Abstract
OBJECTIVE:
To objectively reveal the relationship between tongue coating microbes and bitter taste, sticky and greasy taste in chronic atrophic gastritis (CAG) patients.
METHODS:
16S rRNA high-throughput sequencing was used to detect bacterial diversity and community composition of tongue coating microbes from samples of CAG patients. LEfSe algorithm was used for discovering the different tongue coating microbes in CAG patients with or without bitter taste, also that in CAG patients with or without sticky and greasy taste.
RESULTS:
We respectively compared the features of tongue coating microbes in bitter taste, sticky and greasy taste of CAG patients. At the genus level, 25 tongue coating microbes were significantly different in CAG patients with bitter taste or without bitter taste; 17 tongue coating microbes were significantly different in CAG patients with sticky and greasy taste or without sticky and greasy taste. Campylobacter and Rothia were closely related to CAG patients with bitter taste. Enterococcus, Serratia, Leptotrichia and Selenomonas were closely related to CAG patients with stick and greasy taste.
CONCLUSIONS:
Campylobacter and Rothia possibly contribute to bitter taste of CAG patients, and Enterococcus, Serratia, Leptotrichia and Selenomonas contribute to stick and greasy taste of CAG patients, which is potential for the diagnosis and treatment of CAG.
Keywords: gastritis, atrophic; taste; Campylobacter; Enterococcus; Serratia; Leptotrichia; Selenomonas
1. INTRODUCTION
Chronic atrophic gastritis (CAG) is considered as a precancerous lesion of GC.1 The incidence of CAG in china is about 25.8%.2
The tongue manifestations are a unique diagnostic method in Traditional Chinese Medicine (TCM), and are considered as an external feature of the spleen and stomach.3 Moreover, our previous study4 confirmed that tongue manifestations could classify different TCM subtypes in CAG patients. Recently, more and more researches have investigated the relationship between tongue coating microbes and diseases, such as gastritis,5 CAG,6 GC,7 pancreatic head cancer,8 etc. Therefore, tongue coating microbes play a critical role in human diseases. Furthermore, tongue coating microbes can reflect the mechanism of symptom development. Bitter taste, sticky and greasy taste are common symptoms of CAG patients in the oral cavity. The two symptoms sometimes appear alone or together. However, the detailed mechanism on the tongue coating microbes contributing symptoms such as bitter taste, sticky and greasy taste remains unclear. Whether the two symptoms are caused by the same or different tongue coating microbes remains unclear.
We collected the tongue coating samples of 34 CAG patients. 16S rRNA analysis have shown that tongue coating microbes could be objectively revealed the production mechanisms of bitter taste, sticky and greasy taste in CAG.
2. MATERIALS AND METHODS
2.1. Sample collection
Thirty-four CAG patients who received an endoscopic examination in Shanghai University of TCM affiliated Longhua Hospital and Shuguang Hospital were recruited. The clinical symptoms were recorded using the questionnaire of spleen (stomach) disease. The histological assessment was done by the experienced pathologists following clinical guidelines according to “the updated Sydney System”.9 The inclusion criteria were a confirmed diagnosis of CAG according to pathological examination. This study was approved by the Medical Ethical Committee (2020-834-41-01). All participants signed the informed consent.
2.2. Tongue coating collection
Before collecting, the participants rinsed mouth with saline 3 times. Tongue-coating swabs were used to collect tongue coating of participants. Tongue coating was scraped with swabs from the root to the tip 3 times, the swabs were put into a 2 mL centrifuge tube, then all the tubes were stored at -80 ℃ until analysis.
2.3. DNA extraction and PCR amplification
Microbial community genomic DNA was extracted from 34 samples using the E.Z.N.A.® soil DNA Kit (Omega Bio-tek, Norcross, GA, USA) according to manufacturer’s instructions. The DNA extract was checked on 2% agarose gel, and DNA concentration and purity were determined with NanoDrop 2000 UV-vis spectrophotometer (Thermo Scientific, Wilmington, DE, USA). The hypervariable region V3-V4 of the bacterial 16S rRNA gene were amplified with primer pairs 338 F (5'-ACTCCTACGGGAGGCAGCAG-3') and 806 R (5'-GGACTACHVGGGTWTCTAAT-3') by an ABI GeneAmp® 9700 PCR thermocycler (ABI, Madison, WI, USA). The PCR amplification of 16S rRNA gene was performed. PCR reactions were performed in triplicate. The PCR product was extracted from 2% agarose gel and purified using the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, Union City, CA, USA) according to manufacturer’s instructions and quantified using Quantus™ Fluorometer (Promega, Waltham, MA, USA).
