Abstract
For genetic research to contribute more fully to furthering our knowledge of neuropathic pain we require an agreed, valid and feasible approach to phenotyping, in order to allow collaboration and replication in samples of sufficient size. Results from genetic studies on neuropathic pain have been inconsistent and have met with replication difficulties, in part because of differences in phenotypes used for case ascertainment. Since there is no consensus on the nature of these phenotypes, nor the methods of collecting them, this study aimed to provide guidelines on collecting and reporting phenotypes in cases and controls for genetic studies. Consensus was achieved through a staged approach: (1) systematic literature review to identify all neuropathic pain phenotypes used in previous genetic studies; (2) Delphi survey to identify the most useful neuropathic pain phenotypes, their validity and feasibility; and (3) meeting of experts to reach consensus on the optimal phenotype(s) to be collected from neuropathic pain patients for genetic studies. A basic ‘entry-level’ set of phenotypes was identified for any genetic study of neuropathic pain. This set identifies cases of ‘possible’ neuropathic pain, and controls, and includes: (1) a validated symptom-based questionnaire to determine whether any pain is likely to be neuropathic; (2) body chart or checklist to identify whether the area of pain distribution is neuroanatomically logical; and (3) details of pain history (intensity, duration, any formal diagnosis). This NeuroPPIC ‘entry-level’ set of phenotypes can be expanded by more extensive and specific measures, as determined by scientific requirements and resource availability.
1. Introduction
Genetic analysis of neuropathic pain will illuminate the biological processes underlying the condition and facilitate identification of novel therapeutic and prevention targets. However the clinical utility of findings from current research is questionable, with limited success in identifying the genes contributing to the heritable risk, and difficulties in replicating the documented genes [28,33]. Future genome-wide association and candidate gene studies are likely identify the genetic contribution to neuropathic pain heritability, as has happened with various complex CNS disorders, if sufficiently-powered sample sizes are used [16,42]. While measures to ensure the quality and validity of genetic analysis are well-established [1,10,39], there is little agreement on which neuropathic pain traits should be studied and which phenotyping tools to use. The quality and validity of the phenotype are as important as those of the genetics methodology [50], and perhaps even more important than sample size [30]. A poorly-defined or quantified phenotype will produce errors in effect size estimation, loss of power to identify risk genes, or identification of significant associations with phenotypes with limited clinical relevance [25,34].
An optimally collected phenotype should fully describe the neuropathic pain entity under study every time and capture the same genetic variants in replicated studies. It should be collected with validated instruments (i.e. accurate, precise, reproducible, with known positive and negative predictive values) that are practical (i.e. simple to implement, efficient, cost-effective) and ethical. No single phenotyping instrument will encompass all these requirements. Rather, a number of instruments are required, ranging from ‘highly specific’ tools to ‘very general’. These instruments should be mutually consistent and able to diagnose participants by classifying them as ‘possibly’, ‘probably’ or ‘definitely’ having neuropathic pain [48]. Classification of controls is as important as classification of cases, and both require stringency and clarity [30]. Tailoring the right combination of instruments for any particular study depends on a balance between validity and feasibility.
There is currently no ‘gold standard’ for assessing neuropathic pain, for clinical or research purposes. There is, however, growing consensus, with generally minor variations between approaches [18,45]. Consensus on neuropathic pain phenotyping in genetic research will facilitate: (1) scientific collaboration, increasing the potential to combine cohorts to achieve larger sample sizes; (2) replication of gene discoveries; (3) meta-analyses; and (4) translation from laboratory to general population, and vice-versa [44]. Furthermore, until very large samples are created purely for researching neuropathic pain, population-based research will rely on data from multi-purpose clinical samples. Whilst the lack of a ‘gold standard’ currently precludes calculation of precise positive and negative predictive values, the existence of an agreed standard for neuropathic pain, will facilitate a valid approach to this research.
Led and funded by the International Association for the Study of Pain (IASP) Special Interest Group (SIG) on Neuropathic Pain (NeuPSIG), and in collaboration with the IASP SIG on Genetics and Pain, we staged an approach towards achieving consensus on neuropathic pain phenotyping for human genetic studies. This is intended to inform future genetic research, rather than phenotyping for clinical research or clinical diagnosis.
2. Methods
Our step-wise approach towards this goal had three stages:
A systematic literature review to identify all neuropathic pain phenotypes used in previous genetic studies;
A Delphi survey to determine and rank neuropathic pain phenotypes assessing their validity and feasibility; and
A meeting of experts to reach consensus on ‘ideal’ phenotype(s) to be collected from neuropathic pain patients for genetic studies.
2.1. Systematic review
2.1.1. Aim
To conduct a systematic literature review to identify and compare phenotypes used in genetic studies of non-cancer neuropathic pain in adults for the purposes of informing the phenotypes that may best capture the genetic architecture of neuropathic pain for human genetics studies.
2.1.2. Study selection and data extraction
Electronic databases MEDLINE, EMBASE, SCOPUS, Science Direct, ISI Web of Science and CINAHL were searched from January 1966 to April 2014 for English-language papers. We used the same search strategy as a recent systematic review to identify neuropathic pain [19] and combined this with broad search terms for ‘genetic studies’. These search terms are listed in Supplementary Digital Content 1.
We excluded studies where: (1) it was not possible to distinguish between participants with neuropathic pain and those with non-neuropathic pain (including mixed pain conditions); (2) the pain was cancer-related; (3) the condition was not currently defined by the IASP as neuropathic pain (complex regional pain syndrome (CRPS), temporo-mandibular disorders (TMD), migraine, chronic widespread pain (CWP, and fibromyalgia) [26]; or (4) the study was in children.
Database searches were conducted by one author (OVH). Article titles and then abstracts were reviewed for possible inclusion by two authors (OVH and BHS), before full-text versions of the remaining articles were reviewed and the final selection confirmed. We extracted data on: (1) study characteristics (country, sample population, study design); (2) phenotyping methods described; (3) sample size (cases and controls); (4) the specific neuropathic pain condition(s) under study; and (5) genetic factors investigated.
We differentiated between ‘brief’ and ‘detailed’ phenotype descriptions to give an indication of the level of detail provided by the authors and whether or not it allowed replication by other researchers. For example, ‘Clinical examination by pain specialist’ [14] was regarded as ‘brief’, while ‘Clinical examination: straight leg raising test, manual testing of motor and sensory defects of the lower extremities concordant with MRI findings,’ [36] was regarded as ‘detailed’. This does not necessarily indicate that only a brief examination was conducted in the former case, but indicates the extent to which the phenotyping was described.
In addition to extracting descriptive information, the phenotype information was assessed against the neuropathic pain grading guidelines published by NeuPSIG [48] and approved by IASP [26]. This ranks the diagnostic certainty with which the presence or absence of neuropathic pain can be based on the neuroanatomical distribution of symptoms, patient history and tests confirming the underlying nervous system lesion or disease. The grading system has the following criteria: (1) pain with a neuro-anatomically plausible distribution; (2) a history of a relevant lesion or disease affecting the somatosensory system; (3) confirmatory tests demonstrating presence of negative and positive sensory signs confined to innervation territory of the lesioned nervous structure; and (4) further diagnostic tests confirming a causative lesion or disease entity. Criteria 1 and 2 must be met to allow a working hypothesis of ‘possible’ neuropathic pain. Additionally, criterion 3 or criterion 4 must also be met to reach the grade of ‘probable’ neuropathic pain. If all four criteria are satisfied, the grade of ‘definite’ neuropathic pain is achieved.
2.2. Delphi survey
2.2.1. Aim
To obtain expert consensus on phenotype components that should be used to determine ‘caseness’ in genetic studies of neuropathic pain, and to grade the validity and feasibility of applying these phenotype components in research setting.
2.2.2. Ethics approval
The study had ethics approval from the University of Dundee Research Ethics Committee (UREC 14032).
2.2.3. Participants
Email invitations to take part in a three-round Delphi survey were sent to 28 experts in the field of neuropathic pain phenotyping and/or conducting genetic studies on neuropathic pain. All experts were identified by their publication track record in at least one of the fields. The invitation provided information on the context and objectives of the web-based survey (composed using the SurveyMonkey software application at https://www.surveymonkey.com), and a hyperlink for interested individuals to access the survey). Participation was voluntary, and anonymity was assured.
At the end of the first round of the survey respondents indicated whether they wanted to take part in subsequent rounds. Respondents that elected to continue participating were sent email invitations that included a summary of the results from the previous round, and a hyperlink to a new web-based questionnaire. Round 3 was only completed after the face-to-face consensus meeting (see: 2.3. Consensus meeting)
2.2.4. Questionnaires
In Round 1 expert panelists used a five-point Likert scale (1 = strong disagreement, 3 = no agreement or disagreement, 5 = strong agreement), to rate the level of their agreement with statements regarding: (1) the sensitivity and specificity of symptoms, clinical signs, and additional investigations (e.g. quantitative sensory testing (QST), nerve conduction studies) when diagnosing neuropathic pain; (2) the feasibility of non-expert clinicians and researchers to accurately assess items in the three measurement domains (symptoms, clinical signs, additional investigations); (3) whether symptoms and clinical signs could be self-assessed by study participants; and (4) whether the assessment of medical history, body charts of perceived pain, quality of life, and psychological factors should also be phenotyped in population-based genetic studies. The expert panelists were also asked to list up to four symptoms, signs, and additional investigations that they thought provided the best balance between feasibility and validity when assessing whether a pain was predominantly neuropathic in nature. Finally, respondents rated the level of diagnostic certainty (none, possible, probable, definite) they thought was achieved by nine different assessment combinations. The nine assessment combinations were: symptoms only; symptoms, body chart of perceived pain and pain history; clinical signs only; clinical signs and symptoms; clinical signs, symptoms, body chart of perceived pain and pain history; additional investigations only; additional investigations and clinical signs; additional investigations, clinical signs, and symptoms; and additional investigations, clinical signs, symptoms, and body chart of perceived pain and pain history. These assessments were derived from the neuropathic pain grading system developed by Treede and colleagues [48]. The order of the nine combinations was randomized for each panelist.
In Round 2, following standard Delphi methodology, to allow re-evaluation of responses in light of those of their peers, [32,40], panelists were shown summary results from Round 1. They were asked again for their responses to questions on measurement sensitivity, specificity, feasibility, whether participants could self-complete a diagnostic questionnaire, and on the resultant diagnostic certainty. Based on responses to Round 1, panelists were also provided with a list of 14 verbal descriptors of their pain symptoms (hot/burning, stabbing, itching, numbness, electric shocks/shooting, pricking/tingling/pins-and-needles, pain in an area of numbness, pain in a plausible anatomical distribution, pain evoked by light touch, pain in an area of altered sensation, spontaneous pain, evoked pain, painful cold, and paroxysmal pain), 12 clinical signs (dynamic mechanical allodynia, deep mechanical allodynia, altered sensation to punctate mechanical stimuli, static mechanical hyperalgesia, hypoesthesia to punctate mechanical stimuli, altered reflexes, punctate mechanical hyperalgesia, thermal hyperalgesia, cold allodynia, altered vibration sense, thermal hypoesthesia, and temporal summation), and 5 additional investigations (QST, cerebral evoked potentials, intra-epidermal nerve fiber density, magnetic resonance imaging, and nerve conduction studies). From each of these lists, panelists were asked to rank in descending order of importance the five items they thought provided the best balance between validity and feasibility when making a diagnosis of neuropathic pain. The order of items in each list was randomized for each panelist.
Finally, based on feedback from the first round, panelists were asked to identify up to four additional phenotype components that could be added to the assessment of study participants once ‘caseness’ had been established, which would allow more complex phenotypes to be collected.
In Round 3, respondents were shown summary results from Round 2, and asked to re-rate their ranking of the five most valid and feasible symptoms, clinical signs, and additional investigations to use when making a diagnosis of neuropathic pain. In addition, based on discussions at the face-to-face consensus meeting (see section: 2.3 Consensus meeting), which took place between Round 2 and 3 of the Delphi survey, respondents were asked to rate their agreement on a five-point Likert scale (1 = strong disagreement, 5 = strong agreement) with statements regarding patient history and using a pain body chart. Because these additional questions were only asked once, responses were analyzed separately from the rest of the Delphi survey.
