Abstract
The distribution and genotypic variation of potential microcystin (MC) producers along the southern and eastern shores of Lake Ontario in 2001 and 2003 were examined using a suite of PCR primers. Cyanobacterial, Microcystis sp., and Microcystis-specific toxin primer sets identified shoreline distribution of cyanobacterial DNA (in 97% of the stations) and MC synthetase genes (in 50% of the stations). Sequence analysis of a partial mcyA amplicon targeting Microcystis, Anabaena, and Planktothrix species indicated that the Microcystis sp. genotype was the dominant MC genotype present and revealed a novel Microcystis-like sequence containing a 6-bp insert. Analysis of the same samples with genus-specific mcyE primers confirmed that the Microcystis sp. genotype was the dominant potential MC producer. Genotype compositions within embayments were relatively homogenous compared to those for shoreline and tributary samples. MC concentrations along the shoreline exhibited both temporal and spatial differences as evidenced by the protein phosphatase inhibition assay, at times exceeding the World Health Organization guideline value for drinking water of 1.0 μg MC-LReq liter−1. MC genotypes are widespread along the New York State shoreline of Lake Ontario, appear to originate nearshore, and can be carried through the lake via wind and surface water current patterns.
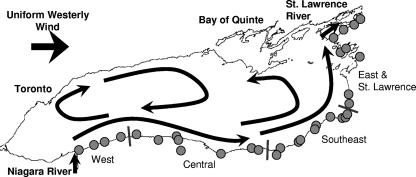
Lake Ontario is the easternmost and smallest of the Great Lakes, bordering Ontario, Canada, and New York, United States. Over 8 million people live in the watershed area and rely on Lake Ontario for recreational and drinking water. Approximately 80% of the water is supplied by the Niagara River, flowing in from Lake Erie, while 14% is delivered by small tributaries. The major outflow (93%) is via the St. Lawrence River (11). Lake Ontario, located last in the chain of Great Lakes, is affected by human activities occurring throughout the water basins of Lakes Superior, Michigan, Huron, and Erie. The Lake Ontario drainage basin is one of the fastest-growing areas in North America in terms of population and one of the most sprawling regions in the world. This growth is concurrent with the alteration of shoreline dynamics, resulting in increased nutrient and contaminant runoff, the effects of which are amplified by the large catchment area (64,030 km2) and long residence time (6 years) (10). Surface water currents controlled by the prevailing winds transport nutrients and particles along the Lake Ontario shoreline in a counterclockwise manner, while a central clockwise gyre mixes particles from the northern to the southern shores (Fig. 1) (3). The combination of runoff and mixing can result in the eutrophication of Lake Ontario embayments needed to trigger massive algal blooms, often occurring in late summer months with concomitant toxin production by cyanobacteria.
FIG. 1.
Sampling locations along the New York State border of Lake Ontario separated into regions. Arrows represent the Lake Ontario average annual surface water current pattern. (Adapted from reference 3 with permission of the publisher.)
Cyanobacteria produce a number of potent toxins. The most common are the microcystins (MCs), hepatotoxins made by a wide range of cyanobacteria, including Microcystis, Anabaena, Nostoc, and Planktothrix species. The generic structure of an MC is cyclo(-d-Ala-X-d-MeAsp-Z-Adda-d-Glu-Mdha-), where Mdha is N-methyldehydro-alanine, d-MeAsp is d- erythro-β-methylaspartic acid, and Adda is the nonproteogenic amino acid 3-amino-9-methoxy-10-phenyl-2,6,8-trimethyl-decca-4,6-dienoic acid (31). Structural variation can occur at all seven positions; however, the X and Z positions represent highly variable amino acids, and demethylation of MeAsp and Mdha is common (42). There are currently more than 80 identified MC variants, each possessing different degrees of toxicity, ranging from 50 to >1,200 μg kg−1 body weight (the 50% lethal dose [intraperitoneal] for mouse) (2, 42, 53).
Toxic and nontoxic populations of cyanobacteria can coexist in a single ecosystem and are indistinguishable by microscopy, rendering this technique unreliable for determining potential MC production. Methods commonly used for toxin monitoring include high-performance liquid chromatography (LC) with photodiode array (PDA) detection and/or mass spectrometry (MS), a protein phosphatase inhibition assay (PPIA), and enzyme-linked immunosorbent assays. These techniques are based on the structures or activities of MCs.
Elucidation of the MC synthetase gene cluster (mcy) encoding the nonribosomal peptide synthetase polyketide synthase enzyme complex has allowed the development of molecular techniques for studying MC-producing genera. The mcy gene clusters are significantly different between the major MC-producing genera Microcystis, Anabaena, and Planktothrix (8, 38, 45). Differences are found in the organization and presence of the mcy genes as well as the nucleotide sequences. The result of this genetic diversity is not fully understood, but its origin may be attributed to the ancestral nature of the genes, transposition between genera, and inter- and intrastrain genetic recombination (27, 28, 35, 44, 45). Multiple studies have focused on detection of the mcy genes using a PCR approach with species-specific primers based on differences within the mcy gene clusters (18, 20, 26, 32, 36, 47) as well as universal primers targeting conserved sequences of the MC operon (14, 17). The common mcy genes targeted are mcyA, -B, -C, -D, and -E, with many studies using a combination of the genes (15, 26, 27, 33, 44, 48). Several studies have indicated that MC-producing Microcystis spp. exhibit substantial genetic diversity within and between lakes located in both similar and different geographic locations (37, 52, 54). Examination of the mcy genes can give information on MC potential, the diversity of toxin producers within a population, and the geographic distribution of MC-producing genotypes.
Here, we investigated the occurrence and diversity of MC genotypes from embayments and lakeside water samples along the southern and eastern shores of Lake Ontario in 2001 and 2003. Previous studies have addressed the occurrence of MCs in Lake Ontario using analytical and biochemical techniques (5, 24); however, this is the first report on the identification of MC producers along the New York shoreline of Lake Ontario using molecular techniques. Multiple MC-specific PCR primers and DNA sequencing were used to determine the potential for MC production and toxin genotype composition in embayments and shoreline samples open to the lake. This was compared to MC concentration determined by PPIA. MC variant, trophic status, and cyanobacterial species composition were explored to determine the relationship with genotype composition and MC production. These data were used to identify areas of potential MC production and to address the contribution of embayments and surface water current patterns to the distribution of MCs along the New York shoreline of Lake Ontario.
MATERIALS AND METHODS
Sample collection.
