Summary
Circulating tumor DNA (ctDNA) carries tumor-specific genetic and epigenetic variations. To identify extranodal natural killer/T cell lymphoma (ENKTL)-specific methylation markers and establish a diagnostic and prognosis prediction model for ENKTL, we describe the ENKTL-specific ctDNA methylation patterns by analyzing the methylation profiles of ENKTL plasma samples. We construct a diagnostic prediction model based on ctDNA methylation markers with both high specificity and sensitivity and close relevance to tumor staging and therapeutic response. Subsequently, we built a prognostic prediction model showing excellent performance, and its predictive accuracy is significantly better than the Ann Arbor staging and prognostic index of natural killer lymphoma (PINK) risk system. Notably, we further establish a PINK-C risk grading system to select individualized treatment for patients with different prognostic risks. In conclusion, these results suggest that ctDNA methylation markers are of great value in diagnosis, monitoring, and prognosis, which might have implications for clinical decision-making of patients with ENKTL.
Keywords: circulating tumor DNA methylation, extranodal natural killer/T cell lymphoma, diagnosis, prognosis, treatment response, surveillance, PINK-C risk grading system
Graphical abstract

Highlights
-
•
ctDNA methylation shows great value in diagnosis, prognosis prediction of ENKTL
-
•
ctDNA methylation can be used as a biomarker to monitor disease progression
-
•
PINK-C risk assessment system could stratify patients for prognostic determinations
-
•
PINK-C risk assessment system might have implications for clinical decision-making
Tian et al. identify and validate a series of ctDNA methylation markers, which can assist in the accurate diagnosis, disease monitoring, and prognosis prediction of ENKTL and may provide therapeutic optimization for early-stage patients.
Introduction
Extranodal natural killer/T cell lymphoma (ENKTL) is a rare subtype of non-Hodgkin’s lymphoma that mainly involves the nasal cavity and nasopharynx.1,2,3 Despite the best currently available therapeutic regimens, approximately half of patients with ENKTL develop progressive disease (PD), and the clinical outcomes remain dismal for patients with advanced ENKTL with a 5-year survival of 50%.4,5
Accurate diagnosis and staging and prompt treatment of ENKTL are critical for an improved clinical outcome.6,7,8 Currently, ENKTL diagnosis mainly depends on tumor biopsy, which, however, is hampered by a low proportion of tumor cells in the biopsy tissues and the presence of necrotic tissues and is further rendered difficult by the abundant vasculature and anatomic complexity of the nasal cavity and nasopharynx.9 It also remains a challenge to discern ENKTL via biopsy from other diseases of the nasopharynx, such as nasopharyngeal carcinoma (NPC) or nasopharyngitis (NPG), resulting in missed or erroneous diagnosis. In addition, almost 70% of patients with ENKTL are classified as early stage per the Ann Arbor staging system. Diverse clinical outcomes are shown in this group of patients, indicating that the Ann Arbor staging system is not efficient enough to perform risk-stratified individual treatment.10 Therefore, it is critical for the management of ENKTL to identify effective diagnostic and prognostics biomarkers of ENKTL.
Circulating tumor DNA (ctDNA) is an extracellular nucleic acid fragment from necrotic and apoptotic tumor cells or is actively released by tumor cells into the peripheral blood.11 It renders feasible noninvasive molecular testing of malignancies using plasma samples, the so-called “liquid biopsy.”12,13 Such an approach also allows for dynamic monitoring of molecular changes in tumors as plasma samples can be acquired at ease at any time during therapy. Aberrant DNA methylation (epigenetic) patterns, especially hypermethylation of tumor suppressor genes, occurs in the early stage of tumor development and can be detected before a tumor is diagnosed in many cancers, such as lung cancer, colorectal cancer, etc.14,15,16,17 ctDNA-bearing, cancer-specific methylation patterns have been studied as candidate biomarkers in liver cancer, colorectal cancer, and primary central nervous system lymphomas (PCNSLs).18,19,20,21 However, ctDNA methylation patterns have not been elucidated in ENKTL.
This study aimed to evaluate the potential value of ctDNA methylation markers in the diagnosis, dynamic monitoring, and prognosis prediction of ENKTL. We applied multiple statistical methods to develop and validate the diagnostic and prognostic prediction model with selected ENKTL-specific ctDNA methylation biomarkers. Moreover, a ctDNA-methylation-based prognostic index of natural killer lymphoma classification (PINK-C) staging scheme was constructed to predict the prognosis of an individual and develop risk-adapted treatment approaches.
Results
ENKTL ctDNA exhibits distinct methylation patterns
The study flowchart is shown in Figure 1. Our sequencing analysis of ctDNA pools of the discovery cohort revealed 75,420 differentially methylated regions (cutoff value of a methylation difference larger than 10% and p value less than 0.05) and 304,843 differentially methylated loci between patients with ENKTL and healthy subjects (Figure 2). As pooled ctDNA may not unravel the actual methylation status of individual samples, we selected the top 1,000 differential methylated loci by fold change and the corresponding p value, dissected these markers in individual samples of the ctDNA EPIC cohort, and identified 982 differential methylation markers. In total, 98 of the ctDNA differential methylation markers were further detected in 61 ENKTL tissue DNA samples and 106 healthy human NK/T cell DNA samples. Pearson correlation analysis demonstrated a strong positive correlation in DNA differential methylation at the 98 differential methylation markers between ctDNA from ENKTL plasma samples and healthy plasma samples and DNA from 61 ENKTL tissue DNA samples and 106 healthy human NK/T cell DNA samples (R = 0.87, p < 0.001) (Figure S1A), indicating that ctDNA differential methylation markers could represent methylation status in ENKTL tissues.
Figure 1.
The study flowchart
ENKTL, extranodal natural killer/T cell lymphoma; ctDNA, circulating tumor DNA methylation markers; EPIC, epigenetic; PINK-C, ctDNA-methylation-based prognostic index of natural killer lymphoma.
Figure 2.
Differential ctDNA methylation patterns between extranodal natural killer T cell lymphoma (ENKTL) and healthy pooled samples
(A) Differentially methylated locations between ENKTL and healthy pooled samples in the chromosomes.
(B) Differentially methylated regions between ENKTL and healthy pooled samples. Differentially methylated CpGs between healthy and tumor plasma samples were identified using DMRfinder v.0.3 with the beta-binomial hierarchical modeling and Wald test for pooled samples.
