Abstract
Methane monooxygenase (MMO) catalyzes the oxidation of methane to methanol as the first step of methane degradation. A soluble NAD(P)H-dependent methane monooxygenase (sMMO) from the type II methanotrophic bacterium WI 14 was purified to homogeneity. Sequencing of the 16S rDNA and comparison with that of other known methanotrophic bacteria confirmed that strain WI 14 is very close to the genus Methylocystis. The sMMO is expressed only during growth under copper limitation (<0.1 μM) and with ammonium or nitrate ions as the nitrogen source. The enzyme exhibits a low substrate specificity and is able to oxidize several alkanes and alkenes, cyclic hydrocarbons, aromatics, and halogenic aromatics. It has three components, hydroxylase, reductase and protein B, which is involved in enzyme regulation and increases sMMO activity about 10-fold. The relative molecular masses of the native components were estimated to be 229, 41, and 18 kDa, respectively. The hydroxylase contains three subunits with relative molecular masses of 57, 43, and 23 kDa, which are present in stoichiometric amounts, suggesting that the native protein has an α2β2γ2 structure. We detected 3.6 mol of iron per mol of hydroxylase by atomic absorption spectrometry. sMMO is strongly inhibited by Hg2+ ions (with a total loss of enzyme activity at 0.01 mM Hg2+) and Cu2+, Zn2+, and Ni2+ ions (95, 80, and 40% loss of activity at 1 mM ions). The complete sMMO gene sequence has been determined. sMMO genes from strain WI 14 are clustered on the chromosome and show a high degree of homology (at both the nucleotide and amino acid levels) to the corresponding genes from Methylosinus trichosporium OB3b, Methylocystis sp. strain M, and Methylococcus capsulatus (Bath).
Methanotrophic bacteria oxidize methane to methanol via the activity of methane monooxygenase (MMO), which, depending on the copper concentration in the cultivation medium, can be found in different fractions. At high copper concentrations, a particulate form of methane monooxygenase (pMMO) with high substrate specificity is expressed. At low copper concentration, the soluble form of the enzyme (sMMO), which possesses a very low substrate specificity and unique properties, is expressed (3). It has been shown that although all methanotrophic bacteria express the pMMO, only some strains, mainly of the type II (34, 39, 43) and type X (6) methanotrophs, are able to express the sMMO. So far, Methylomonas methanica 68-1 (24) and Methylomonas sp. strain GYJ3 (41) are the only type I methanotrophs in which the sMMO has been detected. This nonspecific enzyme oxidizes a variety of hydrocarbons, including aliphatics and aromatics (16), and seems to be highly conserved in its structure (33). Whereas little is known about the structure and mechanism of the pMMO (21, 35), the sMMO has been described and purified from Methylosinus trichosporium OB3b (11), Methylocystis sp. strain M (34), Methylobacterium sp. strain CRL-26 (38), and Methylococcus capsulatus Bath (15). The sMMO from these microorganisms are multicomponent proteins consisting of a hydroxylase, a reductase, and a regulatory protein (protein B, with the exception of the Methylobacterium strain) with similar relative molecular masses.
It was shown that the activity of the hydroxylase component can also be measured in the absence of the other components, with hydrogen peroxide as the oxygen and hydrogen donor (23, 27). The reductase is able to transfer electrons to an acceptor such as dichlorophenolindophenol or cytochrome c (5). In contrast, the activity of the third component, the regulatory protein B, can be measured only if the other components are present.
Methanotrophs able to express the sMMO are of special interest in bioremediation processes due to the extremely high oxidation potential of this enzyme system. Several research groups have studied the distribution of sMMO-possessing strains in various environments by using gene-specific probes (30, 32, 45). The sMMO from newly isolated microorganisms appear to be very highly conserved at the nucleotide sequence level compared to the well-studied sMMO from M. trichosporium and M. capsulatus (31).
The strain studied, WI 14, was isolated from swine manure and was classified as a type II methanotrophic bacterium (19). Originally, it was thought to belong to the genus Methylosinus (20), but subsequent partial sequencing of the 16S rDNA, which can be used for phylogenetic analysis (13, 14), grouped it within the genus Methylocystis (unpublished data).
In this study we describe the expression, purification, and characterization of the sMMO and the complete 16S rDNA and sMMO sequencing of isolate WI 14.
MATERIALS AND METHODS
Chemicals.
DEAE-Sepharose, Blue-Sepharose, and Superose 12 were purchased from Pharmacia (Uppsala, Sweden), and methane and propylene were purchased from Messer Griesheim. Standard proteins for gel filtration and for sodium dodecyl sulfate (SDS)-gel electrophoresis were purchased from Merck (Darmstadt, Germany) and Boehringer (Mannheim, Germany), respectively. All other chemicals used were of analytical grade and were purchased from various manufacturers.
Growth conditions and cell disruption.
Methylocystis sp. strain WI 14 was cultivated on two different mineral salts media. The standard medium contained (in milligrams per liter) the following: KH2PO4, 340; K2HPO4, 435; MgSO4 · 7H2O, 35.6; FeSO4 · 5H2O, 2.5; CaCl2 · 2H2O, 1.85; MnSO4 · H2O, 0.308; ZnSO4 · 7H2O, 0.220; and NaMoO4 · 2H2O, 0.126. (NH4)2SO4 (0.47 g/liter) was used as the nitrogen source. The NMS medium described previously (12) was modified and contained (in milligrams per liter) the following: K2HPO4, 860; KH2PO4, 530; K2SO4, 170; MgSO4 · 7H2O, 37; CaCl2, 7; FeSO4 · 7H2O, 5; ZnSO4 · 7H2O, 0.58; MnSO4 · 7H2O, 0.47; KI, 0.17; H3BO3, 0.12; CoCl2 · 5H2O, 0.10; and NaMoO4 · 2H2O. 0.10. NaNO3 (0.85 g/liter) was used as the nitrogen source. Phosphate and iron solutions were autoclaved separately and added to the final medium. The pH of the complete medium was adjusted to 6.8 with 1 M NaOH. No external copper was added to the medium.
