Abstract
“Orangutan” is derived from the Malay term “man of the forest” and aptly describes the Southeast Asian great apes native to Sumatra and Borneo. The orangutan species, Pongo abelii (Sumatran) and Pongo pygmaeus (Bornean), are the most phylogenetically distant great apes from humans, thereby providing an informative perspective on hominid evolution. Here we present a Sumatran orangutan draft genome assembly and short read sequence data from five Sumatran and five Bornean orangutan genomes. Our analyses reveal that, compared to other primates, the orangutan genome has many unique features. Structural evolution of the orangutan genome has proceeded much more slowly than other great apes, evidenced by fewer rearrangements, less segmental duplication, a lower rate of gene family turnover and surprisingly quiescent Alu repeats, which have played a major role in restructuring other primate genomes. We also describe the first primate polymorphic neocentromere, found in both Pongo species, emphasizing the gradual evolution of orangutan genome structure. Orangutans have extremely low energy usage for a eutherian mammal1, far lower than their hominid relatives. Adding their genome to the repertoire of sequenced primates illuminates new signals of positive selection in several pathways including glycolipid metabolism. From the population perspective, both Pongo species are deeply diverse; however, Sumatran individuals possess greater diversity than their Bornean counterparts, and more species-specific variation. Our estimate of Bornean/Sumatran speciation time, 400k years ago (ya), is more recent than most previous studies and underscores the complexity of the orangutan speciation process. Despite a smaller modern census population size, the Sumatran effective population size (Ne) expanded exponentially relative to the ancestral Ne after the split, while Bornean Ne declined over the same period. Overall, the resources and analyses presented here offer new opportunities in evolutionary genomics, insights into hominid biology, and an extensive database of variation for conservation efforts.
Orangutans are the only primarily arboreal great apes, characterized by strong sexual dimorphism and delayed development of mature male features, a long lifespan (35-45 years in the wild, over 55 years in captivity) and the longest interbirth interval among mammals (8 years on average)2. Orangutans create and adeptly use tools in the wild, and while long presumed socially solitary, dense populations of Sumatran orangutans show complex social structure and geographic variability in tool use indicative of cultural learning3. Both species have been subject to intense population pressure from loss of habitat, deforestation, hunting and disease. A 2004 study estimated 7,000-7,500 Sumatran individuals and 40,000-50,000 Bornean individuals remained in the wild in fragmented subpopulations4,5. The International Union for Conservation of Nature lists Sumatran orangutans as critically endangered and Bornean orangutans as endangered.
We sequenced the genome of a female Sumatran orangutan using a whole-genome shotgun strategy. The assembly provides 5.5-fold coverage on average across 3.08 gigabases (Gb) of ordered and oriented sequence (Table 1)(S1). Accuracy was assessed by several metrics, including comparison to 17 megabases (Mb) of finished bacterial artificial chromosome (BAC) sequences and a novel method of detecting spurious insertions and deletions (S2). Further validation resulted from orangutan-human divergence estimates based on alignment of whole-genome shotgun reads to the human reference (Hs.35)(Fig 1)(S3). We also sequenced the genomes of 10 additional unrelated wild-caught orangutans, five Sumatran and five Bornean, using a short read sequencing platform (297 Gb of data total)(S4). The orangutan gene set was constructed using a combination of human gene models and orangutan cDNA data generated for this project (www.ensembl.org/Pongo_pygmaeus/Info/StatsTable)(S5).
Table 1.
Sumatran orangutan assembly statistics (ponAbe2).
| Total Contig Bases | 3.09 Gb |
| Total Contig Bases >Phred Q20 | 3.05 Gb (98.5%) |
| Ordered/Oriented Contigs & Scaffolds | 3.08 Gb |
| Number of Contigs >1 kb | 410,172 |
| N50 Contig Length | 15.5 kb |
| N50 Contig Number | 55,989 |
| Number of Scaffolds >2 kb | 77,683 |
| N50 Scaffold Length | 739 kb |
| N50 Scaffold Number | 1,031 |
| Average Read Depth | 5.53x |
Figure 1. Divergence among great apes, a lesser ape, and an old world monkey with respect to humans.