Diagnostic performance of a 7-methylation marker signature for ENKTL
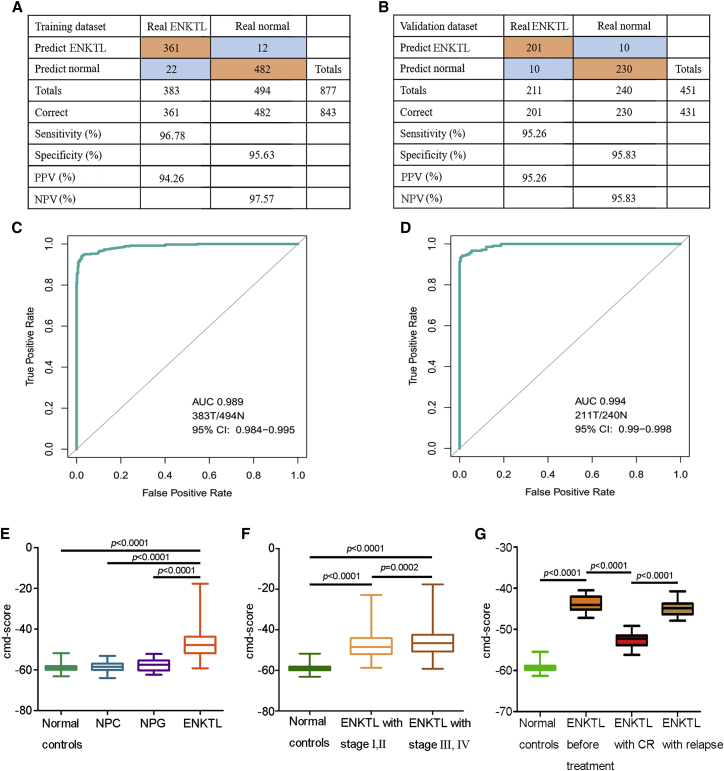
In the training dataset (Table S1), variables were screened using LASSO logistic regression based on the performance of the 98 differentially methylated ctDNA markers. Seven markers were selected to build the ctDNA methylation diagnosis (cmd) score (Table 1); the diagnostic prediction model yielded a sensitivity of 94.26%, a specificity of 97.57%, a positive prediction value (PPV) of 96.78%, and a negative prediction value (NPV) of 95.63%, with an area under the curve (AUC) of 0.989 for the training dataset. The prediction model was validated using the validation dataset, with a sensitivity and PPV of 95.3% and a specificity and NPV of 95.8%, and an AUC of 0.994 (Figures 3A–3D). The prediction model was also validated using an independent held-out cohort, with a sensitivity of 94.3%, a specificity of 94.6%, a PPV of 91.7%, and an NPV of 96.4%, with an AUC of 0.988 (Figures S1B and S1C). Unsupervised hierarchical clustering of the seven markers distinguished ENKTL from healthy controls with a high specificity and sensitivity (Figures S1D and S1E). In addition, each of the seven markers in the prediction model exhibited significant methylation difference between patients with ENKTL and healthy subjects (Figure S2A) and showed a nearly overlapping AUC curve in the training and validation dataset, indicating the absence of bias in both datasets (Figures S2B and S2C).
Table 1.
Characteristics of seven ctDNA methylation markers and their coefficients in ENKTL diagnosis
| Markers | Coefficient | SE | z value | p value | Ref. gene |
|---|---|---|---|---|---|
| Chr1.221290973 | −17.183 | 2.777 | −6.187 | 6.13E−10 | HLX-AS1 |
| Chr8.80200243 | −15.772 | 2.583 | −6.105 | 1.03E−09 | MIR12123 |
| Chr10.125847713 | −13.633 | 2.815 | −4.844 | 1.28E−06 | CHST15 |
| Chr14.101213547 | 5.097 | 1.333 | 3.842 | 0.000131 | DLK1 |
| Chr5.102841867 | −10.91 | 2.909 | −3.751 | 0.000176 | LINC02115 |
| Chr11.36002757 | −8.228 | 1.893 | −4.347 | 1.38E−05 | MIR3973 |
| Chr21.22572637 | −5.152 | 1.512 | −3.407 | 0.000658 | NCAM2 |
ctDNA, circulating tumor DNA; ENKTL, extranodal natural killer/T cell lymphoma; SE, standard errors of coefficients; z value, Wald z-statistic value.
Figure 3.
ctDNA methylation analysis for ENKTL diagnosis in individual samples
(A and B) Confusion tables of binary results of the diagnostic prediction model in the training (A) and validation (B) datasets. Sensitivity = predict ENKTL/total ENKTL × 100%; specificity = predict normal/total normal × 100%; PPV = predict ENKTL/(predict ENKTL + false ENKTL) × 100%; NPV = predict normal/(predict normal + false normal) × 100%.
(C and D) ROC curves of the diagnostic prediction model with ctDNA methylation markers in the training (C) and validation (D) datasets. The area under receiving operator characteristic (ROC) curve, which estimated the proportion of concordant pairs among all actual pairs of observations, with 1 representing perfect prediction, was used to evaluate the predictability of the model.
(E) cmd scores in healthy controls, individuals with NPC and NPG, and patients with ENKTL. The cmd score was calculated by a prognostic model according to individual coefficient listed in Table 1. p value was calculated by t test.
(F) cmd scores in healthy controls and individuals with early stages (Ann Arbor stage I/II) and advanced stages (Ann Arbor stage III/Ⅳ). The cmd score was calculated by a prognostic model according to individual coefficient listed in Table 1. p value was calculated by t test.
(G) cmd score in normal controls and patients with ENKTL before treatment, with CR, and with relapse. The cmd score was calculated by prognostic model according to individual coefficient listed in Table 1. p value was calculated by t test. cmd score, ctDNA methylation diagnosis score; NPC, nasopharyngeal carcinoma; NPG, nasopharyngitis; ENKTL, extranodal natural killer/T cell lymphoma; CR, complete remission; PPV, positive predictive value; NPV, negative predictive value.
Data in boxplot are presented as the median, minimum, maximum, 25th percentile, and 75th percentile.
Furthermore, patients with ENKTL had a significantly higher cmd score than patients with NPC or NPG, two major diagnostic confounders, and healthy subjects (Figure 3E; Table S2). Patients with Ann Arbor stage III or IV ENKTL also had a remarkably higher cmd score than those with Ann Arbor stage I or II ENKTL (Figure 3F). A higher cmd score significantly correlated with elevated lactate dehydrogenase (LDH), the presence of Epstein-Barr virus (EBV)-DNA, number of extranodal involvement ≥2, and distant lymph nodes (all p < 0.05; Figure S3A). In addition, patients who had achieved complete remission had a significantly lower cmd score than patients who had experienced relapse (p < 0.001) (Figure 3G). The probabilities of logistic regression model converted from cmd score among different groups aforementioned are presented in Figures S3B–S3D. These results suggest that the cmd score could be of value in monitoring disease progression and treatment response.
ctDNA prognostic prediction for ENKTL
The characteristics of seven ctDNA methylation markers and their coefficients in ENKTL prognosis prediction are shown in Table 2. By using LASSO-Cox methods, we constructed the 7-marker prognostic ctDNA methylation prognosis (cmp) score for predicting the prognosis of patients with ENKTL (Table 2; Figures S4A and S4B). In the training dataset, at an optimal cmp cutoff of −23.1 (Figure S4C), a cmp score > −23.1 had a significantly higher risk of death (hazard ratio [HR] 5.007, 95% confidence interval [CI] 3.550–7.060; p < 0.001) and progression-free survival (PFS; HR 3.906, 95% CI 2.875–5.308; p < 0.001), and progression versus a cmp score ≤ −23.1 (HR 3.122, 95% CI 2.107–4.625; p < 0.001) (Figures 4A, 4B, S4D, and S4E). By using the same cutoff value in the training dataset, similar findings were observed in the validation dataset.
Table 2.
Characteristics of seven ctDNA methylation markers and their coefficients in ENKTL prognosis prediction
| Markers | Coefficient | SD | z value | p value | Ref. gene |
|---|---|---|---|---|---|
| Chr1.17525460 | −4.131 | 0.777 | −5.316 | 1.06E−07 | PADI1 |
| Chr2.71858769 | −12.53 | 1.148 | −10.910 | <2E−16 | DYSF |
| Chr2.71858749 | −0.774 | 0.952 | −0.813 | 0.416 | DYSF |
| Chr5.82664317 | −2.937 | 0.600 | −0.485 | 9.85E−07 | VCAN |
| Chr10.125847713 | −8.841 | 0.873 | −10.121 | <2E−16 | CHST15 |
| Chr20.56540714 | −1.132 | 0.739 | −1.533 | 0.125 | LINC01742 |
| Chr22.43702154 | −0.116 | 0.695 | −0.166 | 0.868 | LOC101927447 |
ctDNA, circulating tumor DNA; ENKTL, extranodal natural killer/T cell lymphoma; SE, standard errors of coefficients; z value, Wald z-statistic value.
