Abstract
Background
HLA-E interaction with inhibitory receptor, NKG2A attenuates NK-mediated cytotoxicity. NKG2A overexpression by SARS-CoV-2 exhausts NK cells function, whereas virus-induced down-regulation of MHC-Ia reduces its derived-leader sequence peptide levels required for proper binding of HLA-E to NKG2A. This leads HLA-E to become more complex with viral antigens and delivers them to CD8+ T cells, which facilitates cytolysis of infected cells. Now, the fact that alleles of HLA-E have different levels of expression and affinity for MHC Ia-derived peptide raises the question of whether HLA-E polymorphisms affect susceptibility to COVID-19 or its severity.
Methods
104 COVID-19 convalescent plasma donors with/without history of hospitalization and 18 blood donors with asymptomatic COVID-19, all were positive for anti-SARS-CoV-2 IgG antibody as well as a group of healthy control including 68 blood donors with negative antibody were subjected to HLA-E genotyping. As a privilege, individuals hadn’t been vaccinated against COVID-19 and therefore naturally exposed to the SARS-CoV-2.
Results
The absence of HLA-E*01:03 allele significantly decreases the odds of susceptibility to SARS-CoV-2 infection [p = 0.044; OR (95 %CI) = 0.530 (0.286 – 0.983)], suggesting that HLA-E*01:01 + HLA-E*01:01 genotype favors more protection against SARS-CoV-2 infection. HLA-E*01:03 + HLA-E*01:03 genotype was also significantly associated with more severe COVID-19 [p = 0.020; 2.606 (1.163 – 5.844)
Conclusion
Here, our observation about lower susceptibility of HLA-E*01:01 + HLA-E*01:01 genotype to COVID-19 could be clinical evidence in support of some previous studies suggesting that the lower affinity of HLA-E*01:01 to peptides derived from the leader sequence of MHC class Ia may instead shift its binding to virus-derived peptides, which then facilitates target recognition by restricted conventional CD8+ T cells and leads to efficient cytolysis. On the other hand, according to other studies, less reactivity of HLA-E*01:01 with NKG2A abrogates NK cells or T cells inhibition, which may also lead to a greater cytotoxicity against SARS-CoV-2 infected cells compared to HLA-E*01:03. Taken together given HLA-E polymorphisms, the data presented here may be useful in identifying more vulnerable individuals to COVID-19 for better care and management. Especially since along with other risk factors in patients, having HLA-E*01:03 + HLA-E*01:03 genotype may also be associated with the possibility of severe cases of the disease.
Keywords: CD8+ T cell, Coronavirus disease 2019 (COVID-19), HLA-E, NK cell, NKG2A, Polymorphism, SARS-CoV-2, Susceptibility
1. Introduction
The outbreak of the newest strain of SARS-CoV-2, the causative agent of coronavirus disease-2019 (COVID-19), has so far claimed millions of lives [1], however, as with other infections, it is still the adequacy of the innate immune system in the body, that plays a critical role against COVID-19 as the first effective line of defense. In this regard, it is clear that the proper functioning of natural killer (NK) cells, which play a key role in preventing the spread of viruses and disease in the body, is of great importance [2]. Therefore, the presence of NK cells that can both effectively eliminate the virus itself and modulate the inflammatory side effects of the cytokine storm [3], [4] is of particular importance in the pathology of SARS-CoV-2 disease [5].
Previous studies have indicated that the lethal function of NK cells is mainly regulated by one of the most important inhibitory receptors of these cells, NKG2A [6], [7], [8], [9]. The main ligand of this receptor is HLA-E molecule which its interaction with NKG2A is considered as an important axis of inhibitory regulation of NK cell function [10]. However, structurally, HLA-E is a complex molecule associated with a subset of peptides derived from the leader sequence of MHC class I molecules that support its efficient transport to the cell surface and subsequent stabilization for proper interaction with NK cell receptors [11]. This is important in viral infections because while most viruses including SARS-CoV-2 reduce the expression of MHC class I and thus escape the recognition of the cellular immune system and lethality through cytotoxic T cells, on the other hand, decreased MHC class I expression as a supplier of complementary peptides required for HLA-E expression could negatively affect the optimal interaction of this ligand with the NKG2A inhibitory receptor of NK cells, which is essentially associated with an increase in the their lethality against virus-infected cells [12]. In contrast, some viruses, such as SARS-CoV-2, maintain the function level of these cells by increasing the expression of the NKG2A inhibitory receptor [13]. In some other viruses, such as HIV-1, in addition to reducing the expression of MCH class I on infected cells, the virus dramatically increases HLA-E expression and thus escapes the lethal mechanism of NK cells [14]. Similarly, in a recent study in patients with COVID-19, positive regulation of HLA-E in lung epithelial cells was observed in patients with moderate to severe COVID-19 [15]. This, indirectly, could be a sign of double inhibition of NK cells in these patients, since the NK cells in COVID-19 show an exhausted phenotype with high expression of NKG2A inhibitory receptor [16].
Understanding the role of HLA-E in regulating the function of NK cells has so far led to extensive research on this molecule. Various polymorphisms of HLA-E have been identified, of which due to the occurrence of two functional alleles of HLA-E*01:01 and HLA-E*01:03, three genotypes of HLA-E*01:01 + HLA-E*01:01, HLA-E*01:03 + HLA-E*01:03 and HLA-E*01:01 + HLA-E*01:03 are clinically important. Studies have also indicated higher cell surface expression and greater stability of the HLA-E molecule derived from the HLA-E*01:03 allele than the HLA-E*01:01. This difference may lead to a more effective inhibitory function of HLA-E*01:03 in interaction with NKG2A receptor, ultimately resulting in different function of NK cells in response to each of these ligands which suggesting that specific polymorphisms of HLA-E in individuals, can differently affect the function of NK cells [12]. While most previous research has focused on evaluating the expression of the HLA-E molecule in healthy tissues, tumors, and infected cells, as well as identifying different gene alleles of the molecule and structural differences in the products of each allele, recent studies have further examined the relationship between genotypes and alleles with susceptibility to a wide range of diseases and their prognosis or treatments [12], [17], [18], [19]. In this regard, some of the most important studies are those that address the role of HLA-E polymorphism in susceptibility to viral infections such as hepatitis C and HIV or its effect on the response to treatment of these infections [18]. However regardless of evidences that highlight the fundamental role of the HLA-E-NK cell axis in the optimal innate immune response to different health threats, HLA-E-restricted CD8+ T Lymphocytes can also be involved in acquired immune response against some pathogens or tumors [20]. Therefore considering HLA-E as the major modulator of NK cell that its exhaustion plays a key role in pathogenesis of COVID-19 [16], the present study sought to investigate the relationship between the polymorphism of HLA-E and the susceptibility to SARS-CoV2 infection or this disease conditions. In this way, we here compare the genotype of this molecule between healthy blood donors without antiviral antibodies and recovered patients with high antibody titers who were candidates for COVID-19 convalescent plasma (CCP) donation. This novel study is important because it was performed during the period when the vaccine was not available, so that all recovered patients and healthy controls had no history of vaccination against COVID-19, and therefore, any immune response in these individuals was completely natural and in real exposure to the virus.
