Abstract
Small ubiquitin-related modifier (SUMOylation) is a dynamic post-translational modification that maintains cardiac function and can protect against a hypertrophic response to cardiac pressure overload. However, the function of SUMOylation after myocardial infarction (MI) and the molecular details of heart cell responses to SUMO1 deficiency have not been determined. In this study, we demonstrated that SUMO1 protein was inconsistently abundant in different cell types and heart regions after MI. However, SUMO1 knockout significantly exacerbated systolic dysfunction and infarct size after myocardial injury. Single-nucleus RNA sequencing revealed the differential role of SUMO1 in regulating heart cells. Among cardiomyocytes, SUMO1 deletion increased the Nppa+Nppb+Ankrd1+ cardiomyocyte subcluster proportion after MI. In addition, the conversion of fibroblasts to myofibroblasts subclusters was inhibited in SUMO1 knockout mice. Importantly, SUMO1 loss promoted proliferation of endothelial cell subsets with the ability to reconstitute neovascularization and expressed angiogenesis-related genes. Computational analysis of ligand/receptor interactions suggested putative pathways that mediate cardiomyocytes to endothelial cell communication in the myocardium. Mice preinjected with cardiomyocyte-specific AAV-SUMO1, but not the endothelial cell-specific form, and exhibited ameliorated cardiac remodeling following MI. Collectively, our results identified the role of SUMO1 in cardiomyocytes, fibroblasts, and endothelial cells after MI. These findings provide new insights into SUMO1 involvement in the pathogenesis of MI and reveal novel therapeutic targets.
Keywords: Myocardial infarction, SUMO1, SnRNA-seq, Cardiomyocyte
Graphical abstract
Highlights
-
•
SUMO1 deficiency exacerbated systolic dysfunction and infarct size after myocardial injury.
-
•
SnRNA-seq showed cell-specific effects of SUMO1 signaling in infarcted myocardium.
1. Introduction
Ischemic heart disease is caused by complete or partial occlusion of coronary vessels, which leads to myocardial ischemia, hypoxia, and progressive cardiomyocyte (CM) loss in the coronary or microvascular supply area [1]. Loss of functional myocardium initially induces recruitment and infiltration of inflammatory cells, followed by the conversion of fibroblasts (FBs) to myofibroblasts, which deposit large amounts of collagen and form scar in the infarcted area, and endothelial cells (ECs) are involved in neovascularization. Progressive injury after myocardial infarction (MI) triggers pathological cardiac remodeling [2,3]. In addition, persistently activated FBs and accumulation of extracellular matrix lead to abnormal cardiac stiffness and compliance [4]. In short, multiple types of cells are involved in the pathological cardiac response after MI. Identifying the cellular and molecular mechanisms by which coronary ischemia triggers cardiac remodeling will be crucial for identifying new therapeutic approaches.
Small ubiquitin-related modifier (SUMOylation) is a reversible, post-translational modification that modifies protein structures by covalently binding to the lysine residues of the substrate, thereby regulating protein activity, location, and stability [5,6]. SUMOylation is critical for regulating a variety of cellular processes because numerous proteins can be regulated by SUMO molecules in eukaryotic cells [[6], [7], [8], [9]]. The human cell atlas reveals a diversity of cell types in the heart, including CM, FB, and macrophage (MP) subsets and diverse EC clusters in vascular network [[10], [11], [12]]. SUMO molecules play inconsistent roles in different cell types in the heart. For example, regulation of sarcoplasmic/endoplasmic reticulum Ca2+ATPase 2a (SERCA2a) by SUMO1 in CMs is critical for maintaining Ca2+ homeostasis in pressure overload [13,14] or ischemic-induced heart failure [15,16]. In addition, increased SUMOylation affects autophagy by controlling the flux of the ubiquitin-proteasome system to improve protein quality control [17]. However, previous studies have mainly focused on the whole heart or isolated CMs and have not explored the effect of SUMO on different cell types in the heart, which overlooked the effect of SUMO on cardiac pathological processes in distinct cell subsets.
Here, we analyzed the nuclear transcriptome in healthy and MI mouse hearts before and after SUMO1 knockout (SUMO1−/−) using single-nucleus RNA sequencing (snRNA-seq). Our study examined the cell-specific actions of SUMO1 in infarcted myocardium, including CMs, FBs, and ECs. SUMO1 deletion increased the proportion of Nppa+ Nppb+ CM cluster and inhibited the conversion of FB to myofibroblasts. In contrast, SUMO1 deficiency promoted proliferation of the EC clusters responsible for endothelial activation and vascular regeneration, which promoted the revascularization response after MI. In addition, the regulatory effect of vascular endothelial growth factor A (VEGFA) secreted by CMs on EC proliferation was inhibited by SUMO1. Collectively, we demonstrate novel mechanisms of SUMO1 in the regulation of infarcted myocardium based on cell type specificity, which provides a new therapeutic target after MI.
2. Experimental
2.1. Animal experiment
Mice were maintained in a sterile environment under a 12:12 h light/dark cycle (ambient temperature of 23 °C) and fed a standard laboratory diet. All animal studies were approved by the Laboratory Animal Ethics Committee of Tianjin University of Traditional Chinese Medicine (Permit Number: TCM LAEC2014004). The SUMO1−/− mice [18] were backcrossed with C57BL/6 wild-type (WT) mice. SUMO1−/− mice and WT littermates were reproduced by intercrossing the heterozygous mice. Male mice (8–12 weeks old) were used for experiments. The primers used for genotyping were available as follows:
SUMO1-XA-F1: TCCACCTGCCTCTACCTCAAGTGCTG
SUMO1-XA-F2: GGCTGGCTTAACTATGCGGCATCAGAG
SUMO1-XA-R1: CGCCTAAGTCCTCAGTTGAAGGTTTTGC
2.2. Animal models of MI
MI was induced by permanent ligation of the left anterior descending coronary artery (LAD) as we previously described [19]. Briefly, mice were anesthetized with 1% isoflurane supplemented by a one-time administration of 80 mg/kg ketamine and 7 mg/kg xylazine intraperitoneally. Then, mice were intubated for mechanical ventilation with a 16-gauge intravenous catheter. A left thoracotomy was performed to expose the heart and the LAD was ligated with a 6-0 silk ligature. After that, the skin was closed and the mice were placed on a heating plate until they recovered from anesthetic. The sham group went through the same procedure without ligation.
2.3. Echocardiography measurement
To assess cardiac function, Vevo2100 ultrasound imaging system (Visualsonics, Bothell, WA, USA) was used to measure the left ventricular function. Mice were anesthetized with 1% oxygen plus 5% isoflurane. Two-dimensional and M-mode images were observed in the short-axis view. The following parameters were measured: the left ventricular ejection fraction (EF), fractional shortening (FS), the left ventricular systolic diameter (LVIDs), and the left ventricular diastolic diameter (LVIDd).
2.4. Single-cell transcriptome analysis
Raw counts obtained from Singleron Matrix® (Singleron, Nanjing, China) were aligned to the mouse reference (GRCm38version) using the CeleScope pipeline with default parameters (version 1.8.0), provided by Singleronbio company (Nanjing, China) (https://github.com/singleron-RD/CeleScope). The filtered expression matrix outputted by CeleScope for each sample was read in and processed with the R package Seurat (version 4.0.3) [20]. Quality control of the single-cell expression matrix was also performed using Seurat. Cells were first filtered for quality based on the following criteria: having a unique molecular identifier (UMI) count above 500, mapping to over 100 unique genes, and the fraction of unique mitochondrial transcripts less than 15%. Droplets in the cells were detected using the R package Droplet Utils [21], while doublets in the cells were detected using the R package Doublet Finder [22]. Both detections were carried out using default parameters. Supposed droplets and doublets were dropped, remaining cells for all samples were kept, and then they were merged together for following analysis. At last, genes that were expressed in less than 30 cells were also removed.
After quality control, 103,806 cells remained. The expression matrix was normalized and scaled with library size and percentage of mitochondrial read was regressed using Seurat. A total of 2,000 highly variable genes (hvg) were calculated using Find Variable Features from Seurat, and then used for conducting principal component analysis (PCA) with parameter, total number of PCs to compute and store (npcs) = 30. Based on PCA result, the Harmony Program (version 0.1.0) [23] was used to remove potential batches among 12 samples. After correcting batches with Harmony, a new dimensionality reduction named harmony was produced in Seurat single-cell object. The harmony dimensionality reduction, instead of PCA dimensionality reduction, was used to construct a shared nearest neighbor graph by calculating the neighborhood overlap (Jaccard index) between every cell in the single-cell RNA (scRNA) dataset using FindNeighbors from Seurat. And then, the shared nearest neighbor graph was used to identify clusters of cells using FindClusters from Seurat. T-distributed stochastic neighbor embedding (tSNE) dimensionality reduction and uniform manifold approximation and projection (UMAP) dimensionality reduction were performed on the harmony dimensionality reduction for visualization using RunTSNE and RunUMAP from Seurat.
We tested a wide range of resolutions, and the output numbers of identified cell clusters varied with different resolution parameters. To choose a proper resolution parameter, ROGUE method [24] was used to evaluate the purity of output clusters. After ROGUE and manual evaluation, resolution = 0.3 was selected to identify cell clusters. At last, seven cell clusters were obtained.
Wilcoxon rank sum test was performed to identify upregulated genes in each cluster using FindAllMarkers function from Seurat. We found that some clusters might share quite similar upregulated genes. We checked their distribution on tSNE or UMAP. If they were located closely to each other on tSNE or UMAP at the same time, those clusters would be regarded as the same cell type and would be merged into one combined cluster manually. After manually adjustment, all above clusters were merged into four major groups. Wilcoxon rank sum test was performed again to identify significantly upregulated genes in each group using FindAllMarkers. The significantly upregulated genes were used to infer cell types for all the groups. For example, high expressions of Myh7, Tnnt2 and Ttn were used to identify cardiomyocyte cell group, while high expressions of Vwf, Pecam1 and Cdh5 were used to identify endothelial cell group.
2.5. Statistic for cell type enrichment
After identifying cell subtypes using significantly upregulated genes, we calculated the cell type enrichment score (Ro/e) across tissues by comparing the observed and expected cell numbers in each subtype according to the formula, Ro/e = Observed/Expected. The expected cell numbers for each combination of cell subtypes and tissues are obtained from the Chi-square test. The algorithm for quantifying observed and expected cell numbers was described in detail in a previous paper [25]. One cluster was regarded to be enriched in a specific tissue if the corresponding Ro/e is larger than 1.
2.6. Developmental trajectory inference
Monocle (Version 2.18.0) [26] and CytoTrace (version 0.3.3) [27] were used to infer the developmental trajectories of CMs, FBs, and CEs, separately. Each of the interested cell types was separated from Seurat object and applied for trajectory analysis with both Monocle and CytoTrace.
