Abstract
The genome sequence of the solvent-producing bacterium Clostridium acetobutylicum ATCC 824 has been determined by the shotgun approach. The genome consists of a 3.94-Mb chromosome and a 192-kb megaplasmid that contains the majority of genes responsible for solvent production. Comparison of C. acetobutylicum to Bacillus subtilis reveals significant local conservation of gene order, which has not been seen in comparisons of other genomes with similar, or, in some cases closer, phylogenetic proximity. This conservation allows the prediction of many previously undetected operons in both bacteria. However, the C. acetobutylicum genome also contains a significant number of predicted operons that are shared with distantly related bacteria and archaea but not with B. subtilis. Phylogenetic analysis is compatible with the dissemination of such operons by horizontal transfer. The enzymes of the solventogenesis pathway and of the cellulosome of C. acetobutylicum comprise a new set of metabolic capacities not previously represented in the collection of complete genomes. These enzymes show a complex pattern of evolutionary affinities, emphasizing the role of lateral gene exchange in the evolution of the unique metabolic profile of the bacterium. Many of the sporulation genes identified in B. subtilis are missing in C. acetobutylicum, which suggests major differences in the sporulation process. Thus, comparative analysis reveals both significant conservation of the genome organization and pronounced differences in many systems that reflect unique adaptive strategies of the two gram-positive bacteria.
The Clostridia are a diverse group of gram-positive, rod-shaped anaerobes that include several toxin-producing pathogens (notably Clostridium difficile, Clostridium botulinum, Clostridium tetani, and Clostridium perfringens) and a large number of terrestrial species that produce acetone, butanol, ethanol, isopropanol, and organic acids through fermentation of a variety of carbon sources (38, 72, 73, 86). Isolates of Clostridium acetobutylicum were first identified between 1912 and 1914, and these were used to develop an industrial starch-based acetone, butanol, and ethanol (ABE) fermentation process, to produce acetone for gunpowder production, by Chaim Weizmann during World War I (13, 34, 82, 87). During the 1920s and 1930s, increased demand for butanol led to the establishment of large fermentation factories and a more efficient molasses-based process (20, 34). However, the establishment of more cost-effective petrochemical processes during the 1950s led to the abandonment of the ABE process in all but a few countries. The rise in oil prices during the 1970s stimulated renewed interest in the ABE process and in the genetic manipulation of C. acetobutylicum and related species to improve the yield and purity of solvents from a broader range of fermentation substrates (52, 59, 87). This has developed into an active research area over the past two decades.
The type strain, Clostridium acetobutylicum ATCC 824, was isolated in 1924 from garden soil in Connecticut (83) and is one of the best-studied solventogenic clostridia. Strain relationships among solventogenic clostridia have been analyzed (11, 32, 33), and the ATCC 824 strain was shown to be closely related to the historical Weizmann strain. The ATCC 824 strain has been characterized from a physiological point of view and used in a variety of molecular biology and metabolic engineering studies in the United States and in Europe (3, 14, 22–24, 47, 56, 57, 79). This strain is known to utilize a broad range of monosaccharides, disaccharides, starches, and other substrates, such as inulin, pectin, whey, and xylan, but not crystalline cellulose (5, 6, 42, 52, 53). Physical mapping of the genome demonstrated that this strain has a 4-Mb chromosome with 11 ribosomal operons (9) and harbors a large plasmid, about 200 kb in size, which carries the genes involved in solvent formation, hence the name pSOL1 (10). Much work has been done to elucidate the metabolic pathways by which solvents are produced and to isolate solvent-tolerant or solvent-overproducing strains (8, 21, 35, 62, 69, 71, 80). Genetic systems have been developed that allow genes to be manipulated in C. acetobutylicum ATCC824 and related organisms (25, 48–52, 84), and these have been used to develop modified strains with altered solventogenic properties (25, 28, 54, 60).
Knowledge of the complete genome sequence of C. acetobutylicum ATCC 824 is expected to facilitate the further design and optimization of genetic engineering tools and the subsequent development of novel, industrially useful organisms. The sequence also offers the opportunity to compare two moderately related, gram-positive bacterial genomes (C. acetobutylicum and Bacillus subtilis) and to examine the gene repertoire of a mesophile anaerobe with metabolic capacities that were not previously represented in the collection of complete genomes.
MATERIALS AND METHODS
Sequencing.
The genome of C. acetobutylicum ATCC 824 was sequenced by the whole genome shotgun approach (18), using a combination of fluorescence-based and multiplex sequencing approaches (70). The finishing phase involved exhaustive gap closure and quality enhancement work using a variety of biochemical methods and computational tools. Clones from a plasmid library made with randomly sheared 2.0- to 2.5-kb inserts were sequenced from both ends. The sequences were preprocessed and base called with Phred (15), and low-quality reads were removed (multiplex or short-run dye terminator reads with fewer than 100 Phred Q-30 bases [error rate of 10−3], and long-run dye terminator reads with fewer than 175 Q-30 bases). This resulted in 4.9 Mb of multiplex reads and 21.3 Mb of ABI dye-terminator reads (8.3-fold sequence coverage; 51,624 reads in all). The data were assembled using Phrap (University of Washington; http: //bozeman.mbt.washington.edu/phrap.docs/phrap.html), which produced 551 contigs spanning a total of 4.03 Mb. A total of 0.76-fold coverage in paired reads from lambda clones was generated from two genomic lambda libraries (one provided by G. Bennett and one constructed at GTC). These data, together with data from primer-directed sequence walks across all captured gaps (sequence gaps with a bridging clone insert), and second-attempt sequences corresponding to missing mates at the ends of the contigs were reassembled with the original shotgun data to produce a final Phrap assembly. This assembly contained 108 contigs and 88 supercontigs. Further primer-directed sequencing efforts, using plasmid and PCR-generated templates, resulted in the eventual closure of the remaining captured gaps.
Gap closure.
Uncaptured gaps were closed using one of the following methods. The lambda libraries were screened with PCR products designed from the ends of contigs and labeled during the amplification process with digoxigenin. Positive clones from the chemiluminescence screening (Boehringer Mannheim kit) were sequenced from both ends and used as templates for additional primer walks. This resulted in 28 contig joins. Direct genomic sequencing was used to walk into gaps wherever unique primers could be specified near the end of a contig. Primers were identified using GTC's PrimerPicker software and were matched back to the genomic assembly using cross_match (University of Washington; http://bozeman.mbt.washington.edu/phrap.docs/phrap.html). Unique primers were added to 2X Big Dye reactions with 2.5 μg of total genomic DNA; these reactions yielded an 85% success rate with average Q-30 scores of 190. This procedure resulted in 13 contig joins, and multiple walks were performed in many cases. Combinatorial PCR was also used. Initially, a matrix of PCR primers representing all possible combinations in pools of 10 was used to reduce the number of PCRs that had to be performed. This was followed with a “2 × 2” approach using all possible combinations of pairs from the ends of the remaining contigs. Wherever products were observed, the primers contained in the original reaction would be used in combinations of 2 to determine which contig ends belonged together. These primers were then used to amplify the genomic DNA bridging the gap, and the products were used for primer walks. This procedure proved successful in bridging and closing the remaining gaps. The contigs that constituted pSOL1 were identified and linked at an early stage in the project; further work allowed us to produce a finished sequence for the plasmid of 192,000 bp.
Final assembly.
Sequence reads from the above efforts were incorporated into the contigs by means of the custom GTC incremental assembly tools: Inc_Asm, Contig_Merge, CM_calc, CM_auto, and Update_Overlaps. Misassemblies were identified through aberrant coverage or clone tiling and by inappropriate juxtaposition of restriction sites compared to the physical map (9). Each case was successfully resolved using PCR and sequence confirmation. The genome contained 11 rDNA operons, 6 of which occurred in two triplets, approximately 18 kb in length. Each ribosomal operon was independently amplified by PCR utilizing the flanking unique sequences, and the resulting products were sequenced. The operons were then incorporated into the genome assembly at the correct positions. Assembly of the final 13 contigs had to be done manually because of the repetitive elements and the limitations of Phrap and Contig_Merge.
Sequence quality.
The genome sequence was screened for regions of low sequence quality, and 2,883 ‘quality gaps’ were identified. Of these, 2,769 were improved by resequencing of the plasmid template with an alternate chemistry (e.g., energy-transfer dye primer; AP Biotech, Piscataway, N.J.). The remaining quality gaps were improved by means of primer walks. Based on the consensus quality scores generated by Phrap and Contig_Merge, and on the results of systematic quality checks on the lower-quality regions in the final contigs (when it was no longer possible to use the assembly tools because of repeats), we estimate the overall error rate to be substantially less than 1 error in 10,000 bases.
Sequence analysis and annotation.
The genome was analyzed and annotated in context with a large number of finished bacterial and archaeal genomes. Custom Perl scripts were used to automate the execution of similarity search algorithms, and additional scripts were used to filter the results and to create tab-delimited tables and Web pages to summarize the most biologically and functionally relevant information. The program uniorf (a wrapper around ExractOrfs5; GTC) was used to identify open reading frames (ORFs). The coding ORFs were identified using one or more of the three criteria: significant BLASTP2 hit, C. acetobutylicum-specific dicodon usage, or a length of ≥400 residues. Start codons were predicted by their proximity to ribosome binding sequences (67) and by compatibility with BLAST alignment data that minimized or eliminated overlaps. The predicted protein sequences were individually analyzed using sensitive profile-based methods for database searching, including PSI-BLAST (1, 2), IMPALA (64), and SMART (65, 66). All potential frameshifts identified during the analysis phase were investigated in the final sequence assembly. Corrections were made in every case where a probable sequence error could account for the apparent frameshift. In a few cases, genomic PCR amplification and product sequencing was undertaken to evaluate the potential frameshifts. The program tRNAscan was used to identify tRNA genes.
Comparative analysis.
Paralogous families of proteins were identified by comparing the complete set of predicted C. acetobutylicum proteins to itself (after filtering for low-complexity regions with the SEG program (88) using the PSI-BLAST program, which was run for three iterations, and clustering proteins by single-linkage (clustering threshold e value, 0.001) using the GROUPER program (81). Assignment of predicted proteins to clusters of orthologous groups (COGs) (78) was based on the results of the COGNITOR program (78), with manual verification. The functional assignments embedded in the COG database were also used for reconstruction of metabolic pathways and other functional systems in C. acetobutylicum in conjunction with the KEGG (37) and WIT (55) databases. Analysis of the phyletic distribution of the database hits reported by the BLASTP program was performed using the TAX_COLLECTOR program of the SEALS package (81). This was followed by phylogenetic tree construction for selected individual cases. Multiple alignments for phylogenetic reconstruction were generated using the ClustalW program (29) and, when necessary, further adjusted on the basis of the PSI-BLAST search outputs. Phylogenetic trees were constructed using the neighbor-joining method with 1,000 bootstrap replications as implemented in the NEIGHBOR program of the PHYLIP package (16). Evolutionary distance matrices for neighbor-joining tree construction were generated using the PROTDIST program of the PHYLIP package, with Kimura's correction for multiple substitutions.
The PerlTK program Genome_map (70) was used to generate circular genome maps (Fig. 1).
