Abstract
Raman spectroscopy is a rapid analysis method of biological samples without labeling and destruction. At present, the commonly used Raman spectrum classification models include CNN, RNN, etc. The transformer has not been used for Raman spectrum identification. This paper introduces a new method of transformer combined with Raman spectroscopy to identify deep-sea cold seep microorganisms at the single-cell level. We collected the Raman spectra of eight cold seep bacteria, each of which has at least 500 spectra for the training of transformer model. We compare the transformer classification model with other deep learning classification models. The experimental results show that this method can improve the accuracy of microbial classification. Our average isolation level accuracy is more than 97%.
Subject terms: Chemical biology, Microbiology
Introduction
Oceans occupy 70.8 percent of the earth's surface area. Cold seep is fluids from below the seabed sedimentary interface, which will overflow from the seabed in the form of leakage. The main components of these fluids are water, hydrocarbons (natural gas and oil), hydrogen sulfide and fine-grained sediments. These fluids are the source of various dense microbial and animal populations. Microbial populations in this unique environment have been extensively studied. Numerous studies have shown that deep-sea cold seeps, which contain large amounts of combustible ice resources, may be the third ecological environment on Earth found to carry out large-scale nitrogen fixation and provide large amounts of organic matter to deep-sea ecosystems. Therefore, it is very important to conduct research on deep cold seep bacteria and the identification and screening of specific functional bacteria1. The two main types of traditional methods of bacterial detection are bacteriological diagnosis and immunoserological diagnosis. Among them, bacteriological diagnosis is based on the morphology of bacteria (size, shape, arrangement, nucleoplasmic distribution, etc.), bacterial composition, metabolites and nucleic acids. The accuracy of morphological diagnosis is too low, while the study of bacterial composition and metabolism is often too costly, complicated and time-consuming. Immunoserological diagnosis, on the other hand, requires labeling of bacteria and expensive scientific instruments. Most of these methods require bacterial culture to complete microbial identification, delaying the detection process. Therefore, a new label-free, culture-free, non-contact, rapid bacterial identification method at the single-cell level is currently needed for bacteriological studies2–4. In recent years, Raman spectroscopy is often used in the rapid identification and analysis of microorganisms. Raman spectroscopy is an unlabeled5–7, non-invasive, rapid8,9 in situ cell identification method that can be used to identify10 and research11 microbial single-cell species. The single-cell Raman atlas of microorganisms contains a wealth of biochemical data in various physiological states.
Raman spectroscopy can provide biochemical information of bacteria, such as DNA, RNA, proteins, lipids, carbohydrates, etc. Raman spectra are equally capable of providing information about bacterial pigments12–14. For example, the Raman characteristic peaks of carotenoids, which are commonly found in microorganisms, are 1004, 1157, and 1520 cm−1 (C=C stretching vibration)15. In addition, Raman spectroscopy combined with machine learning, deep learning and other classification methods can reflect the differences between different species of microorganisms, thus enabling the identification of bacteria. Raman spectroscopy was able to distinguish Gram-positive from Gram-negative bacteria, and some peaks at 540 and 1380 cm−1 were significantly different for Gram-positive bacteria compared to Gram-negative bacteria16. Ho et al. successfully identified 30 common pathogens using deep learning, achieving an average separation level accuracy of over 82% and antibiotic treatment identification accuracy of 97.0 ± 0.3% on a low signal-to-noise spectrum17.
Current Raman spectral classification algorithms are classified into two types: feature-based classifiers and end-to-end deep learning classifiers. Typically, feature-based approaches derive numerical characteristics from raw signal data18. Partial least squares (PLS), principal component analysis (PCA), independent principal component analysis (ICA), and wavelet analysis (WA), among others, are signal processing algorithms used for feature extraction. The collected characteristics are then subjected to various classification methods based on multiple regression, such as linear discriminant analysis, support vector machine, random forest, and so on19–21. However, when sample sources and spectral acquisition conditions change, the spectral response to substances is not completely linear, which may reduce the model's prediction ability. Although support vector machines (SVM) are said to outperform most multivariate analysis methods21, their classification accuracy will be affected when dealing with large data sets. Traditional analytical modeling methods continue to perform poorly in terms of fit and robustness.