For Monocle analysis, the official tutorial (http://cole-trapnell-lab.github.io/monocle-release/docs/#constructing-single-cell-trajectories) was used. The Monocle object was constructed using new Cell Data Set function in Monocle. EstimateSizeFactors was used to normalize for differences in mRNA recovered across cells, and then Estimate Dispersions was used to help perform differential expression analysis. Genes with mean expression of over 0.1 and expressed in more than 10 cells were selected and applied for differential expression analysis using differential Gene Test from Monocle. Differential expressed genes with qval lower than 0.01 were used to order cells along the trajectory. For CytoTrace analysis, the official tutorial (https://cytotrace.stanford.edu/) was used to predict differentiation states of different cell types separately with default parameters.
2.7. Gene set enrichment analysis
Gene set variation analysis was implemented in the R package ReactomeGSA [28], which is a client to the web-based Reactome Analysis System and performs a gene set analysis using the latest version of the Reactome pathway database as a backend. T cells, myeloid cells, and natural killer (NK) cells were subtracted from Seurat object, separately, and were applied for gene set analysis using analyse_sc_clusters from ReactomeGSA according to the tutor. Enrichment score of gene sets of all reactome pathways was calculated for the cell subtypes. Then, we calculated the standard deviation for enrichment score of each reactome pathway gene set. Among all the reactome pathways, we tried to find the top 20 relevant pathways with the highest standard deviations among cell subtypes.
2.8. Immunofluorescence
Harvested heart tissues were fixed in 4% paraformaldehyde solution for 24 h, embedded in paraffin and sectioned in 4-μm-thick slices. Slices were permeated with 0.1% Triton X-100 for 10 min and then incubated with 3% bovine serum albumin for 1 h. Immunostaining of cTnT, Ankrd1, Top2a, collagen type III, Pecam1, Ki67 and α-SMA using anti-cTnT (# ab91695, 1:300; Abcam, Waltham, MA, USA), anti-Ankrd1 (# 67775-1-Ig, 1:100; Proteintech, Wuhan, China), anti-collagen type III (N-termial) (# 22734-1-AP, 1:200; Proteintech), anti-Pecam1 (# ab24590, 1:200; Abcam), anti-Ki67 (# 27309-1-AP, 1:200; Proteintech), and anti-α-SMA (# ab7817, 1:200; Abcam) primary antibodies were performed on tissue sections overnight at 4 °C. Then the sections were stained with goat anti-rabbit IgG H&L (DyLight® 488) (# ab150077, 1:500; Abcam), and goat anti-mouse IgG H&L (DyLight® 594) (# ab150116, 1:500; Abcam), or goat anti-mouse IgG H&L (DyLight® 488) (# ab150117, 1:500; Abcam), and goat anti-rabbit IgG H&L (DyLight® 594) (# ab150084, 1:500; Abcam) for 40 min at 37 °C, and 4′,6-diamidino-2-phenylindole (# ab104139; Abcam) was used for nuclear localization. Pictures were examined under a confocal laser scanning microscope (LSM 800, ZEISS, Beijing, China).
2.9. Histopathology analysis
Surgically resected hearts were fixed in 4% paraformaldehyde solution and embedded in paraffin. Crosswise 4-μm-thick heart sections were cut, deparaffinized three times in dimethylbenzene, and rehydrated in 100%, 95%, 80% and 70% ethanol, respectively. Then these heart sections were subjected to hematoxylin&eosin staining. Masson trichrome staining was performed in accordance with the manufacturer's protocol (# G1340; Solarbio, Beijing, China). Pictures were taken with an Olympus BX53 microscope (Olympus, Beijing, China).
2.10. Quantitative reverse transcription-polymerase chain reaction (RT-PCR)
Total RNA was extracted from the left ventricle using Trizol reagent (# 15596026; ThermoFisher Scientific, New York, NY, USA). The mRNA levels of heart were detected by quantitative RT–PCR with SYBR Green Master Mix (# 4309155; ThermoFisher Scientific). Relative mRNA levels were determined according to 2-ΔΔCT method, and all data were normalized to Gapdh mRNA level. The PCR primers were obtained from Sangon Biotech (Shanghai, China). The sequences of primers used in the current study are listed as follows:
ANP: F 5′- GTGCGGTGTCCAACACAGAT-3′, R 5′- TCCAATCCTGTCAATCCTACCC-3′;
BNP: F 5′- GAGGTCACTCCTATCCTCTGG-3′, R 5′- GCCATTTCCTCCGACTTTTCTC-3′;
GAPDH: F 5′- AGGTCGGTGTGAACGGATTTG-3′, R 5′-TGTAGACCATGTAGTTGAGG TCA -3′.
2.11. Immunoprecipitation and Western blotting
Fresh heart tissues (around the left ventricular infarct area) were washed with phosphate buffer saline (PBS) twice and lysed with radio immunoprecipitation assay (RIPA) lysis buffer (# R0020; Solarbio), PMSF (# P0100; Solarbio), and N-ethylmaleimide (# 128287; Sigma-Aldrich, Ronkonkoma, NY, USA). Immunoprecipitation was performed in accordance with the manufacturer's protocol (# 88805; ThermoFisher Scientific). The lysate was lysed on ice for 30 min and centrifuged at 14,000 rpm for 15 min. Beads were incubated with antibody against JunD (# ab134067, 5 μg; Abcam) at room temperature on a rotation wheel for 1 h. Then the lysates were incubated with bead JunD complex for 2 h. The resulting protein complexes were separated and Western blotting experimental procedures were performed. Briefly, protein homogenates were separated by sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes, and then blocked with 5% skim milk for 1 h and incubated with anti-SUMO1 (# ab11672, 1:1000; Abcam), anti-JunD (# ab134067, 1:1000; Abcam), and anti-GAPDH (# ab9485, 1:2000; Abcam) at 4 °C overnight. Subsequently, the membrane was incubated with goat anti-rabbit IgG H&L horseradish peroxidase (HRP) (# ab6721, 1:20000; Abcam) and goat anti-mouse IgG H&L HRP (# ab205719, 1:20000; Abcam), and then visualized with the enhanced chemiluminescence reagents (# 34580; ThermoFisher Scientific).
2.12. Human umbilical vein endothelial cells (HUVECs) culture, cytokines, and transfection
HUVECs were purchased from Gibco (# C0035C; Gibco, Billings, MT, USA) and cultured in Dulbecco's modified Eagle medium (# 11995073; Gibco) with 10% fetal bovine serum (FBS) (# 10099141; Gibco) and 1% 10 kU/mL penicillin/10 mg/mL streptomycin at 37 °C in the 5% CO2-95% air atmosphere. Recombinant human VEGFA (# P00063; Solarbio) was used at 10 ng/mL for HUVEC treatment. Plasmid or siRNA transfection of HUVEC was performed by LipofectamineTM 2000 (# 11668019; ThermoFisher Scientific) or LipofectamineTM RNAiMAX (# 13778150; ThermoFisher Scientific) according to the manufacturer's protocol, respectively. Plasmid or siRNA was used to transfect HUVEC when approximately 60%–70% confluent and cells were added VEGFA at 48 h post-transfection for indicated experiments.
2.13. Proliferation and migration of HUVECs
Transfected HUVECs were cultured in 0.5% serum for starvation and then scratched with a 200 μL pipette tip. After that, the medium was replaced with fresh medium and the culture was continued for 16 h. Images were taken with Olympus BX53 microscope and the width of wound healing was calculated by ImageJ software.
2.14. HUVEC tube-formation assay
Twenty-microliters of matrigel (# 354234; Becton Dickinson, Boynton Beach, FL, USA) was added to each well of 96-well plates and incubated in a cell incubator for 30 min. Transfected HUVEC single cell suspension was added on top of matrigel to each well. After 30 min, fresh medium with 10 ng/mL of VEGFA was added and cultured for 6–18 h. Images were taken with Olympus BX53 microscope and numbers of branches and tubes were analyzed by ImageJ software.
2.15. Cell cycle assay
HUVECs were collected and incubated with pre-cooled 70% ethanol for 2 h. The cells were collected by centrifugation, added to PBS with 50 μg/mL propidium iodide (PI) (# ab14083; Abcam), 100 μg/mL RNase A, and 0.2% Triton X-100, and incubated at 4 °C for 30 min in the dark. Flow cytometry was performed with standard procedures, and the results were analyzed by the cell cycle fitting software FlowJo.
2.16. In vivo gene therapy
Recombinant adeno-associated virus (serotype 9) vectors carrying mouse SUMO1 (accession number NM_009460.2) and enhanced green fluorescent protein (EGFP) or ZsGreen with a c-TnT promoter (AAV9-CTnT-SUMO1-EGFP) or TIE promoter (AAV9-TIE-SUMO1-ZsGreen) (HanBio Inc., Shanghai, China) were used. AAV9-CTnT-EGFP or AAV9-TIE-ZsGreen was used as a negative control. Virus was delivered to mouse heart via tail vein injection (viral titer: 1 × 1012 VG/mL, 100 μL/per mouse). Virus infection efficiency was detected after 3 weeks and MI experiments were performed.
2.17. Statistical analysis
All molecular and biochemical data are reported as the mean ± standard deviation (SD). All physiological data are reported as the mean ± SD. Statistical differences between two groups were tested with the Student's t-test. Statistical differences between more than two groups were analyzed with one-way analysis of variance (ANOVA) tests, followed by Tukey's multiple comparison tests using GraphPad prism 7.0 (GraphPad, San Diego, CA, USA). P < 0.05 was considered statistically significant.
3. Results
3.1. SUMO1 is involved in the pathological process of the heart after MI
To determine whether SUMO1 is pathologically relevant to the repair response after MI, we used immunohistochemical staining to detect the SUMO1 level in heart sections from mice after MI. Seven days after MI is within the intermediate phase of the cardiac remodeling response, which can reflect the prognosis of the heart [29]. SUMO1 was not uniformly expressed in the heart sections of healthy mice, and the SUMO1 level was significantly increased 7 days after MI. However, not all heart cells showed high levels of SUMO1 enrichment (Figs. 1A and S1A), which suggested that SUMO1 was involved in the cardiac pathological repair process after MI. Immunofluorescence staining showed SUMO1 was abundantly enriched in the infarcted area. However, the SUMO1 level was significantly decreased in the border zone (mainly CMs) compared with that in the infarct zone (Figs. 1B and S1B), suggesting that SUMO1 levels were altered after MI, but the abundance in different cell types was inconsistent. To further explore the effect of SUMO1 on cardiac function after MI, β-galactosidase-targeted Bay Genomics XA024 ES cell-derived SUMO1−/− mice [18] and littermate WT mice were examined 7 days after MI. Representative echocardiographic images are presented in Fig. 1C. Both WT and SUMO1−/− mice exhibited approximately similar EF%, FS%, LVIDs, and LVIDd values at baseline, suggesting equal cardiac function before MI surgery. However, SUMO1−/− mice displayed significantly worse cardiac function with regard to EF% and FS% compared with those values in WT mice (Figs. 1C–E, S1C, and S1D, and Table S1). 2,3,5-triphenyltetrazolium chloride (TTC) staining and hematoxylin&eosin staining showed that SUMO1−/− mice displayed larger infarct sizes with more severe adverse ventricular remodeling after MI than those in WT mice (Figs. 1F and G, and S1E–S1H). These findings indicated that SUMO1 deficiency in the heart exacerbated MI-induced adverse cardiac remodeling and systolic dysfunction.