Figure 4.
ctDNA methylation analysis for ENKTL prognostic prediction
(A and B) Overall survival curves of patients with ENKTL with low or high risk, according to the ctDNA methylation prognosis score (cmp score) in the training (A) and validation (B) datasets. Survival analysis was performed by Kaplan-Meier method and log-rank test.
(C and D) The ROC curves of the cmp score, Ann Arbor stage, PINK, and cmp score combined with PINK in the training (C) and validation (D) datasets. The area under receiving operator characteristic (ROC) curve, which estimated the proportion of concordant pairs among all actual pairs of observations, with 1 representing perfect prediction, was used to evaluate the predictability of the model. PINK, prognostic index of natural killer lymphoma.
Univariate and multivariate Cox regression analyses revealed that after adjustment, the cmp score remained a significant and independent predictor of both overall survival (OS) and PFS (Tables S3 and S4). Receiver operating characteristic (ROC) analysis showed a significantly better performance of cmp score than the Ann Arbor staging system and the PINK risk system (log-rank test, all p < 0.05; Figures 4C, 4D, S5A, and S5B), which was further enhanced with the addition of the PINK staging system for both the training dataset (AUC 0.794; Figure 4C) and the validation dataset (AUC 0.773; Figure 4D).
Performance of a PINK-C risk stratification system
Using the data from 383 patients with patients with ENKTL in the training dataset, we constructed PINK-C by integrating the cmp score and the PINK system for quantitative prediction of the survival outcomes of individual patients (Figure 5A). Calibration plots showed good agreement between the predicted and observed 3-year OS and 3-year PFS in the two datasets (Figure S5C). The patients were categorized using total PINK-C scores by maximization of prognostic difference (low risk: 0–33; intermediate risk: 66–67; high risk: 99–1166) (Figure S5D and S5E); each subgroup demonstrated a distinct prognosis (log-rank test, p < 0.001; Figures 5B and S6A).
Figure 5.
Establishing PINK-C scoring system to predict OS for patients with ENKTL
(A) PINK-C, comprised of PINK and cmp score status, was constructed to predict the 1-, 3-, and 5-year OSs. Coefficients of the prognostic covariates in the multivariate Cox regression model were used to construct a PINK-C risk system using the “rms” package in R software.
(B) Subgroups of patients with different PINK-C scores (low risk: 0–33; intermediate risk: 66–73; high risk: ≥99) showed distinct OS in the total dataset.
(C and D) OS (C) and progression-free survival (PFS) (D) of intermediate and high-risk Ann Arbor stage I patients receiving RT versus CMT. Survival analysis was performed by Kaplan-Meier method and log-rank test. CMT, combined modality treatment; RT, radiotherapy.
We were interested in whether PINK -C could effectively stratify patients for treatment. Notably, patients with Ann Arbor stage I and intermediate and high PINK-C risk who received combined modality treatment (CMT) had a significantly longer median OS (HR 0.531, 96% CI 0.335–0.843; p = 0.007) and PFS (HR 0.603, 96% CI 0.387–0.940; p = 0.026) than those who received radiotherapy (RT) alone (Figures 5C and 5D). Meanwhile, patients with Ann Arbor stage I and low PINK-C risk who received RT alone had comparable OS (HR 0.721, 96% CI 0.324–1.605; p = 0.423) and PFS (HR 0.719, 96% CI 0.387–1.337; p = 0.298) to their counterparts who received CMT (Figures S6C and S6D). These results indicate that RT could be sufficient for patients with low PINK-C risk, while CMT can be added for intermediate and high PINK-C risk Ann Arbor stage I patients.
Discussion
In this study, we constructed and validated a diagnostic prediction model using a 7-ctDNA methylation marker panel (cmd score) to discriminate ENKTL from NPC or NPG and normal conditions. The cmd score is closely associated with tumor stage, disease progression, and treatment response and could be of good application value for evaluating treatment response and monitoring relapse. Additionally, we constructed a prognosis prediction score (cmp score) with a 7-marker panel, which accurately predicts the clinical outcomes of patients with ENKTL. Our PINK-C risk assessment system based on the cmp score and the PINK system could stratify patients with ENKTL for treatment and prognostic determinations.
ctDNA provides a ready source for the detection and diagnosis of tumors.22 Monitoring somatic changes in ctDNA allows predictive assessment of treatment response of certain solid tumors.20,21,22 One advantage of methylation markers is obviating the need for detecting somatic mutations, as shown in the current study. In addition, we carried out targeted sequencing of specific markers, which can avoid the high cost of deep sequencing and offers greater ease of clinical application. In addition, ctDNA methylation profiling has some advantages over ctDNA mutation analysis for cancer detection, such as higher sensitivity, dynamic range, detectable consist, and multiple altered CpG sites within targeted genomic regions. Although only a fraction of mutations may be detected in ctDNA, it is still worth investigating whether the combination of ctDNA methylation patterns with somatic mutation analysis could provide a “pan-cancer” panel for clinical applications.
ENKTL diagnosis, which mainly relies on tumor biopsy pathologies, is relatively difficult due to the peculiarity of an ENKTL site and heterogeneity of biopsy tissues.2 Approximately 70% of patients with ENKTL were initially diagnosed with sinusitis, nasal polyps, upper respiratory tract infection, etc.23 The misdiagnoses may lead to decreased OS rates, with 1-, 3-, and 5-year OSs of 73.6%, 59.3%, and 43.2% in the misdiagnosed group, respectively, and 88.2%, 82.4%, and 64.2% in the nonmisdiagnosed group, respectively.23 According to NCCN guidelines, standard pathological assessment of ENKTL with immunohistochemistry costs at least $600 USD, while ctDNA methylation tests cost less than $20 USD. In this study, we used ctDNA methylation biomarkers, called the cmd score, to aid in ENKTL diagnosis. The sensitivity and specificity of cmp is considerably high for ENKTL. The cmp score showed good diagnostic performance and represents a cost-effective approach. By simply detecting the cmp score using plasma samples, clinicians can distinguish ENKTL from NPC and NPG. When combined with pathology, ENKTL can be diagnosed quickly and accurately.
To evaluate the generalizability and specificity of the identified ctDNA methylation markers, we assessed the performance of the CpG markers identified in our study with a public dataset. Five of the 14 CpG markers in our diagnostic or prognostic classifier were covered by the 850K Illumina BeadChip in the GEO: GSE169643 dataset. The results are presented in Figure S7. Among 5 of the 14 CpG markers in the GOE: GSE169643 dataset, no methylation ratio difference was detected in the chr1.17525460 and chr1.221290973 sites between the ENKTL ctDNA dataset in our study and tonsils, blood, and bone marrow in the GEO: GSE169643 dataset, while the methylation ratio of chr5.82664317, chr14.101213547, and chr20.56540714 sites is statistically different. The methylation ratio of the 5 sites was similar between the ENKTL ctDNA dataset and NKLGL leukemia samples in the GEO: GSE169643 dataset. Since there are only 3 NKLGL samples, it is not certain that whether the cmp score can distinguish ENKTL from NKLGL. Notably, all the 5 CpG sites showed significant discrepancy between ENKTL ctDNA dataset in our study and the formalin-fixed, paraffin-embedded (FFPE) samples in the GEO: GSE169643 dataset. This indicated that the methylation sites and ratios in tumor samples might not represent that in cell-free DNA. Furthermore, we also assessed the performance of the 3 of the 98 differentially methylated CpG sites covered by the 450K Illumina BeadChip in the 32 tumor types of TCGA dataset (Figure S8). No statistical difference of methylation ratio was observed between the ENKTL ctDNA dataset and most of the 32 tumor types. Considering that our classifiers were constructed based on a specific formula, integration of multiple methylation sites, and usage of an optimal cut-off value, it is reasonable that different tumor types could not be distinguished by a single methylated CpG site. In our study, the cmd score was sufficient to differentiate ENKTL and NPC.