2. Methods and materials
Study cohort: 104 recovered patients who had experienced symptomatic COVID-19 and were eligible to donate convalescent plasma under IBTO regulation [21] were considered as case group. This group was detected by the high titer of anti-SARS-CoV-2 antibody while they also subdivided into two groups based on hospitalization experience; those who were receiving in-home treatment and those who had a history of hospitalization for their more complicated conditions. A control group was selected from blood donors (86 individuals) without any history or suspicion of COVID-19 (according to the interview and routine medical examination before blood donation). However, COVID-19 screening with anti-SARS-CoV-2 ELISA (IgG) kit in these individuals showed a positive titer of antibody in 18 of individuals. Thus, the healthy control group in this study included 68 blood donors with no history of disease and vaccination experience against COVID-19, all of whom lacked detectable antibody against SARS-CoV-2. Although not expected, those blood donors with anti-SARS-CoV-2 IgG antibody who had not reported any sign of disease during the interview were considered as a group of people with asymptomatic COVID-19. Therefore, in summary, this study included four groups of subjects, which were: 1) a healthy control group of blood donors with negative anti-SARS-CoV-2 IgG antibody 2) a group of blood donors with asymptomatic COVID-19 experience with anti-SARS-CoV-2 IgG antibody 3) a COVID-19 convalescent plasma (CCP) donor group with the history of hospitalization due to COVID-19, who had an acceptable titer of anti-SARS-CoV-2 IgG antibody 4) a CCP donor group with a history of COVID-19 without hospitalization who also had an acceptable antibody titer. It is important to note that none of the individuals in these groups had any experience of vaccination against COVID-19. Therefore, all of these individuals were exposed to the SARS-CoV-2 only through natural infection. This is the particular privilege of present research compared to other similar studies, which is made possible by the specific time period of the project. It should be noted that all participants were male and median age in blood donors was 39 (24–54) and in recovered patient was 40 (22–56) years. The informed consent was obtained from all donors and patients involved in this study. It is worth mentioning that all the following methods were performed in accordance with the relevant instructions and regulations.
2.1. Serological test for COVID-19
To rule out previous infection for healthy control group, the serum of volunteer blood donors was subjected to anti-SARS-CoV-2 ELISA IgG assay (EI 2606–9601 G, EUROIMMUN; Lübeck, Germany), while all the CCP donors were also double checked to confirm they were positive for anti-SARS-CoV-2 IgG antibody against SARS-CoV-2. The assay was performed according to the manufacture’s protocol. In brief, the kit contains microplate coated with recombinant S1 domain of the spike protein of SARS-CoV-2. The samples were diluted with buffer and added to each well along with the controls and calibrator. Followed by 60 min incubation at 37 °C, the peroxidase-labeled anti-human IgG was added to detect the bound antibodies. After 30 min of incubation and washing, the chromogen/substrate solution was added while the plate kept at room temperature for 20 min and finally the reaction was stopped by adding stop solution. The color intensity of each well was detected at a wavelength of 450 nm. The results were evaluated semi-quantitatively by calculating the ratio of the extinction of the control or sample over the extinction of calibrator. The ratio < 0.8 was considered as negative results while the ratio ≥ 0.8 to < 1.1 was on borderline and ratio ≥ 1.1 detected as positive.
2.2. Genotyping of HLA-E
Genomic DNA was extracted from patients’ blood samples and then all samples were genotyped for HLA-E locus using a sequence specific primer (SSP)-PCR strategy. Amplification was carried out using the Thermal cycler machine (PEQLAB, Germany), in which the forward primers of HLA-E*01:01 (5′-GCGCCTTTACCCGGTTCTT-3′) and HLA-E*01:03 (5′-GGCTGCGAGATGGGGCCAGCCG-3′) in combination with the reverse primers (5′-GTAGCCCTGTGGACCCTCTTACC-3′) and (5′-CCCGCGGAGGAAGCGACT-3′) were applied to distinguish the HLA-E*01:01 and HLA-E*01:03 alleles. As the internal control, we applied human growth hormone (HGH) in which forward and reverse primers were (5′-TGCCTTCCCAACCATTCCCTT-3) and (5′-CCACTCACGGATTTCTGTTGTGTTTC-3), respectively. All oligonucleotides used in the project were synthesized by Sinaclon (Tehran; Iran). Some PCR products were DNA sequenced by Codon Genetic Group to confirm PCR-SSP results (Supplementary Fig. 1&2).
2.3. Statistical analysis
SPSS software version 23 was used for data analysis. All p values were two-tailed and p-values <0.05 were considered statistically significant. Data were analyzed using Chi-square/ Fisher exact test and logistic regression analysis.
3. Results
3.1. The presence of anti-SARS-CoV-2 IgG antibody in volunteer donors
All CCP donors were positive for anti-SARS-CoV-2 IgG antibody, which confirmed their previous screening results by Iranian blood transfusion organization; however, the volunteer donors eligible for blood donation had not been screened for anti-SARS-CoV-2 IgG antibody. Although this group of donors did not report any sign or obvious risk of disease in their interviews, to confirm they had no previous infection, their serums were examined for the presence of anti-SARS-CoV-2 IgG antibody. Interestingly, of the 86 blood donors studied here, 18 (20.9 %) were positive for anti-SARS-CoV-2 IgG antibody who were placed in the asymptomatic group, while the other 68 donors who were anti-SARS-CoV-2 IgG antibody negative formed the healthy control group. Neither CCP donors nor blood donors were vaccinated against COVID-19 at the time of the study, which rules out the possibility of vaccine-induced immune responses.
