Abstract
The molecular mechanisms that underlie the development of squamous cell skin cancers (SSC) are poorly understood. We have used oligonucleotide microarrays to compare the differences in cellular gene expression between a series of keratinocyte cell that mimic disease progression with the aim of identifying genes that may potentially contribute towards squamous cell carcinoma (SCC) progression in vivo, and in particular to identify markers that may serve as potential therapeutic targets for SCC treatment. Gene expression differences were corroborated by polymerase chain reaction and Western blotting. We identified Axl, a receptor tyrosine kinase with transforming potential that has also been shown to have a role in cell survival, adhesion and chemotaxis, was upregulated in vitro in SCC-derived cells compared to premalignant cells. Extending the investigation to tumour biopsies showed that the Axl protein was overexpressed in vivo in a series of SCCs.
Keywords: Axl, skin cancer, receptor tyrosine kinase
Non-melanoma skin cancer (NMSC) is the most prevalent human cancer, with about 1 million cases in the USA and >60 000 cases occurring annually in the UK. Basal cell carcinoma (BCC) is the most common NMSC, whereas squamous cell carcinoma (SCC) accounts for 20% of all cutaneous malignancies. Squamous cell carcinomas typically arise on sun-exposed body sites, and the incidence in Europe, North America and Australia ranges from 50 to 170 cases per 100 000 individuals (Holme et al, 2000; Diepgen and Mahler, 2002). Cutaneous SCCs are usually treated by excision; however, they do have the potential to recur locally and, in some cases, to metastasize. Currently, the most common prognostic indicators are histologic subtype and tumour size.
In contrast to BCC development, where mutations in the patched signalling pathway are associated with BCC formation (Johnson et al, 1996), the current knowledge on factors that are involved in the de novo development and progression of SCC is relatively sparse.
To identify genes that are differentially expressed in SCC compared to normal skin, we have focused in particular on genes encoding cell surface receptors, as overexpression of the encoded protein may serve either as a useful biomarker or a potential target for therapeutic intervention. We have made use of a unique series of cutaneous SCC cell lines derived from an immunosuppressed patient representing different stages of malignant transformation (Proby et al, 2000), that serve as a model in which to study the changes in gene expression that occur in the transition from premalignant cells to SCC and finally to a metastatic tumour. They were established from forehead skin (PM1), a primary SCC (MET1) and an associated lymph node metastasis (MET4). Affymetrix GeneChip arrays were used to compare the expression profile of PM1 with two malignant cell lines (MET1 and MET4). Most interestingly, the expression analysis revealed upregulation of Axl, a transmembrane receptor tyrosine kinase with transforming potential (O'Bryan et al, 1991) and is an intriguing candidate for involvement in SCC development and progression. Further immunohistochemical analysis of Axl protein expression in a panel of SCCs revealed that Axl was frequently overexpressed in SCCs compared to BCC or normal skin.
MATERIALS AND METHODS
Cell culture
Early passage (less than p16) PM1, MET1 and MET4 keratinocyte cell lines were grown with an irradiated fibroblast feeder layer in Dulbecco's modified Eagle's medium plus HAMS F12 medium containing 10% foetal calf serum essentially as described (Rheinwald and Green, 1975).
Microarray experiments and semi-quantitative reverse transcription–polymerase chain reaction
Briefly, total RNA was extracted from 70–80% confluent PM1, MET1 and MET4 cell lines using TRIZOL Reagent (Invitrogen-15596, Paisley, UK) and was used to generate labelled probes as per manufacturer's instructions. These were used to hybridise the U133A Gene Chips using an Affymetrix Fluidics station 400. Three biological replicates were performed for each of the three cell lines. Full details of the methodology and statistical analysis can be found in Supplementary Material.
Quantitative reverse transcription–polymerase chain reaction (RT—PCR) was performed to validate data from GeneChip experiments and was performed using an OPTICON™ 2 Continuous Fluorescence Detection System (Bio-Rad) on the three replicate samples used in the GeneChip analysis. Full details of the Q–PCR conditions are given in Supplementary Material.