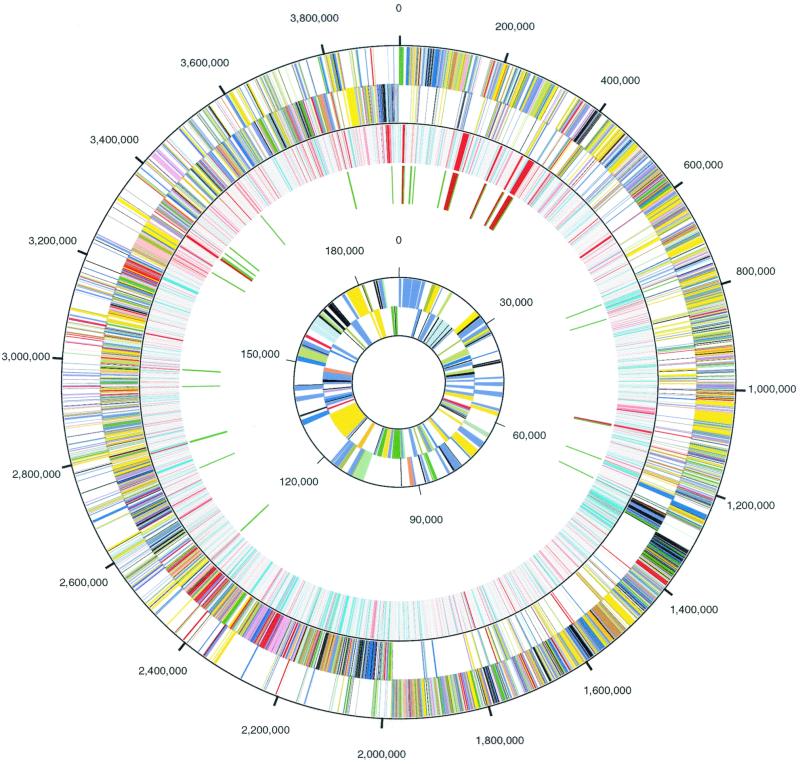
FIG. 1.
Circular representation of the C. acetobutylicum genome and megaplasmid. The outer two rings indicate the positions of genes on the forward and reverse strands of the genome, respectively, color-coded by function. Moving inward, the third ring indicates the G+C content of each putative gene: turquoise (≤27%), gray (27 to 35%), pink-red (>35%); the fourth ring indicates the positions of tRNA (green) and rRNA genes (dark red). The inner rings show the positions of genes on the forward and reverse strands of pSOL1, respectively, color-coded by function (the distance scale for the inner rings differs from the scale of the outer rings, as indicated). The functional color-coding is as follows: energy production and conversion, dark olive; cell division and chromosome partitioning, light blue; amino acid transport and metabolism, yellow; nucleic acid transport and metabolism, orange; carbohydrate transport and metabolism, gold; coenzyme metabolism, tan; lipid metabolism, salmon; translation, ribosome structure, and biogenesis, pink; transcription, olive drab; DNA replication, recombination, and repair, forest green; cell envelope biogenesis, outer membrane, red; cell motility and secretion, plum; posttranslational modification, protein turnover, and chaperones, purple; inorganic ion transport and metabolism, dark sea green; general function prediction only, dark blue; conserved protein, function unknown, medium blue; signal transduction mechanisms, light purple; predicted membrane protein, light green; hypothetical protein, black.
Nucleotide sequence accession numbers.
The sequence of the C. acetobutylicum strain ATCC 824 genome is available in GenBank under the accession number AE001437, and that of the megaplasmid pSOL1 is available under accession number AE001438. Graphical representations of the genome with detailed annotation are available at http://www.ncbi.nlm.nih.gov and http://www.genomecorp.com/programs/sequence_data_clost.shtml.
RESULTS AND DISCUSSION
Genome organization.
The C. acetobutylicum ATCC 824 genome consists of 3,940,880 bp. Genes are distributed fairly evenly, with ∼51.5% being transcribed from the forward strand and ∼49.5% from the complementary strand. A total of 3,740 polypeptide-encoding ORFs and 107 stable RNA genes were identified, accounting for 88% of the genomic DNA, with intergenic regions averaging ∼121 bp. A putative replication origin (base 1) and terminus were identified by GC and AT skew analysis (45); the origin marks a strong inflection point in the coding strand and contains several DnaA boxes, as well as gyrA, gyrB, and dnaA genes that are adjacent to the replication origin in many other bacteria. Another strong inflection in the coding strand occurs at the diametrically opposed putative replication terminus (reminiscent of the Mycoplasma genitalium genome (19) (Fig. 1). The 11 ribosomal operons are clustered in general proximity to the origin of replication and are all oriented in the same direction as the leading replication fork. The megaplasmid, pSOL1, consists of 192,000 bp and appears to encode 178 polypeptides. The single obvious skew inflection was placed at the origin (base 1), although there is no other support for a replication origin at this position (a repA homolog resides ∼2.2 kb away). In contrast to the genome, there is no obvious coding strand bias in the plasmid.
There appear to be two unrelated cryptic prophages in the genome. The first of these spans approximately 90 kb and includes approximately 85 genes (CAC1113 to CAC1197), with 11 phage-related genes, 3 XerC and XerD recombinase-related genes, and a number of DNA processing enzymes. This region contains a strong coding-strand inflection point near its center and has lower-than-average GC content. The second apparent prophage appears to span approximately 60 kb and displays similar coding characteristics in approximately 79 genes (CAC1878 to CAC1957; slightly higher than average in GC content). Genes for three distinct insertion sequence-related proteins (CAC0248, CAC3531, and CAC0656-57) are present on the chromosome. Only one of these is intact; another is a fragment, and the third has a frameshift. Another frameshifted gene coding for a TnpA-related transposase resides on pSOL1 (CAP0095-96). Thus, it appears that there are no active insertion sequence elements in the C. acetobutylicum genome.
There are 73 tRNA genes. The isoleucine tRNA could not be identified using standard search methods; this correlates with the displacement of the typical bacterial form of isoleucyl-tRNA with the eukaryotic version, although for other similarly displaced aminoacyl-tRNA synthetases (see below), the cognate tRNAs were readily identified.
Comparative analysis.
The genome of C. acetobutylicum provides us with at least two unique opportunities: (i) compare, for the first time, two large and moderately related gram-positive bacterial genomes, those of C. acetobutylicum and B. subtilis (41); (ii) investigate the genes that underlie the diverse set of metabolic capabilities so far not represented in the collection of complete genomes.
The median level of sequence similarity (26) between probable orthologs in C. acetobutylicum and B. subtilis was greater than between C. acetobutylicum and any other bacterium, but only by a rather small margin, indicating significant divergence (Table 1). Compared to the other pairs of evolutionarily relatively close genomes, the Clostridium-Bacillus pair is more distant than the species within the gamma-proteobacterial lineage (Escherichia coli, Haemophilus influenzae, Vibrio cholerae, and Pseudomonas aeruginosa) or Helicobacter pylori and Campylobacter jejunii; in contrast, the level of divergence between C. acetobutylicum and B. subtilis is comparable to that between the two spirochetes, Treponema pallidum and Borrelia burgdorferi (Table 1). The comparative analysis of the spirochete genomes has proved to be highly informative for elucidating the functions of many of their genes and predicting previously undetected aspects of the physiology of these pathogens (76).
TABLE 1.
The median identity percentage (± the standard deviation) between orthologous proteins of C. acetobutylicum and those of other bacteria and archaeaa
| Organism | % Protein identity with:
|
||||||||
|---|---|---|---|---|---|---|---|---|---|
| B.s. | E.c. | H.i. | P.a. | V.c. | H.p. | C.j. | T.p. | B.b. | |
| C.a. | 40.3 ± 0.3 | 35.8 ± 0.3 | 35.7 ± 0.3 | 35.5 ± 0.3 | 35.3 ± 0.3 | 35.7 ± 0.3 | 35.8 ± 0.3 | 34.8 ± 0.3 | 35.3 ± 0.4 |
| B.s. | 37.1 ± 0.3 | 36.0 ± 0.3 | 37.3 ± 0.3 | 36.1 ± 0.3 | 36.0 ± 0.3 | 34.9 ± 0.3 | 35.5 ± 0.4 | 35.4 ± 0.3 | |
| E.c. | 49.3 ± 0.7 | 45.4 ± 0.4 | 50.2 ± 0.6 | 35.2 ± 0.3 | 35.6 ± 0.3 | 36.2 ± 0.3 | 34.1 ± 0.4 | ||
| H.i. | 38.1 ± 0.4 | 49.6 ± 0.9 | 37.6 ± 0.6 | 37.6 ± 0.5 | 37.7 ± 0.6 | 36.1 ± 0.4 | |||
| P.a. | 44.8 ± 0.4 | 35.1 ± 0.3 | 35.1 ± 0.3 | 36.1 ± 0.3 | 35.6 ± 0.4 | ||||
| V.c. | 37.3 ± 0.6 | 37.8 ± 0.4 | 35.1 ± 0.3 | 35.8 ± 0.4 | |||||
| H.p. | 46.3 ± 0.6 | 36.7 ± 0.5 | 36.3 ± 0.5 | ||||||
| C.j. | 36.7 ± 0.6 | 37.7 ± 0.5 | |||||||
| T.p. | 38.3 ± 0.6 | ||||||||
Abbreviations of bacterial species are as follows: E.c.; E. coli; B.S., Bacillus subtilis; H.i., Haemophilus influenzae; P.a., Pseudomonas aeruginosa; V.c., Vibrio cholerae; H.p., Helicobacter pylori; C.j., Campylobacter jejuni; B.b., Borrelia burdorferi; T.p., Treponema pallidum. Values in boldface refer to statements made in the text.
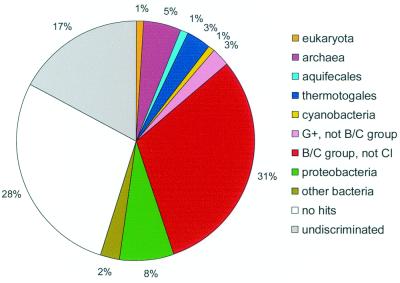
A taxonomic breakdown of the closest homologs for the C. acetobutylicum proteins immediately reveals the specific relationship with the low-GC gram-positive bacteria, with the reliable best hits for 31% of the C. acetobutylicum protein sequences being to this bacterial lineage (Fig. 2). However, nearly as many proteins produced clear best hits to homologs from other taxa (Fig. 2), which emphasizes the likely major role for lateral gene transfer, a hallmark of microbial evolution.
FIG. 2.
Taxonomic distribution of the closest homologs of C. acetobutylicum proteins. Undiscriminated, ORFs whose phylogenetic affinities remained unclear. Abbreviations: G+, gram positive; B/C, Bacillus/Clostridium; Cl, C. acetobutylicum.
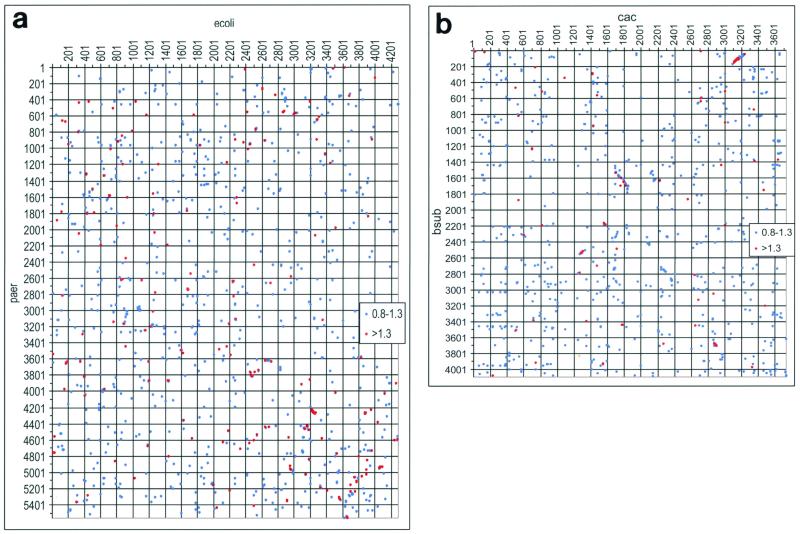
The same trends appear even more notable when the genome organizations of C. acetobutylicum and other bacteria are compared. Gene order is, in general, poorly conserved in the bacteria, with no extended synteny detected even among relatively close genomes, such as those of E. coli and P. aeruginosa or H. influenzae. In contrast, a genomic dot plot comparison of C. acetobutylicum with B. subtilis revealed several regions of colinearity (Fig. 3A and B). Thus, at least some bacterial genomes separated by a moderate evolutionary distance, as exemplified by C. acetobutylicum and B. subtilis, appear to retain the memory of parts of the ancestral gene order. A systematic mapping of conserved gene strings (many of which form known or predicted operons) on the C. acetobutylicum genome shows the clear preponderance of gene clusters shared with B. subtilis but also considerable complementary coverage by conserved operons from other bacterial and even archaeal genomes (Fig. 3C; see supplementary material at ftp://ncbi.nlm.nih.gov/pub /koonin/Clostridium). Altogether, 1,243 Clostridium genes (32% of the total predicted number of genes and 40% of the genes with detectable homologs) belong to conserved gene strings; 779 of these are in 271 predicted operons shared with B. subtilis (Fig. 3C; see supplementary material at ftp://ncbi.nlm.nih.gov/pub /koonin/Clostridium).