2.2.5 Data analysis
Consensus in Delphi surveys is said to have been achieved when a given proportion of participants agree on an item under debate; this proportion varies between studies. For this study, achievement of ‘good’ consensus was assumed when ≥70% of respondents agreed, and ‘strong’ consensus was assumed when there was ≥90% agreement [45].
2.3. Consensus meeting
2.3.1. Aim
To develop a consensus statement on an approach to phenotyping in genetic studies of neuropathic pain in adults, and to identify a basic or ‘entry level’ phenotype for any such study.
2.3.2. Procedure
The meeting was held in Versailles, France, from 12 to 13 June 2014, and consisted of 18 experts, identified by NeuPSIG based on their experience in the fields of pain phenotyping, epidemiology and/or pain genetics. Represented disciplines included: neurology, anaesthesiology, pain medicine, palliative care, primary care, basic neuroscience, and genetics. Activities on the first day included: (1) describing the aims of the meeting, and defining the questions that were to be addressed; (2) presentation of data from the systematic literature review and the results of the first two rounds of the Delphi survey; and (3) short presentations by panelists on the differences and commonalities between ‘phenotyping’ (what information to collect and how to collect it) to collect versus ‘phenomics’ (which pain phenotypes should be used in genetic association analysis), phenotyping of complex diseases, phenotyping by questionnaires, phenotyping by clinical examination, phenotyping using standard and dynamic QST, and phenotyping for clinical trials. Each presentation by a panelist was followed by discussion. Activities on the second day included parallel breakaway discussions followed by plenary-based consensus. These focused on: (1) the availability and use of validated diagnostic neuropathic pain screening tools; (2) the level of diagnostic certainty achieved when defining cases and controls; and (3) generating a consensus definition and the requirements of what should be ‘entry level’ phenotyping requirements for genetic studies on neuropathic pain.
All participants in the consensus meeting contributed as authors of this paper.
3. Results
3.1. Systematic literature review
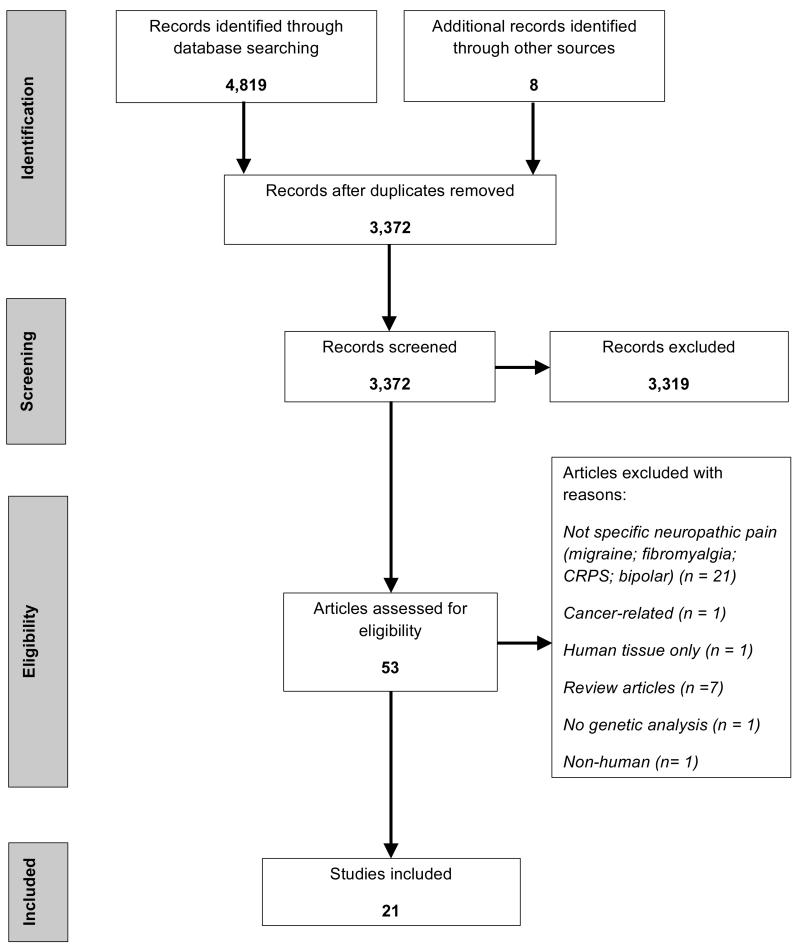
From an initial 4,827 article titles identified through searching electronic databases and hand-searching, 3,372 were identified as unique records. Of these, 21 papers fulfilled the inclusion criteria, and underwent data extraction (Fig. 1).
Figure 1.
Systematic review: PRISMA flow diagram of article identification, assessment, and inclusion.
3.1.2. Characteristics of included studies
Six studies analysed Nordic populations [6,14,22,23,27,36], five analysed other European populations[2,4,15,20,41], three analysed Japanese [37,43,46], two analysed Israeli Jewish [12,35] and South African black, Caucasian and Indian [21,49], one analysed Chinese (Taiwan) [8] and one analysed Caucasian-, African- and Hispanic-American populations [47]. One study included five cohorts with various neuropathic pain conditions from four populations (Danish, Finnish, Israeli-Jewish, and Caucasian-, African- and Hispanic-American) [11]. Two studies on postherpetic neuralgia (PHN) recruited participants from the same Japanese population [37,46]. Several studies reused the same cohorts to investigate different polymorphisms, namely two studies on a South African HIV-positive cohort comprising African black, Caucasian and Indian [21,49], two studies on a Norwegian cohort with discogenic sciatic pain [22,23], two studies on Caucasian-, African- and Hispanic-American patients with persistent pain after surgery for discogenic sciatic pain [11,47], and three studies on a Finnish cohort with discogenic pain [11,27,36].
A variety of causes of neuropathic pain were studied (Table 1). Four study cohorts grouped together a number of causes of neuropathic pain [2,4,11,14], whereas others focused on a solitary cause of neuropathic pain [6,8,12,15,20–23,27,35–37,41,43,46,47,49]. Overall, the majority of associations reported between genotype and risk of neuropathic pain/intensity of neuropathic pain have been isolated findings that have not yet been replicated, or failed to replicate as conflicting findings have been reported. Where there have been consistent reports of genetic associations at specific loci, some of the replication studies included PHN patients recruited from the same study site [37,46] or from the same study cohort [27,36] (i.e. patients having discogenic sciatic pain) (Table 2). Only three studies included a priori replication cohorts into their study designs [11,14,47].
Table 1.
Systematic review: description of cases and controls in included studies
| Study | Sample Size |
Cases | Controls | |
|---|---|---|---|---|
| Cases | Controls | |||
| Armero et al., (2005) [2] | 144 | 139 | Sympathetic reflex dystrophy, complex regional pain syndrome II, failed low-back surgery syndrome, phantom limb pain, peripheral nerve neuralgia, cranial nerve neuralgia, post-herpetic neuralgia | Age-matched healthy subjects without a history of pain |
| Binder et al., (2011) [4] | 371 | 252 | Complex regional pain syndrome, post-herpetic neuralgia, peripheral nerve injury, trigeminal neuropathy, polyneuropathy, central pain, other neuropathies | Healthy German volunteers (all Caucasian) |
| Brasch-Andersen et al., (2011) [6] | 11 | 23 | Polyneuropathy - patients with moderate or better pain relief during escitalopram treatment were categorized as ‘responders’ (cases). | Polyneuropathy - patients with less than moderate pain relief during escitalopram treatment were categorized as ‘non-responders’ (controls) |
| Cheng et al., (2010) [8] | 15 | 50 | Painful diabetic peripheral neuropathy (with foot ulcer) | Painless diabetic peripheral neuropathy (with foot ulcer) |
| Costigan et al., (2010) [11] | 1174 (total from 5 cohorts)a | n/a | Chronic lumbar root pain after surgical discectomy (n = 151), limb amputation (n = 199), sciatica (n = 195), phantom limb pain (n = 100), post-mastectomy pain syndrome (n = 529) | No control group was used (authors modelled the relationship between genotype and pain intensity within the cohorts) |
| Dabby et al., (2011) [12] | 9 | 50 | Chronic severe unexplained neuropathic pain. | Not described. |
| Dominguez et al., (2013) [14] | 325 (total from 2 cohorts) | 731 (total from 3 cohorts) | Persistent postsurgical pain (PPSP) after inguinal hernia repair (n = 94), radicular neuropathy from lumbar disc herniation (n = 231). | No PPSP (n = 95), Swedish national population register for a multiple sclerosis incidence study (n = 213), and the Diabetes Incidence Study in Sweden (n = 423; matched to cases by age, sex, and geographic region). No control group was used for the radicular neuropathy group (authors modelled the relationship between genotype and pain intensity/functional recovery within the cohort). |
| Fernández de-las-Peñas et al., (2013) [15] | 58 | 108 (total from 2 cohorts) | Pain in multiple sclerosis (36 with neurogenic pain). | Pain-free multiple sclerosis (n = 50), and age- and sex-matched, pain-free healthy subjects from the general population with specific exclusion criteria (n = 108). |
| Hegarty and Shorten (2012) [20] | 20 | 33 | Persistent postsurgical pain (PPSP) after lumbar discectomy. | No PPSP (n=33) |
| Hendry et al., (2013) [21] | 158 | n/a | HIV-associated sensory neuropathy (HIV-SN) | No control group was used (authors modelled the relationship between genotype and pain intensity within the cohort). |
| Jacobsen et al., (2012) [22] | 258 | 249 | Lumbar disc herniation and sciatic pain. | Pain-free controls collected from the general health survey Nord-Trøndelag Health Study (HUNT). Controls had no history of back disease, and were matched for age, sex, and smoking status. |
| Jacobsen et al., (2013) [23] | 257 | 253 | Lumbar disc herniation and sciatic pain. | Pain-free controls collected from the general health survey Nord-Trøndelag Health Study (HUNT). Controls had no history of back disease, and were matched for age, sex, and smoking status. |
| Karppinen et al., (2008) [27] | 14 | 139 | GGGA interleukin(IL)-6 haplotype with unilateral discogenic sciatic pain. | Non-GGGA IL-6 haplotype with unilateral discogenic sciatic pain. |
| Nissenbaum et al., (2010) [35] | 21 | 334 | PPSP after unilateral breast mastectomy or lumpectomy. | No PPSP after unilateral breast mastectomy or lumpectomy. |
| Noponen-Hietala et al., (2005) [36] | 155 | 179 | Unilateral discogenic sciatic pain. | Unrelated University of Oulu, Finland, employees and students (ages of 20–69 years), all of whom were Finnish. No data were collected concerning possible musculoskeletal disorders. |
| Ozawa et al., (1999) [37] | 32 | 136 | Post-herpetic neuralgia (PHN) | Unrelated healthy Japanese volunteers. |
| Ramirez et al., (2012) [41] | 6 | 0 | Charcot-Marie-Tooth disease (hereditary painful neuropathy) | No controls reported. |
| Sato et al., (2002) [43] | 40 | 125 | PHN | Healthy Japanese volunteers |
| Sumiyama et al., (2008) [46] | 70 | 220 (total from 2 cohorts) | PHN | Healthy Japanese volunteers with no history of herpes zoster (n =140), herpes zoster patients without PHN (n = 80). |
| Tegeder et al.; (2006) [47] | 168 | 0 | PPSP after lumbar discectomy (the study was replicated in healthy controls with experimentally induced heat, ischaemic and mechanical pain, n = 547). | No controls reported (authors separately modelled the relationship between genotype and pain intensity within the lumbar pain cohort and the experimental pain cohort). |
| Wadley et al., (2012) [49] | 144 | 15 | HIV-SN | Non-painful HIV-SN Authors also modelled relationship between genotype and pain intensity within the 144 with pain. |
Data from a sixth cohort describing the pain sensitivity of healthy volunteers was excluded.
