Abstract
The role(s) of the novel stargazin-like γ-subunit proteins remain controversial. We have shown previously that the neuron-specific γ7 suppresses the expression of certain calcium channels, particularly CaV2.2, and is therefore unlikely to operate as a calcium channel subunit. We now show that the effect of γ7 on CaV2.2 expression is via an increase in the degradation rate of CaV2.2 mRNA, and hence a reduction of CaV2.2 protein level. Furthermore, exogenous expression of γ7 in PC12 cells also decreased the endogenous CaV2.2 mRNA level. Conversely, knockdown of endogenous γ7 with short-hairpin RNAs produced a reciprocal enhancement of CaV2.2 mRNA stability and an increase in endogenous calcium currents in PC12 cells. Moreover, both endogenous and expressed γ7 are present on intracellular membranes, rather than the plasma membrane. The cytoplasmic C-terminus of γ7 is essential for all its effects, and we show that γ7 binds directly via its C-terminus to a ribonucleoprotein (hnRNP A2), which also binds to a motif in CaV2.2 mRNA, and is associated with native CaV2.2 mRNA in PC12 cells. The expression of hnRNP A2 enhances CaV2.2 IBa and this enhancement is prevented by a concentration of γ7 that alone has no effect on IBa. The effect of γ7 is selective for certain mRNAs as it had no effect on α2δ-2 mRNA stability, but it decreased the mRNA stability for the potassium-chloride co-transporter, KCC1, which contains a similar hnRNP A2 binding motif to that in CaV2.2 mRNA. Our results indicate that γ7 plays a role in stabilizing CaV2.2 mRNA.
Keywords: N-type, calcium channel, mRNA stability, gamma7 subunit, hnRNP A2, stargazin
Introduction
Voltage-dependent calcium channels (CaV) are hetero-multimers consisting of a pore-forming α1 subunit, assembled with auxiliary β, α2δ and possibly γ subunits (for review see Catterall, 2000; Dolphin, 2003a; Dolphin, 2003b). The role(s) of the γ subunits in relation to calcium channel function remains unclear. The first CaVγ subunit to be identified was γ1, which co-purifies with the skeletal muscle calcium channel complex (Jay et al., 1991; Powers et al., 1993). In skeletal muscle the γ1 subunit appears to have a suppressive effect, as γ1 knockout mice exhibit increased skeletal muscle calcium currents (Freise et al., 2000). Following the identification of stargazin (γ2) (Letts et al., 1998), subsequent studies have identified six further putative γ subunits (γ3-γ8) (Black and Lennon, 1999; Burgess et al., 1999; Klugbauer et al., 2000; Burgess et al., 2001; Moss et al., 2002). However, it is unclear whether any of these novel stargazin-like γ proteins (γ2-8) play any role as subunits of voltage-gated calcium channels. All members of this family are thought to possess four transmembrane spanning domains with intracellular N- and C-termini. The γ2, γ3, γ4 and γ8 subunits form a sub-family exclusively localized to the central nervous system (Letts et al., 1998; Klugbauer et al., 2000; Sharp et al., 2001; Moss et al., 2003) whose interaction and functional modulation of CaV channels has been investigated in several studies (Letts et al., 1998; Klugbauer et al., 2000; Sharp et al., 2001; Kang et al., 2001; Rousset et al., 2001; Moss et al., 2003), but which are now thought primarily to represent trafficking proteins for the alpha-amino-3-hydroxyl-5-methyl-4-isoxazolepropionate (AMPA) subtype of glutamate receptors (TARPS) (Tomita et al., 2003; Tomita et al., 2004). However, they might also provide a bridge between calcium channels and AMPA receptors (Kang et al., 2006). Furthermore, γ7, despite not having a classical C-terminal PDZ binding domain, has also recently been shown to have effects on AMPA receptor trafficking (Kato et al., 2007).
The γ7 and γ5 proteins are predicted to represent a distinct sub-family of stargazin-related proteins (Burgess et al., 2001; Chu et al., 2001; Moss et al., 2002), with extremely low sequence identity to γ1 and approximately 25% identity to γ2, mainly in the transmembrane domains. We showed that co-expression of the γ7 subunit with CaV2.2 almost abolished the functional expression and markedly suppressed the level of CaV2.2 α1 subunit protein (Moss et al., 2002). It also had smaller suppressive effects on CaV2.1 and CaV1.2 currents (Moss et al., 2002). Our conclusion was that γ7 was not a subunit of these calcium channels. Nevertheless, because of the marked effect of γ7 on calcium channel expression, and since both N-type calcium channels and γ7 are specifically expressed in neuronal tissue, we have now examined the mechanism of action of γ7 in order to probe its physiological function.
The present results indicate that γ7 is involved in regulating the stability of specific mRNAs, including CaV2.2. We propose that the mechanism may involve γ7 sequestering a specific mRNA binding protein, thus compromising the stability of CaV2.2 mRNA.
Materials and Methods
cDNA constructs
The following cDNAs were used: CaV2.2 (D14157) and CaV2.2 δ3′UTR, CaV3.1 (AF027984), β1b (X61394, Dr. T. P. Snutch), α2δ-2 (AF247139, common brain splice variant), γ7 (NM031896), mut-3b Green Fluorescent Protein (GFP, M62653, except S72A and S65G, Dr. T. E. Hughes), mouse KCC1 (AF121118), KV3.1b (M68880), hnRNP A1 (BC070315) and hnRNP A2-ΔRGG (Nichols et al., 2000). Truncated and tagged γ7 constructs and HA-tagged hnRNP A2 were generated using standard molecular biological techniques and confirmed by DNA sequencing. All constructs were cloned into the pMT2 expression vector (Swick et al., 1992), except γ7-HA and γ7-CFP, hnRNP A2-ΔRGG and KCC1, all of which were in pCDNA3.1, except γ7-CFP in pECFP-N1, Kv3.1b in pRC-CMV and hnRNP A1 in pCGT7. pDsRed2-ER plasmid was used in some experiments (Clontech).
Short hairpin RNA design and expression plasmid
siSearch (http://sonnhammer.sbc.su.se/databases.html) and Jura (http://jura.wi.mit.edu/siRNAext/) software was used to design 19-nucleotides sequences corresponding to human and rat γ7 genes. Databases were searched to ensure that these sequences were not homologous to any other known genes. Rat γ7 (AF361345) targets: γ7 96 (5′-CTGGCTGTATATGGAGGAG-3′), γ7 285 (5′-GACAGTACGCACGGCTACA-3′), γ7 500 (5′-CTGAGCAATACTTTCACTA-3′). Human γ7 (AF458897) targets: γ7 96 (5′-CTGGCTGTACATGGAAGAA-3′), γ7 285 (5′-GACAGTGCGCACGGCCACC-3′), γ7 107 (5′-TGGAAGAAGGCACAGTGCT-3′). Then, for each target, 2 oligonucleotides (A and B) were synthesized (Invitrogen, Paisley UK). Oligonucleotide A contains a 5′-overhang (TTTG) for ligation into a BpiI (BbsI) site, the sense and the antisense of these sequences linked by a hairpin loop of 9 bases (TTCAAGAGA (Brummelkamp et al., 2002)), and a TTTTT sequence corresponding to a termination for transcription of small RNAs by RNA polymerase III; oligonucleotide B contains a 5′-overhang (CTAG) for ligation into a XbaI site, and 52 nucleotides corresponding to the reverse complement of the last 52 nucleotides of oligonucleotide A. Forward and reverse strands were annealed and subcloned downstream a U6 promoter into pG418-shRNA-Empty linearized with XbaI and BpiI. pG418-shRNA-Empty is a pBSII plasmid containing a mouse U6 promoter and a neomycin (G418) resistance cassette. A negative control shRNA, directed against drosophila gene gnu (Xu and Shrager, 2005) and a positive control shRNA directed against c-jun (Lingor et al., 2005), were also synthesized. Correct orientation and location of oligonucleotides cloning were confirmed by sequencing the plasmids. Validation of the plasmid in PC12 cells was determined by its ability to knock-down c-jun (see Supplemental Fig. 1).
Antibodies against γ7
The γ7 tail polyclonal rabbit Ab raised against the C-terminal peptide YPPAIKYPDHLHIS has been described previously (Moss et al., 2002). A γ7 loop Ab was also raised against the peptide VASEYFLEPEINLVTEN, in the loop between transmembrane segments 1 and 2 of γ7. These Abs do not recognize any of the other γ proteins tested (γ2, γ3 or γ4, Supplemental Fig. 2A-C).
Cell culture and transfection
COS-7 cells were cultured as previously described (Campbell et al., 1995). The tsA 201 cells were cultured in D-MEM with 10% fetal bovine serum (FBS) and 1% L-glutamine. Cells were transfected using either Geneporter (Qbiogene, Harefield, UK) or Fugene6 (Roche Diagnostics, Lewes, UK), with equivalent results. The cDNAs (all at 1 μg.μl-1) for CaVα1, α2δ-2, β1b, γ7 and GFP when used as a reporter of transfected cells, were mixed in a ratio of 3 : 1.5 : 2 : 1.5 : 1 : 0.2, unless otherwise stated. When particular subunits were not used, the volume was made up with Tris-EDTA (TE, 10 mM Tris, 1 mM EDTA pH7), or blank vector, or the volume of transfection reagent was reduced, with equivalent results. In some experiments cDNA for the non-conducting potassium channel Kir2.1-AAA (Tinker et al., 1996) was used as control for the presence of γ7, also with equivalent results to the use of blank vector.
PC12 cells were grown in D-MEM, 7.5% FBS and 7.5% horse serum. For electrophysiology, PC12 cells were transiently transfected with cDNAs for GFP and/or γ7, using Fugene6. Differentiation was with serum-free medium containing NGF (100 ng.ml-1 murine 7s NGF, Invitrogen, Paisley, UK), replenished every 48 h. Cells were used for recording after 5 -7 days of differentiation. For the generation of stable cell lines, PC12 cells were transfected with γ7-HA or γ7-CFP and clonal cell lines were established by standard techniques, using 400 μg.ml-1 Geneticin (Invitrogen) for selection. The culture medium was subsequently supplemented with 400 μg/ml Geneticin (Invitrogen).
To obtain high transfection rates with shRNAs, PC12 cells were transfected using an Amaxa Nucleofector, according to manufacturer instructions (Amaxa, Cologne, Germany). The DNA mix contained 2 μg of shRNA plasmid and 0.5 μg of GFP or YFP as a reporter of transfected cells. Cells were used 4 to 5 days after transfection with shRNA.
For primary culture of superior cervical ganglion (SCG) neurons, rats were killed by either CO2 inhalation or cervical dislocation, according to UK Home Office Schedule 1 Guidelines. SCGs were dissected from rats at postnatal day 17. Ganglia were desheathed and lightly gashed before successive collagenase (Sigma) and trypsin (Sigma) treatment, both at 3 mg/ml. To produce a single-cell suspension, ganglia were dissociated by trituration and centrifugation. Dissociated cells were plated onto glass-bottomed plates (MatTeK Corp., Ashland, USA) pre-coated with laminin (Sigma), using 1 ganglion per 5 plates. Cells were maintained with Liebovitz L-15 medium (Sigma), supplemented with 24mM NaHCO3, 10% FBS (Gibco), 33mM glucose (Sigma), 20mM L-glutamine, 1000 IU Penicillin, 1000 IU Streptomycin (Gibco) and 50ng/ml NGF.
Microinjection
cDNAs were injected into SCG neurons 18-24h after they were placed in culture. Microinjection was performed using an Eppendorf microinjection system on a Zeiss Axiovert 200M microscope using the following settings: 100-150 hPa injection pressure, an injection time of 0.2 s and constant pressure of between 40-50 hPa. The cDNAs were injected at 50ng/μl diluted in 200mM KCl.
Measurement of mRNA levels by q-PCR
RNA was isolated using RNeasy columns (Qiagen), including an on-column DNase step. Reverse transcription was carried out using random hexamer primers and Moloney Murine Leukemia Virus Reverse Transcriptase (Promega, Southampton, UK) at 37°C for 2 h. The q-PCR was performed with an iCycler (BioRad, Hercules CA, USA) using the iQ SYBR supermix. For each set of primers and for every experiment a standard curve was generated using a serial dilution of reverse-transcribed RNA combined from several samples. For q-PCR in PC12 cells, the following primers were used: rat CaV2.2 (NM147141) 5′-GGCAAGAAGGAGGCAGAG-3′, 5′-GCAGAAGCGACGGAGTAG-3′; rat γ7 (AF361345) 5′-CTACTCGGGCCAGTTTCTGC-3′, 5′-GCCGGAGGGTAATTTTGC-3′; and rat glyceraldehyde-3-phosphate dehydrogenase GAPDH (AF106860) 5′-ATGACTCTACCCACGGCAAG-3′, 5′-CAT ACT CTG CAC CAG CAT CTC-3′; Data were normalized for expression of GAPDH mRNA.