Samples were collected during the summers of 2001 and 2003 from the southern and eastern shores of Lake Ontario (32 stations in 2001, 8 stations in 2003 [Fig. 1]). At each station, water samples were collected from a depth of 1.0 m, filtered, and immediately stored on dry ice for later analysis in the laboratory. Toxin samples (up to 20 liters) were filtered through a 90-mm glass fiber filter (Whatman 934-AH). DNA and chlorophyll samples (up to 1.0 liter) were filtered onto 47-mm glass fiber filters (Whatman 934-AH). Whole water samples were collected for total phosphorus analysis in acid-washed 125-ml bottles and stored on ice. Surface samples of cyanobacteria were collected with a 63-μm plankton net and stored in 0.2% (vol/vol) glutaraldehyde for phycological analysis.
MC analysis.
Toxin filters were extracted by sonication in 10 ml of 50% aqueous methanol acidified to 1% (vol/vol) with acetic acid. Extracts were clarified by centrifugation at 27,000 × g followed by filtration through a 0.45-μm nylon syringe filter and stored at −20°C. MC concentrations were determined using a PPIA, modified from Carmichael and An (7), run in 96-well plates (15). MC variants were identified from concentrated samples (10 to 20×) by high-performance LC (Ace 5 C18 column, 4.6 by 250 mm) with PDA and MS detectors using a gradient of 30 to 70% acetonitrile in water, both acidified to 0.1% (vol/vol) with trifluoroacetic acid. PDA detection was at 239 nm, and MS with electrospray ionization was used to detect molecular ions between 800 and 1,200 AMU (13). Putative MC variants were compared to standards, published molecular weights, and UV spectra (21, 55).
Chlorophyll a, total phosphorous, and phycological analysis.
Chlorophyll a was determined after extraction by sonication in 15 ml of 90% acetone, using UV-visible light spectrometry in 2001 (34) and fluorescence in 2003 (50). Total phosphorous was determined by acid hydrolysis and ammonium peroxydisulfate digestion, followed by color development with ammonium molybdate, potassium tartrate, and ascorbic acid and detection at 650 nm (9). The trophic statuses of the embayments were determined based on the average of the chlorophyll a and total phosphorous trophic status index values calculated using the Carlson trophic status formulas (6).
Cell counts were estimated for samples where we cloned the mcyA amplicon. Cyanobacteria were identified to the genus level based on Whitford and Schumacher (51) using a phase contrast light microscope under ×100 to 500 magnification. Cyanobacterial colonies and filaments were counted using a Palmer-Malony counting chamber. Cell numbers were estimated from colony and filament size classes using the protocol of Watzin et al. (49).
DNA analysis.
DNA was extracted from a 1.1-cm-diameter subsample of the original 47-mm filter using a protocol modified from Rudi et al. (39) described previously (15). Briefly, filters were placed in Tris-EDTA buffer (pH 8.0) and digested with lysozyme and RNase A, followed by a proteinase K digestion. Samples were clarified by two phenol-chloroform-isoamyl alcohol (25:24:1) extractions and a final chloroform-isoamyl alcohol (24:1) purification. DNA was precipitated in 95% ethanol overnight (−20°C), washed with 80% ethanol, and redissolved in 75 μl of Tris-EDTA buffer. Quantitation was done by UV spectroscopy at 260 nm.
Extracted DNA was amplified by PCR, separated by 1.5% (wt/vol) agarose gel electrophoresis, and visualized using ethidium bromide. Primer sequences used for PCR were based on cyanobacterial 16S rRNA (CYA), a Microcystis sp. 16S rRNA (MIC), and four toxin biosynthetic genes (mcyA, mcyB, mcyD, and mcyE) (Table 1). Samples that were negative by both CYA and MIC primer sets were spiked with lambda DNA and amplified with lambda-specific primers as a positive control for amplification. The mcyB and mcyD primer sets were used for an initial screening of all the samples. The mcyA primer set was used to analyze samples collected in August of 2001 and 2003, and stations positive by mcyA were amplified using genus-specific mcyE primers. Each 25-μl PCR contained 1.5 mM MgCl2, 200 μM of each deoxynucleoside triphosphate, 0.04 U μl−1 Taq polymerase (Applied Biosystems), 300 ng μl−1 bovine serum albumin, 5 ng μl−1 DNA, and 400 nM of each primer. Amplification was performed in an MJ Research PTC-100 thermocycler using the following protocol: initial denaturation at 94°C for 2 min, followed by 20 cycles of 94°C for 30 s, 65°C for 45 s (decreasing by 0.5°C each cycle), and 72°C for 1 min, followed by 15 additional cycles with a steady annealing temperature of 55°C, and ending with a final extension of 72°C for 8 min.
TABLE 1.
PCR primer sequences
| Target | Primer | Directiona | Primer sequence (5′→3′) | Reference |
|---|---|---|---|---|
| Cyanobacterial 16S rRNA | CYA | F | ACGGGTGAGTAACRCGTRA | 46 |
| R | CTTCAYGYAGGCGAGTTGCAGC | |||
| Microcystis 16S rRNA | MIC | F | ATGTGCCGCGAGGTGAAACCTAAT | 30 |
| R | TTACAAYCCAARRRCCTTCCTCCC | |||
| Microcystis mcyD | mcyD | F | GGTTCGCCTGGTCAAAGTAA | 18 |
| R | CCTCGCTAAAGAAGGGTTGA | |||
| Microcystis mcyB | mcyB | F | TGGGAAGATGTTCTTCAGGTATCCAA | 32 |
| R | AGAGTGGAAACAATATGATAAGCTAC | |||
| Microcystis, Anabaena, and Planktothrix sp. mcyA | mcyA | F | AAAAGTGTTTTATTAGCGGCTCAT | 14 |
| R | AAAATTAAAAGCCGTATCAAA | |||
| Microcystis, Anabaena, and Planktothrix sp. mcyE | mcyE | F | GAAATTTGTGTAGAAGGTGC | 47 |
| Microcystis sp. mcyE | R | CAATGGGAGCATAACGAG | ||
| Anabaena sp. mcyE | R | CAATCTCGGTATAGCGGC | ||
| Planktothrix sp. mcyE | R | CTCAATCTGAGGATAACGAT | 36 |
F, forward; R, reverse.
The mcyA amplicons were cloned using a TOPO TA cloning kit from Invitrogen Life Technologies with chemically competent pCR 2.1-TOPO TOP10 One Shot Escherichia coli cells. Up to 10 separate colonies were picked and grown overnight in liquid LB plus ampicillin (50 μg ml−1) for plasmid minipreparation. Minipreps were completed via the alkaline lysis method, and the final DNA pellet was dissolved in 50 μl of water (41). Transformants were confirmed by digestion with EcoRI. DNA sequencing of the genetic recombinants was performed at the Biotechnology Resource Center at Cornell University (Ithaca, NY) using an Applied Biosystems automated 3730 DNA analyzer with BigDye Terminator chemistry and AmpliTaq-FS DNA polymerase. DNA sequences were modified using BioEdit (version 7.0.5.3) and Mega 3.1, queried against known sequences in the GenBank database using a nucleotide-nucleotide BLAST (blastn), and aligned to known sequence matches using ClustalX (version 1.81) and Mega 3.1 (1, 12, 16, 19). Neighbor-joining analysis of the mcyA sequence alignment was done with PAUP* 4.0 and Mega 3.1 using p-distance analysis (43). Bootstrap values were obtained for 1,000 replicates.
