcal/day), plant/animal protein (g/day), plant/animal fat (g/day), carbohydrate (g/day), and fiber (g/day). Based on the tree-based structure of dietary intake, we calculated Chao1, Shannon, and Simpson indices for within-subject (alpha)-diversity, which reflected the overall diversity of food source consumption within each patient. In addition, to capture the variation from patient to patient in terms of dietary intake composition, we calculated unweighted and weighted UniFrac distances for between-subject (beta)-diversity of dietary intake (R packages ‘vegan’ and ‘GuniFrac’).
To test for the association of dietary diversity with microbiome variation across subjects, we performed the Procrustes analysis to compare the shapes of two beta-diversity matrixes by translating, rotating, and uniformly scaling the matrixes (R package ‘ape’).
Dietary diversity in associations with lifestyle factors and metabolic diseases
To explore whether the AKP or anti-AKP effect of lifestyle factors and metabolic diseases might be attributed by the dietary diversity,we examined the difference of dietary diversity indices according to lifestyle factors and metabolic diseases. Thus, we applied the generalized linear model to investigate the association of alpha-diversity and the permutational multivariate analysis of variance (PERMANOVA) test to investigate the association of beta-diversity of diet consumption with lifestyle factors and metabolic diseases. Significant differences in the alpha- and beta-diversity according to lifestyle factors and metabolic diseases were visualized as box plot and principal coordinate analysis plots.
The LEfSe analysis was performed in the Galaxy web application (https://huttenhower.sph.harvard.edu/galaxy/) and other statistical analyses were performed in R 3.6.0.
Results
Lifestyle factors and metabolic diseases and microbial variation
Network structure for the partial correlation of the gut microbiome in CRC
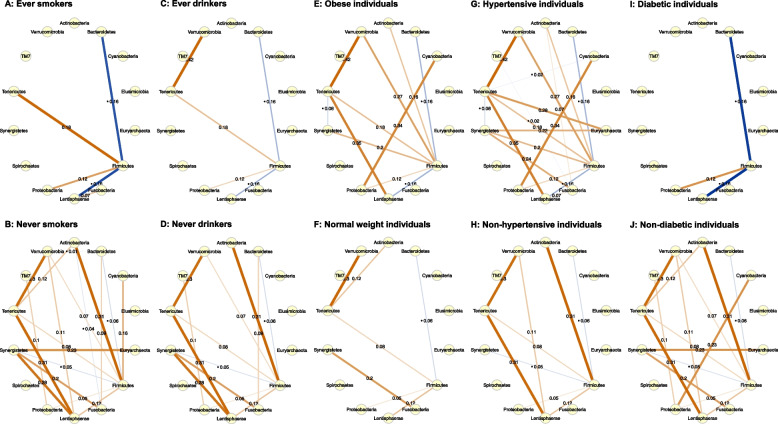
The Spearman partial correlation between phyla across different population groups is shown in Figs. 1A-1J. Significant non-zero edges were identified in the COZINE framework. The networks of phylum abundance in the exposed group of smoking status, alcohol consumption, or diabetes were relatively sparse compared to the non-exposed group, whereas the networks of phylum abundance in the obese or hypertensive group were relatively dense compared to the counterpart. In these groups of patients with more connected microbial communities (never smokers, never drinkers, obese, hypertensive, and non-diabetic individuals), there were strong abundance correlations between Tenericutes and Verrucomicrobia (ρ=0.30 to ρ=0.42), Tenericutes and Lentisphaerae (ρ=0.31 to ρ=0.35). Nevertheless, abundances between Bacteroidetes and Firmicutes phyla, which were the two most abundant phyla, were negatively correlated (ρ=-0.16 to ρ=-0.06). Of all communities, Firmicutes appeared to be the central phylum, which mostly connected with other phyla in the networks.
Fig. 1.
COpositional Zero-Inflated Network Estimation (COZINE) identifies network structure of phylum abundance in (A) ever smoking, (B) never smoking, (C) ever drinking, (D) never drinking, (E) obese, (F) normal weight, (G) hypertensive, (H) non-hypertensive, (I) diabetic, and (J) non-diabetic individuals. Nodes represent the abundance of phyla, and edges represent the partial correlation coefficient between phyla. Brown lines show positive partial correlations, and blue lines show negative partial correlations. The thickness of the edges is proportional to their partial correlations
Assessment of the Anna Karenina principle effect
Table 2 presents the summary statistics of medians and interquartile ranges and p-values from the Wilcoxon test for the difference in intra-sample similarity (C) index. Accordingly, smoking, drinking, and diabetes exhibited the AKP effect (p<0.05 for the null hypothesis that C index in the exposed group is higher than in the non-exposed group), whereas obesity and hypertension exhibit the anti-AKP effect (p<0.05 for the null hypothesis of C index in the exposed group is less than in the non-exposed group). This suggested different responses among individuals who had history of tobacco smoking, alcohol consumption, and diabetes, that not all individuals showed shifts to new microbial compositions, as a result, resulted in an increase in beta-diversity. Under conditions of obesity and hypertension, all individuals were affected and showed shifts to new microbial compositions which were similar from person to person, which resulted in an excessive reduction in microbial compositions and lower beta-diversity compared to their counterparts.
Table 2.