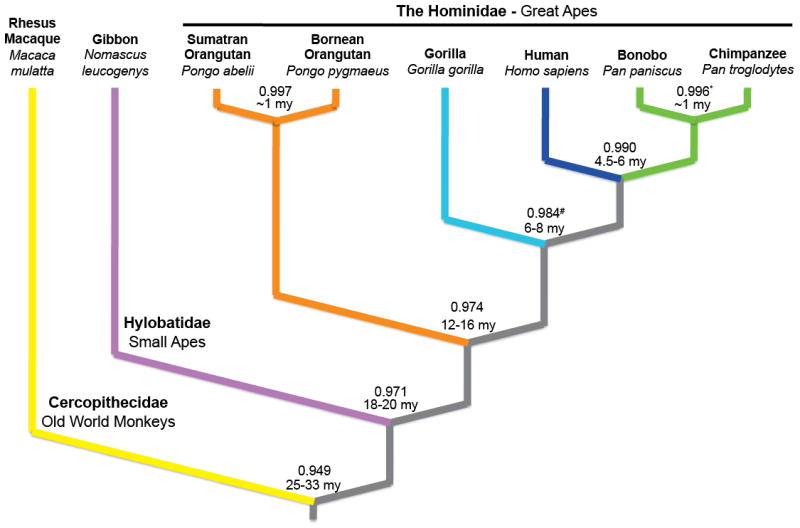
We estimated nucleotide divergence in unique gap-free sequence, indicated at each node, from the alignment of rhesus macaque (yellow), gibbon (purple), orangutan (orange), gorilla (aqua), chimpanzee (green) and human (blue) whole genome shotgun reads to the human reference (Hs.35)(S3). Note that the Bornean (Pongo pygmaeus) and Sumatran (Pongo abelii) orangutan species showed nucleotide identity comparable to that of bonobo (Pan paniscus) and chimpanzee (Pan troglodytes). *Yu et al. 200329, #Chen and Li 200130.
Among hominids, the orangutan karyotype is the most ancestral6, and sequencing the orangutan genome allowed a comprehensive assessment of conservation among the wide range of rearrangement types and sequence classes involved in structural variation. We characterized orangutan synteny breaks in detail cytogenetically in concert with an in silico approach that precisely tracked rearrangements between primate (human, chimpanzee, orangutan and rhesus macaque) and other mammalian assemblies (mouse, rat and dog)(S6). Alignment-level analyses at 100 kb and 5 kb resolution found the orangutan genome underwent fewer rearrangements than the chimpanzee or human genomes, with a bias for large-scale events (>100 kb) on the chimpanzee branch (Table 2). Orangutan large-scale rearrangements were further enriched for segmental duplications (SD)(52%) than for small-scale events (27%), suggesting mechanisms other than non-allelic homologous recombination may have made a greater contribution to small rearrangements. Genome-wide, we estimated less segmental duplication content (3.8% total) in the orangutan genome compared to the chimpanzee and human genomes (5%) using equivalent methods (S11). We also assessed the rate of turnover within gene families as an additional measure of genome restructuring (S12). Our analysis indicated that the human and chimpanzee lineages, as well as their shared ancestral lineage after the orangutan split, had the highest rates of gene turnover among great apes (0.0058 events/gene/my) – over twice the rate of the orangutan and macaque lineages (0.0027) – even as the nucleotide substitution rate decreased7. Collectively these data strongly suggest structural evolution proceeded much more slowly along the orangutan branch, in sharp contrast to the acceleration of structural variation noted for the chimpanzee and human genomes8,9.
Table 2.
Number of genome rearrangements by species.
| Species | Rearrangements >100 kb | Rearrangements >5 kb |
|---|---|---|
| Orangutan | 38 | 861 |
| Chimpanzee | 85 (+124%) | 1095 (+27%) |
| Human | 54 (+42%) | 1238 (44%) |
The number in parentheses indicates the %Δ with respect to the orangutan genome. Note 40 events >100 kb and 532 events >5 kb were assigned to the human-chimpanzee ancestor by ancestral reconstruction (S6).
One structural variant we characterized in detail was a previously described polymorphic “pericentric inversion” of orangutan chromosome 1210. Surprisingly, both forms of this chromosome showed no difference in marker order by fluorescence in situ hybridization (FISH) despite two distinct centromere positions – the hallmark of a neocentromere (Fig 2)(S8). Neocentromere function was confirmed by chromatin immunoprecipitation with antibodies to centromeric proteins CENP-A and CENP-C and subsequent oligo array hybridization (ChIP-on-chip), which narrowed the neocentromere to a ~225 kb gene-free window devoid of alpha satellite-related sequences. Our observations bore similarity to a recently described centromere repositioning event in the horse genome11; however, this is the first observation of such a variant among primates, with the additional complexity of polymorphism in two closely related species. Potentially related, orangutan chromosome 12 did not show any appreciable centromeric alphoid FISH signal in comparison to other autosomes. The neocentromere likely arose prior to the Bornean/Sumatran split as it is found in both species, and represents a unique opportunity to study the initial stages of centromere formation and the impact of such a large chromosomal variant on population variation and recombination.