Nucleotide sequence accession numbers.
Sequences were deposited in GenBank (accession numbers EF424278 through EF424374).
RESULTS
Occurrence of MCs along the New York coast of Lake Ontario.
Low levels of MCs were detected at each station from the sample period of June to August 2001, ranging from 0.002 to 0.070 μg liter−1, with the largest amount occurring at Port Bay East in early August (Table 2). A temporal variation in MC concentrations was observed, with some stations highest in June and others in August, but the highest lakewide average and total MC concentrations occurred during August (data not shown). Stations resampled in August 2003 had MC concentrations ranging from 0.003 to 1.043 μg liter−1, with the largest amount occurring at Selkirk Shores in late August (Table 2). The Oswego Shore station also had a high MC value (0.990 μg liter−1) in early August. In late August 2003, Nine Mile Bay, Little Salmon River, and Selkirk Shores stations all had higher MC concentrations than those observed in late August 2001 (11 to 69 times higher per station). West Sodus Bay, Little Sodus Bay (bayside), and Chaumont Bay all had similar MC concentrations as observed in 2001.
TABLE 2.
Sampling locations along the Lake Ontario shoreline, with numbers of times sampled, maximum MC levels reached, and mcyB and mcyD potentials for each month
| Regional location, yr, and station name | Location typea | No. of times sampled | Maximum MC-LReq (μg/liter)b |
mcyB potential/mcyD potential for indicated mo
|
|||
|---|---|---|---|---|---|---|---|
| June | July | Early August | Late August | ||||
| West, 2001 | |||||||
| Fort Niagara | L | 3 | 0.011 | −/+ | +/+ | −/− | |
| Wilson Tuscarora | E | 3 | 0.014 | −/− | +/+ | −/− | |
| Town of Olcott | L | 2 | 0.005 | −/− | −/− | ||
| Golden Hill | L | 3 | 0.011 | −/− | −/− | −/− | |
| Central, 2001 | |||||||
| Lakeside Beach | L | 1 | 0.002 | −/− | |||
| Oak Orchard Harbor | L | 2 | 0.013 | −/− | −/− | ||
| Hamlin | E | 3 | 0.021 | −/− | −/+ | −/− | |
| Braddock Bay | E | 2 | 0.010 | −/− | −/− | ||
| Eastman Park | L | 2 | 0.010 | −/− | −/− | ||
| Irondequoit Bay | E | 5 | 0.015 | −/− | −/− | −/− | +/+ |
| B. Forman County Park | L | 2 | 0.005 | −/− | −/− | ||
| West Sodus Bay | E | 5 | 0.004 | −/− | −/− | −/− | −/− |
| Southeast, 2001 | |||||||
| Port Bay West | E | 4 | 0.006 | −/− | +/+ | −/− | −/+ |
| Port Bay East | L | 4 | 0.070 | −/− | −/− | −/− | +/− |
| Little Sodus Bay, lakeside | L | 5 | 0.060 | −/− | −/− | −/− | +/+ |
| Little Sodus Bay, bayside | E | 3 | 0.004 | −/− | −/+ | ||
| Fair Haven | E | 2 | 0.005 | −/− | −/− | ||
| Oswego Port Authority | L | 3 | 0.004 | −/− | −/− | −/− | |
| Nine Mile | L | 5 | 0.029 | −/− | −/− | −/− | +/+ |
| Little Salmon River | L | 4 | 0.010 | −/− | −/− | −/− | +/+ |
| Selkirk Shores | L | 4 | 0.015 | −/− | −/− | −/− | +/+ |
| Pine Grove | E | 2 | 0.005 | −/− | −/− | ||
| East and St. Lawrence, 2001 | |||||||
| Sandy Pond | E | 1 | 0.020 | −/− | |||
| Lakeview Mgmt Area | L | 3 | 0.010 | −/− | −/+ | −/− | |
| Lakeview DEC Launch | E | 1 | 0.003 | −/− | |||
| Westcott Beach | E | 3 | 0.006 | −/− | −/− | −/− | |
| Long Point Isthmus | L | 4 | 0.004 | −/− | −/+ | −/− | −/− |
| Chaumont Bay | E | 4 | 0.040 | −/− | −/− | −/− | −/+ |
| Tibbit Point | L | 4 | 0.005 | −/− | −/− | −/− | −/− |
| Cedar Point | E | 4 | 0.005 | −/− | −/− | −/− | −/− |
| Grass Point | E | 3 | 0.003 | −/− | −/− | −/− | |
| Collins Point | E | 1 | 0.003 | −/− | |||
| Southeast and East, 2003 | |||||||
| West Sodus Bay | E | 1 | 0.003 | +/+ | |||
| Little Sodus Bay, bayside | E | 1 | 0.003 | +/+ | |||
| Oswego River | E | 1 | 0.293 | +/+ | |||
| Oswego Shore | L | 2 | 0.990 | +/+ | +/+ | ||
| Nine Mile | L | 1 | 0.321 | +/+ | |||
| Little Salmon River | L | 1 | 0.200 | +/+ | |||
| Selkirk Shores | L | 1 | 1.043 | +/+ | |||
| Chaumont Bay | E | 2 | 0.047 | −/+ | |||
L, lakeside sample; E, embayment sample.
For stations sampled more than once, only the maximum MC value is given.
Presence of mcyB and mcyD genes.
Stations sampled in June, July, and August 2001 were analyzed for MC potential with the mcyB and mcyD primer sets (Table 2) as well as the CYA- and MIC-specific primer sets. In June 2001, the westernmost station, Fort Niagara, was the only station positive by the mcyB or mcyD primer set. Six stations spanning the entire sampling region were positive for MC potential in July. In early August 2001, there were no toxin genes detected from any of the sampling stations. In late August, MC potential increased to include nine stations spanning all regions except the western, which was not sampled at this time. Regardless, the greatest MC potential was observed in late August.
In late August 2003, eight stations were sampled, and each was positive for MC potential by both the mcyB and the mcyD primer sets except Chaumont Bay, which had only an mcyD amplicon (Table 2). Only one sample was collected earlier in the year (from Oswego Shore, collected in early August); therefore, changes in toxin potential throughout the season could not be determined for 2003.
MC potential, as determined by mcyB and mcyD, was detected in 25 of the 107 samples collected in 2001 and 2003. In 9 of those 25 samples, only one of the toxin genes was detected, with 8 of those 9 instances being only the mcyD amplicon.
Determination of genotype composition by mcyA sequencing.