Deep learning-based classification methods have recently received a lot of attention because they perform well in certain classification tasks22–25. Deep learning method has powerful learning function and can eliminate the influence of nonlinearity. Convolutional neural networks (CNN) can accurately solve complex problems involving large amounts of data by simulating the structure and functions of computer biological neural networks26. However, CNN relies heavily on the choice of kernel. It will lose some time series information. If there is no deep structure, it is difficult to perceive the wide internal relationship of the signal, which may lead to a large number of calculations. Various recurrent neural networks (RNNs) have been proposed to learn the time characteristics of Raman signals27. However, the RNN steps cannot be parallelized and the efficiency is low. Another problem is that RNN only works on previous memory and current state. However, the Raman signal is continuous. Neither CNN nor RNN can well perceive the global dependence of Raman signals. In 2017, Google's machine translation team only adopted the attention mechanism to accomplish machine translation jobs, abandoning network architectures like RNN and CNN entirely28. Later, it was used in the field of image classification and achieved good results29–32. This method can calculate the representation of sequences with dependencies between different locations.
In this study, we build a Raman spectral database containing eight species of deep-sea cold seep bacteria and propose a new method for classifying Raman spectra using transformer structures. To the best of our knowledge, this is the first time that a transformer is used to classify Raman spectra. We use the commonly used classification methods and transformer structures to identify Raman spectra of deep-sea cold seep bacteria, and compare the identification results of different methods. The results show that the transformer structure obtained a high accuracy in the task of identifying eight species of cold seep bacteria.
Materials and methods
Sample preparation
We selected 8 pure cultured microorganisms of different species and genera isolated from deep-sea sediments. See Table S1 for the sea area where each microorganism is located. Select a single colony and transfer it to 10 ml 2216E liquid medium for activation. Inoculate into the fresh culture medium in the ratio of 1:100, culture at 150 rpm in a shaking table at 10 °C for 24 h, and then take 2 ml culture medium for centrifugation to collect the bacteria.
Raman spectroscopy acquisition
In this experiment, the confocal Raman spectrometer (Hooke P300, Hooke Instrument Co., Ltd., China) is used to collect the Raman spectrum, equipped with 532 nm solid-state laser (Cobolt 08-DPL, Cobolt, Sweden) and—70 °C cooled CCD detector (PIXIS 100 B, Princeton instruments, USA). The laser is a continuous wave laser with an output power of 50mW, and the spectral bandwidth (FWHM) of the laser is 1 MHz. The laser beam was focused by a 100× objective (LMPlan FLN 100× , Olympus, Japan). The numerical aperture (NA) of the objective is 0.8, and the actual spot size after convergence by the objective is 406 nm.The power irradiated on each sample is 5mW and the exposure time is 5 s. The size of the bacteria was around 1 μm and we measured a Raman spectrum at the middle of each bacteria. At least 500 spectra were collected for each sample to limit the impact of spectral noise (Fig. 1).
