Abstract
Four-way junctions (4Hs) are important intermediates in DNA rearrangements such as genetic recombination. Under the influence of multivalent cations these molecules undergo a conformational change, from an extended planar form to a quasi-continuous stacked X-structure. Recently, a number of X-ray structures and a nuclear magnetic resonance (NMR) structure of 4Hs have been reported and in three of these the position of multivalent cations is revealed. These structures belong to two main families, characterized by the angle between the two co-axial stacked helices, which is either around +40 to +55° or around –70 to –80°. To investigate the role of metal-ion binding on the conformation of folded 4Hs we performed Brownian-dynamics simulations on the set of available structures. The simulations confirm the proposed metal-ion binding sites in the NMR structure and in one of the X-ray structures. Furthermore, the calculations suggest positions for metal-ion binding in the other X-ray structures. The results show a striking dependence of the ion density on the helical environment (B-helix or A-helix) and the structural family.
INTRODUCTION
Branched nucleic acid species play important roles in cellular processes that involve nucleic acid rearrangements. Examples are the DNA Holliday junction in genetic recombination, a four-way junction (4H) and the hammerhead ribozyme, a three-way junction, which is involved in RNA self catalysis (for reviews see 1–4). Metal ions participate in the folding to the three-dimensional structures of those nucleic acid architectures. It has been shown that branch migration in a 4H in the presence of multivalent cations is a factor 1000 slower than when they are absent (5,6), which is thought to be related to the different conformations in the absence or presence of multivalent cations (for review see 4). Detailed knowledge of nucleic acid structures, including their metal-ion binding sites, is therefore of importance to clarify the principles underlying their folding processes. The positions of metal ions in nucleic acid structures can be experimentally determined by X-ray crystallography or by nuclear magnetic resonance (NMR) spectroscopy (7–11). The first technique gives an electron density that subsequently can be attributed to a metal ion. NMR gives the position of the metal ion indirectly from intermolecular nuclear Overhauser effects; these are then used as distance restraints in the structure calculation. Recently it was shown (12) that Brownian-dynamics (BD) simulations of cation diffusion could faithfully predict metal-ion binding sites. In BD simulations of cation binding to nucleic acid molecules, a positively charged test sphere diffuses under the influence of Brownian motions towards the negatively charged molecule according to the non-linear Poisson–Boltzmann equation (13). The solvent is treated as a continuum with dielectric constant of 78 ɛ0 while within the nucleic acid molecule the dielectric constant is set to 4 ɛ0. The nucleic acid molecule is taken as a rigid framework in the BD simulations. By varying the radius and/or charge of the test sphere different cations can be simulated. Possible additional effects of inner-sphere binding or H-bond formation that can further stabilize the cation–nucleic acid complex are not explicitly considered. Recently, several groups observed metal ions in RNA structures (10,14) at positions deduced from BD simulations (12).
In the last decade a considerable amount of structural data has been collected on 4Hs by a variety of biophysical methods. In solution, DNA 4Hs exist as an equilibrium of three conformers (for reviews see 1,2,4). The unfolded form (Fig. 1A, center) is evident under conditions of low ionic strength and in the absence of multivalent cations. Note that such open forms are also observed in crystal structures of the junction in complex with the branch migration-promoting protein RuvA (15) and in complex with a site-specific recombination protein (16). Conditions that stabilize either one of the stacked conformers (Fig. 1A, left or right), such as ionic strength and the sequence around the junction, shift the equilibrium into one or the other direction. Only now, some of the folding rules start to emerge (17; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication). In particular, the penultimate base pair from the junction site has a strong influence on conformer selection (17–19). Already from the early data (20–23) it was reasoned that in the stacked conformation, the two-stacked helices are in an anti-parallel orientation. The angle between the two helical arms was deduced to be around 60° from fluorescence resonance energy transfer (FRET) (24), time resolved (tr) FRET (25), atomic-force microscopy (26) and from NMR using dipolar couplings (27; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication).
Figure 1.
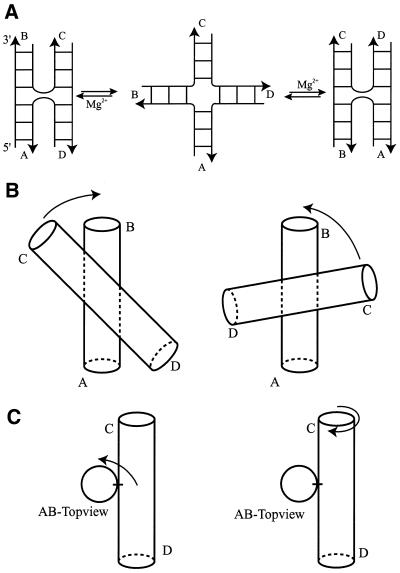
(A) Schematic representation of the conformational equilibria that 4Hs can undergo under the influence of multivalent cations. Under the influence of multivalent cations, the 4H undergoes a conformational transition from an open or extended form (center panel) into either one or a mixture of two alternatively stacked forms. The stacked conformers are denoted as an A/B-stacked conformation (left panel), where arm A stacks on arm B and arm C stacks on arm D, and an A/D-stacked conformation (right panel), in which arm A stacks on arm D and arm B stacks on arm C. The anti-parallel orientation can be seen from the backbones of the crossing strands that make a U-turn at the branch point. (B) Rotations that define the sign of the anti-parallel inter-helix angle between the A/B- and C/D-stacked helices. Left panel, positive inter-helix angle; right panel, negative inter-helix angle. (C) Left panel, rotation that defines the azimuthal rotation of the C/D-stacked helix perpendicular to the helical axis of the A/B-stacked helix; right panel, the same rotation but now seen as a rotation of the A/B-stacked helix with respect to the C/D-stacked helix (see text).