Methylocystis sp. strain WI 14 was maintained on NMS medium containing 50% glycerol under liquid nitrogen. Cells were cultivated in 1-liter Erlenmeyer flasks containing 100 ml of the corresponding medium with a gas-tight seal. One-quarter of the gas phase was displaced by methane, which was used as the sole carbon source. Cultivation was carried out on a rotary shaker (220 rpm) at 30°C. After 44 h, in the middle of the exponential growth phase, cells were harvested by centrifugation at 6,000 × g for 20 min, washed twice in 25 mM Tris-HCl buffer (pH 7.2), and suspended in the same buffer containing 5 mM sodium thioglycolate. The cells were disrupted by being passed twice through a French pressure cell (SLM Instruments, Urbana, Ill.) operating at 20,000 lb/in2. Unbroken cells and debris were removed by centrifugation at 15,000 × g for 30 min at 4°C. The supernatant was further centrifuged at 100,000 × g for 90 min to separate the membrane from the soluble fraction.
Enzyme assays.
The activity of MMO was measured by gas chromatography (in a GC 5890 II instrument [Hewlett Packard] with a 25-m Ultra 2 capillary column 50°C isotherm, hydrogen carrier gas at 2.26 ml/min, and a flame ionization detector) with propylene as the substrate. The reaction mixture contained, in a final volume of 2 ml, 25 mM Tris-HCl buffer (pH 7.2), 5 to 10 mM NADH or NADPH, and a corresponding amount of active enzyme; it was placed in a vial with 10 ml of headspace which was sealed gas-tight. By using a gas-tight syringe, 2 ml of the gas phase was displaced by propylene. After incubation for 15 min at 30°C on a rotary shaker, the propylene oxide formed was quantified by using a calibration curve. Furthermore, the activity of the hydroxylase of sMMO was measured as described above with 10 mM H2O2 instead of the other components and NAD(P)H. The activity of sMMO was also measured with naphthalene. Solid naphthalene was added to the reaction mixture containing whole cells (optical density at 600 nm of 0.2) in 25 mM Tris-HCl buffer (pH 7.2) and incubated for 15 min at 30°C. The α- or β-naphthol formed was measured spectrophotometrically by monitoring the increase of absorption at 525 nm after addition of 0.1 μmol of Fast Blue B salt (ε = 3.81 mmol−1 cm−1 [24]).
The activity of the reductase of sMMO (component C) was assayed in a reaction mixture (1 ml) containing 50 mM sodium phosphate buffer (pH 7.0), 1 mM NAD(P)H, 0.05 mM cytochrome c, and corresponding amounts of enzyme. The increase in the absorbance of cytochrome c at 550 nm and 25°C was monitored. The specific activity was defined as micromoles of cytochrome c reduced (ε = 1 mmol−1 cm−1) per minute (U) per milligram of protein.
Protein determination.
The protein concentration was determined as described by Bradford (2) with bovine serum albumin as the standard. During purification of the components of sMMO, protein was determined by measuring the absorption at 280 nm.
Protein purification.
All purification procedures were performed at 4°C on an LKB II FPLC system (Pharmacia, Uppsala, Sweden).
The soluble cell protein was loaded on a DEAE-Sepharose FF column (20 by 150 mm) previously equilibrated with 25 mM Tris-HCl buffer (pH 7.2) containing 5 mM sodium thioglycolate (buffer A). The flow rate was 1 ml/min. Under these conditions, all the components of sMMO were bound to the column. The hydroxylase, component B, and the reductase were eluated by using 0.15, 0.3, and 0.5 M NaCl in buffer A, respectively. These fractions were concentrated and desalted by ultrafiltration with an ultrafiltration membrane (exclusion size, 100, 10, and 30 kDa, respectively; Amicon, Danvers, Mass.) and reloaded onto a DEAE-Sepharose FF column. Subsequently, the components were eluated by using linear gradients of NaCl in buffer A from 0 to 0.15 M (hydroxylase), 0.15 to 0.3 M (component B), and 0.3 to 0.5 M (reductase).
(i) Hydroxylase and component B.
After concentration and desalting by ultrafiltration, the enriched hydroxylase and component B, respectively, were loaded onto two combined Superose 12 HR gel filtration columns (10 by 300 mm) equilibrated with 50 mM sodium phosphate buffer (pH 7.5) containing 150 mM NaCl. The proteins were eluated with the same buffer and collected in 0.5-ml fractions. The flow rate was 0.4 ml/min. Active fractions were pooled and concentrated by ultrafiltration.
(ii) Reductase (component C).
After concentration and desalting by ultrafiltration, the enriched reductase was loaded onto a Blue Sepharose column (10 by 100 mm) equilibrated with buffer A. The column was washed with 0.5 M NaCl in buffer A until no protein could be detected. Reductase was eluated with a linear gradient between 0.5 and 1 M NaCl in buffer A. Active fractions were pooled and concentrated by ultrafiltration.
Purification control and estimation of molecular mass.
Gel electrophoresis was performed as described previously (40) in 10 or 12% polyacrylamide gels containing 0.1% SDS (25) by using the discontinuous buffer system originally described by Davis (7).
The molecular masses of the native components of sMMO were determined by gel filtration on a Superose 12 HR column by using standard proteins (Merck) as indicated. SDS-gels were used to determine the molecular mass of subunits of the hydroxylase and to screen for purity. The SDS-gels were silver stained (28).
Determination of the amino terminal sequence.
Purified components of sMMO were separated into subunits by SDS-gel electrophoresis and blotted to a polyvinylidene difluoride membrane (Bio-Rad). Amino-terminal amino acid sequence determination was performed with a sequencer (473A; Applied Biosystems) by the Edman method. Phenylthiohydantoin amino acids were analyzed by high-pressure liquid chromatography with a reversed-phase column (9). The determined N-terminal amino acids were used to assign the open reading frames to the appropriate components of sMMO.
Isoelectric focusing.
The isoelectric point of the components of sMMO from Methylocystis sp. strain WI 14 was determined with the Multiphor II system from Pharmacia, using SERVALYT gels (pH 3 to 9) as specified by the manufacturer. The proteins were visualized by staining with Coomassie blue R-250.
Atomic absorption spectrometry.
To determine the iron content of purified hydroxylase (1 mg was diluted in 10 mM HCl) and to determine the copper content of the cultivation media, an atomic absorption spectrum was recorded (Elan 5000; Perkin-Elmer).