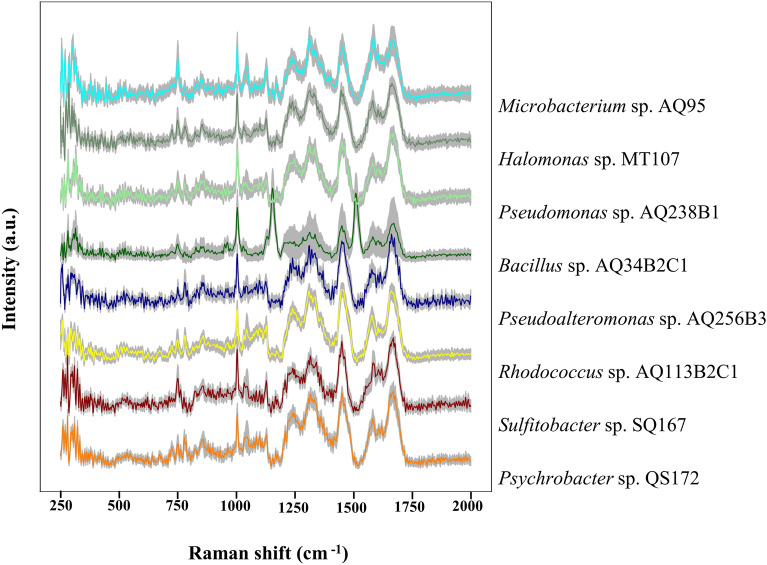
Figure 1.
Raman spectra of eight cold seep microorganisms. Each strain has at least 500 spectra. The solid line represents the average value of Raman spectrum, and the standard deviation is represented by shadow.
Data processing
Each process from the transmitter to the receiver of the spectrometer may interfere with the noise of the obtained signal, which affects the further analysis of Raman spectrum. Therefore, it is very necessary to preprocess the collected spectral data. We removed cosmic rays from the spectrum, corrected the baseline with the Subbackmod function in Biodata's toolbox, and normalized with the Mapminmax function (Fig. 2)33.
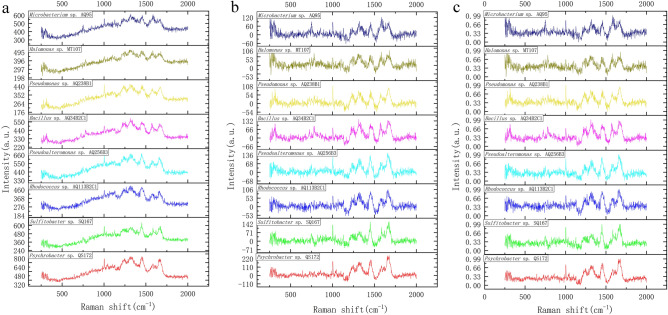
Figure 2.
Examples of raw and preprocessed spectra examples. (a) Untreated spectra of 8 bacteria. (b) Spectra after baseline subtraction. (c) Spectra after normalization. These spectra are input into our model.
Results and discussion
Model evaluation
In this study, we used AlexNet, ResNet models respectively to analyze and verify the feasibility and accuracy of Raman spectroscopy combined with deep learning model to classify cold-seep microorganisms. In order to better apply these methods to this experiment, we fine-tune the above model. For the above model, we use one-dimensional convolution layers instead of two-dimensional convolution layers.
We used the fivefold cross-validation method, to test the classification model of classifying data capabilities, and minimize caused by inappropriate dataset partition problems, such as the fitting model on the training set, the before-fitting results may not be a model, but because the dataset partition is not reasonable. First, we created five data sets by dividing the data for each bacterium into five equal portions. The classification model was divided into four groups for training, and one group was used as test data. In order to avoid over-fitting of the neural network, the four groups of data were randomly reorganized into two parts: 80% data for training and 20% data for verification. In the process of cross-validation, the accuracy of the five optimization models was compared, and the classification model with the best accuracy was selected from the five optimization models.