For measurement of mRNA degradation rates, q-PCR of transcript levels was performed in Xenopus oocytes. Plasmid cDNAs were injected intranuclearly for CaV2.2, β1b, α2δ-2, either with or without γ7. After 24 h, oocytes were incubated for the stated times with actinomycin D (50 μg/ml). RNA extraction, RT and q-PCR were performed as described above. The following primers were used: CaV2.2: 5′-CTCTGCGCTTACTGAGAATC-3′ and 5′-AACAGGAAGAGCAGGAAGAG-3′; 18S: 5′-TGACTCAACACGGGAAACCT-3′ and 5′-AATCGCTCCACCAACTAAGAAC-3′; α2δ2: 5′-GGTATTTGCTGCCACTGATG-3′ and 5′-AGGCTGCGACGGTAGAAG-3′; KCC1: 5′-AGCATAAGGTTTGGAAGAAGTG-3′ and 5′-CAGGCGGAGGTGATACAG-3′. Data were normalized for expression of 18S ribosomal RNA.
Oligoribonucleotide binding to hnRNP A2
This was performed as described previously (Hoek et al., 1998), with the exception that mouse brain, rather than rat brain, was used as a source of hnRNP A2. The following RNA oligonucleotides were used A2RE 5′-Biotin: GCCAAGGAGCCAGAGAGCAUG-3′; CaV2.2 5′-Biotin: GCCAAGGAGCGGGAGCGAGUC-3′; Non-specific 5′-Biotin: CAAGCACCGAACCCGCAACUG-3′. A2RE and non-specific sequences are identical in base composition (Hoek et al., 1998). Proteins were taken up in 200 μl of SDS sample buffer and heated at 65° C for 10 minutes. 20 μl of each was run on a 4-12% Bis-Tris gel and western blotting and immunodetection was performed with anti-hnRNP A2 Ab (Autogen Bioclear, 1:200).
Yeast two hybrid assay
Assays were carried out using the MATCHMAKER GAL4 two hybrid system (Clontech). Fragments of hnRNP A2 (amino acids 1-177 or 178-342), the γ7 C-terminus (201-275), the CaV2.2 I-II loop (360-483) and CaVβ1b were generated by PCR and subcloned in-frame into the vectors pACT2 and pAS2-1. Plasmids were cotransformed into the yeast strain Y190 and transformants were selected by plating onto minimal selective dropout (SD) -Leu, -Trp agar. Protein interactions were identified by restreaking colonies onto SD -Leu, -Trp, -His plates and carrying out colony-lift β-galactosidase assays, or growth assays in the presence of 10 mM 3-amino-1,2,4-triazole, according to the supplied protocol.
Immunoprecipitation
γ7-HA was immunoprecipitated from stably transfected PC12 and transiently transfected tsA 201 cells. Endogenous γ7 and hnRNP A2 proteins were immunoprecipitated from untransfected PC12 cells. The following general method was used. Cells were washed with ice cold phosphate buffered saline (PBS; Sigma-Aldrich, Gillingham, UK) and harvested in PBS with 10 mM EDTA and protease inhibitor cocktail (Roche Diagnostics). Cells were lysed and protein solubilized by agitation with extraction buffer (1% Igepal, 20 mM Tris, 150 mM NaCl, 1 mM EDTA, 0.5% sodium deoxycholate, 0.1% SDS and protease inhibitor cocktail, pH 7.4) for 30 min. at 4°C. Insoluble material was removed by centrifugation at 40,000 × g for 1 h at 4°C. The supernatant was cleared with 50 μg of Protein G linked to agarose beads (Sigma) for 2 h at 4°C. The supernatant was incubated with 2μg of high affinity Anti-HA Ab (Clone 3F10, Roche Diagnostics), or γ7 C-terminus Ab, or hnRNP A2 monoclonal Ab EF67 (Santa Cruz Biotechnology, Santa Cruz, CA), overnight at 4°C with constant agitation. A further 20 μg of Protein G linked to agarose beads was added and incubated for 2 h at 4°C with constant agitation. Beads were washed twice with a high detergent buffer (1% Igepal, 20 mM Tris, 150 mM NaCl, 1 mM EDTA, protease inhibitor cocktail, pH 7.4), twice with a high salt buffer (0.1% Igepal, 20 mM Tris, 500 mM NaCl, 1 mM EDTA, protease inhibitor cocktail, pH 7.4) and twice with a low salt buffer (0.1% Igepal, 20 mM Tris, 1 mM EDTA, protease inhibitor cocktail, pH 7.4). Bound protein was removed from the beads by the addition of lauryl dodecyl sulphate (LDS) sample buffer with reducing agent (Invitrogen), and heating to 65 °C for 10 min. Samples containing immunoprecipitated endogenous γ7 were treated with LDS buffer as above and eluted proteins concentrated by precipitation with ice-cold acetone prior to resuspension in fresh LDS buffer for polyacrylamide gel electrophoresis.
For protein sequencing, samples were separated on 4-12% Bis-Tris gels (Invitrogen) and stained with Coomassie (Simply safe Blue stain, Invitrogen). Bands of interest were excised from the gel and protein identification was performed by the Imperial College Proteomics Facility by tryptic mass fingerprinting, confirmed by MS-MS. The controls were from non-transfected cells, treated identically.
Coimmunoprecipitation of RNAs associated with hnRNP A2
PC12 cells were lysed and protein solubilized by agitation with extraction buffer containing 1% Igepal, 20 mM Tris, 150 mM NaCl, 1 mM EDTA, 0.5% sodium deoxycholate, protease inhibitor cocktail (pH 7.4), supplemented with 5U/ml RNAguard (GE Healthcare), for 30 minutes at 4°C. Samples were centrifuged (50,000 × g, 1 h at 4°C) and the corresponding supernatant was then precleared with 50 μg protein G-sepharose. The supernatant was divided into two and either incubated with 2μg/ml of high affinity anti-HA Ab or an equivalent volume of PBS overnight at 4°C with constant agitation. A further 20 μg of protein G-sepharose was added and incubated for 2 h at 4°C with constant agitation. Beads were washed six times by centrifugation as described in the previous section, except that the wash buffers were supplemented with 5U/ml RNAguard. A small aliquot of beads was removed for protein analysis. The RNA on the remaining beads was extracted with Trizol (Invitrogen). Total RNA was reverse-transcribed with SuperScript III (Invitrogen) using random primers. PCR was performed using the primers described above.
Western blotting
Cells were processed for SDS PAGE as previously described (Raghib et al., 2001). For detection of endogenous γ7 in PC12 cells, cells were lysed on ice by sonication for 3 × 10s, and centrifuged at 3000 × g for 3 min. This supernatant was decanted and centrifuged at 50,000 × g for 4 h at 4°C. The high-speed supernatant and membrane-containing pellet were then taken up in SDS buffer. Samples (50 μg cell lysate protein/lane) or IP samples prepared as above were separated using Novex 4-12% Tris-glycine or 4-12% Bis-Tris NuPAGE gels (Invitrogen) and transferred electrophoretically to polyvinylidene fluoride membranes. The membranes were blocked with 3% BSA / 0.02% Tween 20 and then incubated overnight at room temperature with the relevant primary Ab: rabbit anti-hnRNP A/B Ab (H-200, Autogen Bioclear, Calne, UK or Santa Cruz Biotechnology, Santa Cruz, CA, 1:1000); anti-hnRNP A/B monoclonal Ab from(F16, Autogen Bioclear, 1:1000); anti hnRNP A2 monoclonal Ab (EF-67, 1:1000 (Nichols et al., 2000)); rabbit anti-HA Ab, (Sigma, 1:1000), affinity-purified anti- γ7 Ab (0.4 μg.ml-1) or anti-α2δ-2(102-117) (1 μg.ml-1) (Brodbeck et al., 2002). The γ2, γ3 and γ4 Abs have been described previously (Moss et al., 2003). Detection was performed using anti-rabbit or anti-mouse secondary Ab conjugated to HRP (BioRad, 1μg.ml-1), and bound Abs were detected using enhanced chemiluminescence (ECL) or ECLPlus reagents (Amersham Pharmacia Biotech (APB), Little Chalfont, UK). Chemiluminescence or fluorescence was detected using a Typhoon 9410 Variable Mode Imager (APB), set in chemiluminescence or fluorescence mode, respectively. Protein bands were quantified using Imagequant 5.2, on non-saturated images.
Immunocytochemistry
Cells were fixed and permeablized for immunocytochemistry essentially as previously described (Brice et al., 1997). The primary Abs used were affinity-purified anti- γ7 loop or tail Ab (as stated, 0.8 μg.ml-1), mouse monoclonal anti-PDI (Abcam, Cambridge, UK, 1:100). Alexa Fluor 594-phalloidin (Invitrogen) was also used. Secondary Texas red, FITC or biotin-conjugated goat anti-mouse (Molecular Probes, Eugene, Oregon) or goat anti-rabbit (Sigma) Abs were applied at 10 μg.ml-1 and 5 μg.ml-1 respectively. When used, Texas red- or FITC-conjugated streptavidin were applied at 3.33 μg.ml-1. A Cy3- tyramide signal amplification kit (Perkin Elmer) was used to detect γ7 in SCG neurons. In some experiments, the nuclear dye 4′,6-diamidino-2-phenylindole (DAPI, 300 nM, Molecular Probes) was also used to visualize the nucleus. Cells were mounted in Vectashield (Vector Laboratories, Burlingame, CA) to reduce photobleaching, and examined on a confocal laser scanning microscope (Leica TCS SP or Ziess LSM), using a ×40 (1.3 NA) or ×63 (1.4 NA) oil-immersion objective. Optical sections were 1.5 μm. Photomultiplier settings were kept constant in each experiment and all images were scanned sequentially. In some experiments, where stated, a conventional fluorescence microscope (Axiovert 200, Zeiss) and CCD camera was used; images were captured using Volocity software (Improvision, UK).
Image processing was performed using ImageJ (http://rsb.info.nih.gov/ij/). Co-localization analysis was performed using the co-localization plugin on images converted to 8 bit. Fluorophores are considered as co-localized in each pixel when their respective intensities are higher than the threshold (50) of their channels and when their intensity ratio is greater than 50%.
Electrophysiology
Xenopus oocytes were prepared, injected and utilized for electrophysiology as previously described (Canti et al., 1999), with the following exceptions. Plasmid cDNAs for the different VDCC subunits α1, α2δ-2, β1b and other constructs such as γ7 were mixed in equivalent weight ratios at 1 μg.μl-1, unless otherwise stated, and 9 nl injected intranuclearly, following appropriate dilution. The recording solution for CaV2.2-injected oocytes contained (in mM): Ba(OH)2 10; TEA-OH 80; CsOH 2; Hepes 5 (pH 7.4 with methanesulfonic acid).
Whole-cell patch-clamp recording using tsA 201 or PC12 cells was performed and analyzed as described (Meir et al., 2000), with 1 mM Ba2+ as charge carrier (unless stated) and a holding potential of -100mV. Currents were measured 10 ms after the onset of the test pulse and the average over a 2 ms period was calculated and used for analysis. Data are expressed as mean ± s.e.m and I-V plots were fit with a modified Boltzmann equation as described (Canti et al., 2001).
Results
The reduction of functional CaV2.2 protein by γ7 is mediated via the C-terminus of γ7
These experiments were initiated in the light of our previous finding that co-expression of γ7 reduced CaV2.2 protein level, as well as its functional expression (Moss et al., 2002). We have now dissected the region of γ7 responsible for the inhibitory effect by making constructs lacking most or part of the cytoplasmic C-terminus, γ7(1-217) and γ7(1-238) (Fig. 1A). Following cDNA injection in Xenopus oocytes, when full-length γ7 was co-expressed with CaV2.2 it produced ~90 % suppression of CaV2.2 currents (Fig. 1B, C). In contrast, the shorter γ7(1-217) transmembrane construct had very little influence on the expression of CaV2.2 currents (Fig. 1B, C), whereas the γ7(1-238) construct produced a partial reduction of the CaV2.2 current (Fig. 1C). The C-terminal motif (T/S-SPC) is conserved between γ7 and γ5. Although this does not represent a classical PDZ binding motif, we investigated the importance of this epitope in mediating the effects of γ7 on CaV2.2 currents. A construct lacking these four amino acids (γ7(1-271)) was as effective as γ7 in reducing CaV2.2 IBa (Fig. 1C), indicating that this motif is not involved in the effect of γ7. Furthermore, addition of a C-terminal tag, such as haemaglutinin (HA) or CFP, did not affect the ability of γ7 to suppress the expression of CaV2.2 currents. For γ7-HA, 72.5 ± 10.1 % (n=5) inhibition of CaV2.2 currents was observed. These results indicate that a region of the γ7 C-terminus between R217 and S271 is responsible for inhibiting CaV2.2 expression.