Fig. 1.
SUMO1 is necessary for the pathological repair process after MI. (A, B) Representative images of immunohistochemistry (A) and immunofluorescence (B) of SUMO1 in heart sections from mice after MI. (C) Representative echocardiographic M-mode images of WT and SUMO1−/− mice. (D, E) LVEF (D) and LVFS (E) were quantified via echocardiography (n = 8). (F) Representative images of five heart sections from one heart after MI (n = 6), followed by TTC staining. (G) Representative images of hematoxylin&eosin staining in hearts after 7 days of MI. Mean ± standard error of mean (SEM), ∗∗P < 0.01, ∗∗∗P < 0.001 vs. WT. Sham or WT. MI. MI: myocardial infarction; WT: wild-type; LVEF: left ventricular ejection fraction; LVFS: left ventricular fraction shortening; TTC: 2,3,5-triphenyltetrazolium chloride.
3.2. SnRNA-seq identified four major heart cell populations
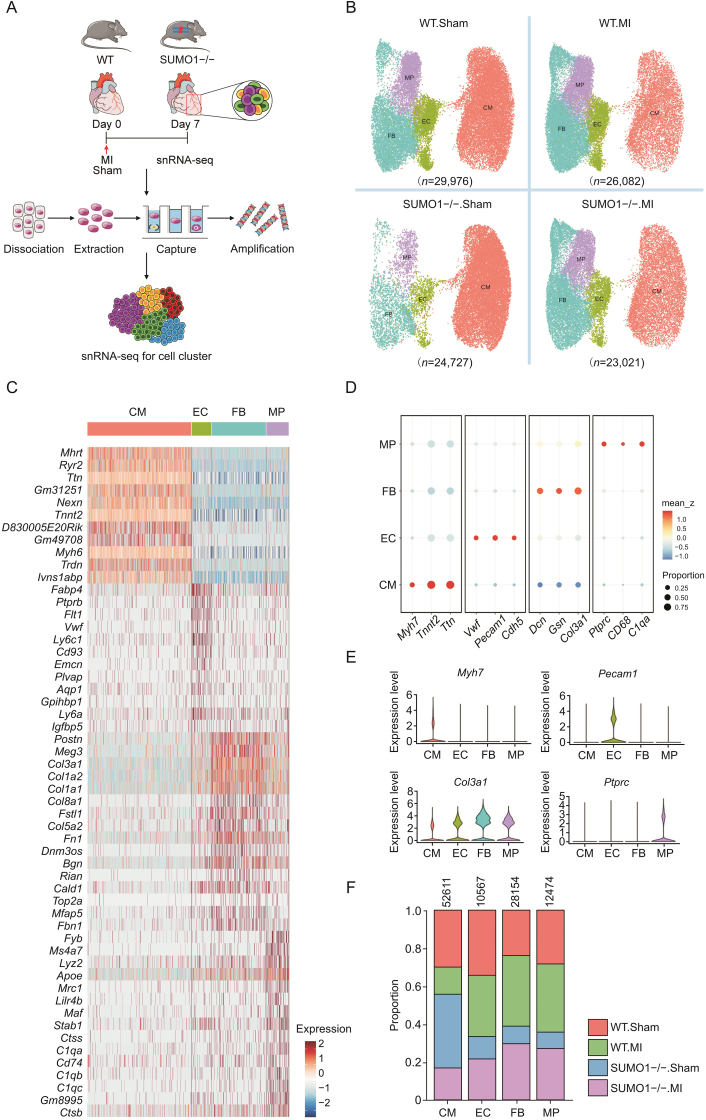
Pathological processes of MI involve multiple cell types [30]. We applied unbiased snRNA-seq to identify cell populations in the hearts of WT and SUMO1−/− mice on day 7 after the sham operation or MI (Fig. 2A). After filtration and quality control, 103,806 cells were classified into four cell types according to specific markers, including CMs, ECs, FBs, and MPs, and these groups displayed specific gene expression signatures (Figs. 2B and C). Additionally, we analyzed the level of SUMO1 in the distinct subset and showed that the expression of SUMO1 was influenced by cell types (Fig. S2). We used classic cell markers to identify specific cell populations (Myh7, Tnnt2, and Ttn for CMs; Pecam1, Cdh5, and Vwf for ECs; Col3a1, Gsn, and Dcn for FBs; CD68, Ptprc, and C1qa for MPs) (Figs. 2D and E). Importantly, based on the relative abundance of sequenced cells from each group within each cell type, we detected an altered abundance of four different cell types in response to MI (Fig. 2F). We compared the proportions of different cell types in WT and SUMO1−/− mice in the Sham and MI groups. CMs were markedly lost after MI, and SUMO1−/− mice showed a greater proportion of CM loss than WT mice, suggesting that SUMO1 deletion led to a decreased compensatory ability of CMs during ischemic-hypoxic injury. In addition, FBs, ECs, and MPs in SUMO1−/− mice had stronger proliferative abilities after MI than those in WT mice (Table S2). These results revealed that SUMO1 exerted different regulatory effects on various cell types in the heart after MI.
Fig. 2.
Overview of the 103,806 single cells isolated from WT mice and SUMO1−/− mice. (A) Schematic overview of cardiac cells isolation for single nuclei RNA sequencing (snRNA-seq). (B) UMAP view of cell types from MI model of WT and SUMO1−/− mice. (C) Heatmap of top marker genes per cell type. (D, E) Dot plot (D) and violin plots (E) showing specificity genes for cell clusters. (F) Fraction of each cardiac cell population relative to all cardiac cell clusters in each group. WT: wild-type; MI: myocardial infarction; UMAP: uniform manifold approximation and projection; CM: cardiomyocytes; EC: endothelial cells; FB: fibroblasts; MP: macrophages.
3.3. SUMO1 deficiency increased Nppa+Nppb+Ankrd1+CM cluster after MI
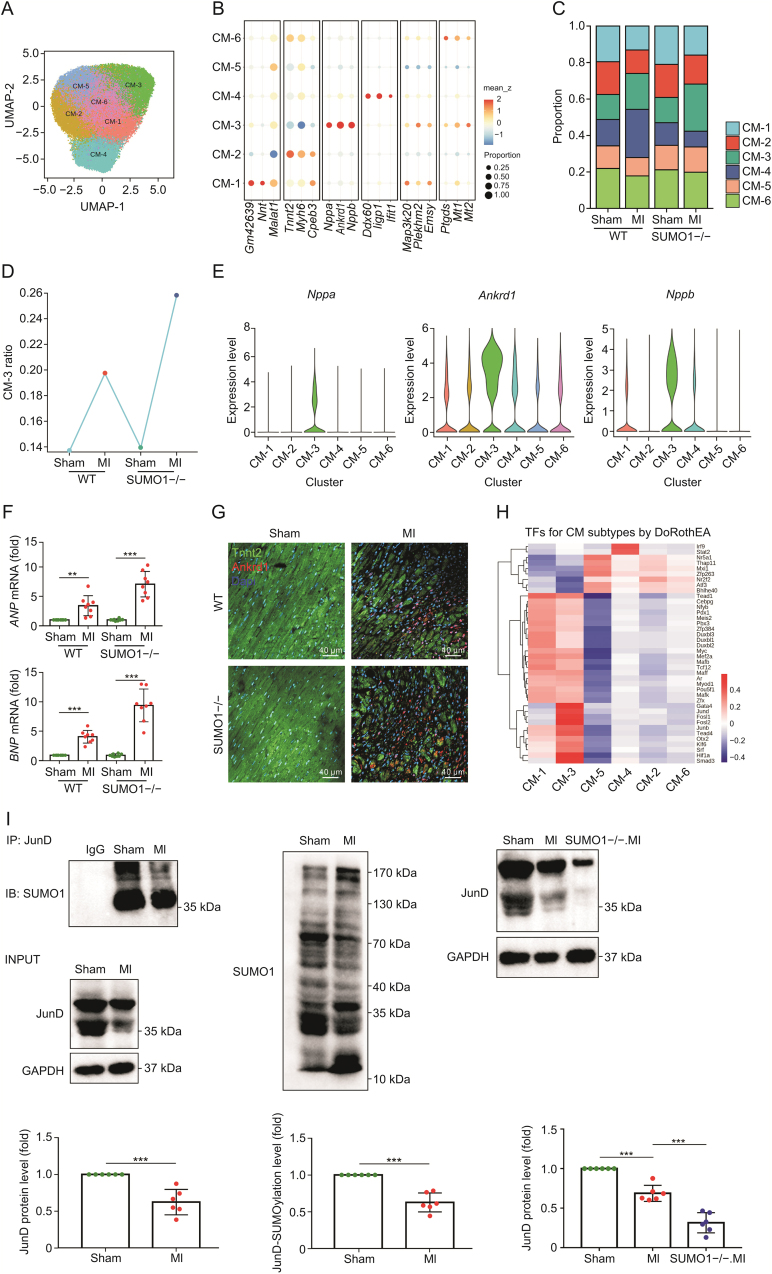
CMs are the predominant cell type that determines cardiac fate after MI; therefore, we explored the key CM subsets affected by SUMO1 after MI by reclustering all CMs. The 52,611 CMs derived from all samples clustered into six distinct CM subsets with a high degree of robustness and displayed individual gene expression profiles (Figs. 3A, and S3A–C, and Table S3). In addition to the canonical marker genes, CM clusters also expressed identity-specific genes. For example, the CM-1 cluster characteristically expressed Nnt (encoding an integral protein in the inner mitochondrial membrane involved in the energy supply) [31]. Substantial loss of CM-1 cluster after MI correlated with impaired cardiac function (Figs. 3B, 3C, S3B and S3C). Importantly, we found that the CM-3 cluster specifically expressed a variety of stress-related genes (Nppa, Nppb, Ankrd1), which are indicators of a poor prognosis of heart disease [32]. Notably, a recent study [33] showed that Ankrd1, a stress marker, was present around the infarcted area, suggesting that the CM-3 cluster might serve as a landmark for CMs around the injured area, and an increased proportion of the CM-3 cluster might indicate injury aggravation. The up-regulated genes in the CM-3 cluster were mainly involved in the left ventricular hypertrophic response pathways by DisGeNet ORA analysis (Figs. S3C–F). The proportion of the CM-3 cluster (Nppa+ Nppb+ Ankrd1+) was significantly increased after MI, and SUMO1−/− mice showed a greater proportion of the CM-3 cluster than WT mice (Figs. 3D and E). SUMO1−/− mice had higher mRNA levels of atrial natriuretic peptide (ANP) and B-type natriuretic peptide (BNP) after MI than those in WT mice (Fig. 3F). Additionally, the ANP and BNP levels positively correlated with local wall stress and resulted in a larger infarct size [34]. Using immunofluorescence staining, we found that SUMO1−/− mice presented more Ankrd1+CMs in the border region after MI than those in WT mice (Fig. 3G), suggesting that SUMO1 deletion increased the proportion of the CM-3 cluster after MI. Surprisingly, we found that MI triggered an increase in the CM-4 cluster proportion, but this was reversed in SUMO1−/− mice. The CM-4 cluster mainly expressed interferon-related genes including Ddx60, Iigp1, and Ifit1, suggesting a transcriptomic response to interferon (Figs. 3B and C, and S3C). The CM-4 cluster might be involved in immune surveillance in the heart, and SUMO1−/− might impair this effect.