Sequencing of the 16S rDNA. (i) Oligonucleotide primers.
Primers were synthesized with a DNA synthesizer (380A; Applied Biosystems). The primers 27f (26), R5 (42), R8 (8), and 1492r (26) were used.
(ii) Isolation and PCR amplification of 16S rDNA.
Total genomic DNA from WI 14 was obtained by boiling the cell suspension in sterilized aqua dest. for 5 min and removing the cell debris by centrifugation (29). PCR was carried out with primers 27f and 1492r in a thermal cycler (480; Perkin-Elmer Cetus). PCR mixtures were prepared in a total reaction volume of 50 μl and consisted of 5 μl of the supernatant fluid from the boiled bacterial suspension, 40 to 60 pmol (2 μl) of each primer, 200 μM (8 μl) deoxynucleoside triphosphates (Pharmacia), and ultrapure water (27.5 μl). The samples were overlaid with 75 μl of mineral oil, and the initial denaturation was done for 2 min at 100°C. Subsequently, after the mixture was chilled on ice, 2.5 U (0.5 μl) of Taq polymerase in 5 μl of 10× Taq DNA polymerase buffer (100 mM Tris-HCl [pH 9.0], 500 mM KCl, 15 mM MgCl2) (Pharmacia) was added to each sample. The thermal profile included 30 cycles of 1 min at 94°C (denaturation), 1 min at 65°C (reannealing), and 1 min at 72°C (DNA extension) and denaturation. All water controls were subjected to the same PCR conditions as the samples. Following the last cycle, the sample block was kept at 4°C until electrophoresis was performed.
(iii) Cloning and DNA sequencing of PCR fragments by the cycle-sequencing reaction.
The PCR product was separated by gel electrophoresis on 1% agarose (TAE 1×), cut out, and extracted from the gel with the QIAEX II gel extraction kit (Qiagen). Purified DNA was cloned in a pCR 2.1-TOPO vector with the TOPO TA cloning kit (Invitrogen) in the competent E. coli strain TOP 10. Plasmid DNA was isolated with the Wizard Plus SV Miniprep system (Promega). A sequencing PCR with the sequencing primers mentioned above was carried out with the ABI PRISM dRhodamine Terminator cycle-sequencing kit (Applied Biosystems) in a GeneAmp PCR system 2400 (Perkin-Elmer). The thermal profile included 25 cycles of 30 s at 96°C (denaturation), 5 s at 50°C (reannealing), and 4 min at 60°C (DNA extension). The product of the sequencing PCR was purified with a CENTRI-SEP column (Princeton Separations) and analyzed in an automated fluorescence sequencer (373A; Applied Biosystems).
(iv) Phylogenetic analysis.
16S rDNAs for comparison were obtained from the GenBank sequence database of the National Center for Biotechnology Information, Bethesda, Md. Sequences were aligned with the ClustalW multiple-sequence alignment of the European Bioinformatics Institute, and the dendrogram was constructed with the programs SEQBOOT, DNAPARS, DNADIST, FITCH, NEIGHBOR, and CONSENSE from the PHYLIP version 3.5c package (10).
PCR amplification and sequencing of sMMO gene cluster.
Total genomic DNA from Methylocystis sp. strain WI 14 was obtained as described above. The sMMO gene cluster was amplified in three discrete fragments. The PCR amplification and sequencing reaction conditions were identical to those described for the 16S rDNA amplification and sequencing reactions, unless stated otherwise. The primer sequences used for the sMMO amplification and sequencing reactions of the individual sMMO genes are listed in Table 1. All assigned nucleotide position numbers correspond to the sMMO gene cluster sequence of Methylocystis sp. strain M (accession no. U81594). The first fragment (2,012 bp), amplified with primers mmoXF2 and mmoYR2, starts at position 540 (the start codon A540TG of mmoX) and ends at position 2551 (which lies within mmoY). The second fragment (1,564 bp), amplified with primers mmoYF and mmoZR2, starts at position 2440 (which lies within mmoY) and ends at position 4003 (which lies within mmoZ). The third fragment (2,349 bp), amplified with primers mmoBF and mmoCR2, starts at position 3523 (which includes mmoB) and ends at position 5871 (which includes the mmoC termination codon TGA5871). An annealing temperature of 50°C was used in the amplification of the first and third fragments, and an annealing temperature of 65°C was used for the second fragment. The PCR-amplified products were cloned as described above.
TABLE 1.
Primers used for sMMO sequencing
| Primer (reference) | Locationa | Sequence (5′ to 3′) |
|---|---|---|
| mmoXF2 | 540–556 | ATGGCGATCAGTCTCGC |
| mmoX1 (32) | 1070–1099 | CGGTCCGCTGTGGAAGGGCATGAAGCGCGT |
| mmoXR2 | 2058–2075 | GATGTCCTCGAGCGTCCA |
| mmoYR2 | 2534–2551 | CGATACCAGTCAGTCGTG |
| mmoYF (30) | 2440–2461 | CCGACTGGATCGCCGGCGGCCT |
| mmoYF2 | 2811–2830 | CTCGACAAGGTCGACAATGC |
| mmoBR2 | 3608–3627 | TGAGCACGATATCCTCGATG |
| mmoBF (30) | 3523–3544 | AGTTCTTCGCCGAGGAGAACCA |
| mmoZR2 | 3982–4003 | AGTCCTGGATGAACTTCGTCGC |
| mmoZF (30) | 4022–4043 | CGCCGTTCCGCAAGAGCTACGA |
| mmoCR3 | 5065–5083 | AGGTGCGGCACAGCAGAAC |
| mmoCR4 | 5453–5472 | GCTCTTGTGCAGCGAGAAGG |
| mmoCR2 | 5848–5871 | TCAGCCGCTCGCCAGGAATTTCTC |
Coding region of Methylocystis sp. strain M (accession no. U81594).
The annealing temperature used for the sequencing PCR was 50°C except when primers mmoXR2 and mmoCR3 were used; then the annealing temperature was 45°C. Comparative analyses of the nucleotide and derived amino acid sequences were performed with MacVector 6.0 (Oxford Molecular).
RESULTS
Sequencing and comparison of 16S rDNA.