Recently, five high-resolution structures of stacked 4Hs became available (18,28–31; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication); four are X-ray structures and one is an NMR structure. The experimental structures mostly confirm the aforementioned general structural features of the 4H, e.g. co-axial stacking across the junction and the anti-parallel orientation of the stacked helices (Fig. 1). However, there is no coherence in the observed inter-helix angle between the five structures, which can be divided into two groups with inter-helix angle of ∼40 and –80°, respectively. Only one experimental structure showed magnesium-binding sites (31). The first 4H hybrid structure was also crystallized in the presence of cobalt(III)hexammine, which was observed in the electron density (28). The 4H NMR structure has also been determined in the presence of cobalt(III)hexammine and its binding site elucidated (18; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication). It has already been shown by two independent groups that DNA 4Hs in solution undergo the same folding process in the presence of cobalt(III)hexammine as in the presence of (hexa-hydrated) magnesium (11,32). It has to be noted, however, that also here the sequence can alter the effects of different cations (33). The two remaining X-ray structures, although at higher resolution than the 4H hybrid structures, did not show evidence for multivalent cation binding sites (29,30). In addition to these DNA or hybrid structures, an RNA 4H structure is present in the crystal structure of the hairpin ribozyme (34). No multivalent cations are observed in this 4H, in accordance with the finding that RNA 4Hs adopt a stacked conformation already in the absence of cations (for review see 4).
In all structures there is considerable clustering of phosphate groups around the point of strand exchange. The electrostatic potential in this central region is therefore highly negative. Previous calculations by other groups (35,36) suggest that there will be an accumulation of counterions in 4Hs in the stacked conformation. It has been shown (32,37) that monovalent ions at sufficiently high concentrations can induce the (partial) folding of DNA 4Hs into the stacked conformation. However, multivalent cations are much more efficient in this and probably have more specific binding sites, mainly thanks to their increased charge. However, their potential to form coordination complexes can also play a role.
We performed BD simulations on the set of DNA and DNA/RNA hybrid 4H structures to investigate and compare the metal-ion binding properties of 4Hs of different sequence and/or conformation. We find a marked dependence of the metal-ion binding on the conformation of the 4H.
MATERIALS AND METHODS
PDB coordinates
Atom coordinates were extracted from the Brookhaven Protein Data Bank (PDB). PDB accession codes are as follows: 1BR3 for the 4H structure in the DNA enzyme (28), 467D for the non-Watson–Crick 4H structure (29), 1DCW for the all DNA X-ray 4H structure (30) and J6 for the DNA 4H NMR structure (18; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication). For the NMR-derived structures, the average structure was calculated and minimized in Xplor (38) with holistic and the experimental restraints. For X-ray structures, hydrogen atoms were added in Xplor. All ligand and water atoms were removed from the coordinate sets prior to the computations.
Electrostatics
Electrostatics were treated with the non-linear Poisson–Boltzmann equation using a continuum model for the solvent with a dielectric constant of 78 ɛ0 for water and 4 ɛ0 for the nucleic acid molecule. Atomic partial charges are taken from the AMBER 4.1 force field (39). The grid spacing for calculation of the electrostatic field was 1.25–1.35 Å.
BD simulations
BD simulations were performed with a rigid nucleic acid target structure and freely diffusing metal ions using the UHBD program (13). The diffusing metal ions were modeled as spherical probes of radii between 1.0 and 2.5 Å and charge ranging between +1 and +3. Trajectories of the diffusing test spheres were initiated at randomly chosen points at a distance of r = 200 Å from the center of mass of the nucleic acid molecule. Trajectories exceeding an outer cut-off sphere of radius q = 300 Å were terminated. Test spheres that stayed within the cut-off sphere were uniformly simulated for 200 ns at a time step of 0.02 ps. Trajectory coordinates were recorded at intervals of 200 ps. For each probe radius, 2000 trajectories were calculated. The number of calculated trajectories did not influence the subsequent determination of metal-ion binding sites, as long as more than several hundred trajectories were used for statistical evaluation (12). A standard run would take 1–2 h on a R10000 Silicon Graphics workstation.
Determination of metal-ion binding sites
From the trajectories that stayed inside the outer cut-off sphere (see BD simulations), the final 100 recorded steps, corresponding to the last 20 ns of the simulation, were used for the evaluation. For each set of BD trajectories, the probability of finding the center of the charged probe within a discrete volume element in space was calculated on a cubic 1 Å grid, as described (12). The probabilities are categorized in terms of occupancy by an approximate ranking of predicted metal-ion binding sites according to the occupation of volume elements (12). The following operational method was applied: from the last 100 steps of the trajectories (2000 in total) that stayed within the cut-off sphere a volume element of 1 Å3 site is considered ‘highly occupied’ if more than 800 spheres are trapped within it. A high-occupancy site is defined when one isolated ‘highly occupied’ 1 Å3-volume element is found or when an isolated group of adjacent ‘highly occupied’ 1 Å3-volume elements is found. In addition, a simulated density site is considered delocalized when the r.m.s.d. from the center of a site is >1 Å. The probabilities are visualized as a density grid superimposed on the nucleic acid structure (see Figs 3 and 4).
Figure 3.
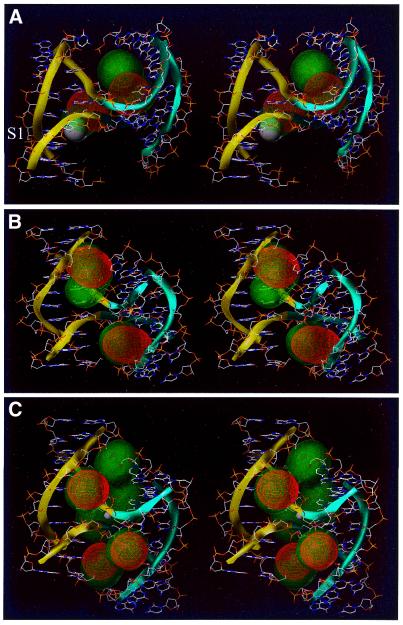
Stereo view of metal-ion binding sites predicted by BD simulations on 4Hs belonging to family I. The yellow and blue ribbons indicate the backbone of the A/B- and the C/D-stacked helix, respectively. Ion density calculated by BD simulations is superimposed on the 4H structure. High-occupancy sites for +3e charged test spheres are visualized as red density grids, while sites of lower occupancy are shown as green translucent densities. (A) 1BR3, crystallographic cobalt(III)hexammine is shown as a white sphere. (B) 467D and (C) 1DCW, crystallographic sodium is shown as a white sphere.