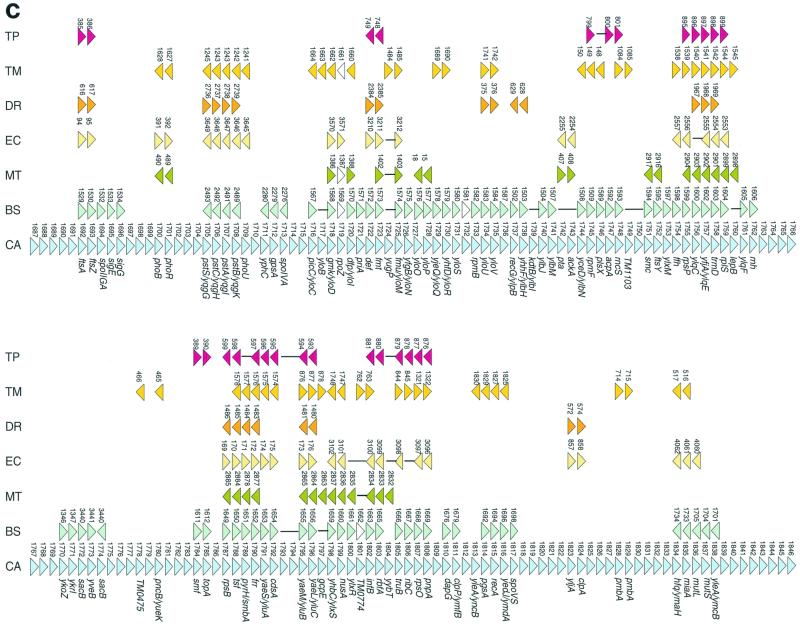
FIG. 3.
Conservation of gene order in C. acetobutylicum and other bacteria and archaea. (a) A genome dot plot comparison of E. coli (ecoli) and P. aeruginosa (paer). The numbers on the axes indicate the gene numbers in the corresponding genome. Each large unit corresponds to 200 genes, and each small unit corresponds to 100 genes. The red dots indicate protein alignments with a score density of >1.3 bit/position, and the blue dots indicate alignments with a score density of 0.8 to 1.3 bit/position. (b) A genome dot plot comparison of C. acetobutylicum (cac) and B. subtilis (bsub). (c) A comparison of genome organization in bacterial and archaeal genomes in the longest region of conserved gene order between C. acetobutylicum and B. subtilis. Abbreviations: TP, T. pallidum; TM, T. maritima; DR, D. radiodurans; EC, E. coli; MT, M. thermoautotrophicum; BS, B. subtilis; CA, C. acetobutylicum. The protein-coding genes in all genomes are denoted by numbers, starting from the first gene in the corresponding GenBank records. The white triangles show genes that are not homologous to the corresponding C. acetobutilicum genes. In gene strings that contain deletions compared to the C. acetobutylicum genome, the missing genes are replaced by lines joining the genes that are adjacent in the given genome. For each gene of C. acetobutylicum, the gene name of the ortholog in B. subtilis (or in another genome if a B. subtilis ortholog was not detectable) is indicated.
The genome region that shows the greatest level of gene order conservation between C. acetobutylicum and B. subtilis includes ∼200 genes and includes primarily (predicted) operons encoding central cellular functions, such as translation and transcription (Fig. 3C). The multiple genome alignment for this region clearly shows numerous rearrangements of gene clusters, with large-scale colinearity seen only between C. acetobutylicum and B. subtilis. The intermediate conservation of gene order seen between C. acetobutylicum and B. subtilis is likely to be particularly informative in terms of complementing functional predictions based on direct sequence conservation. For example, the predicted large “superoperon,” which contains genes for several components of the translation machinery (def, encoding N-formylmethionyl-tRNA deformylase; fmt, encoding methionyl-tRNA formyl transferase; and fmu, encoding a predicted rRNA methylase), transcription, and replication, additionally includes the genes yloO (CAC1727), yloP (CAC1728), and yloQ (CAC1729). These genes encode predicted protein phosphatase, serine-threonine protein kinase, and a GTPase, respectively. Based on the operon context, the readily testable predictions can be made that yloQ is a previously uncharacterized translation factor, whereas yloO and yloP are likely to play a role in the regulation of translation and/or transcription.
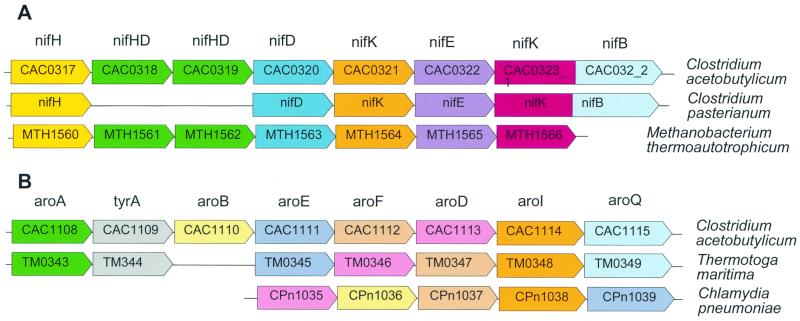
The mosaic picture of operon conservation can be explained by a combination of the processes of horizontal operon transfer, gene (operon) loss, and operon disruption (rearrangement). Distinguishing between these phenomena is, in many cases, difficult, but in certain extreme situations, one of the evolutionary routes is clearly preferable. A striking example is the conservation of the nitrogen fixation operon (six genes in a row) between C. acetobutylicum and another nitrogen fixator, the archaeon Methanobacterium thermoautotrophicum (Fig. 4A). This particular gene organization so far has not been seen in any other genome except for that of another clostridial species, C. pasteurianum, in which, interestingly, two genes of the operon are deleted (Fig. 4A). Similarly, the aromatic amino acid biosynthesis operon is conserved, albeit with local rearrangements, in C. acetobutylicum, Thermotoga maritima, and partially in Chlamydia (Fig. 4B). In these and similar cases, it is hard to imagine an evolutionary scenario that does not involve horizontal mobility of these operons, along with operon disruption in some of the bacterial and archaeal lineages.
FIG. 4.
Horizontally transferred operons in C. acetobutylicum. (a) Conservation of the nitrogenase operon in two species of Clostridium and M. thermoautotrophicum. (b) Conservation of the aromatic amino acid biosynthesis operon in C. acetobutylicum, T. maritima, and Chlamydia pneumoniae. Orthologs are shown by the same color.
In general, C. acetobutylicum carries the typical complement of genes that are conserved in most bacteria. The only gene that is present in all other bacteria (and, in fact, in all genomes sequenced to date) but is missing in C. acetobutylicum is that for thymidylate kinase.
A differential genome display analysis for C. acetobutylicum and B. subtilis, which was performed using the COG system (78), revealed 186 conserved protein families (COGs) that are represented in C. acetobutylicum but not in B. subtilis. Many of these proteins are involved in redox chains that are characteristic of the anaerobic metabolism of Clostridia as opposed to the aerobic metabolism of B. subtilis, as well as oxidation and reduction that are required for assimilation of nitrogen and hydrogen. Another group of enzymes belongs to biosynthetic pathways that are present in C. acetobutylicum but not in B. subtilis, primarily those for certain coenzymes, for example, cyancobalamin (see supplementary material at ftp://ncbi.nlm.nih.gov/pub/koonin /Clostridium). Conversely, 335 COGs were detected in which B. subtilis was represented, whereas C. acetobutylicum was not. An obvious part of this set consists of genes coding for components of aerobic redox chains, such as cytochromes and proteins involved in the assembly of cytochrome complexes. Also missing are a variety of membrane transporters, the glycine cleavage system that is present in the majority of bacteria. Several metabolic pathways are incomplete; for example, a considerable part of the tricarboxylic acid (TCA) cycle and molybdopterin biosynthesis is missing. The TCA cycle is incomplete in many prokaryotes, but in most of these cases, the chain of reactions producing three key precursors, 2-oxoglutarate, succinyl-CoA, and fumarate, can proceed in either the oxidative or the reductive direction (30). In C. acetobutylicum, citrate synthase, aconitase, and isocitrate dehydrogenase are missing. It appears, however, that what remains of the TCA cycle could function in the reductive (counterclockwise in Fig. 5) direction. The counterparts of enzymes involved in succinyl-CoA and 2-oxoglutarate formation in other organisms are missing in C. acetobutylicum. However, the genome encodes acetoacetyl:acyl CoA-transferase that catalyzes butyryl-CoA formation in solventogenesis (CAP0163-0164) and might also utilize succinate for the synthesis of succinyl-CoA and 2-oxoacid:ferredoxin oxidoreductase (CAC2458-2459) that could catalyze 2-oxoglutarate formation from succinyl-CoA (Fig. 5). Succinate dehydrogenase/fumarate reductase, the enzyme that normally catalyzes the reduction of fumarate to succinate, seems to be missing in C. acetobutylicum. However, this reaction is linked to the electron transfer chain and might be supported by another dehydrogenase whose identity could not be easily determined.
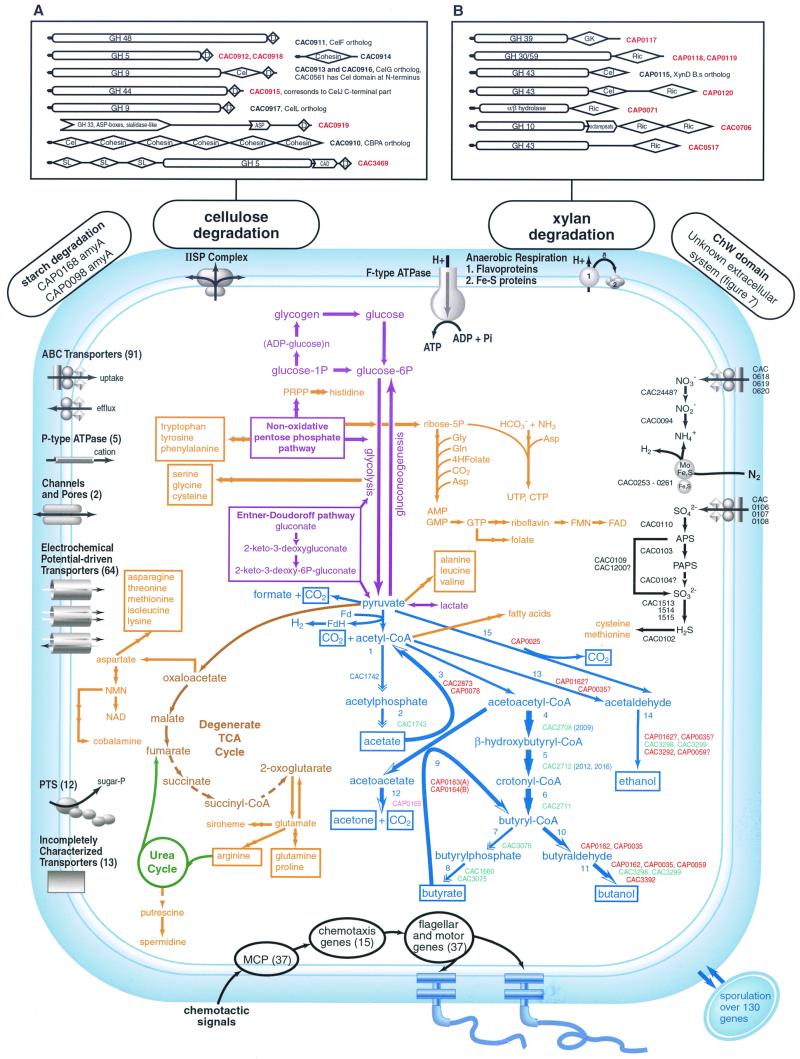
FIG. 5.