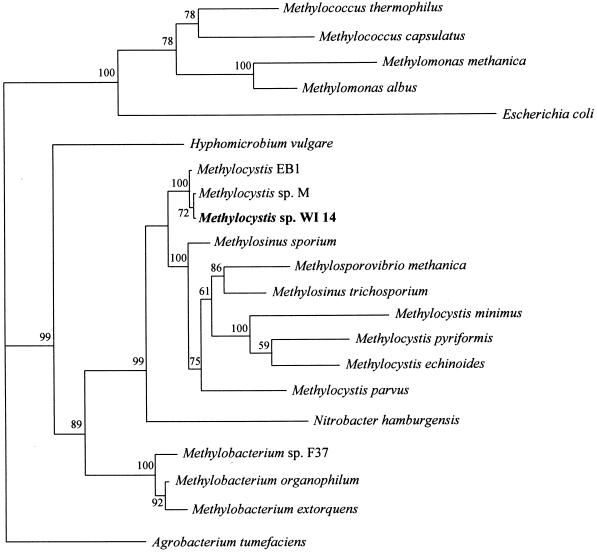
By using the universal primers 27f and 1492r, a DNA fragment from isolate WI 14 was amplified, cloned, and sequenced. Comparison of 16S rDNA revealed that strain WI 14 (accession no. AF153281) was very close to the Methylocystis group, in particular to Methylocystis sp. strain M and Methylocystis strain EB1 (Fig. 1).
FIG. 1.
Phylogenetic analysis of the 16S rDNA from Methylocystis sp. strain WI 14. The BOOTSTRAP values from 100 replicates are indicated.
Expression and properties of sMMO.
Depending on growth conditions, Methylocystis sp. strain WI 14 is able to form pMMO or sMMO. If the strain grows at copper concentrations exceeding 0.1 μM, sMMO expression is strongly repressed (Table 2). The nitrogen source also seems to influence the expression of this enzyme. Table 2 shows that although the microorganism grew on several N sources such as some amino acids, sMMO activity could be detected only if nitrate or ammonium ions were used. As expected, growth did not occur with amino acids as the sole sources of nitrogen and carbon.
TABLE 2.
Expression of sMMO from Methylocystis sp. strain WI 14 depending on the growth conditions
| Copper concn (μM) | N sourceb | OD600c at:
|
Sp act of sMMO (mU/mg)d | |
|---|---|---|---|---|
| 0 h | +24 h | |||
| 0.016a | KNO3 | 0.15 | 0.86 | 12.2 |
| 0.016a | NH4Cl | 0.07 | 0.49 | 7.6 |
| 0.016a | Carnitine | 0.07 | 0.07 | 0 |
| 0.016a | Glycine | 0.07 | 0.11 | 0 |
| 0.016a | Alanine | 0.07 | 0.11 | 0 |
| 0.016a | Cysteine | 0.07 | 0.67 | 0 |
| 0.016a | Serine | 0.07 | 0.60 | 0 |
| 0.1 | KNO3 | 0.15 | 1.16 | 4.4 |
| 0.5 | KNO3 | 0.15 | 0.92 | 1.5 |
| 1.0 | KNO3 | 0.15 | 0.86 | 1.3 |
| 5.0 | KNO3 | 0.15 | 0.23 | 0 |
Copper concentration due to the impurity of the chemicals used.
10 mM.
OD600, optical density at 600 nm.
The activity of sMMO was assayed in whole cells by the naphthalene test.
For activity, the sMMO of Methylocystis sp. strain WI 14 requires NADH or NADPH as an electron donor. In the ultracentrifuged crude extract, sMMO activity can be measured only if large amounts of the appropriate coenzyme (NADH, 0.5 mM [4.5 mU/mg] to 5 mM [19.3 mU/mg]; NADPH, 5 mM [15.5 mU/mg]) are used. The optimum sMMO activity occurs at 30°C and pH 7.2 with Tris-HCl buffer (25 mM). For stabilization of the sMMO, 5 mM sodium thioglycolate was added. In ultracentrifuged crude extracts stored at 25°C, sMMO exhibited about 55% loss of activity after 90 min. The same solutions showed about 15% loss of activity after 90 min at 4°C.
Substrate specificity.
The substrate specificity of the sMMO from Methylocystis sp. strain WI 14 was determined by using ultracentrifuged crude extract and is presented in Table 3. sMMO converts a variety of substrates such as alkanes up to heptane, alkenes, cyclic hydrocarbons, aromatics, and halogenated aromatics to their corresponding hydroxylated or oxidized products. Interestingly, the alkanes were not hydroxylated in the 1 position. All the products detected were n-alcohols (minimum detection limit, 100 pg/μl). For the halogenated substrates (chlorobenzene, fluorobenzene, and bromobenzene), a new product peak, whose retention time did not correspond to phenol, resorcine, or pyrogallol, respectively, could be observed. Furthermore, no chloride, fluoride, or bromide ions were detected, indicating that no oxidative dehalogenation occured.
TABLE 3.
Substrate specificity of the sMMO from Methylocystis sp. strain WI 14
| Substrate | Product(s) | Sp act (mU/mg) |
|---|---|---|
| Ethylene | Ethylene oxide | 33.6 |
| Propylene | Propylene oxide | 25.4 |
| Butylene | Butylene oxide | 19.1 |
| Propane | 2-Propanol | 334.3 |
| Butane | 2-Butanol | NDb |
| Pentane | n-Pentanola | —c |
| Hexane | n-Hexanol | — |
| Heptane | n-Heptanol | — |
| Diethylic ether | Ethanol, acetaldehyde | 205.7 |
| Cyclohexane | Cyclohexanol | ND |
| Benzene | Phenol | ND |
| Toluene | Cresola | — |
| Xylene | Xylenola | — |
| Naphthalene | Naphthol | 1.2 |
| Chloropentane | Chloropentanol | 3.9 |
| Chlorobenzene | Chlorophenola | — |
| Fluorobenzene | Fluorophenola | — |
| Bromobenzene | Bromophenola | — |
| Chloronaphthalene | Chloronaphthol | 0.8 |
The position of hydroxylation could not be determined.
ND, not determined.
—, determination of the specific activity was not possible because the products were not exactly identified.
Purification of sMMO.