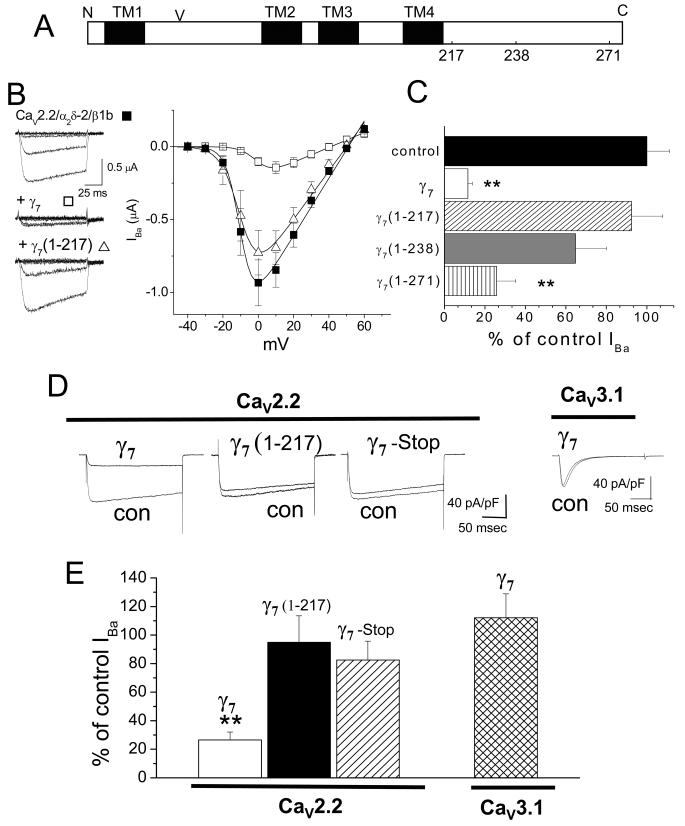
Figure 1. Effect of C-terminal truncation of γ7 on CaV2.2/β1b/α2δ-2 and CaV3.1 currents.
A, Linearized diagram of γ7, indicating the approximate positions of the 4 transmembrane (TM) segments (black bars), the N-glycosylation site (V) at N45, and the position of the truncations at amino acids 217, 238 and 271.
B, Left: example traces elicited following cDNA injection into Xenopus oocytes by 100 ms step depolarizations to between -40 and 0 mV from a holding potential of -100 mV for CaV2.2/β1b/α2δ-2 (top), plus γ7 (center) and plus γ7 (1-217) (bottom). The charge carrier was 10 mM Ba2+. The symbols beside the traces refer to the relevant data in the current-voltage relationship (right); CaV2.2/β1b/α2δ-2 (■, n= 19), plus γ7 (1-217) (Δ, n =18) and plus γ7 (□, n=8). Data are fit by a modified Boltzmann function as described in Methods, with V50, act of -9.9, -9.8 and +1.2 mV respectively and Gmax of 19.6, 16.6 and 5.9 μS, respectively.
C, Mean % of control peak IBa for CaV2.2/β1b/α2δ-2 currents (black bar, n=29) when co-expressed with γ7 (white bar, n=31) or its truncated constructs, γ7(1-217) (hatched bar, n=15), γ7(1-238) (grey bar, n=12) and γ7(1-271) (striped bar, n=16). The statistical significances compared to control are ** P<0.01. There was no effect of any of the constructs on the voltage for 50 % steady-state inactivation, which from combined experiments was -60.2 ± 0.8 mV for controls (n=16), -58.6 ± 1.0 mV in the presence of γ7 (n=7), -61.7 ± 3.6 mV for γ7(1-217) (n=5) and -59.8 ± 1.0 mV for γ7(1-271) (n=5).
D, Representative traces of peak Ba2+ currents in tsA 201 cells, co-transfected with CaV2.2, β1b and α2δ-2 with pMT2 as control (con), compared to γ7 (panel 1), γ7(1-217) (panel 2), γ7(stop) (panel 3). Panel 4 shows CaV3.1 expression with pMT2 as control (con), compared to γ7. Currents were elicited by depolarization to +5 mV (CaV2.2, 1 mM Ba2+) or -10 mV (Cav 3.1, 10 mM Ba2+), from a holding potential of -100 mV. Calibration bars refer to all traces for CaV2.2.
E, Mean inhibition of Ba2+ currents (expressed as % of control ± s.e.m.) induced by co-expression of CaV2.2 with γ7 (white bar, n=21), γ7 (1-217) (black bar, n=15), γ7(stop) (hatched bar, n=28), or CaV3.1 with γ7 (cross-hatched bar, n=29). Statistical significance compared to the current size without γ7 ** P<0.01, Student’s t test.
The role of the C-terminus of γ7 was confirmed in tsA-201 cells. Here again the truncated γ7(1-217) produced no inhibition of CaV2.2 currents (5.1 ± 18.6 %, Fig. 1D, E), whereas full-length γ7 produced 94.9 ± 18.6 % inhibition (Fig. 1D, E). To confirm that the γ7 protein was responsible for this effect, we showed that a γ7 construct containing a stop codon prior to the first transmembrane domain produced no inhibition (Fig. 1D, E). Furthermore, in contrast to the effect of γ7 on CaV2.2 currents, it had no significant effect on CaV3.1 currents (Fig. 1D, E).
We raised two antipeptide Abs against γ7, to unique peptides in the linker between transmembrane segments I and II, and in the C-terminus. Neither Ab recognised any protein bands in untransfected Cos-7 cells (Fig. 2A), and neither Ab cross-reacted with other γ proteins tested (Supplemental Fig. 2A-C). We then confirmed that the truncated γ7 constructs were all expressed at a similar level to full-length γ7, and that the truncated constructs were of the expected size (Fig. 2A). Next, we examined the effect of γ7 and its C-terminal truncations on the level of CaV2.2 protein. Both γ7 and γ7(1-271) produced approximately 70% inhibition of CaV2.2 protein, whereas γ7(1-217) caused no reduction (Fig. 2B). The extent to which CaV2.2 protein expression was suppressed by γ7 and its truncated constructs was closely correlated with the degree of inhibition of CaV2.2 IBa observed in Xenopus oocytes (Fig. 2C). As a control, we showed that expression of an unrelated protein of similar size to γ7 (KV3.1b) had no effect on CaV2.2 protein expression (Fig. 2B).
Figure 2. Effect of C-terminal truncation of γ7 on CaV2.2 protein.
A, γ7 and truncated γ7 constructs were expressed at the expected sizes in Cos-7 cells (upper panel γ7 I-II loop Ab, lower panel γ7 C terminal tail Ab). In each lane, the lower band corresponds to the expected protein molecular weight, and the upper band(s) correspond to either the mature glycosylated form or intermediate glycosylated species. As expected γ7(1-217) and γ7(1-238) were not detected by the γ7(C-terminal tail) Ab. The specificity of the Abs is indicated by the lack of immunostaining in the absence of transfected constructs (-). Similar results were obtained in Xenopus oocytes (data not shown). The same amount of total protein was loaded in each lane (25 μg).
B, Examples of Western blots showing the effect of (i) γ7, and (ii) γ7(1-271), and the lack of effect of (iii) γ7(1-217) and (iv) KV3.1b on the level of CaV2.2 protein expressed in Cos-7 cells. Con = transfection with Kir-AAA cDNA, in place of γ7 (see Methods). The same amount of total protein was loaded in each pair of lanes (25 μg).
C, Correlation between the effect of the γ7 and its various C-terminal truncated constructs on the level of CaV2.2 protein with their effect on CaV2.2 IBa shown in Fig. 1C. The effect of γ7, γ7(1-217), γ7(1-271) and γ7(1-238) on CaV2.2 protein levels represents the mean inhibition observed in 3-10 experiments. The linear fit has a correlation coefficient, r, of 0.936. All error bars are s.e.m.
γ7 markedly reduces the stability of CaV2.2 mRNA
Several mechanisms could underlie the suppression of CaV2.2 currents and CaV2.2 protein by γ7, including suppression of channel translation, more rapid protein degradation, suppression of channel transcription or increased mRNA breakdown. We addressed this question by examining the effect of γ7 on the level and stability of CaV2.2 mRNA. We found that co-expression of γ7 reduced CaV2.2 mRNA levels in two expression systems examined. In Cos-7 cells there was a 64 % reduction in CaV2.2 mRNA level in the presence of γ7, 48 h after transfection (Supplemental Fig. 3). In agreement with this, when CaV2.2 and γ7 were co expressed in individual Xenopus oocytes, there was a 66 % reduction in CaV2.2 mRNA expression after 24 h (Fig. 3A, see time 0) when compared to control. We then examined whether this was due to an increase in the rate of degradation of CaV2.2 mRNA. The transcription inhibitor actinomycin D was applied 24 h after injection of the relevant cDNAs into Xenopus oocytes (at time T0), and mRNA levels were then determined in individual oocytes at times up to 24 h thereafter. The half-life of CaV2.2 mRNA was 7.8 ± 1.5 h under control conditions (in agreement with Schorge et al. (1999)), and it showed more than a two-fold decrease to 3.5 ± 1.1 h in the presence of γ7 (Fig. 3A, B). Importantly, the truncated γ7(1-217) construct, lacking the C-terminus, had no effect on CaV2.2 mRNA degradation rate (Fig. 3A, B), in agreement with its lack of effect on CaV2.2 currents or CaV2.2 protein (Figs. 1 and 2).
Figure 3. Effect of γ7 and C-terminally truncated γ7(1-217) on CaV2.2 mRNA stability.
A, Effect of γ7 on CaV2.2 mRNA degradation rate. Constructs were expressed in Xenopus oocytes either without (■) or with γ7 (Δ) or with γ7(1-217) (□). In the absence of a γ7 construct, an equivalent amount of a similar sized transcript, a non-functional K+ channel Kir-AAA cDNA was used. After 24 h (T0), actinomycin D (50 μg/ml) was added to the medium, and the CaV2.2 mRNA levels measured at the times shown after this. The numbers of determinations from individual oocytes are between 6 and 12 for each data point; * P<0.05 compared to CaV2.2 (Student’s t test). The lines are apparent linear fits using errors as weight.
B, Bar chart showing the % of CaV2.2, α2δ-2 and KCC1 mRNA present at time t after actinomycin D addition at time T0. For CaV2.2 mRNA turnover, t = 9 h. Bars 1 - 3 are control (black bar, n = 11); + γ7, (white bar, n = 7) and +γ7(1-217) (hatched bar, n = 8). Bars 4 -7 show the % of α2δ-2 mRNA and KCC1 mRNA present 9 and 24 h respectively after actinomycin D addition for the control condition (black bar, n = 8 and 9), + γ7 (white bar, n = 9 and 9). These times were chosen as the % of mRNA remaining in control conditions was similar for all mRNA species. * P<0.05, compared to control, Student’s t test.
C, Degradation rate for endogenous CaV2.2 in the γ7-CFP PC12 cell line (○, n = 3), compared to control pcDNA3.1-transfected PC12 cell line (■, n = 5), following differentiation with NGF. The CaV2.2 mRNA level is expressed as % of time 0 (T0), when actinomycin D was added. The half-life for endogenous CaV2.2 mRNA was 19.5 h in the pcDNA3.1 transfected PC12 cell line. In γ7-transfected PC12 cells the half-life was 9.1 h (** P<0.01, Student’s t test). Insert: Reduction of endogenous CaV2.2 mRNA level in γ7-CFP PC12 cell line, compared to control pcDNA3.1-transfected PC12 cell line. *** P<0.001 vs control.
The relative mRNA concentrations at 24 h in the absence and presence of γ7 are in agreement with an effect only on the degradation rate of CaV2.2 mRNA, and not on its synthesis rate. Fitting the data to the equation d[RNA]/dt = kf - kd*[RNA], where [RNA] is the concentration of RNA at time t and kf and kd are the dominant rate constants for RNA synthesis and degradation, respectively, then [RNA] at time t = (kf/kd)(1-exp(-kd*t)). From the calculated values of kd, kf = 10.1 %/h and 8.8 %/h in the absence and presence of γ7, respectively, relative to the control CaV2.2 mRNA level at 24 h, immediately before actinomycin D treatment. Thus the data provides no evidence of a marked effect of γ7 on the synthesis rate (kf) of CaV2.2 mRNA, despite an over 2-fold increase in the measured degradation rate.
The effect of γ7 on mRNA stability does not affect all calcium channel subunits, as the degradation rate of α2δ-2 mRNA was not affected by γ7 co-expression (half-life ~25 h, Fig. 3B, Supplemental Fig. 4A). In agreement with this, the level of α2δ-2 protein was little affected by γ7 (14.9 ± 2.8 % reduction, n=6, Supplemental Fig. 4B).
We also investigated the effect of γ7 on the stability of KCC1 mRNA (a potassium chloride co-transporter), since it is a membrane protein of similar mass to CaV2.2, whose mRNA has some common sequence motifs (see below). Although KCC1 mRNA was much more stable than that of CaV2.2 (half life 50.5 h), γ7 decreased its half-life to 18.3 h, thus increasing its mRNA turnover from 28 ± 11% to 63 ± 5 % in 24 h (n ≥ 9) (Fig. 3B). Using the method described above, again we found no evidence for any effect on the synthesis rate. The kf was calculated to be 5.0 %/h and 4.1 %/h in the absence and presence of γ7, respectively, relative to the control KCC1 mRNA level at 24 h. These results indicate that the effect of γ7 on mRNA degradation rate is selective for certain mRNAs, but not selective for CaVα1 mRNAs.
In addition, we demonstrated that γ7 had the same effect on endogenous CaV2.2 mRNA, since 9 h after actinomycin D treatment, the CaV2.2 mRNA level in a PC12 cell line stably transfected with γ7-CFP was reduced by 74 % compared to a control PC12 cell line stably transfected with pcDNA3.1 (Fig. 3C, insert), and the endogenous CaV2.2 mRNA turnover rate was correspondingly enhanced (Fig. 3C). The half-life of CaV2.2 mRNA was 19.5 h in the pcDNA3.1 transfected PC12 cell line, which was not significantly different from that in control PC12 cells (16.9 h, data not shown). In γ7-transfected PC12 cells the half-life was 9.1 h (Fig. 3C).