Overview of the basic metabolic pathways in C. acetobutylicum. The pathways are color coded as follows: catabolism of hydrocarbohydrates to pyruvate, purple; (incomplete) TCA cycle, brown; solventogenesis, blue; biosynthetic pathways, orange; urea cycle, forest green; nitrate and sulfate reduction and nitrogen fixation, black. Reactions for which no certain candidate enzyme was found are shown by dashed arrows. Phylogenetic affinities of genes of solventogenesis are shown by color: red for proteobacterial affinity; light green for Bacillus/Clostridium group; magenta for archaea. Genes with uncertain affinity are in blue. Different arrow shapes show that the respective genes are organized in operons. Numbers in the solventogenesis pathway correspond to the following enzymes: 1, phosphotransacetylase; 2, acetatekinase; 3, thiolase; 4, beta-hydroxybutyryl-CoA dehydrogenase; 5, crotonase; 6, butyryl-CoA dehydrogenase; 7, phosphotransbutyrylase; 8, butyrate kinase; 9, acetoacetyl-CoA:acyl-CoA transferase; 10, butyraldehyde dehydrogenase; 11, butanol dehydrogenase; 12, acetoacetate decarboxylase; 13, acetaldehyde dehydrogenase; 14, ethanol dehydrogenase; 15, pyruvate decarboxylase. Transporters are grouped by major categories, and the total number of transporters of each group is indicated in parentheses. The number of ABC transporters was estimated as the number of ABC-type ATPases. A more detailed breakdown of the transporters follows. ABC-type uptake transporters: nitrate, sulfate, phosphate, molybdate, ferrichrome, spermidine/putrescine, ribose, peptide, glycerol-3P (one of each); proline/glycine betaine, multidrug/protein/lipid (two paralogs of each); iron, cobalt (three paralogs); sugar, amino acid (five copies); oligopeptide (six copies). ABC-type efflux transporters: polysaccharide, Na+, (one of each), various specificities, homologous to eukaryotic P-glycoprotein (32 paralogs). P-type ATPases: K+, heavy metal (one of each), cation (three paralogs). Channels and pores: chloride, potassium (one of each). Electrochemical-driven transporters: formate/nitrite, ammonium, C4-dicarboxylate, proton/sodium-glutamate, transporter of cations and cationic drugs, 2-oxoglutarate/malate translocator (one of each); Na+/H+ antiporter, gluconate/proton symporter (two paralogs), Mn2+/H+ transporter, NRAMP family, Na:galactoside symporter family, Co/Zn/Cd symporter (four paralogs), amino acid transporters (12 paralogs), sugar-proton symporter (30 paralogs). PTS (phosphoenolpyruvate-dependant phosphotransferase system): mannitol, fructose, cellobiose, fructose (mannose), galactitol/fructose, lactose, N-acetylglucosamine, arbutin (one of each); glucose, beta-glucosides (two paralogs). Incompletely characterized transporters: xanthine, uracil, arsenite efflux pump (one of each); magnesium and cobalt transporter ferrous iron transport FeoA/FeoB (two paralogs), O-antigen transporter family (six paralogs). Abbreviations: IISP, type II general secretory pathway; PRPP, phosphoribosyl-pyrophosphate; 4Hfolate, tetrahydrofolate; APS, adenylylsulphate; PAPS, phosphoadenylylsulfate; MPS, methyl-accepting chemotaxis protein. Domain architectures of proteins involved in cellulose (A) and xylan degradation (B). Domain name abbreviations: D, dockerin; Ric, ricin; Cel, cellulose binding; SL, S layer; CAD, cell adhesion domain; GK, “Greek key” domain. Signal peptide is shown by an arrow. Gene identifiers of proteins with unique domain organizations are in red.
The repertoires of transcriptional regulators in B. subtilis (27) and C. acetobutylicum are very similar. In particular, of the 17 sigma factors predicted in C. acetobutylicum, 11 have readily detectable orthologs in B. subtilis. C. acetobutylicum also encodes numerous predicted specific transcriptional regulators, including 28 members of the AcrR/TetR family, 22 members of the MarR/EmrRs family, 14 members of the LysR family, 14 members of the Xre family, 9 members of the LacI family, and also several smaller sets of paralogous regulators. One-to-one orthologous relationships could be established only for a minority of these proteins (data not shown), and in some cases, such as, for example, that of the MarR/EmrRs family, part of the observed diversity seems to be due to independent family expansion.
The set of sporulation genes in C. acetobutylicum surprisingly differs from the set that has been well studied in B. subtilis (75). The number and diversity of detectable sporulation genes in Clostridium is much smaller. The most dramatic difference was observed among the SpoV genes. C. acetobutylicum does not have orthologs of the spoVF, spoVK, and spoVM genes, the disruption of which in B. subtilis leads to formation of immature spores that are sensitive to heat, organic solvents, and lysozyme (75). The phosphorelay system that functions in phase 0 of sporulation in B. subtilis (7, 31) appears to be missing in C. acetobutylicum, as indicated by the absence of an ortholog of SpoOB (phosphotransferase B) and SpoOF (a response regulator). In contrast, C. acetobutylicum encodes an apparent ortholog of the SpoOA (CAC2071) signaling protein that consists of a CheY domain and DNA-binding HTH domain and three proteins homologous to the ambiactive transcription repressors and activators AbrB and Abh (CAC1941, CAC0310, and CAC3647), also involved in phase 0 in B. subtilis. Interestingly, the SpoOA gene has been shown to control solventogenesis in solvent-forming Clostridia (60). In B. subtilis, sporulation is regulated by opposing activities of a distinct family of histidine kinases, KinA to KinE, and the Rap family phosphatases; orthologs of these genes were not detected in C. acetobutylicum.
B. subtilis has 22 cot genes that are responsible for coat biosynthesis; only 14 of these genes are conserved in C. acetobutylicum. Similarly, B. subtilis has 21 ger genes, 7 of which are represented by orthologs in Clostridium. Many of the missing GER genes encode various receptors of germination, which appear to be different in these bacteria. Furthermore, C. acetobutylicum does not have an ortholog of the cell-division-initiation gene divIC (75), which is essential in B. subtilis, suggesting differences in the mechanism of septum formation.
B. subtilis has a large set of competence genes which are involved in DNA uptake (12). The majority of these genes are represented by orthologs in C. acetobutylicum, but the proteins encoded by these genes in B. subtilis and C. acetobutylicum typically are not the most closely related members of the respective clusters of orthologs (data not shown). Operon disruption and rearrangements are also observed, suggesting a significant functional difference between the two gram-positive bacteria.
Many of the clostridial genes that are missing in B. subtilis seem to show distinct evolutionary affinities and probably have been acquired via horizontal transfer. In particular, a significant number of clostridial genes are conserved in all archaea whose genomes have been sequenced to date but are present in bacteria only sporadically (Table 2). Many of these genes encode various redox proteins, which reflects the similarity between the anaerobic redox chains in archaea and clostridia. For most of these “archaeal” genes found in bacteria, the probable evolutionary model is a single entry into the bacterial world by horizontal transfer from the Archaea, followed by dissemination among the Bacteria. In several cases, however, direct gene transfer from archaea into the clostridial genome seems likely; examples include the genes for a metal-dependent hydrolase of the metallo-beta-lactamase superfamily (CAC0535), a calcineurin-like phosphatase which has undergone duplication in C. acetobutylicum, probably subsequent to the acquisition of an archaeal gene (CAC1010 and CAC1078), and a predicted DNA-binding protein (CAC3166). Another group of clostridial genes includes probable eukaryotic acquisitions (Table 2). As with archaeal genes, the scenario of a single entry into the bacterial world followed by horizontal dissemination is likely for many of these genes, for example, that for the FHA domain discussed below. However, about 50 genes in C. acetobutylicum could have been directly hijacked from eukaryotes (Table 2). An interesting example is the nucleotide pyrophosphatase, which is encoded within one of the gene clusters including genes for FHA-containing proteins (Fig. 6) and therefore may be also implicated in signaling. As noticed previously, lateral acquisition of some of the aminoacyl-tRNA synthetases from eukaryotes, accompanied by displacement of the original copies, seems to have occurred repeatedly in bacterial evolution (85). C. acetobutylicum is no exception, with its arginyl-tRNA synthetase showing a clear eukaryotic affinity. In these cases, horizontal gene transfer from eukaryotes to specific bacterial lineages appears more likely than horizontal transfer in the opposite direction, bacteria to eukaryotes. The latter interpretation would require independent gene loss in multiple bacterial lineages accompanied by multiple instances of nonorthologous displacement.
TABLE 2.
C. acetobutylicum genes missing in B. subtilis and showing apparent evolutionary affinity to distant taxaa
| Type of affinity and C.a. gene ID | Description and comments for gene product |
|---|---|
| Eukaryotic affinity (49 proteins total) | |
| CAC0406 | Predicted membrane protein, containing FHA domain |
| CAC0529 | Acetylxylan esterase-related enzyme |
| CAC0537 | Acetylxylan esterase; acyl-CoA esterase or GDSL lipase family |
| CAC0920 | Protein related to MIFH/DOPD protein family, function in bacteria is unknown |
| CAC1075 | Beta-glucosidase family protein |
| CAC1344 | Sugar kinase, possible xylulose kinase |
| CAC2541 | Reductase/isomerase/elongation factor common domain |
| CAC3373 | Pectin methylesterase |
| CAC3411 | Homolog of plant auxin-responsive GH3-like protein |
| CAP0004 | Cysteine protease |
| CAP0129 | Glycogen-binding regulatory subunit of S/T protein phosphatase I |
| Archaeal affinity (195 proteins total) | |
| CAC0033 | ABC1 family protein kinase |
| CAC0069 | Predicted iron-binding protein, hemerythrin |
| CAC0214 + others | Endoglucanase, aminopeptidase M42 family |
| CAC0474 CAC0478 | ACT domain containing transcriptional regulator |
| CAC0650 | Adenylate cyclase, class 2 (thermophilic) |
| CAC2000 CAC2001 | Indolepyruvate ferredoxin oxidoreductase, subunit beta and alpha |
| CAC2409 | Transglutaminase-like enzyme, putative cysteine protease |
| CAC2520 + others | Multimeric flavodoxin (WrbA) domain-containing protein |
| CAC3110 | L14E ribosomal protein |
| CAC3597 CAC3598 | Rubrerythrin |
| CAC3555 + others | Nitroreductase family protein |
| Affinity to T. maritima (168 proteins total) | |
| CAC0672 | TGS and inactivated HXXXH domain of Thr-RSase fused to uridine kinase |
| CAC0749 + others | HD and HD-GYP hydrolase |
| CAC1319-CAC1326 | Gene cluster with unknown proteins and glycerol uptake facilitator protein |
| CAC2428 CAC2714 | Activator of 2-hydroxyglutaryl-CoA dehydratase |
| CAC0780 | Tyrosyl-tRNA synthetase |
C.a., C. acetobutylicum.
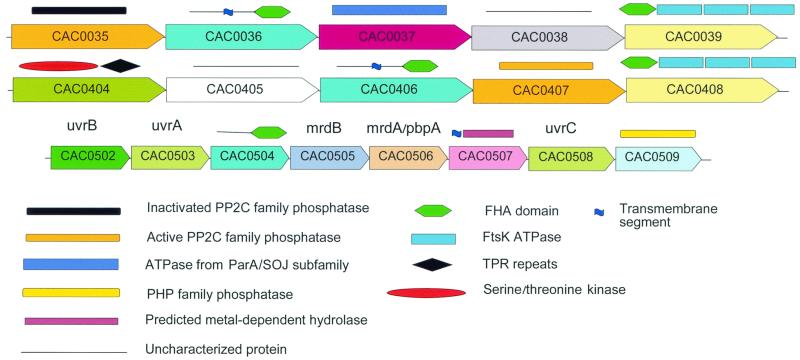
FIG. 6.