The components of the sMMO from Methylocystis sp. strain WI 14 were separated by chromatography on DEAE. Hydroxylase, component B, and reductase were eluted with a step gradient of 0.15, 0.3, and 0.5 M NaCl, respectively.
(i) Component A (hydroxylase).
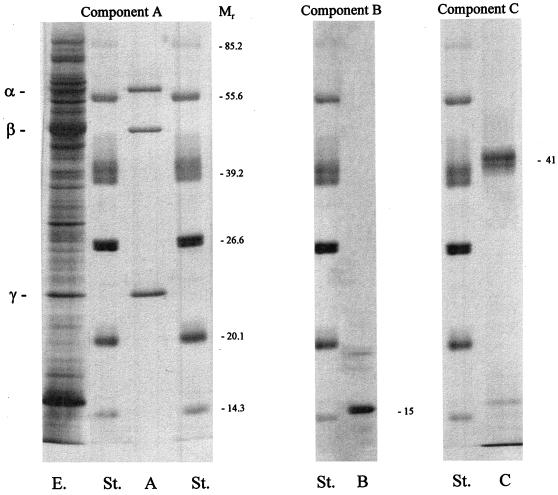
Table 4 shows details of the purification of the hydroxylase. This component could be enriched about 10-fold, resulting in a specific activity of 0.52 U/mg and a yield of 21%. These results suggest that the hydroxylase makes up about 10% of the soluble cell protein. SDS-gel electrophoresis of the purified protein indicated three distinct bands (Fig. 2).
TABLE 4.
Purification of hydroxylase and reductase of sMMO from Methylocystis sp. strain WI 14
| sMMO component | Purification step | Total amt of protein (mg) | Total activity (U) | Sp act (mU/mg) | Purification (fold) | Yield (%) |
|---|---|---|---|---|---|---|
| Hydroxylase (component A) | Ultracentrifuged extract | 487 | 24 | 49 | 1 | 100 |
| DEAE-Sepharose (0.15 M NaCl, step) | 113 | 19 | 167 | 3.4 | 79 | |
| DEAE-Sepharose (0–0.15 M NaCl, linear) | 34 | 15 | 439 | 8.9 | 64 | |
| Superose 12 HR | 10 | 5 | 518 | 10.5 | 21 | |
| Reductase (component C) | Ultracentrifuged extract | 75.1 | 11.3 | 0.15 | 1 | 100 |
| DEAE-Sepharose (0.3–0.5 M NaCl, step) | 6.8 | 12.3 | 1.82 | 12 | 109 | |
| DEAE-Sepharose (0.3–0.5 M NaCl, linear) | 0.36 | 8.5 | 23.86 | 158 | 75 | |
| Blue-Sepharose | 0.11 | 3.3 | 30.64 | 204 | 29 |
FIG. 2.
SDS-gel electrophoresis of the purified components of sMMO from Methylocystis sp. strain WI 14. St., protein standard as indicated; E., crude cell extract; A, B, and C: components of sMMO. Molecular weights are given (in thousands) to the right of the gels.
(ii) Component B.
The activity of component B can be measured only if the other components are present. Since only small amounts of the other components were available in purified form and due to the low stability of the hydroxylase, a purification table could not be generated. However, component B was identified by the increase of propylene oxide formed when it was added to ultracentrifuged extracts (addition of 10 μg of component B increased the oxidation rate by 50%). The final purification step on Superose 12 was necessary to obtain an almost pure protein band in SDS-gel electrophoresis (Fig. 2).
(iii) Component C (reductase).
Reductase was purified almost to completion by ion-exchange and affinity chromatography (Table 4). Gel electrophoresis led to the complete cessation of enzyme activity. In contrast to the hydroxylase, the 204-fold enrichment of reductase suggests that this protein makes up only about 0.5% of the soluble cell protein.
Molecular properties and stability of the components of sMMO.
The molecular properties of the components of sMMO have been determined. The native molecular mass of the hydroxylase was determined by gel filtration to be 229 kDa. SDS-gel electrophoresis shows three distinct bands with molecular masses of 57, 43, and 23 kDa (Fig. 2). Together, these results suggest that the hydroxylase has an α2β2γ2 structure. The isoelectric point was found to be 4.3. Atomic absorption spectroscopy indicated the presence of 3.6 mol of iron per mol of protein. The hydroxylase is very unstable and splits into its inactive subunits if the protein is stored frozen even for short periods (data not shown).
Component B is a single protein with an estimated molecular mass of 15 to 18 kDa. The isoelectric point was found to be 4.6. Unlike the hydroxylase component, protein B was stable when stored frozen.
The reductase appears to be a single protein with an apparent molecular mass of about 41 kDa. The stability of the reductase component was negatively affected by the gel filtration. Consequently, the native molecular mass could not be determined. Like the hydroxylase, the reductase was very unstable. Purified reductase lost about 90% of its activity when stored at 4°C for 120 h. The activity was almost completely restored when Fe2+ ions (2 mM FeSO4) were added.
Although reductase can use both NADH and NADPH as the electron donor with nearly the same activity, the determined Km values (NADPH, 5.2 mM; NADH, 6.4 μM) show that in vivo only NADH can be the physiological electron donor.
Reconstitution of the components of sMMO.
For low-level sMMO activity, only the hydroxylase (MMOH) and the reductase (MMOC) components were required. The presence of component B (MMOB) increased the sMMO activity by about 10-fold but was not essential for enzyme reaction.
Inhibition of sMMO.
Metal ions such as Zn2+, Cu2+, or Ni2+ were particularly strong inhibitors of reductase and sMMO activity. In contrast, EDTA mainly affected the reconstituted sMMO activity and presumably the hydroxylase and component B as well. Hg2+ ions completely eliminated reductase activity and consequently the reconstituted sMMO activity.
sMMO gene sequences.
sMMO genes from strain WI14 are clustered on the chromosome. Table 5 shows the high degree of homology (both at the nucleotide and amino acid level) to the corresponding genes from Methylocystis sp. strain M, Methylosinus trichosporium OB3b, and Methylococcus capsulatus (Bath). Until now, only the complete sMMO nucleotide sequence from these strains was available.
TABLE 5.