3.2. HLA-E genotype association with demographic parameters
Logistic regression model was used to investigate the association between the variables of HLA-E genotype, age, ethnicity and blood group with the possibility of COVID-19. Our univariate analyses showed there was no significant association between these variables and the possibility of COVID-19 except for HLA-E genotype. The analysis was also confirmed by multivariate logistic regression model (using Backward Wald method). Table 1 demonstrates demographic characteristics of CCP donors (case) and healthy blood donors (control).
Table 1.
Demographic characteristics of CCP donors (case) and healthy blood donors (control).
| Variables | Case [N (%)] | Control [N (%)] |
|---|---|---|
| Gender | ||
| Male | 172 | 100 |
| Age (years) | ||
| <35 | 26 (25) | 21 (30.8) |
| ≥35 & <45 | 49 (47.1) | 32 (47.1) |
| ≥45 | 29 (27.9) | 15 (22.1) |
| Ethnicity | ||
| Fars | 57 (54.8) | 43 (63.2) |
| Azerbaijanis | 26 (25) | 18 (26.5) |
| Others | 21 (20.2) | 7 (10.3) |
| Blood group | ||
| A | 34 (32.7) | 28 (41.2) |
| B | 23 (22.1) | 13 (19.1) |
| AB | 9 (8.7) | 9 (13.2) |
| O | 38 (36.5) | 18 (26.5) |
| Total | 104 (1 0 0) | 68 (1 0 0) |
CCP: COVID-19 convalescent plasma donor.
3.3. The distribution of HLA-E genotypes and alleles in healthy control group
According to our results, blood donors (healthy control group; n = 68) showed a comparable frequency with our previous cross-sectional study in Iranian population so that the genotype distribution in the present study was 44.1 % (n = 30) for HLA-E*01:01 + HLA-E*01:01 and 35.3 % (n = 24) for HLA-E*01:01 + HLA-E*01:03 genotype with a lower frequency for HLA-E*01:03 + HLA-E*01:03 genotype (20.6 %; n = 14). Furthermore, in those of blood donors who were positive for anti-SARS-CoV-2 IgG antibody (asymptomatic group; n = 18), 61.1 % (n = 11) had HLA-E*01:01 + HLA-E*01:01 while from the rest, 33.3 % (n = 6) and 5.6 % (n = 1) had HLA-E*01:01 + HLA-E*01:03 and HLA-E*01:03 + HLA-E*01:03 genotypes, respectively. As shown in Table 2 , there was no significant difference between healthy control and asymptomatic groups regarding HLA-E genotype (p = 0.2).
Table 2.
The association between HLA-E genotypes and alleles and the odds of SARS-CoV-2 infection.
 |
p value 0.2 indicates no significant difference of HLA-E genotypes between the healthy control (con.) and asymptomatic (asym.) individuals. p value 0.044 indicates a significant difference of HLA-E genotypes between the healthy control and infected individuals. Comparing healthy control and symptomatic (sym.) individuals, p value 0.006 indicates a significant differences of HLA-E*01:01/*01:01 versus other genotypes while p value 0.016 shows a significant difference of HLA-E*01:03/*01:03 versus others.
3.4. The association between HLA-E genotypes and alleles and the odds of SARS-CoV-2 infection
This study includes 122 SARS-CoV-2 infected cases that were positive for anti-SARS-CoV-2 IgG antibody. Of those, 104 cases were CCP donors with previous experience of symptomatic COVID-19, and 18 blood donors who had passed the regular clinical examination and interview while further evaluation of their antibody status on the time of study showed the experience of previous asymptomatic COVID-19 in these cases. In addition, as healthy control group, this study also includes 68 non-infected blood donors who were negative for anti-SARS-CoV-2 IgG antibody, which means that they had not any exposure to SARS-CoV-2 leading to infection. The comparison between SARS-CoV-2 infected and non-infected individuals showed that the absence of HLA-E*01:03 allele, on the other words HLA-E*01:01 + HLA-E*01:01 genotype, significantly 0.530 times decreases the odds of susceptibility to SARS-CoV-2 infection [p = 0.044; OR (95 %CI) = 0.530 (0.286 – 0.983)]. This indirectly suggests that HLA-E*01:01 + HLA-E*01:01 genotype may favor more protection against SARS-CoV-2 infection (Table 2).
3.5. The association between HLA-E genotypes and alleles and the odds of symptomatic COVID-19
This study includes 104 CCP donors with previous experience of symptomatic COVID-19 who were positive for anti-SARS-CoV-2 IgG antibody. In addition, as healthy control group, this study also includes 68 blood donors who were negative for anti-SARS-CoV-2 IgG antibody. The comparison between these CCP donors and healthy control group showed a significant relevance between HLA-E polymorphisms and COVID-19 (p = 0.021). We also found that the absence of HLA-E*01:03 allele, on the other words HLA-E*01:01 + HLA-E*01:01 genotype, significantly 0.401 times decreases the odds of COVID-19 than those with other genotypes of HLA-E*01:03 + HLA-E*01:03 or HLA-E*01:01 + HLA-E*01:03 [p = 0.006; OR (95 %CI) = 0.401 (0.208 – 0.773)]. Further analysis revealed that the odds of having COVID-19 in individuals with HLA-E*01:03 + HLA-E*01:03 genotype was significantly higher (2.743 times) than those with HLA-E*01:01 + HLA-E*01:01 genotype [p = 0.016; OR (95 %CI) = 2.743 (1.205 – 6.242)]. This indirectly suggests that HLA-E*01:01 + HLA-E*01:01 genotype may favor more protection against COVID-19 (Table 2).
3.6. The relevance of HLA-E genotype/ allele with the severity of COVID-19
According to our study group, the SARS-CoV-2 infected individuals are divided in three groups of asymptomatic infected blood donors (n = 18), CCP donors who experienced symptomatic COVID-19 with in-home treatment (n = 87) and or with hospitalization (n = 17). Given this, our data showed a significant association between HLA-E polymorphisms and severity of COVID-19 (p = 0.004). However, there was no significant difference regarding HLA-E genotype between CCP donors previously hospitalized and CCP donors treated at home. It is noteworthy that, the comparison between each of symptomatic groups with asymptomatic group showed significantly higher frequency of genotype of HLA-E*01:03 + HLA-E*01:03 in symptomatic individuals who were hospitalized [p = 0.030; OR (95 %CI) = 11.90 (1.272 – 111.351)] than those who were non-hospitalized [p = 0.068; OR (95 %CI) = 6.855 (0.865–54.299)] compared to asymptomatic patients.
