SUMMARY
Realizing the full utility of brain organoids to study human development requires understanding whether organoids precisely replicate endogenous cellular and molecular events, particularly since acquisition of cell identity in organoids can be impaired by abnormal metabolic states. We present a comprehensive single-cell transcriptomic, epigenetic, and spatial atlas of human cortical organoid development, comprising over 610,000 cells, from generation of neural progenitors through production of differentiated neuronal and glial subtypes. We show that processes of cellular diversification correlate closely to endogenous ones, irrespective of metabolic state, empowering the use of this atlas to study human fate specification. We define longitudinal molecular trajectories of cortical cell types during organoid development, identify genes with predicted human-specific roles in lineage establishment, and uncover early transcriptional diversity of human callosal neurons. The findings validate this comprehensive atlas of human corticogenesis in vitro as a resource to prime investigation into the mechanisms of human cortical development.
In brief
A resource encompassing single-cell transcriptomic, epigenetic, and spatial atlases of human cortical organoid development shows that processes of cellular diversification in organoids correlate closely to endogenous ones, irrespective of metabolic state, and identifies genes with predicted human-specific roles in lineage establishment.
Graphical Abstract

INTRODUCTION
Development of the human cerebral cortex is a protracted process that spans long periods of in utero development and thus is largely experimentally inaccessible (Silbereis et al., 2016). Although studies using human fetal tissue have proven critical to our current knowledge of human corticogenesis (Florio et al., 2017; Miller et al., 2019), there is an outstanding need for experimental models that can be used to dissect the molecular logic governing human cortical development.
Human brain organoids have emerged as a powerful experimental system to investigate human brain development and neurodevelopmental diseases (Kelley and Pașca, 2022). Organoids generate a large diversity of cell types resembling those that populate the human developing cortex (Camp et al., 2015; Quadrato et al., 2017a; Velasco et al., 2019a; Yoon et al., 2019) in a reproducible fashion (Velasco et al., 2019a). Additionally, organoids broadly recapitulate the transcriptional and epigenetic profile of the endogenous fetal cortex (Velasco et al., 2019a; Trevino et al., 2020; Gordon et al., 2021) and show physiologically-relevant features such as the presence of neuronal activity (Quadrato et al., 2017a; Trujillo et al., 2019, 2020; Miura et al., 2020).
These features make human brain organoids a promising model system for studying poorly-understood processes of human brain development, such as events of human cortical cellular diversification, in particular of populations that have undergone striking species-specific expansion and diversification during human evolution, such as outer radial glia (oRG) and callosal projection neurons (CPN) of the upper cortical layers. However, development in vitro has its challenges, including the possibility of impaired acquisition of cell identity due to abnormal metabolic states not observed in the embryo (Badhuri et al., 2020). Here, we demonstrate through a comprehensive, multi-modal atlas of organoid development across time that proper cell diversity can be established in organoids and that cell identity is largely not affected by metabolic state. We leverage this atlas to analyze cell-type specification and diversification of each cortical lineage, at single-cell resolution, across long timelines of organoid development. These findings validate the utility of this resource to enable investigation of the developmental origin of human-specific cortical features and human neurodevelopmental abnormalities.
RESULTS
Single-cell-resolution transcriptomic, epigenomic, and spatial atlas of human cortical organoid development
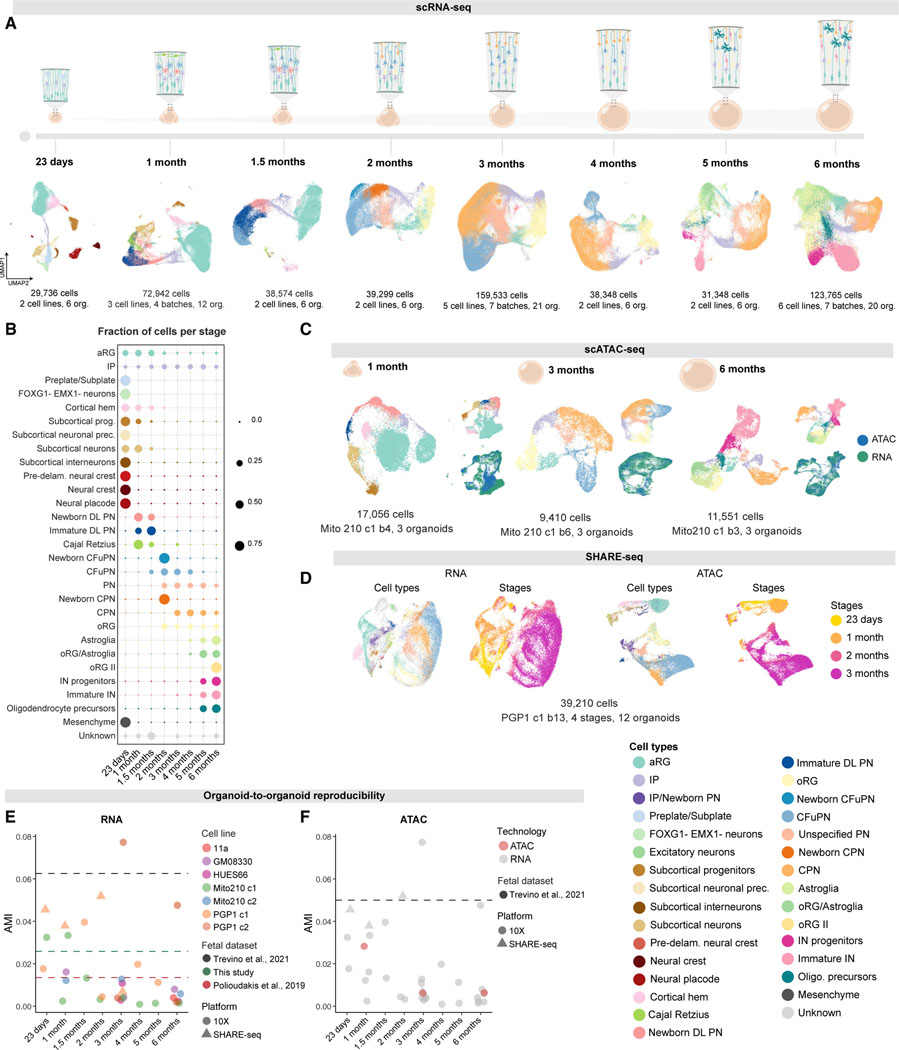
To investigate the molecular characteristics and reproducibility of corticogenesis in human brain organoids, we built a longitudinal single-cell atlas comprising eight timepoints across 6 months of organoid development, spanning processes from early stages of progenitor amplification to later astroglia production (Figure 1A) and profiled at the RNA, chromatin, and spatial transcriptomics levels. Our transcriptomic dataset includes 532,414 cells profiled by single-cell RNA-sequencing (scRNA-seq), combining our previously published datasets (Velasco et al., 2019a; Paulsen et al., 2022) with an additional 218,240 newly profiled cells from multiple timepoints, totaling 83 individually profiled organoids derived from multiple stem cell lines (2–6 lines per stage) and differentiation batches (2–7 batches per line and stage; Figures 1A, 1B, S1, and Data S1 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Our epigenomic dataset comprises 38,017 nuclei profiled by single-cell assay for transposase accessible chromatin using sequencing (scATAC-seq) (Corces et al., 2017; Satpathy et al., 2019), combining our previously published dataset (Paulsen et al., 2022) with an additional 11,551 newly profiled nuclei from organoids after 1, 3, and 6 months in vitro, spanning amplification of neural progenitors, the peak of excitatory neuron diversity, and the emergence of astroglia and interneurons, respectively (Figures 1C and S2A–S2H). In addition, we include 42,810 cells profiled by SHARE-seq, which captures RNA-seq and ATAC-seq profiles simultaneously from the same single cell (Ma et al., 2020) at 4 time points, from 23 days to 3 months (Figures 1D and S2I). Finally, our spatial transcriptomics dataset comprises 10 organoids profiled by Slide-seqV2 (Stickels et al., 2021) at 1, 2, and 3 months in vitro (Figure 2 and S2L–S2Q), spanning the emergence and expansion of excitatory neurons, which achieve their highest diversity in organoids at 3 months in vitro (below). These data provide a comprehensive multiomics molecular map of the development of human cortical organoids. We provide this data as a resource for interactive exploration at the Single Cell Portal: https://singlecell.broadinstitute.org/single_cell/study/SCP1756.
Figure 1. Single-cell transcriptomic and epigenetic landscape of developing cortical organoids.
(A) scRNA-seq of organoids at eight time points. Cells are colored by cell type.
(B) Fraction of cells per cell type at each timepoint.
(C) scATAC-seq of organoids cultured for 1, 3, and 6 months. Insets, scATAC-seq (blue) and scRNA-seq (green) data integration.
(D) SHARE-seq of organoids cultured for 23 days, 1, 2, and 3 months. Left, cell types from all the timepoints. Right, organoid stage.
(E) Adjusted mutual information (AMI) scores between cell types and individual organoids for each scRNA-seq and SHARE-seq dataset, where lower scoresindicate lower variability. Dotted lines represent AMI scores of fetal cortex.
(F) AMI scores between cell types and individual organoids in scATAC-seq (green points). Gray points are repeated from Figure 1E. Dotted lines represent the AMI scores of human fetal cortex. “c”, cell line clone; “b” differentiation batch; org., organoid; aRG: apical radial glia; IP: intermediate progenitor; prec., precursors; DL, deep-layer; PN, projection neurons; CFuPN, corticofugal projection neuron; CPN, callosal projection neuron; oRG, outer radial glia; IN, interneurons. See also Figures S1 and S2 and Data S1 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1.
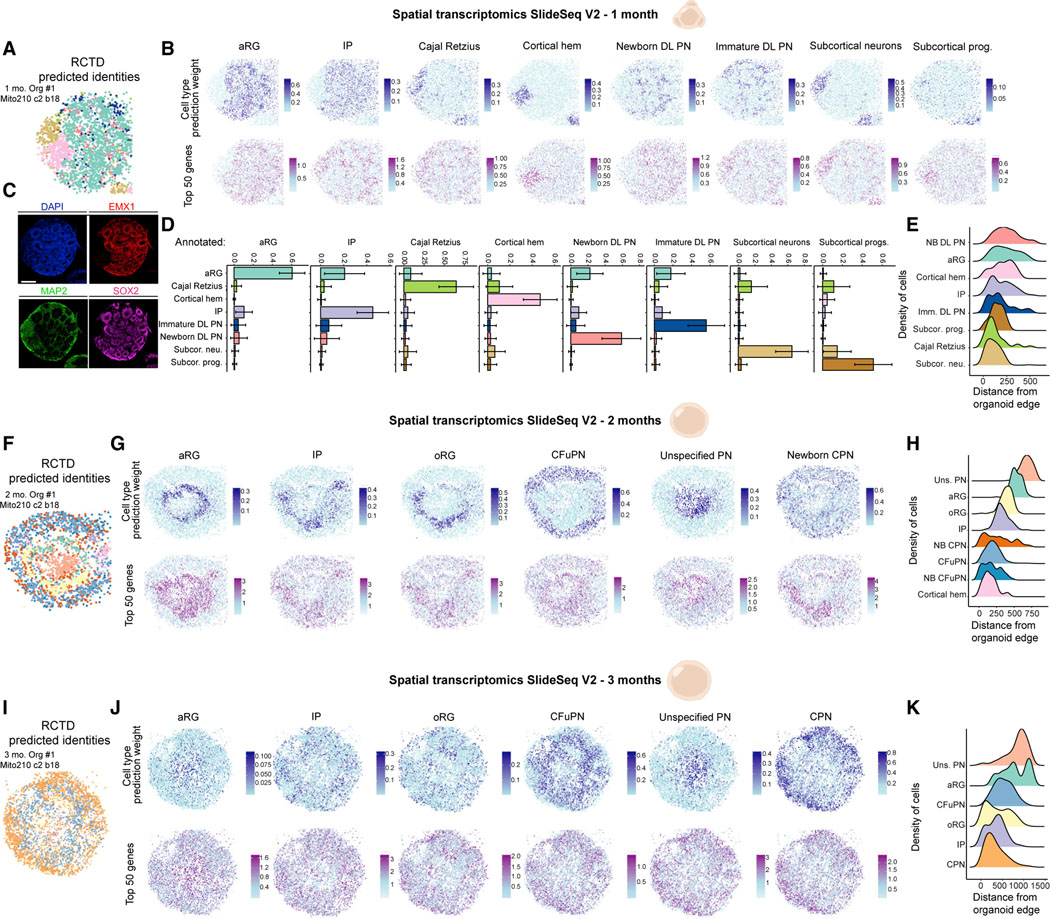
Figure 2. Spatial transcriptomic landscape of developing cortical organoids.
(A) Spatial plot of Slide-seqV2 data from 1-month Organoid #1, colored by RCTD-assigned cell type.
(B) Spatial plots showing RCTD prediction weights (top row) and normalized expression of the top 50 marker genes (bottom row) for each cell type in 1-monthOrganoid #1.
(C) Immunohistochemistry for neuronal (MAP2) and dorsal forebrain progenitor (EMX1, SOX2) types in 1-month Organoid #1, in a section posterior to the one usedfor Slide-seqV2. Scale bar 200 μm.
(D) The mean ± standard deviation of the cell type weights given by RCTD, for beads in 1-month Organoid #1 that were annotated as each cell type.
(E) The distribution of each annotated cell type over the beads’ calculated distance from the edge of the organoid, ordered from top to bottom by median distance.
(F–H) Spatial plots and distributions of cell types from 2-month Organoid #1, as in A, B, and E.
(I–K) Spatial plots of distributions of cell types from 3-month Organoid #1, as in A, B, and E. NB, newborn; Subcor., subcortical; prog., progenitors; neu., neurons; Uns., unspecified. See also Figure S2 and Table S1.
Reproducible cell diversification in organoids over time
At the earliest timepoint characterized (23 days), the organoids contained not only diverse cell types characteristic of the telencephalon, but also a large proportion (~25%–60%) of cells of other regions of the developing nervous system, including putative midbrain, hindbrain, thalamus, neural placode, neural crest, and subcortical interneuron populations (Figures 1A, 1B, S1B, Data S1C and S1K in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). A few non-cortical cells (~8.1%) were still present at 1 month, but from 2 months onwards organoids were exclusively populated by cell types of the cerebral cortex (Figures 1B and S1C–S1I).
Over the course of organoid development, from 1 to 6 months, cellular composition followed differentiation transitions mirroring the order observed in vivo (Figures 1A, 1B, and S1, Data S1 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1) (Angevine and Sidman, 1961; Rakic, 1974; Shen et al., 2006; Yue Huang et al., 2020).
Of interest, at 2 months we also observed emergence of an excitatory neuron population marked by expression of TBR1, NEUROD2, and NEUROD6, but which lacked other canonical markers of subtype identity (labeled here as “unspecified PN”). At 6 months, organoids contained a great abundance of immature interneurons (Figures 1A, 1B, and S1I, Data S1 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1), which were molecularly distinct from the GAD2-positive population observed at 23 days, and, importantly, expressed the telencephalic marker FOXG1, indicating that this later-emerging population has a telencephalic origin. This interneuron population expressed SP8, SCGN, PROX1, and CALB2 (Data S1L in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1), suggesting that these may be olfactory bulb-like interneurons derived from cortical progenitors, as previously described in mice (Kohwi et al., 2007; Young et al., 2007; Fuentealba et al., 2015); alternatively, they may constitute a recently described dorsally-derived interneuron population, which molecularly resembles interneurons derived from the caudal ganglionic eminence (Delgado et al., 2022).
We next assessed if these longitudinal events of cellular production emerged reproducibly across development in all organoids. We calculated the adjusted mutual information (AMI) score for each of the timepoints sampled, measuring the dependence between the proportion of cell types present and the individual organoid of origin. Most replicates, across all ages and cell lines, had AMI scores comparable to those for two published endogenous human cortex datasets (Polioudakis et al., 2019; Trevino et al., 2021) and a newly generated fetal dataset (see below, Figures 1E and S1B–S1I).
Congruent and reproducible cell type-specific epigenomic states
To begin to understand the regulatory logic that underlies this cellular diversification, we investigated gene-regulatory changes across organoid development by relating the 49,568 scATAC-seq profiles from 1-, 3-, and 6-month organoids (Figures 1C and S2A–S2H) to the scRNA-seq atlas.
Overall, there was good agreement between cell types as defined by scRNA-seq and scATAC-seq (Figures 1C and S2A–S2F, Table S2A), and AMI scores indicated that epigenetic variability between individual organoids falls into the same low range as transcriptional variability (Figures 1F and S1J–S1M).
To further investigate the relationship between RNA profiles and underlying chromatin organization in organoids, we applied SHARE-seq to profile 42,810 cells at 4 timepoints, from 23 days to 3 months. Because the non-cortical cell types present at 23 days may have more distinct epigenomic profiles that might not be fully captured in the relatively low number of single cells at this timepoint (1,281 scATAC-seq profiles), we also performed bulk ATAC-seq on organoids at 23 days to generate a comprehensive reference peak set for analysis. These data identified the same cell types at each timepoint as the other modalities( Figures 1D and Table S2B) and showed the same high organoid-to-organoid reproducibility (Figure 1E). Genes with a large number of significant peak-gene associations (DORCs; Ma et al., 2020) included several TFs of known importance during neurodevelopment such as ZIC1, NEUROD2, SOX2, and LHX2, indicating high-precision epigenetic control of these genes (Figure S3D).
Spatial organization of cell types during organoid development
To analyze molecular and cellular events in their native spatial context, we performed spatial transcriptomic profiling using Slide-seqV2 at 1, 2, and 3 months (Figures 2 and S2L–S2N). Due to bead size (10 μm), each bead may capture transcripts from multiple adjacent cells. To resolve cell types from the spatial measurements, we applied RCTD (robust cell type decomposition; Cable et al., 2021) and plotted the expression of genes in the scRNA-seq signatures for each cell type (Figures 2 and S2L–S2N, Table S1).
At 1 month, clusters of apical radial glia (aRG) progenitors occupied both central and peripheral regions of the organoid (Figures 2A–2E, S2L, and S2O), with Cajal Retzius cells, intermediate progenitors (IP), newborn deep layer (DL) projection neurons (PN), and immature DL PN superficially located with respect to aRG. Interestingly, cells of the cortical hem and subcortical regions largely did not intermix with cortical cell types. At 2 months, aRG formed a ring of rosette-like structures around a core that consisted primarily of the “unspecified PN” population (Figures 2F–2H, S2M, and S2P). The remaining cortical cell types were located largely superficial to the aRG. Outer radial glia (oRG) were superficially positioned with respect to aRG (Figures 2H and S2P), reflecting the relative location of these cells in endogenous tissue (Fietz et al., 2010; Hansen et al., 2010). By 3 months, there was a loss of recognizable structures such as rosettes (Figures 2I–2K, S2N, and S2Q), consistent with other studies (Eiraku et al., 2008; Velasco et al., 2019a; Qian et al., 2020). The inner core of the organoids was mainly populated by “unspecified PN” and aRG (Figures 2K and S2Q). These data show dynamic changes and preferential positioning of cell types in cortical organoids that reflect what is observed in vivo.
Cortical organoids capture longitudinal fetal programs of cell identity acquisition with cell-type and temporal specificity
Previous work has shown resemblance between cortical organoids and endogenous human fetal tissue (Camp et al., 2015; Velasco et al., 2019a; Bhaduri et al., 2020; Gordon et al., 2021). However, it remains unexplored whether these similarities extend equally to all cell types and across all steps of development. To address this, we first determined gene expression signatures for all cortical cell types at each stage (Figures 3A and 3B, Table S1, Data S2 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1), by analyzing pseudo-bulk profiles using a model that controls for variability between organoids. We confirmed that the most significant differentially expressed genes (DEGs) in these signatures reflected developmental-stage and cell-type-appropriate expression of known endogenous markers (Data S2 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Importantly, the expression of individual cell type signatures was highly reproducible across organoids; there was no statistically significant difference in the means of any cell-type signature gene set across all organoids assayed at each timepoint (Figures 3A and 3B, Data S2 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1; one-way ANOVA, p > 0.05).
