Abstract
Adaptation to vertebrate blood feeding includes development of a salivary ‘magic potion’ that can disarm host hemostasis and inflammatory reactions. Within the lower Diptera, a vertebrate blood-sucking mode evolved in the Psychodidae (sand flies), Culicidae (mosquitoes), Ceratopogonidae (biting midges), Simuliidae (black flies), and in the frog-feeding Corethrellidae. Sialotranscriptome analyses from several species of mosquitoes and sand flies and from one biting midge indicate divergence in the evolution of the blood-sucking salivary potion, manifested in the finding of many unique proteins within each insect family, and even genus. Gene duplication and divergence events are highly prevalent, possibly driven by vertebrate host immune pressure. Within this framework, we describe the sialome (from Greek sialo, saliva) of the black fly Simulium vittatum and discuss the findings within the context of the protein families found in other blood-sucking Diptera. Sequences and results of Blast searches against several protein family databases are given in Supplemental Tables S1 and S2, which can be obtained from http://exon.niaid.nih.gov/transcriptome/S_vittatum/T1/SV-tb1.zip and http://exon.niaid.nih.gov/transcriptome/S_vittatum/T2/SV-tb2.zip.
Keywords: Simulium vittatum, black fly, sialotranscriptomes, salivary gland transcriptome, sialome, proteome, hematophagy, onchocerciasis
Introduction
The adaptation to blood feeding involves evolution of a complex cocktail of salivary components that help the blood sucker to overcome host defenses against blood loss (hemostasis) as well as inflammatory reactions at the feeding site that disrupt blood flow or cause pain and itching. Accordingly, saliva of blood-sucking arthropods contain anticlotting, antiplatelet, vasodilatory, antiinflammatory, and immunomodulatory components, usually in redundant amounts.1 Blood-feeding Diptera also take sugar meals and, perhaps for this reason, their salivary glands contain glycosidases and antimicrobial polypeptides that help sugar digestion and may prevent microbial growth in crop-stored sugar meals.
Black flies are important nuisance pests of humans and farm animals, as well as being vectors of arboviruses2, 3 and river blindness, caused by the worm Onchocerca volvulus.4 Black fly larvae are adapted to a running-water habitat, creating difficulties in establishing laboratory colonies. In spite of these difficulties, the North American black fly Simulium vittatum has been successfully colonized for over 20 years,5, 6 allowing a supply of standard material for laboratory studies. Salivary anticlotting (both anti-thrombin and anti-Xa) and vasodilatory proteins have been described from this fly.7-11 Salivary apyrase, a common enzymatic activity found in the saliva of hematophagous arthropods, has also been identified in S. vittatum and other Simuliidae.12 Apyrase hydrolyzes ATP and ADP to AMP and orthophosphate, thus eliminating the platelet and neutrophil aggregation properties of these nucleotides.13 Hyaluronidase activity has also been described in S. vittatum salivary gland homogenates.14 This enzyme may aid in diffusion of pharmacologically active compounds into the host skin. The presence of histamine in S. vittatum saliva, first proposed by Hutcheon and Chivers-Wilson,15 was detected in salivary secretions of paleartic black flies.16, 17 Except for the vasodilator SVEP (S. vittatum erythema protein),10 also known as Marydilan,13 and a protein with antithrombin activity (given in a patent application by Cupp and Cupp [2000] 18), no other defined polypeptide has been characterized from black fly salivary glands.