2.4. Illumina MiSeq sequencing
Purified amplicons were pooled in equimolar and paired-end sequenced on an Illumina MiSeq PE300 platform (Illumina, San Diego, CA, USA) according to the standard protocols by Majorbio Bio-Pharm Technology Co. Ltd. (Shanghai, China). The raw reads were deposited into the NCBI Sequence Read Archive (SRA) database (Accession Number: PRJNA764325-SRP337893).
2.5. Processing of sequencing data
The raw 16S rRNA gene sequencing reads were demultiplexed, quality-filtered by fastp version 0.20.0 10 and merged by FLASH version 1.2.7.11
Operational taxonomic units (OTUs) with 97% similarity cutoff12,13 were clustered using UPARSE version 7.1,12 and chimeric sequences were identified and removed. The taxonomy of each OTU representative sequence was analyzed by RDP Classifier version 2.2 14 against the 16S rRNA database (eg. Silva v138) using confidence threshold of 0.7.
2.6. Data processing and statistical analysis
The data were analyzed on the online platform of Majorbio Cloud Platform (www.majorbio.com). LEfSe algorithm was used for discovering significantly different tongue coating microbes from the phylum to the genus level; Spearman analysis was used to calculate the correlation between the symptoms of bitter taste, sticky and greasy taste and tongue coating microbes. The functional prediction of tongue coating microbes was performed using the PICRUSt software package. P < 0.05 was considered statistically significant.
3. RESULTS
3.1. Characteristics of patients with CAG
The average age of 34 patients with CAG was (58 ± 10) years old (age range 36-74), including 14 males and 20 females, male: female = 1:1.43. The average age of male was (57 ± 9) years old and BMI was 22.9 ± 1.3. The average age of female was (58 ± 10) years old and BMI was 20.8± 2.9. 3 CAG patients with Helicobacter pylori infection were diagnosed according to pathological examination.
The clinical symptoms of 34 CAG patients were counted, and the order of frequency was stomach distention (26 cases, 76.47%), bitter taste, (25 cases, 73.53%), belching (22 cases, 64.71%), ungratifying defecation (16 cases, 47.06%), pantothenic acid (12 cases, 35.29%), sticky and greasy taste (10 cases, 29.41%), heavy limbs (10 cases, 29.41%), yellow urine (10 cases, 29.41%), stomach pain (9 cases, 26.47%), dry stool (6 cases, 17.65%).
3.2. Analysis of bitter taste and tongue coating microbes
3.2.1. Comparison on tongue coating microbes in CAG patients with or without bitter taste
We compared the tongue coating microbes between 25 CAG patients with bitter taste and 9 CAG patients without bitter taste. At the genus level, as shown in Figure 1, there were 11 tongue coating microbes relatively enriched in CAG patients with bitter taste, including Campylobacter, Alloprevotella, Selenomonas, F0058, norank_f__norank_o__Clostridia_UCG-014, Catonella, Lachnoanaerobaculum, Megasphaera, unclassified_f__Veillonellaceae, Candidatus_Sacchar-imonas, Treponema. There were 14 tongue coating microbes relatively enriched in CAG patients without bitter taste, including Actinomyces, Parascardovia, Rothia, Peptostreptococcus, Micro-bacterium, Propioni-bacterium, Streptomyces, DNF-00809, Desulfobulbus, Paenibacillus, Blautia, Pajaro-ellobacter, Reyranella, Marinobacter as demonstrated in Figure 1B.
Figure 1. Comparison on tongue coating microbes in CAG patients with or without bitter taste.