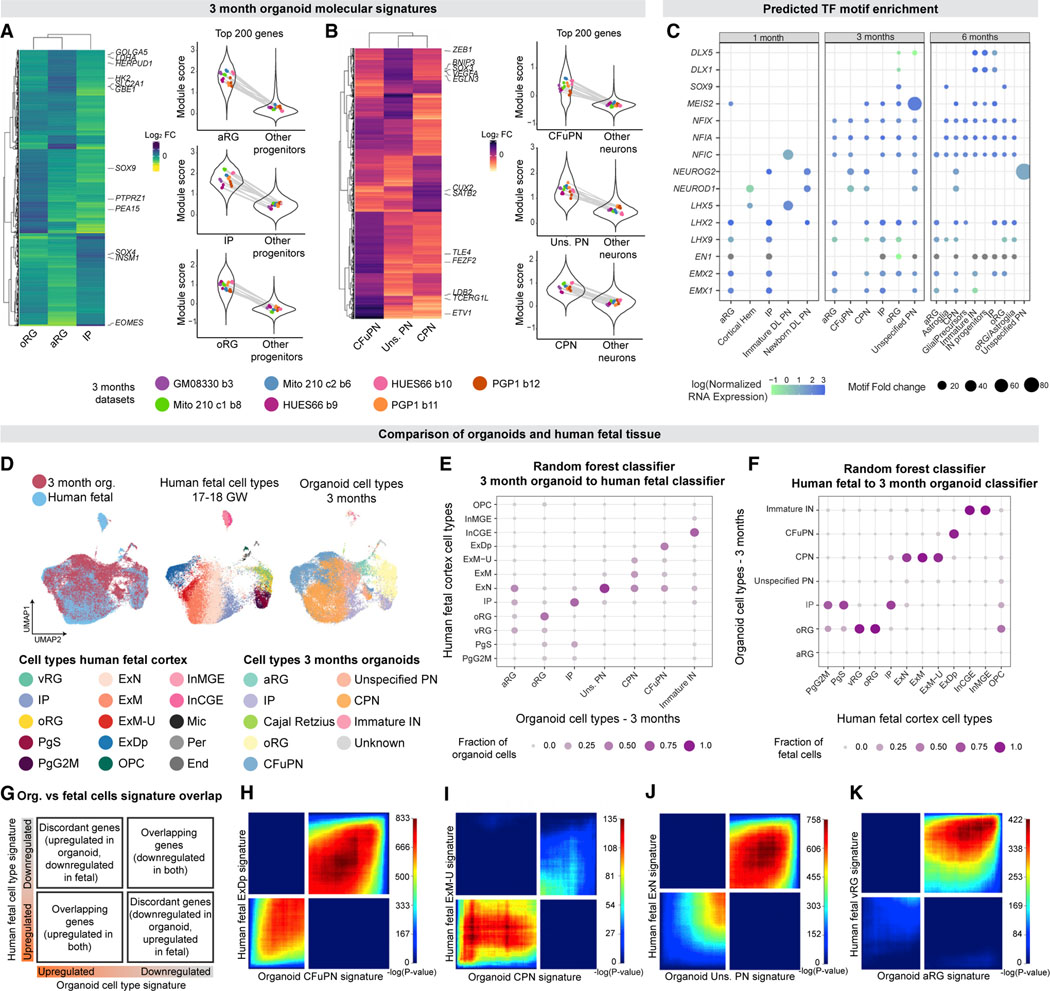
Figure 3. Longitudinal fetal programs of cell identity acquisition are established in cortical organoids with cell type specificity.
(A and B) Left, heatmaps comparing the upregulated genes in each progenitor (A) or neuronal (B) population (genes with adjusted p < 0.0015, log2 fold change>1.5). Right, gene set expression of the top 200 genes in the molecular signatures of each cell type. Violin plots show the module score distribution across single cells; points show the mean module score for each individual organoid, with lines connecting points representing cells from the same organoid. The means within each distribution (within the cell type for that signature, left, or the background cells, right, for each plot) showed no significant difference across individual organoids in all cases (one-way ANOVA, p > 0.05).
(C) TF motifs enriched (p < 1e-10) in peaks with increased accessibility (Bonferroni adjusted p < 0.1) in each cell type, per timepoint. Points are sized by foldchange enrichment of the motif in accessible peaks versus GC-content-matched background regions, and colored by normalized mRNA expression for each TF in the corresponding cell type from scRNA-seq. Gray points indicate zero expression.
(D) Cells from 3-month organoids, integrated with human fetal data (Polioudakis et al., 2019). Left, cells colored by dataset. Middle, fetal cells are colored by cell type as assigned in Polioudakis et al. (2019). Right, organoid cells are colored by cell type.
(E) Assignments of 3-month organoid cells to fetal cell types, via random forest. Points are sized and colored by the fraction of each organoid cell type assigned toeach fetal cell type.
(F) Assignments of fetal cells to 3-month organoid cell types, via random forest. Points are sized and colored by the fraction of each human fetal cell type assignedto each organoid cell type.
(G) Schematic of the RRHO2 plot. Genes from each signature are ordered from most upregulated to most downregulated, with the most upregulated gene of eachsignature in the lower left corner.
(H–K) RRHO2 plots comparing the signatures of organoid CFuPN to fetal ExDp (H), organoid CPN to fetal ExM-U (I), organoid uns. PN to fetal ExN (J), and organoid aRG to fetal vRG (K). Points in the plot are colored by the p value of hypergeometric tests measuring the significance of overlap of gene lists up to that point. Cell types as in Polioudakis et al. (2019): vRG, ventricular radial glia; PgS, progenitors in S phase; PgG2 M, progenitors in G2 M phase; ExN, migrating excitatory neurons; ExM, maturing excitatory neurons; ExM-U, maturing excitatory neurons-upper layer enriched; ExDp, deep layer excitatory neurons; OPC oligodendrocyte precursors; InMGE, interneurons from the medial ganglionic eminence; InCGE, interneurons from the caudal ganglionic eminence; Mic, microglia; Per, pericyte; End, endothelial. See also Tables S1 and S2 and Data S2 and S3 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1.
Next, we examined putative cell type-specific regulatory mechanisms, by identifying transcription factor (TF) motifs enriched in the accessible chromatin regions in each cell type (Heinz et al., 2010) and comparing them to the expression of the cognate TF. We confirmed that known developmental-stage and cell-type appropriate TFs and their motifs were associated with the relevant cell populations (Figure 3C and Table S2C). Finally, we applied the SHARE-seq data to relate the expression of a TF with the chromatin accessibility of its motif within the same cell. The expression of many TFs was positively correlated with accessibility of their motifs across the genome (Figure S2I), suggesting that they are bound at these sites.
We next sought to establish how closely the molecular signatures identifying cortical cell types in organoids corresponded to those defining cortical cell types in the endogenous fetal cortex. We integrated our organoid data with 91,844 profiles from two published scRNA-seq datasets (Figure 3D and Polioudakis et al., 2019; Trevino et al., 2021) and with 60,806 single-nucleus RNA-seq (snRNA-seq) profiles that we newly generated from human fetal cortex at PCW 14, 15, 16, and 18 (Figures S2J–S2K). We assessed similarity in two ways, both of which indicated high agreement between organoid and fetal cortical cell types. First, we applied a random forest classifier-based approach (Shekhar et al., 2016). Organoid cells at each stage were predominantly assigned to the corresponding endogenous cell types by a classifier trained on the fetal cells, and vice versa (Figures 3E and 3F, Table S4A). In the classifier trained on organoid cells, very few fetal cells were assigned to the aRG and “unspecified PN” organoid cells (Figure 3F), suggesting that these two populations are less closely related to endogenous cell types. Classifiers trained on 2, 3, 4, 5, or 6-month organoid cells yielded similar results (Table S4A), but those trained on 1- or 1.5-month organoid cells showed multiple fetal progenitor cell types assigned to organoid aRG (Table S4A), likely pointing to stage-specific differences in the expression profile of the organoid aRG population.
Second, we applied a pairwise comparison of molecular signatures defined separately for each cortical cell type in organoid and fetal cells using rank-rank hypergeometric overlap (RRHO2; Plaisier et al., 2010; Cahill et al., 2018) (Figure 3G). This method also showed a high agreement of expression signatures between organoid and corresponding endogenous fetal cell types (Figures 3H–3K, Data S3 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
Importantly, we found that the molecular signature of the “unspecified PN” of organoids, while most closely matching fetal migrating ExNs from Polioudakis et al. (2019), GluN 1 and 5 clusters from Trevino et al. (2021), and CPN from our fetal dataset, showed a comparatively weak agreement overall, driven more by shared down-regulated genes (Figure 3J, Data S3 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Similarly, the signature of organoid aRG did not show a large overlap of upregulated genes with any fetal signature at any timepoint from 2 months in vitro onwards (Figure 3K, Data S3 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Thus, although these two cell types showed some overall transcriptional resemblance to endogenous cells (Figure 3E), their transcriptional signatures had comparatively weak matching to cell type-specific fetal signatures from this gestational age range.
In sum, using two independent methods we find that organoid cell types closely transcriptionally resemble cells of the endogenous fetal cortex, with only two notable exceptions: aRG and a type of mis-specified PN that does not seem to exist in vivo.
Similar to the transcriptional analysis, we found good concordance of the epigenomic landscape of cell types in cortical organoids with their endogenous counterparts. We compared scATAC-seq data from 1-, 3-, and 6-month organoids to a recently published scATAC-seq dataset of human cortex at PCW 16–24 (Trevino et al., 2021). Again, corresponding organoid and fetal cell types had highly similar epigenomic signatures (Figures S3A–S3C, Data S3 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1) and identified examples of cell type-specific gene regulatoryelementsconservedbetweenorganoidsandfetaltissue (Figures S3E–S3H). Indeed, when the datasets were analyzed at similar total number of reads, few DARs were found between the fetal and organoid samples (Table S2D). Notably, we found substantial overlap between the peaks called in each dataset, with higher concordance at the older organoid timepoints: 73% of peaks called in 1-month organoids overlap with peaks called in fetal cells, while 88% and 85% of peaks called in 3-month and 6-month organoids, respectively, overlap with peaks called in fetal cells. It is possible that 1-month organoids represent an earlier developmental stage than the PCW 16–24 cortex profiled in the Trevino et al. (2021) dataset.
Overall, our analysis demonstrates that most organoid cell types closely replicate the cell type-specific signatures and epigenetic states of the corresponding human fetal cortical cell types, indicating a high degree of fidelity in cell identity.
Human cortical organoid cell type identity is largely unaffected by metabolic state
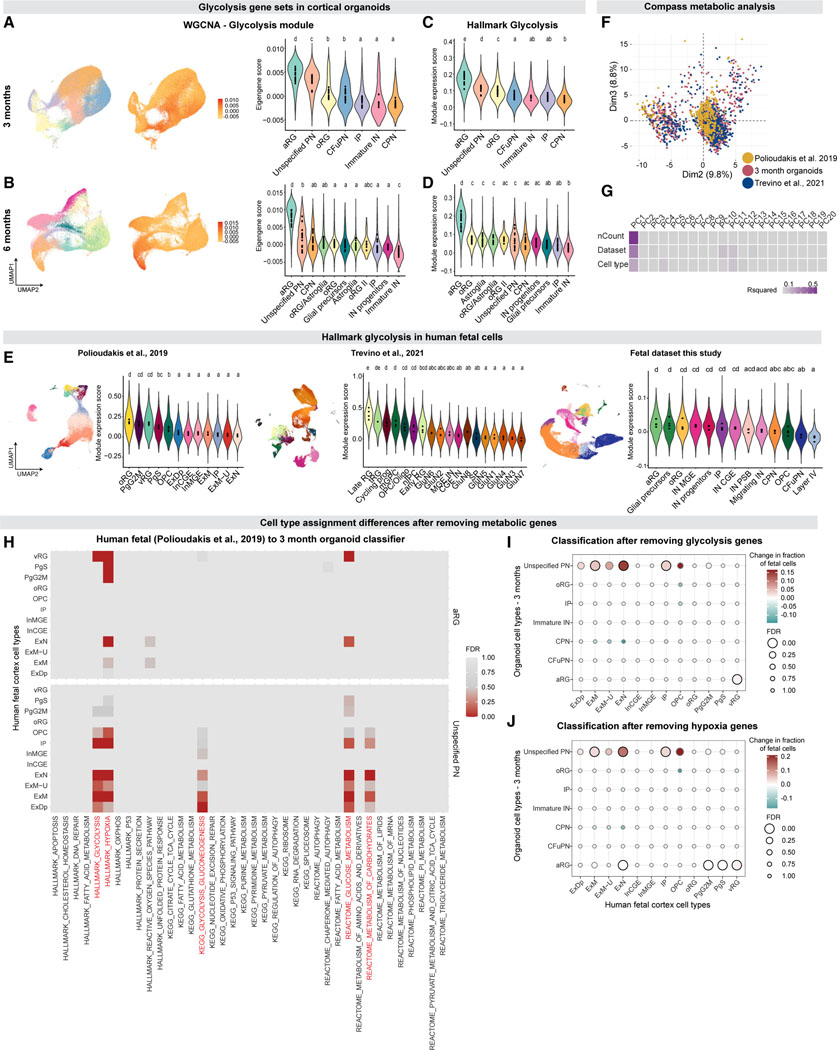
Weighted gene correlation network analysis (WGCNA; Langfelder and Horvath, 2008) (Table S3, Data S4 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1) identified a module including glycolysis-associated genes starting at 1.5 months in vitro, which was specifically enriched in the two populations of cells with the weakest resemblance to fetal cells, namely aRG and “unspecified PN”, from 2 months in vitro onwards (Figures 4A and 4B). This prompted us to examine cell type-specific variation in metabolic processes implicated in organoid biology (Pollen et al., 2019; Bhaduri et al., 2020; Qian et al., 2020; Tanaka et al., 2020) using curated gene sets from MsigDB (Liberzon et al., 2015). “Unspecified PN” and aRG, along with other progenitor types, were enriched for the glycolysis and hypoxia gene sets at multiple stages of organoid development (Figures 4C and 4D, Data S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). This is comparable, however, to an enrichment of these gene sets observed in a subset of endogenous human fetal progenitors (Trevino et al., 2021; Polioudakis et al., 2019; and this study; Figure 4E, Data S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Thus, metabolic states observed during normal development of cortical progenitors are also observed in the corresponding cell types in organoids. Upregulation of metabolic states in organoid cells was also detected on an epigenetic level in the SHARE-seq data; significantly upregulated chromatin regions in cells with high hypoxia module scores were found to be associated with 249 genes, which were enriched for metabolic GO processes including “response to hypoxia” (FDR = 0.02), as well as for neurodevelopmental terms including “forebrain development” (FDR = 0.05, Table S2E).
Figure 4. Human cortical organoid cell type identity is largely unaffected by metabolic state.
(A and B) Enrichment of a WGCNA gene module containing glycolysis genes in 3- and 6-month organoids. Left, cells from all 3-month (A) and 6-month
(B) organoids, downsampled to an equal number of cells per organoid. Cells are colored according to cell type (left) and to eigengene score for the “glycolysis” module (middle). Right, violin plots showing the distribution of eigengene scores across cell types. Points indicate average scores for each individual organoid. Letters above violin plots indicate the results of a one-way ANOVA followed by pairwise TukeyHSD; cell types with the same letter have no significant difference between organoid averages.
(C and D) Violin plots showing the distribution of module scores for the MSigDB Hallmark Glycolysis gene set across cell types in 3-month (C) and 6-month (D) organoids, in the same downsampled data. Points and letters as in (A).
(E) Fetal cells (left, Polioudakis et al., 2019, middle, Trevino et al., 2021, right, fetal dataset from this study) colored by their assigned cell type (top), and violin plot showing the distribution of module scores for the MSigDB Hallmark Glycolysis gene set across cell types (bottom). Points and letters as in (A).
(F) Principal component analysis (PCA) of the Compass matrix of metabolic reaction potential activity scores. Cells are colored by dataset.
(G) Correlation of the top 20 principal components (PCs) of the Compass PCA with nCount (the number of UMIs per cell), the dataset of each cell, and the cell typelabel of each cell. Tiles colored by R-squared values from linear regressions of the PC loadings with each metadata value.
(H) Significance (false discovery rate by Benjamini-Yekutieli) of the increase in Polioudakis et al. (2019) fetal cell assignments to the 3-month organoid aRG label (top) and unspecified PN cell types (bottom) after individually removing each of the 38 gene sets (x axis) from the model.
(I and J) Change in the assignments of Polioudakis et al. (2019) fetal cell types to 3-month organoid cell types, after removing the MSigDB Hallmark Glycolysis
(I) and Hypoxia (J) gene sets from the model. Points are colored by the change in the fraction of cells assigned to each label by the reduced model, compared to the full model. Points are sized by the FDR of the increase in that fraction. Points with FDR<0.05 are outlined in black. Changes were considered significant if they showed an FDR<0.05 and changed at least 1% of assigned cells. Cell types as in Trevino et al. (2021): tRG: truncated radial glia; mGPC, multipotent glial progenitor cell; OPC/Oligo, oligodendrocyte progenitor cell/oligodendrocyte; nIPC, neuronal intermediate progenitor cell; GluN, glutamatergic neuron; CGE IN, caudal ganglionic eminence interneuron; MGE IN, medial ganglionic eminence interneuron; SP, subplate. See also Figures S3, S4, and S5, Tables S3 and S4, and Datas S4 and S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1.
To understand whether these metabolic differences depended on the culture protocol employed, we profiled 69,333 cells from a different organoid model, whole-brain organoids (Quadrato et al., 2017a), at 3.5 months in vitro, and found a similar specific enrichment of glycolysis and hypoxia gene sets in aRG and unspecified PN cell populations (Figures S3L–S3N, Data S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1), showing that multiple organoid models have similarly restricted differences in metabolic gene expression across similar cell types.
In contrast, while oxidative phosphorylation has been reported to be abnormal in organoids (Bhaduri et al., 2020), the MSigDB oxidative phosphorylation signature did not show strong enrichment in any particular cortical cell type in either of our organoid models or in fetal tissue (Data S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
In sum, neural progenitors, especially aRG, in both organoids and endogenous fetal tissue show an enrichment of glycolysis and hypoxia genes, possibly due to the previously reported energetic requirements of these cells (Khacho et al., 2016; Zheng et al., 2016; Namba et al., 2020).
Broader examination of the degree to which organoids may differ from endogenous fetal cells in expression of metabolic genes showed only limited differences across all cell types. We applied three complementary approaches: Compass, which uses flux balance analysis to model the metabolic state of single cells from scRNA-seq data (Wagner et al., 2021); RRHO2, to identify processes enriched among the genes up-regulated in organoids but downregulated in fetal cells; and differential expression across a panel of MSigDB metabolic gene sets. Across all three analyses, organoids and fetal cells showed similar results, with only a few metabolic pathways, in particular glycolysis and oxidative phosphorylation, being enriched in organoids (Figures 4F–4G, S3I–S3K, and S4). Notably, Compass showed that organoid and endogenous fetal cells clustered together well and could not be discriminated based on their metabolic fluxes (Figures 4F–4G and S3I–S3J). The first principal component (PC1) correlated strongly with cell complexity as previously reported (Wagner et al., 2021); however, none of the top 20 principal components (PCs) captured differences between data sets (Figure 4G), indicating that organoid and endogenous cells cluster together well based on their overall metabolic flux.
Finally, we leveraged the RRHO2 analysis to identify processes enriched among the genes up-regulated in organoids relative to fetal cells. In GO analysis, synapse- and cell cycle-related terms were the most prominent biological processes found to differ between organoid and fetal cells (Figure S4C), possibly indicating differences in the maturation of the cells being compared. Among the 38 MSigDB metabolic gene sets, only a few pathways, including oxidative phosphorylation and lipid metabolism, ranked high in organoids and low in human datasets (Figure S4D).
It is possible that the enrichment of metabolic genes in the aRG and “unspecified PN” populations could be interfering with the alignment of organoid cell types to their endogenous counterparts, as has been suggested for other cortical organoid models (Bhaduri et al., 2020). We therefore systematically removed each of the 38 MSigDB metabolic gene from the variable genes used to train the random forest cell-type classifiers and assessed the effect on the resulting cell classifications. For the vast majority of gene sets, removing the set did not significantly change the number of cells assigned to each cell type (Figures 4H–4J and S5A–S5C, Table S4B). Only two categories of metabolic processes affected classification: in the classifier trained with 3-month organoid cells, removing gene sets related to glucose metabolism and hypoxia significantly increased the number of fetal cells assigned to the “unspecified PN” and aRG cell types (FDR <0.0001, Figures 4H–4J). No other cell type showed an increase in assignments. Importantly, genes related to other fundamental subcellular processes such as apoptosis and oxidative phosphorylation did not significantly change the assignment of any fetal cell type to organoid cell types (Figures 4H and S5, Table S4B).