Figure 2. The neocentromere of orangutan chromosome 12.
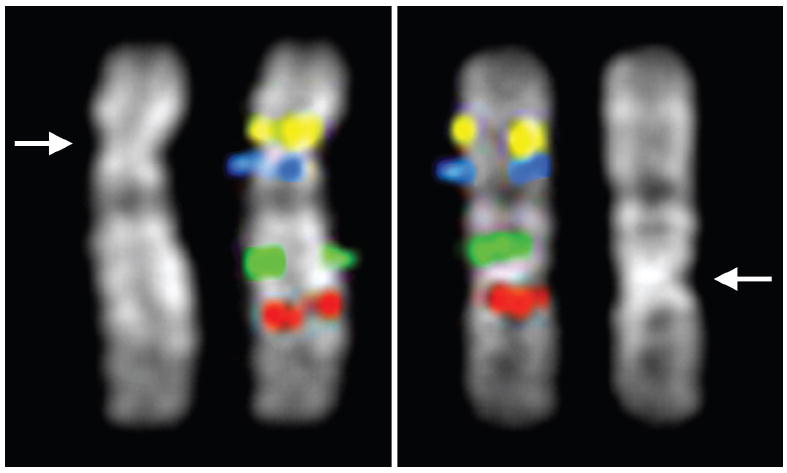
Note the identical order of four BAC-derived FISH probes (IDs in S8) between the normal (left panel) and neocentromere-bearing (right panel) configurations of orangutan chromosome 12, despite discordant centromere positions indicated by arrows on the adjacent DAPI-only images. The neocentromere recruits centromeric proteins CENP-A and CENP-C and lies within a ~225 kb gene-free and alpha satellite-free region. The neocentromere-bearing variant is polymorphic in both Bornean and Sumatran populations, suggesting the neocentromere arose prior to the Bornean/Sumatran split, yet has not been fixed in either species.
The orangutan genome has a comparable cadre of mobile elements to that of other primates, comprising roughly half the genome12,13,14. Orangutan LINE1 (L1) and SVA expansions were expectedly broad, with roughly 5,000 and 1,800 new insertions respectively, consistent with other primates (S9). Surprisingly, Alu elements were relatively quiescent, with only ~250 recent insertions identified by computational and laboratory approaches (Fig 3). By comparison, 5,000 human-specific and 2,300 chimpanzee-specific Alu elements were identified by similar methods. The rate of processed pseudogene formation, which like Alu insertion requires functional L1 machinery, was similar for the human (8.0/my), chimpanzee (12.7/my) and orangutan (11.6/my) lineages (S10). We identified a small number of polymorphic Alu elements exclusive to Pongo abelii (S19), indicating that Alu retroposition has been strongly limited, but not eliminated. This dramatic Alu-specific repression represents an unprecedented change in primate retrotransposition rates16,17. Possible explanations include L1 source mutations that lowered Alu affinity and cis mobilization preference18, pressure against Alu retroposition from the APOBEC RNA editing family19, or fixation of less effectively propagated Alu “master” variants.
Figure 3. Alu quiescence in the orangutan lineage.
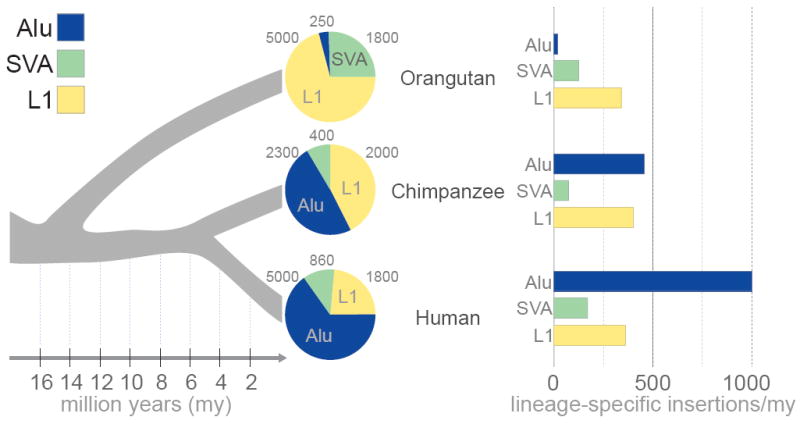
We identified only ~250 lineage-specific Alu retroposition events in the orangutan genome, a dramatically lower rate than that of other sequenced primates, including humans. The total number of lineage-specific L1, SVA and Alu insertions is shown (pie chart), along with the rate of insertion events per element type (bar graph). Reduced Alu retroposition potentially limited the effect of a wide variety of repeat-driven mutational mechanisms in the orangutan lineage that played a major role in restructuring other primate genomes.