The primer pairs used for amplification of the selected targets were:
| Axl: | GAGAACATTAGTGCTACGCGGAA/ |
| CCTTAGCCCTATGTCCATTAGCA | |
| G6PDH: | GTTCCGTGAGGACCAGATCTAC/ |
| GGCTCCTTGAAGGTGAGGATAA |
Antibodies and immunohistochemistry
Antibodies used in this study were anti-Axl (C-20), anti-Gas 6 (N-20), (Santa Cruz Biotechnology Inc., Palo Alto, CA, USA), anti-α tubulin (Ab-1, Oncogene Science, Cambridge, MA, USA).
Archival paraffin blocks were used for immunohistochemistry; ethical approval for this study was obtained from the East London and City Health Authority Research Ethics Committee.
Axl expression was examined using standard immunohistochemical techniques using 4 μm thick sections that had been deparaffinized, then blocked with rabbit serum and incubated with a goat polyclonal anti-Axl antibody. The sections were then processed as described (Jackson et al, 2002). Quantitative analysis of Axl staining was performed using KS400 version 3.0 imaging software (Carl Zeiss Ltd, Welwyn Garden City, UK). The percentage of cells expressing Axl was calculated in four representative high-power fields. Statistical analysis was performed using Arcus Quickstat (Statsdirect, Sale, UK, Biomedical version 1.1) and JMP©, SAS Inc. (Karey, NC, USA). For Western blot analysis was performed as described (Jackson et al, 2002), signals were detected using ECL + (Amersham Biosciences UK Ltd, Bucks, UK).
RESULTS
Gene expression profiles in premalignant and malignant keratinocyte lines
We compared the relative gene expression levels of the MET1 and MET4 lines with PM1, and also compared expression in MET4 with the solid tumour from which it appears to have originated, MET1 (Popp et al, 2000). The profiling revealed that 276 genes were statistically differentially expressed in PM1 cells compared to MET1 and MET4 cells (P = 0.0001). For practical reasons, we have applied an arbitrary filter level of five-fold changes in the ratio of expression levels; this relatively high cutoff point was used with a view to focus on the genes that are most grossly affected. As a result, an overall comparison of transcript levels from PM1 vs MET1, PM1 vs MET4 and MET1 vs MET4 revealed that 82 genes were significantly differentially expressed with a greater than fivefold change across the three tumour-derived cell lines that fell into diverse functional categories potentially affecting extracellular and intracellular signalling, proliferation and adhesion (Table 1). In particular, we noted that the axl tyrosine kinase receptor was significantly overexpressed in the MET1 relative to PM1 cells, and was also overexpressed 4.3-fold in Met4 relative to PM1 cells (Table 1).
Table 1.
Gene expression profile using Affymetrix arrays of genes differentially expressed in MET1 and MET4 vs PM1cell line and MET1 vs MET4.
| MET1 vs PM1 | MET4 vs PM1 | MET1 vs MET4 | |||
|---|---|---|---|---|---|
| Genbank | Common | Function | Fold +/− | Fold +/− | Fold +/− |
| Ubiquitin cycle/proteolysis and peptidolysis | |||||
| AA020826 | CTSB | Cathepsin B | −5 | ||