Intra-sample similarity index and Wilcoxon test for the detection of AKP effects of lifestyle factors and metabolic diseases
| Factor | Exposed (E) group a | Non-exposed (NE) group a | P-value (E≠NE) b | P-value (E>NE) c | P-value (E<NE) d |
|---|---|---|---|---|---|
| Smoking status | 0.812 (0.695-0.877) | 0.790 (0.690-0.870) | 0.001 | <0.001 | >0.99 |
| Alcohol consumption | 0.815 (0.709-0.882) | 0.784 (0.689-0.861) | <0.001 | <0.001 | >0.99 |
| Obesity | 0.778 (0.679-0.855) | 0.807 (0.705-0.877) | <0.001 | >0.99 | <0.001 |
| Hypertension | 0.782 (0.678-0.861) | 0.803 (0.705-0.874) | <0.001 | >0.99 | <0.001 |
| Diabetes | 0.804 (0.694-0.889) | 0.790 (0.690-0.870) | <0.001 | <0.001 | >0.99 |
aData are presented as median (interquartile range)
bP-value less than 0.05 indicates the presence of AKP or anti-AKP effect
cP-value less than 0.05 indicates AKP effect
dP-value less than 0.05 indicates anti-AKP effect
Differentially abundant bacteria
Bacteria that were highly enriched in individuals who have smoked on at least one occasion or have never smoked, who have consumed alcohol on at least one occasion or have never consumed alcohol, who are obese or are of normal weight, who have hypertension or do not have hypertension, and who are diabetic or non-diabetic are presented in (Additional File 1: eFigure1-5). The list of these taxa at different phylum, class, order, family, genus, and species levels is summarized in Table 3. Taxa related to class Bacilli were enriched in smokers, whereas taxa related to order Desulfovibrionales and Synergistales were enriched in non-smokers. Additionally, taxa related to family Micrococcaceae, Enterococcaceae, and Enterobacteriaceae were enriched in non-drinkers, and taxa related to class Betaproteobacteria was enriched in obese individuals. In terms of metabolic diseases, taxa related to phylum Elusimicrobia was enriched in individuals with hypertension and diabetes.
Table 3.
Abundant bacteria identified by linear discriminant analysis effect size analysis in different statuses of lifestyles and metabolic diseases
| Factor | Phylum | Class | Order | Family | Genus | Species |
|---|---|---|---|---|---|---|
| Smoking status | ||||||
| Never | Synergistetes |
Deltaproteobacteria Synergistia |
Desulfovibrionales Synergistales |
Desulfovibrionaceae Synergistaceae |
Bilophila Synergistes |
Desulfovibrio D168 Bilophila sp. Synergistes sp. |
| Ever | Bacilli | Lactobacillales | Streptococcaceae |
Streptococcus Hafnia |
Streptococcus sp. Hafnia alvei |
|
| Alcohol consumption | ||||||
| Never | rc4_4 | rc4_4 sp. | ||||
| Ever | Gammaproteobacteria | Enterobacteriales |
Micrococcaceae Enterobacteriaceae Enterococcaceae |
Rothia Citrobacter Enterococcus |
Citrobacter sp. Enterococcus sp. |
|
| Body mass index | ||||||
| <25.0 kg/m2 | SHA_98 | Fusobacterium |
Ruminococcus albus Peptostreptococcus anaerobius Streptococcus agalactiae Fusobacterium sp. |
|||
| ≥25.0 kg/m2 | Betaproteobacteria | Burkholderiales |
Lachnospiraceae Alcaligenaceae |
Faecalibacterium Sutterella |
Bacteroides coprophilus Clostridium baratii Faecalibacterium prausnitzi Blautia obeum Sutterella sp. |
|
| Hypertension | ||||||
| No | Eubacterium | Eubacterium biforme | ||||
| Yes | Elusimicrobia Synergisteles |
Elusimicrobia Synergistia |
Elusimicrobiales Synergistales |
Elusimicrobiaceae | Desulfovibrio sp. | |
| Diabetes | ||||||
| No | Synergisteles | Synergistia | Synergistales | S24_7 |
Ruminococcus Barnesiella |
Desulfovibrio D168 Prevotella sp. Ruminococcus lactaris Barnesiella intesttinihominis |
| Yes | Elusimicrobia | Elusimicrobia |
Lactobacillales Elusimicrobiales |
Peptostreptococcaceae Streptococcaceae Elusimicrobiaceae |
Veillonella Streptococcus |
Desulfovibrio sp. Ruminococcus gnavus Veillonella dispar Streptococcus sp. |
Dietary diversity and microbial variation
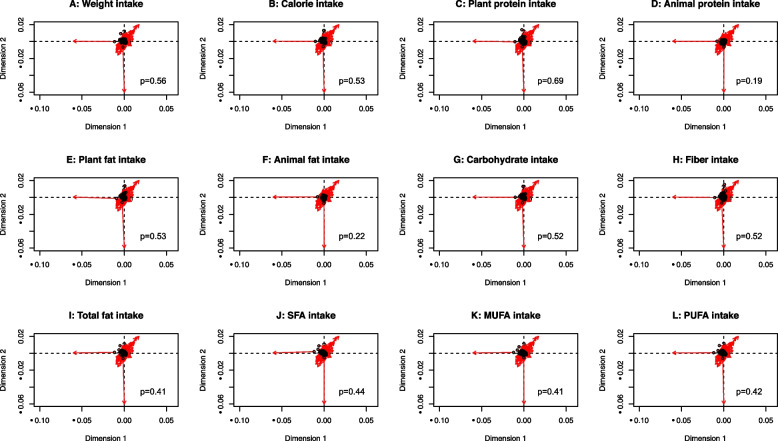
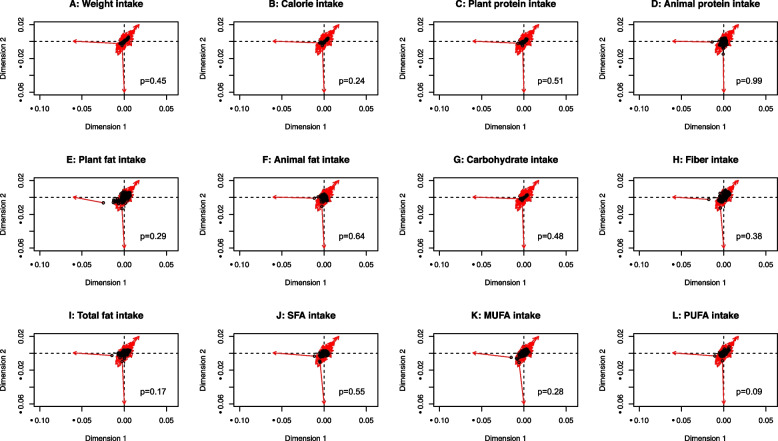
The rotation from the dietary beta-diversity matrix (for weight consumption, energy intake, plant protein, animal protein, plant fat, animal fat, carbohydrates, fiber, total fatty acids, saturated fatty acids, monounsaturated fatty acids (MUFAs), and polyunsaturated fatty acids (PUFAs)) into the microbiome diversity matrix was examined by the Procrustes analysis. We found that the food choice of an individual did not correspond with the microbiome composition of that individual when analyzed using the unweighted (Figs. 2A-2L) and weighted (Figs. 3A-3L) UniFrac-based food distances (p>0.05).