Collectively, in the SARS-CoV-2 infected individuals, HLA-E*01:03 + HLA-E*01:03 genotype was significantly associated with more severe COVID-19 [p = 0.020; 2.606 (1.163 – 5.844)] (Table 3 ).
Table 3.
The relevance of HLA-E genotypes and alleles with the severity of COVID-19.
| Groups | Asym. (n = 18) |
Sym. non-hospitalized (n = 87) |
Sym. hospitalized (n = 17) | p value; OR (95 %CI) |
|---|---|---|---|---|
| Genotype/allele | ||||
| HLA-E*01:01 + HLA-E*01:01 | 11 (61.1 %) | 19 (21.8 %) | 6 (35.3 %) | |
| HLA-E*01:03 + HLA-E*01:03 | 1 (5.6 %) | 25 (28.7 %) | 7 (41.2 %) | 0.020; 2.606 (1.163–5.844) |
| HLA-E*01:01 + HLA-E*01:03 | 6 (33.3 %) | 43 (49.4 %) | 4 (23.5 %) | |
| HLA-E*01:01 | 17 (94.4 %) | 62 (71.3 %) | 10 (58.8 %) | |
| HLA-E*01:03 | 7 (38.9 %) | 68 (78.2 %) | 11 (64.7 %) |
CI, confidence interval; OR, odds ratio. p value indicates a significant difference of HLA-E genotype between the asymptomatic (Asym.) and symptomatic (Sym.) non-hospitalized groups.
categorizing all infected cases in two groups with asymptomatic and symptomatic COVID-19 showed a significant higher frequency of HLA-E*01:03 + HLA-E*01:03 [p = 0.014; OR (95 %CI) = 14.080 (1.702 – 116.474)] and significant lower frequency of HLA-E*01:01 + HLA-E*01:01 [p = 0.028; OR (95 %CI) = 0.290 (0.096 – 0.878)] in the symptomatic groups compared to asymptomatic group (Table 4 ). The results also showed that the odds of being asymptomatic carrier of COVID-19 in the individuals with HLA-E*01:01 allele trends to be more compared to symptomatic ones [p = 0.054; OR (95 %CI) = 7.556 (0.964 – 59.241)]. On the other hand, HLA-E*01:01 + HLA-E*01:01 genotype was significantly higher in those who experienced asymptomatic disease than symptomatic patients [p = 0.028; OR (95 %CI) = 3.447 (1.140 – 10.424)].
Table 4.
The relevance of HLA-E genotypes and alleles with the symptom of disease.
| Groups |
Asym. (n = 18) |
Sym. (n = 104) |
p value; OR (95 %CI) |
|---|---|---|---|
| Genotype/allele | |||
| HLA-E*01:01 + HLA-E*01:01 | 11 (61.1 %) | 25 (24 %) | 0.028; 0.290 (0.096–0.878) |
| HLA-E*01:03 + HLA-E*01:03 | 1 (5.6 %) | 32 (30.8 %) | 0.014; 14.080 (1.702–116.474) |
| HLA-E*01:01 + HLA-E*01:03 | 6 (33.3 %) | 47 (45.2 %) | |
| HLA-E*01:01 | 17 (94.4 %) | 72 (69.2 %) | |
| HLA-E*01:03 | 7 (38.9 %) | 79 (76 %) |
CI, confidence interval; OR, odds ratio. Comparing asymptomatic and symptomatic groups, p value 0.028 indicates a significant difference of HLA-E*01:01 + HLA-E*01:01 versus other genotypes, while p value 0.014 shows a significant difference of HLA-E*01:03 + HLA-E*01:03 versus others.
4. Discussion
This study was performed on two main arms of which one is blood donors with no known history of COVID-19 and the other is CCP donors who had varying degrees of the disease. In addition, this study was performed before the presentation of the anti-SARS-CoV-2 vaccines, which means that none of the participants had any experience of vaccination against disease and thus their immune response was induced naturally only in the face of the virus itself. To establish a group of healthy control with no history of infection, our focus was on the first arm of study, the volunteer blood donors who were passed the clinical examination and reported no signs of COVID-19. This was an important parameter for donor eligibility on the time of sampling for this study prior to vaccine presentation for global use. However, with all precaution, 20.9 % of blood donors of this study were found to be positive for anti-SARS-CoV-2 IgG antibody. Therefore, this part of donors was excluded while the rests of 68 formed a healthy control group. Notably, due to the numerous parameters involved in early COVID-19 epidemic (before global access to vaccines), including the onset stage of the epidemic and domestic disease control policies, in particular lockdown strategies, the seroprevalence of SARS-CoV-2 antibodies in blood donors varied widely from country to country making their generalization almost impossible. In this regard, the closest study to the present independent research conducted by Arabkhazaeli, et al., in which in a period almost similar to ours, they detected anti-SARS-CoV-2 antibodies in 14 % of Iranian asymptomatic donors using SARS-CoV-2 IgG/IgM Rapid Test Dipstick [22]. It should be noted, however, that since the kit used in that study was screening-based, caution should be taken in comparing its results with the present study, which is based on ELISA. Especially since, the mentioned kit screens IgG/IgM antibodies, while the ELISA kit in this study detects specific IgG antibody against S1 domain of the spike protein of SARS-CoV-2. In addition, above-mentioned study has also covered a bigger cohort in the multicenter national research. Thus give all variables, it is not surprising that studies in other countries have also shown varying results based on different disease control strategies and their economic and health status, from 40 % in Pakistani donors to<5 % in more developed countries, all during the period before global vaccination against SARS-CoV-2 [23], [24], [25]. Finally, after determining the status of anti-SARS-CoV-2 antibodies, the three main groups of our study included the healthy control group (negative antibody blood donors), the asymptomatic infected group (positive antibody blood donors) and the group with definite disease history (including two subgroups of CCP donors with or without a history of hospitalization) were identified.