Figure 4.
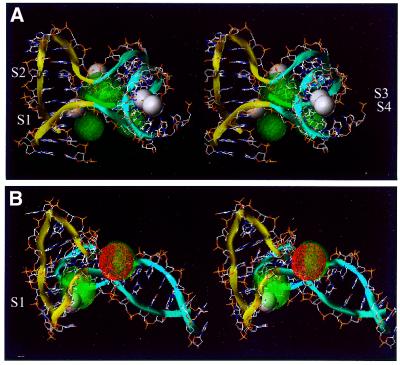
Stereo view of metal-ion binding sites predicted by BD simulations on 4Hs belonging to family II. The yellow and blue ribbons indicate the backbone of the A/B- and the C/D-stacked helix, respectively. Ion density calculated by BD simulations is superimposed on the 4H structure. High-occupancy sites for +3e charged test spheres are visualized as red density grids, while sites of lower occupancy are shown as green translucent densities. (A) 1EGK, crystallographic hexa-hydrated magnesium is shown as white spheres. (B) J6, the averaged position of the NMR cobalt(IIII)hexammine is shown as a white sphere.
RESULTS
General
The secondary structures of the 4Hs under investigation are shown in Figure 2. As noted before, all experimental 4H structures present a different inter-helical angle between the two-stacked helices (Table 1). The hybrid structure (1BR3) of the 4H in the 10–23 DNA enzyme has an inter-helix angle of 55°. Recently, Nowakowski et al. (31) reported an alternative 4H conformation (1EGK) for a hybrid 4H with almost the same sequence as in their first hybrid structure (Fig. 2). Here an inter-helix angle of –80° was observed. The two all DNA X-ray structures (467D and 1DCW) have both an inter-helix angle of ∼40°. This similarity is not surprising as the two sequences are very close (Fig. 2). In the NMR solution structure this angle is around –70°, the RNA 4H in the hairpin structure has a similar value (around –60°).
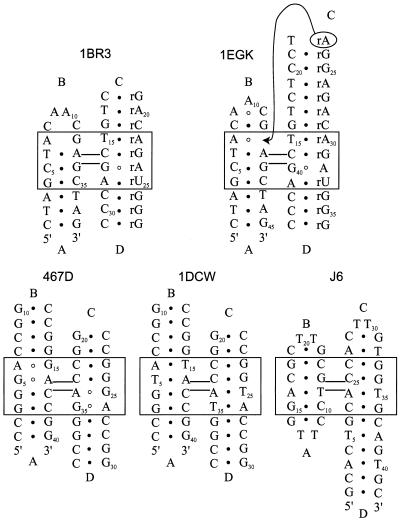
Figure 2.
Secondary structure of the different 4H molecules used in the BD simulations. The molecules are labeled with their PDB accession codes; 1BR3 (30), 1EGK (33), 467D (31), 1DCW (32) and J6 (B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication); helical arms are indicated with capital Arabics. In the original paper J6 was represented as an A/D-stacked conformer, for them sake of comparison we present it here as an A/B-stacked conformer. Non-canonical base pairs are indicated by open circles, standard Watson–Crick base pairs are indicated by closed circles (see text).
Table 1. Summary of characteristics of 4H structures classified in two families.
| Type | Method | Resolution (Å) | Helix angle (°) | Reference | |
|---|---|---|---|---|---|
| 1BR3 | RNA/DNA hybrid, mismatches | X-ray | 3.0 | 55 | (28) |
| 467D | DNA, mismatches | X-ray | 2.0 | 40 | (29) |
| 1DCW | DNA | X-ray | 2.0 | 41 | (30) |
| 1EGK | RNA/DNA hybrid, mismatches | X-ray | 3.1 | –80 | (31) |
| J6 | DNA | NMR | –70 | B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication |
As can be seen from the r.m.s.d. values (Table 2), all structures have similar architectural features around the stacked junction. However, further analysis of the structures shows that they can be classified in two broad families characterized by the inter-helix angle between the two co-axial stacks (see Fig. 1). In family I the inter-helix angle is ∼40–55° and in family II the inter-helix angle is negative and larger, ∼70–85° (see Table 1). The division in the two families was also suggested as right-handed crosses (family I) or left-handed crosses (family II) by Timsit and coworkers on the basis of observed packing interactions in different double-stranded B-DNA crystal structures (3,40,41). The value of the inter-helix angle is related to important stereochemical parameters that are worth discussing before describing the results. The inter-helix angle is first related to the handedness of the base pairs at the junctions (23). With the usual right-handed twist angle for the base pairs at the junctions, the inter-helix angle is <60° and the two exchanging strands cross above each other between the two helical stacks. Larger positive inter-helix angles are prohibited because the phosphates of the crossing strands would intermingle (22). If one helix is rotated in the negative direction with respect to the other, the two exchanging strands will become less and less crossed and tend to become parallel and above each other. This rotation can be accomplished by an under-twisting of the junction base pairs. Alternatively, the under-twisting could be distributed over the 3 bp steps in which the junction base pairs are involved. This is in fact observed in the experimental structures and will be discussed elsewhere (B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication). Much larger negative angles could even require left-handed helical stacks of the junction base pairs. In addition, one helical arm can present azimuthal rotations in a plane perpendicular to the helical axis of the other helical arm (see Fig. 1). These points are observed in the structures that have been used for the calculations.