We then leveraged our spatial transcriptomics developmental atlas to investigate whether there was an association between topographical location and metabolic state of cortical organoid cells. The inner regions of both 2- and 3-month organoids were predominantly composed of aRG and unspecified PN (Figures 2F–2K, 5A, S2M, S2N, S2P, and S2Q), the two populations whose metabolic state affected assignment of cell identity. Notably, aRG became more centrally located between 1 and 2 months, consistent with our finding that aRG identity becomes affected starting at 1.5 months (Data S3 and S5 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
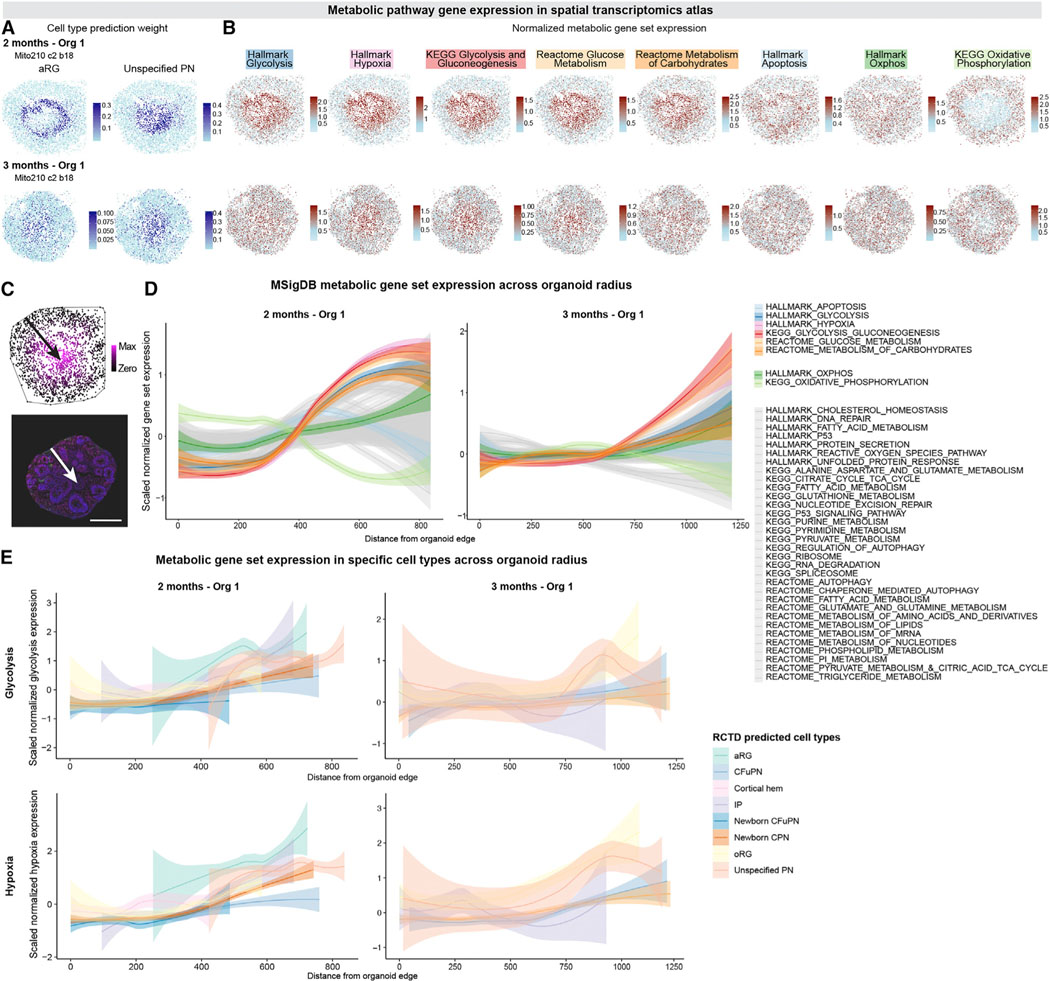
Figure 5. Metabolically-compromised cells reside in a restricted and central region of human cortical organoids.
(A) Spatial plots colored by RCTD prediction weights for aRG (left) and uns. PN (right), for 2-month Organoid #1 (top) and 3-month Organoid #1 (bottom), repeated from Figures 3G and 3J.
(B) Spatial plots showing the summed, normalized expression of genes in the indicated MSigDB gene sets for 2-month Organoid #1 (top) and 3-month Organoid #1 (bottom).
(C) Schematic showing how bead distance from the edge of the organoid was calculated. Top, Slide-seqV2 data from 2-month Organoid #1, with beads colored by distance from organoid edge. Line shows convex hull around organoid. Bottom, IHC of a 1.5-month organoid, showing DAPI (blue), CTIP2 (magenta), HOPX (green), and SOX2 (red). Scale bar 200 μm. Arrows point from edge of organoid to points most distant from the edge.
(D) Distribution of the beads’ scaled expression for each gene set compared to their distance from the edge of the organoid. Solid line shows the smoothed conditional mean values across beads; colored band, 95% confidence interval. Distributions are shown separately for 2-month Organoid #1 (left) and 3-month Organoid #1 (right).
(E) Distributions as in C for the Hallmark Glycolysis (top) and Hallmark Hypoxia (bottom) gene sets, for beads assigned to each cell type. Cell types assigned to more than 8 beads per organoid are shown. Distributions are shown for 2-month Organoid #1 (left) and 3-month Organoid #1 (right). See also Figure S6 and Data S6 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1.
Moreover, while the expression of most of the 38 MSigDB metabolic pathway gene sets was constant across the organoid diameter, pathways associated with hypoxia and glycolysis were enriched in cells toward the center of the organoid (Figures 5C and 5D and S6A–S6C). Notably, cells located in the inner regions of the organoids showed higher expression of glycolysis and hypoxia genes, independent of cell identity (Figures 5E and S6D–S6F). We confirmed these results with the hypoxia detection reagent pimonidazole and by immunohistochemistry for markers of cellular stress, which also showed signal restricted to the organoid core from 2 months in vitro onwards (Figures S6G–S6L, Data S6 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
Overall, our analysis shows that specification of the vast majority of the cortical cell types generated in the present organoid model is not affected by diverging metabolic states, which impact the identity of only two cell types (aRG and “unspecified PN”), accounting for only 3 to 15% of organoid cells at 2–6 months (Figures 1B and S1E–S1I). Notably, affected cells are restricted to the center of the organoid. Furthermore, only glycolysis and hypoxia affected assignment in these two cell types. The data point at cell type- and metabolic pathway-specific effects that are restricted to cells in the organoid core, rather than broad metabolic susceptibility of cells generated in organoids compared to endogenous cortical tissue.
Molecular logic of development of individual human cortical cell types
Our comprehensive, multi-modal atlas of organoid development across time shows that proper cell diversity can be established in organoids and that cell identity is largely not affected by metabolic state. We therefore sought to apply this molecular map of organoid development to understand how cellular lineages are established in the human cortex and the transcriptional events associated with lineage decisions. We inferred developmental trajectories using the 459,711 cortical cells (excluding unspecified PN). To highlight relationships between cell types across time, we connected cell clusters from the same and consecutive timepoints by transcriptional similarity, while preserving the known temporal stage of the cells (Ko et al., 2020) (Figure 6A, Data S7A in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
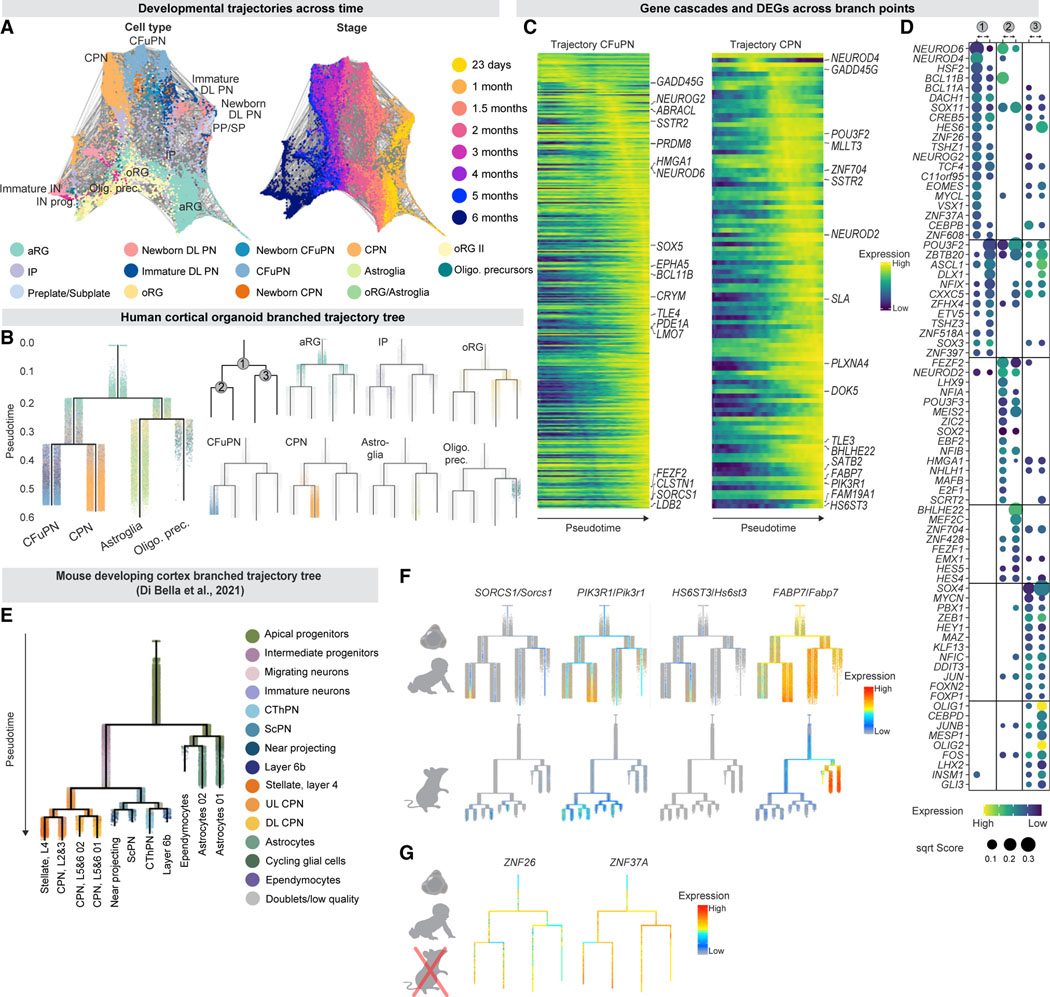
Figure 6. Molecular development of human cortical cell types.
(A) Integrated trajectory of scRNA-seq datasets across time. Force-directed graph created using FLOWMAP. Cell clusters are colored by majority cell type (left, colors as in Figure 1) and by timepoint (right). Gray lines connect cell clusters with high transcriptional similarity.
(B) URD branching tree. Cells colored by identity. Branch points are labeled 1–3.
(C) Heatmap of lineage-specific gene cascades for CFuPN and CPN. Gene expression in each row is scaled to maximum observed expression. Genes are ordered by their onset and peak expression timings.
(D) Top 20 TF per branch. Points are sized by their score (feature importance in a gradient boosting classifier) and colored by their average expression in the corresponding cells (row-scaled). Branch points numbered as in B.
(E) URD branching tree of the mouse developing cortex (embryonic day 10.5 to postnatal day 4) adapted from (Di Bella et al., 2021).
(F) Expression of human lineage-specific genes in human cortical organoid tree (top) and developing mouse cortex tree (bottom, Di Bella et al., 2021).
(G) Expression of two human ZNF genes in the simplified branching tree. See also Figure S7, Table S5 and Data S7.
When visualized as a force-directed graph, this analysis showed lineage relationships reflecting those expected in endogenous tissue. aRG were connected to other progenitor types (IP, oRG), as well as to different types of excitatory neurons, both early- (preplate/subplate, newborn DL PN, immature DL PN) and late-produced (CFuPN, CPN), consistent with the existence of both direct and indirect (i.e., through other progenitor cell types) neurogenesis in organoids. At 5 and 6 months, aRG were also connected to astroglia, oligodendrocyte precursors, and interneurons. The generation of these cell types occurs during perinatal (late) stages of cortical development in vivo (Kriegstein and Alvarez-Buylla, 2009; Obernier and Alvarez-Buylla, 2019). The majority of IP were found within the excitatory neuron lineage (66% of first-neighbor clusters to IP clusters contain majority excitatory projection neurons or other IP, whereas only 33% of all clusters belong to those types), in agreement with the commitment of this progenitor cell type to glutamatergic neuron production in vivo (Hevner, 2019). oRG were connected to postmitotic neurons, consistent with the neurogenic potential of this progenitor cell type (Fietz et al., 2010; Hansen et al., 2010). We also observed a connection between oRG and IP (2.7% of IP neighbors are oRG), suggesting that oRG can give rise to IP in organoids, as described in the endogenous human and non-human primate cortex (Betizeau et al., 2013; Coquand et al., 2022).
Interestingly, at late timepoints, oRG were connected to both neuronal and glial progenitors (Figure 6A). Although oRGs were initially described as neurogenic progenitors (Fietz et al., 2010; Hansen et al., 2010), recent work in vivo has pointed to their possible contribution to astrocyte and oligodendrocyte production in the human and non-human primate brain (Rash et al., 2019; Yue Huang et al., 2020). To further explore the extent to which organoids can model the oRG-to-oligodendrocyte lineage, we searched for a recently-described pre-oligodendrocyte precursor cell (pre-OPC) present in human endogenous cortex at late gestational stages and characterized by the co-expression of oRG (HES1, HOPX, MOXD1) and early oligodendrocyte lineage markers (EGFR, PDGFRA, OLIG1, OLIG2). At 5 and 6 months in vitro, but not at earlier stages, a population expressing these markers was indeed present in organoids (Data S7B–S7C in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). These data support initial findings in the field that human oRG contribute to glial lineages in addition to excitatory neurons during human corticogenesis.
To further investigate how progenitors’ transcriptional profile changes as they develop, we examined differential gene expression in each cell type over time. Overall, we found that progenitors acquire an expression profile reflective of the progeny they produce (Data S7D–S7E in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1 ); the scATAC-seq data showed a similar pattern (Data S7F in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1).
As a complementary approach allowing finer resolution of developmental stage within timepoints, we applied the trajectory-inference algorithm URD to generate a branched trajectory tree based on the transcriptional similarity of pseudotimeordered cells (Figures 6B and S7A–S7C) (Farrell et al., 2018). We defined the root as aRG present at the earliest stage profiled, 23 days in vitro, and the tips as the terminal cell types of each cortical lineage. The order established in the tree reflects the cells’ stage and differentiation status, with basal progenitors following aRG, and then diverging into neuronal (CFuPN, CPN) and glial (astroglia, oligodendrocyte precursors) lineages.
We used the tree to map dynamic expression changes over development (Figures 6C, 6D, S7C, and S7D, Table S5) and to identify lineage-specific genes associated with human cortical neuron subtypes. To this end, we compared this human cortical tree to a published URD tree for mouse developing cortex (Di Bella et al., 2021, Figure 6E). This analysis revealed lineage-specific genes in the human organoid tree that showed a different expression pattern in the mouse tree (Figure 6F). For instance, SORCS1 is associated with the CFuPN lineage in the human organoid tree, but expressed across multiple cortical projection neurons in the mouse. Similarly, PIK3R1 is associated with the CPN lineage in humans, but more broadly expressed in mouse. Other genes associated with human projection neuron lineages are not expressed in the mouse developing cortex (e.g., HS6ST3 for CPN) or are expressed in different cells (e.g., FABP7, associated with the CPN lineage in humans, is exclusively associated with progenitors and glial cells in mouse). We verified these species- and cell type-specific expression patterns using the human endogenous fetal datasets (Figure S7E), and in two additional datasets of the mouse developing cortex (Yuzwa et al., 2017; Telley et al., 2019).
To extract putative molecular programs associated with human cell fate decisions, we identified the top genes and TFs associated with each lineage bifurcation in the human organoid URD tree (Figures 6D and S7F). To distinguish between shared mammalian and human-specific programs governing these bifurcations, we compared the TFs associated with homologous bifurcations in the mouse. Multiple TFs not previously associated with neurogenesis and/or gliogenesis were associated with key branches in the developing human cortex tree, including DACH1 and TSHZ1 in the neuronal branch; CXXC5 and ZFHX4 in the glial branch, and ZNF704 for the CPN branch (Figure S7G). Although orthologs of all of these genes were expressed in the mouse developing cortex, some had distinctive expression in the human vs. mouse tree (ZNF704/Zfp704, Figure S7G), suggesting divergent species-specific functions. Notably, some candidate human regulators predicted to have a role in neuronal commitment in humans had no 1:1 mouse ortholog (Ruan et al., 2008), including ZNF26 and ZNF37A (Figure 6G).
To further elucidate the underlying regulatory mechanisms, we leveraged the SHARE-seq map. We projected the chromatin profiles in the SHARE-seq data onto an URD tree built from the gene expression profiles from the same cells (Figure S7H–S7J) and identified TFs showing concordant motif enrichment and expression for each branch (Figure S7K). The two branching points obtained in this new tree closely resembled branch 1 and 2 of the RNA-only tree, and the gene programs associated with each segment in the RNA-only tree were enriched in the expected segments of the SHARE-seq tree (Figures S7N and S7L). The motif for ZNF264, which has no mouse ortholog (Ruan et al., 2008), was enriched in accessible chromatin in cells in the early steps of the CFuPN lineage. We confirmed the expression of this gene in projection neurons from three human fetal datasets (Polioudakis et al., 2019; Trevino et al., 2021; this study). This suggests a species-specific role for ZNF264 in human projection neuron development, and, more broadly, highlights newly emerging human-specific regulatory processes of neurodevelopment in the cerebral cortex.
Human callosal projection neuron diversity emerges during embryonic development
Recent work has described a higher gene-expression heterogeneity in the CPN population in the adult human cortex compared to mouse, suggesting that the expansion of this neuronal population in humans has been accompanied by increased diversification (Berg et al., 2021). However, how and when these different CPN types arise remains unknown.
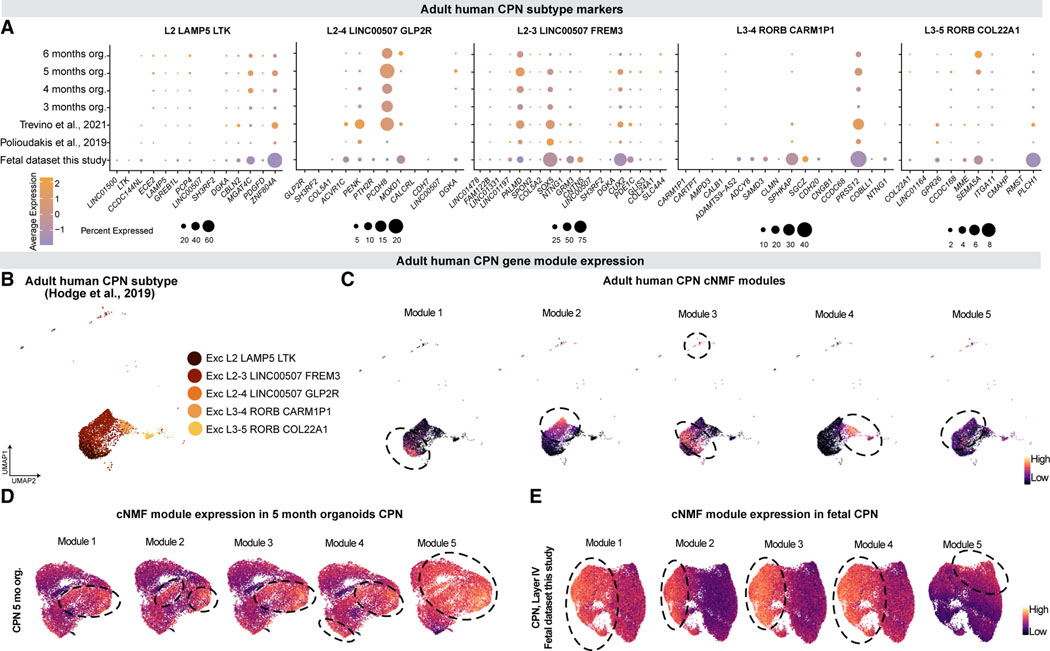
We examined the terminal neuronal cell types of our tree to investigate if the CPN gene-expression heterogeneity described in adulthood (Hodge et al., 2019; Berg et al., 2021) was already detectable during early stages of development. Marker genes of the five different CPN types (Hodge et al., 2019) were expressed from early stages of development in both fetal and organoid CPN; markers highly expressed in fetal CPN were also highly expressed in organoids and vice versa (Figure 7A). To assess if the molecular diversity found in adulthood could also be observed in early CPN, we identified five co-varying gene programs (modules) in adult CPN (Table S6), broadly representing each adult CPN type (Figures 7B and 7C, DataS8A-B inMendeleyData: https://doi.org/10.17632/7cxccpv4hg.1), and scored them in fetal and organoid CPN. All five modules were expressed in fetal and organoid CPN. While their expression did not subdivide developing CPN into five subpopulations as seen in the adult, they began to separate the early developing CPN, with module five showing the most distinctive pattern (Figures 7D and 7E, Data S8C–S8I in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1). Therefore, although the full expression diversity of human adult CPN is not yet present, the data points at CPN divergence as a process that starts at early stages of development.
Figure 7. Callosal projection neuron diversity emerges at early stages of development.
(A) Dotplot showing the percentage of cells (dot size) and average expression (color) of adult CPN subtype markers (Hodge et al., 2019) in human organoid and fetal CPN.
(B) The five transcriptional subtypes of adult CPN described in (Hodge et al., 2019; Berg et al., 2021).
(C) Feature plot of adult CPN showing the expression of consensus cNMF gene modules. Each cNMF module corresponds to a CPN transcriptional subtype.
Dotted circles show areas of high gene module expression.