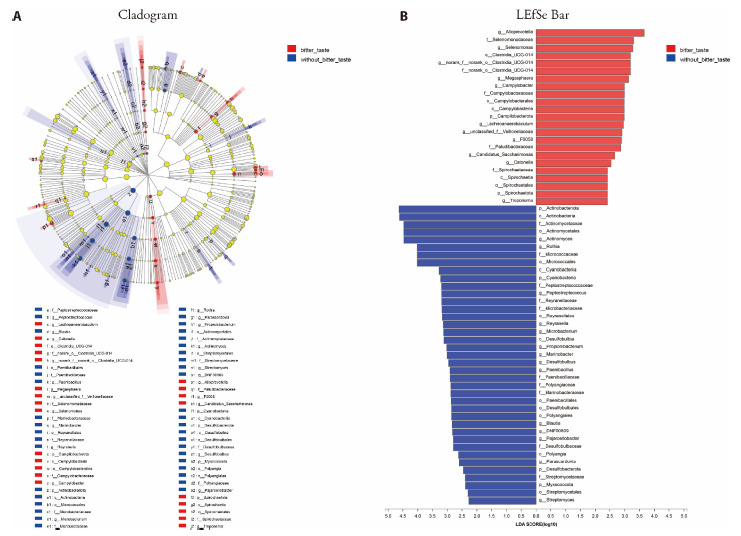
A: cladogram plot. Red nodes indicated significantly enriched bacterial colony with significant impact in CAG patients with bitter taste, and blue nodes indicated significantly enriched bacterial colony with significant impact in CAG patients without bitter taste. Light yellow nodes indicated bacterial colony without significant difference in both CAG patients with bitter taste and without bitter taste. B: LDA discriminant analysis. Red bar represented the bacterial colony enriched in CAG patients with bitter taste; blue bar represented the bacterial colony enriched in the CAG patients without bitter taste. CAG: chronic atrophic gastritis; LDA: linear discriminative analysis.
3.2.2. Correlation between tongue coating microbes and bitter taste
We carried out spearman correlation analysis between the symptom of bitter taste and 287 tongue coating microbes at the genus level.
As shown in Figure 2, 11 genera were positively correlated with bitter taste, including Campylobacter (P < 0.001, R = 0.615), Alloprevotella (P = 0.001, R = 0.561), Selenomonas (P = 0.001, R = 0.544), unclassified_ f_ Veillonellaceae (P = 0.004, R = 0.483), Megasphaera (P = 0.006, R = 0.459), Lachnoanaerobaculum (P = 0.018, R = 0.404), norank_f_norank_o_Clostridia_ UCG-014 (P = 0.028, R = 0.377), F0058 (P = 0.030,R = 0.373), Candidatus_Saccharimonas (P = 0.034, R = 0.364), Treponema (P = 0.037, R = 0.359), Catonella (P = 0.039, R = 0.356); 15 genera were negatively correlated with bitter taste, including Parascardovia (P = 0.002, R = -0.518), Rothia (P = 0.01, R = -0.432), Peptostreptococcus (P = 0.012, R = -0.425), Marinobacter (P = 0.014, R = -0.417), Reyranella (P = 0.014, R = -0.417), Blautia (P = 0.014, R = -0.416), DNF00809 (P = 0.014, R = -0.416), Microbacterium (P = 0.014, R = -0.416), Paenibacillus (P = 0.014, R = -0.416), Pajaroe-llobacter (P = 0.014, R = -0.416), Actinomyces (P = 0.016, R = -0.411), Streptomyces (P = 0.017, R = -0.407), Propi-oniba-cterium (P = 0.040, R = -0.355), Bacteroides (P = 0.041, R = -0.452), Desulfobulbus (P = 0.048, R = -0.342).
Figure 2. Heatmap of spearman correlation between tongue coating microbes and bitter taste.