Table 2.
Systematic review: summary of genetic associations identified with neuropathic pain conditions
| Gene investigated | Loci with at least one significant association |
Cause of neuropathic pain [Reference] |
|---|---|---|
| Multiple concordant studies reporting associations at specific loci | ||
| HLA-A | *33 | PHN [37,43,46]a |
| HLA-B | *44 | PHN [37,43,46]a |
| IL6 GGGA haplotype | rs1800796 rs1800795 rs13306435 rs1800797 |
Discogenic sciatic pain [27,36]b |
| Multiple discordant studies for specific loci | ||
| COMT | rs4680 | Significantly associated with:
Not associated significantly with: |
| GCH1 | rs3783641 rs10483639 rs8007267 |
Significantly associated with:
Not associated significantly with: |
| KCNS1 | rs734784 | Significantly associated with:
Not associated significantly with: |
| HLA-DQB1 | *03:02 | Significantly associated with:
Not associated significantly with:
|
| HLA-DRB1 | *04 | Significantly associated with:
Not associated significantly with: |
| HLA-DRB1 | *13 | Significantly associated with:
Not associated significantly with: |
| HLA-A-B-DRB1 haplotype | *33-*44-*1302 | Significantly associated with:
Not associated significantly with:
|
| OPRM1 | rs1799971 | Significantly associated with:
Not associated significantly with:
|
| Solitary studies reporting associations at novel loci | ||
| CACNG2 | rs4820242 rs2284015 rs2284017 rs2284018 rs1883988 |
PPSP (mastectomy/lumpectomy) [35] |
| HLA-A-B | *33-*44 | PHN [37] |
| HLA-DQB1-DRB1 haplotype | *03:02-*04 | PPSP (inguinal hernia) [14] |
| HLA-C | w3 | PHN [37] |
| HTR2C | rs6318 | Polyneuropathy [6] |
| IL6 d | rs1800796 rs1800795 rs13306435 |
Discogenic sciatic pain [36] |
| KCNS1 haplotype | rs4499491 rs6017486 rs6073643 |
HIV-SN [21] |
| MMP1 | rs1799750 | Discogenic sciatic pain [23] |
| TRPV1 | rs920829 rs8065080 rs222747 |
Multiple aetiologies [4] |
| Solitary studies reporting no associations across multiple locif | ||
| ABCB1 | Polyneuropathy [6] | |
| CYP2C19 | Polyneuropathy [6] | |
| CYP2D6RS | PPSP (lumbar discectomy) [20] | |
| HTR2A | Polyneuropathy [6] | |
| IL1A | Discogenic sciatic pain [36] | |
| IL1B | Discogenic sciatic pain [36] | |
| SLC6A4 | Polyneuropathy [6] | |
| TNFA | Discogenic sciatic pain [36] | |
| TNFA | PHN [43] | |
Note: Data from studies describing a single patient [12] or hereditary neuropathic pain conditions [41] have not been included in the Table;
Participants in Sumiyana et al., 2008 [46] and Ozawa et al., 1999 [37] were recruited from the same study site;
Participants in Karppinen et al., 2008 [27] and Noponen-Hietala et al., 2005 [36] were from the same cohort;
Treatment response to escitalopram;
Association with pain intensity, not risk of neuropathic pain;
Loci not listed for clarity;
DPN: Diabetic polyneuropathy;
HIV-SN: HIV-associated sensory neuropathy;
PHN: Post-herpetic neuralgia;
PPSP: Persistent post-surgical pain
3.1.3. Phenotyping methods
The identified studies could be classified into three broad categories, according to their reason for phenotyping neuropathic pain, to: (1) determine whether any pain was neuropathic (identification of cases and controls) [2,4,8,12,14,15,20,22,23,35–37,43,46,49]; (2) identify endo-phenotypes within cohorts with neuropathic pain (i.e. phenomics) [6,11,21,27,41,47,49]; and (3) identify pain (not specifically neuropathic) in a neurological condition, e.g. multiple sclerosis [15].
Table 3 and Supplementary Digital Content 2 provide summaries of the phenotyping methods described in these articles. Overall, ‘clinical examination’ was the most frequently reported phenotyping method; described in 15/21 studies, with ‘brief examination’ reported in 6 studies, including in total 1,742 cases, and ‘detailed examination’ reported in 9 studies (1,346 cases). ‘Pain-rating scales’ (either visual analogue or numerical rating scales, 2,786 cases) and ‘history’ (2,552 cases) were the next most common phenotyping method described, with 13/21 studies each. With ‘Pain-rating scales’ only half of these articles described what the relevance of the pain score was: for example, to be used as a cut-off for ‘caseness’ to categorize patients as ‘cases’ or ‘controls’. Nine of the articles reporting a ‘history’ provided a ‘brief history’ (2,226 cases), and 4 of the articles provided a ‘detailed history’ (326 cases). All other phenotyping methods were described in fewer than half of the articles, with the least frequently reported phenotyping methods being ‘nerve conduction studies’ (2/21), ‘intra-epidermal nerve fibre density’ (2/21), ‘inflammatory markers’ (2/21), ‘body chart of perceived pain’ (1/20), and related ‘psychiatric measures’ (1/21). In two studies there was phenotyping heterogeneity, manifesting in the use of more than one sample population cohort within the same study [11,14].
Table 3.
Systematic review: phenotyping methods reported in the article text
| Phenotyping method used | Number of studies |
References | Total summed cases/controlsd |
|---|---|---|---|
| Clinical examinationc | |||
| • Total | 15 | ||
| • Brief | 6 | [6,11,14,15,40,46]a,b | Brief: 1,742/862 |
| • Detailed | 9 | [8,14,20–23,27,35,48]b | Detailed: 1,346/1,649 |
| Pain rating scale | 13 | [8,11,14,15,20–23,27,35,42,46,48]a,b | 2,786/1,932 |
| Historyc | |||
| • Total | 13 | ||
| • Brief | 9 | [2,11,14,15,34,37,42,46,47]a,b | Brief: 2,226/1,387 |
| • Detailed | 4 | [8,12,21,48] | Detailed: 326/15 |
| Radiological imaging (MRI) | 9 | [11,12,14,15,20,22,23,27,35]a,b | 2,270/1,742 |
| Neuropathic pain identification questionnaire | 2 | [21,48] | 302/15 |
| Quantitative sensory testing (QST) | 5 | [4,6,12,20,40] | 417/358 |
| Not reported | 4 | [1,5,18,22]a | 175/301 |
| Nerve conduction studies (NCS) | 2 | [12,40] | 15/50 |
| Intra-epidermal nerve fibre density (IENFD) | 2 | [12,40] | 15/50 |
| Inflammatory markers | 2 | [12,20] | 29/53 |
| Body chart of perceived pain | 1 | [15] | 58/108 |
| Psychological measures (HADS) | 1 | [20] | 20/53 |
Costigan et al., 2010 [11]: Six independent cohorts each with different phenotyping methods;
Dominguez et al., 2013 [14]: Two independent cohorts each with different phenotyping methods;
‘brief’ and ‘detailed’ describe the level of detail provided in the text, not necessarily that the assessment was ‘brief’ or ‘detailed’;
cases/controls were not summed across studies if they were taken from the same existing cohort;
MRI: Magnetic resonance imaging;
HADS: Hospital Anxiety and Depression Scale
The only specific neuropathic pain symptom questionnaire reported was the AIDS Clinical Trials Group Brief Peripheral Neuropathy Screen [9,21,49]. No studies reported the use of validated case ascertainment tools such as the Douleur Neuropathique en 4 questions (DN4) [5], Leeds Assessment of Neuropathic Symptoms and Signs (LANSS) [3], or painDETECT [17].
The control groups described in the studies could be divided into three broad categories: (1) healthy volunteers [2,4,15,36,37,43,46]; (2) population-specific national reference cohorts [14,22,23]; and (3) diseased controls [6,8,14,15,20,27,35,49]. Three studies used more than one category of control group [14,15,46]. In the first two categories (healthy controls and population-specific national reference cohorts), little or no information was provided on the phenotyping of controls. Typically the information provided in these studies was restricted to whether the controls were healthy, genetically unrelated, and matched the cases for various demographic variables. Diseased controls were defined as having the same disease/aetiology as the cases, but were not considered to have neuropathic pain at the time of the study.
Four studies modelled genotype against intensity of neuropathic pain and thus did not include control groups [11,21,47,49]. Similarly, two case reports did not include control groups at all [12,41].
3.1.4. Grading framework: ‘possible’, ‘probable’ or ‘definite’ neuropathic pain
Only 1 of the 20 studies cited the NeuPSIG neuropathic pain grading system, and that single study indicated that cases satisfied the criteria for ‘definite’ neuropathic pain [41]. Table 4 summarises the grading criteria met by each study based on the description of the phenotyping methodology in the article text. Only one study did not describe assessing a history [6]. Although only one study reported using a body chart of perceived pain [15] to assess whether pain occurred in a plausible anatomical distribution, the assessment of pain distribution could be inferred from the phenotyping descriptions provided in 13 studies, even if the method of assessment was not explicitly described [8,11,12,14,20–23,27,35,36,49]. Fourteen studies described assessing clinical signs [6,8,12,14,15,20–23,27,36,41,49], and 8 studies described the use of methods that could identify a nerve lesion (either radiological imaging and/or QST) [4,6,12,20,22,23,27,36]. In summary, 4 studies included patients identified as ‘possible’ neuropathic pain cases [11,35,46,47], 5 included patients identified as ‘probable’ neuropathic pain cases [8,14,15,21,49], and only 7 included patients identified as ‘definite’ neuropathic pain cases [12,20,22,23,27,36,41]. Six studies could not be graded according to the criteria proposed by Treede and colleagues [48], and were classified as ‘undefined’.
Table 4.
Systematic review: neuropathic pain grading criteria achieved
| Study | Criterion 1: | Criterion 2: | Criterion 3: | Criterion 4: | Grading |
|---|---|---|---|---|---|
|
| |||||
| Plausible neuro- anatomical distribution |
Appropriate history |
Altered sensory signs |
Evidence of lesion or disease |
||
| Armero et al., (2005) [2] | ● | Undefined *** | |||
| Binder et al., (2011) [4] | ● | ● | Undefined *** | ||
| Brasch-Andersen et al., (2011) [6] | ● | ● | Undefined *** | ||
| Cheng et al., (2010) [8] | ● | ● | ● | Probable | |
| Costigan et al., (2010) [11] * | ● | ● | Possible | ||
| Dabby et al., (2011) [12] | ● | ● | ● | ● | Definite |
| Dominguez et al., (2013) [14]** | ● | ● | ● | Probable | |
| Fernández de-las-Peñas et al., (2013) [15] | ● | ● | ● | Probable | |
| Hegarty and Shorten (2012) [20] | ● | ● | ● | ● | Definite |
| Hendry et al., (2013) [21] | ● | ● | ● | Probable | |
| Jacobsen et al., (2012) [22] | ● | ● | ● | ● | Definite |
| Jacobsen et al., (2013) [23] | ● | ● | ● | ● | Definite |
| Karppinen et al., (2008) [27] | ● | ● | ● | ● | Definite |
| Nissenbaum et al., (2010) [35] | ● | ● | Possible | ||
| Noponen-Hietala et al., (2005) [36] | ● | ● | ● | ● | Definite |
| Ozawa et al., (1999) [37] | ● | Undefined *** | |||
| Ramirez et al., (2012) [41] | ● | ● | ● | ● | Definite |
| Sato et al., (2002) [43] | ● | Undefined *** | |||
| Sumiyama et al., (2008) [46] | ● | Undefined *** | |||
| Tegeder et al., (2006) [47] | ● | ● | Possible | ||
| Wadley et al., (2012) [49] | ● | ● | ● | Probable | |
Costigan et al. (2010): Six independent cohorts each containing different information about phenotyping method, number of cases; The cohort with the lesser probability of neuropathic pain is reflected in Table 4.
