Abstract
The ability of transposons to mobilize to new places in a genome enables them to introgress rapidly into populations. The piRNA pathway has been characterized recently in the germ-line of the fruit fly, Drosophila melanogaster, and is responsible for down-regulating transposon mobility. Transposons have been used as tools in mosquitoes to genetically transform a number of species including Anopheles stephensi, a vector of human malaria. These mobile genetic elements also have been proposed as tools to drive anti-pathogen effector genes into wild mosquito populations to replace pathogen-susceptible insects with those engineered genetically to be resistant to or unable to transmit a pathogen. The piRNA pathway may affect the performance of such proposed genetic engineering strategies. Here we identify and describe the Anopheles stephensi orthologs of the major genes in the piRNA pathway, Ago3, Aubergine (Aub), and Piwi. Consistent with a role in protection from transposon movement, these three genes are expressed constitutively in the germ line cells of ovaries and induced further following a bloodmeal.
Keywords: Piwi, Aubergine, Aub, Argonaut 3, Ago3, transposon, mobility
Introduction
Transposons are mobile genetic elements that potentially can spread rapidly through populations despite fitness costs incurred by excision and insertion (McClintock 1987). The field of vector genetics has benefited from this mobility; genetic modification of transposons to carry exogenous genes into the genome has enabled the transformation of mosquito species, and it is possible that the same tools could be used to drive an anti-pathogen effector gene into a wild population to replace it with one unable to transmit pathogens (Coates et al. 1998; Catteruccia et al. 2000; Grossman et al. 2001; James 2005; Terenius et al. 2008). An RNA interference pathway, called the piRNA pathway, was characterized recently in the fruit fly, Drosophila melanogaster, and is responsible for inhibiting the movement of transposons (Sarot et al. 2004; Vagin et al. 2006; Saito et al. 2006; Brennecke et al. 2007; Pélisson et al. 2007). This pathway employs the Piwi proteins, a subfamily of the Argonautes, along with sequence-specific small RNAs called Piwi-interacting RNA (piRNAs), to target transposon-derived RNAs for degradation in the germ line tissue. This pathway enables organisms to inhibit transposon movement and therefore to prevent or minimize the genetic lesions these events can cause.
The piRNA pathway was proposed to play a role in transposon regulation in mosquitoes of the genus Aedes based on small RNA and genomic sequencing data (Arensburger et al. 2011; Akbari et al. 2013). However, some notable differences are observed between data from Ae. aegypti and those derived from D. melanogaster. The Ae. aegypti genome has a much higher proportion of transposon sequences (47% in Ae. aegypti compared to 15.8% in D. melanogaster) but a smaller representation of piRNAs targeting transposons (~20% in Ae. aegypti compared with 50% in D. melanogaster) (Kaminker et al. 2002; Nene et al. 2007; Smith et al. 2007; Arensburger et al. 2011; Akbari et al. 2013). Furthermore, there are eight putative Piwi family genes in Ae. aegypti annotated in VectorBase, one with homology to DmAgo3 and seven that show homology to DmPiwi and DmAub. Thus, the piRNA pathway may function differently in mosquitoes and fruit flies. However, data collected from the mosquito genus Anopheles look more similar to corresponding data in Drosophila than Aedes. For example, one-to-one orthologs exist in An. gambiae for each of the three D. melanogaster Piwi genes. Additionally, anopheline mosquitoes have a relatively small genome resulting from lower transposon representation compared to the Ae. aegypti genome (Holt et al. 2002; Kaminker et al. 2002; Hoa et al. 2003; Zhou et al. 2007; Nene et al. 2007; Marinotti et al. 2013).
In addition to its function in regulating transposon mobility, the piRNA pathway also may have roles relevant to vector biology. Mosquitoes are majors vectors of viral infections to humans and animals and molecular aspects of mosquito-virus interactions are major foci of research. Viral-specific piRNAs derived from D. melanogaster, Anopheles and Aedes cell lines challenged with viral infection have been isolated, implicating the piRNA pathway in virus control (Chotkowski et al. 2008; Morazzani et al. 2012; Vodovar et al. 2012; Schnettler et al. 2013; Léger et al. 2013). The piRNA pathway also may regulate gene expression since a proportion of piRNAs sequenced in D. melanogaster and Ae. aegypti are specific to endogenous protein-encoding genes (Brennecke et al. 2007; Arensburger et al. 2011; Akbari et al. 2013).