The study was then continued by HLA-E genotyping in healthy control group where similar to our previous cross-sectional study in Iranian population [18], here we also found a higher frequency of HLA-E*01:01 allele. Accordingly, with a bigger cohort here, the genotype distribution for this group was also reported as 44.1 % for HLA-E*01:01 + HLA-E*01:01 and 35.3 % for HLA-E*01:01 + HLA-E*01:03 genotype with a lower frequency for HLA-E*01:03 + HLA-E*01:03 genotype (20.6 %), which almost confirmed our previous results. The genotype analysis did not show any significant difference between asymptomatic blood donors and healthy control group. However, intriguingly, comparing SARS-CoV-2 infected group (who were positive for anti-SARS-CoV-2 IgG antibody, including asymptomatic blood donors and CCP donors who experienced symptomatic COVID-19) with non-infected (as healthy control group; blood donors who were negative for anti-SARS-CoV-2 IgG antibody), our study showed that the absence of HLA-E*01:03 allele significantly decreases the odds of susceptibility to SARS-CoV-2 infection, this indirectly suggests that HLA-E*01:01 + HLA-E*01:01 genotype may favor more protection against SARS-CoV-2 infection. Furthermore, in our study, the comparison between CCP donors who were positive for anti-SARS-CoV-2 IgG antibody and healthy control group (negative for anti-SARS-CoV-2 IgG antibody) indicated a significant association between HLA-E polymorphisms and COVID-19, so that the odds of having COVID-19 in individuals with HLA-E*01:03 + HLA-E*01:03 genotype was significantly higher than those with HLA-E*01:01 + HLA-E*01:01 genotype. This indirectly suggests that HLA-E*01:01 + HLA-E*01:01 genotype may favor more protection against COVID-19.
According to further analysis among three groups of infected individuals with different levels of disease severity including asymptomatic infected blood donors, CCP donors who experienced symptomatic COVID-19 with and without hospitalization, there was a significant association between HLA-E polymorphisms and severity of COVID-19 (p = 0.004). Interestingly, our results showed that the frequency of HLA-E*01:03 + HLA-E*01:03 genotype in CCP donors who were hospitalized for COVID-19 was significantly more than asymptomatic infected blood donors [p = 0.030; OR (95 %CI) = 11.90 (1.272 – 111.351)]. However, the comparison between asymptomatic group and CCP donors who were non-hospitalized and treated at home did not show significant difference [p = 0.068; OR (95 %CI) = 6.855 (0.865 – 54.299)], suggesting that HLA-E*01:03 + HLA-E*01:03 genotype is more associated with hospitalization as a marker of severity of disease. Collectively, in the SARS-CoV-2 infected individuals, HLA-E*01:03 + HLA-E*01:03 genotype was significantly associated with more severe COVID-19 [p = 0.020; 2.606 (1.163 – 5.844)].
In line with these data, several previous studies have highlighted the relevance of HLA-E polymorphisms with different viral infections, of which the association of distinct HLA-E alleles with the status of hepatitis C, its treatment and clearance are the most studied to date [26], [27], [28]. Our recent study showed a significant relevance between homozygous status for HLA-E*01:03 allele with increased resistance to anti-HCV treatments in frequently transfused thalassemia patients, while, this research also confirmed the association between HLA-E*01:03 allele and increased susceptibility to HCV infection as well as higher resistance of individuals with HLA-E*01:01 + HLA-E*01:01 genotype against hepatitis C [18].
Consistently, Schulte et al. also showed that among patients with chronic HCV infection (with genotype 2 or 3), the HLA-ER/ HLA-ER (HLA-E*01:01 + HLA-E*01:01) genotype is significantly lower than those of healthy controls. This group also suggested that the HLA-ER/ HLA-ER genotype may also be related to the spontaneous clearance of HCV infection [27]. This finding might be to some extent relevant to our analysis where we showed that between individuals who were infected with SARS-CoV-2, HLA-E*01:01 + HLA-E*01:01 genotype was much significantly higher in those who experienced asymptomatic disease than symptomatic patients. Accordingly, based on HLA-E allelic variants, in a retrospective follow-up study on patients co-infected with HIV/ HCV viruses, Fulgencio et al. found an association between HLA-E*01:01 allele and increased odds of HCV clearance [28]. This study is somewhat similar to our findings that HLA-E*01:01 allele trends to increase the odds of being asymptomatic in those who infected by SARS-CoV-2.
The association of HLA-E polymorphisms with the status of HCV infection has been so far discussed from various mechanistic aspects. Given the similarities between the findings of this study and what has already been observed with hepatitis C, these mechanisms may also be extended to COVID-19, and it therefore seems useful to refer to them here. On one hand, some studies indicated that homozygous individuals for the HLA-E*01:01 allele may induce less NK cell inhibition by NKG2A due to their decreased expression of HLA-E on target cells. The phenomenon that leads to a stronger immune cytolytic effect against HCV infected cell in these individuals than those with homozygous HLA-E*01:03 allele, who express more stable HLA-E due to its greater affinity to peptide derived from the leader sequence of MHC class Ia [27], [28]. On the other hand, CD8+ T cell activity can also be inhibited through the interaction between HLA-E associated MHC class Ia peptides and NKG2A. However, due to the lower affinity of HLA-E*0101 for MHC-derived peptides, it may bind more efficiently to HCV-derived antigens and instead deliver them to CD8+ T cells and thus making infected cells more susceptible to the cytolysis by the acquired immune system [29], [30]. In general, HLA-E–antigen–specific recognition by T cell receptors (TCRs) usually compensates the failure of T cell mediated cytotoxicity where the down-regulation of MHC class Ia expression by human tumor or viruses facilitates their evasion from classical CD8+ CTL-mediated killing. This is a mechanism by which HLA-E–restricted unconventional CD8+ T cells have been reported to play an important role in recognition of antigenic peptides derived from different tumors and persistent pathogen infections, including human immunodeficiency virus (HIV), cytomegalovirus (CMV), and Mycobacterium tuberculosis (Mtb), Epstein Barr virus (EBV) and Salmonella typhi (2, 3) [31].
Similarly, it is postulated that in COVID-19 patients, HLA-E molecule expressed on the surface of infected cells, can also present SARS-CoV-2-derived peptides to CD8+ T cells with a same manner to be recognised by TCRs and lead to CTL-mediated killing of virus infected cells [32]. Intriguingly, in most recent study, Hammer et.al have shown that due to down-regulation of classical MHC class Ia during SARS-CoV-2 infection, reduced HLA-E access to these natural peptides causes this molecule to bind to viral peptides instead. In this regards, SARS-CoV-2 non-structural protein 13 (Nsp13) encodes for an HLA-E-restricted peptide, Nsp13232–240 that its complex with HLA-E abrogates inhibition of NKG2A+ NK cells which were supposed to naturally interact with HLA-E-associated MHC Ia peptide [33]. As a result, this abrogation leads to a missing self-recognition which allows more efficient cytolytic NK cell activation against infected cells.