It is tempting to propose a correlation between reduced Alu retroposition and the greater structural stability of the orangutan genome. Over one million (M) Alu elements exist within primate genomes. Because of their large copy number and high sequence identity, Alu repeats play a crucial role in multiple forms of structural variation through insertion and post-insertion recombination20. By virtue of reduced Alu retroposition, the orangutan lineage experienced fewer new insertions and a putative decrease in the number of regions susceptible to post-insertion Alu-mediated recombination events genome-wide, limiting the overall mobile element threat to the genome.
The unique phylogenetic position of Pongo species also offered the opportunity to detect signals of positive selection with increased power. We assessed positive selection in 13,872 human genes with high-confidence orthologs in the orangutan genome, and in one or more of the chimpanzee, rhesus macaque and dog genomes, using branch-site likelihood ratio tests (S15)14,21. Two new Gene Ontology (GO) categories were statistically enriched for positive selection in primates: “visual perception” and “glycolipid metabolic processes”22. The enrichment for visual perception includes strong evidence from two major visual signalling proteins: arrestin (SAG, P=0.007) and recoverin (RCVRN, P=0.008), as well as the opsin, OPN1SW1 (P=0.020), associated with blue color vision23. The enrichment for glycolipid metabolism is interesting due to medium-to-strong evidence for positive selection (nominal P<0.05) from six genes expressed in nervous tissue that cluster in the cerebroside-sulfatid region of the sphingolipid metabolism pathway (Fig 4). This pathway is associated with human neurodegenerative diseases such as Gaucher’s, Sandhoff’s, Tay-Sachs, and metachromatic leukodystrophy. Variation in lipid metabolism may have impacted neurological evolution among primates, and diversity of diets and life history strategies, as apes – especially orangutans – have slower rates of reproduction and dramatically lower energy usage than other primates and mammals1.
Figure 4. Enrichment for positive selection in the cerebroside-sulfatid metabolism pathway.
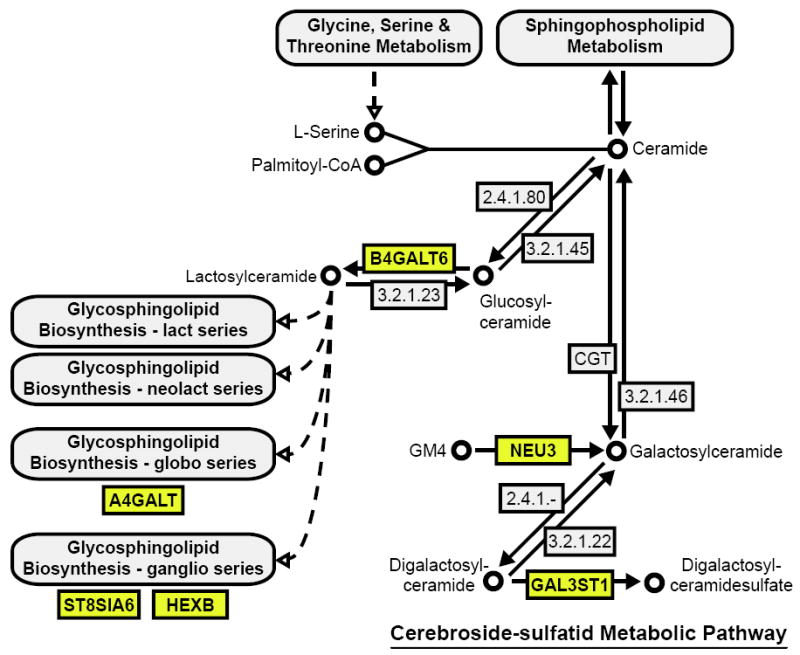
We identified six genes (indicated in yellow) under moderate to strong positive selection in primates (P<0.05) that fall within the cerebroside-sulfatid region of the sphingolipid metabolism pathway (adapted from human KEGG pathway 00600; http://www.genome.jp/kegg/kegg2.html). This pathway is associated with several human lysosomal storage disorders, such as Gaucher’s disease, Sandhoff’s disease, Tay Sachs disease and metachromatic leukodystrophy.