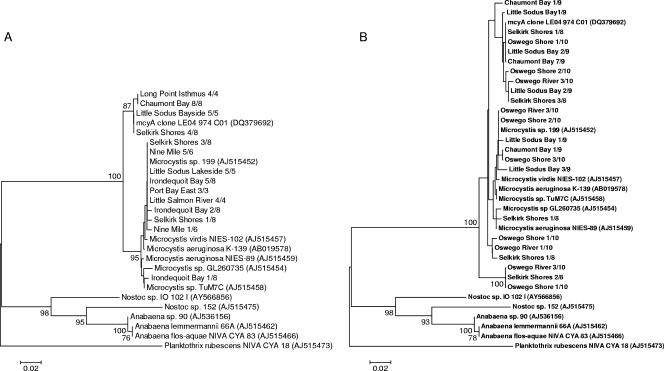
To determine the cyanobacterial genera present that could be responsible for MC production, samples collected in August of 2001 were analyzed using an mcyA primer set targeting the mcyA gene from Microcystis, Anabaena, and Planktothrix species and the resulting amplicons cloned and sequenced (14). The mcyA sequences were confirmed to be from the MC synthetase complex by using the GenBank database BLASTX search. Phylogenetic analysis, utilizing reference mcyA sequences obtained from GenBank, showed that each Lake Ontario station amplicon clustered with Microcystis genera and was distinctly different from Anabaena and Planktothrix sequences (Fig. 2). This large Microcystis clade was split into two smaller clusters, one grouping with known Microcystis mcyA sequences (291 bp) and the other subset clustering with a previously reported Microcystis-like sequence (297 bp) isolated from Lake Erie (37). This unique clade was characterized by a 6-bp insert at positions 258 to 263 in the partial mcyA sequence, corresponding to the same positions where additional residues are found in Planktothrix, Anabaena, and Nostoc partial mcyA sequences. Amplicons from Chaumont Bay (eight of eight), Long Point Isthmus (four of four), Little Sodus Bay (bayside) (five of five), and Selkirk Shores (four of eight) all had clones containing this unique sequence.
FIG. 2.
Neighbor-joining trees made using p-distance analysis to show the relationship between partial mcyA genes (∼300 bp) from Lake Ontario embayments in 2001 (A) and 2003 (B). Trees were generated using the program Mega 3.1. Bootstrap values of >75% are displayed at the nodes (1,000 replicates). The scale bar is a measure of the number of substitutions per site. Clone names indicate the embayment and number of clones represented. Reference sequences were obtained from GenBank, with accession numbers shown in parentheses.
Selkirk Shores, located at the mouth of a tributary running into Lake Ontario, was the only station in 2001 containing clones that clustered in both Microcystis clades. Samples collected from two stations at Little Sodus Bay, bayside and lakeside, had distinctively different genotypic populations. The bayside population, collected from inside the bay, corresponded entirely with the unique Microcystis-like mcyA sequence, while the lakeside population, collected from the side of the isthmus open to Lake Ontario, grouped with the traditional Microcystis mcyA sequence. Sequences obtained from the southern shore of Lake Ontario (Irondequoit, Little Sodus Bay [lakeside], Nine Mile, Port Bay East, Little Salmon River, and Selkirk Shores) formed a large cluster with other known Microcystis mcyA sequences and had comparable similarities.
In 2003, samples from Little Sodus Bay (bayside), Selkirk Shores, Chaumont Bay, Oswego Shore, and Oswego River were amplified by the mcyA primer set and the amplicons cloned and sequenced to determine genotype composition and the influence of lakeside and embayment populations on MC potential within Lake Ontario. A BLASTX search and phylogenetic analysis indicated that the partial mcyA sequences isolated were from the MC synthetase complex and again correlated with other known mcyA sequences from Microcystis genera. The large Microcystis clade was split into two smaller subclades, one corresponding to known Microcystis mcyA sequences and the other to the unique Microcystis-like mcyA sequences isolated in 2001 (Fig. 2). Chaumont Bay (8 of 9), Selkirk Shores (4 of 8), Little Sodus Bay (bayside) (5 of 9), Oswego River (3 of 10), and Oswego Shore (3 of 10) samples all had clones corresponding to the unique Microcystis-like mcyA sequence. Each station had a mixed genotype population.
To ensure that our identification of the genus responsible for MC production was not due to cloning biases, species-specific mcyE primer sets were used. These primer sets have the same forward primer, with specific Microcystis, Anabaena, and Planktothrix sp. reverse primers (Table 1) (36). All stations were positive by the Microcystis-specific mcyE primer set. In 2001, four of the nine stations examined had faint amplification by either the Anabaena or the Planktothrix sp.-specific mcyE primer sets: Little Sodus Bay (lakeside) and Selkirk Shores showed amplification of Planktothrix specific genes, and Irondequoit Bay, Selkirk Shores, and Chaumont Bay amplified Anabaena-specific genes (data not shown).
Genotype composition and MC variants.
MC variants were identified by LC-MS and compared with respect to genotype composition determined by mcyA sequencing (Table 3). In 2001, eight of the nine stations contained detectable levels of MC by LC-MS. The MC congeners identified included RR, YR, LR, LA, and LY (Table 3). All stations with detectable levels of MC contained the LR variant. The LR variant was dominant at all stations except Port Bay East, where the ratio of RR to LR was 51% to 49%. In 2003, four of the five samples examined had detectable levels of MC by LC-MS and included the toxin congeners RR, LR, and LA. As in 2001, all samples examined in 2003 contained the LR variant, which was also the dominant congener (Table 3). Variation in MC congener composition at both Chaumont Bay and Selkirk Shores was higher in 2003 than in 2001. Samples containing the unique Microcystis-like mcyA sequence in 2001 and 2003 had the MC variants LR, LA, and RR. In neither year did there appear to be a pattern between the observed MC variant and the presence or absence of the unique Microcystis-like mcyA sequence (Table 3).
TABLE 3.