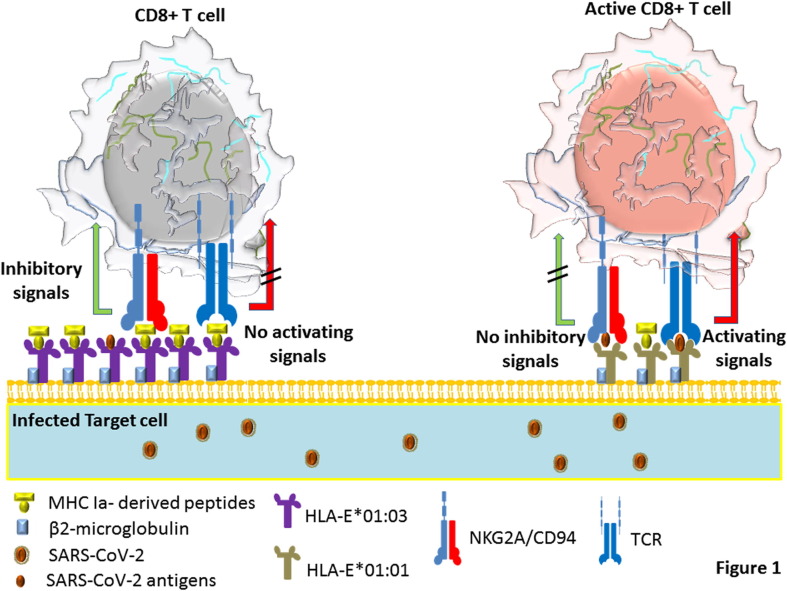
In summary, similar to what was previously suggested for other viral infection, the higher cellular expression of HLA-E*01:03 molecule and its greater affinity for MHC class Ia derived peptides compared to HLA-E*01:01 [34], [35] likely supports its more efficient interaction with NKG2A in COVID-19, which may limit both NK cells and CD8+ T cells mediated cytotoxicity against infected targets. In contrast, considering lower affinity of HLA-E*01:01 to MHC class Ia derived peptides, with the same scenario proposed for HCV infection [29], [30], this molecule may bind viral peptides more efficiently than HLA-E*01:03, especially when MHC class Ia is down-regulated by SARS-CoV-2. On the other word, as a possible mechanism for our study that HLA-E*01:01 + HLA-E*01:01 genotype confers less susceptibility to COVID-19, The possible higher affinity of HLA-E*01:01 for SARS-CoV-2 derived peptide may not only lead to better HLA-E restricted recognition of this peptide by TCR that activates conventional CD8 + T cells, but its less reactivity with NKG2A may abrogate NK cells or T cells inhibition leading to their enhanced cytotoxicity against SARS-CoV-2 infected cell (Fig. 1, Fig. 2 ). Although these speculations may provide some mechanistic explanations for why HLA-E*01:01 + HLA-E*01:01 genotype could better protect individuals against SARS-CoV-2 disease, extensive molecular and cellular studies are required in the future to exclusively evaluate the mentioned mechanisms in the COVID-19.
Fig. 1.
HLA-E polymorphisms and CD8 + T cell function against SARS-CoV-2 infection. Decreased accessibility of HLA-E to its required peptide due to SARS-CoV-2-induced down-regulation of MHC-Ia expression may divert its binding to the viral peptides. The presentation of these peptides to HLA-E-restricted CD8 + T cells through TCR instead of NKG2A induces T cell mediated cytolytic effect against SARS-CoV-2 infected cells. Here, HLA-E*01:03 which was already shown to have higher cellular expression and more affinity to MHC class Ia derived peptides than HLA-E*01:01 may further support the NKG2A-depebdent inhibition of CD8 + T cells cytotoxicity against SARS-CoV-2 infected cells. In contrast, despite its lower expression, the possible higher affinity of HLA-E*01:01 to viral peptide and its presentation to HLA-E-restricted CD8 + T cells may induce its cytolytic activity against SARS-CoV-2 in a higher level than HLA-E*01:03. This may support the observed reduced susceptibility of individuals with HLA-E*01:01 + HLA-E*01:01 genotype to COVID-19.
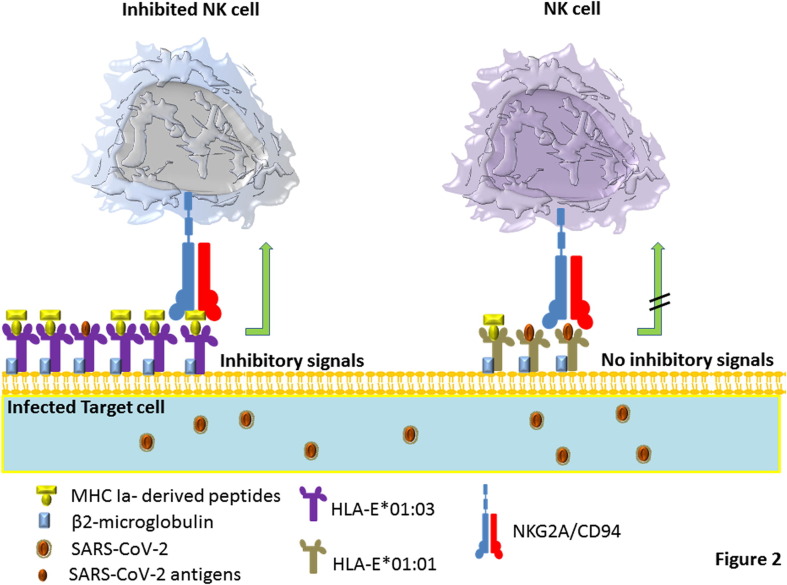
Fig. 2.
HLA-E polymorphisms and NK cell function against SARS-CoV-2 infection. SARS-CoV-2 infection down-regulates the expression of classical MHC class Ia. This decreases HLA-E access to its natural peptides required for the interaction with inhibitory receptor NKG2A on NK cell. Alternatively, HLA-E can bind to viral peptides instead, and since its complex with SARS-CoV-2 antigen cannot interact with NKG2A, the inhibitory function of NK cells is abrogated while the resultant missing self-recognition allows more efficient cytolytic activation against infected cells. Here, compared to HLA-E*01:03, the lower expression of HLA-E*01:01 and its less affinity to peptide derived from the leader sequence of MHC class Ia, shifts this molecule to become more complex with viral peptides, which makes this ligand less reactive to NKG2A. This may favour less NK cells inhibition in individuals with HLA-E*01:01 + HLA-E*01:01 genotype, leading to more potent cytolytic effects against SARS-CoV-2 infected cells.