Table 2. Heavy-atom atomic r.m.s.d.a for superimposition of the different 4H structures used in the simulations.
| A/B- and C/D-stack (Å) | A/B-stack (Å) | C/D-stack (Å) | |
|---|---|---|---|
| 1–5b all | 4.6 | 1.8 | 1.7 |
| 1–4b all X-ray | 3.2 | 1.6 | 1.6 |
| 1, 2b X-ray and DNA | 1.4 | 1.0 | 1.0 |
| 1, 2, 5b all DNA | 4.6 | 1.4 | 1.2 |
| 3, 4b X-ray and hybrid | 7.9 | 3.2 | 3.0 |
aBackbone heavy-atom r.m.s.d., values calculated with Molmol (44).
b1, 1DCW; 2, 467D; 3, 1BR3; 4, 1EGK; 5, J6.
As discussed, the set of available 4H structures shows a multitude of larger and smaller structural differences (e.g. the helix-parameters for all 4H structures considered can be seen in Table S1, available at NAR Online). This can therefore give valuable insights into the metal-ion binding properties of 4Hs and into the role of metal ions in the conformational exchange processes depicted in Figure 1A. To investigate the metal-ion binding sites in the structure of the 4Hs, we performed BD simulations on the set of structures. In the following, the results of the BD simulations are discussed for the different 4H structures. The BD simulations were performed with cationic probe spheres that mimic [Co(NH3)6]3+, [Mg(H2O)6]2+, Mg2+ and Na+, with the respective radii of 2.5 for the ligated cations, which should mimic outer-sphere binding and 1.0 Å for the naked ions which, on the other hand, should mimic inner-sphere binding.
Family I
Hybrid 4H in DNA enzyme (1BR3). Nowakowski et al. (28) reported the presence of a 4H RNA/DNA hybrid in the crystal structure of the 10–23 DNA enzyme at 3.0 Å resolution. This hybrid structure has two stacked A-type helical arms (arms C and D; see Fig. 2); arm A is B-helical whereas its stacking partner arm B is limited to 1 bp and a hexa-loop. Despite the fact that the structure was crystallized in the presence of MgCl2, no metal-ion binding sites could be observed. It was only in the presence of cobalt(III)hexammine that metal-ion binding sites were visible in the electron density. The 38 nt hybrid 4H—corresponding to nucleotides 2–11, 23–40 and 58–67 in the numbering scheme from the original publication (28)—was used as a target in a series of BD simulations (see Fig. 2). In all simulations a large number of diffusing probes were trapped in two main pockets. A high-occupancy site (Materials and Methods) is predicted by the BD simulations for a cobalt(III)hexammine ion, close to the experimental site described by the authors (Fig. 3A, S1; Table 3) (28). This site is located in the major groove of arm B close to the junction. The high r.m.s.d. (2.5 Å) of the center of the BD-calculated density suggests that the site is delocalized (Materials and Methods). The authors do not mention the presence of additional ions in the structure. We find in the BD simulations an additional high-occupancy site in the major groove at the other side of the junction in arm D (Fig. 3A; Table 3). Performing the simulations with probe spheres that mimic hexa-hydrated magnesium ions gives the same core densities. However, the densities are less localized, probably due to the lower charge (2+ versus 3+) of the magnesium ion (data not shown). This finding could well explain that in the 1BR3 4H hybrid structure, binding sites are observed in the presence of cobalt(III)hexammine but not in the presence of (hexa-hydrated) magnesium sites. Even more delocalized ion density is observed when the BD simulation is performed with charged spheres that mimic naked magnesium or sodium ions binding directly to the nucleic acid (data not shown).
Table 3. Centers of BD-predicted and experimental metal-ion binding sites in 4H structures.
| 4H structure | Site | X, Y, Za | r.m.s.d. (Å)c | ||
|---|---|---|---|---|---|
| 1BR3 | 1b | 25.6 | –12.1 | 3.8 | 2.5 |
| 2b | 16.6 | –14.6 | 6.3 | 1.2 | |
| 3b | 17.0 | –12.3 | 12.9 | 1.4 | |
| S1 | 27.2 | –7.4 | 1.8 | n.a. | |
| 467D | 1 | –9.5 | –1.3 | 2.8 | 1.0 |
| 2 | 9.5 | –1.3 | –2.8 | 1.0 | |
| 1DCW | 1 | –17.2 | –3.1 | 16.8 | 1.0 |
| 2 | –12.1 | 3.2 | 22.1 | 0.9 | |
| 3 | –0.4 | 2.9 | 12.9 | 0.7 | |
| 1EGK | 1b | 3.6 | 14.3 | –32.3 | 1.8 |
| 2b | 10.3 | 22.3 | –33.0 | 1.2 | |
| 3b | 12.3 | 13.8 | –23.0 | 1.2 | |
| S1 | 6.8 | 6.9 | –33.5 | 74.5 | |
| S2 | 5.4 | 23.2 | –40.0 | 116.7 | |
| S3 | 4.2 | 12.8 | 24.4 | 122.4 | |
| S4 | –2.3 | 32.1 | 42.8 | 122.6 | |
| J6 | 1b | 18.8 | –53.3 | 43.6 | 1.2 |
| 2 | 24.6 | –42.7 | 46.7 | 1.0 | |
| S1 | 15.0 | –55.2 | 42.3 | 1.8 |
aThe binding sites are those shown in Figures 3 and 4; when high-occupancy sites overlap with lower occupied sites, values are calculated only for the high-occupancy site. Atomic coordinates X, Y, Z refer to the orthogonal coordinate systems of the original PDB files (see Materials and Methods), S indicates experimental sites.
bDelocalized (see Materials and Methods).
cFor BD and NMR sites r.m.s.d. from average position, X-ray sites, B-factor (in Å2). n.a., not available.