Fig. 3.
Deletion of SUMO1 promoted the proportion of Nppa+Nppb+Ankrd1+ CM subcluster. (A) UMAP visualization of clustering revealed six distinct CM populations. (B) Dot plot showing enriched genes for each CM subclusters. (C) Fraction of each CM subcluster relative to all CMs in each group. (D) Line plot showing changes in the relative fractions of CM3 cluster in each group. (E) Violin plots of natriuretic peptide A (Nppa), natriuretic peptide B (Nppb) and ankyrin repeat domain 1 (Ankrd1) in cardiomyocyte clusters. (F) The atrial natriuretic peptide (ANP) and brain natriuretic peptide (BNP) mRNA levels in WT and SUMO1−/− mice after Sham and MI (n = 8). (G) Representative images of immunofluorescence of cardiac troponin T (Tnnt2) (green) and Ankrd1 (red) in heart sections from mice after MI. (H) Heatmap of top 40 highly variable transcription factor among the six cardiomyocyte subclusters. The z-scores of TF activities are colour-coded. (I) The heart homogenates were immunoprecipitated with anti-JunD agarose beads and probed with anti-SUMO1 antibody (n = 6). Mean ± standard error of mean (SEM), ∗∗∗P < 0.001 vs. Sham or MI. UMAP: uniform manifold approximation and projection; CM: cardiomyocyte; MI: myocardial infarction; WT: wild-type; TF: transcription factor.
Gene expression of different CM clusters is regulated by transcription factors (TFs) [35]. TF activity analysis of the snRNA-seq data revealed that the CM-1 and CM-3 clusters had similar binding activity of known CM lineage-specific TFs such as Meis2, Mef2a, and Tead4. However, the CM-3 cluster showed increased binding activity of stress-related TFs, such as the Jun subunit (JunB and JunD) and Fos subunit (Fosl1 and Fosl2) in the AP-1 complex (Fig. 3H). JunD is a negative regulator of CM hypertrophy and reduces ANP and BNP expression in response to hypertrophic stimuli [36]. We examined the binding abundance of JunD to SUMO1 after MI by co-immunoprecipitation. We found that both the SUMOylation level and the expression level of JunD decreased after MI. SUMO1 deletion led to a further decrease in JunD expression after MI compared with that in WT mice (Fig. 3I). Reduced binding of SUMO1 to JunD resulted in decreased JunD stability, which increased Nppa/Nppb transcription after MI. These results revealed that SUMO1 was involved in myocardial injury mainly by regulating the CM-3 cluster that existed in the border zone. Knockout of the SUMO1 gene increased the proportion of the CM-3 cluster and the expression levels of myocardial injury-related genes, and JunD was involved in this process.
3.4. Deletion of the SUMO1 gene inhibited the conversion of proliferative FB clusters to collagen secreting FB clusters after MI
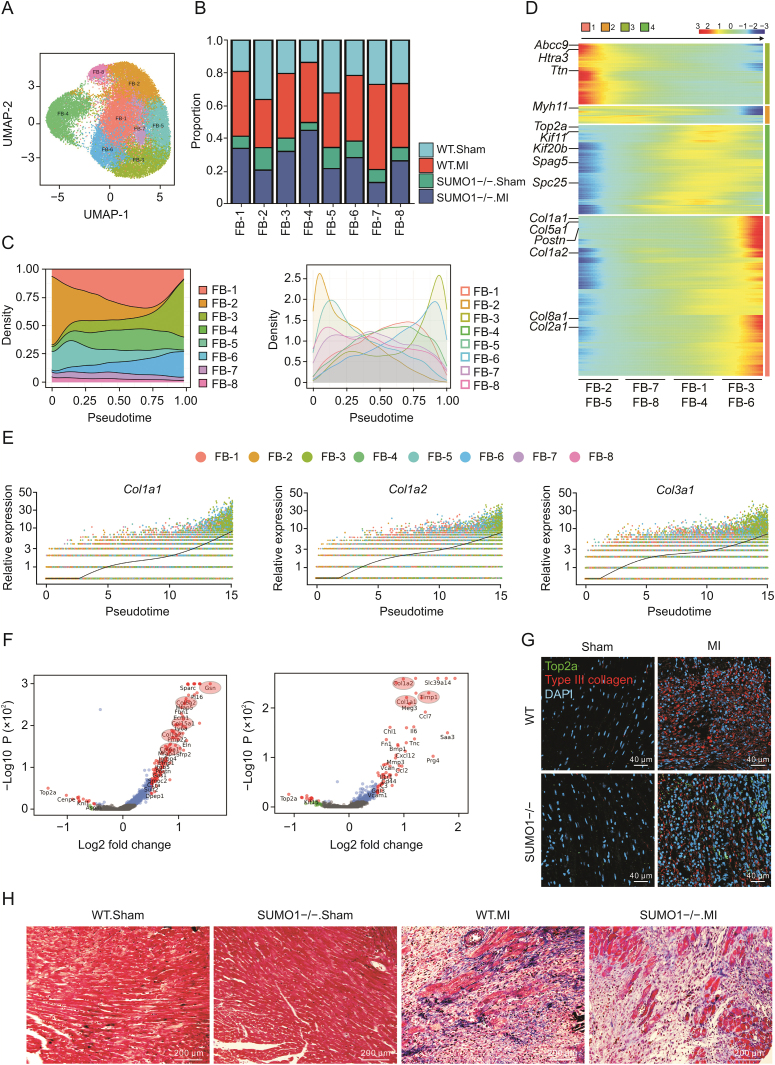
FBs in MI are heterogeneous [37]. To explore the role of FB subsets in cardiac remodeling after MI, we subdivided 28,154 individual FBs into eight clusters (FB1–8) (Figs. 4A, S4A, and S4B, and Table S3). FB subsets substantially proliferated in response to myocardial damage after MI, including the FB-1, FB-3, FB-4, FB-6, FB-7, and FB-8 clusters (Figs. 4B and S4C). However, the FB-2 and FB-5 cluster proportions were no significant difference after MI compared with those in the Sham group. We also analyzed the characteristic genes of each subset to explore the function of each FB cluster in depth (Fig. S4D). Transdifferentiation of FBs to myofibroblasts plays a major role in cardiac repair after MI. We used CytoTrace to analyze the temporal trajectory of FBs to explore FBs differentiation with MI progression (Figs. 4C, and S4E–G). We found that the FB-2 and FB-5 clusters were the initial state FBs subsets (maintained normal cardiac function) because of the low expressions of Postn, Col1a1 and Col1a2 genes, which act as hallmarks of myofibroblast transformation [38]. FBs clusters were ordered based on common kinetics using pseudotime in CytoTrace (Fig. 4D, Tables S4 and S5). The FB-3 and FB-6 clusters were identified as end-stage FBs because of high expression levels of collagen-related genes (Col1a1, Col1a2, and Col3a1) that were mainly involved in extracellular matrix-related pathways (Figs. 4E, 4F, S4H, and S4I). Importantly, we found that the proportions of the FB-3 and FB-6 clusters were reduced in SUMO1−/− mice after MI compared with those in WT mice. However, deletion of the SUMO1 gene significantly increased the proportion of the FB-4 cluster, which served as a subset in the intermediate state of FB differentiation. The FB-4 cluster highly expressed proliferation-related genes (Top2a, Mki67, and Kif11) [39], and Reactome ORA analysis showed that the FB-4 cluster was mainly involved in the cell cycle and mitosis (Figs. S4J and K), indicating that FB-4 was an FB subset involved in the proliferating state. Immunofluorescence staining using Top2a and type III collagen demonstrated that SUMO1−/− resulted in a high proportion of FB proliferation in the infarcted area after myocardial injury but decreased collagen secretion compared with that in WT mice (Fig. 4G). Using Masson staining, immunohistochemical staining for collagen types I and III showed that SUMO1−/− reduced collagen secretion after MI, and SUMO1 deletion triggered partial rupture in the infarct area, indicating insufficient repair capacity (Figs. 4H, S4L, and S4M). These findings suggested that multiple FB subsets were involved in the post-MI repair response, and SUMO1−/− inhibited the differentiation of FBs from a proliferative state to a collagen-secreting state.
Fig. 4.
Knockout of SUMO1 inhibited the conversion of proliferative FB subcluster to collagen-secreting FB subcluster after MI. (A) UMAP visualization of clustering revealed eight distinct FB populations. (B) Fraction of each FB subcluster relative to all FBs in each group. (C) Focused CytoTrace pseudotime analysis including all the FB subclusters. (D) Heatmap of differentially expressed genes, ordered based on their common kinetics through pseudotime CytoTrace. (E) Expressions of Col1a1, Col1a2 and Col3a1 along the pseudotime CytoTrace in all FB subclusters. (F) Volcano plot of specifically expressed genes in FB-3 (left) and FB-6 (right) clusters. (G) Representative images of immunofluorescence of DNA topoisomerase II alpha (Top2a) (green) and type III collagen (red) in heart sections from mice after MI. (H) Representative images of Masson's trichrome staining in hearts after 7 days of MI. UMAP: uniform manifold approximation and projection; FB: fibroblast; MI: myocardial infarction; WT: wild-type; Col1a1: collagen type I alpha 1 chain; Col1a2: collagen type I alpha 2 chain; Col3a1: collagen type III alpha 1 chain; Top2a: DNA topoisomerase II alpha.