Ancestral orangutan species ranged broadly across Southeast Asia, including the mainland, while modern species are geographically restricted to their respective islands due to environmental forces and human population expansion. Historically, protein markers, restriction fragment length polymorphisms, and small sets of mitochondrial and nuclear markers have been used to estimate the divergence and diversity of orangutan species. We employed short read sequencing to address this question from a genome-wide perspective. We first estimated average Bornean/Sumatran nucleotide identity genome-wide (99.68%) based on the alignment of 20-fold coverage of short read data from a Bornean individual to the Sumatran reference (S16). We then called SNPs from the alignment of all short read data from 10 individuals (five Bornean, including the 20-fold coverage mentioned above, and five Sumatran)(S4). We analyzed each species separately using a Bayesian approach with 92% power to detect SNPs (S20). Because of relatively deep sequencing, allele frequency spectra (AFS) were estimated accurately, but with an overestimation of singletons compared to other allele frequency categories of approximately 7.8% based on re-sequencing a subset of SNPs (n=108)(S20). This level of error had only a marginal effect on downstream population genetic analyses (S21). Overall, 99.0% (931/940) of genotypes were accurately called within the re-sequenced subset of SNPs.
In total, we identified 13.2 M putative SNPs across 1.96 Gb of the genome, or 1 SNP every 149 bp on average. Within the Bornean and Sumatran groups we detected 6.69 M (3.80 M Bornean-exclusive) and 8.96 M (5.19 M Sumatran-exclusive) SNPs, respectively (Fig 5). Observing 36% more SNPs among Sumatran individuals strongly supports a larger Ne. In addition, independent analysis of 85 polymorphic retroelement loci among 37 individuals (19 Sumatran, 18 Bornean) also showed more complex Sumatran population structure (S19). Using Watterson’s approach24 we estimated nucleotide diversity from the SNP data as θW = 1.21 and θW = 1.62 per kb for the Bornean and Sumatran species, respectively, and θW = 1.89 per kb for the orangutan species combined, roughly twice the diversity of modern humans25.
Figure 5. Orangutan population genetics and demographics.
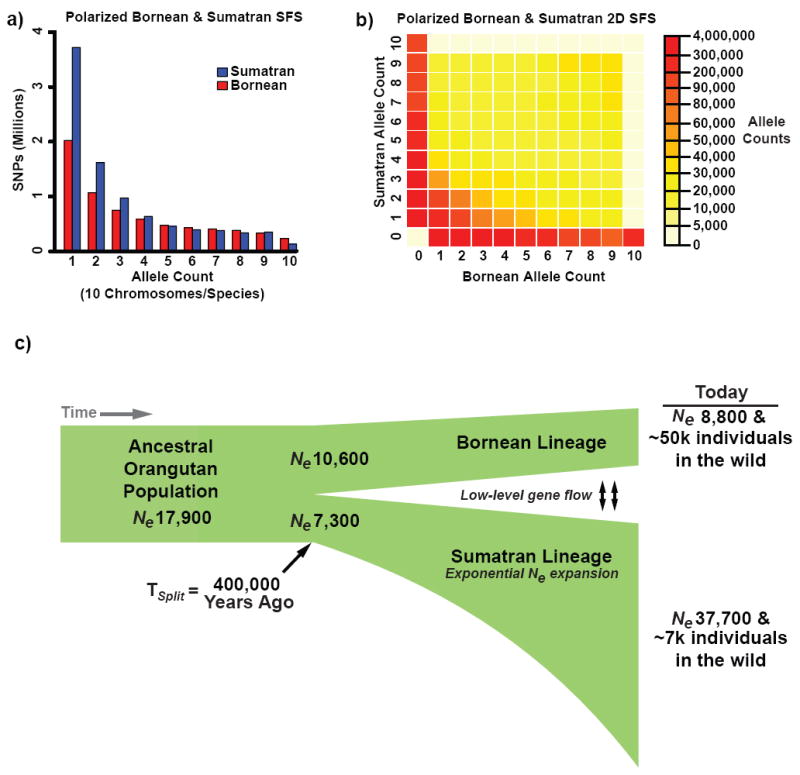
a. Site-frequency spectra (SFS) for 13.2 million Bornean (blue) and Sumatran (red) SNPs are shown, note the enrichment of low-frequency SNPs among Sumatran individuals. b. The majority of SNPs were restricted to their respective island populations as the ‘heat’ of the 2D SFS, representing high allele counts, lay along the axes. c. Our demographic model estimated the ancestral orangutan population (Ne = 17,900) split approximately 400,000 years ago, followed by exponential expansion of Sumatran Ne and a decline of Bornean Ne, culminating in higher diversity among modern Sumatran orangutans despite a lower census population size. The model also supported low-level gene flow (<1 individual/generation) indicated by arrows.
The modal category of SNPs were singletons, with 2.0 M and 3.7 M SNPs observed as single heterozygous sites in a Bornean or Sumatran individual, consistent with the expectation that most genetic variation for an outcrossing population ought to be rare due to mutation drift equilibrium. We observed little correlation between Bornean and Sumatran SNPs in the AFS (i.e., the “heat” of the map is not along the diagonal as expected for populations with similar allele frequencies, but rather along the edges)(Fig 5b). This was further supported by Principal Component Analysis, in which PC1 corresponded to the Bornean/Sumatran population label and explained 36% of the variance (S20).