Dominguez et al. (2013): Two independent cohorts each with different phenotyping method and cases. The cohort with the lesser probability of neuropathic pain is reflected in Table 4.
Grading is undefined in terms of the criteria proposed by Treede et al [48]: Criteria 1 and 2 must be met to allow a working hypothesis of ‘possible’ neuropathic pain. Additionally, criterion 3 or criterion 4 must also be met to reach the grade of ‘probable’ neuropathic pain. If all four criteria are satisfied, the grade of ‘definite’ neuropathic pain is achieved.
3.2. Delphi survey
3.2.1. Participation
Twenty experts, of the 28 approached to take part in the survey, completed Round 1 (responder rate: 20/28, 71%), and 17/20 (85%) indicated that they were willing to participate in subsequent survey rounds. Round 2 invitations were sent to 17 experts, 16 of whom completed the questionnaire (responder rate: 16/17, 94%). Invitations to Round 3 of the survey were sent to the same Round 2 17 experts, 15 of whom completed the questionnaire (responder rate: 15/17, 88%).
3.2.2. Validity and feasibility of assessing symptoms, signs, and completing additional investigations
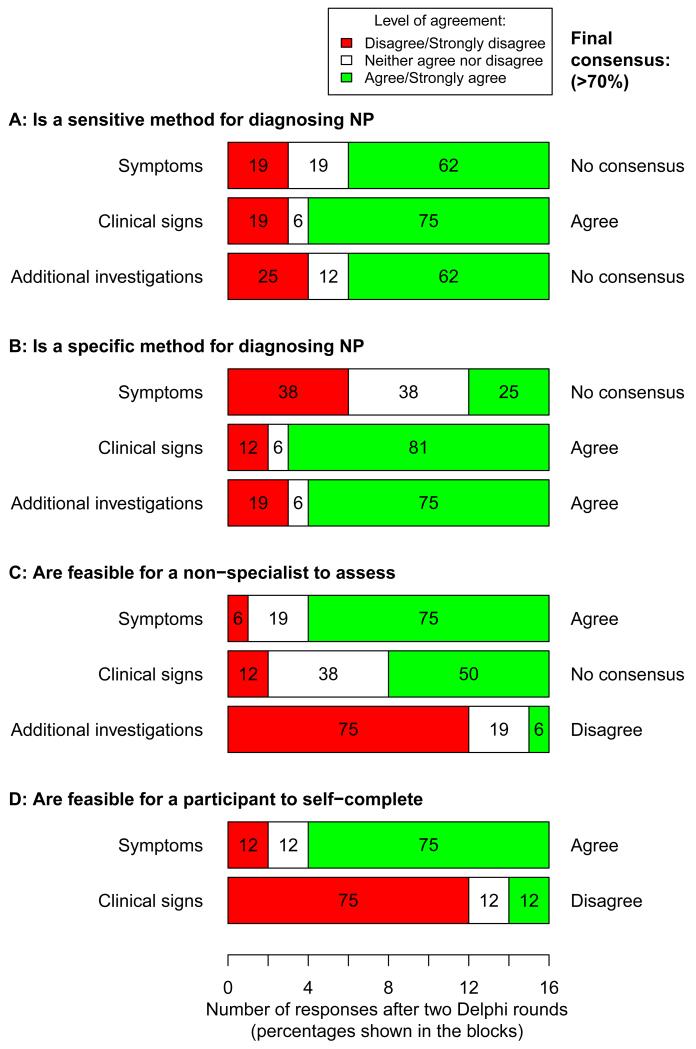
Figure 2 shows the level of agreement of respondents after two survey rounds with regards to whether symptoms, clinical signs, and additional investigations are sensitive and specific assessments of neuropathic pain, and whether they are feasible for non-experts and study participants to complete by themselves. There was good consensus that testing clinical signs was a sensitive method of detecting neuropathic pain as means of identifying true ‘cases’, but poor consensus was achieved for either symptoms or additional investigations alone. There was good consensus that clinical signs and additional investigations (but not symptoms) were specific methods to enable the identification of true ‘controls’. There was good consensus that symptoms could be reliably assessed by non-experts and by study participants, but that additional investigations were not feasible for non-experts or study participants to assess, and that study participants could not reliably self-assess clinical signs. The levels of agreement across each round of the survey are shown in Supplementary Digital Content 3.
Figure 2.
Delphi survey: level of agreement (5-point Likert scale, anchored at 1 = strongly disagree and 5 = strongly agree), and whether consensus was achieved (when ≥ 70% of respondents disagreed/strongly disagreed, or vice versa, agreed/strongly agreed with a statement) after two rounds of the survey (n = 16 in the second round). Panel A: agreement on whether symptoms, clinical signs, and additional investigations are sensitive methods of detecting neuropathic pain. Panel B: agreement on whether the three measurement domains are specific methods for detecting neuropathic pain. Panel C: agreement on whether it is feasible for a non-specialist to assess each of the three measurement domains in a research setting. Panel D: agreement on whether it is feasible for study participants to self-assess symptoms and clinical signs.
3.2.3. Informative symptoms, signs, and additional investigations
Table 5 shows the final consensus on respondents’ choices and rankings of symptoms used in the diagnosis of neuropathic pain for a genetic study. Strong or good consensus was achieved for only two symptoms: ‘hot/burning’, and ‘pain evoked by light touch’. Fewer than one third of respondents included ‘spontaneous pain’, ‘paroxysmal pain’, ‘evoked pain’, ‘painful cold’, ‘itching’ or ‘stabbing pain’ in their top five symptoms for diagnosing neuropathic pain.
Table 5.
Delphi survey: number of participants that ranked symptoms in their top five symptoms for diagnosing neuropathic pain based on the balance between validity and feasibility of measurement
| Symptomsa, b |
Number of times a symptom was listed in the top 5 N (% participants) |
Median rank (inter- quartile range, IQR) in Round 3 |
Good consensus on symptom inclusion in the top 5 (YES: ≥70%) |
|
|---|---|---|---|---|
|
Round 2 (n = 16) |
Round 3: (n = 15) |
|||
| Hot/Burning | 11 (69) | 14 (93) | 2 (1.25 - 4.5) | YES |
| Pain evoked by light touch | 10 (62) | 12 (80) | 3 (2 - 4) | YES |
| Pain in a plausible anatomical distribution | 7 (44) | 10 (67) | 2 (1.25 - 3.75) | -- |
| Pain in an area of numbness | 6 (38) | 8 (53) | 2 (2 - 3.25) | -- |
| Pricking/Tingling/Pins and needles | 11 (69) | 8 (53) | 3 (2.5 - 3.25) | -- |
| Electric shocks/Shooting | 9 (56) | 8 (53) | 4 (3 - 4.25) | -- |
| Pain in an area of altered sensation | 5 (31) | 7 (47) | 2 (1.5 - 3) | -- |
| Numbness | 6 (38) | 5 (33) | 4 (4 - 5) | -- |
| Spontaneous pain | 0 (0) | 2 (13) | 1 (1 - 1) | -- |
| Paroxysmal pain | 3 (19) | 2 (13) | 4.5 (4.25 - 4.25) | -- |
| Evoked pain | 1 (6) | 1 (7) | 3 (3 - 3) | -- |
| Painful cold | 2 (12) | 1 (7) | 3 (3 - 3) | -- |
| Itching | 4 (25) | 1 (7) | 4 (4 - 4) | -- |
| Stabbing pain | 3 (19) | 1 (7) | 4 (4 - 4) | -- |
Symptoms or symptom groups identified by 20 participants in Round 1 in an open question to list four symptoms that provide the best balance between validity and feasibility when diagnosing neuropathic pain;
Symptoms are listed according to number of times they were listed in participants’ top 5 in Round 3, and then by the median rank received by listed symptoms in Round 3 (1 highest ranked, 5 lowest rank);
-- No consensus reached at the ≥70% threshold level after two rounds
Respondents’ choices and rankings of clinical signs used in the diagnosis of neuropathic pain are shown in Table 6. Strong or good consensus was achieved for only two signs: ‘dynamic mechanical allodynia’, and ‘altered sensation to punctate mechanical stimuli’. Fewer than one-third of respondents included ‘temporal summation’, ‘altered vibration sense’, ‘thermal hyperalgesia’, ‘altered reflexes’, ‘static mechanical hyperalgesia’, or ‘deep mechanical hyperalgesia’ in their top five signs to assess.
Table 6.
Delphi survey: number of participants that ranked a clinical sign in their top five signs for diagnosing neuropathic pain based on the balance between validity and feasibility of measurement
| Clinical signa,b |
Number of times a clinical sign was listed in the top 5 (% participants) |
Median rank (IQR) in Round 3 |
Good consensus on clinical sign inclusion in the top 5 (YES: ≥70%) |
|
|---|---|---|---|---|
|
Round 2 (n = 16) |
Round 3: (n = 15) |
|||
| Dynamic mechanical allodynia | 13 (81) | 14 (93) | 1 (1 - 2) | YES |
| Altered sensation to punctate mechanical stimuli | 6 (38) | 12 (80) | 2 (1 - 4.25) | YES |
| Hypoaesthesia to punctate mechanical stimuli | 8 (50) | 10 (67) | 3 (2 - 4) | -- |
| Punctate mechanical hyperalgesia | 7 (44) | 9 (60) | 2 (2 - 4) | -- |
| Cold allodynia | 7 (44) | 9 (60) | 3 (2 - 4) | -- |
| Thermal hypoaesthesia | 5 (31) | 6 (40) | 2.5 (2 - 4.5) | -- |
| Temporal summation | 6 (38) | 4 (27) | 2 (1 - 3.25) | -- |
| Altered vibration sense | 5 (31) | 4 (27) | 3 (2 - 4.25) | -- |
| Thermal hyperalgesia | 2 (12) | 3 (20) | 3 (2.5 - 3.5) | -- |
| Altered reflexes | 2 (12) | 3 (20) | 3 (2.5 - 4) | -- |
| Static mechanical hyperalgesia | 5 (31) | 1 (7) | 3 (3 - 3) | -- |
| Deep mechanical hyperalgesia | 1 (6) | 1 (7) | 4 (4 - 4) | -- |
Clinical signs identified by 20 participants in Round 1 in an open question to list four clinical signs that provide the best balance between validity and feasibility when diagnosing neuropathic pain;
Signs are listed according to number of times they were listed in participants’ top 5 in Round 3, and then by the median rank received by listed signs in Round 3 (1 highest ranked, 5 lowest rank);
-- No consensus reached at the ≥70% threshold level after two rounds
Table 7 shows the final consensus of respondents’ choices and rankings of their top-five additional investigations for diagnosing neuropathic pain. The two top-ranked investigations were, ‘quantitative sensory testing (QST)’, and ‘intra-epidermal nerve fibre density’. In Round 3, when the feasibility of non-specialists making accurate measurements was assessed for each of the five additional investigations, only QST was rated as being feasible by 11/14 (79%) of respondents. More than 70% of respondents agreed that all other methods of assessment were not feasible for non-specialists to assess neuropathic pain.
Table 7.