Patients with ENKTL are mainly evaluated radiologically (computed tomography [CT], MRI, and positron emission tomography [PET]-CT). Despite the use of imaging evaluation criteria (Cheson criteria and others), remission or relapse is not depicted at the molecular level, and minimal residual disease cannot be accurately determined. Besides, these imaging modalities are expensive and cumbersome to perform, leading to inconsistent monitoring.24 Circulating EBV-DNA in blood is a well-known biomarker of tumor burden that is also associated with survival and treatment outcomes in patients with ENKTL.25,26 In this study, our cmd score highly correlates with tumor stage and treatment response and could be used as a tool for monitoring treatment response and occurrences of relapse. Moreover, the cmp score has ease of operation, convenient implementation, and high sensitivity, which is convenient for clinical use. Integration of EBV-DNA status into the cmd score could slightly improve the predictive performance of distinguishing ENKTL from NPC/NPG; however, no statistical difference was observed (Figures S6E and S6F). Therefore, it seems that both EBV-DNA and the ctDNA methylation signatures could distinguish ENKTL from NPG. However, only ctDNA methylation signatures could distinguishing ENKTL from NPC.
The PINK staging system has been recommended for evaluating the prognosis of patients with ENKTL by NCCN guidelines.7 However, the prognosis of patients with the same PINK risk also varies widely.10 In addition, the PINK risk stratification system could not personalize the prognosis of individual patients with ENKTL. In this study, based on the ENKTL-specific ctDNA methylation pattens, we established the cmp score consisting of seven biomarkers that can be used to predict the prognosis of ENKTL. The predictive value of the cmp score was much greater than that of Ann Arbor staging system and the PINK risk system. Our cmp score can complement the PINK risk system. The combinatorial PINK-C system incorporating the cmp score and the PINK system showed a better prognostic value than either scheme alone. Moreover, both physicians and patients could utilize the PINK-C score system for individual survival prediction.
The clinical utility of the cmd score and the cmp score needs to be further demonstrated in future clinical trials, and a prospective cohort study (ClinicalTrials.gov: NCT04423536) is still recruiting. For instance, if a patient has a cmd score > −56.4 in plasma ctDNA, which indicates a higher probability of diagnosis with ENKTL, then further examinations (such as nasopharyngeal endoscope, magnetic resonance scanning of nasopharynx, EBV-DNA) are recommended. Additionally, the therapeutic choice of RT alone or CMT for early-stage ENKTL has not been well defined.27,28,29,30 In this study, using the PINK-C, RT alone provided a comparable survival benefit to patients with a PINK-C score ≤33 in Ann Arbor stage I patients, whereas only patients with a PINK-C score >33 could benefit from CMT. In the design of ENKTL clinical trials, the PINK-C system can help physician design treatment plans. When the PINK-C score was >33 in Ann Arbor stage I patients, patients can be advised to use combined chemoradiotherapy to improve the survival rate; when the PINK-C score is ≤33, patients can be advised to use RT alone to reduce adverse effects with comparable efficacy.
It has been reported that some of the downstream genes of the selected ENKTL-specific ctDNA methylation biomarkers in cmd/cmp scores have known functions. Carbohydrate sulfotransferase 15 (CHST15) is the counterpart of sulfotransferase GalNac4S-6ST and has been identified as a gene co-expressed with the RAG1 gene in B cells.31,32 It is associated with tumor progression, metastasis, and maintenance of stem cell properties, and some studies indicate that the CHST15/ILKAP/CCND1 and CHST15/RABL6/PMAIP1 signaling axes may play a role in cell proliferation and apoptosis, but its molecular mechanisms are largely unexplored.33 Delta-like noncanonical Notch ligand 1 (DLK1) has been reported to be highly expressed in common malignancies. It plays an important role in maintaining the undifferentiated phenotype of cells leading to tumorigenesis, possibly through the NOTCH signaling pathway.34,35 Nerve cell adhesion molecule 2 (NCAM2) is a homologous analog of the nerve cell adhesion molecule NCAM1 (CD56).36 NCAM1 is an important marker for NK cells, mediating homophile binding with neighboring cells (e.g., neurons with astrocytes) and heterophile interactions between cells and extracellular matrix components.37 NCAM2 also plays an important role in modifying the cytoskeleton by interacting with proteins involved in cytoskeleton stability and regulating calcium influx.38 Protein arginine deiminase 1 (PADI1) is associated with tumorigenesis and epithelial-mesenchymal transition (EMT) in multiple cancer types by activating the ERK1/2-p38 signaling pathway.39,40 Dysferlin (Dysf) is mainly known as a membrane repair protein in muscle cells.41 Dysf is opposingly involved in tumor migration and invasion and survival.42 Versican (VCAN) was an EMT-related gene. Decreased VCAN expression suppressed cell migration and invasion capacity through transforming growth factor β (TGF-β)/cPML/Smad signaling; VCAN overexpression renders lymphoid cells more chemosensitive.43,44 The investigation of adhesion molecules, cytoskeleton stability, EMT, and cell proliferation and apoptosis involving NOTCH, TGF-β, and ERK signaling pathways could help understand the pathogenetic mechanism of ENKTL.
The limitations of our study are also worth noting. First of all, we included samples only from the Chinese population. Further investigation is needed as to whether the ctDNA methylation model is applicable to other ethnicities. Second, the biological mechanisms of the ctDNA methylation markers in ENKTL are still unknown and need to be further investigated. Third, this retrospective study could have some selection bias.
In conclusion, of the prediction model constructed with ctDNA methylation markers exhibit good diagnostic and prognostic performance and could be useful in stratifying patients with ENKTL for optimized treatment and dynamic monitoring of disease progression and treatment response in the patients, providing physicians with a much-needed tool in the care of patients with ENKTL.