Novel signal transduction operons in C. acetobutylicum. Paralogs are shown by the same color (pattern). Domain organization is shown above the boxes with gene identifiers. See the key in the bottom of the figure and additional details in the text.
Most of the essential functions in C. acetobutylicum and B. subtilis are associated with readily detectable orthologs, but there are also notable cases of nonorthologous gene displacement (Table 3). Examples include glycyl-tRNA synthetase, which is represented by the typical bacterial, two-subunit form in B. subtilis and by the one-subunit archaeal-eukaryotic version in C. acetobutylicum, and uracil-DNA glycosylase, similarly represented by the classical bacterial enzyme (ortholog of E. coli Ung) and by the archaeal version in C. acetobutylicum (Table 3). In many cases, while an apparent orthologous relationship was detected between a clostridial protein and its counterpart from B. subtilis, there was nevertheless a clear difference in the domain architectures (Table 2). Notable examples of unusual domain organizations from C. acetobutylicum include the FtsK ATPase, which is fused to the FHA domain (see below), a Pkn2 family protein kinase fused to tetratricopeptide repeats (CAC0404), and another ATPase fused to a LexA-like DNA-binding domain (CAC1793). The evolution of another set of genes seems to have involved xenologous gene displacement whereby a gene in one of the compared genomes (C. acetobutylicum or B. subtilis) is displaced by the ortholog from a distant branch of the phylogenetic tree, e.g., eukaryotes (Table 3). Characteristically, this evolutionary pattern was detected for three aminoacyl-tRNA synthetases, those for isoleucine, arginine, and histidine; in each of these cases, C. acetobutylicum possesses the archaeal-eukaryotic version as opposed to the typical bacterial versions found in B. subtilis. Another interesting example of xenologous displacement involves the two forms of clostridial ribonucleotide reductase, neither of which groups with the counterparts from B. subtilis in phylogenetic trees. One of the ribonucleotide reductase genes in B. subtilis contains the single intein in that organism; C. acetobutylicum has no inteins, however. These observations show that there had been a significant horizontal exchange of genes between the Clostridium lineage and certain archaea and/or eucaryotes subsequent to its divergence from the Bacillus lineage.
TABLE 3.
Complex relationships between genes of C. acetobutylicum and B. subtilis—nonorthologous and xenologous gene displacement and differences in domain architecturesa
| C.a. gene ID | B.s. gene product | Commentb |
|---|---|---|
| CAC0039, CAC0408 | YukA | DNA segregation ATPase FtsK/SpoIIIE; both contain 3 ATPase domains; C.a. protein in addition contains C-terminal FHA domain |
| CAC0578 | YitJ | Methionine synthase I; same domain organization as in E.c., D.r., Syn., and M.t. but not B.s. |
| CAC0459, CAC3088 | RocR | Arginine degradation positive regulator; AAA-ATPase domain (NtrC family) fused to PAS domains; C.a. proteins contain an unknown domain at the N terminus similar to the N-terminal domain of PrpR E.c. |
| CAC0538-40, CAC2556 | YdhT | Beta-mannanase ManB; C.a. proteins have more complex domain organization |
| CAC0577 | YxiA | Endo-arabinase-related enzyme (family 43, glycosyl hydrolase domain); C.a. contains additional ricin B-like domain |
| CAP0120 | XynD | Xylanase; C.a. contains additional ricin B-like domain |
| CAC1655 | PurQ and PurL | Phosphoribosylformylglycinamidine synthase; C.a. has PurL/PurQ fusion; B.s.—separate proteins |
| CAC1847 | YqfP and YpfD | C.a. has fusion of LytB protein and S1 RNA-binding domain; B.s.—separate proteins |
| CAC1793 | LexA | C.a. has fusion of P-loop ATPase and LexA; B.s. has only a LexA protein and has no ortholog of the ATPase domain |
| CAC2927 | FolA and FolK | Folate biosynthesis enzymes; C.a. has fusion FolA and FolK; B.s.—separate proteins |
| CAC0404 | YbdM | C.a. has fusion Pkn2 protein kinase with TPR repeats; B.s. has only a Pkn2 ortholog without the TPR domain. |
| CAC3245 | 8 one-domain proteins | C.a. has fusion of two B. subtilis domains; B.s. has only separate proteins |
| CAC3195 | GlyQ, GlyS | Glycyl-tRNA synthetase; typical bacterial version (two subunits) in B.s.; eukaryotic version (one subunit) in C.a. |
| CAC0219, CAC2997 | Ung | Uracil-DNA glycosylase; typical bacterial version in B.s.; archaeal version in C.a. |
| CAC1256, CAC1909 | Rph | C.a. encodes RNase D and RNAse E/G, whereas B.s. encodes the unrelated RNAse PH; it can be predicted that these RNases complement each other's functions. |
| CAC0752 | YoqV, YoqU | ATP-dependent DNA ligases; B.s. encodes two copies of a typical ATP-dependent ligase strongly similar to archaeal homologs (one fused to a eukaryotic-type DNA primase); in contrast, C.a. has an extremely diverged version with significant similarity only to predicted ligases from D.r. |
| CAC1041 | ArgS | Arginyl-tRNA synthetase; eukaryotic version in C.a. |
| CAC0935, CAC2740 | HisS, HisZ | Histidyl-tRNA synthetase; B.s. encodes two recently diverged copies of the typical bacterial enzyme; C.a. possesses an archaeal-eukaryotic version and a distinct bacterial variant shared with D.r. and Syn |
| CAC3038 | IleS | Isoleucyl-tRNA synthetase; archaeal-eukaryotic version in C.a. |
| CAC1047 | YosO | Ribonucleotide reductase in B.s. contains an intein; there is no ortholog of this in C.a.; instead, C.a. contains a distinct ribonucleotide reductase with an archaeal evolutionary affinity |
See the Table 1 a footnote for abbreviations. Additional abbreviations of bacterial species are as follows: D.r., Deinococcus radiodurans; Syn, Synechocystis sp.
The first 12 comparisons relate to different domain organization; the last 8 relate to nonorthologous and xenologous gene displacement.
The results of systematic analysis of protein families that are specifically expanded with C. acetobutylicum are largely compatible with the current knowledge of the physiology of the bacterium (Table 4). For example, distinct families of proteins involved in sporulation, anaerobic energy conversion, and carbohydrate degradation were identified (Table 4). A so far unique feature is the presence of four diverged copies of the single-stranded DNA-binding proteins, an essential component of the replication machinery that is present in one or two copies in all other sequenced bacterial genomes. In addition, this analysis revealed remarkable aspects of the signal transduction system in this bacterium. Of particular interest is the proliferation of the phosphopeptide-specific, protein-protein interaction module, the FHA domain, which is generally rare in the Bacteria (44). C. acetobutylicum encodes five FHA-domain-containing proteins, which is comparable to the number of these domains in other bacteria with versatile Ser/Thr-phosphorylation-based signaling, namely Mycobacterium tuberculosis (10) and Synechocystis sp. (7); most of the other bacteria do not encode FHA domains or possess just one copy (58). Four of the genes coding for FHA-domain-containing proteins in C. acetobutylicum belong to two partially similar gene clusters that are unique for C. acetobutylicum and additionally include genes for other phosphorylation-dependent signaling proteins, namely predicted protein kinases and phosphatases (Fig. 6). The fusion of the FHA domain with the FtsK ATPases, which is involved in chromosome segregation, and the presence, in one of the clusters, of an ATPase of the MinD family, also involved in chromosome partitioning, suggest previously unsuspected regulation of cell division in C. acetobutylicum via reversible protein phosphorylation. The fifth FHA-domain-containing protein seems to belong to yet another predicted operon that is potentially involved in cell division as indicated by the presence of genes for a penicillin-binding protein and another membrane protein implicated in cell division in other bacteria (Fig. 6). These observations are compatible with the hypothesis on the role of phosphorylation in the regulation of this process in C. acetobutylicum. Another signaling system that is predicted to play a prominent role in C. acetobutylicum on the basis of protein family expansion analysis includes the so-called HD-GYP domains (name based on the one-letter code for characteristic amino acids) that are suspected to possess cyclic diguanylate phosphoesterase activity (Table 4); the only comparable expansion of the HD-GYP domain is seen in T. maritima. The HD-GYP proteins could play a major role in sensing the redox state of the environment in C. acetobutylicum (M. Y. Galperin, D. A. Natale, L. Aravind, and E. V. Koonin, Letter, J. Mol. Microbiol. Biotechnol. 1:303–305, 1999).
TABLE 4.
Specific expansion of protein families in C. acetobutylicum
| Function and Protein family definition | No. of proteins found in:
|
||||
|---|---|---|---|---|---|
| C. acetobutylicum | B. subtilis | E. coli | M. tuberculosis | T. maritima | |
| Implicated in spore formation | |||||
| CotS | 6 | 3 | 0 | 0 | 0 |
| GerKC | 4 | 4 | 0 | 0 | 0 |
| GerKB | 5 | 5 | 0 | 0 | 0 |
| GerKA | 4 | 6 | 0 | 0 | 0 |
| SpoVT/AbrB | 5 | 3 | 0 | 2 | 0 |
| Spore-cortex-lytic enzyme, CAC3602 | 3 | 3 | 0 | 0 | 0 |
| Signal transduction/regulation | |||||
| BglG family (sugar-dependent transcription antiterminator) | 6 | 7 | 1 | 0 | 0 |
| Xre-like regulator | 24 | 14 | 5 | 4 | 2 |
| HD-GYP | 9 | 0 | 0 | 0 | 10 |
| FHA domain | 5 | 0 | 0 | 10 | 0 |
| Regulators related to fur | 5 | 1 | 1 | 3 | 2 |
| LRPR/YAEG family | 4 | 1 | 0 | 1 | 0 |
| TPR-repeat | 32 | 17 | 4 | 0 | 5 |
| Activator of 2-hydroxyglutaryl-CoA dehydratase | 4 | 0 | 1 | 0 | 2 |
| D-alanine carboxypeptidase | 17 | 15 | 10 | 8 | 3 |
| Energy metabolism | |||||
| NifD, Nitrogenase iron-molybdenum protein | 4 | 0 | 0 | 0 | 0 |
| 6Fe-6S prismane | 4 | 0 | 1 | 0 | 1 |
| Flavodoxin WrbA family | 11 | 1 | 1 | 1 | 0 |
| Nitroreductase family | 12 | 4 | 4 | 2 | 4 |
| Aldehyde-alcohol dehydrogenase | 8 | 3 | 6 | 0 | 4 |
| MoaA/NirJ Fe-S | 7 | 4 | 1 | 4 | 4 |
| NADH/flavin oxidoreductase, COG1902 | 6 | 2 | 2 | 2 | 1 |
| Sugar metabolism | |||||
| XylR/fructokinase | 5 | 3 | 7 | 3 | 8 |
| Mutarotase/aldose | 3 | 1 | 3 | 0 | 1 |
| Pectate lyase | 7 | 0 | 0 | 0 | 0 |
| Levanase/levansucrase | 5 | 4 | 0 | 0 | 1 |
| Ribose 5-phosphate isomerase RpiB | 5 | 1 | 1 | 1 | 1 |
| NodB family | 13 | 6 | 2 | 1 | 1 |
| Endoglucanase (family 5) | 6 | 1 | 0 | 0 | 2 |
| Beta-mannanase (family 26) | 5 | 1 | 0 | 0 | 0 |
| Pectate lyase | 5 | 0 | 0 | 0 | 0 |
| Extracellular functions | |||||
| Dockerin | 10 | 0 | 0 | 0 | 0 |
| Cell-adhesion domain | 19 | 0 | 0 | 0 | 0 |
| Glycosyltransferase | 54 | 26 | 17 | 17 | 17 |
| CAAX-like peptidase | 6 | 2 | 0 | 4 | 1 |
| Peptodoglycan-binding domain | 13 | 7 | 0 | 1 | 0 |
| MSPA | 42 | 10 | 6 | 0 | 7 |
| ricin-like | 7 | 0 | 0 | 0 | 0 |
| CHW repeats | 20 | 0 | 0 | 0 | 0 |
| Other | |||||
| Ribosomal protein S4 | 3 | 1 | 1 | 1 | 1 |
| 6-pyruvoyl-tetrahydropterin synthase | 6 | 1 | 1 | 0 | 1 |
| DNA invertase Pin | 5 | 2 | 2 | 1 | 0 |
| SSB-like protein | 4 | 2 | 1 | 2 | 1 |
| Unknown | |||||
| YitT family | 4 | 7 | 0 | 0 | 1 |
| YUKE/YFJA family | 9 | 2 | 0 | Many | 0 |
| YcaP family | 5 | 5 | 1 | 0 | 0 |
The solventogenesis pathways of C. acetobutylicum involve the formation of acetone, acetate, butanol, butyrate, and ethanol from acetyl-CoA (52). Two mechanisms of butanol formation have been identified in C. acetobutylicum, one of which is associated with solventogenesis (production of butanol, ethanol, and acetone) and the other with alcohologenesis (production of butanol and ethanol only). The genes involved in solventogenesis have been previously identified on the megaplasmid and sequenced (Galperin et al, letter), but the genes responsible for alcohologenesis were unknown. The genome sequencing allows the identification of a second alcohol-aldehyde dehydrogenase (CAP0035), a pyruvate decarboxylase (CAP0025), and an ethanol dehydrogenase (CAP0059) that are probably involved in this alcohologenic metabolism (Fig. 5) and interestingly are also carried by the megaplasmid. The enzymes involved in the final steps of solvent formation show variable phylogenetic profiles, and in particular, several of them appear to be specifically related to the homologs from the archaeon Archaeoglobus fulgidus (Fig. 5). In contrast, the genes for the two subunits of another key enzyme of the acetone pathway, acetoacetyl-CoA:acyl-CoA transferase, show a clear proteobacterial affinity. Together with the fact that a significant subset of the solventogenesis enzymes is encoded on the clostridial megaplasmid, these observations suggest that these pathways could have evolved via a complex sequence of gene/operon acquisition events. The megaplasmid also carries second copies of genes involved in PTS-type sugar transport (CAP0066-68), glycolysis (aldolase, CAP0064) and central metabolism (thiolase, CAP0078). It would be interesting to determine the expression profiles of the plasmid-encoded and chromosomal copies of these genes to investigate (i) whether these genes and the solventogenic genes are regulated or coregulated and (ii) whether metabolic complementarily exists between the chromosome and the plasmid in C. acetobutylicum.