Construction transformer network framework
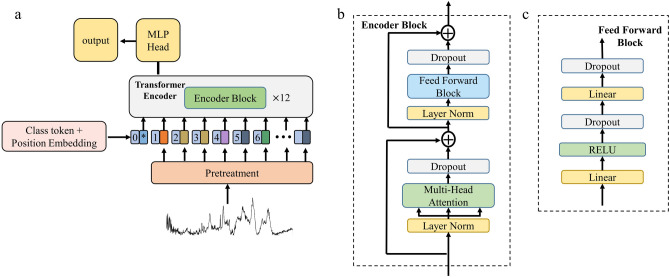
Our network architecture is adapted from the description of Alexey dosovitskiy et al.34 as shown in Fig. 3a. The Raman spectrum is transmitted into the transformer model after pretreatment.。The transformer model is composed of a transformer encoder and a multi-layer perceptron (MLP), which is composed of a linear layer and an active layer. Position encoding is added to the input embedding at the top of the encoder to maintain the sequence's relative or absolute position. In addition, an extra learnable class token is fed to the transformer network that attends to all other tokens. The transformer encoder consists of 12 encoder blocks stacked repeatedly. Encoder block is composed of two blocks. The structure is shown in Fig. 3b. The first block is a multi-head attention block28, which uses 12 attention heads and an embedding dimension of 768. The second block is a simple fully connected feedforward neural network. The module framework of each fully connected feedforward neural network is shown in Fig. 3c. The two blocks are connected by the residual network structure. The Adam optimizer with default settings is used to train the model35. The cross-entropy function is used as the loss function.
Figure 3.
(a) Structure diagram of transformer classification model for Raman Spectrum Classification. (b) Structure diagram of encoder block. (c) Structure diagram of feedforward block.
Classification results of cold seep bacteria by transformer model
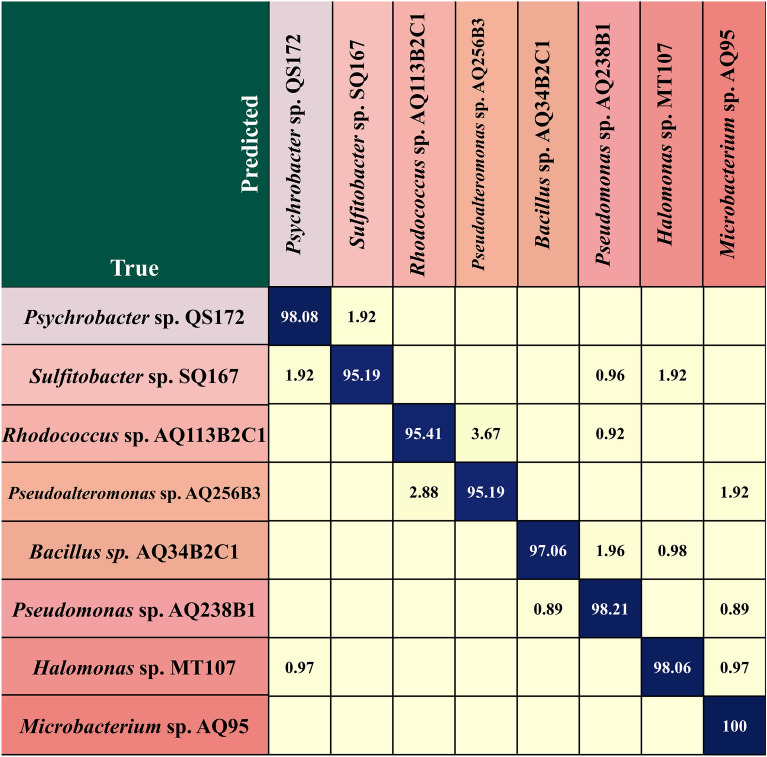
We used a trained transformer classification model to identify the species of each microbial cell based on the flora in the test data set. Each branching group in the test dataset was predicted by the trained Transformer classification model and assigned to a specific category。For identifying different microbial species, our transformer classification model has an average accuracy of 97.3%. As shown in Fig. 4, the classification accuracy of bacterial Microbacterium sp. AQ95 is 100%, the recognition accuracy of Psychrobacter sp. QS172, Bacillus sp. AQ34B2C1, Pseudomonas sp. AQ238B1 and Halomonas sp. MT107 is higher than 97% and the identification accuracy of Sulfitobacter sp. SQ167, Rhodococcus sp. AQ113B2C1, and Pseudoalteromonas sp. AQ256B3 is higher than 95%.
Figure 4.