The biology of the piRNA pathway in mosquitoes is relevant to both applied and basic aspects of disease vector research. Understanding the dynamics of transposon movement in the malaria vector, An. stephensi, and the control of this process is not only an interesting basic question in an organism with a different reproductive strategy than Drosophila species (vector mosquitoes require a blood meal for development of progeny), but also information on transposon control in mosquitoes is essential to proceeding intelligently with the design of gene-drive systems based on transposons. Our observations in An. stephensi support the hypothesis that the piRNA pathway plays a role in transposon control in this species: we find that Piwi, Aub, and Ago3 display expression characteristics appropriate for priming an egg for exposure to paternal transposons: transcripts are detectable in the germ line tissue of adult mosquitoes, become increasingly more abundant in the ovaries with egg development and are found in the embryos.
Results and Discussion
Gene, transcript and putative protein structures of AsAgo3, AsAub and AsPiwi
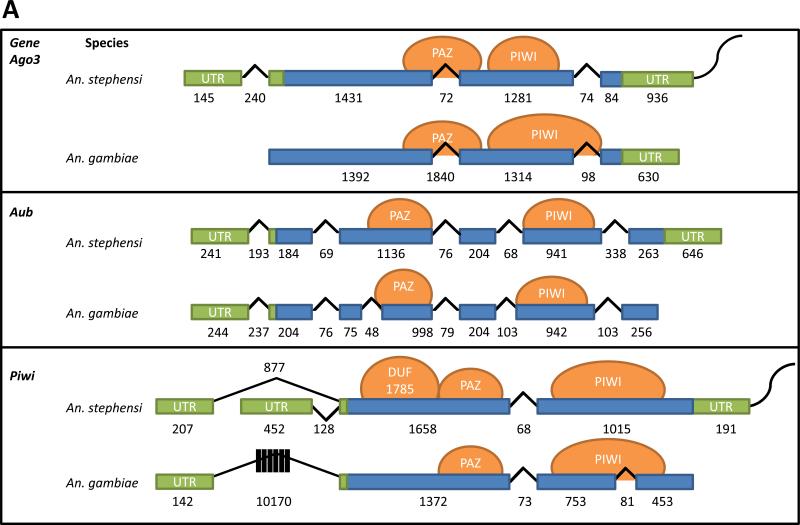
Alignments of the An. gambiae Ago3, Ago4 (Aub) and Ago5 (Piwi) transcript sequences with the An. stephensi genome (Assembly: AsteI2) and transcriptome yielded partial matches that aligned with >78% identity. According to sequence similarity, the genes ASTEI04992, ASTEI03833 and ASTEI06803 were designated AsAgo3, AsAub and AsPiwi, respectively (Figure 1, Table S1). Genomic DNA sequences from the An. stephensi Indian strain (VectorBase.org) were used to design primers for gene amplification studies to determine AsAgo3, AsAub and AsPiwi structures and the complete sequences of their corresponding transcripts. Single transcripts were identified for both AsAgo3 and AsAub, while amplification of the 5’-end of AsPiwi revealed a novel, alternative first exon that aligned to the genome within the first intron (Figure 1A). Both of the AsPiwi transcripts have the same translation start site and are predicted to produce identical proteins. Transcripts of 3895, 3615, and 3071 nucleotides (nt) in length from the start of transcription to the beginning of the poly-adenosine sequences were observed for AsAgo3, and AsPiwi isoforms A and B, respectively. AsAub transcripts extend beyond the first polyadenylation signal in the 3’-end UTR, such that the transcript is larger than 3615 nt. Complete transcripts have been deposited in VectorBase; the transcript sequences reported here supplement previous annotations of these transcripts by providing: 3’- and 5’-end untranslated regions (UTRs) for both AsAgo3 and AsPiwi; an entirely new sixth intron for AsAub and an alternate 5’-end UTR for AsPiwi. Comparisons of the An. stephensi Ago3, Aub and Piwi genes with those of An. gambiae annotated in VectorBase revealed a number of differences: a lack of data supporting a 5’-end UTR for AgAgo3 and 3’-end UTR for AgAub and AgPiwi; AgAub encodes an additional intron compared to AsAub; AsPiwi does not have the large first intron (10,170 nt) found in AgPiwi; and as noted above, AsPiwi encodes an alternate first intron (Figure 1A).