(D and E) Five-month organoid CPN (D) and human fetal CPN (fetal dataset from this study, E) generated using the top 100 genes used in each of the cNMF modules as variable features. Colored by the module scores of each cNMF module. Dotted circles show areas of high gene module expression. See also Table S6 and Data S8 in Mendeley Data: https://doi.org/10.17632/7cxccpv4hg.1.
DISCUSSION
Understanding the mechanisms that govern the generation of cell diversity and overall development of the human brain are major unmet goals, largely impeded by the ethical and practical limitations associated with studying the fetal human brain over extended periods of in utero development. We find that cortical organoids display high levels of developmental fidelity and reproducibility across longitudinal processes of cellular diversification, displaying consistent acquisition of cell identity in each organoid, implemented through reliable molecular programs, irrespective of metabolic state.
Previous work has indicated that the growth environment of organoids can negatively affect their metabolic state (Pollen et al., 2019; Bhaduri et al., 2020). Our work now shows that it is indeed possible to consistently generate accurate cell identities independent of metabolic state in organoids, under specific culture conditions. This said, it is important to note that the identities of two cell types (aRG and “unspecified PN”) were selectively affected by glucose metabolism and hypoxia. This may be related to the localization of these cell types disproportionately towards the more hypoxic center of the organoids. However, the involvement of glucose metabolism is notable, as several prior studies have reported that neuronal progenitors have a predominantly glycolytic metabolism during normal brain development (Namba et al., 2020), and that the transition to a more differentiated state (i.e., from neural progenitor to neurons) is accompanied in vivo by a metabolic switch from glycolysis towards mitochondrial oxidative phosphorylation (Knobloch and Jessberger, 2017; Khacho et al., 2019). In agreement, we find that endogenous fetal progenitors also show an enrichment of glycolytic genes relative to other fetal cell types, suggesting that progenitors in organoids may exhibit an exacerbated alteration of metabolic pathways that are normally more mildly enriched in these same cells during endogenous developmental process in the embryo.
During evolution, callosal projection neurons of the cortical upper layers have undergone a remarkable expansion and diversification in the human cortex relative to other species (Sousa et al., 2017; Berg et al., 2021). Here, we discover that the transcriptional diversity observed for this neuronal population in the adult human cortex (Hodge et al., 2019; Berg et al., 2021) begins to emerge during early stages of cortical development in both organoids and fetal tissue, suggesting that this diversity reflects specific developmental events rather than depending on later factors such as neuronal activity. This finding points at developing organoids as systems to explore the mechanistic underpinning of this critical, human-specific process, and further reinforces the value of organoids as experimental models to investigate otherwise-inaccessible mechanisms of human cortical development, evolution, and disease.
Limitations of the study
There are currently no single-cell resolution molecular datasets of human fetal cortex covering the entire span of cortical development. This limits the power of our study to compare development in organoids to human development in vivo. More complete human endogenous datasets will enable more comprehensive analysis, both to validate the fidelity of organoid development and to enable more accurate matching of developmental timepoints across in vivo and in vitro development. We identified genes with predicted human-specific roles in fate specification. In future work, it will be important to experimentally validate the functional roles of these genes. In addition, these genes were identified in a mouse-human comparison, so to understand the extent to which the roles of these genes are specific to the human lineage, it would be important to evaluate their expression and functions in cortical cells from species more closely related to humans, such as non-human primates. We found evidence of early transcriptional diversity in CPN in both human cortical organoids and endogenous fetal datasets. However, this diversity did not neatly correspond to the CPN transcriptional states reported in the adult human cortex. Further experimentation is required to link the transcriptional variation seen in early CPN to the full diversity seen in adulthood. Finally, in this study we assess fidelity of cell identity and the effects of metabolic state only through transcriptional profile. Metabolic stressors such as hypoxia could potentially affect other cellular features, such as fate potential, that would not emerge from our analyses.
STAR★METHODS
RESOURCE AVAILABILITY
Lead contact
Further information and requests for resources and reagents should be directed to and will be fulfilled by the lead contact, Paola Arlotta (paola_arlotta@harvard.edu).
Materials availability
Cell lines used in this study are available from their respective repositories (see Experimental Model & Subject Details), except for Mito210. There are restrictions to the availability of the Mito210 cell line due to Materials Transfer Agreement conditions, requests will be handled by the lead contact. This study did not generate new unique reagents.
Data and code availability
Read-level data from scRNA-seq, scATAC-seq, SHARE-seq, and Slide-seq data, supporting the findings of this study have been deposited in a controlled access repository at https://www.synapse.org with accession number project ID syn26346373, while count-level data and meta-data have been deposited at the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1756).
Supplemental data related to Figures 1, 3, 4, 5, 6, and 7 has been deposited in Mendeley Data at https://doi.org/10.17632/7cxccpv4hg.1.
Code used during data analysis is available at https://github.com/AmandaKedaigle/OrganoidAtlas. d Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Data from previous publications that was used in this study can be found at the Gene Expression Omnibus under GEO GSE129519, at http://solo.bmap.ucla.edu/shiny/webapp/, and at https://github.com/GreenleafLab/brainchromatin.
EXPERIMENTAL MODEL AND SUBJECT DETAILS
Pluripotent stem cell culture
The female HUES66 embryonic stem cell (ESC) line (Chen et al., 2009) was provided by the Harvard Stem Cell Institute; the male GM08330 iPSC line (aka GM8330–8) was provided by M. Talkowski Lab (MGH) and was originally from the Coriell Institute; the male PGP1 (Personal Genome Project 1) human iPSC line (a.k.a GM23338) was from the laboratory of G. Church (2005); the male 11a human iPSC cell line was from the Harvard Stem Cell Institute; the male Mito210 human iPSC line was provided by B. Cohen Lab (McLean Hospital); and the male H1 parental hESC line (a.k.a. WA01) was purchased from WiCell. Three clones of the Mito210 and PGP1 human iPSC lines were used, and two clones of the H1 hESC line were used. The PGP1 line was authenticated using STR analysis completed by TRIPath (2018); the HUES66 line was authenticated using STR completed by GlobalStem Inc (2008). For authentication of the 11a cell line refer to (Quadrato et al., 2017a). The H1 and GM08330 lines were authenticated using STR analysis completed by WiCell (2021). The Mito210 lines were authenticated by genotyping analysis (Fluidigm FPV5 chip) performed by the Broad Institute Genomics Platform (2017). The GM08330 line has an interstitial duplication in the long (q) arm of chromosome 20. All cell lines tested negative for mycoplasma contamination.
Cell lines were cultured in feeder-free conditions on 1% Geltrex (Gibco) coated cell culture dishes (Corning) using either mTESR1 medium (StemCell Technologies) or mTESR+ medium (StemCell Technologies) with 100 U/mL penicillin and 100 μg/ml streptomycin, at 37°C in 5% CO2. PSC cultures and colonies were dissociated with the Gentle Cell Dissociation Reagent (StemCell Technologies) or ReLeSR (StemCell Technologies). All human PSC cultures were maintained below passage 50, were negative for mycoplasma, and karyotypically normal (analysis performed by WiCell Research Institute). All of the research conducted for this publication was done in accordance with relevant laws and policies and with ethical oversight from the Harvard University Cambridge-area Institutional Review Board and the Harvard Embryonic Stem Cell Research Oversight Committee.
Organoid differentiation
Dorsal organoids protocol
Dorsally patterned forebrain organoids were generated as previously described in (Velasco et al., 2019a, 2019b). Briefly, on day 0, human iPSC or ESC, were dissociated to single cells with Accutase (Gibco), and 9,000 cells per well were reaggregated in ultra-low cell-adhesion 96-well plates with V-bottomed conical wells (sBio PrimeSurface plate; Sumitomo Bakelite) in Cortical Differentiation Medium (CDM) I, containing Glasgow-MEM (Gibco), 20% Knockout Serum Replacement (Gibco), 0.1 mM Minimum Essential Medium non-essential amino acids (MEM-NEAA) (Gibco), 1 mM pyruvate (Gibco), 0.1 mM 2-mercaptoethanol (Gibco), 100 U/mL penicillin, and 100 mg/mL streptomycin (Corning). For the H1 line, Mito210 c3 and PGP1 c3, cells were plated and formed in the same pluripotent medium in which they were maintained for 1 day to better enable embryoid body formation. From day 0–6, ROCK inhibitor Y-27632 (Millipore) was added (final concentration of 20 μM). From day 0–18, Wnt inhibitor IWR1 (Calbiochem) and TGFβ inhibitor SB431542 (Stem Cell Technologies) were added (final concentration of 3 and 5 μM, respectively). From day 18, the aggregates were cultured in ultra-low attachment culture dishes (Corning) under orbital agitation in CDM II, containing DMEM/F12 medium (Gibco), 2 mM Glutamax (Gibco), 1% N2 (Gibco), 1% Chemically Defined Lipid Concentrate (Gibco), 0.25 μg/mL fungizone (Gibco), 100 U/mL penicillin, and 100 μg/mL streptomycin. On day 35, cell aggregates were transferred to spinner-flask bioreactors (Corning) and maintained in CDM III (CDM II supplemented with 10% fetal bovine serum (FBS) (GE-Healthcare), 5 μg/mL heparin (Sigma) and 1% Matrigel (Corning)). From day 70, organoids were cultured in CDM IV (CDM III supplemented with B27 supplement (Gibco) and 2% Matrigel).
Whole brain organoid protocol
Unpatterned brain organoids were generated as previously described (Quadrato et al., 2017a, 2017b). Briefly, on day 0, human iPSC or ESC, were dissociated to single cells with Accutase (Gibco), and 2,500 cells were plated in each well of a 96-well plate and cultured in embryoid body medium as previously described (Lancaster and Knoblich, 2014). On day 6, embryoid bodies were transferred to low attachment, flat-bottom, 24-well plates in 500 μL of intermediate induction medium consisting of DMEM/F12, Knockout Serum Replacement (Gibco), 0.9% FBS GE-Healthcare), 0.7% N2 (Gibco), Glutamax (Gibco), MEM-NEAA (Gibco), and 0.7 μg/mL heparin. On day 8, 500 μL of neural induction medium was added to each well. On day 10, organoids were embedded in Matrigel (Corning), and transferred to cerebral differentiation medium (as described in (Lancaster and Knoblich, 2014)). On day 14, organoids were transferred to spinner flasks. Medium was changed weekly. After 1 month in culture, 14 ng/mL of BDNF (Preprotech, 450–02B) was added to the medium.
Fetal human brain tissue
Human fetal tissue samples were obtained from the Allen Institute for Brain Science in respect of IRB guidelines from Harvard University. Samples processed for single-nuclei RNAseq were obtained from 4 donors (14, 15, 16, and 18 post-conception weeks (PCW)). All of the research conducted for this publication was done in accordance with relevant laws and policies and with ethical oversight from the Harvard University Cambridge-area Institutional Review Board and the Harvard Embryonic Stem Cell Research Oversight Committee.
METHOD DETAILS
Immunohistochemistry
Samples were fixed in 4% paraformaldehyde (PFA) (Electron Microscopy Services). Samples were washed with 1X phosphate buffered saline (PBS) (Gibco), cryoprotected in a 30% sucrose solution, embedded in optimum cutting temperature (OCT) compound (Tissue Tek) and cryosectioned at 12–18 μm thickness. Sections were washed with 0.1% Tween 20 (Sigma) in PBS, blocked for 1 h at room temperature (RT) with 6% donkey serum (Sigma) + 0.3% Triton X-100 (Sigma) in PBS and incubated with primary antibodies overnight diluted with 2.5% donkey serum + 0.1% Triton X-100 in PBS. After washing, sections were incubated at room temperature with secondary antibodies diluted in the same solution as with primary antibodies (1:1000–1:1200; Key Resources Table) for 2 h at room temperature, washed, and incubated with DAPI staining (1:10,000 in PBS +0.1% Tween 20) for 15 min to visualize cell nuclei. Primary antibodies were diluted as specified in the Key Resources Table. The protocol was adapted for immunohistochemistry for SLC16A3, GORASP2, VIMENTIN, HOPX and OLIG2. Briefly, the first wash was followed by incubation of the sections in boiling temperature sodium citrate buffer pH6 (10 mM sodium citrate, 0.05% Tween 20) or 1x IHC Select Citrate Buffer pH 6.0 (Sigma, 21545) for 40 min. Then the sections were extensively washed with 0.1% Tween 20 and the protocol was followed as described above.
KEY RESOURCES TABLE
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
|
| ||
| Antibodies | ||
|
| ||
| Rat anti-CTIP2 1/100 | Abcam | Cat# AB18465; RRID: AB_2064130 |
| Mouse anti-SATB2 1/50 | Abcam | Cat# AB51502; RRID: AB_882455 |
| Chicken anti-MAP2 1/5000 | Abcam | Cat# AB5392; RRID: AB_2138153 |
| Rabbit anti-EMX1 1/50 | Atlas Antibodies | Cat# HPA006421; RRID: AB_1078739 |
| Goat anti-SOX2 1/50 | RB Systems | Cat# AF2018; RRID: AB_355110 |
| Mouse anti-GFAP 1/4000 | Sigma | Cat# G3893; RRID: AB_477010 |
| Rabbit anti-S100B 1/2000 | Abcam | Cat# AB41548; RRID: AB_956280 |
| Rabbit anti-HOPX 1/2500 | Sigma | Cat# HPA 030180; RRID: AB_10603770 |
| Mouse anti-Ki67 1/400 | BD Bioscience | Cat# 550609; RRID: AB_393778 |
| Rabbit anti-SLC16A3 (MCT4) 1/100 | Sigma | Cat# AB3314P; RRID: AB_2286063 |
| Rabbit anti-GORASP2 1/50 | Preoteintech | Cat# 10598-1-AP; RRID: AB_2113473 |
| Chicken anti-VIMENTIN 1/1000 | Millipore | Cat# AB5733; RRID: AB_11212377 |
| Mouse anti-OLIG2 1/100 | Millipore | Cat# AB9610; RRID: AB_570666 |
| Secondary Donkey Alexa Fluor anti-rabbit 647 | Life Technologies | Cat# A31573; RRID: AB_2536183 |
| Secondary Donkey Alexa Fluor anti-rabbit 546 | Life Technologies | Cat# A10040; RRID: AB_2534016 |
| Secondary Donkey Alexa Fluor anti-mouse 546 | Life Technologies | Cat# A10036; RRID: AB_2534012 |
| Secondary Donkey Alexa Fluor anti-mouse 488 | Life Technologies | Cat# A21202; RRID: AB_141607 |
| Secondary Donkey Alexa Fluor anti-goat 647 | Life Technologies | Cat# A21447; RRID: AB_2535864 |
| Secondary Donkey Alexa Fluor anti-rat 488 | Life Technologies | Cat# A21208; RRID: AB_141709 |
| Secondary Donkey Alexa Fluor anti-chicken 488 | Jackson ImmunoResearch Laboratories |
Cat# 703-545-155; RRID: AB_2340375 |
|
| ||
| Biological samples | ||
|
| ||
| Developing human cortex tissue (PCW 14, 15, 16, 18) | Allen Institute for Brain Science | NA |
|
| ||
| Chemicals, peptides, and recombinant proteins | ||
|
| ||
| Geltrex LDEV-Free hESC-Qualified | Life Technologies | A1413301 |
| Rock inhibitor Y-27632 | Millipore | SCM075 |
| IWR-1 | Calbiochem | 681669 |
| SB431542 | Stem Cell Technologies | 72234 |
| Knockout Serum Replacement | Gibco | 10828-028 |
| Non-essential Amino Acids | Gibco | 11140-050 |
| Penicillin/Streptomycin | Corning | 30002CI |
| 2-Mercaptoethanol | Gibco | 21985-023 |
| GlutaMAX | Gibco | 35050-061 |
| StemPro Accutase | Gibco | A1110501 |
| Matrigel | Corning | 356234 |
| B27 Supplement | Gibco | 17504044 |
| Sodium Pyruvate | Gibco | 11360070 |
| N2 Supplement | Gibco | 17502-048 |
| Fungizone | Gibco | 15290018 |
| Heparin | Sigma | H3149 |
| Chemically Defined Lipid Concentrate | Gibco | 11905031 |
| BDNF | Preprotech | 450-02B |
| Paraformaldehyde 16% | Electron Microscopy Services | 15710 |
| Optimum cutting temperature (OCT) compound | Tissue Tek | 4583 |
| Tween-20 | Sigma | P9416 |
| Donkey serum | Sigma | D9663 |
| Triton X-100 | Sigma | T9284 |
| Select Citrate Buffer pH 6.0 | Sigma | 21545 |
| Enzymatics | Sigma | PN-B8667 |
| SUPERase | Qiagen | Y9240L |
| EZ-lysis buffer | Thermo Fisher Scientific | AM2694 |
| Recombinant ribonuclease inhibitor | Takara | 2313B |
| EZ-lysis buffer | Sigma | NUC-101 |
|
| ||
| Critical commercial assays | ||
|
| ||
| Worthington Papain Dissociation System kit | Worthington Biochemical | LK003153 |
| Chromium Single Cell 3′ Library and Gel Bead Kit v3 | 10x Genomics | PN- 1000075 |
| Chromium Single Cell 3′ Library and Gel Bead Kit v2 | 10x Genomics | PN- 120236 |
| Chromium Single Cell ATAC Library And Gel Bead Kit | 10x Genomics | PN- 1000110 |
| Hypoxyprobe kit | Hypoxyprobe | HP1-100kit |
| NucleoSpin Gel and PCR Clean-up | Takara | 740609.250 |
|
| ||
| Deposited data | ||
|
| ||
| Previously published organoid raw sequencing data | Velasco et al. (2019a) | GEO GSE129519 |
| Previously published organoid raw sequencing data | Paulsen et al. (2022) | Synapse syn26346373 |
| Raw sequencing data from organoids | This paper | Synapse syn26346373 |
| Sequencing count data & metadata | This paper | Single Cell Portal SCP1756 |
| Human fetal processed sequencing data | Polioudakis et al. (2019) | Codex http://solo.bmap.ucla.edu/shiny/webapp/ |
| Human fetal processed sequencing data | Trevino et al., 2019 | https://github.com/GreenleafLab/brainchromatin |
| Code used for analysis | This paper | https://github.com/AmandaKedaigle/OrganoidAtlas |
| Supplemental datasets | This paper | https://doi.org/10.17632/7cxccpv4hg.1 |
|
| ||
| Experimental models: Cell lines | ||
|
| ||
| Human HUES66 ESC | Harvard Stem Cell Institute | HSCI HUES 66; NIH Approval Number NIHhESC-10-0057 |
| Human GM08330 iPSC | Coriell Institute | Coriell GM08330 |
| Human PGP1 iPSC | G.Church lab | Coriell GM23338 |
| Human Mito210 iPSC | B. Cohen lab | N/A |
| Human 11a iPSC | Harvard Stem Cell Institute | HSCI 11a |
| H1 (WA01) hESC | WiCell | WiCell, WB34444 |
|
| ||
| Software and algorithms | ||
|
| ||
| ImageJ | Schneider et al. (2012) | https://imagej.nih.gov/ij/ |
| CellRanger | 10x Genomics | https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/what-is-cell-ranger |
| Seurat | Stuart et al. (2019) | https://satijalab.org/seurat/ |
| FLOWMAP | Ko et al. (2020) | https://github.com/zunderlab/FLOWMAP |
| Cytoscape | Cytoscape Consortium | https://cytoscape.org |
| DESeq2 | Love et al. (2014) | https://bioconductor.org/packages/release/bioc/html/DESeq2.html |
| Signac | Stuart et al. (2021) | https://satijalab.org/signac/ |
| Cicero | Pliner et al. (2018) | https://cole-trapnell-lab.github.io/cicero-release/ |
| HOMER | Heinz et al. (2010) | http://homer.ucsd.edu/homer/ |
| Slide-seq pipeline | Macosko Lab | https://github.com/MacoskoLab/slideseq-tools |
| Robust Cell Type Decomposition | Cable et al. (2021) | https://github.com/dmcable/RCTD |
| WGCNA | Langfelder and Horvath (2008) | https://horvath.genetics.ucla.edu/html/CoexpressionNetwork/Rpackages/WGCNA/ |
| MSigDB | Subramanian et al. (2005) | http://www.gsea-msigdb.org/gsea/msigdb/index.jsp |
| RRHO2 | Cahill et al. (2018) | https://github.com/RRHO2/RRHO2 |
| Harmony | Korsunsky et al. (2019) | https://github.com/immunogenomics/harmony |
| Genehancer | Fishilevich et al. (2017) | http://www.genecards.org |
| Scrublet | Wolock et al. (2019) | https://github.com/swolock/scrublet |
| MACS2 | Zhang et al. (2008) | https://github.com/macs3-project/MACS |
| ChromVAR | Schep et al. (2017) | https://www.bioconductor.org/packages/chromVAR |
| GenomicRanges | Lawrence et al. (2013) | https://bioconductor.org/packages/release/bioc/html/GenomicRanges.html |
| Multcomp View | Graves et al. (2019) | https://cran.r-project.org/web/packages/multcompView/index.html |
| Compass | Wagner et al. (2021) | https://yoseflab.github.io/software/compass/ |
| ggplot2 | Wickham (2016) | https://cran.r-project.org/web/packages/ggplot2/index.html |
| aricode | Vinh, 2010 | https://cran.r-project.org/web/packages/aricode/index.html |
| SHARE-seq pipeline | Ma et al. (2020) | https://github.com/masai1116/SHARE-seq-alignment |
| DecontX | Yang et al. (2020) | https://github.com/campbio/celda |
| URD | Farrell et al. (2018) | https://github.com/farrellja/URD |
| Destiny | Angerer et al. (2016) | https://bioconductor.org/packages/release/bioc/html/destiny.html |
| scikit-learn | Pedregosa et al., 2010 | https://scikit-learn.org |
| CIS-BP | Weirauch et al. (2014) | http://cisbp.ccbr.utoronto.ca |
| cNMF | Kotliar et al. (2019) | https://github.com/dylkot/cNMF/ |
Hypoxyprobe
The Hypoxyprobe (100 mg pimonidazole HCl plus 1.0 mL of 4.3.11.3 mouse Mab, Hypoxyprobe kit, HP1–100kit) was added to the medium at a concentration of 100 μM and organoids were incubated with the probe for 1.5 h at 37C and 5% CO2. The organoids were then extensively washed with PBS and fixed in 4% PFA (Electron Microscopy Services) overnight. Immunohistochemistry was performed as described in the above section, including the antigen retrieval step. The primary antibody used at a 1:50 dilution was included in the kit.