| Transport | |||||
| N74607 | AQP3 | Aquaporin 3 | −10 | −6 | |
| AF201942 | NXT2 | Nuclear transport factor 2-like export factor 2 | 8 | ||
| Transcription/replication | |||||
| NM_004364 | CEBPA | CCAAT/enhancer binding protein (C/EBP), alpha | −10 | ||
| H94842 | HOXA11 | Homeo box A11 | 6 | 8 | |
| BC005342 | NSBP1 | Nucleosomal-binding protein 1 | 16 | ||
| NM_016594 | FKBP11 | FK506-binding protein 11, 19 kDa | 5 | ||
| AB046692 | AOX1 | Aldehyde oxidase 1 | 8 | ||
| Signalling | |||||
| NM_006176 | NRGN | Neurogranin (protein kinase C substrate, RC3) | 11 | ||
| NM_001945 | DTR | Diphtheria toxin receptor) | 16 | 22 | |
| AA464753 | PDAP1 | Homo sapiens, clone IMAGE:3457786 | −8 | ||
| M14333 | FYN | FYN oncogene related to SRC, FGR, YES | 5 | ||
| NM_004335 | BST2 | Bone marrow stromal cell antigen 2 | 16 | ||
| NM_013447 | EMR2 | egf-like, hormone receptor-like sequence 2 | −6 | ||
| NM_003020 | SGNE1 | Secretory granule, neuroendocrine protein 1 | |||
| (7B2 protein) | 5 | ||||
| AI439556 | TXNIP | Thioredoxin interacting protein | −5 | ||
| Development | |||||
| NM_000366 | TPM1 | Tropomyosin 1 (alpha) | 6 | ||
| NM_003412 | ZIC1 | Zic family member 1 (odd-paired homolog, Drosophila) | 33 | 39 | |
| U61276 | JAG1 | Jagged 1 (Alagille syndrome) | 5 | ||
| NM_014556 | EVC | Ellis van Creveld syndrome | 10 | ||
| Growth factor/chemokine/cytokine/inflammation/immune response | |||||
| NM_002658 | PLAU | Plasminogen activator, urokinase | 6 | ||
| AL573058 | C1R | Complement component 1, r subcomponent | 13 | 10 | |
| M92934 | CTGF | Connective tissue growth factor | 13 | ||
| NM_003596 | TPST1 | Tyrosylprotein sulfotransferase 1 | 8 | ||
| NM_013314 | BLNK | B-cell linker | 6 | ||
| NM_002964 | S100A8 | S100 calcium-binding protein A8 (calgranulin A) | 5 | ||
| NM_002852 | PTX3 | Pentaxin-related gene, rapidly induced by IL-1 beta | 8 | ||
| M69176 | CEACAM1 | Carcinoembryonic antigen-related cell adhesion molecule 1 | 6 | ||
| NM_006379 | SEMA3C | Semaphorin 3C | 10 | ||
| NM_002053 | GBP1 | Guanylate-binding protein 1, interferon-inducible, 67 kDa | 7 | ||
| NM_014391 | ANKRD1 | Cardiac ankyrin repeat protein | −8 | ||
| NM_000963 | PTGS2 | Prostaglandin-endoperoxide synthase 2 | 5 | 16 | |
| L27624 | TFPI2 | Tissue factor pathway inhibitor 2 | 9 | ||
| BG166705 | CXCL5 | Chemokine (C-X-C motif) ligand 5 | 227 | ||
| Cell cycle/oncogene/tumour suppressor | |||||
| NM_001759 | CCND2 | Cyclin D2 | −7 | 8 | |
| M15329 | IL1A | Interleukin 1, alpha | 5 | ||
| NM_005504 | BCAT1 | Branched chain aminotransferase 1, cytosolic | 11 | ||
| NM_000393 | COL5A2 | Collagen, type V, alpha 2 | 44 | 23 | |
| NM_000584 | IL8 | Interleukin 8 | 112 | 32 | |
| Metabolism/stress response | |||||
| NM_005965 | MYLK | Myosin, light polypeptide kinase | −7 | 7 | |
| NM_016816 | OAS1 | 2′,5′-oligoadenylate synthetase 1, 40/46 kDa | 5 | ||
| AW614435 | UROD | Uroporphyrinogen decarboxylase | 16 | ||
| AV704962 | SC4MOL | Sterol-C4-methyl oxidase-like | 7 | ||