Figure 1. Gene structure comparisons and phylogenetic relationships of the An. stephensi Piwi family genes.
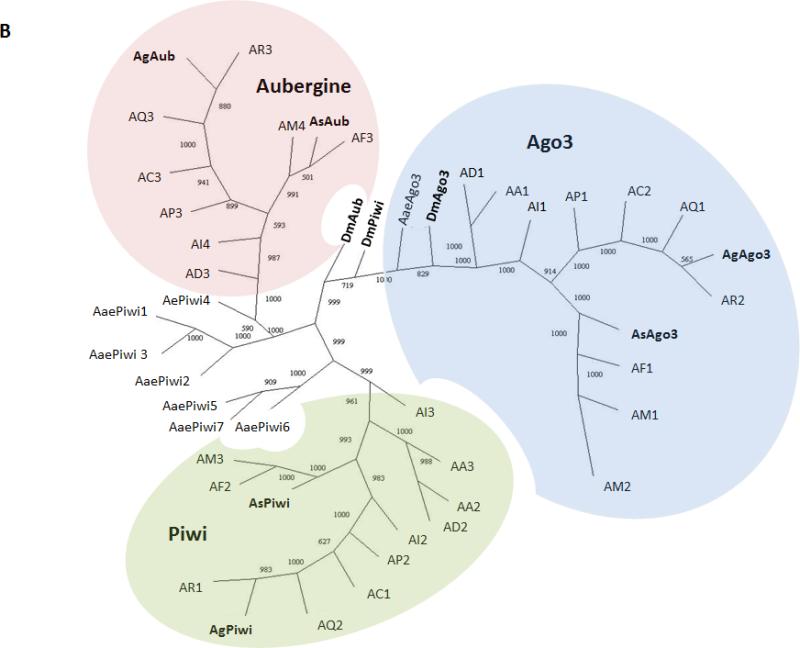
A) Schematic representations of transcription products for An. stephensi Ago3, Aub and Piwi detected in samples of ovaries collected at 48 hr PBM. Exons and introns are represented by boxes and lines, respectively, with the length in nucleotides indicated below each. Untranslated regions are colored green, open-reading frames are blue, predicted protein-binding domains are represented by orange circles and the polyA tail is the curved line at each 3’-end. B) Phylogenetic tree generated from alignment of predicted amino acid sequences for Piwi family proteins in An. stephensi (As), An. gambiae (Ag), Aedes aegypti (Aae), and Drosophila melanogaster (Dm). Bootstrap values between genes are listed between each pair of corresponding nodes. Genes from other mosquito species are represented: AA, Anopheles albimanus; AC, Anopheles christi; AD, Anopheles darlingi; AI, Anopheles dirus; AF, Anopheles funestus; AM, Anopheles minimus; AP, Anopheles epiroticus, AQ, Anopheles quadriannulatus and AR, Anopheles arabiensis. The digits following species name designations are arbitrary and correspond to the gene names listed in Table S1.
The encoded amino acid sequences predicted from all three transcripts using the ExPASy online translate tool (http://web.expasy.org/translate) encode both PAZ and PIWI domains (Höck & Meister 2008) as predicted using and SMART online protein domain prediction tool (http://smart.embl-heidelberg.de/) (Figure 1A). Additionally the AsPiwi predicted proteins have a conserved domain of unknown function (DUF 1785) present in Argonautes and co-occurring with PIWI domains (Su et al. 2009; Kurscheid et al. 2009; Poulsen et al. 2013; Zheng 2013).