Based on these data, our demographic model consisted of a two-population model with divergence and potential migration, growth and difference in population size (S21). Among several models tested we found very strong statistical support (105 log-likelihood units) for the most complex model, which included a split with growth and subsequent low-level migration. We estimated a relative Ne of 210% for Sumatran orangutans relative to the ancestral and 49% for Bornean orangutans, noting a four-fold difference for the derived populations (Fig 5c). Assuming a mutation rate of 2.0×10-8 and 20 years per generation, we estimated an ancestral Ne of 17,900 and a split time of 400k ya.
Parallel to the SNP-based effort, we employed a coalescent hidden Markov model (coal-HMM) approach to estimate speciation time, recombination rate and ancestral Ne from the alignment of 20-fold coverage of a Bornean individual to the Sumatran reference (S17). This method also supported a relatively recent Bornean/Sumatran speciation time (334k ± 145k ya), and estimated a recombination rate of 0.95 ± 0.72 cM/Mb. We independently estimated the ancestral Ne of the autosomes (26,800 ± 6,700) and the X chromosome (20,400 ± 7,400), which was consistent with the theoretical ¾ effective population size of X chromosomes compared to autosomes. The Bornean and Sumatran X chromosome thus diverged as expected, in contrast to the human-chimpanzee speciation process26,27.
The orangutan story is thus a tale of two islands with distinct evolutionary histories. Our high-resolution population studies explored the counter-intuitive nature of orangutan diversity – greater variation among Sumatran orangutans than their Bornean counterparts despite a smaller population size (approximately 7-fold lower by recent estimates). Further dissection of the orangutan speciation process will require a broader survey, incorporating representatives from additional orangutan subpopulations.
Finally, even though we found deep diversity in both Bornean and Sumatran populations, it is not clear whether this diversity will be maintained with continued habitat loss and population fragmentation. Evidence from other species suggests fragmentation is not the death knell of diversity28, but their slow reproduction rate and arboreal lifestyle may leave orangutan species especially vulnerable to rapid dramatic environmental change. It is our hope that the genome assembly and population variation data presented here provide a valuable resource to the community to aid the preservation of these precious species.
Methods Summary
Whole-genome sequencing was performed as described previously12,13,14. The genome assembly was constructed with a custom computational pipeline (S1). Assembly source DNA was derived from a single Sumatran female (Susie; Studbook #1044; ISIS #71), courtesy of the Gladys Porter Zoo, Brownsville, Texas. Short fragment sequencing libraries for population studies (S4) were constructed in accordance with standard Illumina protocols and sequenced on the Illumina GAIIx platform. The resulting data were processed with Illumina base-calling software and analyzed using custom computational pipelines. See Supplemental Information for additional details.
Supplementary Material
Acknowledgments
The orangutan genome project was funded by the National Human Genome Research Institute (NHGRI), including grants U54 HG003079 (R.K.W.) and U54 HG003273 (R.A.G), with further support from National Institutes of Health R01 GM59290 (M.A.B.), PO1 AG022064 (M.A.B.), HG002385 (E.E.E.) and HG002238 (W.M.), National Science Foundation DBI-0644111 (A.S. and B.B.), David and Lucile Packard Foundation (A.S., V.T. and T.V.), Cornell University Provost’s Fellowship (A.L.M.), UK Medical Research Council (C.P.P., G.L., S.M., and A.H.), Marie Curie Fellowship (T.M.-B.), Ministerio de Ciencia e Innovación-Spain (MCI-Spain) and Fundación M. Botín (V.Q., X.P., G.O., and C.L.-O.), MCI-Spain BFU2006-15413-C02-01 and BFU2009-13409-C02-02 (A.N.), Spanish National Institute for Bioinformatics (INAB) and SFRH/BPD/26384/2006 from the Fundação para a Ciência e a Tecnologia (R.F.), PRIN and CEGBA (M.R., N.A. and G.D.V.), and the Commission of the European Communities IRG-224885 (T.V.), IRG-231025 (B.B.). We thank the Gladys Porter Zoo, Brownsville, Texas, and Drs. Stephen O’Brien and Svante Pääbo for use of orangutan samples. D.P.L. would like to thank Sean D. McGrath, Aye Wollam and Rachel M. Abbott for technical assistance. We would also like to recognize all the important work that could not be cited due to space limitations. Resources for exploring the orangutan genome are available at UCSC (http://genome.ucsc.edu), Ensembl (http://www.ensembl.org), NCBI (http://ncbi.nlm.nih.gov) and The Genome Center at WashU (http://genome.wustl.edu/genomes/view/pongo_abelii/).