Confusion matrix of 8 cold seep bacteria. All spectra of each bacterium are classified into the correct category.
Receiver operating characteristic (ROC) curves were used to assess the specificity and sensitivity of five species classifications in the fivefold cross-validation study (Fig. 5). The eight strains' mean AUC (area under the ROC curve) was all greater than 0.97, indicating that our classification model had high specificity and sensitivity for classifying different microbial species.
Figure 5.
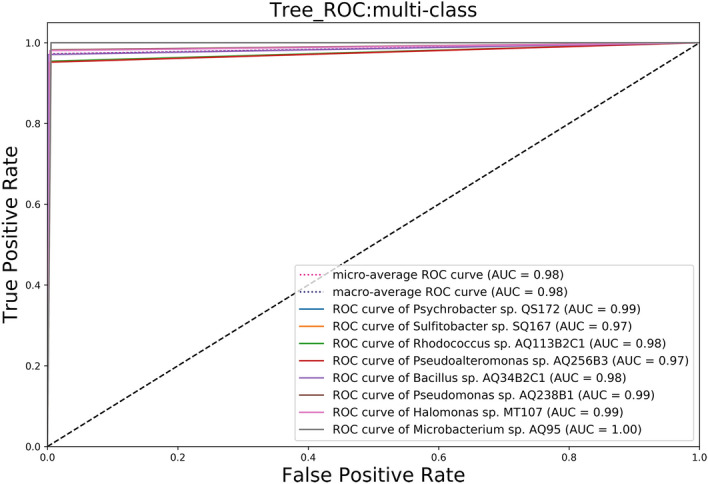
Receiver operating characteristic (ROC) curves of the Transformer model.
In comparison, we used common analytical techniques such as deep learning to predict the types of single cells. We applied the original spectral data to AlexNet and ResNet for prediction, with the accuracy of 96.5% and 95.9% respectively. We plotted the confusion matrix of the above-mentioned classification model for deep-sea bacteria (Fig. S1, S2).
In conclusion, these results suggest that Raman spectroscopy combined with Transformer is a reliable method for the accurate identification of different microorganisms at the single-cell level.
Conclusions
In this study, we analyzed eight kinds of microorganisms obtained from cold seep in different sea areas. We collected their Raman spectra and combined with transformer model to classify cold seep microorganisms. In addition, we used fivefold cross-validation to ensure good robustness of the model. Raman spectroscopy can be easily extended to new microbial applications due to its undamaging and unlabeled advantages. Meanwhile, the specificity, sensitivity, and accuracy of other common deep learning classification methods are compared. In addition, the new method of Raman spectrum classification proposed in this paper can also be applied to the accurate classification of other samples, which provides valuable insights for the accurate analysis of Raman data in the future.
Supplementary Information
Acknowledgements
This research was financially supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (No. XDA22020403).
Author contributions
B.L. and K.L. mainly conceived and listed the outline of the review. X.Q. prepared biological samples. B.L. and K.L. mainly wrote the review. W.Z. and B.L. contributed to review modification and editing. All authors contributed to the article and approved the submitted version.
Data availability
Data underlying the results presented in this paper are not publicly available at this time but may be obtained from the authors upon reasonable request.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Bo Liu and Kunxiang Liu.
Contributor Information
Weijia Zhang, Email: wzhang@idsse.ac.cn.
Bei Li, Email: beili@ciomp.ac.cn.
Supplementary Information
The online version contains supplementary material available at 10.1038/s41598-023-28730-w.