A: twenty-six genera and bitter taste. The horizontal coordinates in the graph was tongue coating microbes, the vertical coordinates were the symptom of bitter taste. Red indicated positive correlation, blue indicated negative correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001). B: eleven positively correlated genera and bitter taste. The horizontal coordinates in the graph was tongue coating microbes, the vertical coordinates were the symptom of bitter taste. Red indicated a strong positive correlation, blue indicated a weak positive correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001). C: fifteen negatively correlated genera and bitter taste. The horizontal coordinates in the graph was tongue coating microbes, the vertical coordinates were the symptom of bitter taste. Red indicated weak negative correlation and blue indicated strong negative correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001).
3.2.3. Functional prediction of tongue coating microbes related to bitter taste
At the genus level, Kyoto Encyclopedia of Genes and Genomes (KEGG) functional prediction was performed on the tongue coating microbes enriched in the group with bitter taste. At the same time, 42 KEGG pathways (level 2) and abundance value were obtained. In descending order of abundance value, the top 10 pathways were Global and overview maps, Carbohydrate metabolism, Amino acid metabolism, Metabolism of cofactors and vitamins, Energy metabolism, Translation, Replication and repair, Nucleotide metabolism, Membrane transport, Glycan biosynthesis and metabolism (Figure S1A).
Furthermore, at the genus level, the KEGG functional prediction was performed on the 14 tongue coating microbes enriched in the group without bitter taste, and a total of 44 KEGG pathways (level 2) and abundance value were obtained. In descending order of abundance, the top 10 pathways were Global and overview maps, Carbohydrate metabolism, Amino acid metabolism, Energy metabolism, Metabolism of cofactors and vitamins, Translation, Replication and repair, Nucleotide metabolism, Membrane transport, Lipid metabolism (Figure S1B).
3.3. Analysis of sticky and greasy taste and tongue coating microbes
3.3.1. Comparison on tongue coating microbes in CAG patients with or without sticky and greasy taste
We compared the tongue coating microbes between 10 CAG patients with sticky and greasy taste and 24 CAG patients without sticky and greasy taste. At the genus level, as shown in Figure 3A, there were 4 tongue coating microbes relatively enriched in CAG patients with sticky and greasy taste, including Serratia, Enterococcu, DNF00809, Enhydrobacter. There were 13 tongue coating microbes relatively enriched in CAG patients without sticky and greasy taste, including Leptotrichia, Fusobacterium, Selenomonas, Eubacterium_saphenum_ group, Aggregatibacter, Mycoplasma, Comamonas, Catonella, Lautropia, Treponema, Fretibacterium, norank_f_norank_o_norank_c_Gracilibacteria, Lachn-oanaerobaculum, as shown in Figure 3B.
Figure 3. Comparison on tongue coating microbes in CAG patients with or without sticky and greasy taste.

A: cladogram plot. red nodes indicated significantly enriched bacterial colony with significant impact in CAG patients with sticky and greasy taste, and blue nodes indicated significantly enriched bacterial colony with significant impact in CAG patients without sticky and greasy taste. light yellow nodes indicated bacterial colony without significant difference in both CAG patients with or without sticky and greasy taste. B: LDA discriminant analysis. red bar represented the bacterial colony enriched in CAG patients with sticky and greasy taste; blue bar represented the bacterial colony enriched in CAG patients with sticky and greasy taste. CAG: chronic atrophic gastritis; LDA: linear discriminative analysis.
3.3.2. Correlation between tongue coating microbes and sticky and greasy taste
We carried out spearman correlation analysis between the symptom of sticky and greasy taste and 287 tongue coating microbes at the genus level. At the same time, we drew the correlation heatmap.
As shown in Figure 4, 4 genera were positively correlated with sticky and greasy taste, including Enterococcus (P = 0.024, R = 0.387), Enhydrobacter (P = 0.024, R = 0.387), DNF00809 (P = 0.024, R = 0.387), Serratia (P = 0.028, R = 0.376). 13 genera were nega-tively correlated with sticky and greasy taste, including Aggregatibacter (P = 0.001, R = -0.528), Freti-bacterium (P = 0.002, R = -0.517), Lachno-anaerobaculum (P = 0.004, R = -0.484), Catonella(P = 0.008, R = -0.448), norank_f_norank_o_norank_ c_Gracilibacteria (P = 0.014,R = -0.417), Seleno-monas (P = 0.023, R =-0.388), Eubacterium_ saphenum_group (P = 0.026, R = -0.382), Fusobac-terium (P = 0.030, R = -0.372), Leptotrichia (P = 0.032, R = -0.369), Mycoplasma (P = 0.040, R = -0.354), Treponema (P = 0.044, -0.348), Comamonas (P = 0.046, R = -0.345), Lautropia (P = 0.049, R = -0.341).