Black flies are classically grouped with the Nematocera suborder of Diptera in the Simuliidae family within the Culicomorpha infraorder. The Nematocera contain several families of blood-sucking flies, including mosquitoes (Culicidae), biting midges (Ceratopogonidae), and sand flies (Psychodidae). Salivary transcriptomes (sialotranscriptomes) have been made and described from several mosquito species, including the genera Anopheles,19-23, Aedes,24, 25 and Culex.26 Similarly, sialotranscriptomes have been described from Old and New World sand flies27-29 and from the biting midge Culicoides sonorensis.30 In these studies, family- and genus-specific proteins or whole protein families have been uncovered, indicating the rapid evolution and divergence of salivary proteins. For example, the powerful vasodilator Maxidilan31 is found only in New World sand flies, while adenosine serves as the main vasodilator in Old World sand flies.32, 33 The anopheline family of antithrombins is uniquely found in anophelines,34 while serpins function as the main anticlotting in Aedes,35 and Culicoides have abundant expression of Kunitz domain-containing salivary peptides.30 Several unique and expanded protein families with unknown function exist within different genera or family, such as the SG1 anopheline family,21 the 16.8-kDa family in Culex,26 and many others. In this work, we sequenced approximately 1,500 expressed sequence tags (ESTs) from a cDNA library made from S. vittatum salivary glands, uncovering for the first time the sialotranscriptome of a member of Simuliidae. Several new protein families were discovered that may have pharmacologic or antimicrobial activities and might serve as epidemiologic markers of Simulium exposure or be used as anti-disease vaccines.36
Materials and Methods
Chemicals
Standard laboratory chemicals were purchased from Sigma Chemicals (St. Louis, MO) if not specified otherwise. Formic acid and trifluoroacetic acid (TFA) were obtained from Fluka (Milwaukee, WI). Trypsin was purchased from Promega (Madison, WI). HPLC-grade acetonitrile was from EM Science (Darmstadt, Germany), and water was purified by a Barnstead Nanopure system (Dubuque, IA).
Black Flies
Colonized black flies were reared according to the protocol described in Bernardo et al.5 The history of the colony is given in Brockhouse et al.;6 the colony is the same as the one used in almost all previous studies of black fly saliva.7-10, 14 Adult females were collected within 4 hours after eclosion and stored at 4°C. To harvest mRNA, salivary glands were dissected in ice cold HEPES saline (10 mM HEPES/ 150 mM NaCl, pH 7.2) within 24 hours of adult eclosion, transferred to RNAlater (Ambion, Austin, TX), and stored at −70°C until use. As S. vittatum is an autogenous species, we also collected salivary glands from adult females 24-48 hours following their first oviposition, as they are competent to blood feed at this time and presumably have synthesized a full complement of salivary proteins. These glands, which were used for protein extraction, were stored in HEPES saline at −70°C until use.
Salivary Gland Isolation and Library Construction
S. vittatum mRNA from 60 pairs of salivary glands was isolated using the Micro-FastTrack mRNA isolation kit (Invitrogen, San Diego, CA). The PCR-based cDNA library was made following the instructions for the SMART cDNA library construction kit (Clontech, Palo Alto, CA). This system utilizes oligoribonucleotide (SMART IV) to attach an identical sequence at the 5/ end of each reverse-transcribed cDNA strand. This sequence is then utilized in subsequent PCR reactions and restriction digests.
First-strand synthesis was carried out using PowerScript reverse transcriptase at 42°C for 1 hour in the presence of the SMART IV and CDS III (3/) primers. Second-strand synthesis was performed using a long distance (LD) PCR-based protocol, using Advantage™ Taq polymerase (Clontech) mix in the presence of the 5/ PCR primer and the CDS III (3/) primer. The cDNA synthesis procedure resulted in creation of SfiI A and B restriction enzyme sites at the ends of the PCR products that are used for cloning into the phage vector. PCR conditions were as follows: 95°C for 20 sec; 24 cycles of 95°C for 5 sec., 68°C for 6 min. A small portion of the cDNA obtained by PCR was analyzed on a 1.1% agarose gel to check quality and range of cDNA synthesized. Double-stranded cDNA was immediately treated with proteinase K (0.8 μg/ml) at 45°C for 20 min, and the enzyme was removed by ultrafiltration though a Microcon (Amicon Inc., Beverly, CA) YM-100 centrifugal filter device. The cleaned, double-stranded cDNA was then digested with SfiI at 50°C for 2 hours, followed by size fractionation on a ChromaSpin– 400 column (Clontech). The profile of the fractions was checked on a 1.1% agarose gel, and fractions containing cDNAs of more than 400 bp were pooled and concentrated using a Microcon YM-100.