Limitations of the study
In this study, we constructed a diagnostic prediction signature using a ctDNA methylation marker panel to discriminate ENKTL from NPC or NPG and healthy conditions. We did not verify whether the signature could be used to distinguish ENKTL from other lymphoma/malignances. Moreover, whether the discrimination power of the ctDNA methylation signature applies to other ethnicities is unclear. We also built a PINK-C risk assessment system based on a prognostic ctDNA methylation signature that could stratify patients with ENKTL for treatment and prognostic determinations. The clinical utility of PINK-C system is still need validated in prospective studies and clinical trials.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Biological samples | ||
| ENKTL patients’ and normal subjects blood samples | Sun Yat-Sen University Cancer Center (SYSUCC); Guangdong Provincial People’s Hospital (GPPH); Peking University Cancer Hospital & Institute (PUCH); Hunan Cancer Hospital (HNCH); Beijing Tongren Hospital (BJTH); Tianjin Medical University Cancer Institute and Hospital (TMUCIH); First people’s Hospital of Foshan, Tongji Hospital, Fudan University Shanghai Cancer Center, and Affiliated Cancer Hospital of Zhengzhou University. | N/A |
| ENKTL tissue DNA samples | Sun Yat-Sen University Cancer Center (SYSUCC) | N/A |
| Nasopharynx cancer (NPC) ctDNA samples | Sun Yat-Sen University Cancer Center (SYSUCC) | N/A |
| Nasopharyngitis (NPG) ctDNA samples | Sun Yat-Sen University Cancer Center (SYSUCC) | N/A |
| Normal human NK/T cell DNA samples | Guangzhou Women and Children Medical Center (GWCMC) | N/A |
| Chemicals, peptides, and recombinant proteins | ||
| Ampligase Enzyme and Buffer | Epicentre | A0102K |
| Exonuclease III | Thermo Fisher SCIENTIFIC | EN0191 |
| Exonuclease I | Thermo Fisher SCIENTIFIC | 70073X5000UN |
| dNTP | Thermo Fisher SCIENTIFIC | AB0196 |
| Phusion High-Fidelity DNA polymerase | Thermo Fisher SCIENTIFIC | F555S |
| KAPA HiFi HotStart Uracil + ReadyMix (2X) | Roche | KK2801 |
| Taq DNA polymerase | Thermo Fisher SCIENTIFIC | EP0404 |
| Probes | Integrated DNA Technologies | N/A |
| T4 polynucleotide kinase | New England Biolabs | M0201S |
| Critical commercial assays | ||
| EZ-96DNA methylation Gold kit | ZYMO RESEARCH | D5008 |
| ctDNA extraction kits | EliteHealth | N/A |
| AllPrep DNA/RNA Mini Kit (50) | QIAGEN | 80204 |
| TruSeq Methyl Capture EPIC Library Prep Kit | illumina | FC-151-1003 |
| CD3+CD56+NK/T Cell Isolation Kit | Miltenyi Biotec | Cat#130-093-064 |
| DNeasy Blood &Tissue Kit | QIAGEN | Cat#69506 |
| Deposited data | ||
| The China National GeneBank DataBase (CNGBdb) | This paper | CNP0003741 |
| Software and algorithms | ||
| R software | R Development Core Team | https://www.r-project.org/ |
| R studio | N/A | https://www.rstudio.com/ |
| X-tile software | Yale School Of Medicine45 | https://medicine.yale.edu/lab/rimm/research/software/ |
| BitmapperBS version 1.0.2.3 | M/A | http://home.ustc.edu.cn/∼chhy/BitMapper.html |
| Fastp version 0.20.0 | HaploX | N/A |
| Bedtools version 2.29.0 | N/A | https://github.com/arq5x/bedtools2/releases |
| Least Absolute Shrinkage and Selection Operator (LASSO) algorithm | Fu46 and Tibshirani47 | Glmnet package |
Resource availability
Lead contract
Further information and resource requests should be directly to and will be fulfilled by the lead contract, Qing-Qing Cai (caiqq@sysucc.org.cn).
Material availability
All unique/stable reagents generated in this study are available from the lead contract.
Experimental model and subject details
Patients and participants
The discovery cohort consisted of 24 ENKTL ctDNA pools of plasma samples from 480 ENKTL patients who received care at Sun Yat-Sen University Cancer Center (SYSUCC) between January 1, 2007 and December 30, 2018 and 60 normal ctDNA pools of 1200 normal plasma samples from SYSUCC (n = 250) and Sun Yat-Sen Memorial Hospital (SYSMH) (n = 950).
The retrospective ctDNA epigenetic (EPIC) cohort consisted of blood samples from 594 newly diagnosed pathologically-confirmed ENKTL patients and 734 healthy subjects from SYSUCC and Guangdong Provincial People’s Hospital (GPPH) between January 1, 2012 and December 30, 2018, from Peking University Cancer Hospital & Institute (PUCH), Hunan Cancer Hospital (HNCH), Beijing Tongren Hospital (BJTH), and Tianjin Medical University Cancer Institute and Hospital (TMUCIH) between January 1, 2012 and December 30, 2018. An independent held-out cohort consisted of blood samples from 35 ENKTL patients and 56 normal subjects from the First people’s Hospital of Foshan, Tongji Hospital, Fudan University Shanghai Cancer Center, and Affiliated Cancer Hospital of Zhengzhou University were further used to validate the diagnostic prediction power. The patients had complete clinical data and had received first-line non-anthracycline-based regimens with or without radiotherapy. These ctDNA methylation data were randomized in a 2:1 ratio into the training dataset (n = 877) and the validation dataset (n = 451) for construction of diagnostic and prognostic models.
In addition, 61 ENKTL tissue DNA samples, 105 nasopharynx cancer (NPC) ctDNA samples and 43 nasopharyngitis (NPG) ctDNA samples were obtained from SYSUCC, and 148 ctDNA samples from normal human NK/T cell DNA samples were obtained from Guangzhou Women and Children Medical Center (GWCMC). We also obtained another 75 ctDNA samples from ENKTL patients at times of pretreatment (n = 25), complete remission (CR) (n = 25) and relapse (n = 25) between January 1, 2019 and December 30, 2020 from SYSUCC (Figure S6B). Incisional or needle biopsy were used to obtain specimens at diagnosis, and were reassessed by two pathologists (HLR and ML) per the 2017 WHO classification criteria.3
The study protocol was approved by the ethics review committees of all participating institutions. Personal data were anonymized in the paper.
Assessment
We retrieved patient demographic and baseline data including Eastern Cooperative Group performance status (ECOG-PS) scores, Ann Arbor stage, extranodal involvement, lactate dehydrogenase (LDH), B symptoms, lymph node involvement, and non-nasal types. Nasal and non-nasal types were determined by physical and radiological (CT, MRI, and PET-CT) examination of upper aerodigestive tract (UADT) involvement.48 Lymph node and extranodal involvement was identified by imaging and/or biopsy. Regional lymph nodes were those in the drainage area of the primary tumor and distant lymph nodes were those beyond the drainage area. A plasma LDH content above 245 U/L was considered elevated. Any detectable Epstein-Barr virus (EBV) DNA in plasma or whole blood was defined as EBV positive. Combined modality treatment (CMT) was defined as chemotherapy with or without radiotherapy (RT). Complete remission and relapse were determined by physical and radiological (CT, MRI, and PET-CT) examination according to Cheson criteria.49 Progressive disease (PD) was defined as a >25% increase in the sum of tumor lesion sizes, or emergence of new lesions. Progression-free survival (PFS) was calculated from the date of initial diagnosis to the first documented progression. Overall survival (OS) was calculated from the date of diagnosis to death from any cause.
Method details
DNA extraction
ctDNA was extracted using commercial ctDNA extraction kits (EliteHealth, Guangzhou Youze, China) following the manufacturer’s instructions and quantified using a Qubit 2.0 fluorometer (Invitrogen, Carlsbad, CA, USA). In addition, human normal NK/T cells were obtained by CD3+CD56+NK/T Cell Isolation Kit (Cat#130-093-064, Miltenyi Biotec, Bergisch Gladbach, Germany). Genomic DNA was extracted from tissue/cell samples using the DNeasy Blood &Tissue Kit (Cat#69506, QIAGEN, GmBH, Germany) following the manufacturer’s instructions and quantified using NanoDrop (Cat# ND-1000, Thermo Fisher, Waltham, MA, USA). Plasma sample volumes were in a range between 1.5-2 mL, cfDNA mass input amounts were larger than 15ng from each sample.
Full epigenetic (EPIC) methylation sequencing
Twenty individual ctDNA samples were pooled into a volume of 50 μL. DNA library pool preparation, hybridization-based target enrichment, bisulfite modification, and library amplification were performed using TruSeq Methyl Capture EPIC Library Prep Kit (Illumina, San Diego, CA, USA) according to the manufacturer’s protocol recommendations. Library concentration was determined using a Qubit 2.0 fluorometer (Invitrogen, Waltham, MA, USA) and library quality was assessed by capillary electrophoresis. Libraries were sequenced with a HiSeq X Ten Systems (Illumina, San Diego, CA, USA).