We found no evidence that the effect of γ7 involves the induction of endoplasmic reticulum (ER) stress or the unfolded protein response, although this has been postulated as a mechanism of action of TARP γ2 (Sandoval et al., 2007). Co-expression of CaV2.2 with γ7 in tsA 201 cells, at a high transfection efficiency using Amaxa nucleofection, did not increase the editing of X binding protein (XBP) or induce C/EBP homologous protein (CHOP) expression, both of which are markers of the induction of the unfolded protein response (Harding et al., 2002), whereas these responses were evoked by exposure of cells to tunicamycin or dithiothreitol, both of which are well-known activators of ER stress (DJC and ACD, unpublished results).
Knockdown of γ7 increases endogenous CaV2.2 mRNA level in PC12 cells
To study the physiological role of endogenous γ7, we chose to silence its expression using RNA interference. We first examined the presence and localization of native γ7 in PC12 cells. Immunocytochemical localization showed that endogenous γ7 is expressed in these cells (Fig. 4A and Supplemental Fig. 5). We therefore used this cell line to examine the effect of knock-down of γ7. We designed three shRNAs specifically complementary to sequences in either rat or human γ7 mRNA. To confirm the effectiveness of the γ7 shRNAs, we made a PC12 cell line over-expressing human γ7-CFP, and showed that 5 days after transfection, a mix of three shRNAs directed against human γ7 mRNA reduced the γ7-CFP protein level in this cell line, by 75% compared to control transfected cells (Fig. 4B, C). We also found the three corresponding rat γ7 shRNAs reduced endogenous γ7 mRNA levels in PC12 cells, either individually or when mixed, whereas the control Drosophila gnu shRNA did not (Fig. 4D). This would represent a larger reduction in individual transfected cells, taking into account the incomplete transfection efficiency (about 30 %). Furthermore, the native γ7 protein present endogenously in PC12 cells (Fig. 4A, E) was depleted by 70 ± 6 % in individual PC12 cells, following transfection with the rat γ7 shRNAs (Fig. 4E, F).
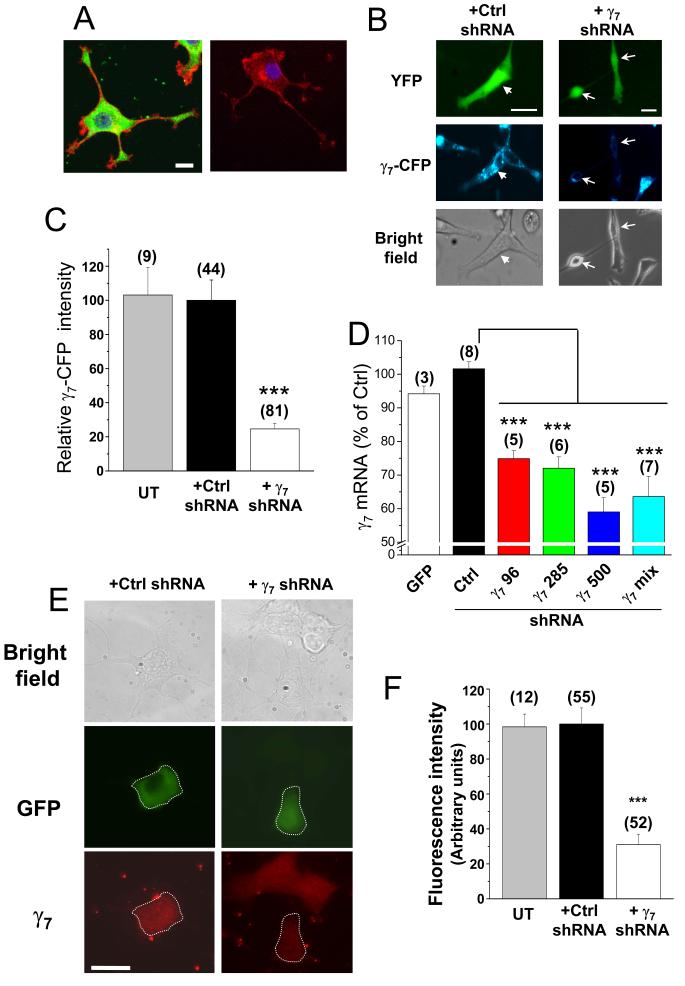
Figure 4. Short hairpin RNA constructs knockdown γ7 levels in PC12 cells.
A, Left: endogenous γ7 (detected using γ7 I-II loop Ab, green) in a differentiated PC12 cell, together with F-actin (Alexa Fluor 594-phalloidin, red) and nuclear staining (DAPI, blue). Right: control in the absence of primary γ7 Ab. Calibration bar 10 μm applies to both images. The images are a Z-stack of 5 to 8 confocal images.
B, Fluorescence microscopy of PC12 cells stably transfected with a human γ7-CFP construct. These cells were transiently transfected with YFP and either negative control gnu shRNA (Ctrl, left panel) or a mixture of three human γ7 shRNAs (γ7, right panel), and examined after 5 days. Upper row: transfected cells are identified with YFP fluorescence. Middle row: CFP fluorescence is almost abolished in cells transfected with γ7 shRNA. Arrowhead in left panel indicates a cell transfected with the control shRNA that shows γ7-CFP fluorescence. Arrows in right panel indicate two cells transfected with γ7 shRNA that do not show γ7-CFP fluorescence. Lower row: bright field. Scale bars 30 μm.
C, Bar chart of the mean percentage of CFP fluorescence in PC12 γ7-CFP cells after transfection with γ7 shRNA (white bar) compared to transfection with Ctrl shRNA (black bar) or untransfected PC12 cells (UT, gray bar). *** P<0.001 vs Ctrl.
D, Effect of rat γ7 shRNA on γ7 mRNA level in PC12 cells. γ7 mRNA was quantified by q-PCR in PC12 cells transfected with GFP (white bar), negative control shRNA (Ctrl, black bar) or human γ7 shRNAs: γ7 96 (red bar), γ7 285 (green bar), γ7 500 (blue bar) or γ7 mix (cyan bar). *** P<0.001 vs Ctrl.
E, Effect of transfection with γ7 shRNA on endogenous γ7 protein levels in PC12 cells. Left panel: cells transfected with control shRNA and GFP, right panel: cells transfected with rat γ7 shRNA and GFP. Top row: bright-field image; middle row: GFP, with transfected cell outlined; bottom row: endogenous γ7 visualized by immunocytochemistry (red), showing reduced fluorescence in γ7 shRNA-transfected cell (outlined). Scale bar 20 μm applies to all images, which were taken on a conventional fluorescence microscope.
F, Quantification of data including that given in E, showing reduction in mean fluorescence intensity (measured as fluorescence density in arbitrary units) in γ7 shRNA transfected cells (white bar), compared to control shRNA (Ctrl, black bar) or untransfected cells (UT, gray bar); *** P<0.001, compared to Ctrl.
If γ7 is playing a physiological role in controlling mRNA stability, its knockdown might be expected to have an effect opposite to γ7 over-expression, to increase the endogenous CaV2.2 mRNA level and enhance its stability. This was indeed the case, as CaV2.2 mRNA levels were increased by about 30 % following rat γ7 shRNA transfection (Fig. 5A), which would represent a larger enhancement in individual transfected cells, taking into account the transfection efficiency. An expected physiological correlate of this would be an increase in calcium channel currents in differentiated PC12 cells. In agreement with this, we recorded a 33.1 ± 4.1% (n = 48) increase in peak somatic calcium channel current density in PC12 cells transfected with rat γ7 shRNAs over the same time scale (Fig. 5B).
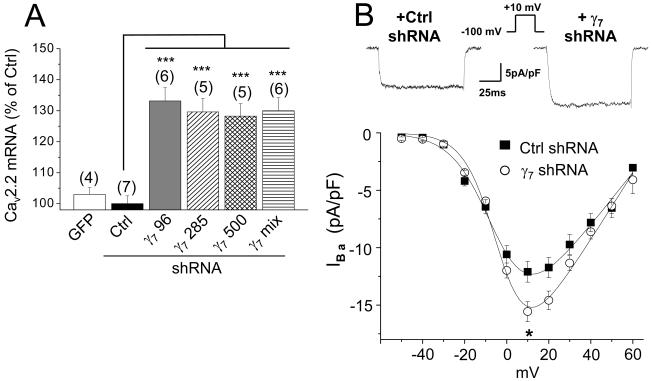
Figure 5. Short hairpin RNA constructs enhance endogenous CaV2.2 mRNA levels and calcium channel currents in PC12 cells.
A, Effect of rat γ7 shRNA on endogenous Cav2.2 mRNA level in PC12 cells. Cav2.2 mRNA was quantified by q-PCR in PC12 cells transfected with GFP (white bar), negative control shRNA (Ctrl, black bar) or γ7 shRNAs: γ7 96 (gray bar), γ7 285 (hatched bar), γ7 500 (cross-hatched bar) or γ7 mix (striped bar). *** P<0.001 vs Ctrl.
B, Upper panel: representative traces of peak calcium channel currents recorded from differentiated PC12 cells transfected with Ctrl shRNA (left panel) and γ7 shRNA (right panel). Currents were elicited by a 100 ms depolarization step to +10 mV from a holding potential of -100 mV. Lower panel: Current-voltage relationships for the two conditions (■, Ctrl shRNA; ○ γ7 shRNA). The charge carrier was 10 mM Ba2+. The knockdown of γ7 induces an increase of the peak current (-16.1 ± 0.9 pA/pF, n = 48) compared with control (-12.5 ± 0.9 pA/pF, n = 17, P<0.05).
γ7 exists in a complex with RNA binding proteins
Since PC12 cells contain endogenous γ7, they represent a suitable model cell type in which to search for protein complexes containing γ7. For this study we used a PC12 cell line stably expressing γ7 with a C-terminal HA tag. Following immunoprecipitation of γ7-HA from the γ7-HA PC12 cell line, and extensive washing, the presence of co-immunoprecipitating proteins, representing proteins in a complex with γ7, was examined by SDS-PAGE (Fig. 6A). Bands of interest, which were present in the γ7-HA immunoprecipitate, but absent from the control, non-transfected PC12 cells (Fig. 6A, left, arrows, representative of 3 experiments), were excised and identified, following tryptic digestion, by peptide mass fingerprinting. The bands labeled (1, 36 kDa) and (2, 37 kDa) were both identified, with 34 and 38 % peptide coverage, respectively, to be heterogeneous ribonuclear protein A2 (hnRNP A2), which has a number of splice variants of 33 - 38 kDa (Hatfield et al., 2002). Band 2 also contained hnRNP A3 (40 % coverage), which shares extensive sequence homology and is found in a complex with hnRNP A2 (Ma et al., 2002). The presence of γ7-HA was confirmed by immunoblotting (Fig. 6A, right). Both hnRNP A2 and hnRNP A3 are members of the hnRNP A/B sub-family.
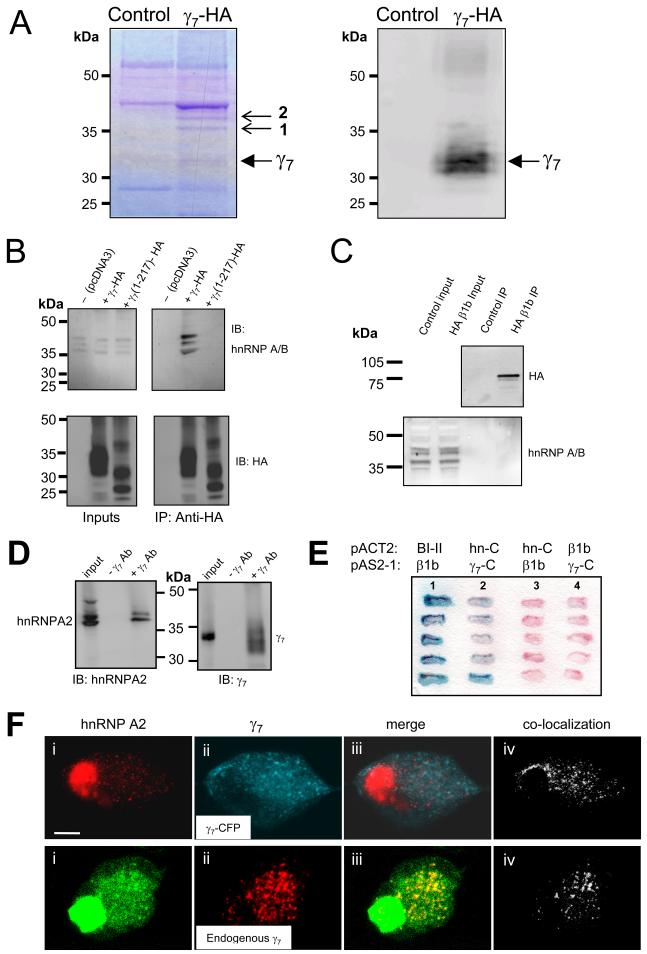
Figure 6. Identification of proteins interacting in a complex with γ7.