Figure 4. Heatmap of the spearman correlation between tongue coating microbes and sticky greasy taste.

A: seventeen genera and sticky greasy taste: The horizontal coordinates in the graph was tongue coating microbes, the vertical coordinates were the symptom of sticky greasy taste. Red indicated positive correlation, blue indicated negative correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001). B: four positively correlated genera and sticky greasy taste. The horizontal coordinates were tongue coating microbes; the vertical coordinates were the symptom of sticky greasy taste. Red indicated a strong positive correlation, blue indicates a weak positive correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001). C: 13 negatively correlated genera and sticky greasy taste. The horizontal coordinates in the graph was tongue coating microbes, the vertical coordinates were the symptoms of sticky greasy taste. Red indicated weak negative correlation and blue indicates strong negative correlation. The darker color indicated stronger correlation. The magnitude of significant P values was marked with a, b, c (0.01 < aP ≤ 0.05, 0.001 < bP ≤ 0.01, cP ≤ 0.001).
3.3.3. functional prediction of tongue coating microbes related to sticky and greasy taste
At the genus level, KEGG functional prediction was performed on the tongue coating microbes enriched in the group of sticky and greasy taste. At the same time, a total of 42 KEGG pathways (Level 2) and abundance value were obtained. In descending order of abundance value, the top 10 pathways were Global and overview maps, Carbohydrate metabolism, Amino acid metabolism, Membrane transport, Metabolism of cofactors and vitamins, Energy metabolism, Signal transduction, Cellular community-prokaryotes, Nucle-otide metabolism, Xenobiotics biodegradation and metabolism (Figure S2A).
At the genus level, the KEGG functional prediction was performed on the 13 tongue coating microbes enriched in the group without sticky and greasy taste, and 42 KEGG pathways (Level 2) and abundance values were obtained. In descending order of abundance, the top 10 pathways were Global and overview maps, Carbohydrate metabolism, Amino acid metabolism, Membrane transport, Metabolism of cofactors and vitamins, Energy metabolism, Translation, Replication and repair, Nucleotide metabolism, Cellular community- prokaryotes (Figure S2B).
4. DISCUSSION
In this study, 16S rRNA high-throughput sequencing was used to investigate the variation of tongue coating microbes in CAG patients with bitter taste, sticky and greasy taste. This study provided reference for clarifying the production mechanism of bitter taste, sticky and greasy taste of CAG patients on tongue coating microbes.
The tongue manifestations are the external signs of internal organs according to the theory of TCM, which like a mirror to show changes in internal organs of Zang and Fu. However, the tongue coating is the external signs most closely associated with the Spleen and Stomach in all organs of Zang and Fu.3 Several studies have proved the relationship between tongue coating microbes and the occurrence of gastritis. There were 123 and 258 species-level OTUs enriched in the tongue coating of gastritis with Cold Syndrome, or with Hot Syndrome, respectively.5 Bacillus was present only in the yellow tongue coating of patients with chronic erosive gastritis.15 Microbial components of tongue coating in CAG patients were significantly different from chronic non-atrophic gastritis patients and healthy people.6 In CAG patients, there were 9 tongue coating microbes in genus namely Streptococcus, Veillonella, Leptotrichia, Prevotella, Rothia, Stomatobaculum, Lachnoana-erobaculum, Solobacterium, Unidentifiedetc,16 signi-ficantly different from healthy people. Contents of dominant microbes (Streptococcus, Prevotella-7, Moraxell) changed in CAG patients with greasy tongue coating group.17
Campylobacter, belonging to Gram-negative bacteria, is one of the main factors inducing human bacterial gastroenteritis.18
Campylobacter concisus and Campylobacter rectus in species were associated with the Hot Syndrome of gastritis in TCM.19 In addition, tongue coating microbes of Campylobacter in genus was potential related with bitter taste in gastritis patients.20 Moreover, TNF which can be induced by Campylobacter concisus was associated with bitter taste.21 Our study also found Campylobacter was significantly enriched and related in CAG patients with bitter taste.