Fig. 2.
Procrustes analysis of tree-based food beta-diversity (unweighted UniFrac) of daily (A) weight consumption, (B) energy intake (C) plant protein, (D) animal protein, (E) plant fat, (F) animal fat, (G) carbohydrates, (H) fiber, (I) total fatty acids, (J) saturated fatty acids, (K) monounsaturated fatty acids, and (L) polyunsaturated fatty acids with microbiome composition beta-diversity (Aitchison’s distance). The plots show the rotation between the two ordinations necessary to make them match as closely as possible. Symbols (in black color) show the position of the samples in the first ordination (tree-based food), and arrows (in red color) point to their positions in the target ordination (microbiome composition)
Fig. 3.
Procrustes analysis of tree-based food beta-diversity (weighted UniFrac) of daily (A) weight consumption, (B) energy intake (C) plant protein, (D) animal protein, (E) plant fat, (F) animal fat, (G) carbohydrates, (H) fiber, (I) total fatty acids, (J) saturated fatty acids, (K) monounsaturated fatty acids, and (L) polyunsaturated fatty acids with microbiome composition beta-diversity (Aitchison’s distance). The plots show the rotation between the two ordinations necessary to make them match as closely as possible. Symbols (in black color) show the position of the samples in the first ordination (tree-based food), and arrows (in red color) point to their positions in the target ordination (microbiome composition)
Dietary diversity in associations with lifestyle factors and metabolic diseases
The association between the diversity of diet consumption and lifestyle factors and metabolic diseases is shown in (Additional File 1: eTable 2). All alpha-diversity indices of PUFA intake are significantly lower in hypertensive individuals than those without history of hypertension (Additional File 1: eFigures 6A-C). Both the distances for beta-diversity measurements showed the significant difference of diverse PUFA intake between smokers and non-smokers. However, the percentages of the variance explained by smoking status were observed to be very low (the R-square values of 1.95% and 1.88% for unweighted and weighted UniFrac distance metrics for PUFA intake diversity, respectively), which food source diversity of PUFA intake appeared not to be distinct (Additional File 1: eFigures 6D-E). The differences in within-subject dietary diversity of PUFA intake by history of diabetes and plant fat intake by history of hypertension were found; whereas between-subject dietary diversity of plant fat, carbohydrates, fiber, total fatty acids, and MUFA intake was associated with smoking status, depending on indices and measurements.
Discussion
In this cohort of CRC patients, we investigated the microbiome variation according to different lifestyles, metabolic diseases, and diet consumption. While smokers, drinkers, and diabetic individuals had an increase in microbiome stochasticity (the AKP effect), the anti-AKP effects were presented in obese and hypertension individuals, compared to their counterparts.
The use of the AKP effects has been well established for the composition of microbiomes affected by external diverse stressors, such as predators, parasites, and social disruption, through the replacement or generation of locally deterministic changes of sensitive bacteria [6, 26]. Previous studies have demonstrated that modifiable factors have a strong effect on the structure and function of human gut microbial communities. Understanding these effects in a cohort of CRC patients is a vital goal in consulting recommendations through microbial ecology-based evidence. Here, we observed the AKP effects of smoking status, alcohol consumption, and diabetes, which indicates that smoking, drinking, and diabetes in CRC patients may cause a more variable and unstable microbiome structure due to the unavailability of the host to modulate their microbiome when disturbed. Consistently, findings from our network analyses indicated increased dispersion of bacteria in smokers, drinkers, and diabetic individuals than those of non-smokers, non-drinkers, and non-diabetics, respectively. In contrast, we observed a more stable microbiome composition among patients with elevated BMI or blood pressure compared to patients with low BMI or blood pressure, which indicated a more stable microbiome in obese or hypertensive patients than their counterparts.