A, Proteins co-immunoprecipitated with γ7-HA from stably transfected PC12 cells. Left: Coomassie staining. Right: western blotting and immunodetection with anti-HA Ab of immunoprecipitated and control samples separated by SDS-PAGE. Solid arrow indicates γ7-HA. Small arrows indicate proteins co-immunoprecipitated with γ7-HA. Bands 1 and 2 were identified by peptide mass fingerprinting to contain hnRNP A2. Band 2 also contained hnRNP-A3. The control lane is untransfected PC12 cells. Position of molecular weight markers is shown on the left. Representative of three experiments.
B, Endogenous hnRNP A2 co-immunoprecipitates with transiently transfected γ7-HA, but not with γ7(1-217)-HA in tsA-201 cells. Western blotting and immunodetection with anti-hnRNP A/B Ab (H-200, upper row) and anti-HA Ab (lower row) of input (left) and HA-immunoprecipitated samples (following 500 mM NaCl wash, right) separated by SDS-PAGE. Position of molecular weight markers is shown on the left. Blots are representative of 3 - 5 independent experiments, using anti-hnRNP A/B Abs from two different sources.
C, CaVβ1b with a C-terminal HA tag was expressed and immunoprecipitated as described for γ7-HA. Its presence in the precipitate is confirmed in the upper blot. The control lane is from cells not transfected with CaVβ1b-HA. Immunoblotting for endogenous hnRNP A2 shows it is present in the input lanes, but absent from the immunoprecipitate.
D, Co-immunoprecipitation of endogenous hnRNP A2 and γ7 proteins from PC12 cells. Western blotting and immunodetection with anti-hnRNP A2 Ab (EF67, left panel) and γ7 C-terminus Ab (right panel) of input (left-hand lane of both panels) and γ7 C-terminus Ab-immunoprecipitated samples (right-hand lane of both panels) separated by SDS-PAGE. Control immunoprecipitations where γ7 Ab was omitted are shown in the center lane of each panel. Position of molecular weight markers is indicated between the panels. Blots are representative of 2 independent experiments.
E, The interaction of the C-terminus of hnRNP A2 with the cytoplasmic C-terminal tail of γ7 was independently confirmed by yeast co-transformation tests. Lane 1: positive control (blue reaction product) showing CaV2.2 I-II loop (BI-II) pACT2 and β1b pAS2-1. Lane 2: interaction between hnRNP A2 C-terminus (hn-C) in pACT2 and γ7 C-terminus (γ7-C) in pAS2-1. Lanes 3 and 4: negative controls showing that the hnRNP A2 C-terminus and the γ7 C-terminus do not interact with β1b in the alternative vector. The filter was incubated with the X-gal (5-bromo-4-chloro-3-indolyl-b-D-galactopyranoside) substrate for 5h.
F, Regions of co-localization of hnRNP A2 with γ7-CFP (upper row), or endogenous γ7 (lower row) in SCG neurons. Panel i: immunolocalization of endogenous hnRNP A2 in SCG neuron cell bodies (upper panel, red; lower panel, green). Panel ii: localization of γ7-CFP (upper panel, blue) or immunolocalization of endogenous γ7 (lower panel, red). Panel iii: merger of images of i and ii, with the co-localized regions shown in white (upper panel) or yellow (lower panel). Panel iv: co-localized γ7 and hnRNP A2 pixels are also shown separately for clarity (white). Scale bar 10 μm applies to all images. No endogenous γ7 staining was observed in the absence of primary Ab (data not shown).
We then confirmed that immunoprecipitation of γ7-HA was able to pull-down endogenous hnRNP A/B proteins in another system, tsA-201 cells transiently transfected with γ7-HA (Fig. 6B). Three bands were detected (36 - 42 kDa), using hnRNP A/B Abs, which, according to the antibody specificity and molecular weights are likely to represent one or more of hnRNP A1, A2 and A3, all of which also have splice variants. No hnRNP A/B immunoreactivity was immunoprecipitated by γ7(1-217)-HA, under the same conditions (Fig. 6B), indicating that these hnRNPs utilize the C-terminus of γ7 for this interaction. This is in agreement with our previous finding that the C-terminus of γ7 is essential for its functional effects. No hnRNP A/B immunoreactive proteins were co-immunoprecipitated with an unrelated HA-tagged protein, HA-CaVβ1b (Fig. 6C), indicating that the HA tag is not responsible for this binding.
We also found that pull-down of endogenous γ7 from untransfected PC12 cells, with the γ7 C-terminal Ab was able to immunoprecipitate endogenous hnRNP A2 (Fig. 6D).
The C-terminus of hnRNP A2 interacts directly with the C terminus of γ7
To determine whether the interaction between γ7 and hnRNP A2 was direct, we employed the yeast two-hybrid technique. When the γ7 C-terminus (residues 201-275) was used as bait, we found an interaction with the C-terminus of hnRNP A2 (residues 178-342, Fig. 6E, lane 2). Because of our previous work in this area, we used the high affinity (~10 nM, Leroy et al., 2005) interaction between the I-II linker of CaV2.2 and the CaVβ1b subunit as a positive control (Fig. 6E, lane 1). According to the relative time taken for colonies to turn blue in a colony-lift filter assay, the interaction between hnRNP A2 and γ7 C-terminus is weaker than that between CaV2.2 and CaVβ1b (data not shown). As negative controls, we showed that there was no interaction between the C-termini of either hnRNP A2 or γ7 and CaVβ1b (Fig. 6E, lanes 3 and 4). Furthermore, the N-terminus of hnRNP A2 (amino acids 1-86) was found not to interact with the C-terminus of γ7 (data not shown). All these data indicate that γ7 is likely to be present in a complex with hnRNP A2. Nevertheless, we cannot rule out that hnRNP A3, which shows marked homology to hnRNP A2, also interacts with γ7, either directly or by binding to hnRNP A2.
hnRNP A2 co-localizes with γ7 in neuronal cytoplasm
We used immunocytochemistry to examine whether γ7 and hnRNP A2 are co-localized in SCG neurons. Although, as expected, most hnRNP A2 is localized in the nucleus, it is also observed in small cytoplasmic granules in SCG neurons (Fig. 6F, i). In the somatic cytoplasm there are clear regions of co-localization of both transfected γ7-CFP and endogenous γ7 with hnRNP A2 (Fig. 6F ii and iii). This is in agreement with our suggestion that γ7 may sequester free hnRNP A2, as outlined in Fig. 9A, B.
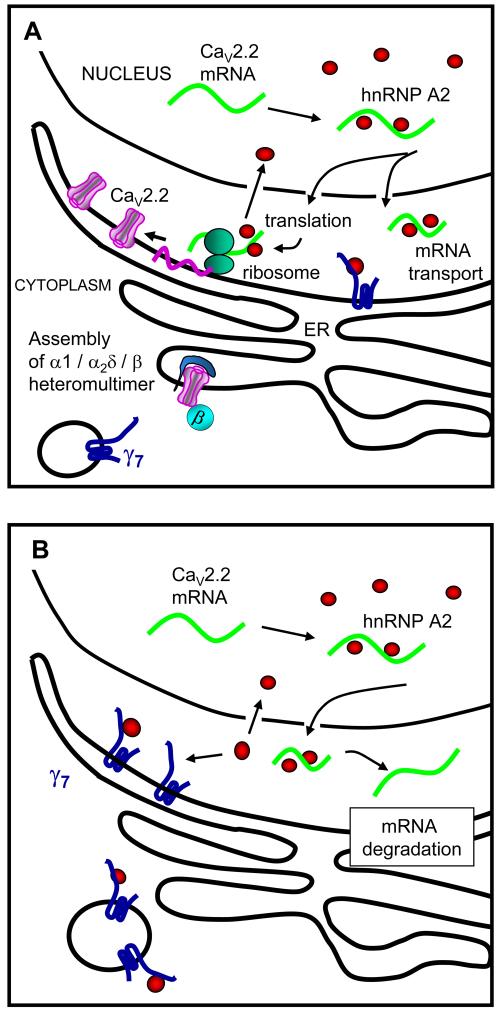
Figure 9. Diagram of proposed function of γ7.
A, The physiological distribution of hnRNP A2 and γ7. The hnRNP A2 (red circles) is localized largely in the nucleus where it binds to A2RE sequences on mRNAs and exits to the cytoplasm with these mRNAs, including that of CaV2.2 (green). This is thought to stabilize certain mRNAs, reducing degradation and therefore enhancing expression. Our results suggest this may be the case for CaV2.2. hnRNP A2 may also stabilize the mRNAs for transport. γ7 (dark blue) is present on the ER and is also associated with motile vesicles.
B, Following overexpression of γ7, it may sequester cytoplasmic hnRNP A2 and therefore increase degradation of CaV2.2 mRNA.
CaV2.2 mRNA binds to hnRNP A2
The hnRNP A2 proteins are highly expressed in brain, and are primarily present in the nucleus, where they bind to particular nucleotide sequences in mRNA. The combination of mRNA with its many bound proteins, including hnRNP A2, then exits the nucleus as ribonucleoprotein particles (Pinol-Roma, 1997). The interaction with hnRNP A2 has been found to stabilize certain mRNAs for trafficking to a remote site of protein synthesis (Hoek et al., 1998), and also to enhance translation (Kwon et al., 1999). In neurons and oligodendrocytes, hnRNP A2 has been implicated in the transport of mRNAs containing an A2 response element (A2RE) sequence, for local protein synthesis (Hoek et al., 1998; Shyu and Wilkinson, 2000).
We therefore examined whether CaV2.2 mRNA could be present in a complex with hnRNP A2. We found that the CaV2.2 mRNA used in this study contains at least two highly conserved sequences predicted to bind hnRNP A2 (Table 1), from the consensus sequence identified in MBP mRNA (Ainger et al., 1997). These two sequences are in constitutive exons, present in all splice variants of CaV2.2. The sequences preserve A and G at positions 8 and 9, previously shown to be essential for hnRNP A2 binding and function (Ainger et al., 1997; Shan et al., 2003), and the sequences are highly conserved in CaV2.2 between species (Table 1), with related sequences being found in CaV2.1 (data not shown). We found that a biotinylated 21 base ribonucleotide corresponding to sequence 1 from CaV2.2 (Table 1) binds endogenous hnRNP A2 from a mouse brain lysate, to a similar extent to the sequence originally described from MBP (Hoek et al., 1998) (Fig. 7A). We further found that an anti-HA Ab co-immunoprecipitated N-terminally HA-tagged hnRNP A2, both with over-expressed full-length rabbit CaV2.2 mRNA in tsA 201 cells (data not shown), and also with endogenous CaV2.2 mRNA in PC12 cells (Fig. 7B). Furthermore an anti-hnRNP A2 monoclonal Ab co-immunoprecipitated endogenous hnRNP A2 together with endogenous CaV2.2 mRNA from PC12 cells (Fig. 7C).
Table 1. hnRNP A2 mRNA binding motifs in CaV2.2 mRNA.
| sequence | similarity to consensus sequence in MBP | Gene | |
|---|---|---|---|
| MBP A2RE | GCCAAGGAGCCAGAGAGCAUG | 21 bases = A2RE | |
| (1) CaV2.2 rabbit mouse human |
GCUAAGGAGCGCGAGAGAGUG GCCAAGGAGCGGGAGCGAGUC GCCAAGGAGCGAGAGAGGGUG |
16/21 identical 15/21 identical 18/21 identical |
Cacna1b Exon 9 |
| (2) CaV2.2 rabbit mouse human |
UCCAAGGAGCUGGAGCGTGAC UCCAAGGAGCUGGAGAGGGAC UCCAAGGAGCUGGAGAGGGAC |
13/21 identical 14/21 identical 14/21 identical |
Cacna1b Exon 27 |
The binding motif identified in MBP (Hoek et al., 1998) is compared to the two found in CaV2.2. The essential conserved AG bases are in bold and the sequences identical to MBP A2RE are underlined. Exon numbering as shown in ENSEMBL.
Figure 7. Binding of hnRNP A2 to CaV2.2 mRNA and effect of hnRNP A2 and γ7 on CaV2.2 currents.
A, Evidence that one of the two potential A2RE sites in CaV2.2 mRNA binds to hnRNP A2. Endogenous hnRNP A2 (indicated by the arrow) from mouse brain lysate was pulled down by biotinylated oligonucleotides containing the consensus A2RE site (lane 2) and the potential site in CaV2.2 (lane 3) but not by beads alone (lane 1) or by a non-specific sequence with the same composition as the consensus A2RE site (lane 4). The hnRNP A2 was detected by immunoblotting with anti-hnRNP A2 Ab (F16). Position of molecular weight markers is shown on the left. Representative of two experiments.
B, Coimmunoprecipitation of endogenous CaV2.2 mRNA associated with hnRNP A2 in PC12 cells. Two days after transfection with HA-hnRNP A2 (HA-A2, lanes 1 and 3) or hnRNP A2 (A2, lanes 2 and 4), hnRNP A2 protein was immunoprecipitated from the whole cell lysate (lanes 1 and 2) with HA antibody (lanes 3 and 4). Immunoblots show that HA Ab pulled-down hnRNP A2 (top row, anti-hnRNP A2 (EF-67 Ab); middle row, anti-HA). Co-immunoprecipitated RNA was extracted, reverse transcribed and amplified by PCR. A specific PCR product corresponding to the endogenous CaV2.2 mRNA was amplified (35 cycles) only in the condition where hnRNP A2 protein was pulled-down (bottom row, lane 3).