In TCM, many factors22,23 are involved in the pathogenesis of bitter taste, such as liver-gallbladder damp-heat, liver depression transforming into fire, up-flaming of gallbladder fire, damp-heat in the stomach and intestines, and heat fire flaming upward. Researchers found that the increased bile acids might be the main theoretical basis of bitter taste. Bitter taste is related to salivary bile acids migrating from the blood.24 Firmicutes in the gut could positively regulate bile acid (BA) metabolism.25 In our results, several genera of tongue coating such as Selenomonas, norank_f__norank_o__ Clostridia_UCG-014, Catonella, Lachnoanaero-baculum, Megasphaera and unclassified_f__Veillone-llaceae belong to Firmicutes were enriched. Whether tongue coating microbes belong to Firmicutes mediate the bitter taste through modulating BA metabolites and enhancing BAs migrating from blood into saliva of CAG patients, need to be confirmed in our further study.
The abundance of Rothia in the tongue coating of liver-fire hyperactivity syndrome in hypertensive patients with bitter taste was significantly higher than that of the healthy people.26 However, in our study, Rothia enriched in CAG patients without bitter taste, indicating that the different tongue coating microbes might mediate bitter taste in different diseases, and possible many factors are involved in the bitter taste.
At the genus level, we found that 4 tongue coating microbes were relatively enriched and related in CAG patients with sticky and greasy taste, including Serratia, Enterococcus, DNF00809 and Enhydrobacter. The Enterococcus,27 belonging to ubiquitous Gram-positive bacteria, is a common opportunistic pathogen in the oral cavity and gastrointestinal tract, which is also the leading cause of health care-associated infections (HAIs) globally.
Serratia 28 is an opportunistic pathogen known to cause an array of infectious presentations including pneumonia, wound infections, skin and soft tissue infections.
Enterococcus and Serratia increase the inflammatory response of the body, thereby leading to decreased salivary gland function and slow salivary flow.29 The saliva viscosity of chronic gastritis patients is significantly higher than that of healthy people, and the saliva flow rate is significantly reduced.30
At the same time, we found that 13 tongue coating microbes were not enriched and related in CAG patients with sticky and greasy taste, such as Leptotrichia and Selenomonas. Leptotrichia, belonging to Gram-negative bacteria, inhabits the oral cavity, intestines, urinary system, and female genital tract of humans,31 and can ferment carbohydrates and produce lactic acid.32 Selenomonas was compatible with periodontal health,33 and related to glucose metabolism.34 Leptotrichia and Selenomonas can decompose carbohydrates and glucose of food. So, the digestive function of CAG patients with sticky and greasy taste may be weaken and enhance the retention of food residues in the oral cavity. Thus, our findings indicated that these above bacteria may increase the viscosity of saliva and the retention of food residue in the oral cavity, finally resulted in the symptoms of sticky and greasy taste.
According to the above results, we can see that the tongue coating microbes related to the symptoms of bitter taste, sticky and greasy taste were completely different. However, based on the functional prediction results, it can be seen that the functions of the tongue coating microbes related to the symptoms of bitter taste, sticky and greasy taste were mostly same, and they were all involved in Carbohydrate metabolism, Amino acid metabolism, Energy metabolism.
The reasons of functional prediction may be that 34 patients belong to CAG disease and had the clinical manifestations of yellow and greasy tongue coating. The theory in TCM believes that yellow and greasy tongue coating is the reaction of damp-heat accumulation in the spleen and stomach. Therefore, the results of functional prediction may reflect the characteristics of CAG diseases and syndromes. However, the in-depth mechanism such as bitter taste, sticky and greasy taste needs to be further explored combing with other technologies.