Padlock probe design and synthesis
Padlock probes covered 3,000,000 CpG markers were designed using the ppDesigner software. In general, the target CpG position was designed to be located at the central portion of a ∼100 bp captured region flanked by a linker sequence containing a primer-binding region separated by multiple Cs to produce probes of equal length. A 6-bp unique molecular identifier (UMI) was incorporated into the probe to accurately estimate the original (before PCR) methylated and unmethylated fragment count of the samples.
Probes were synthesized commercially (Integrated DNA Technologies, Colerville, IA, USA). For capture experiments, probes were mixed, in vitro phosphorylated with T4 polynucleotide kinase (New England Biolabs, Ipswich, MA, USA) according to the manufacturer’s instructions, and purified using P-30 Micro Bio-Spin columns (Bio-Rad, Hercules, CA, USA).
DNA methylation data processing
Raw methylation data were preprocessed using fastp version 0.20.0 with default parameters for both pooled and individual samples. Clean reads were then aligned to human genome build hg19 using bitmapperBS version 1.0.2.3 with default parameters for pooled samples and “pbat” mode for individual samples, and bam format results were sorted by sambamba version 0.7.0. Aligned reads were deduplicated based on UMIs using umi_tools dedup program.
DNA methylation calling was performed using MethylDackel version 0.4.0 extract default parameter for pooled samples and “--keepDups” parameter for individual samples, and DNA methylation calls for methylated and unmethylated controls were extracted from the alignment file. Sequencing depths are not less than 10 reads/sites. The methylated values located in the target regions were extracted using bedtools version 2.29.0. The methylated values, from 0 to 1 were used to quantify melathytion levels, and a threshold value of 0.10 for the methylated values and the associated p < 0.05 were considered significant. Differentially methylated CpGs between normal and tumor plasma samples were identified using DMRfinder version 0.3 with the beta-binomial hierarchical modeling and Wald test for pooled samples.
Building a diagnostic model
By using the Least Absolute Shrinkage and Selection Operator (LASSO) algorithm, 7 significant differential methylation markers were selected in the training dataset.46,47 Seventy-five percent of the dataset was subsampled for 100 iterations without replacement and features with an occurrence frequency >95% were selected. The expected generalization error was estimated by a 10-fold cross-validation to determine the tuning parameters, and the biggest value of lambda (“1-se” lambda) was adopted to keep the error within one standard error of the minimum and the consistency of LASSO selection results were improved after five iterations. A logistic regression model was fitted using the 7 markers identified by the LASSO algorithm as covariates, and a combined methylation diagnosis (cmd) score was calculated by multiplying the estimated unbiased coefficient and the marker methylation ratio matrix in both the training dataset and the validation dataset. The area under receiving operator characteristic (ROC) curve, which estimated the proportion of concordant pairs among all actual pairs of observations, with 1.0 representing perfect prediction, was used to evaluate the predictability of the model. The optimal cmd cutoff of the training dataset was determined by the maximum Youden index and used to generate the confusion tables for both the training dataset and the validation dataset.
Building a prognosis prediction model
We explored a prognosis prediction model using the two datasets. In the training dataset, a similar subsampling strategy was used to shrink the number of markers to a reasonable range in the LASSO-Cox method for prognostic marker selection.47 Briefly, LASSO regression shrinks the coefficient estimate towards zero by using 1-s.e. criteria. We adopted the LASSO regression model to achieve variable selection, and 7 markers with non-zero coefficient associated with OS were screened out to construct the prognostic signature. Subsequently, a Cox proportional hazards model was fitted with the corresponding coefficients for each marker to calculate a combined methylation prognostic (cmp) score for each individual. To validate our prediction model, we calculated the cmp score for each patient in the validation dataset using the same coefficient estimates from the training dataset. The optimal cmp cutoff determined by X-tile software was used to categorize the patients into the high cmp group and the low cmp score group.45
Quantification and statistical analysis
Univariate and multivariate cox proportional hazards models were used to evaluate the independent significance of covariates with the “survival” package in R software. Significant variables (p < 0.15) from univariate analysis were entered into multivariate analysis. We recorded the variables as dichotomous. We used Kaplan-Meier method and log-rank test to preform survival analyses. We use hazards ratio (HR) to compare the risk of survival events between different groups. Patients who were alive or experienced PD were censored at the last date known alive or at their last disease assessment. Coefficients of the prognostic covariates in the multivariate Cox regression model were used to construct a PINK-C risk system using the "rms" package in R software. We used calibration curve, concordance probability (C-index), and time-dependent ROC curve to evaluate the accuracy of the nomogram by using the “survival ROC” package in R software All statistical analyses were done using R software (version 3.5.3). A two-sided p-value < 0.05 was considered statistically significant.
Acknowledgments
We thank Kang Zhang (Center for Biomedicine and Innovations, Faculty of Medicine, Macau University of Science and Technology and University Hospital) and Gen Li (Guangzhou Women and Children’s Medical Center) for manuscript consultation, technical support, and sample processing. This work was supported by grants from the National Key Research and Development Program (2022YFC2502602), the National Natural Science Foundation of China (82230001 and 82270199), the Sun Yat-Sen University Clinical Research 5010 Program (2020009), the Special Support Program of Sun Yat-sen University Cancer Center (PT19020401), and the Clinical Oncology Foundation of Chinese Society of Clinical Oncology (Y-XD2019-124 and Y-SY2021ZD-0110).
Author contributions
Conception and design, Q.-Q.C. and X.-P.T.; development of methodology, X.-P.T.; acquisition of data, all authors; analysis and interpretation of data, X.-P.T.; writing, review, and/or revision of the manuscript, all authors; administrative, technical, or material support, all authors; study supervision, Q.-Q.C. All authors reviewed the manuscript and approved the final revision.
Declaration of interests
The authors declare no competing interests.
Published: February 21, 2023
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.xcrm.2022.100859.
Contributor Information
Rong Tao, Email: rtao@shca.org.cn.
Wen-Yu Li, Email: lwy80411@163.com.
Hui-Qiang Huang, Email: huanghq@sysucc.org.cn.
Qing-Qing Cai, Email: caiqq@sysucc.org.cn.
Supplemental information
Data and code availability
The key raw data have been deposited onto the China National GeneBank DataBase (CNGBdb) with accession number CNP0003741.