| NM_004265 | FADS2 | Fatty acid desaturase 2 | 4 | 5 | |
| NM_018192 | LEPREL1 | Leprecan-like 1 | 10 | 15 | |
| NM_007034 | DNAJB4 | DnaJ (Hsp40) homolog, subfamily B, member 4 | 5 | ||
| AF054841 | TM4SF7 | Transmembrane 4 superfamily member 7 | −5 | −6 | |
| NM_000050 | ASS | Argininosuccinate synthetase | −5 | ||
| Cytoskeleton/cell adhesion | |||||
| NM_002628 | PFN2 | Profilin 2 | 25 | 15 | |
| U76549 | KRT8 | Keratin 8 | −6 | ||
| AI831452 | KRT6B | Keratin 6B | 5 | ||
| NM_004791 | ITGBL1 | Integrin, beta-like 1 (with EGF-like repeat domains) | 7 | 15 | |
| AI806793 | COL8A2 | Collagen, type VIII, alpha 2 | 8 | ||
| AI922599 | VIM | Vimentin | 5 | ||
| NM_000422 | KRT17 | Keratin 17 | −5 | ||
| Apoptosis | |||||
| NM_006290 | TNFAIP3 | Tumor necrosis factor, alpha-induced protein 3 | 7 | ||
| BG260394 | SNCA | Synuclein, alpha | 8 | 5 | |
| Receptor | |||||
| NM_021913 | AXL | AXL receptor tyrosine kinase | 12 | ||
| NM_000142 | FGFR3 | Fibroblast growth factor receptor 3 | −35 | ||
| Others | |||||
| NM_000104 | CYP1B1 | Cytochrome P450, subfamily I | −7 | ||
| NM_003118 | SPARC | Secreted protein, acidic, cysteine-rich (osteonectin) | −7 | ||
| NM_007150 | ZNF185 | Zinc finger protein 185 (LIM domain) | 5 | ||
| BF213279 | FARP1 | FERM, RhoGEF (ARHGEF) pleckstrin domain protein 1 | −8 | ||
| NM_014903 | NAV3 | Neuron navigator 3 | 21 | 18 | |
| M58026 | CALML3 | Calmodulin-like 3 | −11 | ||
| BF217861 | MT1E | Metallothionein 1E | 15 | 23 | |
| NM_020672 | S100A14 | S100 calcium-binding protein A14 | −7 | −12 | |
| NM_024593 | FLJ11767 | Hypothetical protein FLJ11767 | −6 | ||
| NM_000396 | CTSK | Cathepsin K (pycnodysostosis) | 10 | 7 | |
| NM_016081 | KIAA0992 | Palladin | 6 | ||
| NM_005785 | RNF41 | Hypothetical SBBI03 protein | 6 | ||
| NM_004900 | APOBEC3B | Apolipoprotein B | 8 | 7 | |
| AF088867 | AGR2 | Anterior gradient 2 homolog (Xenepus laevis) | 32 | 88 | |
| AF010316 | PTGES | Prostaglandin E synthase | −7 | −6 | |
| NM_002281 | KRTHB1 | Keratin, hair, basic, 1 | −17 | ||
| U10691 | MAGEA3 | Melanoma antigen, family A, 6 | −11 | ||
| AU145365 | — | Clone HEMBA1004629, mRNA sequence | 7 | ||
| AK025430 | — | Hypothetical protein FLJ21777 | −5 | ||
| NM_024554 | PGBD5 | Piggybac transposable element derived 5 | −31 | ||
| NM_018436 | ALLC | Allantoicase | 5 | ||
| NM_025143 | C21orf96 | Hypothetical protein FLJ20856 | 5 | ||
| W72694 | LOC51063 | Hypothetical protein LOC51063 | −8 | ||
Only genes whose expression changed by five-fold or more and P-value of 0.001 or less are listed.
Axl mRNA and protein expression in PM1, MET1 and MET4 cell lines
Quantitative RT–PCR was performed on axl transcripts to support the findings of the expression profiling. The analysis was carried out on the RNA prepared for the three biological replicates used in the Affymetrix analysis. The results shown in Figure 1A support the data from the chip analysis. Western blotting of cell lysates showed that Axl protein was also overexpressed in the MET1 and MET4 lines relative to the PM1 line (Figure 1B).
Figure 1.