Furthermore, there was greater dispersion of the microbiome composition in smoking and alcohol-consuming patients than in their counterparts, which was interpreted as dysbiosis with the less connected network in the dysbiotic group than in the non-dysbiotic group. Several biological mechanisms have been proposed that describe how numerous toxic chemicals in cigarette smoke, such as nicotine, aldehydes, and heavy metals, can affect the bacterial community through the peripheral immune system [27, 28]. Tobacco smoking has been shown to inhibit natural killer cell activities, enhance white blood cell counts, and increase infection susceptibility, which results in the impairment of antimicrobial defenses [27, 28]. Additionally, smoking can alter the gut microbiome by accumulating the gut taxon that promotes inflammation, such as Bacteroides, Lachnospira, Prevotella stercorea, and Ruminococcus [29]. Alcohol dependence has been associated with the onset of an inflammatory environment in the gut and alters the gut microbiome by deriving alcoholic metabolites and several neurotransmitters such as gamma-aminobutyric acid, serotonin, and dopamine [30–32].
The AKP effects of BMI remain controversial. Our study found a decrease in the gut microbiome variation related to obesity in CRC patients. However, Ma et al. observed a significantly higher similarity index in obese than lean individuals but a non-significant difference between overweight and lean subjects, indicating the presence of AKP effects in obesity only [7]. In general, a BMI in the overweight/obese range is related to the development of CRC through the mediators of systemic inflammation such as tumor necrosis factor-alpha and interleukin 6 [33, 34]. Inflammation also contributes to an increased risk of CRC via affecting obesity-related dysbiosis [35]. A meta-analysis of 1,301 participants revealed a lower microbiome diversity in obese compared to non-obese individuals without CRC, however, the diversity did not differ between obese and non-obese CRC patients [36], demonstrating the absence of AKP effects of obesity in individuals with CRC. Considering that Asian populations have a higher body fat percentage than non-Asian populations [37, 38], our study selected the cutoff BMI of 25 kg/m2 for obesity instead of the World Health Organization recommendation and observed the anti-AKP effect of obesity. We hypothesized that the different cutoffs of BMI affected our observation of AKP effects of obesity.
In this study, we found a decrease in the gut microbiome variation related to a history of hypertension but an increased heterogeneity related to underlying diabetes among CRC patients. Such bidirectional effects of the gut microbiome on blood pressure and fasting glucose level have been proposed [39, 40]. The overgrowth of the genus Prevotella and Klebsiella was shown to contribute to pre-hypertension and hypertension, and a large sample of Finns reported a weak association between an increase in several genera in the phylum Firmicutes and a decrease in many distinct Lactobacillus species in patients with blood pressure [41, 42]. The modulation of metformin, as well as other anti-diabetic agents to the microbial community, also received much-deserved interest [43, 44]. Nevertheless, the AKP effects of hypertension and type 2 diabetes have not been investigated before [45].
The concept of the AKP effect was introduced with the more extensive stochasticity and heterogeneity of microbiome composition between individuals, thus, leaded to a decrease in the ability of the immune responding to the exposure [6, 7]. However, the anti-AKP effect was further argued as an extreme case of the AKP effect [6]. Under mild conditions, dysbiosis in some individuals resulted in the difference of their microbial composition and increased between-individual variations [45]. Under severe conditions, dysbiosis occurred in most of individuals, which drastically reduced their microbial communities and made the microbial composition to be more similar between individuals [15].
Associations of food groups and dietary patterns with microbiome composition and diversity have been identified in the Asian population [46–48]. However, how the overall diet shapes the microbiome profile remains unclear. By constructing the tree-based food from the SQFFQ of 106 food items, we observed that diversity in terms of weight, energy, macronutrients, and fatty acid intake did not shape the gut microbial community in our cohort of CRC patients, which might be similar to the result of diets accounting for the only small proportion of microbiome variation in population-level studies [9, 49–51]. Notably, a recent ultra-dense longitudinal study revealed a high interaction of diet and microbiome at the individual level [45], which may explain our non-significant findings.
The concept of dietary diversity has been introduced as an indicator of nutrient adequacy and overall diet quality [52]. Despite the disparity from different calculation methods, pooled results of 16 individual studies did not find any significant associations between dietary diversity score with both overweight/obesity and BMI, which was consistent with our findings [53]. In addition, inverse associations between dietary diversity and the history of hypertension may partially support the reduced stochasticity of the microbiome community among hypertensive individuals due to their reduced composition and quantities of foods. However, the link between dietary diversity and microbiome variation did not show any significant findings.
The strength of the current study includes the use of the hierarchical tree for food-based consumption, which could reduce the dimension of complex dietary intake information. Besides, this study contains some limitations. First, from the nature of observational studies, there could be measurement errors in the assessment of modifiable factors and metabolic diseases due to recall bias. However, the utilization of a validated SQFFQ might minimize this error. Second, more than half of our study participants did not provide information on dietary intake, which may limit us in detecting the diet-microbiome relationship. We assumed those with and without dietary data were comparable because they did not differ in terms of general characteristics. Last, our study was unable to expand the AKP theory in the identification of the diversity-stability association and the assessment of the balance between deterministic and stochastic forces due to the lack of longitudinal data [7].
Conclusion
In summary, our findings suggested an immune dysregulation and a reduced ability of the host and its microbiome in regulating the community composition. History of smoking, alcohol consumption, and diabetes were shown to affect parts of individuals in shifting new microbial communities and exert higher levels of stochasticity, whereas obesity and history of hypertension appeared to affect majority of individuals and shifted to drastic reductions in microbial compositions. Non-significant associations between dietary choices and microbiome diversity suggested the only small proportion of microbiome variation can be explained by dietary intake at the population level. Understanding the contribution of modifiable factors to differentiations of the gut microbiome among individuals may provide insights into how the microbiome regulates the effects of these factors on the health outcomes of CRC patients.