In summary, our findings demonstrated that Campylobacter and Rothia were related to CAG patients with bitter taste; Enterococcus, Serratia, Leptotrichia and Selenomonas were related to CAG patients with stick and greasy taste. The results indicated that the mechanisms of bitter taste, stick and greasy taste caused by tongue coating microbes were completely different. Tongue coating microbes might be objective, sensitive and non-invasive methods for CAG diagnosis with different symptoms in clinic, which should be further investigated in more bigger sample set. However, in the present study, the limitation was that the distribution of symptoms of CAG patients is uneven, and the sample set should be expanded for in-depth research in the future study.
Figure S1. Heatmap of KEGG function on the tongue coating microbes enriched in the group with or without bitter taste.
A: tongue coating microbes enriched in the group with bitter taste; B: the tongue coating microbes enriched in the group without bitter taste. The horizontal coordinates were the name of group, and the vertical coordinates was the name of functional pathway in level 2. The red color block represented the abundance value was high, and blue color represented the abundance value was low. KEGG: Kyoto Encyclopedia of Genes and Genomes.
Figure S2. Heatmap of KEGG function on the tongue coating microbes enriched in the group with or without sticky and greasy taste.
A: tongue coating microbes enriched in the group with sticky and greasy taste; B: tongue coating microbes enriched in the group without sticky and greasy taste. The horizontal coordinates were the name of group, and the vertical coordinates was the name of functional pathway in level 2. The red color block represented the abundance value was high, and blue color represented the abundance value was low. KEGG: Kyoto Encyclopedia of Genes and Genomes.
Contributor Information
Guoping LIU, Email: 13564133728@163.com.
Yu ZHENG, Email: zhengyu2050@163.com.
REFERENCES
- 1. Park YH, Kim N. Review of atrophic gastritis and intestinal metaplasia as a premalignant lesion of gastric cancer. J Cancer Prev 2015; 20: 25-40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Du Y, Bai Y, Xie P, et al. Chronic gastritis in China: a national multicenter survey. BMC gastroenterology 2014; 14: 1-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Wang YQ. Diagnostics of Traditional Chinese Medicine. Beijing: Gao Deng Jiao Yu Chu Ban She, 2016: 28-9. [Google Scholar]
- 4. Zhu CM, Gu WJ, Yang DC, et al. Study on the characteristics of TCM tongue image in patients with chronic ttrophic gastritis. Shi Jie Ke Xue Ji Shu-Zhong Yi Yao Xian Dai Hua 2020; 22: 1595-600. [Google Scholar]
- 5. Jiang B, Liang X, Chen Y, et al. Erratum: integrating next-generation sequencing and traditional tongue diagnosis to determine tongue coating microbiome. Sci Rep 2012; 2: 344-67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Li H, Wang WH, Guo CR, et al. Analysis of microbial community structure of tongue coating at different taxonomic levels. Zhong Hua Zhong Yi Yao Za Zhi 2020; 35: 6298-301. [Google Scholar]
- 7. Hu J, Han S, Chen Y, et al. Variations of tongue coating micro-biota in patients with gastric cancer. Biomed Res Int 2015; 1-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Lu H, Ren Z, Li A, et al. Tongue coating microbiome data distinguish patients with pancreatic head cancer from healthy controls. J Oral Microbiol 2019; 11: 1-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Dixon MF, Genta RM, Yardley JH, et al. Classification and grading of gastritis. The updated Sydney System. International Workshop on the Histopathology of Gastritis, Houston 1994. Am J Surg Pathol 1996; 20: 1161-81. [DOI] [PubMed] [Google Scholar]
- 10. Chen S, Zhou Y, Chen Y, et al. FASTP: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018; 34: i884-90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Magoč T, Salzberg SL. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 2011; 27: 2957-63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Edgar RC. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 2013; 10: 996-8. [DOI] [PubMed] [Google Scholar]