Predicted amino acid sequence alignments show a high percent identity between An. stephensi Ago3, Aub and Piwi and putative orthologous proteins in D. melanogaster (48%, 49% and 44% , respectively) and An. gambiae (77%, 89% and 72%, respectively) (Figure S1) supporting the conclusion that these genes represent one-to-one ortholog pairs (Figure 1B). Interestingly, AsAub and AsPiwi were more similar to each other than to either DmAub or DmPiwi, as reported previously for these proteins in An. gambiae (Hoa et al. 2003). Analysis of other available anopheline genomes (VectorBase) identifies one orthologous gene corresponding to each of the three Piwi proteins in all species, with a few exceptions. Three Piwi genes in An. albimanus are predicted to encode two Piwi orthologs and no Aub ortholog. Four Piwi genes were identified for An. minimus, whose genome encodes two putative Ago3 orthologs in addition to Aub and Piwi orthologs, and An. dirus, whose genome encodes two putative Piwi orthologs in addition to Ago3 and Aub orthologs (Figure 1B) . When amino acid sequences from all seven Aedes aegypti Piwi subfamily members were included in the analysis, Aub-like and Piwi-like orthologs segregated as predicted in earlier reports (Figure 1B; Akbari et al. 2013). A higher amino acid sequence diversity at the N-termini of the An. stephensi piRNA components is consistent with findings in D. melanogaster; therefore these sequences were used as the basis for designing corresponding probes for hybridization in situ (Brennecke et al. 2007).
Stage- and tissue-specific transcript abundance
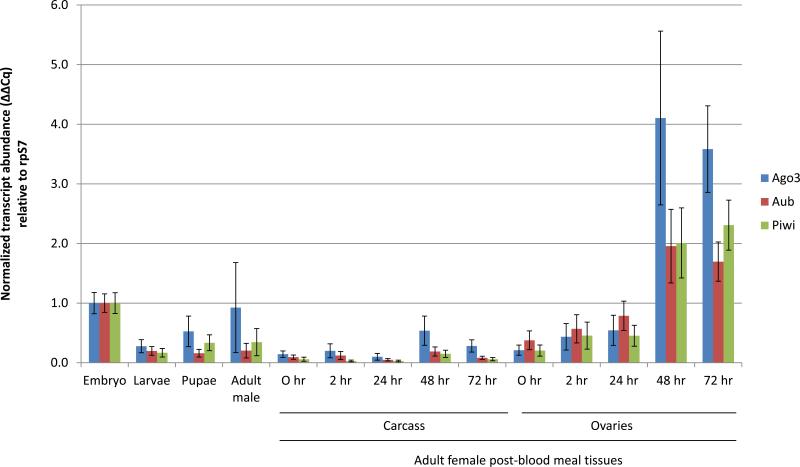
Quantitative real-time PCR was used to detect and measure the accumulation of AsAgo3, AsAub and AsPiwi transcripts at embryonic, larval, pupal and adult developmental stages, and in ovaries and carcasses of adult females (Figure 2). All three piRNA pathway gene transcripts are most abundant at 48-72 hours post blood meal (hr PBM) in the ovaries and also are significantly more abundant in early embryos (0-2 hr) than at any other stage analyzed (Table S2). This expression profile is consistent with microarray data collected for these genes in An. gambiae, although in those experiments, transcript abundance for all three genes increases significantly by 24 hr PBM (p-values=0.00005, 0.0021, 0.006 for AgAgo3, AgAub and AgPiwi, respectively: Marinotti et al. 2005). Transcription of zygotic genes does not occur earlier than 1-2 hours post egg laying in fertilized embryos of D. melanogaster (Zalokar 1976; Pritchard & Schubiger 1996). Assuming a similar regulation of the zygotic genome exists in An. stephensi, transcripts present in early embryos represent those deposited maternally during ovary development. These combined data support the hypothesis that AsAgo3, AsAub, and AsPiwi genes are expressed at the appropriate time and place to repress transposon expression and remobilization.
Figure 2. Abundance profiles of AsAgo3, Aub and Piwi transcripts during development.