Comparison of sMMO from Methylocystis sp. strain WI 14 at the level of nucleotide and amino acid sequences
| Methylocystis sp. strain WI 14 (accession no. AF153282)a | % Similarity to sequence from:
|
|||||
|---|---|---|---|---|---|---|
|
Methylocystis sp. strain M (31) (accession no. U81594)
|
M. trichosporium OB3b (4) (accession no. X55394)
|
M. capsulatus (Bath) (accession no. M90050)
|
||||
| Nucleotide sequence | Amino acid sequence | Nucleotide sequence | Amino acid sequences | Nucleotide sequence | Amino acid sequence | |
| sMMO (mmoXf2-mmoCr2) (5254 nt) | 92 (5,281 nt) | 83 (5,254 nt) | 64 (5,344 nt) | |||
| CDS-mmoX (partial) (1,557 nt) | 95 (1,558 nt) | 99 (517 aa)b | 90 (1,557 nt) | 96 (518 aa) | 77 (557 nt) | 91 (517 aa) |
| CDS-mmoY (1,187 nt) | 91 (1,187 nt) | 97 (395 aa) | 87 (1,187 nt) | 89 (395 aa) | 65 (1,188 nt) | 69 (395 aa) |
| CDS-mmoB (416 nt) | 94 (416 nt) | 99 (138 aa) | 93 (416 nt) | 98 (138 aa) | 67 (427 nt) | 76 (141 aa) |
| CDS-mmoZ (510 nt) | 89 (509 nt) | 95 (169 aa) | 86 (510 nt) | 95 (169 aa) | 62 (520 nt) | 71 (171 aa) |
| CDS-orfY (318 nt) | 85 (336 nt) | 91 (111 aa) | 65 (326 nt) | 66 (106 aa) | 42 (382 nt) | 43 (124 aa) |
| CDC-mmoC (partial) (999 nt) | 93 (997 nt) | 99 (333 aa) | 74 (1,003 nt) | 75 (345 aa) | 60 (1,013 nt) | 69 (341 aa) |
CDS, coding sequence; nt, nucleotide.
aa; amino acid.
DISCUSSION
16S rRNA gene phylogenetic analysis revealed that isolate WI 14 possesses type II methanotroph sequences that are most closely related to the genus Methylocystis, in particular to Methylocystis sp. strain M and Methylocystis strain EB1. This is in agreement with the studies of the morphology (electron microscopy of membranes) and biochemistry (detection of hydroxypyruvate reductase and α-ketoglutarate dehydrogenase activity as key enzymes of the type II methanotrophic bacteria) of strain WI 14 (19).
In methanotrophic bacteria, the oxidation of methane is always catalyzed by an MMO, which requires two reduction equivalents to split the O—O bond. As mentioned above, the enzyme exists in both particulate (pMMO) and soluble (sMMO) forms. Whereas all strains have the pMMO, only a few are able to express the sMMO (1, 33). The Methylocystis sp. strain WI 14 described in this study expresses the sMMO only if grown under copper limitation (0.1 μM causes a loss of about 60% of sMMO activity) and with ammonia or nitrate as the nitrogen source. If serine or cysteine is used instead, the strain grows but does not form the sMMO even though copper is absent. The influence of copper ions on sMMO expression is well known. It could be demonstrated that in M. trichosporium OB3b, the sMMO is completely repressed at copper concentrations of about 0.4 μM (22). It has been suggested that the sMMO-encoding genes localized in an operon are controlled by a regulatory copper-binding protein which interacts with DNA in the promoter region. Copper changes the affinity of the protein to DNA and represses sMMO expression (36, 37). However, the mechanism by which nitrogen regulates expression has not yet been described. Although it is known that strains such as Methylocystis sp. strain GB 25, which are not able to form the sMMO, cannot grow on nitrate as the nitrogen source (18), the effect of nitrogen on sMMO and pMMO expression has not been reported previously.
The sMMO investigated shows a very narrow substrate specificity and oxidizes some aliphatics, aromatics, and halogenated compounds only to their corresponding hydroxylated products. Interestingly, the oxidation of aliphatics always led to the n-alcohol, not the 1-alcohol. In contrast, Green and Dalton (16) showed that the sMMO of Methylococcus capsulatus (Bath) partially oxidizes pentane and hexane to 1-pentanol and 1-hexanol, respectively. Furthermore, in this study, the halogenated hydrocarbons were hydroxylated but not dehalogenated. For the sMMO of M. trichosporium OB3b, Sullivan and Chase (44) obtained similar results. 1,2,3-Trichlorobenzene was oxidized to 2,3,4- and 3,4,5-trichlorophenol.
Similar to nearly all sMMO so far isolated (except for the enzyme from Methylobacterium sp. strain CRL-26 [38]), the sMMO from Methylocystis sp. strain WI 14 has three components: a hydroxylase, a reductase, and a regulatory protein. The molecular properties of these components and the nucleotide sequences of the encoding genes possess particularly strong similarity to the sMMO from M. trichosporium OB3b and Methylocystis sp. strain M (Table 5). Reconstitution of the purified components showed that component B is not necessary for enzyme activity but that its presence results in a 10-fold increase in activity. For the sMMO from M. trichosporium and Methylocystis sp. strain M, it has been demonstrated that component B increases activity 10- and 11-fold, respectively (34), and that in the last strain it could be replaced by Fe2+ ions. In contrast, in the investigated strain, Fe2+ ions did not affect the hydroxylase and were not able to replace component B but were essential for maintenance and recovery of reductase activity after storage.
sMMO from Methylocystis sp. strain WI 14 is strongly inhibited by Cu2+, Zn2+, and Ni2+ ions. Although the inhibition of Zn2+ and Ni2+ ions is due mainly to the influence on the reductase, Cu2+ ions also seem to affect the other components. However, Hg2+ ions lead to a total loss of enzyme activity, suggesting that there are essential sulfhydryl groups involved in substrate hydroxylation and coenzyme oxidation, respectively. Green et al. (17) showed that Cu2+ and Zn2+ ions inhibit the reductase component of the sMMO from Methylococcus capsulatus (Bath), thereby inhibiting sMMO activity. On the other hand, Ni2+ ions had no effect on the reductase or on sMMO activity as a whole. In contrast, Jahng and Wood (22) reported that both Cu2+, Zn2+, and Ni2+ ions influenced sMMO activity.