The cDNA mixture was ligated into the λ TriplEx2 vector (Clontech), and the resulting ligation mixture was packaged using the GigaPack® III Plus packaging extract (Stratagene, La Jolla, CA) according to the manufacturer’s instructions. The packaged library was plated by infecting log-phase XL1- Blue Escherichia coli cells (Clontech). The percentage of recombinant clones was determined by blue-white selection screening on LB/MgSO4 plates containing X-gal/IPTG. Recombinants were also determined by PCR, using vector primers (5/ λ TriplEx2 sequencing primer and 3/ λ TriplEx2 sequencing) flanking the inserted cDNA, with subsequent visualization of the products on a 1.1% agarose/EtBr gel.
Sequencing of the S. vittatum cDNA Library
The S. vittatum salivary gland cDNA library was plated on LB/MgSO4 plates containing X-gal/IPTG to an average of 250 plaques per 150-mm Petri plate. Recombinant (white) plaques were randomly selected and transferred to 96-well MICROTEST ™ U-bottom plates (BD BioSciences, Franklin Lakes, NJ) containing 100 μl of SM buffer [0.1 M NaCl; 0.01 M MgSO4; 7 H2O; 0.035 M Tris-HCl (pH 7.5); 0.01% gelatin] per well. The plates were covered and placed on a gyrating shaker for 30 min at room temperature. The phage suspension was either immediately used for PCR or stored at 4°C for future use.
To amplify the cDNA using a PCR reaction, 4 μl of the phage sample was used as a template. The primers were sequences from the λ TriplEx2 vector and named pTEx2 5seq (5/-TCC GAG ATC TGG ACG AGC-3/) and pTEx2 3LD (5/-ATA CGA CTC ACT ATA GGG CGA ATT GGC-3/), positioned at the 5/ and the 3/ end of the cDNA insert, respectively. The reaction was carried out in 96-well flexible PCR plates (Fisher Scientific, Pittsburgh, PA) using the TaKaRa EX Taq polymerase (TAKARA Mirus Bio, Madison, WI), on a Perkin Elmer GeneAmp® PCR system 9700 (Perkin Elmer Corp., Foster City, CA). The PCR conditions were: one hold of 95°C for 3 min; 25 cycles of 95°C for 1 min, 61°C for 30 sec; 72°C for 6 min. The amplified products were analyzed on a 1.5% agarose/EtBr gel. Approximately 200–250 ng of each PCR product was transferred to Thermo-Fast 96-well PCR plates (ABgene Corp., Epsom, Surrey, UK) and frozen at −20°C before cycle sequencing using an ABI3730XL machine.
Bioinformatic Tools and Procedures Used
ESTs were trimmed of primer and vector sequences, clusterized, and compared with other databases as previously described.22 The CAP3 assembler was used to assemble EST sequences,37 the BLAST tool was used to identify similar sequences in various databases,38 he ClustalW39 tool was used to align multiple sequences and TreeView version 1.6.6 software40 was used to visualize phylogenetic trees. Dendrograms were drawn by the neighbor-joining (NJ) method implemented in MEGA package (version 4.0), and bootstrap pseudoreplicate was performed to evaluate statistical significance of tree topology.41 For functional annotation of transcripts, we used the tool BlastX 42 to identify similar protein sequences to the NR protein database of the National Center for Biotechnology Information (NCBI) and to the Gene Ontology (GO) database.43 The tool, Reverse Position Specific Blast (RPSBlast),42 was used to search for conserved protein domains in the Pfam,44 SMART,45 Kog,46 and conserved domains databases (CDD) 47. The tool Seedtop, included in the stand-alone blast package, was used to search for PROSITE motifs.48 We have also compared the transcripts with other subsets of mitochondrial and rRNA nucleotide sequences downloaded from NCBI and to several organism proteomes downloaded from NCBI (yeast), Flybase (Drosophila melanogaster), or ENSEMBL (Anopheles gambiae). Segments of the three-frame translations of the EST (because the libraries were unidirectional, we did not use six-frame translations), starting with a methionine found in the first 300 predicted amino acids (AA), or to the predicted protein translation in the case of complete coding sequences, were submitted to the SignalP server49 to help identify translation products that could be secreted. O-glycosylation sites on the proteins were predicted with the program NetOGlyc50. Functional annotation of the transcripts was based on all the comparisons above. Following inspection of all these results, transcripts were classified as either Secretory (S), Housekeeping (H) or of Unknown (U) function, with further subdivisions based on function and/or protein families.