Microscopy
Immunofluorescence images were acquired with a Zeiss Axio Imager.Z2 and Lionheart™ FX Automated Microscope (BioTek Instruments). Images were analyzed and processed with the Gen5 (BioTek Instruments) and Zen Blue (Zeiss) image processing software, and ImageJ (Schneider et al., 2012).
Dissociation of brain organoids and scRNA-seq
Individual brain organoids were dissociated into a single-cell suspension using the Worthington Papain Dissociation System kit (Worthington Biochemical). A detailed description of the dissociation protocol is available at Protocol Exchange, with adaptations depending on age and size (Quadrato et al., 2017b; Velasco et al., 2019b). We resuspended dissociated cells in ice-cold PBS containing 0.04% BSA (Sigma, PN-B8667), counted them with Countess II (Thermo Fisher Scientific), and then adjusted the volume to the final concentration of 1,000 cells/μl. We loaded cells onto a Chromium™ Single Cell 3′ Chip (10x Genomics, PN-120236), and processed them through the Chromium Controller to generate single cell GEMs (Gel Beads in Emulsion). scRNA-seq libraries were prepared with the Chromium™ Single Cell 3′ Library and Gel Bead Kit v3 and v3.1 (10x Genomics, PN- 1000075, 1000121), with the exception of a few libraries in the earlier experiments that were prepared with a v2 kit (10x Genomics, PN- 120236). See Table S7 for information on how many cells were estimated to be loaded and what kit was used. We pooled libraries from different samples based on molar concentrations and sequenced them on a NextSeq 500 or NovaSeq 6000 instrument (Illumina) with 28 bases for read 1 (26 bases for v2 libraries), 55 bases for read 2 (57 bases for v2 libraries), and 8 bases for Index 1. If necessary, after the first round of sequencing, we re-pooled libraries based on the actual number of cells in each and re-sequenced with the goal of producing an equal number of reads per cell for each sample.
Single cell RNA-seq data pre-processing
All cortical brain organoid scRNA-seq datasets consisted of three individual organoids, except for the PGP1 c2 batch 17 at 6 months in which, as previously published (Velasco et al., 2019a), one organoid showed a differentiation defect and was excluded. Wholebrain organoid single-cell datasets consisted of four (HUES66) and four (GM8330) individual organoids. ScRNA-seq reads were aligned to the GRCh38 human reference genome and the cell-by-gene count matrices were produced with the Cell Ranger pipeline (10x Genomics, (Satpathy et al., 2019), for version number, see Table S7). Default parameters were used, except for the “−cells” argument. Data was analyzed using the Seurat R package v3.2.2 (Stuart et al., 2019) using R v3.6. Cells expressing a minimum of 500 genes were kept, and UMI counts were normalized for each cell by the total expression, multiplied by 106, and log-transformed.
In the case of GM08330 1-month organoids, cells were demultiplexed using genotype clustering from cells from a different experiment that were sequenced in the same lane. To demultiplex, SNPs were called from Cell Ranger BAM files with the cellSNP tool v0.1.5, and then the vireo function was used with default parameters and n_donor = 2, from the cardelino R library v0.4.0 (Huang et al., 2019) to assign cells to each genotype.
Single cell RNA-seq dimensionality reduction and clustering
For each dataset of organoids, variable genes were found using the “mean.var.plot” method with default parameters, and the ScaleData function was used to regress out variation due to differences in total UMIs per cell and to cell cycle gene expression, following Seurat’s CellCycleScoring method. Principal component analysis (PCA) was performed on the scaled data for the variable genes, and the top 30 PCs were used for downstream analysis. Cells were clustered in PCA space using Seurat’s FindNeighbors on the top 30 PCs, followed by FindClusters with resolution = 1.0, except for the Mito210 c1 23 days data set, in which FindClusters with resolution = 2.0 was used. Cells were visualized by Uniform Manifold Approximation and Projection (UMAP) on the top 30 PCs.
Single cell RNA-seq cluster annotations
For each dataset, upregulated genes in each cluster were identified using the VeniceMarker tool from the Signac package v0.0.7 from BioTuring (https://github.com/bioturing/signac). Cell types were assigned to each cluster by looking at the upregulated genes with lowest Bonferroni adjusted p values. In some cases, clusters were further subclustered to assign identities at higher resolution (See https://github.com/AmandaKedaigle/OrganoidAtlas/blob/master/Subclustering.R for specific clusters that were separated; for cluster numbers, refer to the metadata available for download from the Single Cell Portal).
Adjusted mutual information (Xuan Vinh, Epps and Bailey, 2010) was calculated between cell type assignments and individual organoids with the aricode R package v1.0.0 (Chiquet et al., 2020).
Single cell RNA-seq data integration
For visualization purposes and downstream analyses within each of the timepoints (1, 1.5, 2, 3–3.5, 4, 5 and 5.5–6 months), cortical organoid datasets were merged using Seurat, re-normalized and scaled as above, batch-corrected using Harmony v1.0 with default parameters (Korsunsky et al., 2019), and visualized using UMAP on the first 30 Harmony dimensions.
Single cell ATAC-seq
Prior to processing, organoids were frozen in Recovery cell culture freezing medium (Thermo Fisher Scientific) at −80°C. Nuclei were extracted from 1-, 3-, and 6- month organoids derived from the Mito210 c1 line using two types of procedures, due to their size differences. For the 1-month organoids, nuclei were extracted following a 10x Genomics-provided protocol (Genomics, 2019) to minimize material loss. For the 3- and 6-month organoids, we used a sucrose-based nuclei isolation protocol (Corces et al., 2017) to better remove debris. ScATAC-seq libraries were prepared with the Chromium™ Single Cell ATAC Library & Gel Bead Kit (10x Genomics, PN- 1000110). Approximately 15,300 nuclei were loaded in each channel (to give an estimated recovery of 10,000 nuclei per channel). Libraries were pooled from different samples based on molar concentrations and sequenced with 1% PhiX spike-in on a NextSeq 500 instrument (Illumina) with 33 bases each for read 1 and read 2, 8 bases for Index 1, and 16 bases for Index 2. See Table S7 for sequencing information.
Single cell ATAC-seq data pre-processing
Reads from scATAC-seq libraries were aligned to the GRCh38 human reference genome and the cell-by-peak count matrices were produced with the Cell Ranger ATAC pipeline v1.2.0 (10x Genomics). Default parameters were used, except with ‘−force-cells 5000’ in one organoid (ATAC-Org5) at 3 months. Data were analyzed using the Signac R package v1.1.0 (Stuart et al., 2021) using R v3.6. Annotations from the EnsDb.Hsapiens.v86 genome annotation package (Rainer, 2017) were added to the object. After consideration of the QC metrics recommended as part of Signac, cells were retained that had 1,500–20,000 fragments in peak regions, at least 40% of reads in peaks, less than 5% of reads in blacklisted regions, a nucleosome signal of less than 4, and a TSS Enrichment score greater than 2. Latent semantic indexing (LSI) was performed to reduce the dimensionality of the data (counts were normalized using term frequency inverse document frequency, all features were set as top features, and singular value decomposition (SVD) was performed). The top LSI component was discarded as it correlated strongly with sequencing depth, and components 2–30 were used for downstream analysis.
To overlap accessible regions with annotated regions of the genome, peaks called in each scATAC-seq timepoint were overlapped with promoters and gene bodies (defined using the method in Signac’s GeneActivity function) and with human enhancer regions from Genehancer v4.4 (Fishilevich et al.,”017), using the “overlapsAny” function from the GenomicRanges R package.
Single cell ATAC-seq clustering and cluster annotation
Cells were clustered using Seurat’s FindNeighbors on the top 30 PCs, followed by FindClusters with the smart local moving (SLM) algorithm and otherwise default parameters. Variation in the cells was visualized by UMAP.
Cells were annotated by two parallel methods. In one method, scATAC-seq data were integrated with scRNA-seq data from the corresponding Mito210 c1 dataset for each timepoint, using Seurat’s TransferData to predict cell type labels for the ATAC cells. Separately, differentially accessible regions (DARs) were called per scATAC-seq cluster using FindMarkers with the logistic regression framework, with the number of fragments in peak regions as a latent variable. These DARs were mapped to the closest genes. Top genes per cluster were used to confirm and refine cluster cell type assignments from those based on transferring labels from scRNA-seq.
Adjusted mutual information (Xuan Vinh, Epps and Bailey, 2010) was calculated between cell type assignments and individual organoids with the aricode R package v1.0.0 (Chiquet et al., 2020).
Identification of co-accessible regions from single cell ATAC-seq
To identify co-accessible sites across each scATAC-seq dataset, the Cicero package was employed (Pliner et al., 2018). A Cicero CellDataSet object was created from the Signac object and run_cicero was run with sample_num = 100.
Single cell ATAC-seq motif enrichment
Motif enrichment analysis was performed by first getting a set of differentially accessible peaks per cell type, using FindMarkers as above. For each cell type, peaks with increased accessibility with an adjusted p value (Bonferroni correction) less than 0.1 were then supplied to the HOMER software v4.11.1 (Heinz et al., 2010), using a 300 bp fragment size and masking repeats. The top 5 de novo motifs per cell type found by HOMER, with a p value ≤ 1e-10, are reported in Table S7, along with all TFs whose known binding sites match that motif with a score ≥ 0.59.
SHARE-seq and bulk ATAC-seq
Nuclei were isolated from fresh-frozen organoids in HEPES Lysis Buffer (10 mM HEPES pH 7.3, 10 mM NaCl, 3 mM MgCl2, 0.1% NP-40) for 5 min on ice. Organoids from day 90 were dissociated using spring scissors (Roboz, RS-5650), while all others were dissociated by gentle pipetting. Samples were then diluted in 1 mL HDT Buffer (10 mM HEPES pH 7.3, 10 mM NaCl, 3 mM MgCl2, 0.1% Tween 20, 0.01% digitonin). Nuclei were pelleted at 800xg for 3 min at 4C and resuspended in room temperature HDT Buffer at a concentration of 1 million nuclei/ml. Formaldehyde was added to a 0.2% final concentration and nuclei were fixed for 5 min at RT. Fixation was quenched with glycine (140 mM final concentration), Tris pH 8.0 (50 mM final concentration), and BSA (0.1% final concentration) for 5 min on ice. Nuclei were washed two additional times with HDT Buffer, split into single use aliquots, pelleted and stored at −80C. All buffers were supplemented with Enzymatics (Qiagen, Y9240L) and SUPERase (Thermo Fisher Scientific, AM2694) RNase inhibitors.
SHARE-seq was performed as described in (Ma et al., 2020) with the following modifications: NIDT Buffer (10 mM Tris pH 7.5, 10 mM NaCl, 3 mM MgCl2, 0.1% Tween 20, 0.01% digitonin) was used instead of NIB. For reverse transcription and template switching, 5X Smart-seq3 Buffer (40 mM DTT, 125 mM Tris pH 8.0, 5 mM GTP, 150 mM NaCl, 12.5 mM MgCl2) was used instead of 5X Maxima RT Buffer.
Bulk ATAC seq was performed on day 23 organoids (4 separate organoids, Mito210 cell line, clone 2, batch 20) by isolating nuclei as described above from four fresh-frozen organoids followed by transposition reaction (without fixation) using the same reaction conditions as described above. Following transposition, DNA was purified using NucleoSpin Gel and PCR Clean-up kits (Takara, 740609.250). Libraries were prepared identically to SHARE-seq ATAC libraries.
Both scRNA-seq and scATAC-seq raw sequencing reads were processed as described in (Ma et al., 2020) (pipeline available at https://github.com/masai1116/SHARE-seq-alignment).
SHARE-seq scRNA-seq dimensionality reduction & clustering
Cell barcodes with a minimum of 1,000 UMIs were retained. Two rounds of filtering were performed to computationally remove doublets and aggregated cells. First, the UMI count matrix was normalized to the average number of UMIs detected across all cells. The 5,000 genes with the highest variance were log2+1 transformed and PCA was performed on this set of genes. The top 20 components were used to find the 20 k-nearest neighbors (KNN) and the estimated library size for each cell barcode smoothened across KNN. Cell barcodes with disproportionately high smoothened library size (identified as outlier from the distribution of library sizes per experiment) were identified as “clumps” and computationally removed. Next, UMI counts for the remaining cell barcodes were processed using Scrublet (Wolock et al., 2019) and the top scoring barcodes were identified as “doublets” and removed. The top 20 principal components were used for UMAP visualization, identifying KNN and Louvain clustering.
SHARE-seq scATAC-seq dimensionality reduction and clustering
Fragment files from all single cell sequencing runs were combined and peaks were called using MACS2 (Zhang et al., 2008). For bulk ATAC on day 23 samples, data was processed as described in (Buenrostro et al., 2015). The peak sets were combined and filtered as described in (Lareau et al., 2019). Briefly, +/− 400 bp windows were extended from each peak summit and sorted according to MACS2 significance scores. Starting with the most significant peak window, all other overlapping windows with lower significance were removed. This was repeated until no overlapping windows were left. The resulting peaks were resized to 300 bp and these intervals were used for subsequent analysis.
Reads in peaks were counted using Signac (Stuart et al., 2021) and cell barcodes were filtered for a minimum fraction of reads in peaks of 0.2, and estimated library size of at least 2000 reads. Cell “clumps” were identified similarly to what described for scRNA-seq; topics were identified using cisTopic (Bravo González-Blas et al., 2019) and the top 20 topics were used for finding KNN. Library sizes were smoothened over these KNN and the top barcodes were removed. Cell barcodes identified as “doublets” in the scRNA-seq analysis above were also removed. Topics were identified on the remaining cells and the top 20 were used for UMAP visualization, KNN identification and Louvain clustering.
SHARE-seq motif enrichment
Motif scores were computed on all scATAC cell barcodes, filtered as described above, using chromVAR (Schep et al., 2017). For subsequent analysis, data was then subsetted to cell barcodes shared between the RNA and ATAC datasets. For each TF, the RNA-motif correlation was calculated as the Spearman correlation coefficient across all cells between the TF’s motif scores and the same TF’s gene expression values.
SHARE-seq gene-peak associations and DORC scores
Gene-peak associations were found on the set of all filtered cells shared between the scATAC-seq and scRNA-seq datasets using the method described in (Ma et al., 2020). Briefly, for each gene, peaks within +/− 50 kb of their TSS were considered for possible associations (median 9 peaks per gene). Spearman correlations were calculated between normalized scATAC counts and normalized scRNA UMI counts. A set of 50 background peaks per tested region was selected using chromVAR, controlling for GC content and accessibility across all cells. The mean and standard deviation of this background set was used to calculate a Z-score and p value for the observed association. Significant associations were called as those with p value less than 0.05.
Domains of regulatory chromatin (DORCs) were called as genes with greater than 5 associated peaks and DORC scores were the total number of reads within these associated peaks. For visualization, DORC scores were smoothened across the KNN identified in the total scATAC dataset and normalized RNA expression values were smoothened across KNN identified in the total scRNA-seq dataset. After subsetting to cells shared in both datasets, both DORC scores and RNA expression values were quantile normalized prior to visualization. The residual was defined as the (normalized smoothened DORC score) – (normalized smoothened RNA expression value).
Slide-seqV2
For Slide-seqV2, organoids were embedded in OCT (Tissue Tek) and cryosectioned at 10 μm thickness. Sections were transferred to a custom-made array of densely packed barcoded beads (termed “pucks”) for Slide-seqV2 experiments. Library construction was performed as described (Stickels et al., 2021). Briefly, first-strand synthesis was performed by incubating the puck with tissue sections in the reverse-transcription solution followed by tissue digestion, second-strand synthesis, library amplification, and purification. The Slide-seqV2 libraries were sequenced with a NextSeq500 with an estimated ~100 million reads per puck. See Table S7 for sequencing information.
Slide-seqV2 data pre-processing
Sequencing reads were aligned to the GRCh38 human reference genome and followed by the Slide-seq pipeline (https://github.com/MacoskoLab/slideseq-tools) and as previously described (Stickels et al., 2021). Since organoid sizes were much smaller than the Slide-seq pucks, images were manually cropped to the edges of the organoids and beads outside of the images were excluded.
Slide-seqV2 cell type decomposition
Cell type decomposition was performed by RCTD (Cable et al., 2021). Briefly, cell type profiles learned from scRNA-seq data corresponding to the same timepoints and cell line were used to decompose mixtures from the Slide-seqV2 data. First, for each timepoint, a reference was made by using the age-matched scRNA-seq dataset. Next, the RTCD package fits a statistical model to estimate the mixture and cell type identities at each bead of the Slide-seqV2 data. We restricted our analysis to beads with more than 200 UMIs. Finally, gene set expression of the top 50 genes from each cell type signature, as previously calculated from scRNA-seq data (Table S1), was visualized by plotting 100x the sum of the reads for all genes in the gene set that appeared in the Slide-seqV2 data, normalized by the number of UMI for each bead.
Cell type signature analysis
To calculate lists of genes defining cell subtype identity from scRNA-seq data, we partitioned each dataset into a group containing cortical neurons and progenitors separately: for 1-month organoids, neurons (newborn DL PN and immature DL PN), and progenitors (aRG and IP); for 2-month organoids, excitatory neurons (i.e., CFuPN, CPN, and “unspecified PN”; Newborn CFuPN and CFuPN were merged into one cell type for this analysis), and progenitors (i.e. aRG, IP and oRG); for 3-month organoids, excitatory neurons (i.e., CFuPN, CPN, and “unspecified PN”) and progenitors (i.e., aRG, IP, and oRG); for 4-month organoids, excitatory neurons (i.e., CFuPN, CPN, and “unspecified PN”) and progenitors (i.e., aRG, IP, and oRG); for 5-month organoids, neurons (CPN, “unspecified PN”, and immature IN) and progenitors-glia (aRG, IP, oRG & Astroglia, oligodendrocyte precursors, and IN progenitors); for 6-month organoids, neurons (CPN, “unspecified PN”, and immature IN) and progenitors-glia (aRG, IP, oRG & Astroglia, oligodendrocyte precursors, and IN progenitors), the very few oRG II cells were excluded from the 6-month analysis. For the purpose of plotting signatures of all cell types present in the organoid in Slide-seq data, the 1-month signatures were recalculated including Cajal Retzius, Cortical hem and subcortical cells (Table S1). To examine processes enriched among genes up-regulated in organoids in the RRHO2 analysis, the neuron signatures from (Trevino et al., 2021) were re-calculated, combining all GluN cell types.
Next, in order to control for variation in the datasets, final signatures were calculated using a paired analysis, pairing cell type clusters occurring in the same individual organoids. Reads were summed across cells in each cell type in each organoid in all cell lines. Genes with less than 10 total UMI were excluded, and then DESeq2 (Love et al., 2014) was used to calculate DEGs between each cell type and all other cell types in its group. The DESeq2 design formula was “~ organoid + CellType”, so that DESeq2 found differentially expressed genes between cell types (defined as the cell type of interest vs Other for each analysis), controlling for the effect of the individual organoid.
To examine whether expression of signature genes was consistent across individual organoids, module scores for the top 200 up-regulated genes in each signature were calculated using Seurat’s AddModuleScore function on the same Seurat objects used as input to the DESeq2 method. For each cell type signature, datasets were split into the cell type of interest and remaining cells, and the average module score across each individual organoid was calculated. These averages were then submitted to a one-way ANOVA using the base R function aov, using the formula AverageValue ~ organoid. None of these tests were significant for an overall difference of means (significance reported as Pr(>F) > 0.05).