Delphi survey: number of participants that ranked an additional investigation in their top five symptoms for diagnosing neuropathic pain based on the balance between validity and feasibility of measurement
| Additional investigationsa,b |
Number of times an investigation was listed (% participants) |
Median rank (IQR) in Round 3 |
|
|---|---|---|---|
| Round 2 (n = 16) | Round 3 (n = 15) | ||
| Quantitative sensory testing | 12 (75) | 14 (93) | 1 (1 - 1.75) |
| Intra-epidermal nerve fibre density | 12 (75) | 13 (87) | 2 (2 - 3) |
| Nerve conduction studies | 8 (50) | 13 (87) | 3 (2 - 3) |
| Magnetic resonance imaging | 6 (38) | 12 (80) | 5 (4 - 5) |
| Evoked potentials | 7 (44) | 11 (73) | 4 (3.5 - 4) |
Additional investigations identified by 20 participants in Round 1 in an open question to list four additional investigations that provide the best balance between validity and feasibility when diagnosing neuropathic pain. NOTE: only 5 additional investigations were provided for participants to rank in Round 2 and 3, so the number of times an investigation was listed was not used to assess consensus;
Additional investigations are listed according to number of times they were listed in participants’ top 5 in Round 3, and then by the median rank received by listed items in Round 3 (1 highest ranked, 5 lowest rank)
3.2.4. Perceived diagnostic certainty
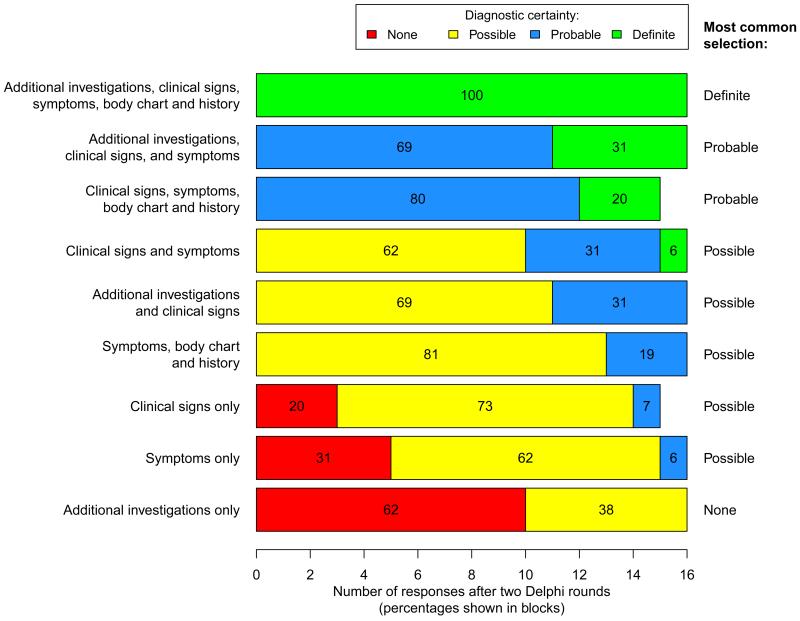
Figure 3 shows the level of perceived diagnostic certainty, according to the neuropathic pain grading system [48], indicated by respondents for various assessment modalities used alone or in combination with each other [2]. There was unanimous consensus that a combination of ‘additional investigations, clinical signs, symptoms, body chart and patient history’ provided a diagnosis of ‘definite’ neuropathic pain. All respondents also indicated that combinations of: ‘additional investigations, clinical signs and symptoms’, and ‘clinical signs, symptoms, body chart and patient history’ provided a diagnosis of at least ‘probable’ neuropathic pain. The poorest perceived diagnostic certainty was associated with assessments that only included ‘additional investigations’. The levels of agreement across each round of the survey are shown in Supplementary Digital Content 4.
Figure 3.
Delphi survey: level of diagnostic certainty achieved by individual assessment types and combinations of these assessments when assessing whether a pain is ‘definite’, ‘probable’ or ‘possible’ neuropathic pain [2]. Results shown are after two rounds of the Delphi survey (n =16 in the second round).
3.2.5. Additional phenotyping measures
The level of agreement on which measurements should be included in an assessment of neuropathic pain (in addition to symptoms, signs, and additional investigations) is shown in Table 5 (and Supplementary Digital Content 5). After two rounds of the survey, there was strong consensus that assessment should include a history and a body chart to indicate pain distribution. Prompted by discussions at the consensus meeting (see below), in Round 3 of the survey we probed what the body chart and history should include. Having ≥ 70% of respondents in agreed or strong agreement, the strongest consensus was for the following items assessing patient history: ‘a previous diagnosis of neuropathic pain by another clinician’, and ‘risk factors for having neuropathic pain’ (see Supplementary Digital Content 6). On the body chart of perceived pain, 75% (12/16) of respondents indicated that, in the event of more than one pain site, the ‘main pain and any other pains’ should be identified. Almost two-thirds (64%, 9/14) thought that a checklist of body regions was a suitable substitute for a body chart cartoon.
3.3. Consensus meeting
The primary goal of the meeting was the development of a consensus statement on ‘entry-level’ phenotyping requirements for genetic studies of neuropathic pain in human adults’.
3.3.1. Consensus statement development
Participants agreed that there is an inverse relationship between accurate case ascertainment of neuropathic pain and large-scale implementation thereof in population-based studies. We agreed that the entry-level phenotyping requirements included in the statement should form the basis of establishing neuropathic pain ‘caseness’ for genetic studies. This basic phenotyping must allow the addition of more in-depth measures for higher level phenotyping, to allow the identification of genetic loci for specific aspects of neuropathic pain other than just presence or absence of neuropathic pain. The entry-level requirements should provide a framework to guide researchers on study design, and provide a platform to appraise their findings to allow feasible and valid assessment for the presence or absence of neuropathic pain in large, population-based genetic studies.
3.3.2. ‘Entry level’ phenotyping for genetic studies, a consensus
Box 1 shows the agreed NeuroPPIC entry-level requirements that would allow the classification of participants as having ‘possible’ neuropathic pain. Consistent with the findings of the Delphi survey, a neuropathic pain ‘case’ must have:
-
(1)
pain with neuropathic characteristics (i.e. positive on a validated screening questionnaire; or pain that is ‘hot/burning’, and/or ‘pain evoked by light touch’);
-
(2)
a distribution or location of this pain that is anatomically consistent with an underlying somatosensory lesion or disease (as indicated by a body chart or body parts checklist); and
-
(3)
further characterisation in the form of a clinical history (duration and intensity of pain, presence of other pains) and demographic information relevant to the population/disease being studied.
Box 1. NeuroPPIC consensus statement on entry-level phenotyping for genetic studies of neuropathic pain in humans.
Entry-level phenotyping
| |
|---|---|
| 1. Symptom assessment using neuropathic pain screening tools |
|
| 2. Anatomical distribution of pain |
|
| 3. Historyd |
|
If there is a discrepancy between the outcome of the screening tool, pain history and body chart, it might be appropriate to exclude the participant from both case and control groups or to stratify the genetic analysis with a weighted importance on each of the entry level criteria;
Participants in the meeting were most familiar with the DN4-interview: Douleur Neuropathique en 4 questions-interview [5], S-LANSS: Self-report Leeds assessment of neuropathic symptoms and signs [3], and painDETECT [17];
Round 3 of the Delphi survey identified (≥ 70% of respondents) that ‘main pain and any other pains’ should be recorded, with the site of the main pain being noted (64% of respondents agreed that a checklist of body parts could be used instead of an body chart cartoon);
Round 3 of the Delphi survey identified (≥ 70% of respondents) that ‘a previous diagnosis of neuropathic pain by another clinician’, and ‘risk factors for having neuropathic pain’ should be included in the history;
Pain intensity should not be used as a pre-determined cut-off value for case ascertainment.
A neuropathic pain ‘control’ would be negative for (1) and (2), but will be characterised as (3) as clinically relevant.
The panel agreed that if there are discrepancies between the findings from the screening tool, anatomical distribution, and patient history, then the participant should be excluded from the study. Overall, these three components only satisfy criteria 1 and 2 of the NeuPSIG grading system, and thus the level of diagnostic certainty achieved is ‘possible’ neuropathic pain [48].
Whilst acknowledging the lack of a ‘gold standard, we agreed that achieving greater diagnostic certainty (i.e. ‘probable’ or ‘definite’ neuropathic pain) is desirable. Such greater diagnostic certainty requires additional assessments that confirm the presence of positive and/or negative sensory signs (for ‘probable’ neuropathic pain) or provide direct evidence of a lesion to the somatosensory nervous system (for ‘definite’ neuropathic pain) in the affected area. More extensive phenotyping could allow case stratification into additional pain phenotypes based on sensory and psychological parameters (e.g., pain catastrophizers versus non-catastrophizers; pin-prick hypoaesthesia versus pin-prick hyperaesthesia).
The panel additionally agreed that it could not currently be prescriptive in the selection of additional phenotyping measures to use beyond determination of 'caseness’, as the choice was dependent on factors such as the feasibility of making measurements based on cost and practicality, the population/disease being studied (e.g., central versus peripheral neuropathic pain), and the research question being addressed. However, choice of additional measures to use when assessing ‘caseness’, should be consistent with the NeuPSIG grading system for neuropathic pain. See Supplementary Digital Content 7 for examples of additional phenotypes voiced at the meeting.
3.3.3. Misclassification
We agreed that misclassification of cases and controls significantly affects the power of studies to detect true genetic associations and the reproducibility of findings. We identified several key factors that may contribute to misclassification of cases and controls (Box 2). These factors can be divided into two broad categories: (1) misclassification due to on incomplete pain history; and (2) misclassification due to an imperfect sensitivity and specificity of available assessments for neuropathic pain. Ideally, where reasonable doubt arises after applying the phenotyping criteria proposed here, an individual should be considered as neither a case nor a control, and removed from analysis.
Box 2. Possible factors that may contribute to misclassification of neuropathic pain.
| History |
|---|
| Past episodes of neuropathic pain, but neuropathic pain not currently active |
| Changing neuropathic pain phenotype over time |
| Multiple chronic pain conditions |
| Measurement |
| The specificity of questionnaires (i.e. ability to correctly identify controls) |
| Heterogeneity of endo-phenotypesa within neuropathic pain syndromes |
| Similarities in endo-phenotypesa across neuropathic pain syndromes |
| Inconsistent mode of measurement (e.g., face-to-face, postal, web-based) |
| Cross-cultural and language validity of screening tools and questionnaires |
| Assessment of the ‘main pain’ when there are multiple pain diagnoses or sitesb |
Sub-phenotypes within the broader classification of having neuropathic pain (e.g., neuropathic pain associated with hypoaesthesia to pin-prick versus neuropathic pain associated with hyperaesthesia to pin-prick);
Study participants with multiple pain sites of mixed origin (neuropathic and non-neuropathic) may be misclassified as controls if only their ‘main pain’ is characterised and that pain is non-neuropathic in origin. Nevertheless, the panel agreed that asking participants to characterise their ‘main pain’ was the most practical approach to assessing patients with multiple pain sites (especially if using a self-assessment approach), but were possible, a more complete assessment of such participants should be made. The issue is less of problem when investigating pain of known aetiology (e.g., post-herpetic neuralgia, post-amputation pain), and patients can be directed to focus on the pain produced (or not) by this aetiology.
4. Discussion
The unequivocal identification of genetic sequence variants correlating with complex diseases requires sample sizes ranging from large cohorts (including hundreds to thousands of participants) when interrogating a few candidate genes, to much larger sample sizes that include tens of thousands of participants when studying the whole genome, [31,51] and extensive phenotyping [50]. Unlike Mendelian disorders, the genetic contribution to complex diseases such as chronic pain is generally attributed to contributions of a high number of risk variants, including single nucleotide polymorphisms (SNPs), most with a small effect. In neuropathic pain, a number of relevant candidate genes have been identified in animal studies, and an even smaller number show some association in humans, as shown above. However, none of these SNPs has been consistently replicated in large human cohorts [33]. This is an essential step in confirming the validity of these genetic findings [7]. Failure to replicate, in humans, a finding originally made in humans may be because the original finding was spurious, study cohorts were underpowered, there was heterogeneity in the studied pain condition or study populations, or there were differences in the phenotyping of cases and/or controls. The ideal control in a genetic study would be an age- and sex-matched individual who was exposed to the same lesion or disease of the somatosensory system as the case, but who did not develop neuropathic pain [30]. Rigorous, standardised phenotyping is therefore crucial. Failure to translate a finding from rodent models to humans may likewise be because the original finding was spurious, the human cohort was underpowered, the phenotypes were different, or the gene under study does not play a role in the tested human condition. Pain geneticists who seek candidate neuropathic pain genes in animal models should try to approximate their phenotypes to those used by human pain geneticists, although animal phenotypes also require considerable further refinement in terms of both internal and external validity [38].