Supplementary Information
Additional file 1: eTable 1. Hierarchical classification of foods at different levels. eTable 2. P-values from the generalized linear model for associations of the dietary alpha-diversity (Chao1, Shannon, Simpson indices) and from the permutational of variance tests for associations of dietary beta-diversity (unweighted UniFrac and weighted UniFrac distances) with lifestyle factors and metabolic diseases. eFigure 1. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in smokers and non-smokers. Yellowish circles indicate taxon which are not enriched in either smoking or non-smoking group. The diameter of each circle is proportional to the relative abundance. eFigure 2. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in drinkers and non-drinkers. Yellowish circles indicate taxon which are not enriched in either drinking or non-drinking group. The diameter of each circle is proportional to the relative abundance. eFigure 3. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in obese and normal weight individuals. Yellowish circles indicate taxon which are not enriched in either obesity or normal weight group. The diameter of each circle is proportional to the relative abundance. eFigure 4. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in hypertensive and nonhypertensive individuals. Yellowish circles indicate taxon which are not enriched in either hypertensive or non-hypertensive group. The diameter of each circle is proportional to the relative abundance. eFigure 5. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in diabetic and non-diabetic individuals. Yellowish circles indicate taxon which are not enriched in either diabetes or non-diabetes group. The diameter of each circle is proportional to the relative abundance. eFigure 6. Within- and between-subject dietary diversity of polyunsaturated fatty acid (PUFA) intake according to history of hypertension and smoking status. Box plots show dietary diversity indices [(A) Chao1, (B) Shannon, (C) Simpson] of PUFA supply within hypertensive and non-hypertensive individuals. Principal coordinate analysis plots based on (D) Unweighted UniFrac and (E) Weighted UniFrac show variation in food composition of PUFA supply between smoking and non-smoking individuals.
Acknowledgements
Not applicable.
Abbreviations
- CRC
Colorectal cancer
- SCFA
Short-chain fatty acid
- AKP
Anna Karenina principle
- BMI
Body mass index
- AJCC
American Joint Committee on Cancer
- SQFFQ
Semi-quantitative food frequency questionnaire
- CAN-Pro
Computer-Aided Nutritional Analysis Program
- COZINE
COpositional Zero-Inflated Network Estimation
- LEfSe
Linear discrimination analysis effect size
- MUFA
Monounsaturated fatty acid
- PERMANOVA
Permutational multivariate analysis
- PCR
Polymerase chain reaction
- PUFA
Polyunsaturated fatty acid
Authors’ contributions
TH made contributions to study conceptualization, data analysis, interpretation the results, and was a major contributor in writing the manuscript. MK made contributions to study conceptualization and design, data interpretation, revising the manuscript critically for intellectual content, and funding acquisition. JWP and SYJ contributed to data collection, funding acquisition, and involved in revising the manuscript critically for intellectual content. JL made substantial contributions to data analysis and involved in revising the manuscript. AS made contributions to study conceptualization and design, data interpretation, and revising the manuscript critically for intellectual content. All authors critically reviewed this manuscript and approved the final version to be published.
Funding
This study was supported by grants from Seoul National University Hospital (No. 0420190530) and the National Research Foundation of Korea (NRF) grant funded by the Korea goverment (MSIT) (No. 2022R1A2C1004608). The funding bodies have no role in study design, collection, analysis, interpretation of data, or writing the manuscript. The authors declare no conflicts of interest. Authors Minjung Kim and Aesun Shin had full access to all the data in the study and took responsibility for the integrity of the data and the accuracy of the data analysis.
Availability of data and materials
The sequencing dataset supporting the conclusions of this article is available from the NCBI Sequence Read Archive (SRA) repository, [PRJNA797640, in https://www.ncbi.nlm.nih.gov/sra/PRJNA797640]. All other data are available from the corresponding author by reasonable request.
Declarations
Ethics approval and consent to participate
All participants provided written informed consent prior to stool sample collection. The study was approved by the Institutional Review Board of Seoul National University (No. 2205-105-1325). All methods were performed in accordance with the relevant guidelines and regulations.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Footnotes
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Minjung Kim, Email: minjungkim@snuh.org.
Aesun Shin, Email: shinaesun@snu.ac.k.