References
- 1.Yang S, et al. Genomic and enzymatic evidence of acetogenesis by anaerobic methanotrophic archaea. Nat. Commun. 2020;11:3941. doi: 10.1038/s41467-020-17860-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schroder UC, et al. Combined dielectrophoresis-Raman setup for the classification of pathogens recovered from the urinary tract. Anal. Chem. 2013;85:10717–10724. doi: 10.1021/ac4021616. [DOI] [PubMed] [Google Scholar]
- 3.Maquelin K, Dijkshoorn L, van der Reijden TJ, Puppels GJ. Rapid epidemiological analysis of acinetobacter strains by Raman spectroscopy. J. Microbiol. Methods. 2006;64:126–131. doi: 10.1016/j.mimet.2005.04.028. [DOI] [PubMed] [Google Scholar]
- 4.Rosch P, et al. On-line monitoring and identification of bioaerosols. Anal. Chem. 2006;78:2163–2170. doi: 10.1021/ac0514974. [DOI] [PubMed] [Google Scholar]
- 5.Frosch T, Popp J. Relationship between molecular structure and Raman spectra of quinolines. J. Mol. Struct. 2009;924:301–308. doi: 10.1016/j.molstruc.2008.12.019. [DOI] [Google Scholar]
- 6.Song Y, et al. Single-cell genomics based on Raman sorting reveals novel carotenoid-containing bacteria in the Red Sea. Microb. Biotechnol. 2017;10:125–137. doi: 10.1111/1751-7915.12420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jochum T, Michalzik B, Bachmann A, Popp J, Frosch T. Microbial respiration and natural attenuation of benzene contaminated soils investigated by cavity enhanced Raman multi-gas spectroscopy. Analyst. 2015;140:3143–3149. doi: 10.1039/c5an00091b. [DOI] [PubMed] [Google Scholar]
- 8.Domes C, Domes R, Popp J, Pletz MW, Frosch T. Ultrasensitive detection of antiseptic antibiotics in aqueous media and human urine using deep UV resonance Raman Spectroscopy. Anal. Chem. 2017;89:9997–10003. doi: 10.1021/acs.analchem.7b02422. [DOI] [PubMed] [Google Scholar]
- 9.Keiner R, Gruselle MC, Michalzik B, Popp J, Frosch T. Raman spectroscopic investigation of 13CO 2 labeling and leaf dark respiration of Fagus sylvatica L. (European beech) Anal. Bioanal. Chem. 2015;407:1813–1817. doi: 10.1007/s00216-014-8446-8. [DOI] [PubMed] [Google Scholar]
- 10.Strola SA, et al. Single bacteria identification by Raman spectroscopy. J. Biomed. Opt. 2014;19:111610. doi: 10.1117/1.JBO.19.11.111610. [DOI] [PubMed] [Google Scholar]
- 11.Jing X, et al. Raman-activated cell sorting and metagenomic sequencing revealing carbon-fixing bacteria in the ocean. Environ. Microbiol. 2018;20:2241–2255. doi: 10.1111/1462-2920.14268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ashton L, Lau K, Winder CL, Goodacre R. Raman spectroscopy: Lighting up the future of microbial identification. Future Microbiol. 2011;6:991–997. doi: 10.2217/fmb.11.89. [DOI] [PubMed] [Google Scholar]
- 13.Puppels G, et al. Studying single living cells and chromosomes by confocal Raman microspectroscopy. Nature. 1990;347:301–303. doi: 10.1038/347301a0. [DOI] [PubMed] [Google Scholar]
- 14.Huang WE, Griffiths RI, Thompson IP, Bailey MJ, Whiteley AS. Raman microscopic analysis of single microbial cells. Anal. Chem. 2004;76:4452–4458. doi: 10.1021/ac049753k. [DOI] [PubMed] [Google Scholar]
- 15.Marshall CP, et al. Carotenoid analysis of halophilic archaea by resonance Raman spectroscopy. Astrobiology. 2007;7:631–643. doi: 10.1089/ast.2006.0097. [DOI] [PubMed] [Google Scholar]
- 16.de Siqueira EOFS, da Silva AM, Pacheco MTT, Giana HE, Silveira L., Jr Biochemical characterization of pathogenic bacterial species using Raman spectroscopy and discrimination model based on selected spectral features. Lasers Med. Sci. 2021;36:289–302. doi: 10.1007/s10103-020-03028-9. [DOI] [PubMed] [Google Scholar]