MC concentration and variants, PCR results, abundance of potential MC-producing cyanobacterial cells, and trophic status at stations analyzed for genotype composition by mcyA sequencinga
| Yr and site | MC-LReq concn (μg/liter)b | MC variant ratio (%) | PCR results for CYA/MIC | % abundance of potential MC-producing cyanobacterial cellsc of indicated genus
|
Carlson's trophic status index for:
|
Trophic status | |||
|---|---|---|---|---|---|---|---|---|---|
| Microcystis | Anabaena | Aphanizomenon | Chl-a | Total P | |||||
| 2001 | |||||||||
| Irondequoit Bay | 0.015 | LR (50)/RR (25)/LY (25) | +/+ | 85.9 | 0.0 | 14.1 | 40 | 34 | Oligo |
| Port Bay East | 0.007 | RR (51)/LR (49) | +/+ | 0.0 | 30.9 | 69.1 | 40 | 58 | Meso |
| Little Sodus Bay, bayside | 0.004 | LR (80)/LA (20) | +/+ | 86.0 | 0.0 | 14.0 | 48 | 18 | Oligo |
| Little Sodus Bay, lakeside | 0.060 | LR (81)/YR (13)/RR (6) | +/+ | 93.7 | 0.5 | 5.8 | 38 | 43 | Meso |
| Salmon River | 0.004 | LR (100) | +/+ | 0.0 | 0.0 | 0.0 | 42 | 63 | Eu |
| Nine Mile | 0.013 | LR (100) | +/+ | 0.0 | 0.0 | 0.0 | 46 | 35 | Meso |
| Selkirk Shores | 0.015 | LR (100) | +/+ | 0.0 | 0.0 | 0.0 | 41 | 24 | Oligo |
| Long Point Isthmus | 0.004 | ND | +/+ | 100.0 | 0.0 | 0.0 | 59 | 64 | Eu |
| Chaumont Bay | 0.040 | LR (100) | +/+ | 0.0 | 0.0 | 0.0 | 33 | 28 | Oligo |
| 2003 | |||||||||
| Little Sodus Bay, bayside | 0.003 | ND | +/− | 18.5 | 71.2 | 10.3 | 36 | 58 | Meso |
| Chaumont Bay | 0.047 | LR (72)/LA (28) | +/+ | 5.1 | 94.9 | 0.0 | 46 | 47 | Meso |
| Selkirk Shores | 1.043 | LR (78)/RR (22) | +/+ | 100.0 | 0.0 | 0.0 | 50 | 76 | Eu |
| Oswego Shore | 0.769 | LR (72)/RR (28) | +/+ | 100.0 | 0.0 | 0.0 | 48 | 68 | Eu |
| Oswego River | 0.239 | LR (100) | +/+ | 100.0 | 0.0 | 0.0 | 42 | 61 | Meso |
Samples listed in bold have the novel mcyA insert. ND, not detected; Chl-a, chlorophyll a; eu, eutrophic; meso, mesotrophic; oligo, oligotrophic.
MC concentration from the sample cloned.
No Planktothrix cells were observed.
Trophic status and cyanobacterial species composition.
The Carlson trophic status index and cyanobacterial species composition were compared to those for samples targeted by mcyA sequencing in 2001 and 2003. Trophic status, based on average chlorophyll a and total phosphorous trophic status indices, ranged from oligotrophic to eutrophic in 2001 and mesotrophic to eutrophic in 2003 (Table 3). In 2001, there was no relationship between MC concentration and trophic status (r2 = 0.16, n = 9), while in 2003, the samples examined exhibited higher MC production with increased trophic state (r2 = 0.99, n = 5). Stations containing the unique Microcystis-like mcyA sequence in 2001 and 2003 ranged from oligotrophic to eutrophic.
Cyanobacterial cell numbers fluctuated widely between sampling stations in both 2001 and 2003. Several stations had few or no visible Microcystis cells (five of nine stations in 2001), but amplicons were obtained by the cyanobacterial and Microcystis sp.-specific primer sets. In 2003, Selkirk Shores and Oswego Shore, two stations corresponding to particulate MC concentrations near the WHO guideline value for MC in drinking water, had few potentially toxic cyanobacteria present by microscopy in the net-collected samples. There were Anabaena filaments present in 4 of the 13 stations counted, but no Planktothrix filaments were observed. In general, the abundance of potentially toxic cyanobacterial genera was low. Stations containing the unique mcyA genotype contained either no visible toxigenic cyanobacterial cells (Selkirk Shores and Chaumont Bay in 2001), only Microcystis cells (Long Point Isthmus in 2001 and Selkirk Shores, Oswego River, and Oswego Shore in 2003), a combination of Microcystis and Anabaena cells (Chaumont Bay in 2003), a combination of Microcystis and Aphanizomenon cells (Little Sodus Bay [bayside] in 2001), or a combination of Microcystis, Anabaena, and Aphanizomenon cells (Little Sodus Bay [bayside] in 2003).
DISCUSSION
Embayments such as Irondequoit, Port, Little Sodus, Chaumont, and the Oswego River are significantly separated from each other and the main body of Lake Ontario and are considered separate systems. They are affected by cultural eutrophication and provide protected mesocosms for cyanobacterial growth and potential MC production (23). The eutrophication is in part due to sewage effluent (Port Bay), high turbidity (Little Sodus Bay), river discharge (Oswego Harbor), or agricultural runoff and inadequate septic systems (Chaumont Bay). Makarewicz et al. (24) explored MC production along the southern shore of Lake Ontario from the Niagara River to Chaumont Bay and reported levels well below the WHO guideline value for drinking water of 1.0 μg MC-LReq liter−1 (maximum MC concentration of 0.795 μg liter−1 at Long Pond North), with the highest MC values from bays and rivers (24). More recently, Makarewicz et al. (unpublished) have reported an algal scum (Sackets Harbor) and shoreside sample (Oak Orchard Creek) containing higher MC concentrations. Here, we report MC levels from Oswego Shore (near the site of the Onondaga County water intake pipes) and Selkirk Shores (a public park area) with MC values near 1.0 μg MC-LReq liter−1. MC values reported for the 32 other stations (Fig. 1) were well below 1.0 μg MC-LReq liter−1 (Table 2); however, the potential for MC production was widespread (see below).
Cyanobacteria and Microcystis spp. were present in nearly all of the samples collected in 2001 and 2003 as determined by molecular analysis. Microscopic examination revealed few cyanobacteria, including Microcystis spp., at several sites. This may be a reflection of low cell abundance and phycological sampling technique in areas corresponding to high sediment loading and water mixing. Cyanobacterial cells and small colonies can have diameters of less than 63 μm, thereby passing through plankton nets, leading to underestimates of the community population. The lack of visual confirmation emphasizes the need for molecular analysis to detect potentially toxic cyanobacteria in low concentrations prior to formation of blooms with elevated toxin levels. PCR amplicons from the mcyB and mcyD primer sets indicated the potential for toxin formation along the southern and eastern shores of Lake Ontario (17 of 34 stations). The lack of toxin genes in early August 2001 indicated periodicity in toxin blooms similar to previously reported bloom dynamics in New York State (15). The mcyD primer set detected MC genes more often than the mcyB primer set and was more applicable to detecting toxin genes at low concentrations. Sensitivity studies comparing the mcyD and mcyB primer sets confirm this finding and support the use of primers against mcyD as an early indicator of toxin potential (A. M. Hotto, unpublished data).
Microcystis was the major MC producer in Lake Ontario based on mcyA sequences. It is possible that the mcyA primers reflected only the dominant MC sequences from each station, thereby underestimating the genotypic diversity within the population. However, use of mcyE species-specific primers confirmed the presence of Microcystis spp. and indicated that the presence of potentially toxic Planktothrix and Anabaena species was not widespread. Anabaena and Oscillatoria spp. have been previously detected in offshore waters of Lake Ontario; however, they did not make up a large proportion of the phytoplankton biomass (<15%) (22). This was in agreement with our current observations.