References
- 1.DeGruttola AK, Low D, Mizoguchi A, Mizoguchi E. Current understanding of dysbiosis in disease in human and animal models. Inflamm Bowel Dis. 2016;22(5):1137–1150. doi: 10.1097/MIB.0000000000000750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ji BW, Sheth RU, Dixit PD, Tchourine K, Vitkup D. Macroecological dynamics of gut microbiota. Nat Microbiol. 2020;5(5):768–775. doi: 10.1038/s41564-020-0685-1. [DOI] [PubMed] [Google Scholar]
- 3.Zheng D, Liwinski T, Elinav E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020;30(6):492–506. doi: 10.1038/s41422-020-0332-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sheflin AM, Whitney AK, Weir TL. Cancer-promoting effects of microbial dysbiosis. Curr Oncol Rep. 2014;16(10):406. doi: 10.1007/s11912-014-0406-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Cheng Y, Ling Z, Li L. The intestinal microbiota and colorectal cancer. Front Immunol. 2020;11:615056. doi: 10.3389/fimmu.2020.615056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zaneveld JR, McMinds R, Vega Thurber R. Stress and stability: applying the Anna Karenina principle to animal microbiomes. Nat Microbiol. 2017;2:17121. doi: 10.1038/nmicrobiol.2017.121. [DOI] [PubMed] [Google Scholar]
- 7.Ma ZS. Testing the Anna Karenina principle in human microbiome-associated diseases. iScience. 2020;23(4):101007. doi: 10.1016/j.isci.2020.101007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Deschasaux M, Bouter KE, Prodan A, Levin E, Groen AK, Herrema H, Tremaroli V, Bakker GJ, Attaye I, Pinto-Sietsma SJ, et al. Depicting the composition of gut microbiota in a population with varied ethnic origins but shared geography. Nat Med. 2018;24(10):1526–1531. doi: 10.1038/s41591-018-0160-1. [DOI] [PubMed] [Google Scholar]
- 9.Falony G, Joossens M, Vieira-Silva S, Wang J, Darzi Y, Faust K, Kurilshikov A, Bonder MJ, Valles-Colomer M, Vandeputte D, et al. Population-level analysis of gut microbiome variation. Science. 2016;352(6285):560–564. doi: 10.1126/science.aad3503. [DOI] [PubMed] [Google Scholar]
- 10.Conlon MA, Bird AR. The impact of diet and lifestyle on gut microbiota and human health. Nutrients. 2014;7(1):17–44. doi: 10.3390/nu7010017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rebersek M. Gut microbiome and its role in colorectal cancer. BMC Cancer. 2021;21(1):1325. doi: 10.1186/s12885-021-09054-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Zhao L, Cho WC, Nicolls MR. Colorectal cancer-associated microbiome patterns and signatures. Front Genet. 2021;12:787176. doi: 10.3389/fgene.2021.787176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Feng Q, Xu L, Li L, Qiu J, Huang Z, Jiang Y, Wen T, Lu S, Meng F, Shu X. Risk of death in colorectal cancer patients with multi-morbidities of metabolic syndrome: a retrospective multicohort analysis. Cancer Res Treat. 2021;53(3):714–723. doi: 10.4143/crt.2020.481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Skyrud KD, Myklebust TA, Bray F, Eriksen MT, de Lange T, Larsen IK, Moller B. How many deaths from colorectal cancer can be prevented by 2030? a scenario-based quantification of risk factor modification, screening, and treatment in Norway. Cancer Epidemiol Biomarkers Prev. 2017;26(9):1420–1426. doi: 10.1158/1055-9965.EPI-17-0265. [DOI] [PubMed] [Google Scholar]
- 15.Johnson AJ, Zheng JJ, Kang JW, Saboe A, Knights D, Zivkovic AM. A Guide to diet-microbiome study design. Front Nutr. 2020;7:79. doi: 10.3389/fnut.2020.00079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Zhao J, Li Z, Gao Q, Zhao H, Chen S, Huang L, Wang W, Wang T. A review of statistical methods for dietary pattern analysis. Nutr J. 2021;20(1):37. doi: 10.1186/s12937-021-00692-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li H, Li T, Beasley DE, Hedenec P, Xiao Z, Zhang S, Li J, Lin Q, Li X. Diet diversity is associated with beta but not alpha diversity of Pika gut microbiota. Front Microbiol. 2016;7:1169. doi: 10.3389/fmicb.2016.01169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ahn Y, Kwon E, Shim JE, Park MK, Joo Y, Kimm K, Park C, Kim DH. Validation and reproducibility of food frequency questionnaire for Korean genome epidemiologic study. Eur J Clin Nutr. 2007;61(12):1435–1441. doi: 10.1038/sj.ejcn.1602657. [DOI] [PubMed] [Google Scholar]
- 19.Hoang T, Lee J, Kim J. Differences in dietary patterns identified by the Gaussian graphical model in Korean adults with and without a self-reported cancer diagnosis. J Acad Nutr Diet. 2021;121(8):1484–1496 e1483. doi: 10.1016/j.jand.2020.11.006. [DOI] [PubMed] [Google Scholar]
- 20.Hoang T, Lee J, Kim J, Park B. Food intake behavior in cancer survivors in comparison with healthy general population; from the Health Examination center-based cohort. J Cancer Prev. 2019;24(4):208–216. doi: 10.15430/JCP.2019.24.4.208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Ha MJ, Kim J, Galloway-Pena J, Do KA, Peterson CB. Compositional zero-inflated network estimation for microbiome data. BMC Bioinformatics. 2020;21(Suppl 21):581. doi: 10.1186/s12859-020-03911-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Matchado MS, Lauber M, Reitmeier S, Kacprowski T, Baumbach J, Haller D, List M. Network analysis methods for studying microbial communities: A mini review. Comput Struct Biotechnol J. 2021;19:2687–2698. doi: 10.1016/j.csbj.2021.05.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ning D, Deng Y, Tiedje JM, Zhou J. A general framework for quantitatively assessing ecological stochasticity. Proc Natl Acad Sci U S A. 2019;116(34):16892–16898. doi: 10.1073/pnas.1904623116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Hoang T, Kim MJ, Park JW, Jeong SY, Lee J, Shin A. Nutrition-wide association study of microbiome diversity and composition in colorectal cancer patients. BMC Cancer. 2022;22(1):656. doi: 10.1186/s12885-022-09735-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kable ME, Chin EL, Storms D, Lemay DG, Stephensen CB. Tree-based analysis of dietary diversity captures associations between fiber intake and gut microbiota composition in a healthy US adult cohort. J Nutr. 2022;152(3):779–788. doi: 10.1093/jn/nxab430. [DOI] [PubMed] [Google Scholar]