The need for a consensus-based, rather than purely evidence-based approach to phenotyping neuropathic pain for genetic studies reflects the absence of ‘gold standards’ for both case definition (what is neuropathic pain?) and case attribution (how do we know who has neuropathic pain?). Through a process of systematic literature review, Delphi survey and expert consensus meeting, we have identified an agreed ‘entry level’ approach to phenotyping cases and controls for studies of genetic associations with neuropathic pain in adults. This phenotype is based on establishing the presence or absence of ‘possible’ neuropathic pain, according to the most internationally agreed system for classifying neuropathic pain clinically [48]. This uses validated symptom-based screening tools and a body chart or checklist to determine neuroanatomical relevance. Further details, also collected by questionnaire, include pain history (duration, intensity, formal diagnoses) and demographic information (Box 1). We present this ‘entry level’ approach as the minimum phenotyping standard for genetic association studies on neuropathic pain in adults, with the expectation that more extensive phenotyping should build on this ‘entry level’ approach, as required and as resources allow. Adoption of these NeuroPPIC phenotyping guidelines will facilitate comparison and collaboration between researchers, including replication studies and meta-analyses. The guidelines are applicable to different types of genetic studies of neuropathic pain, from targeted studies interrogating a few genetic loci that use specific neuropathic pain traits, to hypothesis-free genome-wide studies with large sample sizes. They will also apply to studies with a larger array of phenotypes, including more detailed pain traits, psychosocial traits and pharmacological screens. Our systematic review identified the lack of a consistent approach to phenotyping neuropathic pain, poor reporting of phenotyping methodologies, and failure to adequately describe control groups (especially when ‘healthy controls’ were used). These limitations are likely to have contributed to inconsistent and non-replicated findings, and highlight the need for the NeuroPPIC process we have completed.
Although we have recommended the use of validated assessment instruments where possible, our guidelines are not intended to guide case definition, attribution or classification for clinical research or neurological practice, where more individual certainty may be required, based on empirical evidence of the presence or absence of neuropathic pain and the known likelihood of identifying this through any assessment. This will require separate work, and is the subject of an on-going NeuPSIG project. Instead, our consensus-based development provides a standardised approach for use by researchers in different settings internationally, in the expectation that it will allow a common approach to identifying the presence or absence of ‘possible’ neuropathic pain for genetic research. However, future research should validate the ‘entry level’ phenotype against best available clinical assessments, with a view to calculating sensitivity, specificity and positive and negative predictive values.
Delphi surveys are an accepted method of achieving consensus on complex issues in science and medicine [32,40], including pain medicine, where a ‘gold standard’ does not exist (as in neuropathic pain [26]). For example previous studies have used Delphi methods to develop a standard definition for back pain [13] and neuropathic pain [45] for use in prevalence studies, and to develop guidelines for the management of hip and knee osteoarthritis [52]. Identified limitations of the method include its basis in the opinions of a selected group of survey participants, and the potential for the outcome to be a diluted version of ‘best opinion’ [40], criticisms that could also be levelled at consensus meetings. There is no agreed optimum number or composition of panellists or rounds for Delphi surveys or consensus meetings. Our intention was to include a range of disciplines rather than to be comprehensive in our inclusion, and to include a similar number of participants and rounds as in previous successful studies [13,45,52]. This would allow the processes to remain manageable, while ensuring a contribution of expertise from each relevant discipline and consistency with existing research. Contributors were identified on the basis of their published track record in neuropathic pain phenotyping and/or genetics. It is possible that a different group of contributors would have produced different outcomes, and unanimity was not obtained for any item in the survey. It is, though, notable that good or very good consensus was achieved in many useful aspects of this Delphi survey, and agreement on the NeuroPPIC ‘entry level’ phenotyping approach was achieved after intense deliberations. We are, therefore, confident that our findings reflect the views of the relevant research community, resulting in adoption of NeuroPPIC’s approach in future genetic studies.
NeuroPPIC has only provided a consensual statement on the nature of ‘entry level’ phenotyping and instruments that can be used to accomplish successful classification of participants as having versus not having neuropathic pain. These NeuroPPIC guidelines are for the basic phenotyping requirements for any genetic study of neuropathic pain in adults. It is notable that this ‘entry level’ phenotyping is based only on study participants’ responses to questions and does not include clinical examination or further investigations. This ‘entry level’ phenotyping approach will allow inclusion in large-scale questionnaire-based surveys. Our Delphi survey confirms that augmenting a simple symptom checklist by questions on pain bodily distribution and history will allow the identification of ‘possible’ neuropathic pain. Although symptoms could be confined to a smaller list of verbal descriptors (e.g. ‘hot/burning pain’ and ‘pain evoked by light touch’), the use of a slightly longer but well-validated screening instrument will prevent the need for validation of a new questionnaire instrument. Available screening instruments include DN4, S-LANSS and PainDETECT, but we are making no statement on which of these, if any, is to be preferred. This approach conforms to the ‘very general’ approach outlined in the introduction, and will be suitable for large-scale population-based studies, including those in which neuropathic pain is just one of a number of conditions to be phenotyped. More extensive phenotyping of additional aspects of neuropathic pain will require a more detailed set of assessments, to allow for inclusion of ‘very specific’ phenotypes. While NeuroPPIC was able to provide some pointers towards these levels, we have not yet furnished agreed guidelines on the nature of these phenotypes and the instruments to collect them, and this will require further work, building on the ‘entry level’ approach. Any additional, more extensive phenotyping should, however, be consistent with the ‘entry level’ approach, to allow specificity to be built in to the basic model. This will allow incremental approaches to phenotyping within a large cohort, and the testing for replication of associations identified in extensively phenotyped cohorts.
4.1. Reporting recommendations
The STREGA [29] and GRIPS [24,25] guidelines provide a framework for reporting genetic epidemiology studies, and indicate that participant eligibility criteria, and methods of participant selection and screening, including subsets of participants, should be reported. Moreover, in case-control studies, this information should be reported for both cases and controls. To facilitate clear and open reporting of phenotyping in genetic studies of neuropathic pain, we propose the following NeuroPPIC guidelines for describing study participants: (1) a description of sampling methodology and participant eligibility criteria; (2) an unambiguous description of the case definition used to define the presence or absence of neuropathic pain in cases and controls; (3) a description of the methods used to assess each component of the case definition, and the criteria used to define abnormal function for each method used; and (4) a summative statement of whether the case definition satisfies the criteria for ‘possible’, ‘probable’ or ‘definite’ neuropathic pain based on the latest grading guidelines (currently Treede and colleagues [48]). These recommendations are consistent with the recommendations made by STREGA and GRIPS, and should be used in addition to those two guidelines.
By providing an ‘entry-level’ phenotype and making recommendations on the reporting of the criteria and methods used for caseness and its ascertainment we believe that greater consistency and transparency can be achieved in studies on the genetics of neuropathic pain in adult humans. These improvements will facilitate advancements in the field by enabling collaboration between research groups, replication of discoveries of contributing genetic variants, meta-analyses, and translation from the laboratory to the general population, and back again.
Supplementary Material
Acknowledgements
We thank the 20 responders to the Delphi survey for their input, and Dr Harriet Wordsworth for taking comprehensive minutes of the expert panel meeting. This work was entirely funded by the Neuropathic Pain Special Interest Group (NeuPSIG) of the International Association for the Study of Pain. DLHB is a Wellcome Senior Clinical Scientist (Ref 095698z/11/z). NA, DB, DLHB, RB, NF, TSJ, ASCR, BHS, and DY are members of the DOLORisk consortium funded by European Commission Horizon 2020 (ID633491).
Conflicts of interest
PRK, NA, MH, SNR, ASCR and BHS were members of the NeuPSIG Management Committee at the time of the research. OvH, DLHB, LD, RFreeman, MH, ZS and DY declared no other conflicts of interest. NA declared consultancy or lecture fees from Pfizer, Astra Zeneca, Lilly, Grunenthal, Johnson and Johnson Sanofi Pasteur Mćrieux outside the scope of this work. PRK declared consultancy fees from Reckitt Benckiser, lecture fees from Pfizer and Novartis, and travel support from Janssen. BHS declared occasional consultancy and lecture fees, on behalf of his institution, from Pfizer, Napp and Grunenthal. He is the Scottish Government Lead Clinician for Chronic Pain. RB has received grants / research support from Pfizer, Genzyme, Grünenthal, Mundipharma. He is a member of the IMI “Europain” collaboration and industry members of this are: Astra Zeneca, Pfizer, Esteve, UCB-Pharma, Sanofi Aventis, Grünenthal, Eli Lilly and Boehringer Ingelheim. German Federal Ministry of Education and Research (BMBF): German Research Network on Neuropathic Pain, NoPain system biology. German Research Foundation (DFG). He has received speaking fees from Pfizer, Genzyme, Grünenthal, Mundipharma, Sanofi Pasteur, Medtronic, Eisai, Lilly, Boehringer Ingelheim, Astellas, Desitin, Teva Pharma, Bayer-Schering, MSD. He has been a consultant for Pfizer, Genzyme, Grünenthal, Mundipharma, Allergan, Sanofi Pasteur, Medtronic, Eisai, Lilly, Boehringer Ingelheim, Astellas, Novartis, Bristol-Myers-Squibb, Biogenidec, AstraZeneca, Merck, Abbvie, Daiichi Sankyo and Glenmark Pharmaceuticals. GB and TET are employed by deCODE Genetics/Amgen Inc, and have received funding from the European Commission (HEALTH-FP7-2013-Innovation-602891 to the NeuroPain Consortium) and the NIH (R01DE022905). DLHB declared occasional consultancy and lecture fees, on behalf of his institution, from Pfizer. MIB declared consultancy and lecture fees from Pfizer, Bayer and Grunenthal. RFreynhagen declared research support, consulting, or lecture fees in the past 2 years from Astellas, Grünenthal, Janssen, Lilly and Pfizer. TSJ declared consultancy fees for Pfizer, Grünenthal, Orion and Astellas and lecture fees from Pfizer. SNR declared a research grant from Medtronic and that he has served as an Advisor for Mitsubishi Tanabe Pharma Inc. ASCR declared Imperial College Consultants Consultancy work and Scientific Board Memberships contracted by Imperial College Consultants. In the last 3 years these have included remunerated work for the following companies: Spinifex, Merck, Astellas, Medivir, Asahi Kasei Pharma, Servier, Relmada, Abide, Neusentis (Pfizer), Aquilas (Neuromax) and Mitsubishi; and owner ship of share options in Spinifex. He also declared receipt of research grant support in the last 3 years, on behalf of his institution, from Pfizer (as part of London Pain Consortium and Neuropain) and Astellas (as part of EU Innovative Medicines Initiative grant- EUROPAIN). He is a member of the UK Department of Health (non-financial) Joint Committee on Vaccination and Immunisation, Varicella/Herpes Zoster subgroup. MH declared consultancy fees for AbbVie, Allergan, Astellas, Eli Lilly, Janssen Cilag, Pfizer and Sanofi-Aventis, and lecture fees from Astellas, Eli Lilly, Janssen Cilag, MSD, Mundipharma, Orion and Pfizer.