Each histogram represents data (average ± SEM) of three biological replicates normalized to the embryo sample. Adult female 0 hr PBM samples were collected before blood-feeding. Embryos were collected between 0 and 2 hours following oviposition. p-values for all comparisons are listed in Table S2.
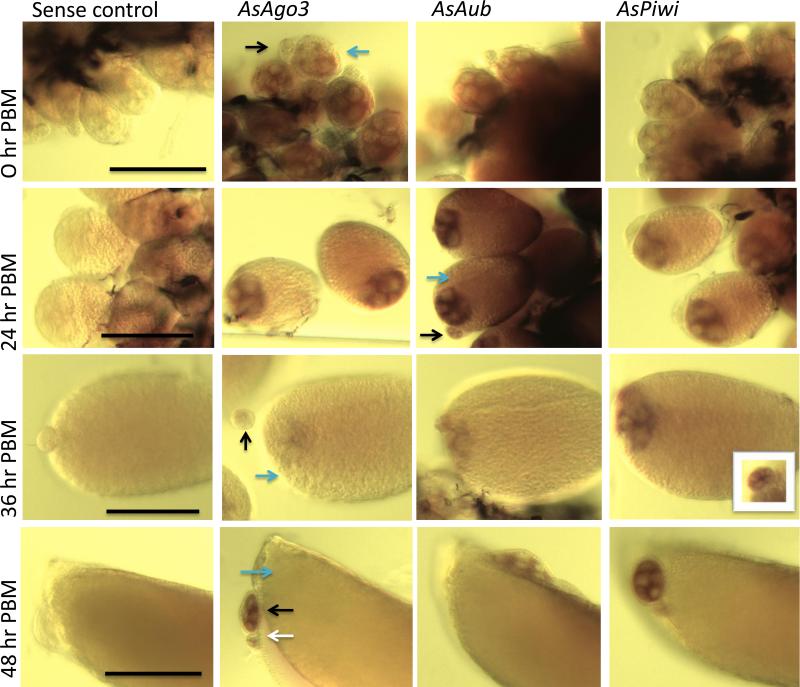
The localization of Piwi gene transcripts was determined at different times during ovary development. AsAgo3, AsAub and AsPiwi antisense RNA probes hybridized to transcripts in the cytoplasm of the nurse cells and oocytes of primary follicles, collectively the germ-line tissue, and in the previtellogenic secondary and tertiary follicles, which represent the earliest visible stages of oocyte development, that do not progress further until subsequent blood meals (Figure 3). Diffuse signals corresponding to the three piRNA pathway gene transcription products are seen in oocytes at 24 hr PBM. These signals are barely distinguishable visibly at 36 hr PBM and undetectable at 48 hr PBM. Since transcript abundance measured by quantitative gene amplification (qPCR) at this stage is 2-4 fold higher than in ovaries from unfed females, we speculate that the transcripts in the oocytes at 48 hr PBM are still present but either too diffuse to detect using hybridization in situ or that the endochorion at this stage is developed enough so that the hybridization and/or detection components of the assay could not penetrate the primary follicle. Furthermore, no transcripts were detected in either experimental or control groups of embryos collected 0-2 hours following oviposition, although this stage also has abundant transcripts based on qPCR analyses. These transcripts likely are those deposited into the oocyte by nurse cells during development. It is also evident from these images that by 48 hrs PBM, the secondary follicles represent an important contributor to the transcript quantities measured in qPCR, indicating that new transcript present at this time point is not likely in the primary follicle. It is reasonable, based on the data presented here, to hypothesize that AsAgo3, AsAub and AsPiwi are expressed early in the primary, secondary and tertiary follicles following a blood meal and that their transcripts are accumulated and present in the oocytes throughout development. Strong, nonspecific background staining was seen at 48 hr PBM superficially around the egg floats and at all stages in the trachea in both control and experimental groups.
Figure 3. Spatial localization of AsAgo3, Aub and Piwi transcripts in ovaries.
Whole-mount hybridization in situ of An. stephensi ovaries in sugar-fed (0 hours post-blood meal [hr PBM]) and at 24, 36 and 48 hr PBM. Blue, black and white arrows indicate the primary, secondary and tertiary follicles, respectively. Scale bar depicts 100 micrometers for each temporal group.