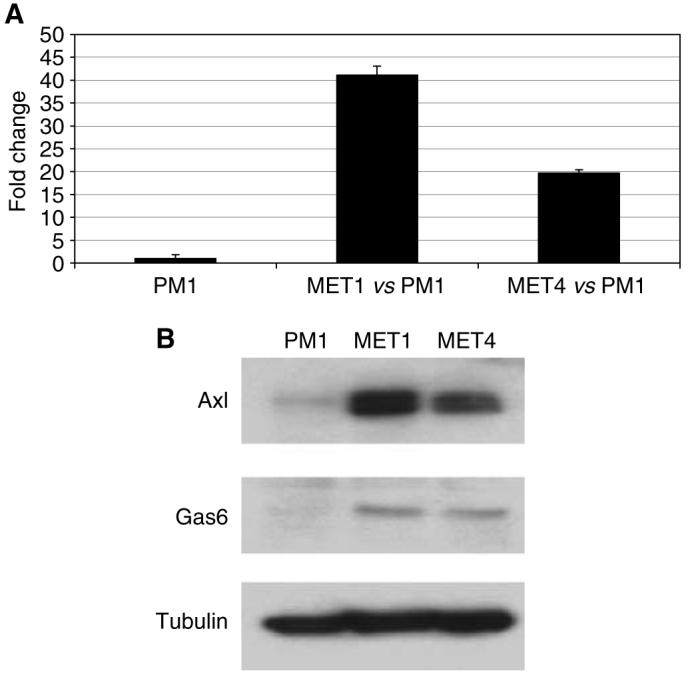
(A) Quantitative RT–PCR of axl gene expression in PM1, MET1 and MET4 cells. (B) Expression of Axl and Gas6, in PM1, MET1 and MET4 cells. Protein extracts were prepared from the different cell lines, separated by SDS–PAGE and Western blotted using specific monoclonal antibodies as described in Materials and Methods. Equal loading of proteins was verified by Western blotting of tubulin.
Immunohistochemical analysis of Axl expression in SCCs
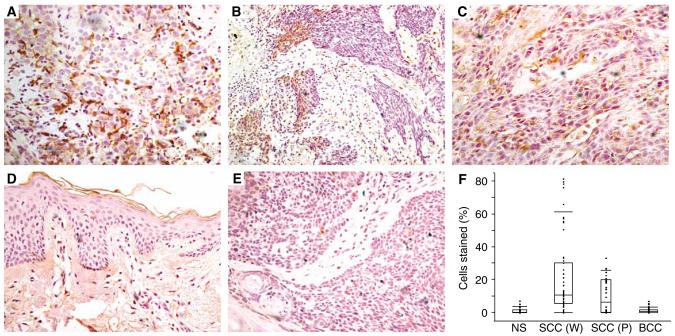
To evaluate the expression of Axl in tumours, we performed an immunohistochemical study on a panel of SCCs, BCCs and normal skin biopsies using anti-Axl-specific antibodies. Axl expression was examined in 17 SCCs (11 well-differentiated and six poorly differentiated) from 16 individuals (Figure 2). Axl expression in 10 BCCs and nine normal skin samples was also investigated. Mast cells that showed consistent, strong, cytoplasmic staining were used in all sections as a positive internal control (data not shown). Goat IgG, at the same concentration as the anti-Axl goat IgG, served as a negative control. Normal epidermis had almost no staining (see Figure 2D) with a mean of 1.3% (95% confidence interval (CI): 0.3 – 2.3) of epidermal cells staining in each section examined. The mean percentage of cells staining with Axl in BCC was 1.3% (95% CI: 0.5 – 2.1%), suggesting that Axl does not have a significant role in cell signalling in BCC (see Figure 2E).
Figure 2.
Immunohistochemistry with anti-Axl antibody demonstrates that Axl expression is increased in SCC. (A) Membranous and cytoplasmic staining in well-differentiated SCC. (B) Heterogeneity of Axl staining in well-differentiated SCC. (C) Axl expression in poorly differentiated SCC. (D) Axl expression in normal skin. (E) Axl expression in BCC. (F) Percentage of cells staining with Axl was counted in four high-power fields in each tumour section. The box and whisker plots represent 5th, 25th, 50th, 75th and 95th centiles.