References
- 1.Harabuchi Y., Takahara M., Kishibe K., Nagato T., Kumai T. Extranodal natural killer/T-cell lymphoma, nasal type: Basic science and clinical progress. Front. Pediatr. 2019;7:141. doi: 10.3389/fped.2019.00141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kwong Y.L. Natural killer-cell malignancies: diagnosis and treatment. Leukemia. 2005;19:2186–2194. doi: 10.1038/sj.leu.2403955. [DOI] [PubMed] [Google Scholar]
- 3.Swerdlow S.H.C.E., Harris N.L., et al. Fourth edition. IARC; 2017. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues, Revised. [Google Scholar]
- 4.Tse E., Kwong Y.L. The diagnosis and management of NK/T-cell lymphomas. J. Hematol. Oncol. 2017;10:85. doi: 10.1186/s13045-017-0452-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Allen P.B., Lechowicz M.J. Management of NK/T-Cell lymphoma, nasal type. J. Oncol. Pract. 2019;15:513–520. doi: 10.1200/JOP.18.00719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Fox C.P., Civallero M., Ko Y.H., Manni M., Skrypets T., Pileri S., Kim S.J., Cabrera M.E., Shustov A.R., Chiattone C.S., et al. Survival outcomes of patients with extranodal natural-killer T-cell lymphoma: a prospective cohort study from the international T-cell Project. Lancet. Haematol. 2020;7:e284–e294. doi: 10.1016/S2352-3026(19)30283-2. [DOI] [PubMed] [Google Scholar]
- 7.Kim S.J., Yoon D.H., Jaccard A., Chng W.J., Lim S.T., Hong H., Park Y., Chang K.M., Maeda Y., Ishida F., et al. A prognostic index for natural killer cell lymphoma after non-anthracycline-based treatment: a multicentre, retrospective analysis. Lancet Oncol. 2016;17:389–400. doi: 10.1016/S1470-2045(15)00533-1. [DOI] [PubMed] [Google Scholar]
- 8.Zhang Y., Ma S., Cai J., Yang Y., Jing H., Shuang Y., Peng Z., Li B., Liu P., Xia Z., et al. Sequential P-GEMOX and radiotherapy for early-stage extranodal natural killer/T-cell lymphoma: a multicenter study. Am. J. Hematol. 2021;96:1481–1490. doi: 10.1002/ajh.26335. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Win K.T., Liau J.Y., Chen B.J., Takata K., Chen C.Y., Li C.C., Hsiao C.H., Chuang S.S. Primary cutaneous extranodal natural killer/T-cell lymphoma misdiagnosed as peripheral T-cell lymphoma: the importance of consultation/referral and inclusion of EBV in situ hybridization for diagnosis. Appl. Immunohistochem. Mol. Morphol. 2016;24:105–111. doi: 10.1097/PAI.0000000000000162. [DOI] [PubMed] [Google Scholar]
- 10.Tian X.P., Ma S.Y., Young K.H., Ong C.K., Liu Y.H., Li Z.H., Zhai Q.L., Huang H.Q., Lin T.Y., Li Z.M., et al. A composite single-nucleotide polymorphism prediction signature for extranodal natural killer/T-cell lymphoma. Blood. 2021;138:452–463. doi: 10.1182/blood.2020010637. [DOI] [PubMed] [Google Scholar]
- 11.Stroun M., Maurice P., Vasioukhin V., Lyautey J., Lederrey C., Lefort F., Rossier A., Chen X.Q., Anker P. The origin and mechanism of circulating DNA. Ann. N. Y. Acad. Sci. 2000;906:161–168. doi: 10.1111/j.1749-6632.2000.tb06608.x. [DOI] [PubMed] [Google Scholar]
- 12.Newman A.M., Bratman S.V., To J., Wynne J.F., Eclov N.C.W., Modlin L.A., Liu C.L., Neal J.W., Wakelee H.A., Merritt R.E., et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat. Med. 2014;20:548–554. doi: 10.1038/nm.3519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ignatiadis M., Sledge G.W., Jeffrey S.S. Liquid biopsy enters the clinic - implementation issues and future challenges. Nat. Rev. Clin. Oncol. 2021;18:297–312. doi: 10.1038/s41571-020-00457-x. [DOI] [PubMed] [Google Scholar]
- 14.Esteller M. Epigenetics in cancer. N. Engl. J. Med. 2008;358:1148–1159. doi: 10.1056/NEJMra072067. [DOI] [PubMed] [Google Scholar]
- 15.Schuebel K.E., Chen W., Cope L., Glöckner S.C., Suzuki H., Yi J.M., Chan T.A., Van Neste L., Van Criekinge W., van den Bosch S., et al. Comparing the DNA hypermethylome with gene mutations in human colorectal cancer. PLoS Genet. 2007;3:1709–1723. doi: 10.1371/journal.pgen.0030157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Baylin S.B., Jones P.A. A decade of exploring the cancer epigenome - biological and translational implications. Nat. Rev. Cancer. 2011;11:726–734. doi: 10.1038/nrc3130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Brock M.V., Hooker C.M., Ota-Machida E., Han Y., Guo M., Ames S., Glöckner S., Piantadosi S., Gabrielson E., Pridham G., et al. DNA methylation markers and early recurrence in stage I lung cancer. N. Engl. J. Med. 2008;358:1118–1128. doi: 10.1056/NEJMoa0706550. [DOI] [PubMed] [Google Scholar]
- 18.Xu R.H., Wei W., Krawczyk M., Wang W., Luo H., Flagg K., Yi S., Shi W., Quan Q., Li K., et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat. Mater. 2017;16:1155–1161. doi: 10.1038/nmat4997. [DOI] [PubMed] [Google Scholar]
- 19.Luo H., Zhao Q., Wei W., Zheng L., Yi S., Li G., Wang W., Sheng H., Pu H., Mo H., et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci. Transl. Med. 2020;12:eaax7533. doi: 10.1126/scitranslmed.aax7533. [DOI] [PubMed] [Google Scholar]
- 20.Downs B.M., Ding W., Cope L.M., Umbricht C.B., Li W., He H., Ke X., Holdhoff M., Bettegowda C., Tao W., Sukumar S. Methylated markers accurately distinguish primary central nervous system lymphomas (PCNSL) from other CNS tumors. Clin. Epigenetics. 2021;13:104. doi: 10.1186/s13148-021-01091-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shen S.Y., Singhania R., Fehringer G., Chakravarthy A., Roehrl M.H.A., Chadwick D., Zuzarte P.C., Borgida A., Wang T.T., Li T., et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature. 2018;563:579–583. doi: 10.1038/s41586-018-0703-0. [DOI] [PubMed] [Google Scholar]
- 22.Pessoa L.S., Heringer M., Ferrer V.P. ctDNA as a cancer biomarker: a broad overview. Crit. Rev. Oncol. Hematol. 2020;155 doi: 10.1016/j.critrevonc.2020.103109. [DOI] [PubMed] [Google Scholar]
- 23.Liu P.L., Cheng Z.X., Lu M.P., Zhang L.Q., Chen H.B. [Misdiagnosis analysis: 120 patients with nasal extranodal NK/T cell lymphoma in head and neck] Lin chuang er bi yan hou tou jing wai ke za zhi = Journal of clinical otorhinolaryngology, head, and neck surgery. 2020;34:73–78. doi: 10.13201/j.issn.1001-1781.2020.01.018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Xu P., Guo R., You J., Cheng S., Li J., Zhong H., Sun C., Xu H., Huang H., Li B., Zhao W. Dynamic evaluation of the prognostic value of (18)F-FDG PET/CT in extranodal NK/T-cell lymphoma, nasal type. Ann. Hematol. 2021;100:1039–1047. doi: 10.1007/s00277-021-04466-3. [DOI] [PubMed] [Google Scholar]
- 25.Suzuki R., Yamaguchi M., Izutsu K., Yamamoto G., Takada K., Harabuchi Y., Isobe Y., Gomyo H., Koike T., Okamoto M., et al. Prospective measurement of Epstein-Barr virus-DNA in plasma and peripheral blood mononuclear cells of extranodal NK/T-cell lymphoma, nasal type. Blood. 2011;118:6018–6022. doi: 10.1182/blood-2011-05-354142. [DOI] [PubMed] [Google Scholar]