ACKNOWLEDGMENTS
We thank J. Bär (Institut für Biochemie, Universitätsklinikum, Universität Leipzig) for the determination of the N-terminal amino acid sequence of the proteins.
This work was supported by the Umweltforschungszentrum, Leipzig-Halle GmbH (grant UFZ-24196).
REFERENCES
- 1.Bowman J P, Jimenez L, Rosario I, Hazen T C, Sayler G S. Characterization of the methanotrophic bacterial community present in a trichloroethylene-contaminated groundwater site. Appl Environ Microbiol. 1993;59:2380–2387. doi: 10.1128/aem.59.8.2380-2387.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bradford M M. A rapid sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- 3.Burrows K J, Cornish A, Scott D, Higgins I J. Substrate specificities of the soluble and particulate methane mono-oxygenases of Methylosinus trichosporium OB3b. J Gen Microbiol. 1984;5:335–342. [Google Scholar]
- 4.Cardy D L N, Laidler V, Salmond G P C, Murrell J C. Molecular analysis of the methane monooxygenase (MMO) gene cluster of Methylosinus trichosporium OB3b. Mol Microbiol. 1991;5:335–342. doi: 10.1111/j.1365-2958.1991.tb02114.x. [DOI] [PubMed] [Google Scholar]
- 5.Colby J, Dalton H. Characterization of the second prosthetic group of the flavoenzyme NADH-acceptor reductase (component C) of the methane monooxygenase from Methylococcus capsulatus (Bath) Biochem J. 1979;177:903–908. doi: 10.1042/bj1770903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Colby J, Stirling D I, Dalton H. The soluble methane mono-oxygenase of Methylococcus capsulatus Bath: its ability to oxygenate n-alkanes, n-alkenes, ethers, and alicyclic, aromatic and heterocyclic compounds. Biochem J. 1977;165:395–402. doi: 10.1042/bj1650395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Davis B J. Disc electrophoresis. II. Method and application to human serum proteins. Ann N Y Acad Sci. 1964;121:404–427. doi: 10.1111/j.1749-6632.1964.tb14213.x. [DOI] [PubMed] [Google Scholar]
- 8.Dorsch M, Stackebrandt E. Some modifications in the procedure of direct sequencing of PCR amplified 16S rDNA. J Microbiol Methods. 1992;16:271–279. [Google Scholar]
- 9.Edman P, Henschen A. Sequence determination. In: Needleman S B, editor. Protein sequence determination. A source book of methods and techniques. 2nd ed. Berlin, Germany: Springer-Verlag KG; 1975. pp. 232–279. [Google Scholar]
- 10.Felsenstein J. Phylogenies from molecular sequences: interference and reliability. Annu Rev Genet. 1988;22:521–565. doi: 10.1146/annurev.ge.22.120188.002513. [DOI] [PubMed] [Google Scholar]
- 11.Fox B G, Froland W A, Dege J E, Lipscomb D. Methane monooxygenase from Methylosinus trichosporium OB3b: purification and properties of a three-component system with high specific activity from a type II methanotroph. J Biol Chem. 1989;264:10023–10033. [PubMed] [Google Scholar]
- 12.Fox B G, Froland W A, Jollie D R, Lipscomb J D. Methane monooxygenase from Methylosinus trichosporium OB3b. Methods Enzymol. 1990;188:191–202. doi: 10.1016/0076-6879(90)88033-7. [DOI] [PubMed] [Google Scholar]
- 13.Fox G E, Pechman K R, Woese C R. Comparative cataloguing of 16S ribosomal ribonucleic acid: a molecular approach to procaryotic systematics. Int J Syst Bacteriol. 1977;27:44–57. [Google Scholar]
- 14.Fox G E, Stackebrandt E. The application of 16S rRNA cataloguing and 5S rRNA sequencing in bacterial systematics. Methods Microbiol. 1987;19:405–458. [Google Scholar]
- 15.Green J, Dalton H. Protein B of soluble methane monooxygenase from Methylococcus capsulatus (Bath) J Biol Chem. 1985;260:15795–15801. [PubMed] [Google Scholar]
- 16.Green J, Dalton H. Substrate specificity of soluble methane monooxygenase: mechanistic implications. J Biol Chem. 1989;264:17698–17703. [PubMed] [Google Scholar]
- 17.Green J, Prior S D, Dalton H. Copper ions as inhibitors of protein C of soluble methane monooxygenase of Methylococcus capsulatus (Bath) Eur J Biochem. 1985;153:137–144. doi: 10.1111/j.1432-1033.1985.tb09279.x. [DOI] [PubMed] [Google Scholar]
- 18.Grosse S. Methanabbau in Methylocystis sp. GB 25. Master dissertation. Leipzig, Germany: Universität Leipzig; 1994. [Google Scholar]
- 19.Grosse S. Methanmonooxygenase—Charakterisierung aus Typ II-Methanotrophen. Ph.D. thesis. Leipzig, Germany: Universität Leipzig; 1998. [Google Scholar]
- 20.Grosse S, Voigt C, Wendlandt K-D, Kleber H-P. Purification and properties of methanol dehydrogenase from Methylosinus sp. WI 14. J Basic Microbiol. 1998;38:189–196. doi: 10.1002/(sici)1521-4028(199807)38:3<189::aid-jobm189>3.0.co;2-s. [DOI] [PubMed] [Google Scholar]
- 21.Hanson R S, Hanson T E. Methanotrophic bacteria. Microbiol Rev. 1996;60:439–471. doi: 10.1128/mr.60.2.439-471.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jahng D, Wood T K. Metal ions and chloamphenicol inhibition of soluble methane monooxygenase from Methylosinus trichosporium OB3b. Appl Microbiol Biotechnol. 1996;45:744–749. [Google Scholar]
- 23.Jiang Y, Wilkins P C, Dalton H. Activation of the hydroxylase of sMMO from Methylococcus capsulatus (Bath) by hydrogen peroxide. Biochim Biophys Acta. 1993;1163:105–112. doi: 10.1016/0167-4838(93)90285-y. [DOI] [PubMed] [Google Scholar]