5. Conclusion
Taken together given HLA-E polymorphisms, the data presented here may be useful to identify individuals more vulnerable to COVID-19 for better care and management. Especially, along with other risk factors in patients, having HLA-E*01:03 + HLA-E*01:03 genotype may also be associated with the possibility of severe cases of the disease. In this regard, according to our findings on HLA-E- based severity or susceptibility to disease, this study can be useful in reducing the risk of contracting COVID-19 in certain groups such as health care workers who are in constant and close contact with patients, or individuals with underlying diseases. It is noteworthy that even with the advent of various vaccines at this stage of the epidemic; our study could also be useful in providing recommendations on vaccination protocols and their intervals for vulnerable individuals.
Declaration of Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Acknowledgments
Acknowledgements
The authors wish to thank Dr. Faranak Etesam and Dr. Majid Moslemi from Tehran Blood Transfusion Center.
Availability of data and material.
The corresponding author can make available some dataset upon reasonable request.
Authors’ contributions
EH designed the methodology, performed the experiments, analyzed the data, performed the statistics and co-wrote the manuscript. AM collected sample, fulfilled the forms and helped with conducting the experiments. MG designed the study, wrote the manuscript, helped with data analysis, conducting experiments and acted as corresponding authors. AA acted as statistical consultant. AG depicted figures and wrote their legends. All authors reviewed the manuscript.
Funding
This work was part of Dr. Hosseini’s approved projects (No. 1118) supported by High Institute for Research and Education in Transfusion Medicine in Iran.
Ethics approval and consent to participate.
Ethics approval (IR.TMI.REC.1399.015.) for this study was obtained from the appropriate ethics committees of High Institute for Research and Education in Transfusion Medicine.
Consent for publication
Not applicable.
Competing interests
All authors have approved the final article and declare that they have no competing interests.
Footnotes
Supplementary data to this article can be found online at https://doi.org/10.1016/j.humimm.2023.02.002.
Appendix A. Supplementary data
The following are the Supplementary data to this article:
References
- 1.Acter T, Uddin N, Das J, Akhter A, Choudhury TR, Kim S: Evolution of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) as coronavirus disease 2019 (COVID-19) pandemic: A global health emergency. Science of the Total Environment 2020:138996. [DOI] [PMC free article] [PubMed]
- 2.Vidal S.M., Khakoo S.I., Biron C.A. Natural killer cell responses during viral infections: flexibility and conditioning of innate immunity by experience. Current opinion in virology. 2011;1:497. doi: 10.1016/j.coviro.2011.10.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Henderson L.A., Canna S.W., Schulert G.S., Volpi S., Lee P.Y., Kernan K.F., et al. On the alert for cytokine storm: Immunopathology in COVID-19. Arthritis & Rheumatology. 2020 doi: 10.1002/art.41285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lam V.C., Lanier L.L. NK cells in host responses to viral infections. Current opinion in immunology. 2017;44:43. doi: 10.1016/j.coi.2016.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Yao Z., Zheng Z., Wu K., Junhua Z. Immune environment modulation in pneumonia patients caused by coronavirus: SARS-CoV, MERS-CoV and SARS-CoV-2. Aging (Albany NY) 2020;12:7639. doi: 10.18632/aging.103101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Ghasemzadeh M., Hosseini E., Schwarer A.P., Pourfathollah A.A. NK cell maturation to CD56dim subset associated with high levels of NCRs overrides the inhibitory effect of NKG2A and recovers impaired NK cell cytolytic potential after allogeneic hematopoietic stem cell transplantation. Leukemia research. 2016;43:58. doi: 10.1016/j.leukres.2015.12.002. [DOI] [PubMed] [Google Scholar]
- 7.Nguyen S., Dhedin N., Vernant J.-P., Kuentz M., Jijakli A.A., Rouas-Freiss N., et al. NK-cell reconstitution after haploidentical hematopoietic stem-cell transplantations: immaturity of NK cells and inhibitory effect of NKG2A override GvL effect. Blood. 2005;105:4135. doi: 10.1182/blood-2004-10-4113. [DOI] [PubMed] [Google Scholar]
- 8.André P., Denis C., Soulas C., Bourbon-Caillet C., Lopez J., Arnoux T., et al. Anti-NKG2A mAb is a checkpoint inhibitor that promotes anti-tumor immunity by unleashing both T and NK cells. Cell. 2018;175:1731. doi: 10.1016/j.cell.2018.10.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Moser J.M., Gibbs J., Jensen P.E., Lukacher A.E. CD94-NKG2A receptors regulate antiviral CD8+ T cell responses. Nature immunology. 2002;3:189. doi: 10.1038/ni757. [DOI] [PubMed] [Google Scholar]
- 10.Lee N., Llano M., Carretero M., Ishitani A., Navarro F., López-Botet M., et al. HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proceedings of the National Academy of Sciences. 1998;95:5199. doi: 10.1073/pnas.95.9.5199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ulbrecht M., Couturier A., Martinozzi S., Pla M., Srivastava R., Peterson P.A., et al. Cell surface expression of HLA-E: interaction with human β2-microglobulin and allelic differences. European journal of immunology. 1999;29:537. doi: 10.1002/(SICI)1521-4141(199902)29:02<537::AID-IMMU537>3.0.CO;2-6. [DOI] [PubMed] [Google Scholar]
- 12.Hosseini E., Schwarer A.P., Ghasemzadeh M. Do human leukocyte antigen E polymorphisms influence graft-versus-leukemia after allogeneic hematopoietic stem cell transplantation? Experimental hematology. 2015;43:149. doi: 10.1016/j.exphem.2014.11.007. [DOI] [PubMed] [Google Scholar]
- 13.Yaqinuddin A., Kashir J. Innate immunity in COVID-19 patients mediated by NKG2A receptors, and potential treatment using Monalizumab, Cholroquine, and antiviral agents. Medical Hypotheses. 2020;109777 doi: 10.1016/j.mehy.2020.109777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Nattermann J., Nischalke H.D., Hofmeister V., Kupfer B., Ahlenstiel G., Feldmann G., et al. HIV-1 infection leads to increased HLA-E expression resulting in impaired function of natural killer cells. Antivir Ther. 2005;10:95. doi: 10.1177/135965350501000107. [DOI] [PubMed] [Google Scholar]