The appearance of a unique Microcystis-like mcyA sequence in Lake Ontario is still perplexing. The sequence was characterized by a 6-bp insert (TTTGCG), corresponding to the amino acids Phe-Ala (FA). This is similar to the insertion in the partial mcyA sequences of Anabaena and Nostoc spp. (TTTGGT; FG) and Planktothrix spp. (TTGGGT; LG) and observed in samples from Lake Erie (TTTGCG; FA) (37). It is unknown whether this genotype represents a novel Microcystis sp. or a genus closely related to Microcystis that has not been previously reported to produce MCs. Efforts to isolate the responsible species are currently in progress. The presence of this genetic element may be due to a recombination event, a mutational insertion, or an ancestral relic. It is most closely related to a similar insertion in the partial mcyA sequences of Anabaena and Nostoc spp., lending support to reports that the MC biosynthetic gene cluster originated in an Anabaena-like sp. due to the colinearity between the mcy genes and MC structure (4, 25, 38). It is unknown how the extra base pairs and their corresponding amino acid residues affect MC production or toxicity.
Variations in trophic status and cyanobacterial species composition were examined to determine if they corresponded to the presence or absence of the unique mcyA genotype. The unique mcyA genotype was present in samples ranging from oligotrophic to eutrophic, in stations containing few or no visible potentially toxic cyanobacterial cells, and in stations containing Microcystis, Anabaena, and Aphanizomenon species. There was increased genotypic diversity in Chaumont Bay and Little Sodus Bay (bayside) from 2001 to 2003 concurrent with the visible appearance of Anabaena filaments. However, there was no indication of toxic Anabaena in 2003 by either mcyA sequencing or mcyE species-specific amplification. The lack of congruity between nutrients, biomass, cyanobacterial composition, and toxicity emphasizes the unpredictable nature of algal blooms.
Variation within the MC congener profile was examined to determine its relationship with the presence/absence of the unique mcyA sequence. Previously, only the MC-RR, -YR, and -LR variants have been detected in Lake Ontario (29). Here, we also identified MC-LA and -LY, but the most common congener was MC-LR, corresponding to both mcyA genotypes. Variations were not observed in the Ala residue encoded by the mcyA2 gene targeted by our mcyA primer set. Increased genetic diversity from 2001 to 2003 coincided with the appearance of MC congeners MC-LA (Chaumont Bay) and -RR (Selkirk Shores).
One of the key questions for the management of MC-producing species is whether a toxic bloom originates in an embayment and is transported to the lake or initiates offshore and is transported through the lake via the lakewide circulation pattern. The sporadic distribution of mcy potential along the shoreline (Table 2) indicates that separate stations harbor potential MC-producing populations. The separation of genotype and phenotype between most embayments and lakeside samples confirms that the water current pattern did not have a strong effect on MC distribution. This is also supported by the spatial differences in MC concentration in both this study and reports by Makarewicz et al. (unpublished). Water exchange between the embayment and the main lake may result in the export of organisms to the lake and their transport along the shoreline by prevailing water currents. However, embayments such as Little Sodus Bay are separated from Lake Ontario such that water exchange is negligible (40).
Lakeside genetic populations examined along the southern shore of Lake Ontario in 2001 (Little Sodus Bay [lakeside], Nine Mile, Port Bay East, Little Salmon River, and Selkirk Shores) clustered together, representing low genotypic diversity. The absence of the unique mcyA genotype from the lakeside samples indicated that this genotype originated in selected embayments. Selkirk Shores was the only station with a mixed genotypic population in 2001. The traditional mcyA genotype could have originated upstream in the embayment or been carried in from the west, while the unique mcyA genotype likely originated in the protected embayment. The disparity between genotype populations at Little Sodus Bay in 2001 and 2003 may be a result of water runoff carrying another toxin population into the embayment. Storm-induced currents have the ability to flush toxic genera into the lake from nearby ponds and small lakes due to the strong affect on particle flow (up to several tens of cm s−1), resulting in unusual mixing patterns and the introduction of different MC genotypes (3). Makarewicz et al. (unpublished) also noted higher MC concentrations in ponds, rivers, lakes, and embayments draining into Lake Ontario, suggesting that these sources serve as a reservoir for MC production.
In 2003, the Oswego River strongly influenced the MC-producing population. The Oswego River and Oswego Shore samples had similar genotype compositions, indicating that the toxin population originated upstream or at the mouth of the Oswego River and was carried into the lake. The same traditional and novel mcyA sequences have been detected in Oneida Lake, which could seed Lake Ontario through the Erie Canal and Oswego River (A. M. Hotto, unpublished). We hypothesize that the cyanobacterial population was then transported eastward to the Oswego Shore site. This circulation pattern may also have transported the population to Selkirk Shores and contributed to the high MC content observed there. It is possible that transportation of this unique genotype between lakes may occur through external factors, such as fishing boat transfers or animal carriers. The observed novel sequence has been noted only in the northeastern United States, giving support for the formation of a unique ecotype.
Although MC concentrations at most stations were below the WHO guideline value for drinking water, MC-producing genotypes were widespread along the New York shoreline of Lake Ontario. Distribution of a unique MC-producing genotype indicated that Lake Ontario embayments harbor toxigenic cyanobacteria. Toxin populations from lakeside stations add to the cyanobacterial populations along the shoreline, and river inputs have a large influence on mixed genotypic populations. MC production and genotype distribution were not correlated with nutrients or algal biomass. Continued alteration of shoreline habitat, introduction of nonnative species, and decline of native community populations in Lake Ontario, where the genes for toxin production have been shown to exist, may all lead to increased occurrence of cyanobacterial blooms and MC production in embayments with elevated nutrient levels, increased light penetration due to zebra mussel introduction, and protection from wave turbulence. It is essential that monitoring programs be aware of the presence of these potential MC producers and the novel genotypes found in these lakes.
Acknowledgments
We thank Xingye Yang, Elizabeth Patchett, and Nick Smith for their help in sample collection and water chemistry analysis. Caitlynn Armstrong assisted in DNA extraction, and Anthony Ouellette and Steven Wilhelm provided assistance with the PCR protocols.
This work was funded by New York Sea Grant under NOAA grant no. NA16RG1645 and by the NOAA Merhab program under grant no. NA16OP2788.
Footnotes
Published ahead of print on 25 May 2007.