- 17.Ho CS, et al. Rapid identification of pathogenic bacteria using Raman spectroscopy and deep learning. Nat. Commun. 2019;10:4927. doi: 10.1038/s41467-019-12898-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Phinyomark A, Phukpattaranont P, Limsakul C. Feature reduction and selection for EMG signal classification. Expert Syst. Appl. 2012;39:7420–7431. doi: 10.1016/j.eswa.2012.01.102. [DOI] [Google Scholar]
- 19.Chan JW, et al. Nondestructive identification of individual leukemia cells by laser trapping Raman spectroscopy. Anal. Chem. 2008;80:2180–2187. doi: 10.1021/ac7022348. [DOI] [PubMed] [Google Scholar]
- 20.Notingher I, et al. Multivariate analysis of Raman spectra for in vitro non-invasive studies of living cells. J. Mol. Struct. 2005;744–747:179–185. doi: 10.1016/j.molstruc.2004.12.046. [DOI] [Google Scholar]
- 21.Pan L, et al. Method for classifying a noisy Raman spectrum based on a wavelet transform and a deep neural network. IEEE Access. 2020;8:202716–202727. doi: 10.1109/access.2020.3035884. [DOI] [Google Scholar]
- 22.Mikołajczyk, A. & Grochowski, M. in 2018 International interdisciplinary PhD workshop (IIPhDW). 117–122 (IEEE).
- 23.Affonso C, Rossi ALD, Vieira FHA, de Leon Ferreira ACP. Deep learning for biological image classification. Expert Syst. Appl. 2017;85:114–122. doi: 10.1016/j.eswa.2017.05.039. [DOI] [Google Scholar]
- 24.Perez L, Wang J. The effectiveness of data augmentation in image classification using deep learning. arXiv. 2017 doi: 10.48550/arXiv.1712.04621. [DOI] [Google Scholar]
- 25.Liu B, et al. Laser tweezers Raman spectroscopy combined with deep learning to classify marine bacteria. Talanta. 2022;244:123383. doi: 10.1016/j.talanta.2022.123383. [DOI] [PubMed] [Google Scholar]
- 26.Krizhevsky A, Sutskever I, Hinton GE. ImageNet classification with deep convolutional neural networks. Commun. ACM. 2017;60:84–90. doi: 10.1145/3065386. [DOI] [Google Scholar]
- 27.He, K., Zhang, X., Ren, S. & Sun, J. in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 770–778.
- 28.Vaswani, A. et al. Attention is all you need. Adv. Neural Inf. Proc. Sys.30 (2017).
- 29.Sun, J., Xie, J. & Zhou, H. in 2021 IEEE 3rd Global Conference on Life Sciences and Technologies (LifeTech). 92–93 (IEEE).
- 30.Song, Y., Jia, X., Yang, L. & Xie, L. (2021) Transformer-based spatial-temporal feature learning for eeg decoding. arXiv preprint arXiv:2106.11170. 10.1109/LifeTech52111.2021.9391844.
- 31.Wang, X. et al. in International Conference on Medical Image Computing and Computer-Assisted Intervention. 186–195 (Springer).
- 32.Meng L, et al. Enhancing dynamic ECG heartbeat classification with lightweight transformer model. Artif. Intell. Med. 2022;124:102236. doi: 10.1016/j.artmed.2022.102236. [DOI] [PubMed] [Google Scholar]
- 33.De Gussem K, De Gelder J, Vandenabeele P, Moens L. The biodata toolbox for MATLAB. Chemom. Intell. Lab. Syst. 2009;95:49–52. doi: 10.1016/j.chemolab.2008.08.003. [DOI] [Google Scholar]
- 34.Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. 10.48550/arXiv.2010.11929 (2020).
- 35.Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980, 10.48550/arXiv.1412.6980 (2014).
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Data underlying the results presented in this paper are not publicly available at this time but may be obtained from the authors upon reasonable request.