467D and 1DCW. Ortiz-Lombardía et al. (29) reported the first all DNA 4H structure (henceforth 467D) at 2.16 Å resolution. In the sequence (Fig. 2), two non-canonical (G·A) base pairs are present. One is located at the junction and the other one penultimate to the junction in the same arm, the structure is symmetric around the junction. In another structure (30), the mismatch is replaced by standard T·A base pairs (Fig. 2). This structure (henceforth 1DCW) is also symmetric around the junction and is solved at 2.0 Å resolution. The backbone r.m.s.d.s for the two structures is 1.0 Å (see Table 2). In both structures no divalent cations could be observed, although both are crystallized in their presence (MgCl2 and CaCl2 for 467D and 1DCW, respectively). However, in 1DCW an electron density peak is observed that was assigned to a Na+ ion (30).
We performed BD simulations on both structures with different charge and radii spheres. The simulations with a sphere mimicking the cobalt(III)hexammine ion yielded high-occupancy sites in both molecules (Fig. 3B and C). They are symmetrically located in the minor grooves adjacent to the junction. As expected from the structural similarities, the densities observed in the two structures are similar. However, in the 1DCW structure they are slightly more localized than in the 467D structure (Table 3). In addition, a delocalized ion density (Materials and Methods) is calculated at the junction site of 1DCW. This ion density corresponds well with the experimentally observed Na+ ion (see Fig. 3C), although the experimentally observed Na+ ion has penetrated more into the interior of the junction, probably due to its smaller sphere than the simulated cobalt(III)hexammine sphere.
As no multivalent cations are observed in the experimental structures 467D and 1DCW, a comparison between predicted and experimental multivalent cation sites cannot be made here. However, a comparison of the calculated densities in these structures with the site experimentally observed in the 1BR3 hybrid structure (Fig. 3A, S1) shows that in the 1DCW and 467D structure (Fig. 3B and C) no density is observed in the major grooves close to the junction. The ion densities have moved to the minor grooves in the 1DCW and 467D structures, despite the similar inter-helix angles. However, note that in the 1BR3 structure the minor groove at the corresponding locations is either perturbed or very narrow, which can explain this difference (see Discussion). The same delocalization effect with respect to lower charge and/or sphere radius as mentioned above is also observed here.
Family II
1EGK second hybrid 4H structure. The secondary structure of this hybrid 4H (1EGK) has three arms that are very similar to the ones in the 1BR3 structure (see Fig. 2), except for the following differences: (i) the third base pair from the junction in arm A is switched from rCG in 1BR3 to rGC in 1EGK; (ii) residue A7 (see Fig. 2), which has a deoxyribose sugar in the 1EGK sequence whereas it has a ribose sugar in the 1BR3 structure. The fourth arm (arm D) consists in the 1BR3 structure of a junction base pair capped by a hexa-loop. In the 1EGK structure, this hexa-loop structure is no longer present. Arm D now consists of the junction base pair, the six repositioned residues and in addition residue A23 (see Fig. 2), which makes a long-range interaction. This residue intercalates at the penultimate position in arm C, where it forms a non-canonical base pair with A7.
The 46 nt 1EGK hybrid 4H—corresponding to nucleotides 1–14, 27–36 and 133–154 in the numbering scheme from the original publication (31), was used as a target in a series of BD simulations (see Fig. 2). Four of the experimentally observed hexa-hydrated magnesium sites are inside this part of the molecule. Delocalized densities are observed for the simulations for all ionic spheres tested. Because of the negative inter-helix angle, a pocket is created on one side of the junction where the major grooves of the helical arms face each other. The main calculated ion densities with a charged test sphere mimicking the cobalt(III)hexammine ion spread out in this pocket (Fig. 4A). The ion density is delocalized as follows from the r.m.s.d. values in Table 3. Three of the experimental sites (Fig. 4A, S1, S2 and S3) are located on the edge of this pocket. The high B-factors (Table 3) of the crystallographic magnesium sites also suggest a delocalized binding mode. The fourth site (Fig 4A, S4) resides at the outside of the 1EGK hybrid 4H structure and no density is calculated from the BD simulations (this site has also a high B-factor). Again the density becomes more delocalized when test spheres are used with a lower charge and/or smaller radii.
NMR 4H structure. van Buuren et al. (18; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication) reported the first solution structure of an intra-molecular DNA 4H. Because of the negative helix angle, on one side of the junction the major grooves of the helical arms face each other and form a cavity. In the NMR experiments, cobalt(III)hexammine was used as a probe to determine the location of metal-ion binding site(s). Only one binding pocket was observed in that study. It is located adjacent to the junction in the major groove of arm A, which forms one side of the pocket formed by the two facing major grooves (note that for convenience of comparison with the other 4Hs, the arm notation has here been circularly rotated compared with the one used by van Buuren et al. (18; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication). In the study, a fast chemical exchange (lifetimes smaller than milliseconds) between free and bound cobalt(III)hexammine was found (11). In the BD simulations, using a sphere that mimics the cobalt(III)hexammine ion, ion density was calculated between the two facing major grooves of arms A and D. Figure 4B shows the ion density calculated by the BD simulations superimposed on the experimental NMR structure. As can be seen, the calculated density overlaps with the experimentally derived cobalt(III)hexammine ion binding site (Fig. 4B, S1; Table 3). The r.m.s.d. values (>1 Å) of both the BD-calculated density and the average experimental site (S1) suggest that the metal-ion binding site is delocalized. The ion density of this site becomes more delocalized when using a charged test sphere mimicking hexa-hydrated magnesium, an even more delocalized density is observed with spheres mimicking lower charge and smaller sphere cations (data not shown). A high-occupancy density is calculated on the other side of the junction. It is highly localized as it is coordinated with the crossing phosphates and directly faces the solution, i.e. no pocket is formed here in which the ion is captured. The simulations were run on the averaged structure of the NMR ensemble, performing the simulation on another structure of the ensemble this ion density shifts with respect to its location in the average structure. Over the ensemble of NMR structures this solvent-exposed density will smear out and become less occupied.