C, Left panel: Endogenous hnRNP A2 proteins were immunoprecipitated from PC12 cells with anti-hnRNP A2 Ab EF-67 (upper row, lane 2). In this condition, a specific PCR product corresponding to the endogenous CaV2.2 mRNA was also amplified (35 cycles, lower row). CaV2.2 mRNA was not detected in the control where antibody was omitted (lane 3). Right panel: The immunoprecipitation was repeated, including an acetone precipitation step, to confirm the presence of endogenous hnRNP A2.
D, Enhancement by hnRNP A2 but not hnRNP A1 of CaV2.2 currents recorded from Xenopus ooctyes. Left panel: example traces elicited by 50 ms step depolarizations to between -40 and 0 mV in 10 mV steps, from a holding potential of -100 mV for CaV2.2/β1b/α2δ-2 (upper traces) or CaV2.2/β1b/α2δ-2 plus hnRNP A2 (lower traces). The charge carrier was 10 mM Ba2+. Right panel: bar chart shows the effect of hnRNP A2 co-expression with CaV2.2/β1b/α2δ-2 on peak CaV2.2 IBa amplitude, expressed relative to the mean peak control current in each experiment. Control (black bar, n = 18), for both groups taken from experiments directly comparing hnRNP A1 and hnRNP A2: + hnRNP A2 (white bar, n = 21) and + hnRNP A1 (hatched bar, n = 19). These data are pooled from two separate experiments showing similar results. Statistical significance * P<0.05, ** P<0.01 (one way ANOVA and Bonferroni’s post test).
E, Inhibition of the hnRNP A2-mediated enhancement of IBa in Xenopus oocytes by a low concentration of γ7. Bar chart compares the effect of two concentrations of γ7 cDNA (γ7: CaV2.2 ratio 1 and 0.25), and also shows the lack of enhancement by hnRNP A2 on peak CaV2.2 IBa amplitude, when co-expressed with the lower concentration of γ7 (γ7: CaV2.2 ratio 0.25). Data are expressed as a % of the mean peak control CaV2.2 current in each experiment, and all data were recorded 2 days after cDNA injection. Control in the absence of γ7 or hnRNP A2 (white bars); + γ7 (ratio 1; hatched bar), + γ7 (ratio 0.25; cross-hatched bar) + hnRNP A2 (black bar, from data obtained in the same experiments), + hnRNP A2 and γ7 (ratio 0.25) (gray bar). Number of determinations given above each bar. These data are pooled from 5 different batches of oocytes, in all of which similar results were observed. Statistical significance ** P<0.01 (one way ANOVA and Bonferroni’s post test).
hnRNP A2 enhances the expression of CaV2.2 currents, and this is counteracted by γ7
Since hnRNP A2 binds to CaV2.2 mRNA, we wished to examine the consequences on calcium channel expression of co-expression with hnRNP A2. When this construct was co-expressed with CaV2.2/β1b/α2δ-2 in Xenopus oocytes, it produced a significant enhancement of the peak calcium current amplitude, without affecting the current kinetics or voltage-dependence (Fig. 7D). In experiments performed in parallel, an unrelated hnRNP (A1), that is involved in mRNA processing and export but does not bind to A2RE sequences (Cullen, 2000), did not enhance CaV2.2 calcium currents (Fig. 7D). Thus hnRNP A2 may enhance CaV2.2 mRNA stability in Xenopus oocytes or increase its transport to sites of translation. This is compatible with the role previously attributed to hnRNP A2, that it enhances both transport and translation of specific mRNAs (Kwon et al., 1999). We also found that the effect of hnRNP A2 was able to generalize to other CaV2 channels, as enhancement of CaV2.1/β4/α2δ-2 currents was also observed, the peak IBa at +5 mV being increased to 194.8 ± 19.9 % of control (n = 28, P = 0.0004). CaV2.1 mRNA contains A2RE sequences that are homologous to those of CaV2.2 (data not shown).
We then investigated whether there was an interaction between the effects of hnRNP A2 and γ7 on calcium channel current expression. The inhibitory effect of γ7 on CaV2.2 calcium channel currents, shown in Fig. 1B, was found to be concentration-dependent (data not shown), and a low concentration of γ7 was chosen (1:4 dilution of γ7 cDNA), that did not inhibit CaV2.2 currents in the Xenopus oocyte expression system (Fig. 7E), in order to examine its interaction with the effect of hnRNP A2. We found that the enhancement of CaV2.2 currents by hnRNP A2 was prevented by co-expression of this low concentration of γ7 (Fig. 7E). This indicates that γ7 opposes the effect of hnRNP A2, supporting the hypothesis that the binding of hnRNP A2 to its binding site on the C-terminus of γ7 reduces the amount available to interact with CaV2.2 mRNA in the cytosol. In agreement with this interpretation, expression of the cytosolic C-terminus of γ7 (γ7 (201-275)), which our yeast two hybrid data shows to be able to bind hnRNP A2 (Fig. 6E), was itself able to reduce the amplitude of CaV2.2 currents. In experiments similar to those shown in Fig. 1B, the peak CaV2.2 IBa current amplitude was reduced by 70 .6 ± 4.5 % (n = 19) relative to control currents, in the presence of the C-terminus of γ7 (data not shown), compared to an 87.4 % reduction by full-length γ7 in the same experiment.
Investigation of the subcellular localization of γ7
The topology of γ1 and the TARP γ proteins indicates that the linker between the first and second transmembrane domains is extracellular (Chu et al., 2001) and the same might be anticipated for γ7, which has a predicted glycosylation site in this loop (see Fig. 1A). We obtained mutational evidence that γ7 is N-glycosylated on N45 in the I-II loop, supporting the proposed topology (Fig. 8A). However, heterologously expressed γ7 is not inserted in the plasma membrane of tsA 201 cells, since it was not detected with an Ab to this I-II loop in non-permeabilized cells (Fig. 8B).
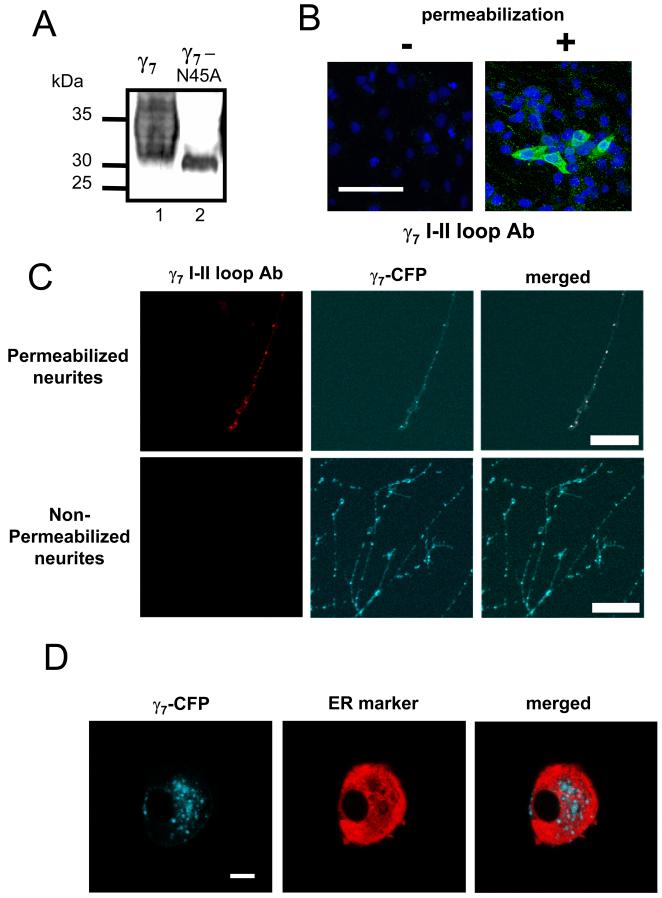
Figure 8. Subcellular localization of γ7.
A, Western blotting of a γ7 mutant in which the potential glycosylation site, N45 is mutated to A, and immuno-detection with anti-γ7 I-II loop Ab. Lane 1: γ7; lane 2: γ7N45A. The reduction in mass and sharpening of the band indicates that γ7 is normally glycosylated at this site. Positions of molecular weight markers are shown on the left. The data are representative of 2 independent experiments.
B, Immunodetection of transiently transfected γ7 (green) in non-permeabilized (left) and permeabilized (right) tsA 201 cells, using γ7 I-II loop Ab. The nuclear stain DAPI (blue) was used as a cell marker. Scale bar: 100 μm, applies to both images. No immunostaining was observed in any field in the absence of permeabilization.
C, Immunostaining for γ7 using I-II loop Ab (Texas red secondary Ab, left panel) co-localizes completely with γ7-CFP (middle panel), as shown by yellow regions in merged image (right), in permeabilized (upper row), but not non-permeabilized (lower row) SCG neurons. Scale bar: 40 μm, applies to all images.
D, Partial co-localization of γ7-CFP and an ER marker in SCG neurons. Left: γ7-CFP; center: ER marker (ER-DsRed), right: overlay showing co-localization of DsRed with γ7-CFP (white). Scale bar: 10 μm and applies to all images.
We have observed that γ7-CFP, when heterologously expressed in cultured SCG neurons, is present, in part, in motile intracellular vesicles (Fig. 8C and data not shown). These co-localize with γ7 I-II loop Ab immunoreactivity only when the cells are permeabilized (Fig. 8C). No plasma membrane staining was detected in the absence of cell permeabilization (Fig. 8C). We also found that neither endogenous γ7 (Fig. 4A), nor γ7-CFP (Fig. 4B and data not shown) localized to the plasma membrane in PC12 cells. The localization of γ7 is therefore on intracellular membranes in all the cells we have examined. We found that the distribution of γ7-CFP partially overlapped with that of an ER marker in the somata of SCG neurons microinjected with γ7-CFP and pDsRed2-ER (Fig. 8D). It was also found in vesicular structures, which were negative for the ER marker (Fig. 8D). Similar results were obtained in PC12 cells using a different ER marker, protein disulfide isomerase (data not shown).
Discussion
We have previously described the identification of two genes that encode γ5 and γ7, by their homology with the mouse stargazin gene (cacng2), and have cloned and expressed the cDNA for both human and mouse γ7 (Moss et al., 2002). The γ7 protein contains 275 amino acids with an estimated protein mass of 31 kDa, and has four predicted transmembrane-spanning domains with intracellular N- and C-termini and a consensus N-glycosylation sequence in the loop between the first two transmembrane segments (Fig. 1A). Together with the other γ-like proteins, it belongs to the claudin super-family, members of which play diverse roles in cellular physiology (Sanders et al., 2001). Initially the novel γ1-like proteins (γ2-8) were investigated as potential calcium channel subunits (Letts et al., 1998; Klugbauer et al., 2000; Sharp et al., 2001; Kang et al., 2001; Rousset et al., 2001; Moss et al., 2003). However, we have shown that while the human γ2 and γ4 TARPS are expressed in Purkinje neurons, there was no effect of these proteins on calcium currents composed of CaV2.1/β4/α2δ-2, a combination which mimics the major calcium channel complement in Purkinje cells (Moss et al., 2003). More recently these proteins have been shown to have roles in trafficking and localization of AMPA glutamate receptors (Tomita et al., 2003; Tomita et al., 2004; Fukata et al., 2005; Kato et al., 2007). Together with the data presented here, these findings suggest that γ-proteins may play diverse and possibly multiple roles in intracellular trafficking.
In our original study, we showed that co-expression with γ7 almost completely abolished Ba2+ current through N-type CaV2.2 channels expressed from cDNA in both Xenopus oocyte and Cos-7 cell expression systems (Moss et al., 2002). Several mechanisms could underlie the suppression of CaV2.2 currents by γ7 protein, but we have now identified that a major mechanism of suppression of CaV2.2 currents by γ7 results from a decrease in CaV2.2 mRNA stability. This process requires the cytoplasmic C-terminus of γ7, since all its effects, to inhibit CaV2.2 current and protein expression and to increase the rate of mRNA degradation, are completely prevented by the removal of most of the C-terminus of γ7. It is well known that the effect of over-expression of a protein does not necessarily reflect its physiological function, since an over-expressed protein may act to sequester interacting partners. However, our evidence indicates that the regulation of the stability of CaV2.2 mRNA, and potentially other mRNAs, represents an important role for native γ7. This evidence stems from the finding that knockdown of endogenous γ7 in PC12 cells, using γ7 shRNA, substantially enhances their endogenous CaV2.2 mRNA level and enhances endogenous somatic calcium currents following differentiation.