3.5. Loss of SUMO1 promoted proliferation of EC subsets with revascularization ability after MI
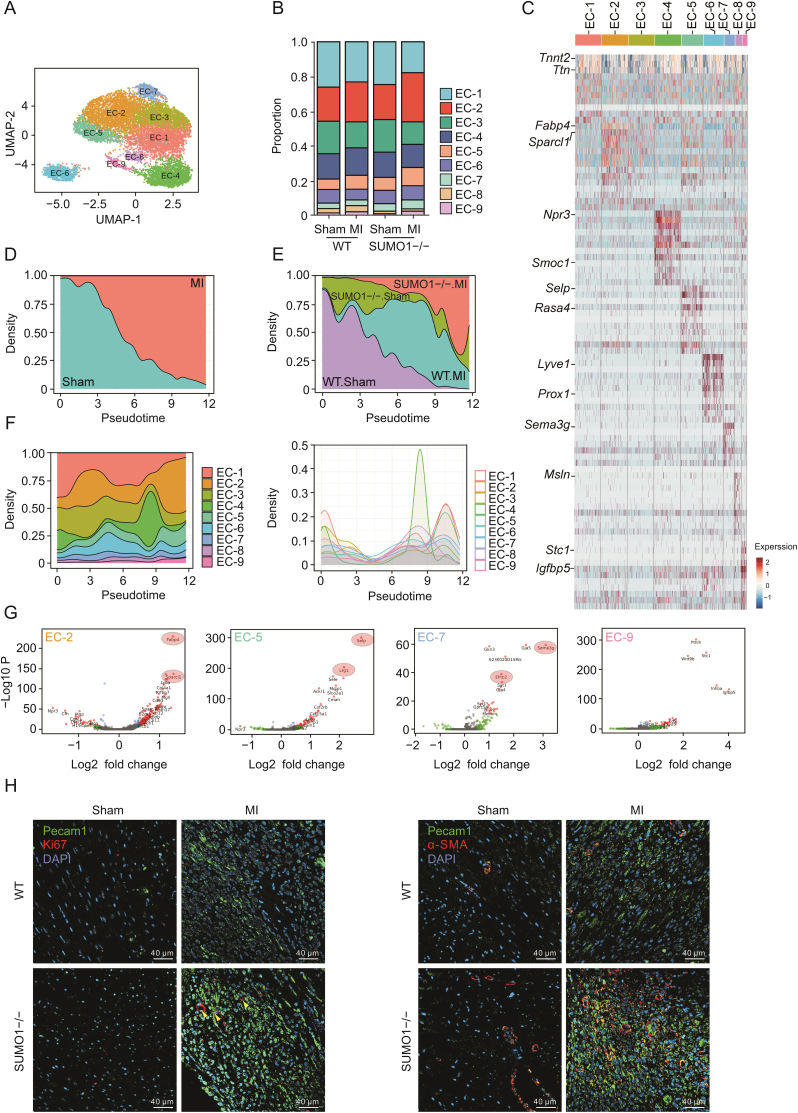
Reconstruction and recovery of the neovascular network at the infarct site after myocardial injury is critical to improve the prognosis of MI [3]. Vascular homeostasis and neovascularization after MI are regulated by ECs. The 10,567 ECs derived from all samples clustered into nine distinct EC subsets with a high degree of robustness (Figs. 5A, S5A, and S5B). We found that the EC-2, EC-5, EC-8, and EC-9 proportions increased after MI, whereas the EC-1 and EC-3 proportions decreased relative to those in the Sham group, indicating the heterogeneity of ECs and the involvement of EC subsets in different functions after MI (Fig. 5B). We identified EC clusters by characteristically expressed genes (Fig. 5C). The EC-1 cluster highly expressed CM-like signature genes (Tnnt2 and Ttn). This EC cluster was previously demonstrated among adult cardiac ECs in response to myocardial contraction and cardiac pumping [10]. The EC-2 cluster strongly expressed Fabp4, a coronary vascular EC marker [40]. The EC-2 cluster also expressed Sparcl1, a tissue repair and angiogenesis marker previously identified in the injury response, suggesting that the EC-2 cluster might act as coronary ECs during revascularization in response to myocardial injury [41]. The EC-4 cluster characteristically expressed Npr3 and Smoc1, which were detected in the endocardium of mice, indicating an endocardial-derived EC subset [10,40]. The EC-5 cluster mainly expressed Selp and Rasa4, which were previously identified as markers expressed by subsets of activated ECs [42]. The EC-6 cluster mainly expressed characteristic lymphatic vessel genes (Lyve1 and Prox1) [43]. Lymphatic vessels served as conduits for the maturation and delivery of immune cells, promoting the clearance of inflammatory cells during the remodeling phase after MI. In addition to canonical EC marker genes, the EC-7 cluster also expressed Sema3g, which was involved in vascular development, angiogenesis, and remodeling of new blood vessels in response to myocardial ischemia and hypoxia [44,45]. Additionally, Sema3g + ECs were recently shown to be arterial/arteriole ECs and involved in vascular remodeling [46,47]. Using Monocle pseudotime to analyze the temporal trajectory of EC subsets, we found that MI induced EC differentiation and that SUMO1−/− promoted EC differentiation because the SUMO1−/− group aggregated in the terminal state (Figs. 5D and E, S5C, and S5D). We showed the transition from basal states (EC-1 and EC-3 clusters) to intermediate states (EC-4 cluster) and terminal states (EC-2, EC-5, and EC-9 clusters) by pseudo-temporal analysis of EC subsets (Fig. 5F). Importantly, we found that SUMO1 loss increased the proportions of the EC-2, EC-5, EC-7, and EC-9 clusters after MI, most of which were in a terminal state. These clusters were mainly responsible for EC activation, coronary/arteriole revascularization, and vascular regeneration, which were beneficial for cardiac function after MI (Figs. 5G, S5E, S5F, and S6). To further explore the effect of SUMO1 deletion on ECs after MI, we performed immunofluorescence staining with Pecam1 and Ki67 (cell proliferation indicators). We found that loss of the SUMO1 gene promoted the distribution of Ki67+ ECs in the infarcted area after MI. Furthermore, Pecam1 and α-SMA are typical biomarkers of vascular ECs and parietal cells, respectively [48]. Immunofluorescence results showed that SUMO1 gene loss induced the formation of numerous small blood vessels in the infarcted area (Fig. 5H). These results revealed that SUMO1−/− was beneficial to EC proliferation, thereby promoting neovascularization and revascularization in the infarcted area.
Fig. 5.
Loss of SUMO1 promoted the proliferation of EC subcluster with revascularization ability after MI. (A) UMAP visualization of clustering revealed nine distinct ECs populations. (B) Fraction of each EC subcluster relative to all ECs in each group. (C) Heatmap showing gene expression signatures for each EC subset. (D, E) Focused CytoTrace pseudotime analysis including pre-MI (D) and post-MI and each group (E). (F) Focused CytoTrace pseudotime analysis including all the FB subclusters. (G) Volcano plot of specifically expressed genes in EC-2, EC-5, EC-7 and EC-9 cluster. (H) Representative images of immunofluorescence of Pecam1 (green), Ki67 (red) (left) and Pecam1 (green), α-SMA (red) (right) in heart sections from mice after MI (n = 6). UMAP: uniform manifold approximation and projection; EC: endothelial cells; MI: myocardial infarction; WT: wild-type; α-SMA: alpha smooth muscle actin; Ki67: marker of proliferation Ki-67; Pecam1: platelet and endothelial cell adhesion molecule 1.
3.6. SUMO1 blocked neovascularization by inhibiting VEGFA signaling
Communication between cells in cardiac tissue through cytokines and membrane proteins ensures efficient and orderly cardiac function. Receptor-ligand-mediated intercellular communication is critical for coordinating multiple biological processes including development, differentiation, and stress [49]. To further investigate the communication network between different cell types in the heart, we performed a comprehensive and systematic analysis of cell-to-cell communication molecules using CellPhoneDB (Fig. 6A). We found complex regulatory interactions between various cell types, including autocrine and paracrine effects (Fig. 6B). Interestingly, the EC2, EC-5, and EC-9 clusters, as EC subsets responsible for EC activation, coronary/arteriole revascularization, and revascularization, were regulated by VEGFA secreted by CMs (Fig. 6C). Notably, the proportions of EC-2, EC-5, and EC-9 clusters were increased in SUMO1−/− mice after MI compared with those in WT mice. VEGFA induces angiogenesis by stimulating EC activation, migration, and capillary-like structure formation [50,51]. To investigate whether SUMO1−/− promotes VEGFA-mediated EC proliferation, we transfected SUMO1 silencing (siSUMO1) and SUMO1 overexpression plasmids into human umbilical vein ECs (HUVECs) (Figs. 6D and E, and S7). SUMO1 silencing significantly enhanced the proliferation and migration ability of VEGFA-induced HUVECs in vitro (Figs. 6F and G). To further determine whether SUMO1 mediates the angiogenic signal of VEGFA, a matrigel tube formation assay was performed to examine the effect of SUMO1 on EC cord formation. Silencing of SUMO1 in HUVECs significantly promoted VEGFA-induced EC tubule formation, whereas overexpression of SUMO1 resulted in a reduction in the number of tubes and branches (Figs. 6H and I). In addition, SUMO1 gene silencing increased the VEGFA-mediated G2/M phase ratio in HUVECs (Figs. 6J and K). These findings suggested that the effect of VEGFA on promoting EC proliferation was inhibited by SUMO1, which was detrimental to angiogenesis and revascularization after MI.
Fig. 6.
SUMO1 inhibited VEGFA signaling in endothelial cells and restrained neovascularization. (A) Heatmap showing logarithmic interaction scores between all cell subsets. (B, C) Unique ligand-receptor interactions (B), and the specific ligand-receptor pair (C) for all cell subclusters. (D) Schematic overview of SUMO1 knockdown/overexpression experiments in proliferative HUVEC. (E) The SUMO1 mRNA levels after transfected SUMO1 plasmid (n = 6). Data expressed as fold change of SUMO1/GAPDH vs. Sham. (F, G) Cell scratch assay was used to measure EC migration (F), and wound healing (% closure) was quantified (n=6) (G). (H, I) Representative images of EC tube formation (H), and number of branches (left) and tubes (right) were quantified (n=6) (I). (J , K) Cell cycle analysis by propidium iodide staining and flow cytometry for HUVEC (J), and percentage of HUVEC in G2/M phase of the cell cycle (n=6) (K). Mean ± SEM, ∗P < 0.05, ∗∗P < 0.01, ∗∗∗P < 0.001 vs. Ctrl or siCtrl. VEGFA: vascular endothelial growth factor A; HUVEC: human umbilical vein endothelial cell; CM: cardiomyocyte; EC: endothelial cell; Ctrl: Control.