DISCUSSION
All the calculated densities, either with ions mimicking outer-sphere binding or inner-sphere binding, are rather smeared out, suggesting multiple or delocalized binding sites for the ions, which would point to weakly specific ion binding to the 4H structures. Dynamically delocalized ion binding, probably also in fast exchange with bulk (as observed experimentally in J6), should appear prevalent from the calculated ion densities. However, some sites are more frequently occupied than others, leading to higher ionic density levels. Above a given cut-off value (see Materials and Methods), we considered those sites as ‘high-occupancy’ binding sites. Some of those ‘high-occupancy’ sites might, but not necessarily, constitute specific ion binding sites. In any case, without considering partially dehydrated ions the specificity of recognition cannot be properly assessed. This is emphasized by the fact that with decreasing sphere size and charge the delocalization in ion binding becomes even more pronounced, indicating that (naked) divalent and monovalent cations are less frequently bound to 4Hs than trivalent cations and that the importance of inner-sphere binding is limited. In accordance with this is the absence of monovalent cations in most experimental 4H structures, whereas in one case (1EGK) binding sites for (hexa-hydrated) magnesium ions and in two cases for trivalent cations are detected (1BR3 and J6).
For all 4H structures the main BD-obtained density is observed in the grooves of the structures in proximity to the junction. In a recent publication, monovalent ion binding in the minor groove of B-DNA was reported from a molecular dynamics study (42), the ion binding affected the time-averaged minor groove width. Indeed, in structures containing a B-type helix (e.g. 1DCW) this was also observed in this study. However, we note that the nucleic acid structure is kept fixed in the BD calculations, and variability in nucleic acid structure can therefore not be observed here. Generally speaking, for structures belonging to family I, the ion densities are observed in the minor grooves facing the junction. Simulations on the 4H model by von Kitzing et al. (22), which belongs to family I, also yield minor groove binding (data not shown). In the 1BR3 structure an intermediate pattern is observed for the calculated ion densities. This deviating pattern can be explained by the fact that most arms deviate from B-DNA in geometry. The RNA helix has a narrower major groove than B-DNA, preventing the spheres penetrating. The minor groove of the DNA helix in arm B is perturbed by the hexa-loop and, thus, this site merges with the major groove of the facing A-helix (arm C), creating a large pocket where the main density is observed.
The structures belonging to family II have facing major grooves at the junction, and this geometrical arrangement creates a large cavity suitable for cation binding. This is indeed where the main ion densities are observed in the BD simulations. Simulations performed on an RNA 4H model from the U1 snRNA molecule (23) mainly confirm this finding (data not shown). Generally, the observed densities in family II structures present a lower occupancy than the densities in structures belonging to family I. The smeared out density in the large cavity suggests that alterations in sequence in this part of the molecule will only have small consequences for the metal-ion binding pocket. However, in both families the exact nature of the metal-ion binding site can be affected by sequence variation. In this way, the bias of the conformational equilibrium (Fig. 1A) might be influenced. The main contribution of the metal ion in this conformational exchange process occurs through the preferred binding to one of the stacked conformations, thereby stabilizing it. Indeed, although non-electrostatic interactions play an important role here (11,18; B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication), the local sequence around the stacked junction can influence the conformational equilibria via the electrostatic field generated by the base pair stacks close to the junction.
Another question that still remains, and which is not addressed here, is the reason why certain 4H molecules prefer family I, while others prefer family II. Geometry suggests that RNA 4Hs belong to family II (23). This is confirmed by the RNA 4H in the hairpin ribozyme structure (34). For DNA or DNA/RNA hybrid 4H structures both families are experimentally observed. That both families can occur was already indicated by the packing interactions seen in crystals of double-stranded DNA molecules (3,21,41). Family I requires that the positive helix angle remains relatively small (less than ∼60°), otherwise the crossing phosphates would come too close (22). When the helix angle is in the negative domain (family II), the crossing strands become less crossed and large negative inter-helix angles are allowed. These larger angles are experimentally observed in the crystalline environment as well as in solution (Table 2). In the crystal structure (1EGK) this conformation is stabilized by intercalation and base pairing of residue A23 (see above). It is interesting to note that in solution the DNA 4H has the relatively large negative helix angle. One would indeed expect that in the absence of specific interactions, in solution the less compact conformation is favored (B.N.M.van Buuren, J.Schleucher and S.S.Wijmenga, submitted for publication).
In conclusion, we report the location of metal-ion binding sites in a set of 4H structures, as derived by BD simulations. The results indicate that multivalent cations stabilize the stacked 4H conformation by mainly non-specific electrostatic interactions, neutralizing the negative charge of the phosphates that cluster in the grooves surrounding the branch point in the stacked 4H. The 4H structures can be divided into two main families: family I has an inter-helix angle of around +40 to +55° (‘right-handed cross’) while family II presents an angle around –70 to –80° (‘left-handed cross’). Thus, in family I, the main sites are localized in the minor grooves, whereas they are preferentially localized in the major grooves in family II. Structures built from short DNA helices can be of any family, but all-RNA 4Hs appear to adopt preferentially family II structures. The global architectures of the two families are such that the potential binding pockets have different sizes and are placed differently with respect to the grooves. The results show that the global conformation of the stacked 4H structure and the location of its metal-ion binding site(s) are intimately related parameters. Interestingly, theoretical calculations on rod-like polyions with condensed counterions show that two molecules can attract each other even if they have the same net charge (43). Furthermore, it is found that at short distances the parallel orientation of the rods (similar to family I) corresponds to the minimum of the free energy, while the perpendicular orientation (akin to family II) is the free energy minimum at large separations, becoming meta-stable for short distances (43). Thus, in all DNA 4Hs, the transition from family II to family I should critically depend on environmental factors like ionic strength, ion type and thermal fluctuations.