In summary, these combined data support the conclusions that piRNA pathway gene transcripts in An. stephensi are abundant and enriched in the germline tissue during follicle development and are present in newly-laid embryos. These expression properties are consistent with a role in germ-line protection from transposon mobilization. Future experiments will focus on addressing whether transposon repression is indeed a role of the piRNA pathway in this important vector mosquito species.
Experimental Procedures
An Anopheles stephensi strain maintained in our laboratory since 2004 was founded with mosquitoes provided by Dr. Marcelo Jacobs-Lorena (Johns Hopkins University). This line was used for all experiments reported here. Mosquitoes were maintained at 27°C and 77% humidity with 12:12 daily light-dark cycles and 30 minute dusk and dawn transitions. Larvae were fed a diet of powdered fish food (Tetramin) mixed with yeast. Adults were provided 10% sucrose ad libitum. Anesthetized mice were used for blood feeding adult females.
Samples for qPCR analyses were prepared from whole mosquitoes, dissected ovaries or carcasses (all tissues excluding the ovaries). RNA was extracted from 50 individuals for each sample (except embryos, where ~300 were used) by homogenization in Trizol reagent (Invitrogen) followed by chloroform extraction and RNA purification using the Zymo Clean and Concentrator 25 (Zymo Research). Samples were treated with DNase RQ1 (Promega) and tested for genomic DNA contamination. cDNA was synthesized using the iScript kit (Biorad) and used directly for qPCR reactions with Kapa Sybrfast supermix (Kapa Biosystems). All primers were optimized for annealing temperatures and cDNA concentrations; at least three technical and biological replicates each were used for each data point. Primers and corresponding amplification efficiencies are listed in Table S3. Biorad software (Version 3.0) was used for statistical analysis with the default two-sided t-test with a p-value threshold of 0.025 set for significance.
Rapid Amplification of cDNA Ends (RACE) was performed with the SMARTer RACE kit (Clontech). cDNA was synthesized using the Clontech reagents and RNA collected from ovaries dissected 48 hr PBM. Gene amplification reactions were performed using Phusion High Fidelity Master Mix from New England Biosystems. Nested RACE reactions were performed with touchdown-PCR cycles on separate preparations of 5’- and 3’-end cDNA templates as described in the Clontech protocol. RACE and nested-RACE products were run on agarose gels and selected amplicons were cloned into the pCR-Blunt II TOPO plasmid. Plasmids were transformed into and amplified in chemically-competent TOP10 E. coli, and sequenced using M13 forward and reverse primers. Consensus mature transcript sequences have been deposited in Genbank with accession codes KJ808821 (PiwiA), KJ808822 (PiwiB), KJ808823 (Ago3) and KJ808824 (Aubergine).
Predicted amino acid sequences of Ago3, Aubergine and Piwi were downloaded from Vectorbase.com for all the Anopheles species available and for Aedes aegypti. Orthologous sequences of Drosophila melanogaster were obtained from NCBI (http://www.ncbi.nlm.nih.gov). The resulting alignment was fed to MrBayes to simultaneously test for 10 models of amino acids evolution (Ronquist et al. 2012). Phylogeny analyses were performed with RAxML through the Cipres Gateway Portal imposing the WAG mutation model and 1000 bootstrap resamplings of the original datasets. CONSENSE (PHYLIP version 3.5c, Felsenstein, 1993) was used to generate an unrooted consensus tree that was visualized by Treeview (http://taxonomy.zoology.gla.ac.uk/rod/treeview.html).
Hybridization in situ of ovaries was performed according to the protocol described by Juhn and James (2012). Briefly, ovaries were dissected from 5-10 mosquitoes and fixed in a 4% formaldehyde solution for 1 hr. Samples were treated with proteinase K, post-fixed and hybridized using digoxigenin (DIG)-labeled antisense and sense-RNA probes generated with the DIG RNA-labeling kit (Roche). The 5’ RACE product clones were used as a template for amplification with M13 forward and reverse primers and each PCR product was used as a substrate for the RNA-labeling reactions. Following overnight hybridization with the labeled probe, samples were treated with RNase A to remove unbound probe and incubated with anti-DIG-alkaline phosphatase (AP)-conjugated antibody (Roche) overnight at 4°C. Colorimetric detection of probe localization was performed by incubation with NBT/BCIP (Roche) as a substrate for 5-7 hours in the dark. Samples were incubated overnight in glycerol, mounted and visualized using bright-field microscopy.