The cellulosome, the macromolecular complex for cellulose degradation, has been genetically and biochemically characterized in four Clostridium species (C. thermocellum, C. cellulovorans, C. cellulolyticum, and C. josui) but not in C. acetobutylicum (which is able to hydrolyze carboxy-methyl cellulose but not amorphous or crystalline cellulose (68). The proteins of the cellulosome contain a C-terminal Ca2+-binding dockerin domain, which is required for the binding to the cohesin domains of a scaffolding protein (36, 40). Genome sequence analysis revealed at least 11 proteins that are confidently identified as cellulosome components (Fig. 5A). Most of these genes are organized in an operon-like cluster (CAC910 to CAC919) with a gene order similar to that of those in mesophilic C. cellulolyticum and C. cellulovorans, as distinct from the more dispersed organization in the thermophile C. thermocellum (4, 77). The large glycohydrolase CAC3469 is the homolog of EngE of C. cellulovorans, which is also encoded away from the main cellulosome gene cluster. Unlike EngE, CAC3469 possesses an additional cell adhesion domain (Fig. 5A). This protein contains S-layer homology domains and cell adhesion domains similar to those of SlpA, one of the anchoring proteins of C. thermocellum. The presence of the short cohesion domain protein CAC914 suggests a role in cellulosome function related to that of the HbpA protein of C. cellulovorans (77). The other dockerin-domain containing proteins, those of the GH48, GH5, and GH9 families, might interact with CAC910, the ortholog of the scaffolding protein CbpA. Generally, although the cellulosome has not been detected in C. acetobutylicum, the number of relevant proteins and domains would seem sufficient to encode the various combinations of cellulose-binding and hydrolytic proteins found in this complex. An interaction between CAC3469 and CAC910 could be speculatively proposed as a means of anchoring a potential cellulosome-like structure to the peptidoglycan.
In work analyzing the cellulolytic activities of C. acetobutylicum strains, it was found that NRRL B 527 could hydrolyze Avicel and acid-swollen cellulose but C. acetobutylicum ATCC 824 could not (42). The subsequent taxonomic and historical analyses of these strains (32, 33) indicate a close relationship and suggest that further investigation of the cluster from strain B 527 would be informative in elucidating the reason for the different cellulolytic activities of the two strains. Further work is required to resolve these issues and to determine the exact functions of the cellulosome subunits in C. acetobutylicum.
In addition to the known cellulosome components, C. acetobutylicum encodes numerous other proteins that are predicted to be involved in the degradation of xylan, levan, pectin, starch, and other polysaccharides. Altogether, there seem to be over 90 genes encoding proteins implicated in these processes, including representatives of at least 14 distinct families of glycosyl hydrolases. In particular, a predicted operon located on the C. acetobutylicum megaplasmid (CAP0114 to CAP0120) consists mostly of genes encoding xylan degradation enzymes. Similarly to the cellulosome components, these enzymes possess complex domain architectures, with the oligosaccharide-binding ricin domain (74) typically present at the C terminus; the addition of ricin domain is (so far) a unique feature of this postulated novel system for xylan degradation in Clostridium (Fig. 5B). Two of the putative xylanases presumably correspond to previously reported enzymes of xylan degradation isolated from C. acetobutylicum ATCC 824 (43).
A number of sugar PTS transport system genes,as well as the corresponding regulatory system analogs (e.g., Hpr, ptsK, and CcpA), have been found which couple transport signals to genetic regulation of degradative operons (61, 63). Non-PTS-mediated uptake of certain sugars, especially pentoses, has been found in several clostridial species (52). Many primary active transporters, including ABC-type transporters and P-type ATPases, electrochemical potential-driven transporters, channels and pores, and uncharacterized transporters were detected among the gene products of C. acetobutylicum (Fig. 5; see details in the figure legend). There is, however, no ortholog of the glucose facilitator of B. subtilis (17).
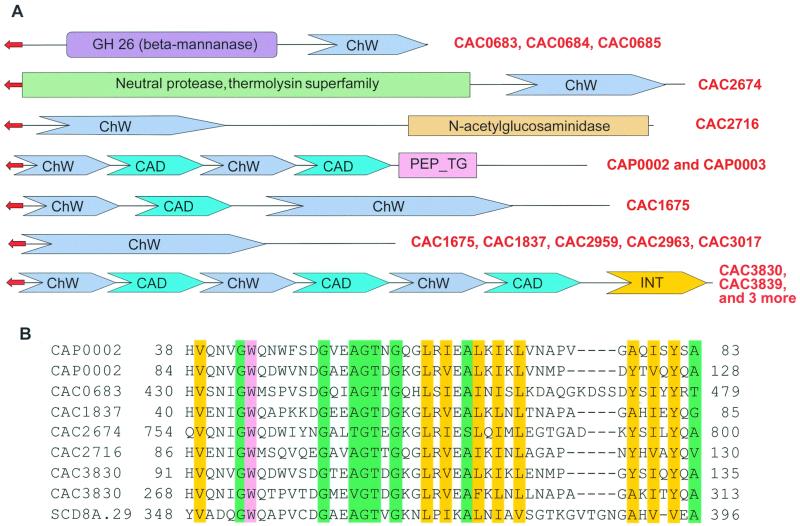
Along with previously characterized molecular complexes involved in extracellular hydrolysis of organic polymers, a novel system possibly related to these processes was detected. The signature of this system is a previously undetected domain with a distinct repetitive structure, which we designated as “ChW repeats” (clostridial hydrophobic, with a conserved W, tryptophan) (Fig. 7B). So far, the only nonclostridial protein containing similar repeats was detected in Streptomyces coelicolor (Fig. 7B). All proteins containing ChW repeats contain confidently predicted signal peptides at their N termini and do not contain predicted transmembrane helices, which suggests that all of them are secreted (Fig. 7A). Some of the ChW-repeat proteins contain additional enzymatic domains, such as glycosyl hydrolases or proteases, which implicates them in the degradation of polysaccharides and proteins. Several ChW-repeat proteins also contain domains that are involved in cell interactions, such as the cell adhesion domain (39) and the leucine-rich repeat (internalin) domain (46) (Fig. 7A). The internalin domain has been shown to play a critical role in the host cell invasion by the bacterial pathogen Listeria monocytogenes (46). In C. acetobutylicum, these domains might be responsible for interactions with plant cells. ChW repeats also could function in either substrate-binding or protein-protein interactions. The specific expansion of this domain in C. acetobutylicum suggests the existence of a novel molecular system, which partially resembles the cellulosome and could also form structurally distinct multisubunit complexes involved in polymer degradation and interaction with the environment. Elucidation of the function of this system is expected to shed light on the unique physiology of C. acetobutylicum.
FIG. 7.
A predicted novel extracellular macromolecular system based on proteins containing the previously uncharacterized ChW repeats. Domain name abbreviations: CAD, cell adhesion domain; INT, intrenalin-related domain; PEP_TG, predicted peptidase of transglutaminase family. Signal peptide is shown by a red arrow. Gene identifiers of proteins with unique domain organizations are in red. (A) The domain architectures of the proteins with ChW repeats. (B) Multiple alignment of ChW repeats in selected proteins from C. acetobutylicum; SCD8A0.29 is an S. coelicolor protein. The highlighting shows conserved amino acid residues. A yellow background indicates hydrophobic residues (A, C, F, I, L, M, V, W, Y, G), a green background indicates small residues (A, C, S, T, D, N, V, G, P), and magenta color indicates aromatic residues (W, Y, F). The numbers indicate the positions of the first and last residues of the aligned region in each protein sequence.
The extreme diversity of the domain architectures of the proteins that comprise the cellulosome and other predicted polymer degradation systems suggests that such complexes are highly dynamic not only in terms of the subunit stoichiometry (68) but also with respect to the genetic organization, with horizontal gene transfer, domain shuffling, and nonorthologous gene displacement playing pivotal roles in their evolution. C. acetobutylicum is the first sequenced bacterial genome with such a remarkable abundance of polymer degradation systems, which makes it a model for future studies on other bacteria with similar lifestyles. In addition, the sequencing of the C. acetobutylicum genome will offer perspectives in future comparative genomic studies concerning pathogenic bacteria, e.g., C. difficile, C. tetani, and C. perfringens, which are currently being sequenced by other groups.
ACKNOWLEDGMENTS
This work was supported by research grants DE-FG02-95ER-61967 (D.R.S.), DE-FG02-00ER629 (M.J.D.), NSF BES0001288 (G.N.B.), and USDA 00-35504-9269 (G.N.B.).
We are grateful to Guy Plunkett (University of Wisconsin) for performing the skew analyses and to John Reeve for helpful discussions and for suggesting C. acetobutylicum as a target for genomic sequencing.