- 26.Ito Y., Kimura H., Maeda Y., Hashimoto C., Ishida F., Izutsu K., Fukushima N., Isobe Y., Takizawa J., Hasegawa Y., et al. Pretreatment EBV-DNA copy number is predictive of response and toxicities to SMILE chemotherapy for extranodal NK/T-cell lymphoma, nasal type. Clin. Cancer Res. 2012;18:4183–4190. doi: 10.1158/1078-0432.CCR-12-1064. [DOI] [PubMed] [Google Scholar]
- 27.Li Y.X., Yao B., Jin J., Wang W.H., Liu Y.P., Song Y.W., Wang S.L., Liu X.F., Zhou L.Q., He X.H., et al. Radiotherapy as primary treatment for stage IE and IIE nasal natural killer/T-cell lymphoma. J. Clin. Oncol. 2006;24:181–189. doi: 10.1200/JCO.2005.03.2573. [DOI] [PubMed] [Google Scholar]
- 28.Kim G.E., Lee S.W., Chang S.K., Park H.C., Pyo H.R., Kim J.H., Moon S.R., Lee H.S., Choi E.C., Kim K.M. Combined chemotherapy and radiation versus radiation alone in the management of localized angiocentric lymphoma of the head and neck. Radiother. Oncol. 2001;61:261–269. doi: 10.1016/s0167-8140(01)00428-5. [DOI] [PubMed] [Google Scholar]
- 29.Ma H.H., Qian L.T., Pan H.F., Yang L., Zhang H.Y., Wang Z.H., Ma J., Zhao Y.F., Gao J., Wu A.D. Treatment outcome of radiotherapy alone versus radiochemotherapy in early stage nasal natural killer/T-cell lymphoma. Med. Oncol. 2010;27:798–806. doi: 10.1007/s12032-009-9288-7. [DOI] [PubMed] [Google Scholar]
- 30.Yang Y., Zhu Y., Cao J.Z., Zhang Y.J., Xu L.M., Yuan Z.Y., Wu J.X., Wang W., Wu T., Lu B., et al. Risk-adapted therapy for early-stage extranodal nasal-type NK/T-cell lymphoma: analysis from a multicenter study. Blood. 2015;126:1424–1432. doi: 10.1182/blood-2015-04-639336. [DOI] [PubMed] [Google Scholar]
- 31.Ohtake S., Ito Y., Fukuta M., Habuchi O. Human N-acetylgalactosamine 4-sulfate 6-O-sulfotransferase cDNA is related to human B cell recombination activating gene-associated gene. J. Biol. Chem. 2001;276:43894–43900. doi: 10.1074/jbc.M104922200. [DOI] [PubMed] [Google Scholar]
- 32.Verkoczy L.K., Guinn B.a., Berinstein N.L. Characterization of the human B cell RAG-associated gene, hBRAG, as a B cell receptor signal-enhancing glycoprotein dimer that associates with phosphorylated proteins in resting B cells. J. Biol. Chem. 2000;275:20967–20979. doi: 10.1074/jbc.M001866200. [DOI] [PubMed] [Google Scholar]
- 33.Wang X., Cheng G., Zhang T., Deng L., Xu K., Xu X., Wang W., Zhou Z., Feng Q., Chen D., et al. CHST15 promotes the proliferation of TE1 cells via multiple pathways in esophageal cancer. Oncol. Rep. 2020;43:75–86. doi: 10.3892/or.2019.7395. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Macedo D.B., Kaiser U.B. DLK1, Notch signaling and the timing of puberty. Semin. Reprod. Med. 2019;37:174–181. doi: 10.1055/s-0039-3400963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pittaway J.F.H., Lipsos C., Mariniello K., Guasti L. The role of delta-like non-canonical Notch ligand 1 (DLK1) in cancer. Endocr. Relat. Cancer. 2021;28:R271–R287. doi: 10.1530/ERC-21-0208. [DOI] [PubMed] [Google Scholar]
- 36.Winther M., Berezin V., Walmod P.S. NCAM2/OCAM/RNCAM: cell adhesion molecule with a role in neuronal compartmentalization. Int. J. Biochem. Cell Biol. 2012;44:441–446. doi: 10.1016/j.biocel.2011.11.020. [DOI] [PubMed] [Google Scholar]
- 37.Sasca D., Szybinski J., Schüler A., Shah V., Heidelberger J., Haehnel P.S., Dolnik A., Kriege O., Fehr E.M., Gebhardt W.H., et al. NCAM1 (CD56) promotes leukemogenesis and confers drug resistance in AML. Blood. 2019;133:2305–2319. doi: 10.1182/blood-2018-12-889725. [DOI] [PubMed] [Google Scholar]
- 38.Parcerisas A., Ortega-Gascó A., Pujadas L., Soriano E. The hidden side of NCAM family: NCAM2, a key cytoskeleton organization molecule regulating multiple neural functions. Int. J. Mol. Sci. 2021;22:10021. doi: 10.3390/ijms221810021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ji T., Ma K., Chen L., Cao T. PADI1 contributes to EMT in PAAD by activating the ERK1/2-p38 signaling pathway. J. Gastrointest. Oncol. 2021;12:1180–1190. doi: 10.21037/jgo-21-283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Coassolo S., Davidson G., Negroni L., Gambi G., Daujat S., Romier C., Davidson I. Citrullination of pyruvate kinase M2 by PADI1 and PADI3 regulates glycolysis and cancer cell proliferation. Nat. Commun. 2021;12:1718. doi: 10.1038/s41467-021-21960-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Han R., Campbell K.P. Dysferlin and muscle membrane repair. Curr. Opin. Cell Biol. 2007;19:409–416. doi: 10.1016/j.ceb.2007.07.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Cox A., Zhao C., Tolkach Y., Nettersheim D., Schmidt D., Kristiansen G., Hauser S., Müller S.C., Ritter M., Ellinger J. The contrasting roles of Dysferlin during tumor progression in renal cell carcinoma. Urol. Oncol. 2020;38 doi: 10.1016/j.urolonc.2020.04.021. 687.e1-687687.e11. [DOI] [PubMed] [Google Scholar]
- 43.Yang L., Wang L., Yang Z., Jin H., Zou Q., Zhan Q., Tang Y., Tao Y., Lei L., Jing Y., et al. Up-regulation of EMT-related gene VCAN by NPM1 mutant-driven TGF-beta/cPML signalling promotes leukemia cell invasion. J. Cancer. 2019;10:6570–6583. doi: 10.7150/jca.30223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Fujii K., Karpova M.B., Asagoe K., Georgiev O., Dummer R., Urosevic-Maiwald M. Versican upregulation in Sezary cells alters growth, motility and resistance to chemotherapy. Leukemia. 2015;29:2024–2032. doi: 10.1038/leu.2015.103. [DOI] [PubMed] [Google Scholar]
- 45.Camp R.L., Dolled-Filhart M., Rimm D.L. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin. Cancer Res. 2004;10:7252–7259. doi: 10.1158/1078-0432.CCR-04-0713. [DOI] [PubMed] [Google Scholar]
- 46.Fu W.J. Penalized estimating equations. Biometrics. 2003;59:126–132. doi: 10.1111/1541-0420.00015. [DOI] [PubMed] [Google Scholar]
- 47.Tibshirani R. The lasso method for variable selection in the Cox model. Stat. Med. 1997;16:385–395. doi: 10.1002/(sici)1097-0258(19970228)16:4<385::aid-sim380>3.0.co;2-3. [DOI] [PubMed] [Google Scholar]
- 48.Suzuki R., Suzumiya J., Yamaguchi M., Nakamura S., Kameoka J., Kojima H., Abe M., Kinoshita T., Yoshino T., Iwatsuki K., et al. Prognostic factors for mature natural killer (NK) cell neoplasms: aggressive NK cell leukemia and extranodal NK cell lymphoma, nasal type. Ann. Oncol. 2010;21:1032–1040. doi: 10.1093/annonc/mdp418. [DOI] [PubMed] [Google Scholar]
- 49.Cheson B.D., Fisher R.I., Barrington S.F., Cavalli F., Schwartz L.H., Zucca E., Lister T.A., et al. Alliance, Australasian Leukaemia and Lymphoma Group. Eastern Cooperative Oncology Group. European Mantle Cell Lymphoma Consortium. Italian Lymphoma Foundation Recommendations for initial evaluation, staging, and response assessment of Hodgkin and non-Hodgkin lymphoma: the Lugano classification. J. Clin. Oncol. 2014;32:3059–3068. doi: 10.1200/JCO.2013.54.8800. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
The key raw data have been deposited onto the China National GeneBank DataBase (CNGBdb) with accession number CNP0003741.