- 26.Lavrinienko A, Tukalenko E, Kesaniemi J, Kivisaari K, Masiuk S, Boratynski Z, Mousseau TA, Milinevsky G, Mappes T, Watts PC. Applying the Anna Karenina principle for wild animal gut microbiota: temporal stability of the bank vole gut microbiota in a disturbed environment. J Anim Ecol. 2020;89(11):2617–2630. doi: 10.1111/1365-2656.13342. [DOI] [PubMed] [Google Scholar]
- 27.Gui X, Yang Z, Li MD. Effect of cigarette smoke on gut microbiota: state of knowledge. Front Physiol. 2021;12:673341. doi: 10.3389/fphys.2021.673341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Huang C, Shi G. Smoking and microbiome in oral, airway, gut and some systemic diseases. J Transl Med. 2019;17(1):225. doi: 10.1186/s12967-019-1971-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yan S, Ma Z, Jiao M, Wang Y, Li A, Ding S. Effects of smoking on inflammatory markers in a healthy population as analyzed via the gut microbiota. Front Cell Infect Microbiol. 2021;11:633242. doi: 10.3389/fcimb.2021.633242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Qamar N, Castano D, Patt C, Chu T, Cottrell J, Chang SL. Meta-analysis of alcohol induced gut dysbiosis and the resulting behavioral impact. Behav Brain Res. 2019;376:112196. doi: 10.1016/j.bbr.2019.112196. [DOI] [PubMed] [Google Scholar]
- 31.Dubinkina VB, Tyakht AV, Odintsova VY, Yarygin KS, Kovarsky BA, Pavlenko AV, Ischenko DS, Popenko AS, Alexeev DG, Taraskina AY, et al. Links of gut microbiota composition with alcohol dependence syndrome and alcoholic liver disease. Microbiome. 2017;5(1):141. doi: 10.1186/s40168-017-0359-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mutlu EA, Gillevet PM, Rangwala H, Sikaroodi M, Naqvi A, Engen PA, Kwasny M, Lau CK, Keshavarzian A. Colonic microbiome is altered in alcoholism. Am J Physiol Gastrointest Liver Physiol. 2012;302(9):G966–978. doi: 10.1152/ajpgi.00380.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Loke YL, Chew MT, Ngeow YF, Lim WWD, Peh SC. Colon carcinogenesis: the interplay between diet and gut microbiota. Front Cell Infect Microbiol. 2020;10:603086. doi: 10.3389/fcimb.2020.603086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zheng R, Du M, Zhang B, Xin J, Chu H, Ni M, Zhang Z, Gu D, Wang M. Body mass index (BMI) trajectories and risk of colorectal cancer in the PLCO cohort. Br J Cancer. 2018;119(1):130–132. doi: 10.1038/s41416-018-0121-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Mahmoud KF, Griffith KJ, Hayden A, Fogger SA, Kameg BN, Mitchell AM. Women of childbearing age and fetal alcohol spectrum disorder prevention: a position paper. J Addict Nurs. 2020;31(4):302–306. doi: 10.1097/JAN.0000000000000371. [DOI] [PubMed] [Google Scholar]
- 36.Greathouse KL, White JR, Padgett RN, Perrotta BG, Jenkins GD, Chia N, Chen J. Gut microbiome meta-analysis reveals dysbiosis is independent of body mass index in predicting risk of obesity-associated CRC. BMJ Open Gastroenterol. 2019;6(1):e000247. doi: 10.1136/bmjgast-2018-000247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Seo MH, Lee WY, Kim SS, Kang JH, Kang JH, Kim KK, Kim BY, Kim YH, Kim WJ, Kim EM, et al. 2018 Korean society for the study of obesity guideline for the management of obesity in Korea. J Obes Metab Syndr. 2019;28(1):40–45. doi: 10.7570/jomes.2019.28.1.40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Seo MH, Kim YH, Han K, Jung JH, Park YG, Lee SS, Kwon HS, Lee WY, Yoo SJ. Prevalence of obesity and incidence of obesity-related comorbidities in Koreans based on National Health Insurance Service health checkup data 2006–2015. J Obes Metab Syndr. 2018;27(1):46–52. [DOI] [PMC free article] [PubMed]
- 39.Al Khodor S, Reichert B, Shatat IF. The microbiome and blood pressure: can microbes regulate our blood pressure? Front Pediatr. 2017;5:138. doi: 10.3389/fped.2017.00138. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Anhe FF, Barra NG, Schertzer JD. Glucose alters the symbiotic relationships between gut microbiota and host physiology. Am J Physiol Endocrinol Metab. 2020;318(2):E111–E116. doi: 10.1152/ajpendo.00485.2019. [DOI] [PubMed] [Google Scholar]