In contrast to normal skin and BCC, 13 out of 17 SCCs (76%) had significant Axl expression. The mean percentage of well-differentiated SCC (SCCW) cells staining with Axl was 21.5 (95% CI: 5.2 – 37.8%). In general, SCC tumour cells exhibited cytoplasmic staining, although there were a few SCC sections where membranous staining of individual cells was detectable (see Figure 2A). Furthermore, one section showed clear heterogeneity in staining within the SCCW (Figure 2B). The poorly differentiated SCC (SCCP) (Figure 2C) group displayed less Axl staining than SCCW, with a mean percentage of cells staining of 10.7% (95% CI: 1.2 – 22.6%). Statistical analysis was performed using Dunnett's Method to compare Axl staining in normal skin and tumours. There was a statistically significant difference between well- and poorly differentiated SCC compared to normal skin (P<0.01). There was no significant difference between BCC and normal skin staining with Axl, suggesting that Axl signaling may be restricted to SCCs.
DISCUSSION
The deregulation of cellular signalling networks underpins much of the basic framework of carcinogenic processes. A striking feature of the expression analysis was the finding that the axl gene was greatly upregulated in the MET1 cells compared to PM1. Overexpression of both the mRNA and protein was confirmed in MET1 cells in subsequent experiments. Our results are supported by previous studies in murine SCC where increased axl expression was also noted (Loercher et al, 2004). For this analysis, we have exclusively focused on genes whose expression was altered more than five-fold. We cannot rule out at this stage that genes whose expression was not changed by more than five-fold between the different cell lines used in this study may play an important role in determining not only the phenotype of these cells, but also in tumour progression in vivo.
We then extended the observations on the cell lines to investigate Axl protein expression in a pilot study of SCC biopsies. This analysis showed that Axl expression was increased in a significant proportion of the tumours analysed relative to normal skin. Although this is a small series of tumours, it not only validates the approach of using the PM and MET cell lines as a model system, but also suggests that Axl may be a novel marker whose overexpression is frequently associated with SCC. Whether Axl is also overexpressed in other precursor lesions such as actinic keratoses remains to be investigated. Axl expression was not observed in BCC biopsies, suggesting that Axl is not involved in altering signal transduction pathways in these tumours. Axl overexpression has also been noted previously in a variety of other cancers including ovarian (Sun et al, 2004), ocular melanoma (van Ginkel et al, 2004), osteosacroma (Nakano et al, 2003) and renal (Chung et al, 2003) tumours. Axl has been shown recently to play an important role in cell migration and proliferation of human endothelial cells (Holland et al, 2005). Receptors such as Axl that modulate a number of cellular processes such as growth, adhesion and migration and that are overexpressed on cancer cells, makes them targets for the development of novel therapeutics. Multiple clinical trials have employed novel strategies including antibodies that are antagonistic to such receptors (Finn and Slamon, 2003; Emens and Davidson, 2004; Mineo et al, 2004), or alternatively, low molecular weight inhibitors of the kinase activity have also been used, including imatinib mesylate (Gleevec) (Druker, 2004; Pulsipher, 2004; Ross and Hughes, 2004) and gefitinib (Iressa) (Wakeling et al, 2002; Blackledge, 2003; Blackledge and Averbuch, 2004; Park et al, 2004; Tanno et al, 2004). Combination therapy, using both monoclonal antibody together with drug treatment, has also been evaluated (Huang et al, 2004). As yet, no specific inhibitors of Axl activity have been described. Our results suggest that further studies aimed at further elucidating the potential role of Axl in SCC are merited, as it may represent a potential therapeutic target for intervention in skin cancer development.
ACKNOWLEDGEMENTS
We thank B Young for critical reading of the manuscript and helpful suggestions. This research was supported by funding from Cancer Research UK and AICR Grant 03-315 to EA O'Toole.