- 24.Koh S-C, Bowman J P, Sayler G S. Soluble methane monooxygenase production and trichloroethylene degradation by a type I methanotroph Methylomonas methanica 68-1. Appl Environ Microbiol. 1993;59:960–967. doi: 10.1128/aem.59.4.960-967.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Laemmli U K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 26.Lane D J. 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, editors. Nucleic acid techniques in bacterial systematics. J. Chichester. United Kingdom: Wiley & Sons, Ltd.; 1991. pp. 115–175. [Google Scholar]
- 27.Liu Y, Nesheim J C, Paulsen K E, Stankovich M T, Lipscomb J D. Roles of methane monooxygenase reductase component in the regulation of catalysis. Biochemistry. 1997;36:5223–5233. doi: 10.1021/bi962743w. [DOI] [PubMed] [Google Scholar]
- 28.Marcinka K, Röhring C, Kluge S. Changes in protein patterns of pea plants systemically infected with red clover mottle virus. Biochem Physiol Pflanz. 1992;188:187–193. [Google Scholar]
- 29.Martin-Kearley J, Gow J A, Peloquin M, Greer C W. Numerical analysis and the application of random amplified DNA polymerase chain reaction to the differentiation of Vibrio strains from a seasonally cold ocean. Can J Microbiol. 1994;40:446–455. doi: 10.1139/m94-073. [DOI] [PubMed] [Google Scholar]
- 30.McDonald I R, Kenna E M, Murrell J C. Detection of methanotrophic bacteria in environmental samples with the PCR. Appl Environ Microbiol. 1995;61:116–121. doi: 10.1128/aem.61.1.116-121.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.McDonald I R, Uchiyma H, Kambe S, Yagi O, Murrell J C. The soluble methane monooxygenase gene cluster of the trichloroethylene-degrading methanotroph Methylocystis sp. strain M. Appl Environ Microbiol. 1997;63:1898–1904. doi: 10.1128/aem.63.5.1898-1904.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Miguez C B, Bourque D, Sealy J A, Greer C W, Groleau D. Detection and isolation of methanotrophic bacteria possessing soluble methane monooxygenase (sMMO) genes using the polymerase chain reaction (PCR) Microb Ecol. 1997;33:21–31. doi: 10.1007/s002489900004. [DOI] [PubMed] [Google Scholar]
- 33.Murrell C J. Genetics and molecular biology of methanotrophs. FEMS Microbiol Rev. 1992;88:233–248. doi: 10.1111/j.1574-6968.1992.tb04990.x. [DOI] [PubMed] [Google Scholar]
- 34.Nakajima T, Uchiyama H, Yagi O, Nakahara T. Purification and properties of a soluble methane monooxygenase from Methylocystis sp. M. Biosci Biotechnol Biochem. 1992;56:736–740. doi: 10.1271/bbb.56.736. [DOI] [PubMed] [Google Scholar]
- 35.Nguyen H-H T, Elliott S J, Yip J H-K, Chan S I. The particulate methane monooxygenase from Methylococcus capsulatus (Bath) is a novel copper-containing three-subunit enzyme: isolation and characterization. J Biol Chem. 1998;273:7957–7966. doi: 10.1074/jbc.273.14.7957. [DOI] [PubMed] [Google Scholar]
- 36.Nielsen A K, Gerdes K, Degn H, Murrell J C. Regulation of bacterial methane oxidation: transcription of the soluble methane monooxygenase operon of Methylococcus capsulatus (Bath) is repressed by copper ions. Microbiology. 1996;142:1289–1296. doi: 10.1099/13500872-142-5-1289. [DOI] [PubMed] [Google Scholar]
- 37.Nielsen A K, Gerdes K, Murrell J C. Copper-dependent reciprocal transcriptional regulation of methane monooxygenase genes in Methylococcus capsulatus and Methylosinus trichosporium. Mol Microbiol. 1997;25:399–409. doi: 10.1046/j.1365-2958.1997.4801846.x. [DOI] [PubMed] [Google Scholar]
- 38.Patel R N, Savas J C. Purification and properties of the hydroxylase component of methane monooxygenase. J Bacteriol. 1987;169:2313–2317. doi: 10.1128/jb.169.5.2313-2317.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Pilkington S J, Dalton H. Purification and characterisation of the soluble methane monooxygenase from Methylosinus sporium 5 demonstrates the highly conserved nature of this enzyme in methanotrophs. FEMS Microbiol Lett. 1991;78:103–108. [Google Scholar]
- 40.Sambrook J, Fritsch E F, Maniatis T. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, N.Y: Cold Spring Harbor Laboratory Press; 1989. pp. 18.47–18.59. [Google Scholar]
- 41.Shen R N, Yu C L, Ma Q Q, Li S B. Direct evidence for a soluble methane monooxygenase from type I methanotrophic bacteria: purification and properties of a soluble methane monooxygenase from Methylomonas sp. GYJ3. Arch Biochem Biophys. 1997;345:223–229. doi: 10.1006/abbi.1997.0239. [DOI] [PubMed] [Google Scholar]
- 42.Stackebrandt E, Charfreytag O. Partial 16S rRNA primary structure of five Actinomyces species: phylogenetic implications and development of an Actinomyces israelii-specific oligonucleotide probe. J Gen Microbiol. 1990;136:37–43. doi: 10.1099/00221287-136-1-37. [DOI] [PubMed] [Google Scholar]
- 43.Stirling D I, Dalton H. Properties of the methane monooxygenase from extracts of Methylosinus trichosporium OB3b and evidence for its similarity to the enzyme from Methylococcus capsulatus (Bath) Eur J Biochem. 1979;96:205–221. doi: 10.1111/j.1432-1033.1979.tb13030.x. [DOI] [PubMed] [Google Scholar]
- 44.Sullivan J P, Chase H A. 1,2,3-Trichlorobenzene transformation by Methylosinus trichosporium OB3b expressing soluble methane monooxygenase. Appl Microbiol Biotechnol. 1996;45:427–433. [Google Scholar]
- 45.Tsien H-C, Hanson R S. Soluble methane monooxygenase component B gene probe for identification of methanotrophs that rapidly degrade trichloroethylene. Appl Environ Microbiol. 1992;58:953–960. doi: 10.1128/aem.58.3.953-960.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]