- 41.Palmu J, Salosensaari A, Havulinna AS, Cheng S, Inouye M, Jain M, Salido RA, Sanders K, Brennan C, Humphrey GC, et al. Association between the gut microbiota and blood pressure in a population cohort of 6953 individuals. J Am Heart Assoc. 2020;9(15):e016641. doi: 10.1161/JAHA.120.016641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Li J, Zhao F, Wang Y, Chen J, Tao J, Tian G, Wu S, Liu W, Cui Q, Geng B, et al. Gut microbiota dysbiosis contributes to the development of hypertension. Microbiome. 2017;5(1):14. doi: 10.1186/s40168-016-0222-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Gurung M, Li Z, You H, Rodrigues R, Jump DB, Morgun A, Shulzhenko N. Role of gut microbiota in type 2 diabetes pathophysiology. EBioMedicine. 2020;51:102590. doi: 10.1016/j.ebiom.2019.11.051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Whang A, Nagpal R, Yadav H. Bi-directional drug-microbiome interactions of anti-diabetics. EBioMedicine. 2019;39:591–602. doi: 10.1016/j.ebiom.2018.11.046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Johnson AJ, Vangay P, Al-Ghalith GA, Hillmann BM, Ward TL, Shields-Cutler RR, Kim AD, Shmagel AK, Syed AN, Personalized Microbiome Class S et al. Daily sampling reveals personalized diet-microbiome associations in humans. Cell Host Microbe. 2019;25(6):789–802 e785. doi: 10.1016/j.chom.2019.05.005. [DOI] [PubMed] [Google Scholar]
- 46.Wu X, Unno T, Kang S, Park S. A Korean-style balanced diet has a potential connection with Ruminococcaceae enterotype and reduction of metabolic syndrome incidence in Korean adults. Nutrients 2021;13(2):495. [DOI] [PMC free article] [PubMed]
- 47.Noh H, Jang HH, Kim G, Zouiouich S, Cho SY, Kim HJ, et al. Taxonomic composition and diversity of the gut microbiota in relation to habitual dietary intake in Korean adults. Nutrients. 2021;13(2):366. [DOI] [PMC free article] [PubMed]
- 48.Yu D, Nguyen SM, Yang Y, Xu W, Cai H, Wu J, Cai Q, Long J, Zheng W, Shu XO. Long-term diet quality is associated with gut microbiome diversity and composition among urban Chinese adults. Am J Clin Nutr. 2021;113(3):684–694. doi: 10.1093/ajcn/nqaa350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Vangay P, Johnson AJ, Ward TL, Al-Ghalith GA, Shields-Cutler RR, Hillmann BM, Lucas SK, Beura LK, Thompson EA, Till LM, et al. US immigration westernizes the human gut microbiome. Cell. 2018;175(4):962–972 e910. [DOI] [PMC free article] [PubMed]
- 50.Rothschild D, Weissbrod O, Barkan E, Kurilshikov A, Korem T, Zeevi D, Costea PI, Godneva A, Kalka IN, Bar N, et al. Environment dominates over host genetics in shaping human gut microbiota. Nature. 2018;555(7695):210–215. doi: 10.1038/nature25973. [DOI] [PubMed] [Google Scholar]
- 51.Wang J, Thingholm LB, Skieceviciene J, Rausch P, Kummen M, Hov JR, Degenhardt F, Heinsen FA, Ruhlemann MC, Szymczak S, et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat Genet. 2016;48(11):1396–1406. doi: 10.1038/ng.3695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Habte TY, Krawinkel M. Dietary diversity score: a measure of nutritional adequacy or an indicator of healthy diet? J Nutri Health Sci. 2016;3(3):303. [Google Scholar]
- 53.Salehi-Abargouei A, Akbari F, Bellissimo N, Azadbakht L. Dietary diversity score and obesity: a systematic review and meta-analysis of observational studies. Eur J Clin Nutr. 2016;70(1):1–9. doi: 10.1038/ejcn.2015.118. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Additional file 1: eTable 1. Hierarchical classification of foods at different levels. eTable 2. P-values from the generalized linear model for associations of the dietary alpha-diversity (Chao1, Shannon, Simpson indices) and from the permutational of variance tests for associations of dietary beta-diversity (unweighted UniFrac and weighted UniFrac distances) with lifestyle factors and metabolic diseases. eFigure 1. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in smokers and non-smokers. Yellowish circles indicate taxon which are not enriched in either smoking or non-smoking group. The diameter of each circle is proportional to the relative abundance. eFigure 2. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in drinkers and non-drinkers. Yellowish circles indicate taxon which are not enriched in either drinking or non-drinking group. The diameter of each circle is proportional to the relative abundance. eFigure 3. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in obese and normal weight individuals. Yellowish circles indicate taxon which are not enriched in either obesity or normal weight group. The diameter of each circle is proportional to the relative abundance. eFigure 4. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in hypertensive and nonhypertensive individuals. Yellowish circles indicate taxon which are not enriched in either hypertensive or non-hypertensive group. The diameter of each circle is proportional to the relative abundance. eFigure 5. (A) Linear discriminant analysis (LDA) effect size (LEfSe) analysis and (B) cladogram for abundant bacteria in diabetic and non-diabetic individuals. Yellowish circles indicate taxon which are not enriched in either diabetes or non-diabetes group. The diameter of each circle is proportional to the relative abundance. eFigure 6. Within- and between-subject dietary diversity of polyunsaturated fatty acid (PUFA) intake according to history of hypertension and smoking status. Box plots show dietary diversity indices [(A) Chao1, (B) Shannon, (C) Simpson] of PUFA supply within hypertensive and non-hypertensive individuals. Principal coordinate analysis plots based on (D) Unweighted UniFrac and (E) Weighted UniFrac show variation in food composition of PUFA supply between smoking and non-smoking individuals.
Data Availability Statement
The sequencing dataset supporting the conclusions of this article is available from the NCBI Sequence Read Archive (SRA) repository, [PRJNA797640, in https://www.ncbi.nlm.nih.gov/sra/PRJNA797640]. All other data are available from the corresponding author by reasonable request.