Analysis of previously published human fetal data
The dataset from (Polioudakis et al., 2019) was used, which contains Drop-seq data for 33,986 cells from human fetal cortex samples obtained at gestational week 17 and 18, and that had been aligned to the Ensembl release 87 Homo sapiens genome. Raw count data and associated metadata were downloaded from the CoDEx viewer (http://solo.bmap.ucla.edu/shiny/webapp/). Count data from all cells were loaded into a Seurat object and normalized using the same methods as described above for organoid data. To integrate with organoid data, objects were merged in Seurat, and batch-corrected using Harmony with default parameters.
The dataset from (Trevino et al., 2021) was also used. This dataset contains 57,868 single-cell transcriptomes and 31,304 single-cell epigenomes (ATAC-seq) from human fetal cortex samples from gestational week 16, 20, 21 and 24, generated using the 10x Genomics Chromium platform and aligned to the Cell Ranger GRCh38 genome. ScRNA-seq count data and metadata were downloaded from the authors’ GitHub (https://github.com/GreenleafLab/brainchromatin/blob/main/links.txt). Count data was normalized and integrated with organoid data as above. ScATAC-seq counts data and metadata were downloaded from GEO (GSE162170), and fragments files were downloaded from the authors’ GitHub. Cell types that would not be expected to be present in organoids (endothelial cells, pericytes, and microglia) were removed. MACS2 was used to rerun peak calling on the remaining cells via Signac’s CallPeaks function, grouped by cell type, yielding 326,515 peaks. A shared peak space was generated for each organoid timepoint using the reduce function from the GenomicRanges R package (Lawrence et al., 2013) to merge peaks called in the two datasets. This yielded 379,920 shared peaks with 1-month organoids, 351,207 shared peaks with 3-month organoids, and 358,664 shared peaks with 6-month organoids. Integration of fetal and organoid scATAC-seq data was performed using Signac.
We used the cell type labels supplied in the previously published endogenous datasets. However, these different labels match well with our own cell type labels. For instance, a match between fetal deep-layer neurons and organoid CFuPN was considered correct; other fetal excitatory neurons (i.e., migrating, maturing, and upper-layer enriched in Polioudakis et al., 2019, and Glutamatergic neurons 1–8 in Trevino et al., 2021) were matched to organoid CPN; fetal IP/nIPC to organoid IP; and both subtypes of fetal interneurons (InMGE/MGE-IN and InCGE/CGE-IN) to organoid IN. Fetal cycling progenitors from both datasets were split between organoid oRG and IP, while fetal oRG were matched to organoid oRG.
Fetal human brain tissue dissection and single nucleus RNA seq
Cryostat coronal sections (70–100 microns) were produced from human fetal somatosensory cortices and deposited on non-chargeable slides. The cortical plate (CP) was further dissected using single-use micro stab knifes (F.S.T. #72–2201). Following dissection, CP sections were gathered with a p1000 pipette in cold EZ-lysis buffer (Sigma, #NUC-101). For snRNA-seq, nuclei were isolated following the (Habib et al., 2017) protocol. In brief, tissue was transferred to a chilled dounce tube containing 2 mL of ice-cold EZ-lysis buffer. Gently, tissue was homogenized with 20 strokes of pestle A followed by pestle B. The suspension was transferred to a chilled conical tube and incubated on ice for 5 min. Tissue suspension was then centrifuged at 500xg, 4°C for 5 min and resuspended in 1 mL of Nuclei Suspension Buffer (NSB): RNAse-free molecular biology grade PBS, 0.1%BSA (NEB #B9000 S), 0.2 U/ml Recombinant ribonuclease inhibitor (Takara, #2313B). A second centrifugation (same conditions) was performed before final resuspension in 500 mL chilled NSB. Nuclei yield was quantified by Trypan blue staining using disposable Hemacytometer chambers (NCYTO C-Chip™, SKC - DHCN015) prior to Chromium Single Cell 10x V3 assay.
Fetal human brain tissue single nucleus RNA-seq pre-processing
SnRNA-seq reads were aligned to the GRCh38 human reference genome and the cell-by-gene count matrices were produced with the Cell Ranger pipeline version 3.0.2 using default parameters. Data was analyzed using the Seurat R package v3.2.0 using R v3.6.0. Cells expressing a minimum of 500 genes were kept, and UMI counts were normalized and scaled using the SCTransfrom function from Seurat (with vars.to.regress parameter set to the number of genes, number of UMIs, and percent mitochondrial gene expression). This function internally computes variable genes and scales the data on these genes.
Fetal brain tissue single nucleus RNA-seq dimensionality reduction and clustering
PCA was performed on the scaled data of the “SCT” assay created during the SCTransform step. Cells were clustered in PCA space using Seurat’s FindNeighbors on the top 30 PCs, followed by FindClusters with resolution = 0.3. Cells were visualized by UMAP on the top 30 PCs. Upregulated genes in each cluster were identified using the VeniceMarker tool from the Signac package v0.0.7 from BioTuring (https://github.com/bioturing/signac). Cell types were assigned to each cluster by looking at the upregulated genes with lowest Bonferroni adjusted p values. In some cases, clusters were further sub-clustered to assign identities at higher resolution (See https://github.com/AmandaKedaigle/OrganoidAtlas/blob/master/Subclustering.R for specific clusters that were split; for cluster numbers, refer to the metadata available for download from the Single Cell Portal).
Matching of human organoid and fetal cells
To assign organoid cells to fetal cell categories, as defined in (Polioudakis et al., 2019; Trevino et al., 2021), we used the fetal categories to train a random forest (RF) classifier to distinguish between cell types. This was done with the tuneRF function in the randomForest R package (Liaw and Wiener, 2002), with doBest = T, after removing cell types not found in organoids (microglia, endothelial cells, and pericytes), and downsampling the data to have an equal number of cells per cell type. Variable genes from each fetal cell dataset were used to train the model. For validation, 20% of the cells were reserved as a cross-validation set, and compared to a null model prepared by shuffling the cell type labels on the training data set. The classifier was then applied to the organoid cell profiles to assign organoid cells to one of the human cortex cell types. Organoid cells at 1 and 1.5 months were subsetted to exclude cells which are not expected to have corresponding cells in the fetal data (Cajal Retzius cells, cortical hem, and subcortical cells).
To classify fetal cells to organoid cell categories, we trained a RF classifier using organoid data and applied it to classify human fetal cells. The data were then downsampled to have an equal number of cells per cell type, and the same cell types were excluded at 1 month. Variable genes from the organoid dataset were used to train the model. For validation, RF models trained on organoid data were tested on a cross validation held-out set of organoid cells (20% of total dataset), compared to null models.
Matching of accessible chromatin in human organoids and fetal cortex
Shared peaks between fetal (Trevino et al., 2021) scATAC-seq fragments and fragments from each organoid timepoint (1, 3, and 6 months) were derived using the reduce function from the GenomicRanges R package (see above). Fetal scATAC-seq counts matrices were recalculated using the shared feature spaces. LSI and SVD were performed on processed fetal scATAC-seq counts matrices, and SVD components 2–30 were used to calculate UMAP embeddings. Seurat’s FindTransferAnchors function (with parameters k.anchor = 7, max.features = 250, and k.filter = NA) and MapQuery functions were used to transfer fetal cell type labels onto organoid cells, and to project organoid scATAC-seq data into the fetal UMAP space. Cajal Retzius, Cortical hem, and subcortical cells, which are not expected to have corresponding cells in the fetal dataset, were excluded from the 1-month organoid.
Fetal cells were down-sampled to a maximum of 450 cells per cluster, and DARs were calculated for each fetal cell type using Seurat’s FindMarkers with a logistic regression framework, with the number of fragments in peak regions as a latent variable. Organoid DARs for each cell type were recalculated at each organoid timepoint (1, 3, and 6 months) using the new shared peaks calculated for organoid and fetal cells. DAR signatures were ranked by degree of differentiation [-log10(p value) * sign(effect)] and then compared using the improved rank-rank hypergeometric overlap test as described above. Cajal Retzius, Cortical hem, and subcortical cells were once again excluded from the 1-month organoid.
Comparison of cell type signatures between human fetal cells and cortical brain organoids
Organoid DEG signatures (above) were compared to fetal DEG signatures, calculated using the same DESeq2-based method as for organoids. Cells originally labeled as cycling progenitors in the human datasets were reassigned to reflect their cell types (oRG and IP, in (Polioudakis et al., 2019), Early_RG, nIPC, mGPC and IN progenitors in (Trevino et al., 2021)) rather than proliferative state, by using FindMarkers with default parameters to examine upregulated genes in clusters containing these cells. In the human fetal dataset presented here, progenitor cells were directly classified based on their identity rather than cell cycle state. DEG signatures were ranked by degree of differentiation [-log10(p value) * sign(effect)] and then compared using the improved rank-rank hypergeometric overlap test [RRHO2 R package v1.0; (Plaisier et al., 2010; Cahill et al., 2018)].
To identify processes enriched in genes that differ between signatures, overlaps between the discordant genes as identified by the RRHO2 test and published gene sets were evaluated. The metabolic genes lists used in the section below, “Impact of metabolic genes on cell classification”, as well as 42 gene sets relevant to neurobiology (see Figure S4C) were downloaded from the Molecular Signatures Database (MSigDB; (Subramanian et al., 2005; Liberzon et al., 2011), gsea-msigdb.org), and were compared to the discordant genes. The statistical significance of gene overlap between two gene lists was calculated by hypergeometric test, and p values were adjusted using the Bonferroni correction.
Weighted Gene Correlation Network Analysis
In order to learn patterns of coordinated gene regulation across the cortical organoid scRNA-seq datasets, WGCNA (Langfelder and Horvath, 2008) was applied to each dataset with multiple batches: 1, 1.5, 2, 3, 4, 5 and 6 months. The Seurat objects were down-sampled to have an equal number of cells per organoid prior to applying WGCNA. Normalized gene expression data were further filtered to remove outlying genes, mitochondrial, and ribosomal genes. Outliers were identified by setting the upper (>9) and lower (<0.15) thresholds to the average normalized expression per gene. After processing, blockwiseModules function from the WGCNA v1.69 library was performed in R with the parameters networkType = “signed”, minModuleSize = 4, corType = “Bicor”, maxPOutliers = 0.1, deepSplit = 3,trapErrors = T, and randomSeed = 59069. Other than power, the remaining parameters were left as the default setting. The power parameter that determines the resolution of gene module output was chosen separately for each dataset. To pick an adequate parameter, we used the pickSoftThreshold function from WGCNA to test values from 1 to 30. Final resolution was determined by studying the output modules and choosing the resolution that captured the most variation in the fewest total number of modules - this resulted in a power of 8 for the 1-month harmonized dataset, 9 for the 1.5-month harmonized dataset, 14 for the 2-month harmonized dataset, 4 for the 3-month harmonized dataset, 6 for the 4-month harmonized dataset, 9 for the 5-month harmonized dataset, and 5 for the 6-month harmonized dataset.
Module scores
Expression of gene sets identified in MSigDB (Subramanian et al., 2005; Liberzon et al., 2011) compared to random gene lists with similar average expression, was evaluated using Seurat’s AddModuleScore function using the downsampled objects used in WGCNA. The distribution of module scores between cell types was visualized using Seurat’s VlnPlot function. To assess the differences between module scores across cell types at each timepoint, the average of module scores was calculated in each cell type per individual organoid. These averages were then submitted to a one-way ANOVA using the base R function aov, with the formula AverageValue ~ CellType. Since all tests were significant for an overall difference of means (significance Pr(>F) < 0.05), post-hoc comparisons were done with the TukeyHSD function. Results from pairwise comparisons were visualized with a compact letter display using the multcompLetters function from the multcompView R package v0.1–8 (Graves et al., 2019) with default parameters.
For module score analysis of MSigDB metabolic pathways in human fetal cortical cells (Polioudakis et al., 2019), cell types that do not appear in the organoids were excluded from the analysis: microglia (48 cells), pericytes (117), and endothelial cells (237). For module score expression of MSigDB metabolic pathways in 3-month organoids cells, Cajal Retzius cells were excluded from the analysis (99 cells). Remaining cells from each dataset were grouped into broad cell type categories: Cells called “CPN”, “GluN1′′, “GluN2′′, “GluN3′′, “GluN4′′, “GluN5′′, “GluN6′′, “GluN7′′, “GluN8′′, “ExN”, “ExM”, and “ExM-U” were all labeled “UL Neurons.” Cells called “CFuPN”, “SP”, “ExDp”, “ExDp1”, and “ExDp2” were all labeled “DL Neurons.” Cells called “iP” and “nIPC” were labeled “IPs.” Cells called “CGE_IN”, “MGE_IN”, “Immature IN”, “InMGE”, and “InCGE” were labeled “INs.” Cells called “tRG”, “Early_RG”, “Late_RG”, “Cyc_Prog”, “aRG”, “oRG”, “PgS”, “PgG2M”, “vRG” were labeled “RG.” Unspecified PN from the organoids retained their unique label.
Evaluation of metabolic activity with compass
Activity scores for metabolic reactions in the RECON2 database (Swainston et al., 2016) were assigned to each cell from the organoid 3-month dataset, and two human fetal datasets (Polioudakis et al., 2019; Trevino et al., 2021), using the Compass tool (Wagner et al., 2021).
To calculate the PCA, datasets were each down-sampled to 1000 cells, and gene counts were normalized to Counts Per Million. This matrix was used as an input to Compass using default parameters and the homo_sapiens species. Downstream analysis was done as in the original Compass paper (Wagner et al., 2021). Briefly, the output penalties were set to zero if they were below 0.0004, and reactions which had a total score of 0 across all cells or that showed a total range across all cells within 0.001 were eliminated from the analysis. Hierarchical clustering was then performed on the penalty matrix, based on Spearman correlation, and reactions were clustered into metareactions based on cutting the hierarchical clustering tree at a Spearman correlation of 0.95. Metareactions were given penalty scores as the mean of their component reactions. The metareaction penalty matrix was then transformed by log1p. PCA was performed on this transformed data. The top 20 principal components were correlated with the number of UMI per cell, the cell type labels of each cell, and the dataset each cell derived from, using linear regression, with the lm R function. The R squared value from the resulting function was reported.
For differential activity analysis, cells from 3-month organoids and the (Trevino et al., 2021) fetal dataset were grouped into broad cell type categories as in the “Module Scores” section above. Datasets were then downsampled to have 50 cells from each broad cell type category. Only genes involved in one or more metabolic reactions (found using Compass’ −list-genes function) were retained, and then gene counts were normalized to Counts Per Million. Normalizing only metabolic-related genes should remove variability due to overall up- or down-regulation of metabolism across the board, emphasizing variable use of specific pathways. Counts were then submitted to Compass as above, and resulting penalty scores were processed as above, without clustering into metareactions. As in the original Compass original publication, reactions were then submitted to a two-sided Wilcoxon test, using the datasets as the two groups. False discovery rate was calculated using the Benjamini Yekutieli method (Yekutieli and Benjamini, 1999).
Comparison of cortical and whole-brain organoids
To assign high-resolution cell types to cortical cells from whole-brain organoids, cortical cells were extracted from those datasets, i.e. clusters with high expression of FOXG1 and EMX1, and submitted to a RF classifier created from 3- and 5-month cortical organoid datasets, using the same process as used above when classifying fetal cells. Module scores for metabolic gene sets were calculated using the process described in the Module scores section above.
Impact of metabolic genes on cell classification
In order to evaluate the effect of metabolic genes on fetal cell classification to organoid cell types and vice versa, 38 gene sets were downloaded from the MSigDB v7.2 (Liberzon et al., 2011). For each gene set, genes in the set were excluded from the list of genes used to build the RF model. The proportion of fetal cell types assigned to each organoid cell type was then compared to those from the model using all organoid variable genes. To assess which changes were higher than may be expected by chance, a background distribution was built. For each timepoint and pathway, for 10,000 repetitions, a randomly selected list of genes was removed from the variable genes in the RF model and the model was re-built. The background sets of genes were the same size as the MSigDB gene set, and were expression matched, using the same method as used by Seurat’s AddModuleScore function to choose background gene sets (briefly, genes are grouped into 24 bins by average expression in the organoid dataset, and for each gene in the MSigDB gene set, a randomly selected gene from the same bin is added to the background set). p values for each fetal to organoid cell type pair were calculated by evaluating the fraction of times the change in cell type proportions in the background distribution were higher than or equal to the change induced by removing metabolic pathway genes. False discovery rate was calculated using the Benjamini Yekutieli method (Yekutieli and Benjamini, 1999).
To evaluate the expression of metabolism related gene sets across topological location, gene set expression of the MSigDB gene lists was evaluated in the Slide-seqV2 data by plotting 100x the sum of the reads for all genes in the gene set that appeared in the data, normalized by the number of UMI for each bead. Expression scores for each bead were then plotted against distance to the edge of the organoid. The edge distance was produced by calculating the convex hull of the set of beads using the chull R function and for each bead, calculating the minimum distance to the convex hull. This was done by calculating the perpendicular distance from the bead to each line segment of the convex hull and taking the minimum distance of this set. Finally, the relationship of edge distance to expression scores was plotted with geom_smooth from R’s ggplot2 library v3.3.3 (Wickham, 2016), using the Loess smoothing function and default parameters.
Epigenetic changes associated with metabolism
To determine epigenetic changes in cells with high expression of metabolic gene sets, we use the SHARE-seq data. Module scores for the Hallmark Glycolysis and Hypoxia gene sets were assigned to cells based on RNA expression, as above. For differential metabolic peak analysis, high scoring cells were identified as those with a hypoxia score > 0.2. For each peak, the test statistic was defined as (fraction accessible in high-hypoxia cells) – (fraction accessible in low-hypoxia cells). A null distribution was calculated by permuting cell labels 1,000 times. The mean and standard deviation from this were used to calculate a Z-score and p value for each peak. Significant peaks were called those with an FDR-adjusted p value less than 0.1. This resulted in 3,189 DARs, out of 216,877 regions tested (1.5% of detected peaks, Table S2E). We matched these regions to genes using the gene-peak associations we had previously calculated for the SHARE-seq cells, and submitted the gene lists to gene ontology. There were 320 genes regulated by significantly downregulated DARs in the high-hypoxia-scored cells, but these genes were not enriched for any GO terms.
Trajectory inference across time
Because organoids were grown with a protocol to pattern dorsal telencephalon, we excluded cell types known to originate in other regions of the developing neural tube (i.e., subcortical cells and cortical hem, in addition to unspecified PN). All objects were then merged, and then we performed PCA on the resulting object. The first 15 PCs were then used as an input for the FLOWMAP algorithm (Ko et al., 2020) to build a force-directed graph of the datasets, by first clustering cells from each timepoint into 3,000 clusters and then building a graph connecting these clusters, using the “single” mode, the k-means clustering algorithm, Euclidean distance between clusters, and allotting 2–5 edges during each edge-building step of the algorithm. FLOWMAP only allows edges between clusters belonging to the same or consecutive timepoints. The graph, with node coordinates calculated by FLOWMAP, was visualized in Cytoscape v.3.8.2 (Shannon et al., 2003).
Differential expression over time
Because ambient RNA could differ between timepoints and contribute to differential expression results, ambient RNA was estimated and removed from organoids using the decontX function from the Celda R package, version 1.6.1 (Yang et al., 2020), with each organoid treated as its own sequencing batch. For each cell type, DESeq2 was used to identify genes that are differentially expressed across organoid age, in days. Significant genes (those with FDR-adjusted p values of less than 0.05) for each cell type were split into two lists based on whether their log-fold-change was positive or negative. In cases where these lists included more than 1000 significant genes, they were truncated to the 1000 genes with the lowest p values. GO term enrichment was calculated for these lists using the enrichGO function from the clusterProfiler R package (Yu et al., 2012). In order to visualize terms with minimal overlaps, significant GO terms were run through the simplify function with a similarity cutoff of 0.4.
Lineage branching tree trajectory
Developmental trajectories were built in organoids in a form of a branching tree using R package URD v1.1.1 (Farrell et al., 2018). The diffusion map was computed using the R package Destiny v2.14.0 (Angerer et al., 2016) by invoking calcDM function from URD on normalized counts from the organoid datasets (KNN = 100 and sigma.use = 30 for 10x data, and KNN = 100 and sigma.use = 30 for SHARE-seq data). The root cells were assigned to be the aRG cells from our youngest sample (i.e., 23 days aRG). Simulating diffusion from the root to each cell, cells were ordered in pseudotime by floodPseudotime and floodPseudotimeProcess functions with the default parameters. Terminal states of the tree (“tips”) were manually selected leveraging biological knowledge. Astroglia, oligodendrocyte precursors, CPN and CFuPN (from all timepoints at which they were present) were selected as tips for scRNAseq data, and CPN and CFuPN for SHARE-seq data. To recover the developmental trajectories in the data, we performed biased random walks starting from each tip. This walk is biased because the transitions are only permitted for cells with younger or similar pseudotimes. Parameters of the logistic function used to bias the transition probabilities were determined using the pseudotimeDetermineLogistic function (optimal.cells.forward = 40 and max.cells.back = 80). A biased transition matrix was obtained from the pseudotimeWeight-TransitionMatrix function. Ten thousand simulated random walks were performed per tip using simulateRandomWalksFromTips function. Walks were then processed into visitation frequencies using processRandomWalksFromTips function. Finally trees were built using the buildTree function with the following parameters for both trees: visit.threshold = 0.7, minimum.visits = 10, bins.per.pseudotime.window = 5, cells.per.pseudotime.bin = 80, divergence.method = “preference”, p.thresh = 0.05.