Footnotes
Supplementary Information accompanies the paper on British Journal of Cancer website (http://www.nature.com/bjc)
REFERENCES
- Blackledge G. Cancer drugs: the next 10 years. Eur J Cancer. 2003;39:273. [PubMed] [Google Scholar]
- Blackledge G, Averbuch S. Gefitinib (‘Iressa’, ZD1839) and new epidermal growth factor receptor inhibitors. Br J Cancer. 2004;90:566–572. doi: 10.1038/sj.bjc.6601550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung BI, Malkowicz SB, Nguyen TB, Libertino JA, McGarvey TW. Expression of the proto-oncogene Axl in renal cell carcinoma. DNA Cell Biol. 2003;22:533–540. doi: 10.1089/10445490360708946. [DOI] [PubMed] [Google Scholar]
- Diepgen TL, Mahler V. The epidemiology of skin cancer. Br J Dermatol. 2002;146(Suppl 61):1–6. doi: 10.1046/j.1365-2133.146.s61.2.x. [DOI] [PubMed] [Google Scholar]
- Druker BJ. Imatinib as a paradigm of targeted therapies. Adv Cancer Res. 2004;91:1–30. doi: 10.1016/S0065-230X(04)91001-9. [DOI] [PubMed] [Google Scholar]
- Emens LA, Davidson NE. Trastuzumab in breast cancer. Oncology (Huntington) 2004;18:1117–1128. discussion 1131–1132, 1137–1138. [PubMed] [Google Scholar]
- Finn RS, Slamon DJ. Monoclonal antibody therapy for breast cancer: herceptin. Cancer Chemother Biol Response Modif. 2003;21:223–233. doi: 10.1016/s0921-4410(03)21010-3. [DOI] [PubMed] [Google Scholar]
- Holland SJ, Powell MJ, Franci C, Chan EW, Friera AM, Atchison RE, McLaughlin J, Swift SE, Pali ES, Yam G, Wong S, Lasaga J, Shen MR, Yu S, Xu W, Hitoshi Y, Bogenberger J, Nor JE, Payan DG, Lorens JB. Multiple roles for the receptor tyrosine kinase axl in tumor formation. Cancer Res. 2005;65:9294–9303. doi: 10.1158/0008-5472.CAN-05-0993. [DOI] [PubMed] [Google Scholar]
- Holme SA, Malinovszky K, Roberts DL. Changing trends in non-melanoma skin cancer in South Wales, 1988–98. Br J Dermatol. 2000;143:1224–1229. doi: 10.1046/j.1365-2133.2000.03892.x. [DOI] [PubMed] [Google Scholar]
- Huang S, Armstrong EA, Benavente S, Chinnaiyan P, Harari PM. Dual-agent molecular targeting of the epidermal growth factor receptor (EGFR): combining anti-EGFR antibody with tyrosine kinase inhibitor. Cancer Res. 2004;64:5355–5362. doi: 10.1158/0008-5472.CAN-04-0562. [DOI] [PubMed] [Google Scholar]
- Jackson S, Ghali L, Harwood C, Storey A. Reduced apoptotic levels in squamous but not basal cell carcinomas correlates with detection of cutaneous human papillomavirus. Br J Cancer. 2002;87:319–323. doi: 10.1038/sj.bjc.6600431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson RL, Rothman AL, Xie J, Goodrich LV, Bare JW, Bonifas JM, Quinn AG, Myers RM, Cox DR, Epstein EH, Jr, Scott MP. Human homolog of patched, a candidate gene for the basal cell nevus syndrome. Science. 1996;272:1668–1671. doi: 10.1126/science.272.5268.1668. [DOI] [PubMed] [Google Scholar]
- Loercher A, Lee TL, Ricker JL, Howard A, Geoghegen J, Chen Z, Sunwoo JB, Sitcheran R, Chuang EY, Mitchell JB, Baldwin AS, Jr, Van Waes C. Nuclear factor-kappaB is an important modulator of the altered gene expression profile and malignant phenotype in squamous cell carcinoma. Cancer Res. 2004;64:6511–6523. doi: 10.1158/0008-5472.CAN-04-0852. [DOI] [PubMed] [Google Scholar]