REFERENCES
- 1.Altschul, S. F., W. Gish, W. Miller, E. W. Meyers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215:403-410. [DOI] [PubMed] [Google Scholar]
- 2.Babica, P., L. Blaha, and B. Marsalek. 2006. Exploring the natural role of microcystins—a review of effects on photoautotrophic organisms. J. Phycol. 42:9-20. [Google Scholar]
- 3.Beletsky, D., J. H. Saylor, and D. J. Schwab. 1999. Mean circulation in the Great Lakes. J. Great Lakes Res. 25:78-93. [Google Scholar]
- 4.Börner, T., and E. Dittmann. 2005. Molecular biology of cyanobacterial toxins, p. 25-40. In J. Huisman, H. Matthijs, and P. Visser (ed.), Harmful cyanobacteria, vol. 3. Springer, Norwell, MA. [Google Scholar]
- 5.Boyer, G. L. Cyanobacterial toxins in New York and the Lower Great Lakes ecosystems. Adv. Exp. Med. Biol., in press. [DOI] [PubMed]
- 6.Carlson, R. E., and J. Simpson. 1996. A coordinator's guide to volunteer lake monitoring methods. North American Lake Management Society, Madison, WI.
- 7.Carmichael, W. W., and J. An. 1999. Using an enzyme linked immunosorbent assay (ELISA) and a protein phosphatase inhibition assay (PPIA) for the detection of microcystins and nodularins. Nat. Toxins 7:377-385. [DOI] [PubMed] [Google Scholar]
- 8.Christiansen, G., J. Fastner, M. Erhard, T. Börner, and E. Dittmann. 2003. Microcystin biosynthesis in Planktothrix: genes, evolution, and manipulation. J. Bacteriol. 185:564-572. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Clesceri, L. S., A. E. Greenberg, and A. D. Eaton. 1995. Standard methods for the examination of water and wastewater, 19th ed. American Public Health Association, Washington, DC.
- 10.Environment Canada and USEPA. 2005. State of the Great Lakes 2005, p. 15-19. State of the Lakes Ecosystem Conference. Environment Canada, Quebec, Canada, and U.S. Environmental Protection Agency, Washington, DC.
- 11.Fuller, K., H. Sherar, and J. Wittig (ed.). 1995. The Great Lakes: an environmental atlas and resource book, 3rd ed. Government of Canada, Toronto, Ontario, Canada, and U.S. Environmental Protection Agency, Great Lakes National Program Office, Chicago, IL. http://www.epa.gov/glnpo/atlas/index.htm.
- 12.Hall, T. A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41:95-98. [Google Scholar]
- 13.Harada, K.-I., H. Murata, Z. Qiang, M. Suzuki, and F. Kondo. 1996. Mass spectrometric screening method for microcystins in cyanobacteria. Toxicon 34:701-710. [DOI] [PubMed] [Google Scholar]
- 14.Hisbergues, M., G. Christiansen, L. Rouhiainen, K. Sivonen, and T. Börner. 2003. PCR-based identification of microcystin-producing genotypes of different cyanobacterial genera. Arch. Microbiol. 180:402-410. [DOI] [PubMed] [Google Scholar]
- 15.Hotto, A., M. Satchwell, and G. Boyer. 2005. Seasonal production and molecular characterization of microcystins in Oneida Lake, New York, USA. Environ. Toxicol. 20:243-248. [DOI] [PubMed] [Google Scholar]
- 16.Jeanmougin, F., J. D. Thompson, M. Gouy, D. G. Higgins, and T. J. Gibson. 1998. Multiple sequence alignment with Clustal X. Trends Biochem. Sci. 23:403-405. [DOI] [PubMed] [Google Scholar]
- 17.Jungblut, A.-D., and B. A. Neilan. 2006. Molecular identification and evolution of the cyclic peptide hepatotoxins, microcystin and nodularin, synthetase genes in three orders of cyanobacteria. Arch. Microbiol. 185:107-114. [DOI] [PubMed] [Google Scholar]
- 18.Kaebernick, M., B. A. Neilan, T. Börner, and E. Dittmann. 2000. Light and the transcriptional response of the microcystin biosynthesis gene cluster. Appl. Environ. Microbiol. 66:3387-3392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Kumar, S., K. Tamura, and M. Nei. 2004. MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 5:150-163. [DOI] [PubMed] [Google Scholar]
- 20.Kurmayer, R., G. Christiansen, M. Gumpenberger, and J. Fastner. 2005. Genetic identification of microcystin ecotypes in toxic cyanobacteria of the genus Planktothrix. Microbiology 151:1525-1533. [DOI] [PubMed] [Google Scholar]
- 21.Lawton, L. A., and C. Edwards. 2001. Purification of microcystins. J. Chromatogr. A 912:191-209. [DOI] [PubMed] [Google Scholar]
- 22.Makarewicz, J. C., T. W. Lewis, and P. Bertram. 1995. Epilimnetic phytoplankton and zooplankton biomass and species composition in Lake Ontario, 1986 to 1992. U.S. Environmental Protection Agency, Great Lakes National Program Office, Chicago, IL. http://www.epa.gov/glnpo/monitoring/plankton/ont86-92/new_ont86-92_index.html.
- 23.Makarewicz, J. C. 2000. New York's north coast: a troubled coastline. Lake Ontario Embayments Initiative report. Center for Environmental Information, Rochester, NY.
- 24.Makarewicz, J. C., G. L. Boyer, W. Guenther, M. Arnold, and T. W. Lewis. 2006. The occurrence of cyanotoxins in the nearshore and coastal embayments of Lake Ontario. Great Lakes Res. Rev. 7:25-29. [Google Scholar]
- 25.Marahiel, M. A., T. Stachelhaus, and H. D. Mootz. 1997. Modular peptide synthetases involved in nonribosomal peptide synthesis. Chem. Rev. 97:2651-2673. [DOI] [PubMed] [Google Scholar]
- 26.Mbedi, S., M. Welker, J. Fastner, and C. Wiedner. 2005. Variability of the microcystin synthetase gene cluster in the genus Planktothrix (Oscillatoriales, Cyanobacteria). FEMS Microbiol. Lett. 245:299-306. [DOI] [PubMed] [Google Scholar]
- 27.Mikalsen, B., G. Boison, O. M. Skulberg, J. Fastner, W. Davies, T. M. Gabrielsen, K. Rudi, and K. S. Jakobsen. 2003. Natural variation in the microcystin synthetase operon mcyABC and impact on microcystin production in Microcystis strains. J. Bacteriol. 185:2774-2785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Moffitt, M. C., and B. A. Neilan. 2004. Characterization of the nodularin synthetase gene cluster and proposed theory of the evolution of cyanobacterial hepatotoxins. Appl. Environ. Microbiol. 70:6353-6362. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Murphy, T. P., K. Irvine, J. Guo, J. Davies, H. Murkin, M. Charlton, and S. B. Watson. 2003. New microcystin concerns in the lower Great Lakes. Water Qual. Res. J. Canada 38:127-140. [Google Scholar]
- 30.Neilan, B. A., D. Jacobs, T. Del Dot, L. L. Blackall, P. R. Hawkins, P. T. Cox, and A. E. Goodman. 1997. rRNA sequences and evolutionary relationships among toxic and nontoxic cyanobacteria of the genus Microcystis. Int. J. Syst. Bacteriol. 47:693-697. [DOI] [PubMed] [Google Scholar]
- 31.Neilan, B. A., E. Dittmann, L. Rouhiainen, R. A. Bass, V. Schaub, K. Sivonen, and T. Börner. 1999. Nonribosomal peptide synthesis and toxigenicity of cyanobacteria. J. Bacteriol. 181:4089-4097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Nonneman, D., and P. Zimba. 2002. A PCR-based test to assess the potential for microcystin occurrence in channel catfish production ponds. J. Phycol. 30:230-233. [Google Scholar]
- 33.Ouellette, A., S. M. Handy, and S. W. Wilhelm. 2006. Toxic Microcystis is widespread in Lake Erie: PCR detection of toxin genes and molecular characterization of associated cyanobacterial communities. Microb. Ecol. 51:154-165. [DOI] [PubMed] [Google Scholar]
- 34.Parsons, T. R., Y. Maita, and C. M. Lalli. 1984. Plant pigments, p. 100-112. In T. R. Parsons, Y. Maita, and C. M. Lalli (ed.), A manual of chemical and biological methods for seawater analysis. Pergamon Press, New York, NY.