Gene trajectories from the branching tree
To identify marker genes for each trajectory, we used the aucprTestAlongTree function from the URD package. Starting from a tip, this function compares cells in each segment pairwise with cells from each of that segment’s siblings and children (cropped to the same pseudotime limits as the segment under consideration). Genes were considered differentially expressed if they were expressed in at least 10% of cells in the trajectory segment under consideration (frac.must.express = 0.1), their mean expression was upregulated 1.5x compared to the sibling and the gene was 1.25x better than a random classifier for the population as determined by the area under a precision-recall curve. Genes were considered part of a population’s cascade if, at any given branch point, they were considered differentially expressed against at least 50% of their siblings (must.beat.sibs = 0.5), and they were not differentially upregulated in a different trajectory downstream of the branch point. To determine gene expression onset and offset, we used geneSmoothFit function. Looking at the averages gene expression for a group of genes and cells in a moving window through pseudotime (moving.window = 5, cells.per.window = 25), it applied smoothing algorithms (method = spline) to describe the expression of each gene. Genes were then ordered by the pseudotime value at which they enter and then leave “peak” expression (expression 50% higher than minimum value), and enter and then leave “expression” (expression 20% higher than minimum value), in that order.
Genes associated with tree branching points
To define branch-point-associated genes, we selected cells adjacent to the branch points (0.04 pseudotime units before and after) and calculated differentially expressed genes between parent and sibling branches (FindAllMarkers function from Seurat R package, min.pct = 0.1, logfc = 0.25, test.use = wilcox). To select for the genes that vary in pseudotime, we first identified variable genes in the data (Seurat’s FindVariableFeatures function) and then fitted a Lasso regression model using glmnet function from the R package glmnet 2.0–16. (Friedman et al., 2010) (family = “Gaussian”, type.measure = “mse”, nfolds = 10). The best lambda value that minimizes mean squared error (MSE) was obtained using cv.glmnet function. To find the top distinguishing features at a given branch point, a Gradient Boosting Classifier was trained using scikit-learn 0.22.1 (Pedregosa et al., 2011), with the union of genes from the differential expression and regression analyses. A grid search was performed with tenfold cross-validation to optimize the maximum depth (3, 4, 5) and number of estimators (25, 50, 75, 100), which resulted in the best depth of 4 and the best number of estimators 100. Feature importance score was calculated on the basis of maximal estimated improvement by splitting on the feature under consideration against not-splitting (measured in terms of MSE), using the default option in sklearn, “friedman_mse”. The expected amount of improvement is summed over all internal nodes (where splitting occurs) of a single tree, and then summed over all trees in the gradient boosted tree model to get a single number per gene. TFs were identified using the CIS-BP database (Weirauch et al., 2014).
Branching tree trajectory from SHARE-seq data
For the SHARE-seq tree, a URD tree was constructed as above using the RNA gene count data from cells that contained both RNA and ATAC data. After constructing the tree, the underlying gene count data was substituted with the ATAC peak data, while maintaining the RNA tree embedding. To find enriched TF motifs along pseudotime, cells were cut into seven sections along the tree. The parental branch was cut into three equal divisions (pseudotime from 0 to 0.1057075, thento 0.211415, then to 0.3171224), the CPN branchinto two equal divisions (from 0.3171224 to 0.44005, then to 0.5629775) and the CFuPN branch into two equal divisions (from 0.3171224 to 0.4151327, then to 0.5131430). Cells from each of these sections were grouped, and DARs were calculated for each group using Signac’s FindMarkers function, with the logistic regression framework, with the number of fragments in peak regions as a latent variable, and with min.pct = 0.05. Enriched motifs per group were calculated using Homer as in the “ScATAC-seq motif enrichment” section above.
Human callosal projection neuron diversity
Data from adult CPN from (Hodge et al., 2019) were downloaded, and the exon count matrix was used to create a Seurat object, along with their metadata. Counts were normalized using the same method as for the organoid scRNA-seq data. The five CPN populations were used to define gene modules that are specific to each CPN subtype using consensus NMF (cNMF) (Kotliar et al., 2019). The k parameter of cNMF was set to 5 since there are five different CPN clusters in the original dataset. With five cNMF modules identified, the specificity of each module to a CPN subtype was visualized by plotting the cell loading (H matrix). Gene modules were then curated by taking the top 100 genes with the highest feature loading (W matrix) for each cNMF module. To assess CPN diversity in the organoid and fetal datasets, CPN were isolated from each dataset and a UMAP was calculated for each of them, using the module genes as the variable genes. The expression of the gene modules was visualized using Seurat’s AddModuleScore function. The cNMF algorithm (originally written in Python) was implemented in R by Sean K. Simmons and can be found in our GitHub.
QUANTIFICATION AND STATISTICAL ANALYSIS
Sequencing data was pre-processed using CellRanger (Satpathy et al., 2019), Seurat v3.2.2 (Stuart et al., 2019), and Signac v1.1.0 (Stuart et al., 2021) as described in Method Details. Statistical analyses are performed in R v3.6. No specific methods were used to determine if data met assumptions of statistical approaches.
For cell type signature calculations, data was partitioned into cortical neurons and progenitors, and then reads were summed across cells in each cell type in each organoid in all cell lines. Final number of samples equaled the number of organoids in each batch, usually 3 per group. Genes with less than 10 total UMI were excluded, and then DESeq2 (Love et al., 2014) was used to calculate DEGs between each cell type and all other cell types in its group, using a paired analysis between cells from the same organoid.
To calculate DEGs across time, ambient RNA was first estimate and removed from organoids using decontX 1.6.1 (Yang et al., 2020), because ambient RNA could differ between timepoints and contribute to differential expression results. Next, DESeq2 was used as above to identify genes that are differentially expressed across organoid age for each cell type. In this case, the analysis was not paired, and the number of samples equaled the number of individual organoids at each timepoint.
To train random forest classifiers to compare organoid and human cell types, we used the tuneRF function in the randomForest R package (Liaw and Wiener, 2002), with doBest = T, after downsampling the data to have an equal number of cells per cell type – so in each case, the number of cells per cell type was equal to the cell type with the smallest number of cells. For validation, 20% of the cells were reserved as a cross-validation set, and compared to a null model prepared by shuffling the cell type labels on the training data set.
To evaluate the effect of metabolic genes on the above classifiers, genes in metabolic gene sets from MSigDB v7.2 (Liberzon et al., 2011) were excluded from the list of genes used to build the RF model. The proportion of fetal cell types assigned to each organoid cell type was then compared to those from the model using all variable genes. To assess which changes were higher than may be expected by chance, a background distribution was built. For each timepoint and pathway, for 10,000 repetitions, a randomly selected list of genes was removed from the variable genes in the RF model and the model was re-built. The background sets of genes were the same size as the MSigDB gene set, and were expression matched, using the same method as used by Seurat’s AddModuleScore function to choose background gene sets. p values for each fetal to organoid cell type pair were calculated by evaluating the fraction of times the change in cell type proportions in the background distribution were higher than or equal to the change induced by removing metabolic pathway genes. False discovery rate was calculated using the Benjamini Yekutieli method (Yekutieli and Benjamini, 1999).
Developmental trajectories were built in organoids using a force-directed graph across time using FLOWMAP (Ko et al., 2020), and in a form of a branching tree using R package URD v1.1.1 (Farrell et al., 2018). In the former case, cells were grouped into 3,000 clusters per timepoint. In the later case, data from individual cells is retained without being grouped.
Further information on specific statistical tests is listed in Method Details.
ADDITIONAL RESOURCES
Interactive portal for exploring scRNA-seq, scATAC-seq, SHARE-seq, and Slide-seq data: https://singlecell.broadinstitute.org/single_cell/study/SCP1756.
Supplementary Material
Highlights.
Single cell atlas of human brain organoids reveals trajectories of development
Metabolic states do not affect identity acquisition for most cells in organoids
Analysis of pseudo-time trajectories in organoids predicts human-specific regulation
Transcriptional diversity of callosal neurons emerges at early stages of development
ACKNOWLEDGMENTS
We thank J. R. Brown and S. Simmons for editing the manuscript, and the entire Arlotta Lab for support and insightful discussions; A. Shetty and D. Di Bella for help with scRNA-seq cell type classification; V. Vuong, C. Abbate, R. Sartore for technical support in organoid culture and characterization; the Broad Genomics Platform for sequencing; the Talkowski laboratory for the GM08330 line; the Cohen laboratory for the Mito210 line. This work was supported by grants from the Chan Zuckerberg Initiative to P.A., the Stanley Center for Psychiatric Research to P.A. and J.Z.L., the Broad Institute of MIT and Harvard to P.A. and J.Z.L., the National Institutes of Health (R01MH112940 to P.A. and J.Z.L., and P50MH094271, U01MH115727, and RF1MH123977 to P.A.), the Klarman Cell Observatory to J.Z.L. and A.R., and the Howard Hughes Medical Institute to A.R. A.R. was a Howard Hughes Medical Institute and a Koch Institute extramural member while conducting this study.
Footnotes
DECLARATION OF INTERESTS
P.A. is a SAB member at Herophilus, Rumi Therapeutics, and Foresite Labs, and is a co-founder of Vesalius and a co-founder and equity holder at Foresite Labs. A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics and until August 31, 2020, was a SAB member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and Thermo Fisher Scientific. From August 1, 2020, A.R. has been an employee of Genentech. M.P. is an employee of Roche.
SUPPLEMENTAL INFORMATION
Supplemental information can be found online at https://doi.org/10.1016/j.cell.2022.09.010.
REFERENCES
- Angerer P, Haghverdi L, Büttner M, Theis FJ, Marr C, and Buettner F. (2016). destiny: diffusion maps for large-scale single-cell data in R. Bioinformatics 32, 1241–1243. 10.1093/BIOINFORMATICS/BTV715. [DOI] [PubMed] [Google Scholar]
- Angevine JB, and Sidman RL (1961). Autoradiographic Study of Cell Migration during Histogenesis of Cerebral Cortex in the Mouse. Nature 192, 766–768. 10.1038/192766b0. [DOI] [PubMed] [Google Scholar]
- Di Bella DJ, Habibi E, Stickels RR, Scalia G, Brown J, Yadollahpour P, Yang SM, Abbate C, Biancalani T, Macosko EZ, et al. (2021). Molecular logic of cellular diversification in the mouse cerebral cortex. Nature 595, 554–559. 10.1038/s41586-021-03670-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg J, Sorensen SA, Ting JT, Miller JA, Chartrand T, Buchin A, Bakken TE, Budzillo A, Dee N, Ding SL, et al. (2021). Human neocortical expansion involves glutamatergic neuron diversification. Nature 598, 151–158. 10.1038/s41586-021-03813-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Betizeau M, Cortay V, Patti D, Pfister S, Gautier E, Bellemin-Ménard A, Afanassieff M, Huissoud C, Douglas RJ, Kennedy H, and Dehay C. (2013). Precursor diversity and complexity of lineage relationships in the outer subventricular zone of the primate. Neuron 80, 442–457. 10.1016/j.neuron.2013.09.032. [DOI] [PubMed] [Google Scholar]
- Bhaduri A, Andrews MG, Mancia Leon W, Jung D, Shin D, Allen D, Jung D, Schmunk G, Haeussler M, Salma J, et al. (2020). Cell stress in cortical organoids impairs molecular subtype specification. Nature 578, 142–148. 10.1038/s41586-020-1962-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bravo González-Blas C, Minnoye L, Papasokrati D, Aibar S, Hulselmans G, Christiaens V, Davie K, Wouters J, and Aerts S. (2019). cisTopic: cis-regulatory topic modeling on single-cell ATAC-seq data. Nat. Methods 16, 397–400. 10.1038/s41592-019-0367-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buenrostro JD, Wu B, Litzenburger UM, Ruff D, Gonzales ML, Snyder MP, Chang HY, and Greenleaf WJ (2015). Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490. 10.1038/nature14590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cable DM, Murray E, Zou LS, Goeva A, Macosko EZ, Chen F, and Irizarry RA (2021). Robust decomposition of cell type mixtures in spatial transcriptomics’. Nat. Biotechnol 40, 517–526. 10.1038/S41587-021-00830-W. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cahill KM, Huo Z, Tseng GC, Logan RW, and Seney ML (2018). Improved identification of concordant and discordant gene expression signatures using an updated rank-rank hypergeometric overlap approach. Sci. Rep 8, 9588. 10.1038/s41598-018-27903-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camp JG, Badsha F, Florio M, Kanton S, Gerber T, Wilsch-Bräuninger M, Lewitus E, Sykes A, Hevers W, Lancaster M, et al. (2015). Human cerebral organoids recapitulate gene expression programs of fetal neocortex development. Proc. Natl. Acad. Sci. USA 112, 15672–15677. 10.1073/pnas.1520760112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen AE, Egli D, Niakan K, Deng J, Akutsu H, Yamaki M, Cowan C, Fitz-Gerald C, Zhang K, Melton DA, and Eggan K. (2009). Optimal Timing of Inner Cell Mass Isolation Increases the Efficiency of Human Embryonic Stem Cell Derivation and Allows Generation of Sibling Cell Lines. Cell Stem Cell, 103–106. 10.1016/j.stem.2008.12.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiquet J, Rigaill G, and Sundqvist M. (2020). Aricode: Efficient Computations of Standard Clustering Comparison Measures. R Package Version 1.0. [Google Scholar]
- Chou SJ, and Tole S. (2019). Lhx2, an evolutionarily conserved, multifunctional regulator of forebrain development. Brain Res. 1705, 1–14. 10.1016/j.brainres.2018.02.046. [DOI] [PubMed] [Google Scholar]
- Church GM (2005). The personal genome project. Mol. Syst. Biol 1. 10.1038/msb4100040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coquand L, Macé AS, Farcy S, Avalos CB, Di Cicco A, Lampic M, Bessières B, Attie-Bitach T, Fraisier V, Guimiot F, and Baffet A. (2022). A cell fate decision map reveals abundant direct neurogenesis in the human developing neocortex. Preprint at bioRxiv. 10.1101/2022.02.01.478661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corces MR, Trevino AE, Hamilton EG, Greenside PG, Sinnott-Arm-strong NA, Vesuna S, Satpathy AT, Rubin AJ, Montine KS, Wu B, et al. (2017). An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962. 10.1038/nmeth.4396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delgado RN, Allen DE, Keefe MG, Mancia Leon WR, Ziffra RS, Crouch EE, Alvarez-Buylla A, and Nowakowski TJ (2022). Individual human cortical progenitors can produce excitatory and inhibitory neurons. Nature 601, 397–403. 10.1038/S41586-021-04230-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eckler MJ, Larkin KA, McKenna WL, Katzman S, Guo C, Roque R, Visel A, Rubenstein JLR, and Chen B. (2014). Multiple conserved regulatory domains promote Fezf2 expression in the developing cerebral cortex. Neural Dev. 9, 6. 10.1186/1749-8104-9-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eckler MJ, and Chen B. (2014). Fez family transcription factors: Controlling neurogenesis and cell fate in the developing mammalian nervous system. Bioessays 36, 788–797. 10.1002/bies.201400039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eiraku M, Watanabe K, Matsuo-Takasaki M, Kawada M, Yonemura S, Matsumura M, Wataya T, Nishiyama A, Muguruma K, and Sasai Y. (2008). Self-Organized Formation of Polarized Cortical Tissues from ESCs and Its Active Manipulation by Extrinsic Signals. Cell Stem Cell 3, 519–532. 10.1016/J.STEM.2008.09.002. [DOI] [PubMed] [Google Scholar]
- Farrell JA, Wang Y, Riesenfeld SJ, Shekhar K, Regev A, and Schier AF (2018). Single-cell reconstruction of developmental trajectories during zebrafish embryogenesis. Science, 360. 10.1126/SCIENCE.AAR3131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fietz SA, Kelava I, Vogt J, Wilsch-Bräuninger M, Stenzel D, Fish JL, Corbeil D, Riehn A, Distler W, Nitsch R, and Huttner WB (2010). OSVZ progenitors of human and ferret neocortex are epithelial-like and expand by integrin signaling. Nat. Neurosci 13, 690–699. 10.1038/nn.2553. [DOI] [PubMed] [Google Scholar]
- Fishilevich S, Nudel R, Rappaport N, Hadar R, Plaschkes I, Iny Stein T, Rosen N, Kohn A, Twik M, Safran M, and Lancet D. (2017). GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database : the journal of biological databases and curation, bax028. 10.1093/database/bax028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Florio M, Borrell V, and Huttner WB (2017). Human-specific genomic signatures of neocortical expansion. Curr. Opin. Neurobiol 42, 33–44. 10.1016/J.CONB.2016.11.004. [DOI] [PubMed] [Google Scholar]
- Friedman J, Hastie T, and Tibshirani R. (2010). Regularization Paths for Generalized Linear Models via Coordinate Descent. J. Stat. Softw 33, 1–22. 10.18637/JSS.V033.I01. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuentealba LC, Rompani SB, Parraguez JI, Obernier K, Romero R, Cepko CL, and Alvarez-Buylla A. (2015). Embryonic Origin of Postnatal Neural Stem Cells. Cell 161, 1644–1655. 10.1016/J.CELL.2015.05.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genomics, 10x (2019). Nuclei Isolation from Mouse Brain Tissue for Single Cell ATAC Sequencing · Rev B, Demonstrated Protocol. Available at. https://assets.ctfassets.net/an68im79xiti/zZh2iRV5TgWP8A96Easg6/2286fafe2eae406b031567a16272b8ab/CG000212_SingleCellATAC_Nuclei_Isolation_MouseBrain_DemonstratedProtocol_RevB.pdf. [Google Scholar]
- Gordon A, Yoon SJ, Tran SS, Makinson CD, Park JY, Andersen J, Valencia AM, Horvath S, Xiao X, Huguenard JR, et al. (2021). Longterm maturation of human cortical organoids matches key early postnatal transitions. Nat. Neurosci 24, 331–342. 10.1038/s41593-021-00802-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves S, Piepho H-P, and Selzer L. (2019). multcompView: Visualizations of Paired Comparisons. Available at. https://cran.r-project.org/web/packages/multcompView/index.html.