References
- [1].Anderson C a, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT. Data quality control in genetic case-control association studies. Nat. Protoc. 2010;5:1564–1573. doi: 10.1038/nprot.2010.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [2].Armero P, Muriel C, Santos J, Sànchez-Montero FJ, Rodríguez RE, González-Sarmiento R. COMT (Val158Met) polymorphism is not associated to neuropathic pain in a Spanish population. Eur. J. Pain. 2005;9:229–232. doi: 10.1016/j.ejpain.2004.06.005. [DOI] [PubMed] [Google Scholar]
- [3].Bennett M. The LANSS Pain Scale: The Leeds assessment of neuropathic symptoms and signs. Pain. 2001;92:147–157. doi: 10.1016/s0304-3959(00)00482-6. [DOI] [PubMed] [Google Scholar]
- [4].Binder A, May D, Baron R, Maier C, Tölle TR, Treede RD, Berthele A, Faltraco F, Flor H, Gierthmühlen J, Haenisch S, Huge V, Magerl W, Maihöfner C, Richter H, Rolke R, Scherens A, Üçeyler N, Ufer M, Wasner G, Zhu J, Cascorbi I. Transient receptor potential channel polymorphisms are associated with the somatosensory function in neuropathic pain patients. PLoS One. 2011;6 doi: 10.1371/journal.pone.0017387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Bouhassira D, Attal N, Alchaar H, Boureau F, Brochet B, Bruxelle J, Cunin G, Fermanian J, Ginies P, Grun-Overdyking A, Jafari-Schluep H, Lantćri-Minet M, Laurent B, Mick G, Serrie A, Valade D, Vicaut E. Comparison of pain syndromes associated with nervous or somatic lesions and development of a new neuropathic pain diagnostic questionnaire (DN4) Pain. 2005;114:29–36. doi: 10.1016/j.pain.2004.12.010. [DOI] [PubMed] [Google Scholar]
- [6].Brasch-Andersen C, Møller MU, Christiansen L, Thinggaard M, Otto M, Brøsen K, Sindrup SH. A candidate gene study of serotonergic pathway genes and pain relief during treatment with escitalopram in patients with neuropathic pain shows significant association to serotonin receptor2C (HTR2C) Eur. J. Clin. Pharmacol. 2011;67:1131–1137. doi: 10.1007/s00228-011-1056-x. [DOI] [PubMed] [Google Scholar]
- [7].Chanock SJ, Manolio T, Boehnke M, Boerwinkle E, Hunter DJ, Thomas G, Hirschhorn JN, Abecasis G, Altshuler D, Bailey-Wilson JE, Brooks LD, Cardon LR, Daly M, Donnelly P, Fraumeni JF, Freimer NB, Gerhard DS, Gunter C, Guttmacher AE, Guyer MS, Harris EL, Hoh J, Hoover R, Kong CA, Merikangas KR, Morton CC, Palmer LJ, Phimister EG, Rice JP, Roberts J, Rotimi C, Tucker MA, Vogan KJ, Wacholder S, Wijsman EM, Winn DM, Collins FS. Replicating genotype-phenotype associations. Nature. 2007;447:655–60. doi: 10.1038/447655a. [DOI] [PubMed] [Google Scholar]
- [8].Cheng KI, Lin SR, Chang LL, Wang JY, Lai CS. Association of the functional A118G polymorphism of OPRM1 in diabetic patients with foot ulcer pain. J. Diabetes Complications. 2010;24:102–108. doi: 10.1016/j.jdiacomp.2009.02.003. [DOI] [PubMed] [Google Scholar]
- [9].Cherry CL, Wesselingh SL, Lal L, McArthur JC. Evaluation of a clinical screening tool for HIV-associated sensory neuropathies. Neurology. 2005;65:1778–1781. doi: 10.1212/01.wnl.0000187119.33075.41. [DOI] [PubMed] [Google Scholar]
- [10].Clarke GM, Anderson C a, Pettersson FH, Cardon LR, Morris AP, Zondervan KT. Basic statistical analysis in genetic case-control studies. Nat. Protoc. 2011;6:121–133. doi: 10.1038/nprot.2010.182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [11].Costigan M, Belfer I, Griffin RS, Dai F, Barrett LB, Coppola G, Wu T, Kiselycznyk C, Poddar M, Lu Y, Diatchenko L, Smith S, Cobos EJ, Zaykin D, Allchorne A, Shen PH, Nikolajsen L, Karppinen J, Männikkö M, Kelempisioti A, Goldman D, Maixner W, Geschwind DH, Max MB, Seltzer Z, Woolf CJ. Multiple chronic pain states are associated with a common amino acid-changing allele in KCNS1. Brain. 2010;133:2519–2527. doi: 10.1093/brain/awq195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [12].Dabby R, Sadeh M, Gilad R, Lampl Y, Cohen S, Inbar S, Leshinsky-Silver E. Chronic non-paroxysmal neuropathic pain - Novel phenotype of mutation in the sodium channel SCN9A gene. J. Neurol. Sci. 2011;301:90–92. doi: 10.1016/j.jns.2010.10.006. [DOI] [PubMed] [Google Scholar]
- [13].Dionne CE, Dunn KM, Croft PR, Nachemson AL, Buchbinder R, Walker BF, Wyatt M, Cassidy JD, Rossignol M, Leboeuf-Yde C, Hartvigsen J, Leino-Arjas P, Latza U, Reis S, Gil Del Real MT, Kovacs FM, Oberg B, Cedraschi C, Bouter LM, Koes BW, Picavet HSJ, van Tulder MW, Burton K, Foster NE, Macfarlane GJ, Thomas E, Underwood M, Waddell G, Shekelle P, Volinn E, Von Korff M. A consensus approach toward the standardization of back pain definitions for use in prevalence studies. Spine (Phila. Pa. 1976) 2008;33:95–103. doi: 10.1097/BRS.0b013e31815e7f94. [DOI] [PubMed] [Google Scholar]
- [14].Dominguez C a, Kalliomaki M, Gunnarsson U, Moen a, Sandblom G, Kockum I, Lavant E, Olsson T, Nyberg F, Rygh LJ, Roe C, Gjerstad J, Gordh T, Piehl F. The DQB1( *)03:02 HLA haplotype is associated with increased risk of chronic pain after inguinal hernia surgery and lumbar disc herniation. Pain. 2013;154:427–433. doi: 10.1016/j.pain.2012.12.003. [DOI] [PubMed] [Google Scholar]
- [15].Fernández-De-Las Peñas C, Ambite-Quesada S, Ortíz-Gutićrrez R, Ortega-Santiago R, Gil-Crujera A, Caminero AB. Catechol-O-methyltransferase val158met polymorphism (rs4680) Is associated with pain in multiple sclerosis. J. Pain. 2013;14:1719–1723. doi: 10.1016/j.jpain.2013.09.007. [DOI] [PubMed] [Google Scholar]
- [16].Ferrari MD, Klever RR, Terwindt GM, Ayata C, van den Maagdenberg AMJM. Migraine pathophysiology: lessons from mouse models and human genetics. Lancet Neurol. 2015;14:65–80. doi: 10.1016/S1474-4422(14)70220-0. [DOI] [PubMed] [Google Scholar]
- [17].Freynhagen R, Baron R, Gockel U, Tölle TR. painDETECT: a new screening questionnaire to identify neuropathic components in patients with back pain. Curr. Med. Res. Opin. 2006;22:1911–1920. doi: 10.1185/030079906X132488. [DOI] [PubMed] [Google Scholar]
- [18].Haanpää M, Attal N, Backonja M, Baron R, Bennett M, Bouhassira D, Cruccu G, Hansson P, Haythornthwaite JA, Iannetti GD, Jensen TS, Kauppila T, Nurmikko TJ, Rice ASC, Rowbotham M, Serra J, Sommer C, Smith BH, Treede RD. NeuPSIG guidelines on neuropathic pain assessment. Pain. 2011;152:14–27. doi: 10.1016/j.pain.2010.07.031. [DOI] [PubMed] [Google Scholar]
- [19].Van Hecke O, Austin SK, Khan R a., Smith BH, Torrance N. Neuropathic pain in the general population: A systematic review of epidemiological studies. Pain. 2014;155:654–662. doi: 10.1016/j.pain.2013.11.013. [DOI] [PubMed] [Google Scholar]
- [20].Hegarty D, Shorten G. Multivariate prognostic modeling of persistent pain following lumbar discectomy. Pain Physician. 2012;15:421–34. [PubMed] [Google Scholar]
- [21].Hendry L, Lombard Z, Wadley A, Kamerman P. KCNS1, but not GCH1, is associated with pain intensity in a black southern African population with HIV-associated sensory neuropathy: a genetic association study. J. Acquir. Immune Defic. Syndr. 2013;63:27–30. doi: 10.1097/QAI.0b013e318285cf36. [DOI] [PubMed] [Google Scholar]
- [22].Jacobsen LM, Schistad EI, Storesund a., Pedersen LM, Rygh LJ, Røe C, Gjerstad J. The COMT rs4680 Met allele contributes to long-lasting low back pain, sciatica and disability after lumbar disc herniation. Eur. J. Pain (United Kingdom) 2012;16:1064–1069. doi: 10.1002/j.1532-2149.2011.00102.x. [DOI] [PubMed] [Google Scholar]
- [23].Jacobsen LM, Schistad EI, Storesund A, Pedersen LM, Espeland A, Rygh LJ, Røe C, Gjerstad J. Increased Low Back Pain , Sciatica , and Disability After Lumbar Disk Herniation. Clin. J. Pain. 2013;29:967–971. doi: 10.1097/AJP.0b013e31827df7fd. [DOI] [PubMed] [Google Scholar]
- [24].Janssens a. CJW, Ioannidis JP a, Van Duijn CM, Little J, Khoury MJ. Strengthening the reporting of genetic risk prediction studies: The GRIPS statement. Eur. J. Clin. Invest. 2011;41:1004–1009. doi: 10.1111/j.1365-2362.2011.02494.x. [DOI] [PubMed] [Google Scholar]
- [25].Janssens a. CJW, Ioannidis JP a., Bedrosian S, Boffetta P, Dolan SM, Dowling N, Fortier I, Freedman AN, Grimshaw JM, Gulcher J, Gwinn M, Hlatky M a., Janes H, Kraft P, Melillo S, O’Donnell CJ, Pencina MJ, Ransohoff D, Schully SD, Seminara D, Winn DM, Wright CF, van Duijn CM, Little J, Khoury MJ. Strengthening the reporting of genetic risk prediction studies (GRIPS): explanation and elaboration. Eur. J. Clin. Invest. 2011;41:1010–1035. doi: 10.1111/j.1365-2362.2011.02493.x. [DOI] [PubMed] [Google Scholar]
- [26].Jensen TS, Baron R, Haanpää M, Kalso E, Loeser JD, Rice ASC, Treede RD. A new definition of neuropathic pain. Pain. 2011;152:2204–2205. doi: 10.1016/j.pain.2011.06.017. [DOI] [PubMed] [Google Scholar]
- [27].Karppinen J, Daavittila I, Noponen N, Haapea M, Taimela S, Vanharanta H, Ala-Kokko L, Männikkö M. Is the interleukin-6 haplotype a prognostic factor for sciatica? Eur. J. Pain. 2008;12:1018–1025. doi: 10.1016/j.ejpain.2008.01.009. [DOI] [PubMed] [Google Scholar]
- [28].Kim H, Clark D, Dionne R a. Genetic Contributions to Clinical Pain and Analgesia: Avoiding Pitfalls in Genetic Research. J. Pain. 2009;10:663–693. doi: 10.1016/j.jpain.2009.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Little J, Higgins JPT, Ioannidis JPA, Moher D, Gagnon F, Von Elm E, Khoury MJ, Cohen B, Davey-Smith G, Grimshaw J, Scheet P, Gwinn M, Williamson RE, Zou GY, Hutchings K, Johnson CY, Tait V, Wiens M, Golding J, Van Duijn C, McLaughlin J, Paterson A, Wells G, Fortier I, Freedman M, Zecevic M, King R, Infante-Rivard C, Stewart A, Birkett N. STrengthening the REporting of genetic association studies (STREGA)- An extension of the STROBE statement. Genet. Epidemiol. 2009;33:581–598. doi: 10.1002/gepi.20410. [DOI] [PubMed] [Google Scholar]