- Mineo JF, Bordron A, Quintin-Roue I, Loisel S, Ster KL, Buhe V, Lagarde N, Berthou C. Recombinant humanised anti-HER2/neu antibody (Herceptin) induces cellular death of glioblastomas. Br J Cancer. 2004;91:1195–1199. doi: 10.1038/sj.bjc.6602089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakano T, Tani M, Ishibashi Y, Kimura K, Park YB, Imaizumi N, Tsuda H, Aoyagi K, Sasaki H, Ohwada S, Yokota J. Biological properties and gene expression associated with metastatic potential of human osteosarcoma. Clin Exp Metastasis. 2003;20:665–674. doi: 10.1023/a:1027355610603. [DOI] [PubMed] [Google Scholar]
- O'Bryan JP, Frye RA, Cogswell PC, Neubauer A, Kitch B, Prokop C, Espinosa R, III, Le Beau MM, Earp HS, Liu ET. axl, a transforming gene isolated from primary human myeloid leukemia cells, encodes a novel receptor tyrosine kinase. Mol Cell Biol. 1991;11:5016–5031. doi: 10.1128/mcb.11.10.5016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park JK, Lee SH, Kang JH, Nishio K, Saijo N, Kuh HJ. Synergistic interaction between gefitinib (Iressa, ZD1839) and paclitaxel against human gastric carcinoma cells. Anticancer Drugs. 2004;15:809–818. doi: 10.1097/00001813-200409000-00011. [DOI] [PubMed] [Google Scholar]
- Popp S, Waltering S, Holtgreve-Grez H, Jauch A, Proby C, Leigh IM, Boukamp P. Genetic characterization of a human skin carcinoma progression model: from primary tumor to metastasis. J Invest Dermatol. 2000;115:1095–1103. doi: 10.1046/j.1523-1747.2000.00173.x. [DOI] [PubMed] [Google Scholar]
- Proby CM, Purdie KJ, Sexton CJ, Purkis P, Navsaria HA, Stables JN, Leigh IM. Spontaneous keratinocyte cell lines representing early and advanced stages of malignant transformation of the epidermis. Exp Dermatol. 2000;9:104–117. doi: 10.1034/j.1600-0625.2000.009002104.x. [DOI] [PubMed] [Google Scholar]
- Pulsipher MA. Treatment of CML in pediatric patients: should imatinib mesylate (STI-571, Gleevec) or allogeneic hematopoietic cell transplant be front-line therapy? Pediatr Blood Cancer. 2004;43:523–533. doi: 10.1002/pbc.20062. [DOI] [PubMed] [Google Scholar]
- Rheinwald JG, Green H. Serial cultivation of strains of human epidermal keratinocytes: the formation of keratinizing colonies from single cells. Cell. 1975;6:331–343. doi: 10.1016/s0092-8674(75)80001-8. [DOI] [PubMed] [Google Scholar]
- Ross DM, Hughes TP. Cancer treatment with kinase inhibitors: what have we learnt from imatinib? Br J Cancer. 2004;90:12–19. doi: 10.1038/sj.bjc.6601507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun W, Fujimoto J, Tamaya T. Coexpression of Gas6/Axl in human ovarian cancers. Oncology. 2004;66:450–457. doi: 10.1159/000079499. [DOI] [PubMed] [Google Scholar]
- Tanno S, Ohsaki Y, Nakanishi K, Toyoshima E, Kikuchi K. Small cell lung cancer cells express EGFR and tyrosine phosphorylation of EGFR is inhibited by gefitinib (‘Iressa’, ZD1839) Oncol Rep. 2004;12:1053–1057. [PubMed] [Google Scholar]
- van Ginkel PR, Gee RL, Shearer RL, Subramanian L, Walker TM, Albert DM, Meisner LF, Varnum BC, Polans AS. Expression of the receptor tyrosine kinase Axl promotes ocular melanoma cell survival. Cancer Res. 2004;64:128–134. doi: 10.1158/0008-5472.can-03-0245. [DOI] [PubMed] [Google Scholar]
- Wakeling AE, Guy SP, Woodburn JR, Ashton SE, Curry BJ, Barker AJ, Gibson KH. ZD1839 (Iressa): an orally active inhibitor of epidermal growth factor signaling with potential for cancer therapy. Cancer Res. 2002;62:5749–5754. [PubMed] [Google Scholar]