3.7. CM-specific SUMO1 expression via gene therapy attenuated pathological responses after MI
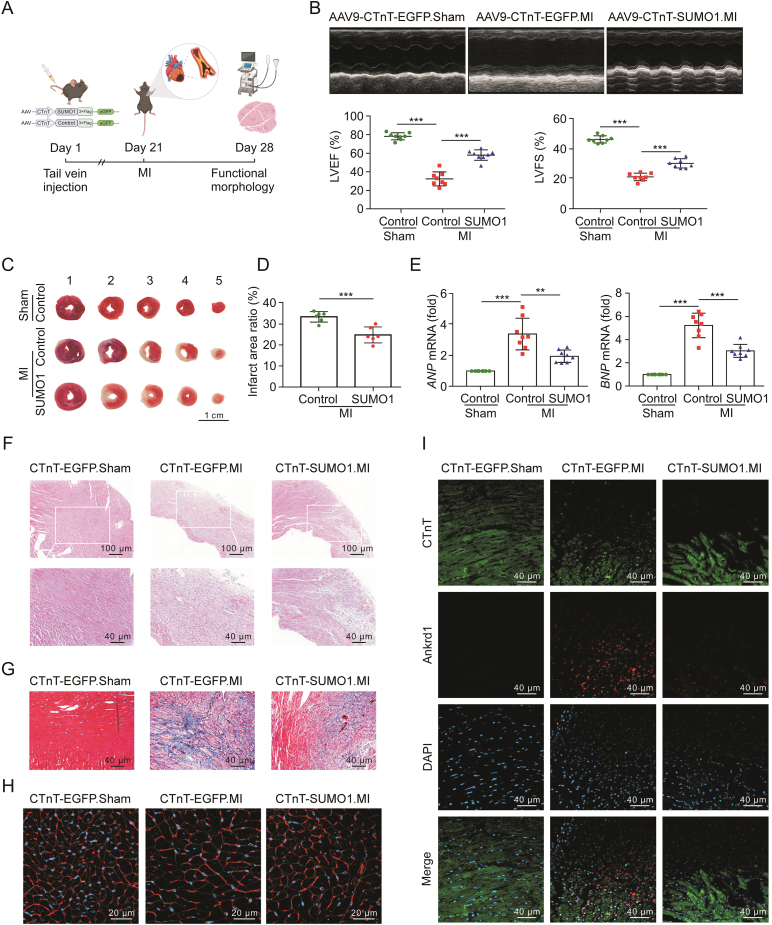
SUMO1 exerted different effects on various types of heart cells, and SUMO1 deletion triggered CM damage and cardiac dysfunction after MI. Therefore, to determine whether CM-specific SUMO1 gene therapy is effective as a preventative therapeutic strategy, we injected adeno-associated virus (AAV) into the tail vein of 8-week-old mice and performed MI or sham surgery 21 days later (Figs. 7A, and S8A). Under MI conditions that promote cardiac dysfunction, AAV-CTnT-enhanced green fluorescent protein (eGFP)-infected mice displayed typical MI-induced responses including contractile dysfunction (EF%, FS%) and increased infarct size (Figs. 7B–D, and S8B, and Table S6). AAV-CTnT-SUMO1 transduced mice were more resistant to MI-induced abnormal heart function and had significantly less cardiac dysfunction and a lesser degree of myocardial damage (Figs. 7B and C and S8B, and Table S6) relative to AAV-CTnT-EGFP controls. CM-specific overexpression of SUMO1 significantly reduced ANP and BNP mRNA levels after MI compared with those in the control group (Fig. 7E). Hematoxylin&eosin and Masson staining showed that AAV-CTnT-SUMO1 transduced mice displayed a smaller infarct size with improved ventricular remodeling and significantly reduced fibrosis levels after MI compared with those in AAV-CTnT-EGFP mice (Figs. 7F, 7G, S8C, and S8D). AAV-CTnT-SUMO1 transduced mice showed a significantly reduced CM area in the border zone compared with that in AAV-CTnT-EGFP controls (Figs. 7H and S8E). Immunofluorescence showed that AAV-CTnT-SUMO1 transduced mice displayed fewer Ankrd1+CMs (and higher JunD+CMs) distributed around the infarct area than those in AAV-CTnT-EGFP controls (Figs. 7I and S8F). These findings indicated that overexpression of SUMO1 in CMs alleviated cardiac remodeling and enhanced cardiac function under pathological conditions.
Fig. 7.
Cardiomyocyte-specific SUMO1 gene therapy attenuated MI-induced cardiac dysfunction and fibrosis pathological process. (A) Schematic of experimental design of cardiomyocyte-specific SUMO1 delivery and MI surgery on mice. (B) Representative echocardiographic M-mode images of AAV-CTnT-EGFP-infected mice and AAV-CTnT-SUMO1-infected mice. LVEF and LVFS were quantified via echocardiography (n = 8). (C, D) Representative images of five heart sections from one heart after MI (C) and image-based quantification analysis of infarct size (n = 6 for each group) (D), followed by 2,3,5-triphenyltetrazolium chloride (TTC) staining. (E) The atrial natriuretic peptide (ANP) and brain natriuretic peptide (BNP) mRNA levels in AAV-CTnT-EGFP-infected mice and AAV-CTnT-SUMO1-infected mice after Sham and MI (n = 8). Data expressed as fold change of ANP/GAPDH and BNP/GAPDH vs. Sham. (F) Representative images of hematoxylin&eosin in hearts after 7 days of MI. (G) Representative images of Masson's trichrome staining in hearts after 7 days of MI. (H) Representative images of wheat germ agglutinin (WGA) in heart sections from mice after MI. (I) Representative images of immunofluorescence of CTnT (green) and Ankrd1 (red) in heart sections from mice after MI. Mean ± standard error of mean (SEM), ∗∗P < 0.01, ∗∗∗P < 0.001 vs. Control or MI. MI: myocardial infarction; AAV: adeno-associated virus; CTnT: cardiac troponin; EGFP: enhanced green fluorescent protein; LVEF: left ventricular ejection fraction; LVFS: left ventricular fraction shortening.
Based on the above findings, to further explore the effect of EC-specific SUMO1 gene therapy, we used AAV viruses for EC-specific SUMO1 delivery and performed MI surgery on mice (Figs. S9A and B). AAV-TIE-SUMO1 transduced mice showed more severe adverse cardiac function after MI relative to the AAV-TIE controls (Figs. S9C–E, and Table S7). Immunofluorescence showed that AAV-TIE-SUMO1 transduced mice displayed fewer small blood vessels around the infarct area than AAV-TIE controls (Fig. S9F). In summary, mice preinjected with CM-specific AAV-SUMO1instead of EC-specific AAV-SUMO1, exhibited improved cardiac function following MI.
4. Discussion
SUMOylation is a type of reversible post-translational modification that modifies protein structure by covalently binding to lysine residues of substrates, thereby regulating enzymatic activity, protein stability, and localization [6]. In recent years, the number of studies of SUMOylation in heart disease have increased, and many SUMOylation-dependent regulatory mechanisms have been described in the context of cardiac hypertrophy [52], heart failure [13,16], and myocardial ischemia-reperfusion injury [53]. However, studies on SUMOylation of the whole heart neglected the effect of SUMO on different cell subsets during the remodeling phase after MI. Here, we found that SUMO1 was critical for post-infarction heart repair and that deletion of the SUMO1 gene aggravated myocardial injury after MI. Differential regulatory effects of SUMO1 on various cell types in the heart were discriminated using snRNA-seq. We identified a subset of CMs representing cardiac injury and characterized their gene expression signature. We further analyzed the gene expression signature of FBs and ECs during cardiac repair after MI. Importantly, depletion of SUMO1 had different effects on CMs, FBs, and ECs, suggesting the potential of SUMO1 as a therapeutic target after MI based on heart cell specificity. Our findings demonstrated the cell-specific actions of SUMO1-mediated signaling in injury and repair processes and highlighted the crucial role of SUMO1 in protecting the infarcted heart (Fig. 8).
Fig. 8.
Potential mechanisms of SUMO1 in different heart cell types after myocardial infarction (MI). In cardiomyocytes, SUMO1 represses Nppa and Nppb transcription and reduces the distribution of Nppa+ Nppb+cardiomyocyte (CM) in injured myocardium in a JunD-dependent manner after MI. In fibroblasts, SUMO1 mediated cardiac repair, promoting the conversion of proliferating fibroblasts to collagen-secreting fibroblasts after myocardial damage. In endothelial cells, SUMO1 inhibited vascular endothelial growth factor A (VEGFA)-mediated angiogenic signaling and inhibited FABP+ endothelial cell (EC) proliferation and revascularization after MI.
Recently, several scRNA sequencing (scRNA-seq) studies have examined the response of heart cells at different stages after MI, including the inflammatory, repair, and healing phases. For example, scRNA-seq of >25,000 cardiac collagen1α1-GFP+ FBs from sham-operated and post-infarct hearts revealed FB heterogeneity and dynamics and determined that the CTHRC1+ FB cluster was a key subset for scar healing [54]. In another study, scRNA-seq of Cdh5+Pecam1+ ECs after MI showed hypoxia-induced activation of the mesenchymal state of ECs with metabolic adaptation, highlighting the heterogeneity of cardiac ECs [55]. Our previous study identified MP heterogeneity and predominant cardiac infiltrating MP subsets by snRNA-seq analysis of CD45+ immune cells in mouse hearts after MI [19]. Here, we focused on non-immune cells and identified CMs, FBs, and ECs subset heterogeneity and multiple functions of different subsets during post-MI repair. For example, the Nppa+ Nppb+ Ankrd1+ CM cluster (CM-3 cluster) is an indicator subpopulation of myocardial injury, and the expansion of its proportion can reflect a worse prognosis after MI. Ankrd1 was previously found to be expressed in the border zone of the infarcted heart, and an increased in Ankrd1+CMs indicated an enhanced response to stress and myocardial injury [32]. Deletion of the SUMO1 gene increased the proportion of the CM-3 cluster and resulted in a larger infarct size after MI, suggesting that SUMO1 antagonized CM damage and stress responses after myocardial injury. Using DoRothEA for transcription factor activity analysis, we found that specific transcription factors were strongly associated with the CM-3 cluster, such as JUN (JunB and JunD) and FOS (Fosl1 and Fosl2), which were involved in AP-1 signaling. Importantly, SUMOylation of JunD enhanced JunD stability and inhibited transcription of Nppa and Nppb, which was critical for reducing the proportion of the CM-3 cluster after MI. The specific overexpression of SUMO1 in CMs via gene therapy resulted in an improved prognosis after MI. Mice preinjected with CM-specific AAV-SUMO1 had suppressed expressions of the ANP and BNP genes and the distribution of Ankrd1 in the border zone, suggesting a focus on SUMO1 expression strategies targeting CMs to treat post-MI injury.