Supplementary Material
Acknowledgments
We thank Aniko Fazekas for assistance with mosquito rearing. This research was supported by a grant from the NIH NIAID (AI29746).
References
- Akbari OS, et al. The Developmental Transcriptome of the Mosquito Aedes aegypti, an Invasive Species and Major Arbovirus Vector. G3 (Bethesda, Md.) 2013;3(9):1493–509. doi: 10.1534/g3.113.006742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arensburger P, et al. The mosquito Aedes aegypti has a large genome size and high transposable element load but contains a low proportion of transposon-specific piRNAs. BMC genomics. 2011;12:606. doi: 10.1186/1471-2164-12-606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brennecke J, et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell. 2007;128(6):1089–103. doi: 10.1016/j.cell.2007.01.043. [DOI] [PubMed] [Google Scholar]
- Catteruccia F, et al. Stable germline transformation of the malaria mosquito Anopheles stephensi. Nature. 2000;405(6789):959–62. doi: 10.1038/35016096. [DOI] [PubMed] [Google Scholar]
- Chotkowski HL, et al. West Nile virus infection of Drosophila melanogaster induces a protective RNAi response. Virology. 2008;377(1):197–206. doi: 10.1016/j.virol.2008.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coates CJ, et al. Mariner transposition and transformation of the yellow fever mosquito, Aedes aegypti. Proceedings of the National Academy of Sciences of the United States of America. 1998;95(7):3748–51. doi: 10.1073/pnas.95.7.3748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goujon M, et al. A new bioinformatics analysis tools framework at EMBL-EBI. Nucleic acids research. 2010;38(Web Server issue):W695–9. doi: 10.1093/nar/gkq313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grossman GL, et al. Germline transformation of the malaria vector, Anopheles gambiae, with the piggyBac transposable element. Insect molecular biology. 2001;10(6):597–604. doi: 10.1046/j.0962-1075.2001.00299.x. [DOI] [PubMed] [Google Scholar]
- Hoa N, et al. Characterization of RNA interference in an Anopheles gambiae cell line. Insect Biochemistry and Molecular Biology. 2003;33(9):949–957. doi: 10.1016/s0965-1748(03)00101-2. [DOI] [PubMed] [Google Scholar]
- Höck J, Meister G. The Argonaute protein family. Genome biology. 2008;9(2):210. doi: 10.1186/gb-2008-9-2-210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holt RA, et al. The genome sequence of the malaria mosquito Anopheles gambiae. Science (New York, N.Y.) 2002;298(5591):129–49. doi: 10.1126/science.1076181. [DOI] [PubMed] [Google Scholar]
- James AA. Gene drive systems in mosquitoes: rules of the road. Trends Parasitol. 2005;21(2):64–67. doi: 10.1016/j.pt.2004.11.004. [DOI] [PubMed] [Google Scholar]
- Kaminker J, et al. The transposable elements of the Drosophila melanogaster euchromatin: a genomics perspective. Genome Biology. 2002;3(12) doi: 10.1186/gb-2002-3-12-research0084. research0084.1–0084.20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurscheid S, et al. Evidence of a tick RNAi pathway by comparative genomics and reverse genetics screen of targets with known loss-of-function phenotypes in Drosophila. BMC molecular biology. 2009;10(1):26. doi: 10.1186/1471-2199-10-26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Léger P, et al. Dicer-2- and Piwi-mediated RNA interference in Rift Valley fever virus-infected mosquito cells. Journal of virology. 2013;87(3):1631–48. doi: 10.1128/JVI.02795-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marinotti O, et al. Microarray analysis of genes showing variable expression following a blood meal in Anopheles gambiae. Insect molecular biology. 2005;14(4):365–73. doi: 10.1111/j.1365-2583.2005.00567.x. [DOI] [PubMed] [Google Scholar]
- Marinotti O, et al. The genome of Anopheles darlingi, the main neotropical malaria vector. Nucleic acids research. 2013;41(15):7387–400. doi: 10.1093/nar/gkt484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClintock B. The Discovery and Characterization of Tranposable Elements: The Collected Papers of Barbara McClintock. Garland Publishing; New York: 1987. [Google Scholar]