REFERENCES
- 1.Altschul S F, Koonin E V. Iterated profile searches with PSI-BLAST—a tool for discovery in protein databases. Trends Biochem Sci. 1998;23:444–447. doi: 10.1016/s0968-0004(98)01298-5. [DOI] [PubMed] [Google Scholar]
- 2.Altschul S F, Madden T L, Schaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bahl H, Mueller H, Behrens S, Joseph H, Narberhaus F. Expression of heat shock genes in Clostridium acetobutylicum. FEMS Microbiol Rev. 1995;17:341–348. doi: 10.1111/j.1574-6976.1995.tb00217.x. [DOI] [PubMed] [Google Scholar]
- 4.Bayer E A, Shimon L J, Shoham Y, Lamed R. Cellulosomes—structure and ultrastructure. J Struct Biol. 1998;124:221–234. doi: 10.1006/jsbi.1998.4065. [DOI] [PubMed] [Google Scholar]
- 5.Blanchet, D., R. Marchal, and J. P. Vandecasteele. 1985. Acetone and butanol by fermentation of inulin. French patent 2559160.
- 6.Bronnenmeier K, Staudenbauer W L. Molecular biology and genetics of substrate utilization in Clostridia. In: Woods D, editor. The Clostridia and bio/technology. Reading, Mass: Butterworth-Heinemann; 1993. p. 443. [PubMed] [Google Scholar]
- 7.Burbulys D, Trach K A, Hoch J A. Initiation of sporulation in B. subtilis is controlled by a multicomponent phosphorelay. Cell. 1991;64:545–552. doi: 10.1016/0092-8674(91)90238-t. [DOI] [PubMed] [Google Scholar]
- 8.Clark S W, Bennett G N, Rudolph F B. Isolation and characterization of mutants of Clostridium acetobutylicum ATCC 824 deficient in acetoacetyl-coenzyme A:acetate/butyrate:coenzyme A-transferase (EC 2.8.3.9) and other solvent pathway enzymes. Appl Environ Microbiol. 1989;55:970–976. doi: 10.1128/aem.55.4.970-976.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Cornillot E, Croux C, Soucaille P. Physical and genetic map of the Clostridium acetobutylicum ATCC 824 chromosome. J Bacteriol. 1997;179:7426–7434. doi: 10.1128/jb.179.23.7426-7434.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cornillot E, Nair R V, Papoutsakis E T, Soucaille P. The genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 reside on a large plasmid whose loss leads to degeneration of the strain. J Bacteriol. 1997;179:5442–5447. doi: 10.1128/jb.179.17.5442-5447.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cornillot E, Soucaille P. Solvent forming genes in Clostridia. Nature. 1996;380:489. [Google Scholar]
- 12.Dubnau D. DNA uptake in bacteria. Annu Rev Microbiol. 1999;53:217–244. doi: 10.1146/annurev.micro.53.1.217. [DOI] [PubMed] [Google Scholar]
- 13.Durre P. New insights and novel developments in Clostridial acetone/butanol/isopropanol fermentation. Appl Microbiol Biotechnol. 1998;49:639–648. [Google Scholar]
- 14.Durre P, Fischer R J, Kuhn A, Lorenz K, Schreiber W, Sturzenhofecker B, Ullmann S, Winzer K, Sauer U. Solventogenic enzymes of Clostridium acetobutylicum: catalytic properties, genetic organization, and transcriptional regulation. FEMS Microbiol Rev. 1995;17:251–262. doi: 10.1111/j.1574-6976.1995.tb00209.x. [DOI] [PubMed] [Google Scholar]
- 15.Ewing B, Hillier L, Wendl M C, Green P. Base-calling automated sequencer traces using phred. I. Accuracy assessment. Genome Res. 1998;8:175–185. doi: 10.1101/gr.8.3.175. [DOI] [PubMed] [Google Scholar]
- 16.Felsenstein J. Inferring phylogenies from protein sequences by parsimony, distance, and likelihood methods. Methods Enzymol. 1996;266:418–427. doi: 10.1016/s0076-6879(96)66026-1. [DOI] [PubMed] [Google Scholar]
- 17.Fiegler H, Bassias J, Jankovic I, Bruckner R. Identification of a gene in Staphylococcus xylosus encoding a novel glucose uptake protein. J Bacteriol. 1999;181:4929–4936. doi: 10.1128/jb.181.16.4929-4936.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Fleischmann R D, Adams M D, White O, Clayton R A, Kirkness E F, Kerlavage A R, Bult C J, Tomb J F, Dougherty B A, Merrick J M, et al. Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269:496–512. doi: 10.1126/science.7542800. [DOI] [PubMed] [Google Scholar]
- 19.Fraser C M, Gocayne J D, White O, Adams M D, Clayton R A, Fleischmann R D, Bult C J, Kerlavage A R, Sutton G, Kelley J M, et al. The minimal gene complement of Mycoplasma genitalium. Science. 1995;270:397–403. doi: 10.1126/science.270.5235.397. [DOI] [PubMed] [Google Scholar]
- 20.Gabriel C L. Butanol Fermentation Process. Ind Eng Chem. 1928;20:1063–1067. [Google Scholar]
- 21.Girbal L, Croux C, Vasconcelos I, Soucaille P. Regulation of metabolic shifts in Clostridium acetobutylicum ATCC 824. FEMS Microbiol Rev. 1995;17:287–297. [Google Scholar]
- 22.Girbal L, Soucaille P. Regulation of Clostridium acetobutylicum metabolism as revealed by mixed-substrate steady-state continuous cultures: role of NADH/NAD ratio and ATP pool. J Bacteriol. 1994;176:6433–6438. doi: 10.1128/jb.176.21.6433-6438.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Girbal L, Soucaille P. Regulation of solvent production in Clostridium acetobutylicum. Trends Biotechnol. 1998;16:11–16. [Google Scholar]
- 24.Gottwald M, Gottschalk G. The internal pH of Clostridium acetobutylicum and its effect on the shift from acid to solvent formation. Arch Microbiol. 1985;143:42–46. [Google Scholar]
- 25.Green E M, Boynton Z L, Harris L M, Rudolph F B, Papoutsakis E T, Bennett G N. Genetic manipulation of acid formation pathways by gene inactivation in Clostridium acetobutylicum ATCC 824. Microbiology (Reading, United Kingdom) 1996;142:2079–2086. doi: 10.1099/13500872-142-8-2079. [DOI] [PubMed] [Google Scholar]
- 26.Grishin N V, Wolf Y I, Koonin E V. From complete genomes to measures of substitution rate variability within and between proteins. Genome Res. 2000;10:991–1000. doi: 10.1101/gr.10.7.991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Haldenwang W G. The sigma factors of Bacillus subtilis. Microbiol Rev. 1995;59:1–30. doi: 10.1128/mr.59.1.1-30.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Harris L M, Desai R P, Welker N E, Papoutsakis E T. Characterization of recombinant strains of the Clostridium acetobutylicum butyrate kinase inactivation mutant: need for new phenomenological models for solventogenesis and butanol inhibition? Biotechnol Bioeng. 2000;67:1–11. [PubMed] [Google Scholar]
- 29.Higgins D G, Thompson J D, Gibson T J. Using CLUSTAL for multiple sequence alignments. Methods Enzymol. 1996;266:383–402. doi: 10.1016/s0076-6879(96)66024-8. [DOI] [PubMed] [Google Scholar]
- 30.Huynen M A, Dandekar T, Bork P. Variation and evolution of the citric-acid cycle: a genomic perspective. Trends Microbiol. 1999;7:281–291. doi: 10.1016/s0966-842x(99)01539-5. [DOI] [PubMed] [Google Scholar]
- 31.Jiang M, Shao W, Perego M, Hoch J A. Multiple histidine kinases regulate entry into stationary phase and sporulation in Bacillus subtilis. Mol Microbiol. 2000;38:535–542. doi: 10.1046/j.1365-2958.2000.02148.x. [DOI] [PubMed] [Google Scholar]
- 32.Johnson J L, Chen J S. Taxonomic relationships among strains of Clostridium acetobutylicum and other phenotypically similar organisms. FEMS Microbiol Rev. 1995;17:233–240. [Google Scholar]
- 33.Jones D T, Keis S. Origins and relationships of industrial solvent-producing clostridial strains. FEMS Microbiol Rev. 1995;17:223–232. [Google Scholar]
- 34.Jones D T, Woods D R. Acetone-butanol fermentation revisited. Microbiol Rev. 1986;50:484–524. doi: 10.1128/mr.50.4.484-524.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Junelles A M, Janati-Idrissi R, El Kanouni A, Petitdemange H, Gay R. Acetone-butanol fermentation by mutants selected for resistance to acetate and butyrate halogen analogs. Biotechnol Lett. 1987;9:175–178. [Google Scholar]
- 36.Kakiuchi M, Isui A, Suzuki K, Fujino T, Fujino E, Kimura T, Karita S, Sakka K, Ohmiya K. Cloning and DNA sequencing of the genes encoding Clostridium josui scaffolding protein CipA and cellulase CelD and identification of their gene products as major components of the cellulosome. J Bacteriol. 1998;180:4303–4308. doi: 10.1128/jb.180.16.4303-4308.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kanehisa M, Goto S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30. doi: 10.1093/nar/28.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Keis S, Bennett C F, Ward V K, Jones D T. Taxonomy and phylogeny of industrial solvent-producing clostridia. Int J Syst Bacteriol. 1995;45:693–705. doi: 10.1099/00207713-45-4-693. [DOI] [PubMed] [Google Scholar]
- 39.Kelly G, Prasannan S, Daniell S, Fleming K, Frankel G, Dougan G, Connerton I, Matthews S. Structure of the cell-adhesion fragment of intimin from enteropathogenic Escherichia coli. Nat Struct Biol. 1999;6:313–318. doi: 10.1038/7545. [DOI] [PubMed] [Google Scholar]
- 40.Kruus K, Lua A C, Demain A L, Wu J H. The anchorage function of CipA (CelL), a scaffolding protein of the Clostridium thermocellum cellulosome. Proc Natl Acad Sci USA. 1995;92:9254–9258. doi: 10.1073/pnas.92.20.9254. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kunst F, Ogasawara N, Moszer I, Albertini A M, Alloni G, Azevedo V, Bertero M G, Bessieres P, Bolotin A, Borchert S, Borriss R, Boursier L, Brans A, Braun M, Brignell S C, Bron S, Brouillet S, Bruschi C V, Caldwell B, Capuano V, Carter N M, Choi S-K, Codani J-J, Connerton I F, Cummings N J, Daniel R A, Denizot F, Devine K M, Dusterhoft A, Ehrlich S D, Emmerson P T, Entian K D, Errington J, Fabret C, Ferrari E, Foulger D, Fritz C, Fujita M, Fujita Y, Fuma S, Galizzi A, Galleron N, Ghim S-Y, Glaser P, Goffeau A, Golightly E J, Grandi G, Guiseppi G, Guy B J, Haga K, et al. The complete genome sequence of the gram-positive bacterium Bacillus subtilis. Nature (London) 1997;390:249–256. doi: 10.1038/36786. [DOI] [PubMed] [Google Scholar]
- 42.Lee S F, Forsberg C W, Gibbins L N. Xylanolytic activity of Clostridium acetobutylicum. Appl Environ Microbiol. 1985;50:1068–1076. doi: 10.1128/aem.50.4.1068-1076.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lee S F, Forsberg C W, Rattray J B. Purification of characterization of two endoxylanases from Clostridium acetobutylicum ATCC 824. Appl Environ Microbiol. 1987;53:644–650. doi: 10.1128/aem.53.4.644-650.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Leonard C J, Aravind L, Koonin E V. Novel families of putative protein kinases in bacteria and archaea: evolution of the “eukaryotic” protein kinase superfamily. Genome Res. 1998;8:1038–1047. doi: 10.1101/gr.8.10.1038. [DOI] [PubMed] [Google Scholar]
- 45.Lobry J R. Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol. 1996;13:660–665. doi: 10.1093/oxfordjournals.molbev.a025626. [DOI] [PubMed] [Google Scholar]
- 46.Marino M, Braun L, Cossart P, Ghosh P. Structure of the lnlB leucine-rich repeats, a domain that triggers host cell invasion by the bacterial pathogen L. monocytogenes. Mol Cell. 1999;4:1063–1072. doi: 10.1016/s1097-2765(00)80234-8. [DOI] [PubMed] [Google Scholar]
- 47.Mattson D M, Rogers P. Analysis of Tn916-induced mutants of Clostridium acetobutylicum altered in solventogenesis and sporulation. J Ind Microbiol. 1994;13:258–268. doi: 10.1007/BF01569758. [DOI] [PubMed] [Google Scholar]
- 48.Mermelstein L D, Welker N E, Bennett G N, Papoutsakis E T. Expression of cloned homologous fermentative genes in Clostridium acetobutylicum ATCC 824. Bio/Technology. 1992;10:190–195. doi: 10.1038/nbt0292-190. [DOI] [PubMed] [Google Scholar]
- 49.Mermelstein L D, Welker N E, Petersen D J, Bennett G N, Papoutsakis E T. Genetic and metabolic engineering of Clostridium acetobutylicum ATCC 824. Ann N Y Acad Sci. 1994;721:54–68. doi: 10.1111/j.1749-6632.1994.tb47376.x. [DOI] [PubMed] [Google Scholar]
- 50.Minton N P, Brehm J K, Oultram J D, Thompson D E, Swinfield T J, Pennock A, Schimming S, Whelan S M, Vetter U, et al. Clinical and molecular aspects of anaerobes. Proceedings of the 6th Biennial Anaerobe Discussion Group International Symposium. 1990. Vector systems for the genetic analysis of Clostridium acetobutylicum; pp. 187–201. [Google Scholar]
- 51.Minton, N. P., T. J. Swinfield, J. K. Brehm, S. M. Whelan, and J. D. Oultram. 1993. Vectors for use in Clostridium acetobutylicum. Genet. Mol. Biol. Anaerobic Bact. 120–140.