- 35.Rantala, A., D. P. Fewer, M. Hisbergues, L. Rouhiainen, J. Vaitomaa, T. Börner, and K. Sivonen. 2004. Phylogenetic evidence for the early evolution of microcystin synthesis. Proc. Natl. Acad. Sci. USA 101:568-573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Rantala, A., P. Rajaniemi-Wacklin, C. Lyra, L. Lepistö, J. Rintala, J. Mankiewicz-Boczek, and K. Sivonen. 2006. Detection of microcystin-producing cyanobacteria in Finnish lakes with genus-specific microcystin synthetase gene E (mcyE) PCR and associations with environmental factors. Appl. Environ. Microbiol. 72:6101-6110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Rinta-Kanto, J. M., and S. W. Wilhelm. 2006. Diversity of microcystin-producing cyanobacteria in spatially isolated regions of Lake Erie. Appl. Environ. Microbiol. 72:5083-5085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Rouhiainen, L., T. Vakkilainen, B. L. Siemer, W. Buikema, R. Haselkorn, and K. Sivonen. 2004. Genes coding for hepatotoxic heptapeptides (microcystins) in the cyanobacterium Anabaena strain 90. Appl. Environ. Microbiol. 70:686-692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rudi, K., F. Larsen, and K. S. Jakobsen. 1998. Detection of toxin-producing cyanobacteria by use of paramagnetic beads for cell concentration and DNA purification. Appl. Environ. Microbiol. 64:34-37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Rueda, F. J., and E. A. Cowen. 2005. The residence time of a freshwater embayment connected to a large lake. Limnol. Oceanogr. 50:1638-1653. [Google Scholar]
- 41.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- 42.Sivonen, K., and G. Jones. 1999. Cyanobacterial toxins, p. 41-111. In I. Chorus and J. Bartram (ed.), Toxic cyanobacteria in water: a guide to their public health consequences, monitoring and management. E & FN Spon, New York, NY.
- 43.Swofford, D. L. 1998. PAUP* phylogenetic analysis using parsimony (*and other methods), version 4. Sinauer Associates, Sunderland, MA.
- 44.Tanabe, Y., K. Kaya, and M. M. Watanabe. 2004. Evidence for recombination in the microcystin synthetase (mcy) genes of toxic cyanobacteria Microcystis spp. J. Mol. Evol. 58:633-641. [DOI] [PubMed] [Google Scholar]
- 45.Tillett, D., E. Dittmann, M. Erhard, H. von Döhren, T. Börner, and B. A. Neilan. 2000. Structural organization of microcystin biosynthesis in Microcystis aeruginosa PCC7806: an integrated peptide-polyketide synthetase system. Chem. Biol. 7:753-764. [DOI] [PubMed] [Google Scholar]
- 46.Urbach, E., D. Robertson, and S. Chisholm. 1992. Multiple evolutionary origins of prochlorophytes within the cyanobacterial radiation. Nature 355:267-270. [DOI] [PubMed] [Google Scholar]
- 47.Vaitomaa, J., A. Rantala, K. Halinen, L. Rouhiainen, P. Tallberg, L. Mokelke, and K. Sivonen. 2003. Quantitative real-time PCR for determination of microcystin synthetase E copy numbers for Microcystis and Anabaena in lakes. Appl. Environ. Microbiol. 69:7289-7297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Via-Ordorika, L., J. Fastner, R. Kurmayer, M. Hisbergues, E. Dittmann, J. Komarek, M. Erhard, and I. Chorus. 2004. Distribution of microcystin-producing and non-microcystin-producing Microcystis sp. in European freshwater bodies: detection of microcystins and microcystin genes in individual colonies. Syst. Appl. Microbiol. 27:592-602. [DOI] [PubMed] [Google Scholar]
- 49.Watzin, M. C., E. B. Miller, A. D. Shambaugh, and M. A. Kreider. 2006. Application of the WHO alert level framework to cyanobacterial monitoring of Lake Champlain, Vermont. Environ. Toxicol. 21:278-288. [DOI] [PubMed] [Google Scholar]
- 50.Welschmeyer, N. A. 1994. Fluorometric analysis of chlorophyll-a in the presence of chlorophyll-b and pheopigments. Limnol. Oceanogr. 39:1985-1992. [Google Scholar]
- 51.Whitford, L. A., and G. J. Schumacher. 1969. A manual of the fresh-water algae in North Carolina. The North Carolina Agricultural Experiment Station, Raleigh, NC.
- 52.Wilson, A. E., O. Sarnelle, B. A. Neilan, T. P. Salmon, M. M. Gehringer, and M. E. Hay. 2005. Genetic variation of the bloom-forming cyanobacterium Microcystis aeruginosa within and among lakes: implications for harmful algal blooms. Appl. Environ. Microbiol. 71:6126-6133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.World Health Organization. 2004. Guidelines for drinking-water quality, 3rd ed., vol. 1, p. 407-408. World Health Organization, Geneva, Switzerland. [Google Scholar]
- 54.Yoshida, M., T. Yoshida, Y. Takashima, R. Kondo, and S. Hiroishi. 2005. Genetic diversity of the toxic cyanobacterium Microcystis in Lake Mikata. Environ. Toxicol. 20:229-234. [DOI] [PubMed] [Google Scholar]
- 55.Zurawell, R. W., H. Chen, J. M. Burke, and E. E. Prepas. 2005. Hepatotoxic cyanobacteria: a review of the biological importance of microcystins in freshwater environments. J. Toxicol. Environ. Health B 8:1-37. [DOI] [PubMed] [Google Scholar]