Regarding FBs, transdifferentiation into myofibroblasts after MI is crucial for FB to exert post-infarction repair effects [56]. The conversion process of FBs from a basal state (maintaining the extracellular matrix homeostasis) to a terminal state (secreting collagen to stabilize the heart) is accompanied by distinct gene expression patterns. For example, the transition from low-expressing collagen-related genes clusters (FB-2 and FB-5) to Col1a1+Col3a1+ clusters (FB-3 and FB-6) indicated a dynamic transformation process of cardiac FBs after MI. However, deletion of the SUMO1 gene blocked the transition from FBs to myofibroblasts. We found an expansion of the proportion of the FB-4 subset in SUMO1−/−.MI mice. FB-4, as a precursor cell subset of myofibroblasts, is a prerequisite for cardiac collagen secretion and deposition in the late stage of myocardial infarction. SUMO1 deletion blocked the transition from FBs to myofibroblasts by inhibiting conversion of the FB-4 cluster to the FB-3 cluster and FB-6 cluster, which resulted in insufficient cardiac collagen secretion and cardiac rupture in post-MI mice. These findings were confirmed by experiments examining fiber deposition and collagen expression. Previously, the type I transforming growth factor-beta (TGFβ) receptor was shown to be modified by SUMO1 in response to TGFβ stimulation, thereby enhancing Smad family member (Smad) activation and downstream transcriptional responses [57]. TGFβ is known to stimulate progression of FBs to the myofibroblast transformation and involved in the development of myocardial fibrosis [58]. Our results indicated that SUMO1 loss inhibited the transition of FBs from a proliferative state to a collagen-secreting state and that the FB-4 cluster was the critical FB cluster that responds to the arrest of FB transformation triggered by SUMO1 loss. Future studies should focus on key factors other than TGFβ that drive the transformation of FBs to myofibroblasts and the effect of SUMO1 on these factors.
Surprisingly, the regulatory effect of SUMO1 on ECs subset was identified. We found that SUMO1−/− increased the proportion of EC-2, EC-5, EC-7, and EC9 clusters after MI, which were broadly involved in EC activation and angiogenesis after myocardial injury, favoring vascular remodeling in the infarcted heart region. In particular, the EC-2 cluster, a subpopulation of coronary/arteriole ECs that respond to revascularization after MI, was significantly expanded after SUMO1 depletion. In addition, cell-to-cell communication analysis showed that CMs regulated EC clusters (mainly EC-2, EC-5, and EC-9) proliferation through VEGFA-mediated paracrine action. Previous studies have shown that EC-specific loss of SUMO specific peptidase 1 (SENP1), a de-SUMOylation enzyme, inhibits pathological angiogenic responses and tissue repair processes [59]. SUMOylation of kinase insert domain receptor (KDR) inhibited VGEF-KDR signaling in ECs and led to angiogenesis dysfunction Here, we found that SUMO1 deletion promoted VEGFA-mediated proliferation of ECs. Excessive SUMOylation is detrimental to the proliferation of the ECs responsible for neovascularization of infarct sites and the construction of vascular networks. A previous study [60] showed that ginkgolic acid (GA) reversed accumulation of s-PML and abolished PML/Pin1 interactions induced by MI, which inhibited TGFβ1-mediated cardiac fibrotic responses. Notably, the inhibitory effect of GA on SUMOylation above ∼130 ka (e.g. s-PML) led to increased SUMO1 expression in the free pool, which resulted in a compensatory increase in SUMOylation levels of the remaining proteins (e.g., s-JunD) and ultimately improved cardiac function after MI. However, deletion of the SUMO1 gene not only led to a decrease in the overall protein SUMOylation level, but also reduced the SUMO1 level in the free pool, which triggered the impairment of the entire SUMO cycle, and the persistent impairment of SUMOylation (e.g., JunD) led to the deterioration of cardiac function after MI. Moreover, the deletion of SUMO1 gene occurred in the early embryonic stage. Although the deletion of SUMO1 gene did not cause changes in cardiac function, but SUMO1 was also involved in cardiac development during embryonic development, which might trigger changes in heart cells at baseline. Therefore, the construction of cell-based specific SUMO1 deletion mice will be necessary in the future.
5. Conclusions
In summary, our analysis determined the role of SUMO1 in the cardiac repair process in response to myocardial injury and described the function of SUMO1 in different types of heart cells, including CMs, FBs, and ECs, after MI. We determined at high resolution that SUMO1 loss triggered transcriptional profiles of distinct cell types and showed that SUMO1 was involved in cardiac remodeling after MI through distinct signaling processes. This dataset also provides an invaluable resource for further studies on the regulatory role of SUMO1 on different heart cell subsets after MI.
CRediT author statement
Zhihao Liu: Project administration, Methodology, Validation, Investigation, Writing - Original draft preparation; Xiaozhi Liu: Project administration, Methodology, Validation, Investigation, Writing - Original draft preparation; Li Liu: Project administration, Methodology, Validation, Investigation, Writing - Original draft preparation; Ying Wang: Investigation, Formal analysis, Software, Validation; Jie Zheng: Investigation, Software and Validation; Lan Li: Data analysis; Sheng Li: Data analysis; Han Zhang: Validation, Formal analysis; Jingyu Ni: Validation, Formal analysis; Chuanrui Ma: Writing - Reviewing & Editing; Xiumei Gao: Writing - Reviewing & Editing; Xiyun Bian: Conceptualization, Supervision, Resources, Project administration; Guanwei Fan: Conceptualization, Supervision, Resources, Project administration.
Declaration of competing interest
The authors declare that there are no conflicts of interest.
Acknowledgments
The authors gratefully acknowledge Xiaohui Fan, PhD (Pharmaceutical Informatics Institute, College of Pharmaceutical Sciences, Zhejiang University, China) for assisting with snRNA-seq experiments. The authors thank Wei Yang, PhD (Duke University Medical Center, USA) for providing SUMO1−/− mice. This study was supported by the Innovation Team and Talents Cultivation Program of National Administration of Traditional Chinese Medicine (Grant No.: ZYYCXTD-D-202207), the National Natural Science Foundation of China (Grant Nos.: 82270304, 81774050, and 81901526), the Tianjin Special Project of New Generation Artificial Intelligence Technology (Project No.: 18ZXZNSY00260), and the Ministry of Education of People's Republic of China “Program for Innovative Research Team in University” (Project No.: IRT_16R54).
Footnotes
Peer review under responsibility of Xi'an Jiaotong University.
Supplementary data to this article can be found online at https://doi.org/10.1016/j.jpha.2022.11.010.
Contributor Information
Xiyun Bian, Email: bblmyx@163.com.
Guanwei Fan, Email: fgw1005@163.com.
Appendix A. Supplementary data
The following is the Supplementary data to this article:
References
- 1.Kaski J.C., Crea F., Gersh B.J., et al. Reappraisal of ischemic heart disease. Circulation. 2018;138:1463–1480. doi: 10.1161/CIRCULATIONAHA.118.031373. [DOI] [PubMed] [Google Scholar]
- 2.Prabhu S.D., Frangogiannis N.G. The biological basis for cardiac repair after myocardial infarction: From inflammation to fibrosis. Circ. Res. 2016;119:91–112. doi: 10.1161/CIRCRESAHA.116.303577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wu X., Reboll M.R., Korf-Klingebiel M., et al. Angiogenesis after acute myocardial infarction. Cardiovasc. Res. 2021;117:1257–1273. doi: 10.1093/cvr/cvaa287. [DOI] [PubMed] [Google Scholar]
- 4.Frangogiannis N.G. The extracellular matrix in myocardial injury, repair, and remodeling. J. Clin. Invest. 2017;127:1600–1612. doi: 10.1172/JCI87491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chang H., Yeh E.T.H. SUMO: From bench to bedside. Physiol. Rev. 2020;100:1599–1619. doi: 10.1152/physrev.00025.2019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Gareau J.R., Lima C.D. The SUMO pathway: Emerging mechanisms that shape specificity, conjugation and recognition. Nat. Rev. Mol. Cell Biol. 2010;11:861–871. doi: 10.1038/nrm3011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Flotho A., Melchior F. Sumoylation: A regulatory protein modification in health and disease. Annu. Rev. Biochem. 2013;82:357–385. doi: 10.1146/annurev-biochem-061909-093311. [DOI] [PubMed] [Google Scholar]
- 8.Zhao X. SUMO-mediated regulation of nuclear functions and signaling processes. Mol. Cell. 2018;71:409–418. doi: 10.1016/j.molcel.2018.07.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Keiten-Schmitz J., Wagner K., Piller T., et al. The nuclear SUMO-targeted ubiquitin quality control network regulates the dynamics of cytoplasmic stress granules. Mol. Cell. 2020;79:54–67.e7. doi: 10.1016/j.molcel.2020.05.017. [DOI] [PubMed] [Google Scholar]
- 10.Litviňuková M., Talavera- López C., Maatz H., et al. Cells of the adult human heart. Nature. 2020;588:466–472. doi: 10.1038/s41586-020-2797-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Wang L., Yu P., Zhou B., et al. Single-cell reconstruction of the adult human heart during heart failure and recovery reveals the cellular landscape underlying cardiac function. Nat. Cell Biol. 2020;22:108–119. doi: 10.1038/s41556-019-0446-7. [DOI] [PubMed] [Google Scholar]
- 12.Mendler L., Braun T., Müller S. The ubiquitin-like SUMO system and heart function: From development to disease. Circ. Res. 2016;118:132–144. doi: 10.1161/CIRCRESAHA.115.307730. [DOI] [PubMed] [Google Scholar]
- 13.Kho C., Lee A., Jeong D., et al. SUMO1-dependent modulation of SERCA2a in heart failure. Nature. 2011;477:601–605. doi: 10.1038/nature10407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Oh J.G., Watanabe S., Lee A., et al. miR-146a suppresses SUMO1 expression and induces cardiac dysfunction in maladaptive hypertrophy. Circ. Res. 2018;123:673–685. doi: 10.1161/CIRCRESAHA.118.312751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.McCrink K.A., Maning J., Vu A., et al. B-Arrestin2 improves post-myocardial infarction heart failure via sarco(endo)plasmic reticulum Ca2+-ATPase-dependent positive inotropy in cardiomyocytes. Hypertension. 2017;70:972–981. doi: 10.1161/HYPERTENSIONAHA.117.09817. [DOI] [PubMed] [Google Scholar]
- 16.Tilemann L., Lee A., Ishikawa K., et al. SUMO-1 gene transfer improves cardiac function in a large-animal model of heart failure. Sci. Transl. Med. 2013;5 doi: 10.1126/scitranslmed.3006487. [DOI] [PubMed] [Google Scholar]