- Morazzani EM, et al. Production of virus-derived ping-pong-dependent piRNA-like small RNAs in the mosquito soma. PLoS pathogens. 2012;8(1):e1002470. doi: 10.1371/journal.ppat.1002470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nene V, et al. Genome sequence of Aedes aegypti, a major arbovirus vector. Science (New York, N.Y.) 2007;316(5832):1718–23. doi: 10.1126/science.1138878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pélisson A, et al. A novel repeat-associated small interfering RNA-mediated silencing pathway downregulates complementary sense gypsy transcripts in somatic cells of the Drosophila ovary. Journal of virology. 2007;81(4):1951–60. doi: 10.1128/JVI.01980-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poulsen C, Vaucheret H, Brodersen P. Lessons on RNA silencing mechanisms in plants from eukaryotic argonaute structures. The Plant cell. 2013;25(1):22–37. doi: 10.1105/tpc.112.105643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pritchard DK, Schubiger G. Activation of transcription in Drosophila embryos is a gradual process mediated by the nucleocytoplasmic ratio. Genes & development. 1996;10(9):1131–42. doi: 10.1101/gad.10.9.1131. [DOI] [PubMed] [Google Scholar]
- Ronquist F, et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematic biology. 2012;61(3):539–42. doi: 10.1093/sysbio/sys029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saito K, et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes & development. 2006;20(16):2214–22. doi: 10.1101/gad.1454806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarot E, et al. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics. 2004;166(3):1313–21. doi: 10.1534/genetics.166.3.1313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnettler E, et al. Knockdown of piRNA pathway proteins results in enhanced Semliki Forest virus production in mosquito cells. The Journal of general virology. 2013;94(Pt 7):1680–9. doi: 10.1099/vir.0.053850-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sievers F, et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular systems biology. 2011;7:539. doi: 10.1038/msb.2011.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith CD, et al. The Release 5.1 annotation of Drosophila melanogaster heterochromatin. Science (New York, N.Y.) 2007;316(5831):1586–91. doi: 10.1126/science.1139815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Su J, et al. Isolation and characterization of Argonaute 2: A key gene of the RNA interference pathway in the rare minnow, Gobiocypris rarus. Fish & Shellfish Immunology. 2009;26(1):164–170. doi: 10.1016/j.fsi.2008.10.002. [DOI] [PubMed] [Google Scholar]
- Terenius O, et al. Molecular genetic manipulation of vector mosquitoes. Cell Host Microbe. 2008;4(5):417–423. doi: 10.1016/j.chom.2008.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vagin V, et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science (New York, N.Y.) 2006;313(5785):320–4. doi: 10.1126/science.1129333. [DOI] [PubMed] [Google Scholar]
- Vodovar N, et al. Arbovirus-derived piRNAs exhibit a ping-pong signature in mosquito cells. PloS one. 2012;7(1):e30861. doi: 10.1371/journal.pone.0030861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zalokar M. Autoradiographic study of protein and RNA formation during early development of Drosophila eggs. Developmental Biology. 1976;49(2):425–437. doi: 10.1016/0012-1606(76)90185-8. [DOI] [PubMed] [Google Scholar]
- Zheng Y. Phylogenetic analysis of the Argonaute protein family in platyhelminths. Mol Phylogenet Evol. 2013;66(3):1050–4. doi: 10.1016/j.ympev.2012.11.014. [DOI] [PubMed] [Google Scholar]
- Zhou X, et al. Identification and characterization of Piwi subfamily in insects. Biochemical and biophysical research communications. 2007;362(1):126–31. doi: 10.1016/j.bbrc.2007.07.179. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.