Footnotes
Author Contributions
D.P.L. led the project and manuscript preparation. D.P.L., A.S., T.M.-B., C.P.P., M.A.B., A.N., E.E.E., M.W.H., C.L.-O., C.D.B., J.M. and M.H.S. led the analyses. Sanger data production, assembly construction, testing and submission: L.W.H, W.C.W., S.-P.Y., Z.W., A.T.C., P.M., M.M., L.A.F., R.A.F., J.O.N., C.P., K.C.W, L.V.N., D.M.M., A.C., H.H.D., J.H., C.L.K., G.R.F. and J.R. BAC sequencing: T.A.G. 454 cDNA sequencing: V.M. and C.M. Illumina sequencing: L.C., K.D.D. and C.F. SNP validation: H.S. Indel assessment: G.L., S.M., A.H. and C.P.P. Segmental duplication, divergence and structural variation studies: T.M.-B., C.A., L.C., Z.C., J.M.K. and E.E.E. Gene models: S.W., S.S. and A.J.V. Assembly-based SNPs: Y.C. and P.F. Ancestral reconstruction and rearrangement analyses: J.M., B.R., B.S., R.B., J.H., D.H., R.S.H. and W.M. Regional variation in nucleotide divergence analyses: R.F., O.F., F.D., D.F., E.G., M.O. and A.N. Cytogenetics and neocentromere characterization: R.R., O.C., N.A., G.D.V., S.P and M.R. Repeat analyses: M.K.K., J.A.W., B.U., M.A.B., A.F.A.S., and R.H. Gene family evolution analyses: C.C., D.R.S. and M.W.H. Protease gene family studies: V.Q., X.S.P., G.R.O. and C.L.-O. Ortholog and defensin analyses: T.V., B.B., A.R., W.M. Positive selection analyses: C.K., T.V., H.A.L., V.T., A.L.M. and A.S. Short read alignments, SNP calling and population genetics: A.R., X.M., J.D. and C.D.B. Demographic analyses: R.N.G. Coalescent-HMM analyses: T.M., J.Y.D., A.H. and M.H.S. Orangutan samples for diversity sequencing: L.C. and O.A.R. BAC library construction: Y.Y. and P.J.dJ. Principle investigators: G.M.W., E.R.M., R.A.G. and R.K.W.
Author Information The Pongo abelii whole-genome shotgun project has been deposited in DDBJ/EMBL/GenBank under the project accession ABGA00000000. The version described in this paper is ABGA00000000.1. Assembly-based SNPs and SNPs derived from short read sequence data have been deposited in dbSNP. All short read data have been deposited into the Short Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accessions listed in supplemental files.
The authors declare no competing financial interests.
References
- 1.Pontzer H, Raichlen DA, Shumaker RW, Ocobock C, Wich SA. Metabolic adaptation for low energy throughput in orangutans. Proc Natl Acad Sci U S A. 107:14048–14052. doi: 10.1073/pnas.1001031107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.van Noordwijk MA, van Schaik CP. Development of ecological competence in Sumatran orangutans. Am J Phys Anthropol. 2005;127:79–94. doi: 10.1002/ajpa.10426. [DOI] [PubMed] [Google Scholar]
- 3.van Schaik CP, et al. Orangutan cultures and the evolution of material culture. Science. 2003;299:102–105. doi: 10.1126/science.1078004. [DOI] [PubMed] [Google Scholar]
- 4.Singleton I, Wich SA, Husson S, Atmoko SU, Leighton M, Rosen N, Traylor-Holzer K, Lacy R, Byers O. Orangutan Population and Habitat Viability Assessment: Final Report. Apple Valley, MN, USA: 2004. [Google Scholar]
- 5.Meijaard E, Wich S. Putting orang-utan population trends into perspective. Curr Biol. 2007;17:R540. doi: 10.1016/j.cub.2007.05.016. [DOI] [PubMed] [Google Scholar]
- 6.Stanyon R, et al. Primate chromosome evolution: ancestral karyotypes, marker order and neocentromeres. Chromosome Res. 2008;16:17–39. doi: 10.1007/s10577-007-1209-z. [DOI] [PubMed] [Google Scholar]
- 7.Yi S, Ellsworth DL, Li WH. Slow molecular clocks in Old World monkeys, apes, and humans. Mol Biol Evol. 2002;19:2191–2198. doi: 10.1093/oxfordjournals.molbev.a004043. [DOI] [PubMed] [Google Scholar]
- 8.Hahn MW, Demuth JP, Han SG. Accelerated rate of gene gain and loss in primates. Genetics. 2007;177:1941–1949. doi: 10.1534/genetics.107.080077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Marques-Bonet T, et al. A burst of segmental duplications in the genome of the African great ape ancestor. Nature. 2009;457:877–881. doi: 10.1038/nature07744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Seuanez H, Fletcher J, Evans HJ, Martin DE. A chromosome rearrangement in orangutan studied with Q-, C-, and G-banding techniques. Cytogenet Cell Genet. 1976;17:26–34. doi: 10.1159/000130684. [DOI] [PubMed] [Google Scholar]