- 17.Gupta M.K., Robbins J. Making the connections: Autophagy and post-translational modifications in cardiomyocytes. Autophagy. 2016;12:2252–2253. doi: 10.1080/15548627.2016.1215384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Evdokimov E., Sharma P., Lockett S.J., et al. Loss of SUMO1 in mice affects RanGAP1 localization and formation of PML nuclear bodies, but is not lethal as it can be compensated by SUMO2 or SUMO3. J. Cell Sci. 2008;121:4106–4113. doi: 10.1242/jcs.038570. [DOI] [PubMed] [Google Scholar]
- 19.Jin K., Gao S., Yang P., et al. Single-cell RNA sequencing reveals the temporal diversity and dynamics of cardiac immunity after myocardial infarction. Small Methods. 2022;6 doi: 10.1002/smtd.202100752. [DOI] [PubMed] [Google Scholar]
- 20.Butler A., Hoffman P., Smibert P., et al. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018;36:411–420. doi: 10.1038/nbt.4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lun A.T.L., Riesenfeld S., Andrews T., et al. EmptyDrops: Distinguishing cells from empty droplets in droplet-based single-cell RNA sequencing data. Genome Biol. 2019;20 doi: 10.1186/s13059-019-1662-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.McGinnis C.S., Murrow L.M., Gartner Z.J. DoubletFinder: Doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 2019;8:329–337.e4. doi: 10.1016/j.cels.2019.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Korsunsky I., Millard N., Fan J., et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods. 2019;16:1289–1296. doi: 10.1038/s41592-019-0619-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Liu B., Li C., Li Z., et al. An entropy-based metric for assessing the purity of single cell populations. Nat. Commun. 2020;11 doi: 10.1038/s41467-020-16904-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Guo X., Zhang Y., Zheng L., et al. Global characterization of T cells in non-small-cell lung cancer by single-cell sequencing. Nat. Med. 2018;24:978–985. doi: 10.1038/s41591-018-0045-3. [DOI] [PubMed] [Google Scholar]
- 26.Trapnell C., Cacchiarelli D., Grimsby J., et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 2014;32:381–386. doi: 10.1038/nbt.2859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Gulati G.S., Sikandar S.S., Wesche D.J., et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science. 2020;367:405–411. doi: 10.1126/science.aax0249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Griss J., Viteri G., Sidiropoulos K., et al. ReactomeGSA-efficient multi-omics comparative pathway analysis. Mol. Cell. Proteomics. 2020;19:2115–2125. doi: 10.1074/mcp.TIR120.002155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yokota T., McCourt J., Ma F., et al. Type V collagen in scar tissue regulates the size of scar after heart injury. Cell. 2020;182:545–562. doi: 10.1016/j.cell.2020.06.030. e23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wang Z., Cui M., Shah A.M., et al. Cell-type-specific gene regulatory networks underlying murine neonatal heart regeneration at single-cell resolution. Cell Rep. 2020;33 doi: 10.1016/j.celrep.2020.108472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Francisco A., Figueira T.R., Castilho R.F. Mitochondrial NAD(P)+ transhydrogenase: from molecular features to physiology and disease. Antioxid. Redox Signal. 2022;36:864–884. doi: 10.1089/ars.2021.0111. [DOI] [PubMed] [Google Scholar]
- 32.Man J., van Duijvenboden K., Krijger P., et al. Genetic dissection of a super enhancer controlling the nppa-nppb cluster in the heart. Circ. Res. 2021;128:115–129. doi: 10.1161/CIRCRESAHA.120.317045. [DOI] [PubMed] [Google Scholar]
- 33.Kuppe C., Ramirez Flores R.O., Li Z., et al. Spatial multi-omic map of human myocardial infarction. Nature. 2022;608:766–767. doi: 10.1038/s41586-022-05060-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hama N., Itoh H., Shirakami G., et al. Rapid ventricular induction of brain natriuretic peptide gene expression in experimental acute myocardial infarction. Circulation. 1995;92:1558–1564. doi: 10.1161/01.cir.92.6.1558. [DOI] [PubMed] [Google Scholar]
- 35.Rubin A.J., Parker K.R., Satpathy A.T., et al. Coupled single-cell CRISPR screening and epigenomic profiling reveals causal gene regulatory networks. Cell. 2019;176:361–376.e17. doi: 10.1016/j.cell.2018.11.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Hilfiker-Kleiner D., Hilfiker A., Castellazzi M., et al. JunD attenuates phenylephrine-mediated cardiomyocyte hypertrophy by negatively regulating AP-1 transcriptional activity. Cardiovasc. Res. 2006;71:108–117. doi: 10.1016/j.cardiores.2006.02.032. [DOI] [PubMed] [Google Scholar]
- 37.Tallquist M.D. Cardiac fibroblast diversity. Annu. Rev. Physiol. 2020;82:63–78. doi: 10.1146/annurev-physiol-021119-034527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Nikoloudaki G., Snider P., Simmons O., et al. Periostin and matrix stiffness combine to regulate myofibroblast differentiation and fibronectin synthesis during palatal healing. Matrix Biol. 2020;94:31–56. doi: 10.1016/j.matbio.2020.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Chen L., Chou C., Knepper M.A. Targeted single-cell RNA-seq identifies minority cell types of kidney distal nephron. J. Am. Soc. Nephrol. 2021;32:886–896. doi: 10.1681/ASN.2020101407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Tang J., Zhang H., He L., et al. Genetic fate mapping defines the vascular potential of endocardial cells in the adult heart. Circ. Res. 2018;122:984–993. doi: 10.1161/CIRCRESAHA.117.312354. [DOI] [PubMed] [Google Scholar]
- 41.Bradshaw A.D., Sage E.H. SPARC, a matricellular protein that functions in cellular differentiation and tissue response to injury. J. Clin. Invest. 2001;107:1049–1054. doi: 10.1172/JCI12939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Depuydt M.A.C., Prange K.H.M., Slenders L., et al. Microanatomy of the human atherosclerotic plaque by single-cell transcriptomics. Circ. Res. 2020;127:1437–1455. doi: 10.1161/CIRCRESAHA.120.316770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Jackson D.G. Hyaluronan in the lymphatics: The key role of the hyaluronan receptor LYVE-1 in leucocyte trafficking. Matrix Biol. 2019;78–79:219–235. doi: 10.1016/j.matbio.2018.02.001. [DOI] [PubMed] [Google Scholar]
- 44.Wang Y., Nakayama M., Pitulescu M.E., et al. Ephrin-B2 controls VEGF-induced angiogenesis and lymphangiogenesis. Nature. 2010;465:483–486. doi: 10.1038/nature09002. [DOI] [PubMed] [Google Scholar]
- 45.Zhang L., Jambusaria A., Hong Z., et al. SOX17 regulates conversion of human fibroblasts into endothelial cells and erythroblasts by dedifferentiation into CD34+ progenitor cells. Circulation. 2017;135:2505–2523. doi: 10.1161/CIRCULATIONAHA.116.025722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Chen D., Sun N., Chen X., et al. Endothelium-derived semaphorin 3G attenuates ischemic retinopathy by coordinating β-catenin-dependent vascular remodeling. J. Clin. Invest. 2021;131 doi: 10.1172/JCI135296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Voigt A.P., Mulfaul K., Mullin N.K., et al. Single-cell transcriptomics of the human retinal pigment epithelium and choroid in health and macular degeneration. Proc. Natl. Acad. Sci. USA. 2019;116:24100–24107. doi: 10.1073/pnas.1914143116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jiang Z., Lu Z., Kou S., et al. Overexpression of Kdr in adult endocardium induces endocardial neovascularization and improves heart function after myocardial infarction. Cell Res. 2021;31:485–487. doi: 10.1038/s41422-020-00436-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Martins-Marques T., Hausenloy D.J., Sluijter J.P.G., et al. Intercellular communication in the heart: Therapeutic opportunities for cardiac ischemia. Trends Mol. Med. 2021;27:248–262. doi: 10.1016/j.molmed.2020.10.002. [DOI] [PubMed] [Google Scholar]
- 50.Napione L., Pavan S., Veglio A., et al. Unraveling the influence of endothelial cell density on VEGF-A signaling. Blood. 2012;119:5599–5607. doi: 10.1182/blood-2011-11-390666. [DOI] [PubMed] [Google Scholar]
- 51.Zeng L., Xiao Q., Chen M., et al. Vascular endothelial cell growth-activated XBP1 splicing in endothelial cells is crucial for angiogenesis. Circulation. 2013;127:1712–1722. doi: 10.1161/CIRCULATIONAHA.112.001337. [DOI] [PubMed] [Google Scholar]
- 52.Lee A., Jeong D., Mitsuyama S., et al. The role of SUMO-1 in cardiac oxidative stress and hypertrophy. Antioxid. Redox Signal. 2014;21:1986–2001. doi: 10.1089/ars.2014.5983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Bian X., Xu J., Zhao H., et al. Zinc-induced SUMOylation of dynamin-related protein 1 protects the heart against ischemia-reperfusion injury. Oxid. Med. Cell. Longev. 2019;2019 doi: 10.1155/2019/1232146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ruiz-Villalba A., Romero J.P., Hernández S.C., et al. Single-cell RNA sequencing analysis reveals a crucial role for CTHRC1 (collagen triple helix repeat containing 1) cardiac fibroblasts after myocardial infarction. Circulation. 2020;142:1831–1847. doi: 10.1161/CIRCULATIONAHA.119.044557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Tombor L.S., John D., Glaser S.F., et al. Single cell sequencing reveals endothelial plasticity with transient mesenchymal activation after myocardial infarction. Nat. Commun. 2021;12:681. doi: 10.1038/s41467-021-20905-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Frangogiannis N.G. Cardiac fibrosis: cell biological mechanisms, molecular pathways and therapeutic opportunities. Mol. Aspect. Med. 2019;65:70–99. doi: 10.1016/j.mam.2018.07.001. [DOI] [PubMed] [Google Scholar]
- 57.Kang J.S., Saunier E.F., Akhurst R.J., et al. The type I TGF-beta receptor is covalently modified and regulated by sumoylation. Nat. Cell Biol. 2008;10:654–664. doi: 10.1038/ncb1728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Katsuno Y., Derynck R. Epithelial plasticity, epithelial-mesenchymal transition, and the TGF-β family. Dev. Cell. 2021;56:726–746. doi: 10.1016/j.devcel.2021.02.028. [DOI] [PubMed] [Google Scholar]
- 59.Zhou H.J., Xu Z., Wang Z., et al. SUMOylation of VEGFR2 regulates its intracellular trafficking and pathological angiogenesis. Nat. Commun. 2018;9 doi: 10.1038/s41467-018-05812-2. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 60.Qiu F., Dong C., Liu Y., et al. Pharmacological inhibition of SUMO-1 with ginkgolic acid alleviates cardiac fibrosis induced by myocardial infarction in mice. Toxicol. Appl. Pharmacol. 2018;345:1–9. doi: 10.1016/j.taap.2018.03.006. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.