- 15.Maucourant C, Filipovic I, Ponzetta A, Aleman S, Cornillet M, Hertwig Let al. : Natural killer cell immunotypes related to COVID-19 disease severity. Science immunology 2020;5:eabd6832. [DOI] [PMC free article] [PubMed]
- 16.Ghasemzadeh M., Ghasemzadeh A., Hosseini E. Exhausted NK cells and cytokine storms in COVID-19: Whether NK cell therapy could be a therapeutic choice. Human immunology. 2021 doi: 10.1016/j.humimm.2021.09.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Prasetyo A.A., Dharmawan R., Irvan Raharjo H. Human leukocyte antigen-e alleles are associated with hepatitis c virus, torque teno virus, and toxoplasma co-infections but are not associated with hepatitis b virus, hepatitis d virus, and GB virus c co-infections in human immunodeficiency virus patients. Journal of Global Infectious Diseases. 2016;8:75. doi: 10.4103/0974-777X.182121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hosseini E., Kazerooni E.S., Azarkeivan A., Sharifi Z., Shahabi M., Ghasemzadeh M. HLA-E* 01: 01 allele is associated with better response to anti-HCV therapy while homozygous status for HLA-E* 01: 03 allele increases the resistance to anti-HCV treatments in frequently transfused thalassemia patients. Hum Immunol. 2022 doi: 10.1016/j.humimm.2022.04.010. [DOI] [PubMed] [Google Scholar]
- 19.Kanevskiy L., Erokhina S., Kobyzeva P., Streltsova M., Sapozhnikov A., Kovalenko E. Dimorphism of HLA-E and its Disease Association. International journal of molecular sciences. 2019;20:5496. doi: 10.3390/ijms20215496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pietra G., Romagnani C., Manzini C., Moretta L., Mingari M.C. The Emerging Role of HLA-E-Restricted CD8+ T Lymphocytes in the Adaptive Immune Response to Pathogens and Tumors. Journal of Biomedicine and Biotechnology. 2010;2010:8. doi: 10.1155/2010/907092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fazeli A., Sharifi S., Mohammadi S., Bahraini M., Arabkhazaeli A., Jelveh N., et al. The demographic and serological characteristics of COVID-19 convalescent plasma donors: Identification of basic criteria for optimal donor selection. Transfusion and Apheresis Science. 2021;103302 doi: 10.1016/j.transci.2021.103302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Arabkhazaeli A., Maghsudlu M., Mohammadi S., Eshghi P. Positive anti-SARS-CoV-2 rapid serological test results among asymptomatic blood donors. Transfusion Clinique et Biologique. 2022;29:24. doi: 10.1016/j.tracli.2021.08.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Slot E., Hogema B.M., Reusken C.B., Reimerink J.H., Molier M., Karregat J.H., et al. Low SARS-CoV-2 seroprevalence in blood donors in the early COVID-19 epidemic in the Netherlands. Nature communications. 2020;11:1. doi: 10.1038/s41467-020-19481-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Saeed S., Drews S., Pambrun C., Yi Q., Osmond L., O'Brien S. Tracking SARS-CoV-2 seroprevalence among Canadian blood donors. Transfusion medicine reviews. 2022;36:67. [Google Scholar]
- 25.Younas A., Waheed S., Khawaja S., Imam M., Borhany M., Shamsi T. Seroprevalence of SARS-CoV-2 antibodies among healthy blood donors in Karachi. Pakistan. Transfusion and Apheresis Science. 2020;59 doi: 10.1016/j.transci.2020.102923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Araújo R.C., Bertol B.C., Dias F.C., Debortoli G., Almeida P.H., Souza F.F., et al. HLA-E gene polymorphisms in chronic hepatitis C: Impact on HLA-E liver expression and disease severity. Human immunology. 2021;82:177. doi: 10.1016/j.humimm.2021.01.018. [DOI] [PubMed] [Google Scholar]
- 27.Schulte D., Vogel M., Langhans B., Krämer B., Körner C., Nischalke H.D., et al. The HLA-ER/HLA-ER Genotype Affects the Natural Course of Hepatitis C Virus (HCV) Infection and Is Associated with HLA-E–Restricted Recognition of an HCV-Derived Peptide by Interferon-γ–Secreting Human CD8+ T Cells. The Journal of infectious diseases. 2009;200:1397. doi: 10.1086/605889. [DOI] [PubMed] [Google Scholar]
- 28.Guzmán-Fulgencio M., Berenguer J., Rallon N., Fernández-Rodríguez A., Miralles P., Soriano V., et al. HLA-E variants are associated with sustained virological response in HIV/hepatitis C virus-coinfected patients on hepatitis C virus therapy. AIDS. 2013;27:1231. doi: 10.1097/QAD.0b013e32835f5b9c. [DOI] [PubMed] [Google Scholar]
- 29.Pietra G., Romagnani C., Mazzarino P., Falco M., Millo E., Moretta A., et al. HLA-E-restricted recognition of cytomegalovirus-derived peptides by human CD8+ cytolytic T lymphocytes. Proceedings of the National Academy of Sciences. 2003;100:10896. doi: 10.1073/pnas.1834449100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pietra G., Romagnani C., Manzini C., Moretta L., Mingari M.C. The emerging role of HLA-E-restricted CD8+ T lymphocytes in the adaptive immune response to pathogens and tumors. Journal of Biomedicine and Biotechnology. 2010;2010 doi: 10.1155/2010/907092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Ottenhoff T.H., Joosten S.A. Mobilizing unconventional T cells. Science. 2019;366:302. doi: 10.1126/science.aay7079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Caccamo N., Sullivan L.C., Brooks A.G., Dieli F. Harnessing HLA-E-restricted CD8 T lymphocytes for adoptive cell therapy of patients with severe COVID-19. British journal of haematology. 2020;190:e185. doi: 10.1111/bjh.16895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hammer Q., Dunst J., Christ W., Picarazzi F., Wendorff M., Momayyezi P., et al. SARS-CoV-2 Nsp13 encodes for an HLA-E-stabilizing peptide that abrogates inhibition of NKG2A-expressing NK cells. Cell reports. 2022;38 doi: 10.1016/j.celrep.2022.110503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Strong RK, Holmes MA, Li P, Braun L, Lee N, Geraghty DE: HLA-E allelic variants. Correlating differential expression, peptide affinities, crystal structures, and thermal stabilities. J Biol Chem 2003;278:5082. [DOI] [PubMed]
- 35.Maier S., Grzeschik M., Weiss E.H., Ulbrecht M. Implications of HLA-E allele expression and different HLA-E ligand diversity for the regulation of NK cells. Hum Immunol. 2000;61:1059. doi: 10.1016/s0198-8859(00)00190-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.