We have subsequently identified by co-immunoprecipitation from a PC12 cell line stably transfected with γ7-HA, that the RNA binding protein hnRNP A2 is co-immunoprecipitated with γ7. hnRNP A2 has been found to be involved in the stability, trafficking and localization of particular mRNAs and has been identified to bind to several different mRNA sequences including CGG repeats (Sofola et al., 2007), AU-rich elements (Hamilton et al., 1999), and a well-characterized binding motif termed A2RE (Ainger et al., 1997; Shan et al., 2003; Fahling et al., 2006). Most hnRNP A2 is localized in the nucleus, where it binds to specific sequences in transcribed mRNA, and is involved in mRNA export into the cytoplasm (for review see Shyu and Wilkinson, 2000), and subsequently in trafficking and enhancement of translation for those mRNAs containing an A2RE sequence (Kwon et al., 1999). Our yeast two-hybrid data indicates that γ7 C-terminus binds directly to hnRNP A2, and our subcellular localization data for γ7 indicates that this interaction will occur on intracellular membranes within the cytoplasm (as depicted in Fig. 9B).
There are two highly conserved consensus hnRNP A2-binding A2RE motifs in the CaV2.2 mRNA used in this study (Table 1), that contain the conserved A and G at positions 8 and 9, which have been found to be a prerequisite for binding of hnRNP A2 (Ainger et al., 1997). There are also several other, less well-conserved, A2RE motifs in the CaV2.2 mRNA sequence, that may also be functional (data not shown). Here we have demonstrated that hnRNP A2 binds to one of the conserved sequences (Table 1, sequence 2), and by homology is highly likely to bind to the other sequence. Furthermore hnRNP A2 co-immunoprecipitates endogenous CaV2.2 mRNA from PC12 cells. We suggest that hnRNP A2 normally binds to sequences on CaV2.2 mRNA in the nucleus and when the ribonucleoprotein particle so formed exits the nucleus, it is transported to its site of translation, and also protected from degradation (as depicted in Fig. 9A). A similar mechanism of interaction with hnRNP A2 has been proposed for MBP mRNA which contains a canonical A2RE sequence (Hoek et al., 1998). It has also been shown that hnRNP A2 itself is subject to transport (Brumwell et al., 2002). Indeed, the hnRNP-A/B proteins have been identified as components in isolated RNA-transporting granules (Kanai et al., 2004). Furthermore, the Drosophila homolog Hrp48 is involved in the localization of oskar mRNA (Yano et al., 2004; Huynh et al., 2004). Other RNA binding proteins, such as Staufen have been shown to have an effect on both localization and decay of specific mRNAs (Broadus et al., 1998; Kim et al., 2005).
Our results provoke the hypothesis that γ7 is involved in the regulation of hnRNP A2 function, as depicted in Fig. 9B. γ7 may sequester hnRNP A2, and this may be the mechanism whereby the stability of specific mRNAs, including that of CaV2.2, are compromised. Furthermore, our results indicate that native, as well as heterologously expressed γ7 both influence the physiological regulation of CaV2.2 mRNA stability, since knock-down of native γ7 increased endogenous CaV2.2 mRNA and CaV2.2 current levels. This indicates that endogenous γ7 may function to limit the availability of cytoplasmic hnRNP A2. A related pathological mechanism of hnRNP A2 sequestration by binding to expanded CGG repeats of FMR1 mRNA has recently been proposed to occur in Fragile X-associated tremor/ataxia syndrome (Sofola et al., 2007). It will be of interest to examine whether a similar interaction occurs with other triplet repeat diseases.
The expression of CaV2.2 channel proteins may be particularly vulnerable to a reduction in available hnRNP A2 because the mRNA degradation rate of CaV2.2 is relatively high. Nevertheless, other mRNAs are also likely to be similarly affected, and we show that the degradation of KCC1 mRNA, which also contains an A2RE sequence, is also enhanced by γ7.
Both CaV2.2 (Mori et al., 1991) and γ7 (Moss et al., 2002) are selectively expressed in neurons. CaV2.2 channel expression and function is dynamically regulated both physiologically (Pravettoni et al., 2000; Inchauspe et al., 2004), and in pathology (Hendriksen et al., 1997). It is of interest that CaV2.2 channels are functionally most important in early development, and in many instances their role is later substituted by P/Q type calcium channels (CaV2.1), both in terms of somatic currents (Salgado et al., 2005) and in terms of synaptic transmission (Iwasaki et al., 2000). Over a similar time period we have found that γ7 expression is strongly up-regulated (M. Nieto-Rostro and ACD, unpublished results). It is now recognized from many studies that regulation of mRNA stability is a very important part of the post-transcriptional control of expression of numerous genes (Wilusz and Wilusz, 2004).
There are indications that CaV2.2 mRNA is subject to transport, as it has been identified in dendritic growth cones (Crino and Eberwine, 1996) and in processes of motor neurons (Jablonka et al., 2007). Furthermore, N-type calcium channels are localized in the presynaptic terminals of peripheral neurons, such as DRG neurons, and local synthesis of transmembrane proteins has recently been demonstrated in axons and axonal growth cones (Brittis et al., 2002). It will be of great interest in the future to examine whether hnRNP A2 affects the transport of CaV2 family mRNAs, to regions distinct from the cell soma.
Supplementary Material
Acknowledgements
We thank the Wellcome Trust, BBSRC and MRC for support. LF held a fellowship from Fondation pour la Recherche Medicale and J.L. held a Wellcome Trust International fellowship. DJC was supported by a BBSRC PhD studentship and DW by a British Heart Foundation PhD studentship. We are grateful to Dr. WJ Frith for mathematical advice, Kanchan Chaggar for technical assistance, Dr. R Nichols for the hnRNP A2 constructs, Dr J Caceres for hnRNPA1 cDNA, Dr. S Alper for KCC1 cDNA, Dr. TJ Shafer for PC12 cells and Drs. A Cahill and A. Fox for pG418-shRNA-Empty vector.
References
- Ainger K, Avossa D, Diana AS, Barry C, Barbarese E, Carson JH. Transport and localization elements in myelin basic protein mRNA. J Cell Biol. 1997;138:1077–1087. doi: 10.1083/jcb.138.5.1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Black JL, Lennon VA. Identification and cloning of putative human neuronal voltage-gated calcium channel gamma-2 and gamma-3 subunits: Neurologic implications. Mayo Clin Proc. 1999;74:357–361. doi: 10.4065/74.4.357. [DOI] [PubMed] [Google Scholar]
- Brice NL, Berrow NS, Campbell V, Page KM, Brickley K, Tedder I, Dolphin AC. Importance of the different β subunits in the membrane expression of the α1A and α2 calcium channel subunits: studies using a depolarisation-sensitive α1A antibody. Eur J Neurosci. 1997;9:749–759. doi: 10.1111/j.1460-9568.1997.tb01423.x. [DOI] [PubMed] [Google Scholar]
- Brittis PA, Lu Q, Flanagan JG. Axonal protein synthesis provides a mechanism for localized regulation at an intermediate target. Cell. 2002;110:223–235. doi: 10.1016/s0092-8674(02)00813-9. [DOI] [PubMed] [Google Scholar]
- Broadus J, Fuerstenberg S, Doe CQ. Staufen-dependent localization of prospero mRNA contributes to neuroblast daughter-cell fate. Nature. 1998;391:792–795. doi: 10.1038/35861. [DOI] [PubMed] [Google Scholar]
- Brodbeck J, Davies A, Courtney J-M, Meir A, Balaguero N, Canti C, Moss FJ, Page KM, Pratt WS, Hunt SP, Barclay J, Rees M, Dolphin AC. The ducky mutation in Cacna2d2 results in altered Purkinje cell morphology and is associated with the expression of a truncated a2d-2 protein with abnormal function. J Biol Chem. 2002;277:7684–7693. doi: 10.1074/jbc.M109404200. [DOI] [PubMed] [Google Scholar]
- Brummelkamp TR, Bernards R, Agami R. A system for stable expression of short interfering RNAs in mammalian cells. Science. 2002;296:550–553. doi: 10.1126/science.1068999. [DOI] [PubMed] [Google Scholar]
- Brumwell C, Antolik C, Carson JH, Barbarese E. Intracellular trafficking of hnRNP A2 in oligodendrocytes. Exp Cell Res. 2002;279:310–320. doi: 10.1006/excr.2002.5604. [DOI] [PubMed] [Google Scholar]
- Burgess DL, Davis CF, Gefrides LA, Noebels JL. Identification of three novel Ca2+ channel gamma subunit genes reveals molecular diversification by tandem and chromosome duplication. Genome Research. 1999;9:1204–1213. doi: 10.1101/gr.9.12.1204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burgess DL, Gefrides LA, Foreman PJ, Noebels JL. A cluster of three novel Ca2+ channel gamma subunit genes on chromosome 19q13.43: Evolution and expression profile of the gamma subunit gene family. Genomics. 2001;71:339–350. doi: 10.1006/geno.2000.6440. [DOI] [PubMed] [Google Scholar]
- Campbell V, Berrow N, Brickley K, Page K, Wade R, Dolphin AC. Voltage-dependent calcium channel β-subunits in combination with alpha1 subunits have a GTPase activating effect to promote hydrolysis of GTP by G alphao in rat frontal cortex. FEBS Lett. 1995;370:135–140. doi: 10.1016/0014-5793(95)00813-o. [DOI] [PubMed] [Google Scholar]
- Canti C, Davies A, Berrow NS, Butcher AJ, Page KM, Dolphin AC. Evidence for two concentration-dependent processes for β subunit effects on α1B calcium channels. Biophys J. 2001;81:1439–1451. doi: 10.1016/S0006-3495(01)75799-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Canti C, Page KM, Stephens GJ, Dolphin AC. Identification of residues in the N-terminus of α1B critical for inhibition of the voltage-dependent calcium channel by Gβγ. J Neurosci. 1999;19:6855–6864. doi: 10.1523/JNEUROSCI.19-16-06855.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Catterall WA. Structure and regulation of voltage-gated Ca2+ channels. Annu Rev Cell Dev Biol. 2000;16:521–555. doi: 10.1146/annurev.cellbio.16.1.521. [DOI] [PubMed] [Google Scholar]
- Chu PJ, Robertson HM, Best PM. Calcium channel gamma subunits provide insights into the evolution of this gene family. Gene. 2001;280:37–48. doi: 10.1016/s0378-1119(01)00738-7. [DOI] [PubMed] [Google Scholar]
- Crino PB, Eberwine J. Molecular characterization of the dendritic growth cone: regulated mRNA transport and local protein synthesis. Neuron. 1996;17:1173–1187. doi: 10.1016/s0896-6273(00)80248-2. [DOI] [PubMed] [Google Scholar]
- Cullen BR. Connections between the processing and nuclear export of mRNA: evidence for an export license? Proc Natl Acad Sci U S A. 2000;97:4–6. doi: 10.1073/pnas.97.1.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dolphin AC. G protein modulation of voltage-gated calcium channels. Pharmacol Rev. 2003a;55:607–627. doi: 10.1124/pr.55.4.3. [DOI] [PubMed] [Google Scholar]
- Dolphin AC. β subunits of voltage-gated calcium channels. J Bioeng Biomemb. 2003b;35:599–620. doi: 10.1023/b:jobb.0000008026.37790.5a. [DOI] [PubMed] [Google Scholar]
- Fahling M, Mrowka R, Steege A, Martinka P, Persson PB, Thiele BJ. hnRNP-A2/B1 modulate collagen prolyl 4-hydroxylase alpha (I) mRNA-stability. J Biol Chem. 2006 doi: 10.1074/jbc.M510925200. [DOI] [PubMed] [Google Scholar]
- Freise D, Held B, Wissenbach U, Pfeifer A, Trost C, Himmerkus N, Schweig U, Freichel M, Biel M, Hofmann F, Hoth M, Flockerzi V. Absence of the gamma subunit of the skeletal muscle dihydropyridine receptor increases L-type Ca2+ currents and alters channel inactivation properties. J Biol Chem. 2000;275:14476–14481. doi: 10.1074/jbc.275.19.14476. [DOI] [PubMed] [Google Scholar]
- Fukata Y, Tzingounis AV, Trinidad JC, Fukata M, Burlingame AL, Nicoll RA, Bredt DS. Molecular constituents of neuronal AMPA receptors. J Cell Biol. 2005;169:399–404. doi: 10.1083/jcb.200501121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamilton BJ, Nichols RC, Tsukamoto H, Boado RJ, Pardridge WM, Rigby WF. hnRNP A2 and hnRNP L bind the 3′UTR of glucose transporter 1 mRNA and exist as a complex in vivo. Biochem Biophys Res Commun. 1999;261:646–651. doi: 10.1006/bbrc.1999.1040. [DOI] [PubMed] [Google Scholar]