- 13. Stackebrandt E, Goebel BM. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 1994; 44: 846-9. [Google Scholar]
- 14. Wang Q, Garrity GM, Tiedje JM, et al. Naive bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 2007; 73: 5261-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Ye J, Cai X, Yang J, et al. Bacillus as a potential diagnostic marker for yellow tongue coating. Sci Rep 2016; 6: 1-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Wang HW, Li L, Li FF, et al. Specific bacteria of tongue coating in patients with chronic atrophic gastritis. Zhong Guo Wei Sheng Tai Xue Za Zhi 2018; 30: 497-501. [Google Scholar]
- 17. Wang HW, Li L, Li FF, et al. Study on the compositions of dominant bacteria and determination of E-adhesion factor and ICAM-1 in chronic atrophic gastritis. Zhong Guo Zhong Xi Yi Jie He Za Zhi 2018; 38:790-4. [Google Scholar]
- 18. O'Brien SJ. The consequences of campylobacter infection. Curr Opin Gastroenterol 2017; 33: 14-20. [DOI] [PubMed] [Google Scholar]
- 19. Li R, Ma T, Gu J, et al. Imbalanced network biomarkers for Traditional Chinese Medicine syndrome in gastritis patients. Sci Rep 2013; 3: 1-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Cui J, Cui H, Yang M, et al. Tongue coating microbiome as a potential biomarker for gastritis including precancerous cascade. Protein Cell 2019; 10: 496-509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Feng P, Jyotaki M, Kim A, et al. Regulation of bitter taste responses by tumor necrosis factor. Brain Behav Immunol 2015; 49: 32-42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Zhang K, Kang JY, Chang RH, et al. Discussion on the pathogenesis of bitter taste in mouth. Zhejiang Zhong Yi Yao Da Xue Xue Bao 2019; 43: 653-6. [Google Scholar]
- 23. Si P, Fu Y, Ni S, et al. Epidemiological survey on prevalence and associated risk factors of bitter taste among inpatients from four grade 3A hospitals in Beijing. J Trad Chin Med Sci 2017; 4: 31-8. [Google Scholar]
- 24. Chen F, Fu YL, Zhang JJ. Experimental analysis of the correlation between bitter taste in liver disease and bile acid from oral saliva. Zhong Guo Shi Yong Nei Ke Za Zhi 2001; 21: 365-6. [Google Scholar]
- 25. Jiang T, Xu C, Liu H, et al. Linderae radix ethanol extract alleviates diet-induced hyperlipidemia by regulating bile acid metabolism through gut microbiota. Front Pharmacol 2021; 12: 1-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Luo JW, Lin CH, Zhu YB, et al. Association of tongue bacterial flora and subtypes of liver-fire hyperactivity syndrome in hypertensive patients. Evid Based Complement Alternat Med 2018; 2018: 1-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. García-Solache M, Rice LB. The enterococcus: a model of adaptability to its environment. Clin Microbiol Rev 2019; 32: e00058-18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Gupta V, Sharma S, Pal K, et al. Serratia, No longer an uncommon opportunistic pathogen - case series & review of literature. Infect Disord Drug Targets 2021; 21: e300821191666. [DOI] [PubMed] [Google Scholar]
- 29. Correia PN, Carpenter GH, Osailan SM, et al. Acute salivary gland hypofunction in the duct ligation model in the absence of inflammation. Oral Dis 2008; 14: 520-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Tong NN. Preliminary study on saliva metabolome of dampness-heat accumulation of spleen in chronic gastritis. Guangzhou: Guangzhou TCM University, 2016: 25. [Google Scholar]
- 31. Eribe ERK, Paster BJ, Caugant DA, et al. Genetic diversity of Leptotrichia and description of Leptotrichia goodfellowii sp. nov. Int J Syst Evol Microbiol 2004; 54: 583-92. [DOI] [PubMed] [Google Scholar]
- 32. Eribe ERK, Olsen I. Leptotrichia species in human infections II. J Oral Microbiol 2017; 9: 1-31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Mcdaniel J, Mcdaniel S, Samiano B J, et al. Microbial screening reveals oral site-specific locations of the periodontal pathogen selenomonas noxia. Curr Issues Mol Biol 2021; 43: 353-64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Melville SB, Michel TA, Macy JM. Pathway and sites for energy conservation in the metabolism of glucose by selenomonas ruminantium. J Bacteriol 1988; 170: 5298-304. [DOI] [PMC free article] [PubMed] [Google Scholar]