- 11.Wade CM, et al. Genome sequence, comparative analysis, and population genetics of the domestic horse. Science. 2009;326:865–867. doi: 10.1126/science.1178158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Initial sequence of the chimpanzee genome and comparison with the human genome. Nature. 2005;437:69–87. doi: 10.1038/nature04072. [DOI] [PubMed] [Google Scholar]
- 13.Lander ES, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- 14.Gibbs RA, et al. Evolutionary and biomedical insights from the rhesus macaque genome. Science. 2007;316:222–234. doi: 10.1126/science.1139247. [DOI] [PubMed] [Google Scholar]
- 15.Esnault C, Maestre J, Heidmann T. Human LINE retrotransposons generate processed pseudogenes. Nat Genet. 2000;24:363–367. doi: 10.1038/74184. [DOI] [PubMed] [Google Scholar]
- 16.Liu G, et al. Analysis of primate genomic variation reveals a repeat-driven expansion of the human genome. Genome Res. 2003;13:358–368. doi: 10.1101/gr.923303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee J, et al. Different evolutionary fates of recently integrated human and chimpanzee LINE-1 retrotransposons. Gene. 2007;390:18–27. doi: 10.1016/j.gene.2006.08.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Kulpa DA, Moran JV. Cis-preferential LINE-1 reverse transcriptase activity in ribonucleoprotein particles. Nat Struct Mol Biol. 2006;13:655–660. doi: 10.1038/nsmb1107. [DOI] [PubMed] [Google Scholar]
- 19.Bogerd HP, et al. Cellular inhibitors of long interspersed element 1 and Alu retrotransposition. Proc Natl Acad Sci U S A. 2006;103:8780–8785. doi: 10.1073/pnas.0603313103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Cordaux R, Batzer MA. The impact of retrotransposons on human genome evolution. Nat Rev Genet. 2009;10:691–703. doi: 10.1038/nrg2640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kosiol C, et al. Patterns of positive selection in six Mammalian genomes. PLoS Genet. 2008;4:e1000144. doi: 10.1371/journal.pgen.1000144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ashburner M, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Makino CL, et al. Recoverin regulates light-dependent phosphodiesterase activity in retinal rods. J Gen Physiol. 2004;123:729–741. doi: 10.1085/jgp.200308994jgp.200308994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Watterson GA. On the number of segregating sites in genetical models without recombination. Theor Popul Biol. 1975;7:256–276. doi: 10.1016/0040-5809(75)90020-9. 0040-5809(75)90020-9 [pii] [DOI] [PubMed] [Google Scholar]
- 25.Li WH, Sadler LA. Low nucleotide diversity in man. Genetics. 1991;129:513–523. doi: 10.1093/genetics/129.2.513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hobolth A, Christensen OF, Mailund T, Schierup MH. Genomic relationships and speciation times of human, chimpanzee, and gorilla inferred from a coalescent hidden Markov model. PLoS Genet. 2007;3:e7. doi: 10.1371/journal.pgen.0030007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Patterson N, Richter DJ, Gnerre S, Lander ES, Reich D. Genetic evidence for complex speciation of humans and chimpanzees. Nature. 2006;441:1103–1108. doi: 10.1038/nature04789. [DOI] [PubMed] [Google Scholar]
- 28.Alcaide M, et al. Population fragmentation leads to isolation by distance but not genetic impoverishment in the philopatric Lesser Kestrel: a comparison with the widespread and sympatric Eurasian Kestrel. Heredity. 2009;102:190–198. doi: 10.1038/hdy.2008.107. [DOI] [PubMed] [Google Scholar]
- 29.Yu N, et al. Low nucleotide diversity in chimpanzees and bonobos. Genetics. 2003;164:1511–1518. doi: 10.1093/genetics/164.4.1511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Chen FC, Li WH. Genomic divergences between humans and other hominoids and the effective population size of the common ancestor of humans and chimpanzees. Am J Hum Genet. 2001;68:444–456. doi: 10.1086/318206. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