SUPPLEMENTARY MATERIAL
A table containing the helix-parameters for all 4H structures considered is available as supplementary material at NAR Online.
Acknowledgments
ACKNOWLEDGEMENTS
We thank Dr Nowakowski for providing unpublished coordinates of cobalt(III)hexammine ions and Dr Youri Timsit for useful discussions. B.N.M.V.B. was supported by an EMBO short-term fellowship. S.S.W. was supported by grants from the Swedish National Research Council and the Bioteknik Medel Umeå University.
REFERENCES
- 1.Seeman N.C. and Kallenbach,N.R. (1994) DNA branched junctions. Annu. Rev. Biophys. Biomol. Struct., 23, 53–86. [DOI] [PubMed] [Google Scholar]
- 2.Altona C., Pikkemaat,J.A. and Overmars,F.J.J. (1996) Three-way and four-way junctions in DNA: a conformational viewpoint. Curr. Opin. Struct. Biol., 6, 305–316. [DOI] [PubMed] [Google Scholar]
- 3.Timsit Y. and Moras,D. (1996) Cruciform structures and functions. Q. Rev. Biophys., 29, 279–307. [DOI] [PubMed] [Google Scholar]
- 4.Lilley D.M.J. (2000) Structures of helical junctions in nucleic acids. Q. Rev. Biophys., 33, 109–159. [DOI] [PubMed] [Google Scholar]
- 5.Panyutin I.G. and Hsieh,P. (1994) The kinetics of spontaneous DNA branch migration. Proc. Natl Acad. Sci. USA, 91, 2021–2025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Panyutin I.G., Biswas,I. and Hsieh,P. (1995) A pivotal role for the structure of the Holliday junction in DNA branch migration. EMBO J., 14, 1819–1826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hud N.V. and Feigon,J. (1997) Localization of divalent metal ions in the minor groove of DNA A-tracts. J. Am. Chem. Soc., 119, 5756–5757. [Google Scholar]
- 8.Kieft J.S. and Tinoco,I.,Jr (1997) Solution structure of a metal-binding site in the major groove of RNA complexed with cobalt (III) hexammine. Structure, 5, 713–721. [DOI] [PubMed] [Google Scholar]
- 9.Hud N.V., Schultze,P. and Feigon,J. (1998) Ammonium ion as an NMR probe for monovalent cation coordination sites of DNA quadruplexes. J. Am. Chem. Soc., 120, 6403–6404. [Google Scholar]
- 10.Gonzalez R.L. Jr, and Tinoco,I.,Jr (1999) Solution structure and thermodynamics of a divalent metal ion binding site in an RNA pseudoknot. J. Mol. Biol., 289, 1267–1282. [DOI] [PubMed] [Google Scholar]
- 11.van Buuren B.N.M., Schleucher,J. and Wijmenga,S.S. (2000) NMR structural studies on a DNA four-way junction: stacking preferences and localization of the metal-ion binding site. J. Biomol. Struct. Dyn., S2, 237–244. [DOI] [PubMed] [Google Scholar]
- 12.Hermann T. and Westhof,E. (1998) Exploration of metal ion binding sites in RNA folds by Brownian-dynamics simulations. Structure, 6, 1303–1314. [DOI] [PubMed] [Google Scholar]
- 13.Madura J.D., Davis,M.E., Gilson,M.K., Wade,R.C., Luty,B.A. and McCammon,J.A. (1994) Biological applications of electrostatic calculations and Brownian-dynamics simulations. Rev. Comp. Chem., 5, 229–267. [Google Scholar]
- 14.Hansen M.R., Simorre,J.P., Hanson,P., Mokler,V., Bellon,L., Beigelman,L. and Pardi,A. (1999) Identification and characterization of a novel high affinity metal-binding site in the hammerhead ribozyme. RNA, 5, 1099–1104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hargreaves D., Rice,D.W., Sedelnikova,S.E., Artymiuk,P.J., Lloyd,R.G. and Rafferty,J.B. (1998) Crystal structure of E.coli RuvA with bound DNA Holliday junction at 6 Å resolution. Nature Struct. Biol., 5, 441–446. [DOI] [PubMed] [Google Scholar]
- 16.Gopaul D.N., Guo,F. and Van Duyne,G.D. (1998) Structure of the Holliday junction intermediate in Cre-loxP site-specific recombination. EMBO J., 14, 4175–4187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.van Buuren B.N.M., Overmars,F.J.J., Ippel,J.H., Altona,C. and Wijmenga,S.S. (2000) Solution structure of a DNA three-way junction containing two unpaired thymidine bases; identification of sequence features that decide conformer selection. J. Mol. Biol., 304, 371–383. [DOI] [PubMed] [Google Scholar]
- 18.van Buuren B.N.M. (2001) Unwinding the structure of branched DNA: structural studies on three- and four-way junctions by NMR and molecular modeling. Thesis, Umea University, Umea, Sweden.