- Güven A, Kalebic N, Long KR, Florio M, Vaid S, Brandl H, Stenzel D, and Huttner WB (2020). Extracellular matrix-inducing Sox9 promotes both basal progenitor proliferation and gliogenesis in developing neocortex. Elife 9, e49808. 10.7554/eLife.49808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Habib N, Avraham-Davidi I, Basu A, Burks T, Shekhar K, Hofree M, Choudhury SR, Aguet F, Gelfand E, Ardlie K, et al. (2017). Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958. 10.1038/nmeth.4407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen DV, Lui JH, Parker PRL, and Kriegstein AR (2010). Neurogenic radial glia in the outer subventricular zone of human neocortex. Nature 464, 554–561. 10.1038/nature08845. [DOI] [PubMed] [Google Scholar]
- Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, and Glass CK (2010). Simple Combinations of Lineage-Determining Transcription Factors Prime cis-Regulatory Elements Required for Macrophage and B Cell Identities. Mol. Cell 38, 576–589. 10.1016/j.molcel.2010.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hevner RF (2019). Intermediate progenitors and Tbr2 in cortical development. J. Anat 235, 616–625. 10.1111/JOA.12939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodge RD, Bakken TE, Miller JA, Smith KA, Barkan ER, Graybuck LT, Close JL, Long B, Johansen N, Penn O, et al. (2019). Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68. 10.1038/s41586-019-1506-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang Y, McCarthy DJ, and Stegle O. (2019). Vireo: Bayesian demultiplexing of pooled single-cell RNA-seq data without genotype reference. Genome Biol. 20, 273. 10.1186/s13059-019-1865-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kang P, Lee HK, Glasgow SM, Finley M, Donti T, Gaber ZB, Graham BH, Foster AE, Novitch BG, Gronostajski RM, and Deneen B. (2012). Sox9 and NFIA Coordinate a Transcriptional Regulatory Cascade during the Initiation of Gliogenesis. Neuron 74, 79–94. 10.1016/j.neuron.2012.01.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelley KW, and Pașca SP (2022). Human brain organogenesis: Toward a cellular understanding of development and disease. Cell 185, 42–61. 10.1016/J.CELL.2021.10.003. [DOI] [PubMed] [Google Scholar]
- Khacho M, Clark A, Svoboda DS, Azzi J, MacLaurin JG, Meghaizel C, Sesaki H, Lagace DC, Germain M, Harper ME, et al. (2016). Mitochondrial Dynamics Impacts Stem Cell Identity and Fate Decisions by Regulating a Nuclear Transcriptional Program. Cell Stem Cell 19, 232–247. 10.1016/j.stem.2016.04.015. [DOI] [PubMed] [Google Scholar]
- Khacho M, Harris R, and Slack RS (2019). Mitochondria as central regulators of neural stem cell fate and cognitive function’. Nat. Rev. Neurosci 20, 34–48. 10.1038/s41583-018-0091-3. [DOI] [PubMed] [Google Scholar]
- Knobloch M, and Jessberger S. (2017). Metabolism and neurogenesis. Curr. Opin. Neurobiol 42, 45–52. 10.1016/j.conb.2016.11.006. [DOI] [PubMed] [Google Scholar]
- Ko ME, Williams CM, Fread KI, Goggin SM, Rustagi RS, Fragiadakis GK, Nolan GP, and Zunder ER (2020). FLOW-MAP: a graph-based, force-directed layout algorithm for trajectory mapping in single-cell time course datasets. Nat. Protoc 15, 398–420. 10.1038/s41596-019-0246-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohwi M, Petryniak MA, Long JE, Ekker M, Obata K, Yanagawa Y, Rubenstein JLR, and Alvarez-Buylla A. (2007). A Subpopulation of Olfactory Bulb GABAergic Interneurons Is Derived from Emx1- and Dlx5/6-Expressing Progenitors. J. Neurosci 27, 6878–6891. 10.1523/JNEURO-SCI.0254-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korsunsky I, Millard N, Fan J, Slowikowski K, Zhang F, Wei K, Baglaenko Y, Brenner M, Loh PR, and Raychaudhuri S. (2019). Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296. 10.1038/s41592-019-0619-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kotliar D, Veres A, Nagy MA, Tabrizi S, Hodis E, Melton DA, and Sabeti PC (2019). Identifying gene expression programs of cell-type identity and cellular activity with single-cell RNA-Seq. Elife 8, e43803. 10.7554/ELIFE.43803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kriegstein A, and Alvarez-Buylla A. (2009). The Glial Nature of Embryonic and Adult Neural Stem Cells. Annu. Rev. Neurosci 32, 149–184. 10.1146/annurev.neuro.051508.135600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lancaster MA, and Knoblich JA (2014). Organogenesis in a dish: Modeling development and disease using organoid technologies. Science 345, 1247125. 10.1126/science.1247125. [DOI] [PubMed] [Google Scholar]
- Langfelder P, and Horvath S. (2008). WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559. 10.1186/1471-2105-9-559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lareau CA, Duarte FM, Chew JG, Kartha VK, Burkett ZD, Kohlway AS, Pokholok D, Aryee MJ, Steemers FJ, Lebofsky R, and Buenrostro JD (2019). Droplet-based combinatorial indexing for massive-scale single-cell chromatin accessibility. Nat. Biotechnol 37, 916–924. 10.1038/s41587-019-0147-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence M, Huber W, Pagè s H, Aboyoun P, Carlson M, Gentleman R, Morgan MT, and Carey VJ (2013). Software for Computing and Annotating Genomic Ranges. PLoS Comput. Biol 9, e1003118. 10.1371/JOURNAL.PCBI.1003118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liaw A, and Wiener M. (2002). Classification and Regression by Random Forest. R. News 2, 18–22. [Google Scholar]
- Liberzon A, Subramanian A, Pinchback R, Thorvaldsdóttir H, Tamayo P, and Mesirov JP (2011). Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740. 10.1093/bioinformatics/btr260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liberzon A, Birger C, Thorvaldsdó ttir H, Ghandi M, Mesirov JP, and Tamayo P. (2015). The Molecular Signatures Database Hallmark Gene Set Collection. Cell Syst. 1, 417–425. 10.1016/j.cels.2015.12.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Love MI, Huber W, and Anders S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. 10.1186/s13059-014-0550-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lukaszewicz A, Savatier P, Cortay V, Giroud P, Huissoud C, Berland M, Kennedy H, and Dehay C. (2005). G1 phase regulation, area-specific cell cycle control, and cytoarchitectonics in the primate cortex. Neuron 47, 353–364. 10.1016/j.neuron.2005.06.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma S, Zhang B, LaFave LM, Earl AS, Chiang Z, Hu Y, Ding J, Brack A, Kartha VK, Tay T, et al. (2020). Chromatin Potential Identified by Shared Single-Cell Profiling of RNA and Chromatin. Cell 183, 1103–1116.e20. 10.1016/J.CELL.2020.09.056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Markenscoff-Papadimitriou E, Whalen S, Przytycki P, Thomas R, Binyameen F, Nowakowski TJ, Kriegstein AR, Sanders SJ, State MW, Pollard KS, and Rubenstein JL (2020). A Chromatin Accessibility Atlas of the Developing Human Telencephalon. Cell 182, 754–769.e18. 10.1016/j.cell.2020.06.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsunaga E, Araki I, and Nakamura H. (2000). Pax6 defines the di-mesencephalic boundary by repressing En1 and Pax2. Development 127, 2357–2365. Available at. internal-pdf://183.74.63.230/Matsunaga-2000-Pax6definesthedi-mesencephal.pdf. [DOI] [PubMed] [Google Scholar]
- Miller DJ, Bhaduri A, Sestan N, and Kriegstein A. (2019). Shared and derived features of cellular diversity in the human cerebral cortex. Curr. Opin. Neurobiol 56, 117–124. 10.1016/J.CONB.2018.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miura Y, Li MY, Birey F, Ikeda K, Revah O, Thete MV, Park JY, Puno A, Lee SH, Porteus MH, and Pașca SP (2020). Generation of human striatal organoids and cortico-striatal assembloids from human pluripotent stem cells. Nat. Biotechnol 38, 1421–1430. 10.1038/s41587-020-00763-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molyneaux BJ, Arlotta P, Hirata T, Hibi M, and Macklis JD (2005). Fezl Is Required for the Birth and Specification of Corticospinal Motor Neurons. Neuron 47, 817–831. 10.1016/j.neuron.2005.08.030. [DOI] [PubMed] [Google Scholar]
- Molyneaux BJ, Goff LA, Brettler AC, Chen HH, Hrvatin S, Rinn JL, and Arlotta P. (2015). DeCoN: genome-wide analysis of in vivo transcriptional dynamics during pyramidal neuron fate selection in neocortex. Neuron 85, 275–288. 10.1016/j.neuron.2014.12.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Namba T, Nardelli J, Gressens P, and Huttner WB (2020). Metabolic Regulation of Neocortical Expansion in Development and Evolution’. Neuron 109, 408–419. 10.1016/j.neuron.2020.11.014. [DOI] [PubMed] [Google Scholar]
- Nowakowski TJ, Pollen AA, Sandoval-Espinosa C, and Kriegstein AR (2016). Transformation of the Radial Glia Scaffold Demarcates Two Stages of Human Cerebral Cortex Development. Neuron 91, 1219–1227. 10.1016/j.neuron.2016.09.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Obernier K, and Alvarez-Buylla A. (2019). Neural stem cells: origin, heterogeneity and regulation in the adult mammalian brain. Development 146, dev156059. 10.1242/dev.156059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paulsen B, Velasco S, Kedaigle AJ, Pigoni M, Quadrato G, Deo AJ, Adiconis X, Uzquiano A, Sartore R, Yang SM, et al. (2022). Autism genes converge on asynchronous development of shared neuron classes. Nature 602, 268–273. 10.1038/s41586-021-04358-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, Blondel M, Prettenhofer P, Weiss R, Dubourg V, and Vanderplas J. (2011). Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res 12, 2825–2830. http://jmlr.org/papers/v12/pedregosa11a.html. [Google Scholar]
- Peukert D, Weber S, Lumsden A, and Scholpp S. (2011). Lhx2 and Lhx9 determine neuronal differentiation and compartition in the caudal forebrain by regulating Wnt signaling. PLoS Biol. 9, e1001218. 10.1371/journal.pbio.1001218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piper M, Barry G, Harvey TJ, McLeay R, Smith AG, Harris L, Mason S, Stringer BW, Day BW, Wray NR, et al. (2014). NFIB-mediated repression of the epigenetic factor Ezh2 regulates cortical development. J. Neurosci 34, 2921–2930. 10.1523/jneurosci.2319-13.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plaisier SB, Taschereau R, Wong JA, and Graeber TG (2010). Rank-rank hypergeometric overlap: identification of statistically significant overlap between gene-expression signatures. Nucleic Acids Res. 38, e169. 10.1093/nar/gkq636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pliner HA, Packer JS, McFaline-Figueroa JL, Cusanovich DA, Daza RM, Aghamirzaie D, Srivatsan S, Qiu X, Jackson D, Minkina A, et al. (2018). Cicero Predicts cis-Regulatory DNA Interactions from Single-Cell Chromatin Accessibility Data. Mol. Cell 71, 858–871.e8. 10.1016/j.molcel.2018.06.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Polioudakis D, de la Torre-Ubieta L, Langerman J, Elkins AG, Shi X, Stein JL, Vuong CK, Nichterwitz S, Gevorgian M, Opland CK, et al. (2019). A Single-Cell Transcriptomic Atlas of Human Neocortical Development during Mid-gestation. Neuron 103, 785–801.e8. 10.1016/j.neuron.2019.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollen AA, Bhaduri A, Andrews MG, Nowakowski TJ, Meyerson OS, Mostajo-Radji MA, Di Lullo E, Alvarado B, Bedolli M, Dougherty ML, et al. (2019). Establishing Cerebral Organoids as Models of Human-Specific Brain Evolution. Cell 176, 743–756.e17. 10.1016/j.cell.2019.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qian X, Su Y, Adam CD, Deutschmann AU, Pather SR, Goldberg EM, Su K, Li S, Lu L, Jacob F, et al. (2020). Sliced Human Cortical Organoids for Modeling Distinct Cortical Layer Formation. Cell Stem Cell 26, 766–781.e9. 10.1016/j.stem.2020.02.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quadrato G, Nguyen T, Macosko EZ, Sherwood JL, Min Yang S, Berger DR, Maria N, Scholvin J, Goldman M, Kinney JP, et al. (2017a). Cell diversity and network dynamics in photosensitive human brain organoids. Nature 545, 48–53. 10.1038/nature22047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quadrato G, Sherwood JL, and Arlotta P. (2017b). Long term culture and electrophysiological characterization of human brain organoids. Protocol Exchange. 10.1038/protex.2017.049. [DOI] [Google Scholar]
- Rainer J. (2017). EnsDb.Hsapiens.v86: Ensembl Based Annotation Package. R package [Google Scholar]
- Rakic P. (1974). Neurons in rhesus monkey visual cortex: systematic relation between time of origin and eventual disposition. Science 183, 425–427. 10.1126/science.183.4123.425. [DOI] [PubMed] [Google Scholar]
- Rash BG, Duque A, Morozov YM, Arellano JI, Micali N, and Rakic P. (2019). Gliogenesis in the outer subventricular zone promotes enlargement and gyrification of the primate cerebrum. Proc. Natl. Acad. Sci. USA 116, 7089–7094. 10.1073/pnas.1822169116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruan J, Li H, Chen Z, Coghlan A, Coin LJM, Guo Y, Hériché JK, Hu Y, Kristiansen K, Li R, et al. (2008). TreeFam: 2008 Update. Nucleic Acids Res. 36, D735–D740. 10.1093/NAR/GKM1005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satpathy AT, Granja JM, Yost KE, Qi Y, Meschi F, McDermott GP, Olsen BN, Mumbach MR, Pierce SE, Corces MR, et al. (2019). Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol 37, 925–936. 10.1038/s41587-019-0206-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schep AN, Wu B, Buenrostro JD, and Greenleaf WJ (2017). chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978. 10.1038/nmeth.4401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, and Ideker T. (2003). Cytoscape: A software Environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. 10.1101/gr.1239303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schneider CA, Rasband WS, and Eliceiri KW (2012). NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675. 10.1038/nmeth.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shekhar K, Lapan SW, Whitney IE, Tran NM, Macosko EZ, Kowalczyk M, Adiconis X, Levin JZ, Nemesh J, Goldman M, et al. (2016). Comprehensive Classification of Retinal Bipolar Neurons by Single-Cell Transcriptomics. Cell 166, 1308–1323.e30. 10.1016/j.cell.2016.07.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shen Q, Wang Y, Dimos JT, Fasano CA, Phoenix TN, Lemischka IR, Ivanova NB, Stifani S, Morrisey EE, and Temple S. (2006). The timing of cortical neurogenesis is encoded within lineages of individual progenitor cells. Nat. Neurosci 9, 743–751. 10.1038/nn1694. [DOI] [PubMed] [Google Scholar]
- Silbereis JC, Pochareddy S, Zhu Y, Li M, and Sestan N. (2016). The Cellular and Molecular Landscapes of the Developing Human Central Nervous System. Neuron 89, 248–268. 10.1016/j.neuron.2015.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sousa AMM, Meyer KA, Santpere G, Gulden FO, and Sestan N. (2017). Evolution of the Human Nervous System Function, Structure, and Development. Cell 170, 226–247. 10.1016/J.CELL.2017.06.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stickels RR, Murray E, Kumar P, Li J, Marshall JL, Di Bella DJ, Arlotta P, Macosko EZ, and Chen F. (2021). Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat. Biotechnol 39, 313–319. 10.1038/s41587-020-0739-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuart T, Butler A, Hoffman P, Hafemeister C, Papalexi E, Mauck WM 3rd, Hao Y, Stoeckius M, Smibert P, and Satija R. (2019). Comprehensive Integration of Single-Cell Data. Cell 177, 1888–1902.e21. 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuart T, Srivastava A, Madad S, Lareau CA, and Satija R. (2021). Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341. 10.1038/S41592-021-01282-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, and Mesirov JP (2005). Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550. 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swainston N, Smallbone K, Hefzi H, Dobson PD, Brewer J, Hanscho M, Zielinski DC, Ang KS, Gardiner NJ, Gutierrez JM, et al. (2016). Recon 2.2: from reconstruction to model of human metabolism. Metabolomics 12, 109. 10.1007/S11306-016-1051-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka Y, Cakir B, Xiang Y, Sullivan GJ, and Park IH (2020). Synthetic Analyses of Single-Cell Transcriptomes from Multiple Brain Organoids and Fetal Brain. Cell Rep. 30, 1682–1689.e3. 10.1016/j.celrep.01.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Telley L, Agirman G, Prados J, Amberg N, Fièvre S, Oberst P, Bartolini G, Vitali I, Cadilhac C, Hippenmeyer S, et al. (2019). Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science 364, eaav2522. 10.1126/science.aav2522. [DOI] [PubMed] [Google Scholar]
- Trevino AE, Müller F, Andersen J, Sundaram L, Kathiria A, Shcherbina A, Farh K, Chang HY, Pașca AM, Kundaje A, et al. (2021). Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 184, 5053–5069.e23. 10.1016/J.CELL.07.039. [DOI] [PubMed] [Google Scholar]
- Trevino AE, Sinnott-Armstrong N, Andersen J, Yoon SJ, Huber N, Pritchard JK, Chang HY, Greenleaf WJ, and Pașca SP (2020). Chromatin accessibility dynamics in a model of human forebrain development. Science 367, eaay1645. 10.1126/science.aay1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trujillo CA, Gao R, Negraes PD, Gu J, Buchanan J, Preissl S, Wang A, Wu W, Haddad GG, Chaim IA, et al. (2019). Complex Oscillatory Waves Emerging from Cortical Organoids Model Early Human Brain Network Development. Cell Stem Cell 25, 558–569.e7. 10.1016/j.stem.2019.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trujillo CA, Adams JW, Negraes PD, Carromeu C, Tejwani L, Acab A, Tsuda B, Thomas CA, Sodhi N, Fichter KM, et al. (2020). Pharmacological reversal of synaptic and network pathology in human MECP2-KO neurons and cortical organoids’. EMBO Mol. Med 13, e12523. 10.15252/emmm.202012523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velasco S, Kedaigle AJ, Simmons SK, Nash A, Rocha M, Quadrato G, Paulsen B, Nguyen L, Adiconis X, Regev A, et al. (2019a). Individual brain organoids reproducibly form cell diversity of the human cerebral cortex. Nature 570, 523–527. 10.1038/s41586-019-1289-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Velasco S, Paulsen B, and Arlotta P. (2019b). Highly reproducible human brain organoids recapitulate cerebral cortex cellular diversity. Protocol Exchange. 10.21203/rs.2.9542/v1. [DOI] [Google Scholar]
- Wagner A, Wang C, Fessler J, DeTomaso D, Avila-Pacheco J, Kaminski J, Zaghouani S, Christian E, Thakore P, Schellhaass B, et al. (2021). Metabolic modeling of single Th17 cells reveals regulators of autoimmunity. Cell 184, 4168–4185.e21. 10.1016/J.CELL.2021.05.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weirauch MT, Yang A, Albu M, Cote AG, Montenegro-Montero A, Drewe P, Najafabadi HS, Lambert SA, Mann I, Cook K, et al. (2014). Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443. 10.1016/J.CELL.2014.08.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickham H. (2016). ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag; ). [Google Scholar]
- Xuan Vinh N, Epps J, and Bailey J. (2010). Information Theoretic Measures for Clusterings Comparison: Variants, Properties, Normalization and Correction for Chance. J. Mach. Learn. Res 11, 2837–2854. [Google Scholar]
- Wolock SL, Lopez R, and Klein AM (2019). Scrublet: Computational Identification of Cell Doublets in Single-Cell Transcriptomic Data. Cell Syst. 8, 281–291.e9. 10.1016/j.cels.2018.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang S, Corbett SE, Koga Y, Wang Z, Johnson WE, Yajima M, and Campbell JD (2020). Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol. 21. 57–15. 10.1186/S13059-020-1950-6/FIGURES/6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yekutieli D, and Benjamini Y. (1999). Resampling-based false discovery rate controlling multiple test procedures for correlated test statistics. J. Stat. Plann. Inference 82, 171–196. 10.1016/s0378-3758(99)00041-5. [DOI] [Google Scholar]
- Yoon SJ, Elahi LS, Pașca AM, Marton RM, Gordon A, Revah O, Miura Y, Walczak EM, Holdgate GM, Fan HC, et al. (2019). Reliability of human cortical organoid generation. Nat. Methods 16, 75–78. 10.1038/s41592-018-0255-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yoshida M, Suda Y, Matsuo I, Miyamoto N, Takeda N, Kuratani S, and Aizawa S. (1997). Emx1 and Emx2 functions in development of dorsal telencephalon. Development 124, 101–111. internal-pdf://159.191.232.224/Yoshida-1997-Emx1andEmx2functionsindevelo.pdf. [DOI] [PubMed] [Google Scholar]
- Young KM, Fogarty M, Kessaris N, and Richardson WD (2007). Subventricular Zone Stem Cells Are Heterogeneous with Respect to Their Embryonic Origins and Neurogenic Fates in the Adult Olfactory Bulb. J. Neurosci 27, 8286–8296. 10.1523/JNEUROSCI.0476-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu G, Wang LG, Han Y, and He QY (2012). clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS A J. Integr. Biol 16, 284–287. 10.1089/omi.2011.0118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yue Huang A, Li P, Rodin RE, Kim SN, Dou Y, Kenny CJ, Akula SK, Hodge RD, Bakken TE, Miller JA, et al. (2020). Parallel RNA and DNA analysis after deep sequencing (PRDD-seq) reveals cell type-specific lineage patterns in human brain. Proc. Natl. Acad. Sci. USA 117, 13886–13895. 10.15154/1503337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuzwa SA, Borrett MJ, Innes BT, Voronova A, Ketela T, Kaplan DR, Bader GD, and Miller FD (2017). Developmental Emergence of Adult Neural Stem Cells as Revealed by Single-Cell Transcriptional Profiling. Cell Rep. 21, 3970–3986. 10.1016/J.CELREP.2017.12.017. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, and Liu XS (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137. 10.1186/gb-2008-9-9-r137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zheng X, Boyer L, Jin M, Mertens J, Kim Y, Ma L, Ma L, Hamm M, Gage FH, and Hunter T. (2016). Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation. Elife 5, e13374. 10.7554/eLife.13374. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
Read-level data from scRNA-seq, scATAC-seq, SHARE-seq, and Slide-seq data, supporting the findings of this study have been deposited in a controlled access repository at https://www.synapse.org with accession number project ID syn26346373, while count-level data and meta-data have been deposited at the Single Cell Portal (https://singlecell.broadinstitute.org/single_cell/study/SCP1756).
Supplemental data related to Figures 1, 3, 4, 5, 6, and 7 has been deposited in Mendeley Data at https://doi.org/10.17632/7cxccpv4hg.1.
Code used during data analysis is available at https://github.com/AmandaKedaigle/OrganoidAtlas. d Any additional information required to reanalyze the data reported in this paper is available from the lead contact upon request.
Data from previous publications that was used in this study can be found at the Gene Expression Omnibus under GEO GSE129519, at http://solo.bmap.ucla.edu/shiny/webapp/, and at https://github.com/GreenleafLab/brainchromatin.