- 52.Mitchell W J. Physiology of carbohydrate to solvent conversion by Clostridia. Adv Microb Physiol. 1998;39:31–130. doi: 10.1016/s0065-2911(08)60015-6. [DOI] [PubMed] [Google Scholar]
- 53.Morris J G. History and future potential for the Clostridia in bio/technology. In: Woods D, editor. The Clostridia and bio/technology. Reading, Mass: Butterworth-Heinemann; 1993. p. 443. [Google Scholar]
- 54.Nair R V, Green E M, Watson D E, Bennett G N, Papoutsakis E T. Regulation of the sol locus genes for butanol and acetone formation in Clostridium acetobutylicum ATCC 824 by a putative transcriptional repressor. J Bacteriol. 1999;181:319–330. doi: 10.1128/jb.181.1.319-330.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Overbeek R, Larsen N, Pusch G D, D'Souza M, Selkov E, Jr, Kyrpides N, Fonstein M, Maltsev N, Selkov E. WIT: integrated system for high-throughput genome sequence analysis and metabolic reconstruction. Nucleic Acids Res. 2000;28:123–125. doi: 10.1093/nar/28.1.123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Papoutsakis E T, Bennett G N. Molecular regulation and metabolic engineering of solvent production by Clostridium acetobutylicum. Bioprocess Technol. 1999;24:253–279. [Google Scholar]
- 57.Petitdemange H, Cherrier C, Bengone J M, Gay R. Study of the NADH and NADPH-ferredoxin oxidoreductase activities in Clostridium acetobutylicum. Can J Microbiol. 1997;23:152–160. [PubMed] [Google Scholar]
- 58.Ponting C P, Aravind L, Schultz J, Bork P, Koonin E V. Eukaryotic signalling domain homologues in archaea and bacteria. Ancient ancestry and horizontal gene transfer. J Mol Biol. 1999;289:729–745. doi: 10.1006/jmbi.1999.2827. [DOI] [PubMed] [Google Scholar]
- 59.Qureshi N, Blaschek H P. Production of acetone butanol ethanol (ABE) by a hyper-producing mutant strain of Clostridium beijerinckii BA101 and recovery by pervaporation. Biotechnol Prog. 1999;15:594–602. doi: 10.1021/bp990080e. [DOI] [PubMed] [Google Scholar]
- 60.Ravagnani A, Jennert K C, Steiner E, Grunberg R, Jefferies J R, Wilkinson S R, Young D I, Tidswell E C, Brown D P, Youngman P, Morris J G, Young M. Spo0A directly controls the switch from acid to solvent production in solvent-forming Clostridia. Mol Microbiol. 2000;37:1172–1185. doi: 10.1046/j.1365-2958.2000.02071.x. [DOI] [PubMed] [Google Scholar]
- 61.Reizer J, Bachem S, Reizer A, Arnaud M, Saier M H, Jr, Stulke J. Novel phosphotransferase system genes revealed by genome analysis—the complete complement of PTS proteins encoded within the genome of Bacillus subtilis. Microbiology. 1999;145:3419–3429. doi: 10.1099/00221287-145-12-3419. [DOI] [PubMed] [Google Scholar]
- 62.Rogers P G, Gottschalk C. Biochemistry and regulation of acid and solvent formation in Clostridia. In: Woods D, editor. The Clostridia and bio/technology. Reading, Mass: Butterworth-Heinemann; 1993. p. 443. [Google Scholar]
- 63.Saier M H, Jr, Chauvaux S, Cook G M, Deutscher J, Paulsen I T, Reizer J, Ye J J. Catabolite repression and inducer control in Gram-positive bacteria. Microbiology. 1996;142:217–230. doi: 10.1099/13500872-142-2-217. [DOI] [PubMed] [Google Scholar]
- 64.Schaffer A A, Wolf Y I, Ponting C P, Koonin E V, Aravind L, Altschul S F. IMPALA: matching a protein sequence against a collection of PSI-BLAST-constructed position-specific score matrices. Bioinformatics. 1999;15:1000–1011. doi: 10.1093/bioinformatics/15.12.1000. [DOI] [PubMed] [Google Scholar]
- 65.Schultz J, Copley R R, Doerks T, Ponting C P, Bork P. SMART: a web-based tool for the study of genetically mobile domains. Nucleic Acids Res. 2000;28:231–234. doi: 10.1093/nar/28.1.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Schultz J, Milpetz F, Bork P, Ponting C P. SMART, a simple modular architecture research tool: identification of signaling domains. Proc Natl Acad Sci USA. 1998;95:5857–5864. doi: 10.1073/pnas.95.11.5857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Shine J, Dalgarno L. Terminal-sequence analysis of bacterial ribosomal RNA. Correlation between the 3′-terminal-polypyrimidine sequence of 16-S RNA and translational specificity of the ribosome. Eur J Biochem. 1975;57:221–230. doi: 10.1111/j.1432-1033.1975.tb02294.x. [DOI] [PubMed] [Google Scholar]
- 68.Shoham Y, Lamed R, Bayer E A. The cellulosome concept as an efficient microbial strategy for the degradation of insoluble polysaccharides. Trends Microbiol. 1999;7:275–281. doi: 10.1016/s0966-842x(99)01533-4. [DOI] [PubMed] [Google Scholar]
- 69.Sierra J, Acousta R, Montoya D, Buitrago G, Silva E. Isolation of spontaneous butanol-resistant mutants of Clostridium acetobutylicum. Rev Colomb Cienc Quimico-Farm. 1996;25:26–35. [Google Scholar]
- 70.Smith D R, Doucette-Stamm L A, Deloughery C, Lee H, Dubois J, Aldredge T, Bashirzadeh R, Blakely D, Cook R, Gilbert K, Harrison D, Hoang L, Keagle P, Lumm W, Pothier B, Qiu D, Spadafora R, Vicaire R, Wang Y, Wierzbowski J, Gibson R, Jiwani N, Caruso A, Bush D, Reeve J N, et al. Complete genome sequence of Methanobacterium thermoautotrophicum deltaH: functional analysis and comparative genomics. J Bacteriol. 1997;179:7135–7155. doi: 10.1128/jb.179.22.7135-7155.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Soucaille P, Joliff G, Izard A, Goma G. Butanol tolerance and autobacteriocin production by Clostridium acetobutylicum. Curr Microbiol. 1987;14:295–299. [Google Scholar]
- 72.Stackebrandt E, Kramer I, Swiderski J, Hippe H. Phylogenetic basis for a taxonomic dissection of the genus Clostridium. FEMS Immunol Med Microbiol. 1999;24:253–258. doi: 10.1111/j.1574-695X.1999.tb01291.x. [DOI] [PubMed] [Google Scholar]
- 73.Stackebrandt E, Rainey F A. Phylogenetic relationships. In: Rood J, McClane B A, Songer J G, Titball R W, editors. The Clostridia: molecular biology and pathogenesis. San Diego, Calif: Academic Press; 1997. [Google Scholar]
- 74.Steeves R M, Denton M E, Barnard F C, Henry A, Lambert J M. Identification of three oligosaccharide binding sites in ricin. Biochemistry. 1999;38:11677–11685. doi: 10.1021/bi990493o. [DOI] [PubMed] [Google Scholar]
- 75.Stragier P, Losick R. Molecular genetics of sporulation in Bacillus subtilis. Annu Rev Genet. 1996;30:297–341. doi: 10.1146/annurev.genet.30.1.297. [DOI] [PubMed] [Google Scholar]
- 76.Subramanian G, Koonin E V, Aravind L. Comparative genome analysis of the pathogenic spirochetes Borrelia burgdorferi and Treponema pallidum. Infect Immun. 2000;68:1633–1648. doi: 10.1128/iai.68.3.1633-1648.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Tamaru Y, Karita S, Ibrahim A, Chan H, Doi R H. A large gene cluster for the clostridium cellulovorans cellulosome. J Bacteriol. 2000;182:5906–5910. doi: 10.1128/jb.182.20.5906-5910.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Tatusov R L, Galperin M Y, Natale D A, Koonin E V. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000;28:33–36. doi: 10.1093/nar/28.1.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Terraciano J S, Rapaport E, Kashket E R. Stress and growth-phase associated proteins of Clostridium acetobutylicum. Appl Environ Microbiol. 1988;54:1989–1995. doi: 10.1128/aem.54.8.1989-1995.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Vasconcelos I, Girbal L, Soucaille P. Regulation of carbon and electron flow in Clostridium acetobutylicum grown in chemostat culture at neutral pH on mixtures of glucose and glycerol. J Bacteriol. 1994;176:1443–1450. doi: 10.1128/jb.176.5.1443-1450.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Walker D R, Koonin E V. 5th International Conference on Intelligent Systems for Molecular Biology. Menlo Park, Calif: AAAI Press; 1997. SEALS: a system for easy analysis of lots of sequences; pp. 333–339. [PubMed] [Google Scholar]
- 82.Weitzmann, C. 1915. Improvements in the bacterial fermentation of carbohydrates and in bacterial culture for same. Great Britain patent 4845.
- 83.Weyer E R, Rettger L F. A comparative study of six different strains of the organism commonly concerned in large-scale production of butyl alcohol and acetone by the biological process. J Bacteriol. 1927;14:399–424. doi: 10.1128/jb.14.6.399-424.1927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Wilkinson S R, Young M. Targeted integration of genes into the Clostridium acetobutylicum chromosome. Microbiology (Reading, United Kingdom) 1994;140:89–95. [Google Scholar]
- 85.Wolf Y I, Aravind L, Grishin N V, Koonin E V. Evolution of aminoacyl-tRNA synthetases—analysis of unique domain architectures and phylogenetic trees reveals a complex history of horizontal gene transfer events. Genome Res. 1999;9:689–710. [PubMed] [Google Scholar]
- 86.Woods D R, editor. The Clostridia and bio/technology. Boston, Mass: Butterworth-Heinemann; 1993. [Google Scholar]
- 87.Woods D R. The genetic engineering of microbial solvent production. Trends Biotechnol. 1995;13:259–264. doi: 10.1016/S0167-7799(00)88960-X. [DOI] [PubMed] [Google Scholar]
- 88.Wootton J C. Non-globular domains in protein sequences: automated segmentation using complexity measures. Comput Chem. 1994;18:269–285. doi: 10.1016/0097-8485(94)85023-2. [DOI] [PubMed] [Google Scholar]