- 19.Miick S.M., Fee,R.S., Millar,D.P. and Chazin,W.J. (1997) Crossover isomer bias is the primary sequence-dependent property of immobilized Holliday junctions. Proc. Natl Acad. Sci. USA, 94, 9080–9084. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Duckett D.R., Murchie,A.I., Diekmann,S., von Kitzing,E., Kemper,B. and Lilley,D.M.J. (1988) The structure of the Holliday junction and its resolution. Cell, 55, 79–89. [DOI] [PubMed] [Google Scholar]
- 21.Timsit Y., Westhof,E., Fuchs,R.P.P. and Moras,D. (1989) Unusual helical packing in crystals of DNA bearing a mutation hot spot. Nature, 341, 459–462. [DOI] [PubMed] [Google Scholar]
- 22.von Kitzing E., Lilley,D.M.J. and Diekmann,S. (1990) The stereochemistry of a four-way DNA junction: a theoretical study. Nucleic Acids Res., 18, 2671–2683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Krol A., Westhof,E., Bach,M., Luhrmann,R., Ebel,J. and Carbon,P. (1990) Solution structure of human U1 snRNA. Derivation of a possible three-dimensional model. Nucleic Acids Res., 18, 3803–3811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Murchie A.I., Clegg,R.M., von Kitzing,E., Duckett,D.R., Diekmann,S. and Lilley,D.M.J. (1989) Fluorescence energy transfer shows that the four-way DNA junction is a right-handed cross of antiparallel molecules. Nature, 341, 763–766. [DOI] [PubMed] [Google Scholar]
- 25.Eis P.S. and Millar,D.P. (1993) Conformational distributions of a four-way DNA junction revealed by time-resolved fluorescence resonance energy transfer. Biochemistry, 32, 13852–13860. [DOI] [PubMed] [Google Scholar]
- 26.Mao C.D., Sun,W.Q. and Seeman,N.C. (1999) Designed two-dimensional DNA Holliday junction arrays visualized by atomic 7 force microscopy. J. Am. Chem. Soc., 121, 5437–5443. [Google Scholar]
- 27.Cromsigt J., van Buuren,B., Schleucher,J. and Wijmenga,S. (2001) Resonance assignment and structure determination for RNA. Methods Enzymol., 338, 371–399. [DOI] [PubMed] [Google Scholar]
- 28.Nowakowski J., Shim,P.J., Prasas,G.S., Stout,C.D. and Joyce,G.F. (1999) Crystal structure of an 82-nucleotide RNA–DNA complex formed by th 10–23 DNA enzyme. Nature Struct. Biol., 6, 151–156. [DOI] [PubMed] [Google Scholar]
- 29.Ortiz-Lombardía M., González,A., Eritja,R., Aymamí,J., Azorín,F. and Coll,M. (1999) Crystal structure of a DNA Holliday junction. Nature Struct. Biol., 6, 913–917. [DOI] [PubMed] [Google Scholar]
- 30.Eichman B.F., Vargason,J.M., Mooers,B.H.M. and Ho,P.S. (2001) The Holliday junction in an inverted repeat DNA sequence: sequence effects on the structure of four-way junctions. Proc. Natl Acad. Sci. USA, 97, 3971–3976 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nowakowski J., Shim,P.J., Stout,C.D. and Joyce,G.F. (2000) Alternative conformation of a nucleic acid four-way junction. J. Mol. Biol., 300, 93–102. [DOI] [PubMed] [Google Scholar]
- 32.Duckett D.R., Murchie,A.I.H. and Lilley,D.M.J. (1990) The role of metal ions in the conformation of the four-way DNA junction. EMBO J., 9, 583–590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Grainger R.J., Murchie,A.I.H. and Lilley,D.M.J. (1998) Exchange between stacking conformers in a four-way DNA junction. Biochemistry, 37, 23–32. [DOI] [PubMed] [Google Scholar]
- 34.Rupert P.B. and Ferre-D’Amare,A.R. (2001) Crystal structure of a hairpin ribozyme-inhibitor complex with implications for catalysis. Nature, 410, 780–786. [DOI] [PubMed] [Google Scholar]
- 35.Olmsted M.C. and Hagerman,P.J. (1994) Excess counterion accumulation around branched nucleic-acids. J. Mol. Biol., 243, 919–929. [DOI] [PubMed] [Google Scholar]
- 36.Fenley M.O., Manning,G.S., Marky,N.L. and Olson,W.K. (1998) Excess counterion binding and ionic stability of kinked and branched DNA. Biophys. Chem., 74, 135–152. [DOI] [PubMed] [Google Scholar]
- 37.Clegg R.M., Murchie,A.I., Zechel,A., Carlberg,C., Diekmann,S. and Lilley,D.M.J. (1992) Fluorescence resonance energy transfer analysis of the structure of the four-way DNA junction. Biochemistry, 31, 4846–4856. [DOI] [PubMed] [Google Scholar]
- 38.Brunger A.T. (1993) X-PLOR Version 3.1: A System for X-ray Crystallography and NMR. Yale University Press, New Haven, CT.
- 39.Cornell W.D. and Kollman,P.A. (1995) A second generation force field for the simulation of proteins, nucleic acids and organic molecules. J. Am. Chem. Soc., 117, 5179–5197. [Google Scholar]
- 40.Timsit Y. and Moras,D. (1991) Groove-backbone interactions in B-DNA implications for DNA condensation and recombination. J. Mol. Biol., 221, 919–940. [DOI] [PubMed] [Google Scholar]
- 41.Timsit Y., Shatzky-Schwartz,M. and Shakked,Z. (1999) Left-handed DNA crossovers. Implications for DNA–DNA recognition and structural alterations. J. Biomol. Struct. Dyn., 16, 775–785. [DOI] [PubMed] [Google Scholar]
- 42.Hamelberg D., Williams,L.D. and Wilson,W.D. (2001) Influence of the dynamic positions of cations on the structure of the DNA minor groove: sequence-dependent effects. J. Am. Chem. Soc., 123, 7745–7755. [DOI] [PubMed] [Google Scholar]
- 43.Stilck J.F., Levin,Y. and Arenzon,J.J. (2002) Thermodynamic properties of a simple model of like-charged attracting rods. J. Stat. Phys., 106, 287–293. [Google Scholar]
- 44.Koradi R., Billeter,M. and Wüthrich,K. (1996) MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph., 14, 51–55. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.