- [30].Manchia M, Cullis J, Turecki G, Rouleau GA, Uher R, Alda M. The Impact of Phenotypic and Genetic Heterogeneity on Results of Genome Wide Association Studies of Complex Diseases. PLoS One. 2013;8:e76295. doi: 10.1371/journal.pone.0076295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [31].Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TFC, McCarroll SA, Visscher PM. Finding the missing heritability of complex diseases. Nature. 2009;461:747–53. doi: 10.1038/nature08494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [32].De Meyrick J. The Delphi method and health research. Health Educ. 2003;103:7–16. [Google Scholar]
- [33].Mogil JS. Pain genetics: Past, present and future. Trends Genet. 2012;28:258–266. doi: 10.1016/j.tig.2012.02.004. [DOI] [PubMed] [Google Scholar]
- [34].Nielsen CS. Definiting human pain phenotypes for genetic association studies. In: Belfer I, Diatchencko L, editors. Pain genetics: basic to translational science. Wiley-Blackwell; Oxford: 2014. pp. 37–50. [Google Scholar]
- [35].Nissenbaum J, Devor M, Seltzer Z, Gebauer M, Michaelis M, Tal M, Dorfman R, Abitbul-Yarkoni M, Lu Y, Elahipanah T, DelCanho S, Minert A, Fried K, Persson AK, Shpigler H, Shabo E, Yakir B, Pisantć A, Darvasi A. Susceptibility to chronic pain following nerve injury is genetically affected by CACNG2. Genome Res. 2010;20:1180–1190. doi: 10.1101/gr.104976.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [36].Noponen-Hietala N, Virtanen I, Karttunen R, Schwenke S, Jakkula E, Li H, Merikivi R, Barral S, Ott J, Karppinen J, Ala-Kokko L. Genetic variations in IL6 associate with intervertebral disc disease characterized by sciatica. Pain. 2005;114:186–194. doi: 10.1016/j.pain.2004.12.015. [DOI] [PubMed] [Google Scholar]
- [37].Ozawa a, Sasao Y, Iwashita K, Miyahara M, Sugai J, Iizuka M, Kawakubo Y, Ohkido M, Naruse T, Anzai T, Takashige N, Ando a, Inoko H. HLA-A33 and -B44 and susceptibility to postherpetic neuralgia (PHN) Tissue Antigens. 1999;53:263–268. doi: 10.1034/j.1399-0039.1999.530306.x. [DOI] [PubMed] [Google Scholar]
- [38].Percie du Sert N, Rice ASC. Improving the translation of analgesic drugs to the clinic: animal models of neuropathic pain. Br. J. Pharmacol. 2014;171:2951–63. doi: 10.1111/bph.12645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [39].Pettersson FH, Anderson C a, Clarke GM, Barrett JC, Cardon LR, Morris AP, Zondervan KT. Marker selection for genetic case-control association studies. Nat. Protoc. 2009;4:743–752. doi: 10.1038/nprot.2009.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [40].Powell C. The Delphi technique: myths and realities. J. Adv. Nurs. 2003;41:376–82. doi: 10.1046/j.1365-2648.2003.02537.x. [DOI] [PubMed] [Google Scholar]
- [41].Ramirez JD, Barnes PRJ, Mills KR, Bennett DLH. Intermediate Charcot-Marie-Tooth disease due to a novel Trp101Stop myelin protein zero mutation associated with debilitating neuropathic pain. Pain. 2012;153:1763–1768. doi: 10.1016/j.pain.2012.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [42].Ripke S, Neale BM, Corvin A, Walters JTR, Farh K-H, Holmans PA, Lee P, Bulik-Sullivan B, Collier DA, Huang H, Pers TH, Agartz I, Agerbo E, Albus M, Alexander M, Amin F, Bacanu SA, Begemann M, Belliveau RA, Jr, Bene J, Bergen SE, Bevilacqua E, Bigdeli TB, Black DW, Bruggeman R, Buccola NG, Buckner RL, Byerley W, Cahn W, Cai G, Campion D, Cantor RM, Carr VJ, Carrera N, Catts SV, Chambert KD, Chan RCK, Chen RYL, Chen EYH, Cheng W, Cheung EFC, Ann Chong S, Robert Cloninger C, Cohen D, Cohen N, Cormican P, Craddock N, Crowley JJ, Curtis D, Davidson M, Davis KL, Degenhardt F, Del Favero J, Demontis D, Dikeos D, Dinan T, Djurovic S, Donohoe G, Drapeau E, Duan J, Dudbridge F, Durmishi N, Eichhammer P, Eriksson J, Escott-Price V, Essioux L, Fanous AH, Farrell MS, Frank J, Franke L, Freedman R, Freimer NB, Friedl M, Friedman JI, Fromer M, Genovese G, Georgieva L, Giegling I, Giusti-Rodríguez P, Godard S, Goldstein JI, Golimbet V, Gopal S, Gratten J, de Haan L, Hammer C, Hamshere ML, Hansen M, Hansen T, Haroutunian V, Hartmann AM, Henskens FA, Herms S, Hirschhorn JN, Hoffmann P, Hofman A, Hollegaard MV, Hougaard DM, Ikeda M, Joa I, Julià A, Kahn RS, Kalaydjieva L, Karachanak-Yankova S, Karjalainen J, Kavanagh D, Keller MC, Kennedy JL, Khrunin A, Kim Y, Klovins J, Knowles JA, Konte B, Kucinskas V, Ausrele Kucinskiene Z, Kuzelova-Ptackova H, Kähler AK, Laurent C, Lee Chee Keong J, Hong Lee S, Legge SE, Lerer B, Li M, Li T, Liang K-Y, Lieberman J, Limborska S, Loughland CM, Lubinski J, Lönnqvist J, Macek M, Jr, Magnusson PKE, Maher BS, Maier W, Mallet J, Marsal S, Mattheisen M, Mattingsdal M, McCarley RW, McDonald C, McIntosh AM, Meier S, Meijer CJ, Melegh B, Melle I, Mesholam-Gately RI, Metspalu A, Michie PT, Milani L, Milanova V, Mokrab Y, Morris DW, Mors O, Murphy KC, Murray RM, Myin-Germeys I, Müller-Myhsok B, Nelis M, Nenadic I, Nertney DA, Nestadt G, Nicodemus KK, Nikitina-Zake L, Nisenbaum L, Nordin A, O’Callaghan E, O’Dushlaine C, O’Neill FA, Oh S-Y, Olincy A, Olsen L, Van Os J, Endophenotypes International Consortium P. Pantelis C, Papadimitriou GN, Papiol S, Parkhomenko E, Pato MT, Paunio T, Pejovic-Milovancevic M, Perkins DO, Pietiläinen O, Pimm J, Pocklington AJ, Powell J, Price A, Pulver AE, Purcell SM, Quested D, Rasmussen HB, Reichenberg A, Reimers MA, Richards AL, Roffman JL, Roussos P, Ruderfer DM, Salomaa V, Sanders AR, Schall U, Schubert CR, Schulze TG, Schwab SG, Scolnick EM, Scott RJ, Seidman LJ, Shi J, Sigurdsson E, Silagadze T, Silverman JM, Sim K, Slominsky P, Smoller JW, So H-C, Spencer CA, Stahl EA, Stefansson H, Steinberg S, Stogmann E, Straub RE, Strengman E, Strohmaier J, Scott Stroup T, Subramaniam M, Suvisaari J, Svrakic DM, Szatkiewicz JP, Söderman E, Thirumalai S, Toncheva D, Tosato S, Veijola J, Waddington J, Walsh D, Wang D, Wang Q, Webb BT, Weiser M, Wildenauer DB, Williams NM, Williams S, Witt SH, Wolen AR, Wong EHM, Wormley BK, Simon Xi H, Zai CC, Zheng X, Zimprich F, Wray NR, Stefansson K, Visscher PM, Trust Case-Control Consortium W. Adolfsson R, Andreassen OA, Blackwood DHR, Bramon E, Buxbaum JD, Børglum AD, Cichon S, Darvasi A, Domenici E, Ehrenreich H, Esko T, Gejman PV, Gill M, Gurling H, Hultman CM, Iwata N, Jablensky AV, Jönsson EG, Kendler KS, Kirov G, Knight J, Lencz T, Levinson DF, Li QS, Liu J, Malhotra AK, McCarroll SA, McQuillin A, Moran JL, Mortensen PB, Mowry BJ, Nöthen MM, Ophoff RA, Owen MJ, Palotie A, Pato CN, Petryshen TL, Posthuma D, Rietschel M, Riley BP, Rujescu D, Sham PC, Sklar P, St Clair D, Weinberger DR, Wendland JR, Werge T, Daly MJ, Sullivan PF, O’Donovan MC. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–7. doi: 10.1038/nature13595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [43].Sato M, Ohashi J, Tsuchiya N, Kashiwase K, Ishikawa Y, Arita H, Hanaoka K, Tokunaga K, Yabe T. Association of HLA-A*3303-B*4403-DRB1*1302 haplotype, but not of TNFA promoter and NKp30 polymorphism, with postherpetic neuralgia (PHN) in the Japanese population. Genes Immun. 2002;3:477–481. doi: 10.1038/sj.gene.6363890. [DOI] [PubMed] [Google Scholar]
- [44].Smith BH, Macfarlane GJ, Torrance N. Epidemiology of chronic pain, from the laboratory to the bus stop: time to add understanding of biological mechanisms to the study of risk factors in population-based research? Pain. 2007;127:5–10. doi: 10.1016/j.pain.2006.11.001. [DOI] [PubMed] [Google Scholar]
- [45].Smith BH, Torrance N, Ferguson J a., Bennett MI, Serpell MG, Dunn KM. Towards a definition of refractory neuropathic pain for epidemiological research. An international Delphi survey of experts. BMC Neurol. 2012;12:29. doi: 10.1186/1471-2377-12-29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [46].Sumiyama D, Kikkawa EF, Kita YF, Shinagawa H, Mabuchi T, Ozawa A, Inoko H. HLA alleles are associated with postherpetic neuralgia but not with herpes zoster. Tokai J. Exp. Clin. Med. 2008;33:150–153. [PubMed] [Google Scholar]
- [47].Tegeder I, Costigan M, Griffin RS, Abele A, Belfer I, Schmidt H, Ehnert C, Nejim J, Marian C, Scholz J, Wu T, Allchorne A, Diatchenko L, Binshtok AM, Goldman D, Adolph J, Sama S, Atlas SJ, Carlezon W a, Parsegian A, Lötsch J, Fillingim RB, Maixner W, Geisslinger G, Max MB, Woolf CJ. GTP cyclohydrolase and tetrahydrobiopterin regulate pain sensitivity and persistence. Nat. Med. 2006;12:1269–1277. doi: 10.1038/nm1490. [DOI] [PubMed] [Google Scholar]
- [48].Treede RD, Jensen TS, Campbell JN, Cruccu G, Dostrovsky JO, Griffin JW, Hansson P, Hughes R, Nurmikko T, Serra J. Neuropathic pain: Redefinition and a grading system for clinical and research purposes. Neurology. 2008;70:1630–1635. doi: 10.1212/01.wnl.0000282763.29778.59. [DOI] [PubMed] [Google Scholar]
- [49].Wadley AL, Lombard Z, Cherry CL, Price P, Kamerman PR. Analysis of a Previously Identified “Pain-Protective” Haplotype and Individual Polymorphisms in the GCH1 Gene in Africans With HIV-Associated Sensory Neuropathy. J. Acquir. Immune Defic. Syndr. 2012;60:20–23. doi: 10.1097/QAI.0b013e31824bcc17. [DOI] [PubMed] [Google Scholar]
- [50].Wong MY, Day NE, Luan JA, Chan KP, Wareham NJ. The detection of gene-environment interaction for continuous traits: Should we deal with measurement error by bigger studies or better measurement? Int. J. Epidemiol. 2003;32:51–57. doi: 10.1093/ije/dyg002. [DOI] [PubMed] [Google Scholar]
- [51].Wright AF, Carothers AD, Campbell H. Gene-environment interactions--the BioBank UK study. Pharmacogenomics J. 2002;2:75–82. doi: 10.1038/sj.tpj.6500085. [DOI] [PubMed] [Google Scholar]
- [52].Zhang W, Moskowitz RW, Nuki G, Abramson S, Altman RD, Arden N, Bierma-Zeinstra S, Brandt KD, Croft P, Doherty M, Dougados M, Hochberg M, Hunter DJ, Kwoh K, Lohmander LS, Tugwell P. OARSI recommendations for the management of hip and knee osteoarthritis, Part II: OARSI evidence-based, expert consensus guidelines. Osteoarthritis Cartilage. 2008;16:137–62. doi: 10.1016/j.joca.2007.12.013. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.