- Harding HP, Calfon M, Urano F, Novoa I, Ron D. Transcriptional and translational control in the Mammalian unfolded protein response. Annu Rev Cell Dev Biol. 2002;18:575–599. doi: 10.1146/annurev.cellbio.18.011402.160624. [DOI] [PubMed] [Google Scholar]
- Hatfield JT, Rothnagel JA, Smith R. Characterization of the mouse hnRNP A2/B1/B0 gene and identification of processed pseudogenes. Gene. 2002;295:33–42. doi: 10.1016/s0378-1119(02)00800-4. [DOI] [PubMed] [Google Scholar]
- Hendriksen H, Kamphuis W, Lopes da Silva FH. Changes in voltage-dependent calcium channel alpha1-subunit mRNA levels in the kindling model of epileptogenesis. Brain Res Mol Brain Res. 1997;50:257–266. doi: 10.1016/s0169-328x(97)00196-4. [DOI] [PubMed] [Google Scholar]
- Hoek KS, Kidd GJ, Carson JH, Smith R. hnRNP A2 selectively binds the cytoplasmic transport sequence of myelin basic protein mRNA. Biochemistry. 1998;37:7021–7029. doi: 10.1021/bi9800247. [DOI] [PubMed] [Google Scholar]
- Huynh JR, Munro TP, Smith-Litiere K, Lepesant JA, St Johnston D. The Drosophila hnRNPA/B homolog, Hrp48, is specifically required for a distinct step in osk mRNA localization. Dev Cell. 2004;6:625–635. doi: 10.1016/s1534-5807(04)00130-3. [DOI] [PubMed] [Google Scholar]
- Inchauspe CG, Martini FJ, Forsythe ID, Uchitel OD. Functional compensation of P/Q by N-type channels blocks short-term plasticity at the calyx of held presynaptic terminal. J Neurosci. 2004;24:10379–10383. doi: 10.1523/JNEUROSCI.2104-04.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iwasaki S, Momiyama A, Uchitel OD, Takahashi T. Developmental Changes in Calcium Channel Types Mediating Central Synaptic Transmission. J Neurosci. 2000;20:59–65. doi: 10.1523/JNEUROSCI.20-01-00059.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jablonka S, Beck M, Lechner BD, Mayer C, Sendtner M. Defective Ca2+ channel clustering in axon terminals disturbs excitability in motoneurons in spinal muscular atrophy. J Cell Biol. 2007;179:139–149. doi: 10.1083/jcb.200703187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jay SD, Sharp AH, Kahl SD, Vedvick TS, Harpold MM, Campbell KP. Structural characterization of the dihydropyridine-sensitive calcium channel α2-subunit and the associated δ peptides. J Biol Chem. 1991;266:3287–3293. [PubMed] [Google Scholar]
- Kanai Y, Dohmae N, Hirokawa N. Kinesin transports RNA: isolation and characterization of an RNA-transporting granule. Neuron. 2004;43:513–525. doi: 10.1016/j.neuron.2004.07.022. [DOI] [PubMed] [Google Scholar]
- Kang MG, Chen CC, Felix R, Letts VA, Frankel WN, Mori Y, Campbell KP. Biochemical and biophysical evidence for gamma2 subunit association with neuronal voltage-activated Ca2+ channels. J Biol Chem. 2001;276:32917–32924. doi: 10.1074/jbc.M100787200. [DOI] [PubMed] [Google Scholar]
- Kang MG, Chen CC, Wakamori M, Hara Y, Mori Y, Campbell KP. A functional AMPA receptor-calcium channel complex in the postsynaptic membrane. Proc Natl Acad Sci U S A. 2006;103:5561–5566. doi: 10.1073/pnas.0601289103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato AS, Zhou W, Milstein AD, Knierman MD, Siuda ER, Dotzlaf JE, Yu H, Hale JE, Nisenbaum ES, Nicoll RA, Bredt DS. New transmembrane AMPA receptor regulatory protein isoform, gamma-7, differentially regulates AMPA receptors. J Neurosci. 2007;27:4969–4977. doi: 10.1523/JNEUROSCI.5561-06.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim YK, Furic L, DesGroseillers L, Maquat LE. Mammalian Staufen1 recruits Upf1 to specific mRNA 3′UTRs so as to elicit mRNA decay. Cell. 2005;120:195–208. doi: 10.1016/j.cell.2004.11.050. [DOI] [PubMed] [Google Scholar]
- Klugbauer N, Dai SP, Specht V, Lacinová L, Marais E, Bohn G, Hofmann F. A family of gamma-like calcium channel subunits. FEBS Lett. 2000;470:189–197. doi: 10.1016/s0014-5793(00)01306-5. [DOI] [PubMed] [Google Scholar]
- Kwon S, Barbarese E, Carson JH. The cis-acting RNA trafficking signal from myelin basic protein mRNA and its cognate trans-acting ligand hnRNP A2 enhance cap-dependent translation. J Cell Biol. 1999;147:247–256. doi: 10.1083/jcb.147.2.247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leroy J, Richards MS, Butcher AJ, Nieto-Rostro M, Pratt WS, Davies A, Dolphin AC. Interaction via a key tryptophan in the I-II linker of N-type calcium channels is required for beta1 but not for palmitoylated beta2, implicating an additional binding site in the regulation of channel voltage-dependent properties. J Neurosci. 2005;25:6984–6996. doi: 10.1523/JNEUROSCI.1137-05.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Letts VA, Felix R, Biddlecome GH, Arikkath J, Mahaffey CL, Valenzuela A, Bartlett FS, Mori Y, Campbell KP, Frankel WN. The mouse stargazer gene encodes a neuronal Ca2+ channel gamma subunit. Nature Genet. 1998;19:340–347. doi: 10.1038/1228. [DOI] [PubMed] [Google Scholar]
- Lingor P, Koeberle P, Kugler S, Bahr M. Down-regulation of apoptosis mediators by RNAi inhibits axotomy-induced retinal ganglion cell death in vivo. Brain. 2005;128:550–558. doi: 10.1093/brain/awh382. [DOI] [PubMed] [Google Scholar]
- Ma AS, Moran-Jones K, Shan J, Munro TP, Snee MJ, Hoek KS, Smith R. Heterogeneous nuclear ribonucleoprotein A3, a novel RNA trafficking response element-binding protein. J Biol Chem. 2002;277:18010–18020. doi: 10.1074/jbc.M200050200. [DOI] [PubMed] [Google Scholar]
- Meir A, Bell DC, Stephens GJ, Page KM, Dolphin AC. Calcium channel β subunit promotes voltage-dependent modulation of α1B by Gβγ. Biophys J. 2000;79:731–746. doi: 10.1016/S0006-3495(00)76331-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mori Y, Friedrich T, Kim M-S, Mikami A, Nakai J, Ruth P, Bosse E, Hofmann F, Flockerzi V, Furuichi T, Mikoshiba K, Imoto K, Tanabe T, Numa S. Primary structure and functional expression from complementary DNA of a brain calcium channel. Nature. 1991;350:398–402. doi: 10.1038/350398a0. [DOI] [PubMed] [Google Scholar]
- Moss FJ, Dolphin AC, Clare JJ. Human neuronal stargazin-like proteins, gamma2, gamma3 and gamma4; an investigation of their specific localization in human brain and their influence on CaV2.1 voltage-dependent calcium channels expressed in Xenopus oocytes. BMC Neurosci. 2003;4:23. doi: 10.1186/1471-2202-4-23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss FJ, Viard P, Davies A, Bertaso F, Page KM, Graham A, Canti C, Plumpton M, Plumpton C, Clare JJ, Dolphin AC. The novel product of a five-exon stargazin-related gene abolishes CaV2.2 calcium channel expression. EMBO J. 2002;21:1514–1523. doi: 10.1093/emboj/21.7.1514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nichols RC, Wang XW, Tang J, Hamilton BJ, High FA, Herschman HR, Rigby WF. The RGG domain in hnRNP A2 affects subcellular localization. Exp Cell Res. 2000;256:522–532. doi: 10.1006/excr.2000.4827. [DOI] [PubMed] [Google Scholar]
- Pinol-Roma S. HnRNP proteins and the nuclear export of mRNA. Semin Cell Dev Biol. 1997;8:57–63. doi: 10.1006/scdb.1996.0122. [DOI] [PubMed] [Google Scholar]
- Powers PA, Liu S, Hogan K, Gregg RG. Molecular characterization of the gene encoding the gamma subunit of the human skeletal muscle 1,4-dihydropyridine-sensitive Ca2+ channel (CACNLG), cDNA sequence, gene structure, and chromosomal location. J Biol Chem. 1993;268:9275–9279. [PubMed] [Google Scholar]
- Pravettoni E, Bacci A, Coco S, Forbicini P, Matteoli M, Verderio C. Different localizations and functions of L-type and N-type calcium channels during development of hippocampal neurons. Developmental Biology. 2000;227:581–594. doi: 10.1006/dbio.2000.9872. [DOI] [PubMed] [Google Scholar]
- Raghib A, Bertaso F, Davies A, Page KM, Meir A, Bogdanov Y, Dolphin AC. Dominant-negative synthesis suppression of voltage-gated calcium channel Cav2.2 induced by truncated constructs. J Neurosci. 2001;21:8495–8504. doi: 10.1523/JNEUROSCI.21-21-08495.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rousset M, Cens T, Restituito S, Barrere C, Black JL, III, McEnery MW, Charnet P. Functional roles of gamma2, gamma3 and gamma4, three new Ca2+ channel subunits, in P/Q-type Ca2+ channel expressed in Xenopus oocytes. J Physiol. 2001;532:583–593. doi: 10.1111/j.1469-7793.2001.0583e.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salgado H, Tecuapetla F, Perez-Rosello T, Perez-Burgos A, Perez-Garci E, Galarraga E, Bargas J. A Reconfiguration of CaV2 Ca2+ Channel Current and Its Dopaminergic D2 Modulation in Developing Neostriatal Neurons. J Neurophysiol. 2005;94:3771–3787. doi: 10.1152/jn.00455.2005. [DOI] [PubMed] [Google Scholar]
- Sanders CR, Ismail-Beigi F, McEnery MW. Mutations of peripheral myelin protein 22 result in defective trafficking through mechanisms which may be common to diseases involving tetraspan membrane proteins. Biochemistry. 2001;40:9453–9459. doi: 10.1021/bi010894f. [DOI] [PubMed] [Google Scholar]
- Sandoval A, Andrade A, Beedle AM, Campbell KP, Felix R. Inhibition of recombinant N-type Ca(V) channels by the gamma 2 subunit involves unfolded protein response (UPR)-dependent and UPR-independent mechanisms. J Neurosci. 2007;27:3317–3327. doi: 10.1523/JNEUROSCI.4566-06.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schorge S, Gupta S, Lin ZX, McEnery MW, Lipscombe D. Calcium channel activation stabilizes a neuronal calcium channel mRNA. Nat Neurosci. 1999;2:785–790. doi: 10.1038/12153. [DOI] [PubMed] [Google Scholar]
- Shan J, Munro TP, Barbarese E, Carson JH, Smith R. A molecular mechanism for mRNA trafficking in neuronal dendrites. J Neurosci. 2003;23:8859–8866. doi: 10.1523/JNEUROSCI.23-26-08859.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp AH, Black JL, III, Dubel SJ, Sundarraj S, Shen JP, Yunker AMR, Copeland TD, McEnery MW. Biochemical and anatomical evidence for specialized voltage-dependent calcium channel gamma isoform expression in the epileptic and ataxic mouse, stargazer. Neuroscience. 2001;105:599–617. doi: 10.1016/s0306-4522(01)00220-2. [DOI] [PubMed] [Google Scholar]
- Shyu AB, Wilkinson MF. The double lives of shuttling mRNA binding proteins. Cell. 2000;102:135–138. doi: 10.1016/s0092-8674(00)00018-0. [DOI] [PubMed] [Google Scholar]
- Sofola OA, Jin P, Qin Y, Duan R, Liu H, de Haro M, Nelson DL, Botas J. RNA-Binding Proteins hnRNP A2/B1 and CUGBP1 Suppress Fragile X CGG Premutation Repeat-Induced Neurodegeneration in a Drosophila Model of FXTAS. Neuron. 2007;55:565–571. doi: 10.1016/j.neuron.2007.07.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swick AG, Janicot M, Cheneval-Kastelic T, McLenithan JC, Lane DM. Promoter-cDNA-directed heterologous protein expression in Xenopus laevis oocytes. Proc Natl Acad Sci USA. 1992;89:1812–1816. doi: 10.1073/pnas.89.5.1812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinker A, Jan YN, Jan LY. Regions responsble for assembly of inwardly rectifying potassium channels. Cell. 1996;87:857–868. doi: 10.1016/s0092-8674(00)81993-5. [DOI] [PubMed] [Google Scholar]
- Tomita S, Chen L, Kawasaki Y, Petralia RS, Wenthold RJ, Nicoll RA, Bredt DS. Functional studies and distribution define a family of transmembrane AMPA receptor regulatory proteins. J Cell Biol. 2003;161:805–816. doi: 10.1083/jcb.200212116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomita S, Fukata M, Nicoll RA, Bredt DS. Dynamic interaction of stargazin-like TARPs with cycling AMPA receptors at synapses. Science. 2004;303:1508–1511. doi: 10.1126/science.1090262. [DOI] [PubMed] [Google Scholar]
- Wilusz CJ, Wilusz J. Bringing the role of mRNA decay in the control of gene expression into focus. Trends Genet. 2004;20:491–497. doi: 10.1016/j.tig.2004.07.011. [DOI] [PubMed] [Google Scholar]
- Xu X, Shrager P. Dependence of axon initial segment formation on Na+ channel expression. J Neurosci Res. 2005;79:428–441. doi: 10.1002/jnr.20378. [DOI] [PubMed] [Google Scholar]
- Yano T, Lopez dQ, Matsui Y, Shevchenko A, Shevchenko A, Ephrussi A. Hrp48, a Drosophila hnRNPA/B homolog, binds and regulates translation of oskar mRNA. Dev Cell. 2004;6:637–648. doi: 10.1016/s1534-5807(04)00132-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.