Proteomic characterization using one-dimensional gel electrophoresis and tandem mass spectrometry (MS)
The soluble protein fraction from salivary gland homogenates from S. vittatum corresponding to approximately 50 μg of protein was brought up in reducing Laemmli gel-loading buffer. The sample was boiled for 10 min and resolved on a_NuPAGE 4-12% Bis-Tris precast gel. The separated proteins were visualized by staining with SimplyBlue (Invitrogen). The gel was sliced into 32 individual sections that were destained and digested overnight with trypsin at 37°C. Peptides were extracted and desalted using ZipTips (Millipore, Bedford, MA) and resuspended in 0.1% TFA prior to MS analysis.
Nanoflow reversed-phase liquid chromatography tandem MS (RPLC-MS/MS) was performed using an Agilent 1100 nanoflow LC system (Agilent Technologies, Palo Alto, CA) coupled online with a linear ion-trap (LIT) mass spectrometer (LTQ, ThermoElectron, San José, CA). NanoRPLC columns were slurry-packed in-house with 5 μm, 300-Å pore size C-18 phase (Jupiter, Phenomenex, CA) in a 75-μm i.d. × 10-cm fused silica capillary (Polymicro Technologies, Phoenix, AZ) with a flame-pulled tip. After sample injection, the column was washed for 30 min with 98% mobile phase A (0.1% formic acid in water) at 0.5 μl/min, and peptides were eluted using a linear gradient of 2% mobile phase B (0.1% formic acid in acetonitrile) to 42% mobile phase B in 40 min at 0.25 μl/min, then to 98% B for an additional 10 min. The LIT-mass spectrometer was operated in a data-dependent MS/MS mode in which each full MS scan was followed by seven MS/MS scans where the seven most abundant molecular ions were dynamically selected for collision-induced dissociation (CID) using a normalized collision energy of 35%. Dynamic exclusion was applied to minimize repeated selection of peptides previously selected for CID.
Tandem mass spectra were searched using SEQUEST on a 20-node Beowulf cluster against an S. vittatum proteome database with methionine oxidation included as dynamic modification. Only tryptic peptides with up to two missed cleavage sites meeting a specific SEQUEST scoring criteria [delta correlation (ΔCn) ≥ 0.08 and charge-state-dependent cross correlation (Xcorr) ≥ 1.9 for [M+H]1+, ≥ 2.2 for [M+2H]2+ and ≥ 3.5 for [M+3H]3+] were considered as legitimate identifications. The peptides identified by MS were converted to Prosite block format 48 by a program written in Visual Basic. This database was used to search matches in the Fasta-formatted database of salivary proteins, using the poorly documented program Seedtop, which is part of the Blast package. The result of the Seedtop search is piped into the hyperlinked spreadsheet to produce a text file, such as the one shown for the apyrase proteins SV-2008, shown as a hyperlink here. Notice that the ID lines indicate, for example, BF18_73, which means that one match was found for fragment number 73 from gel band 18. Because the same tryptic fragment can be found in many gel bands, another program was written to count the number of fragments for each gel band, displaying a summarized result in an Excel table, as seen here, on cell AJ77 of Supplemental Table S2. The summary in the form of BF11 → 18| BF12 → 18| BF13 → 2| indicates that 18 fragments were found in Band 11, while 18 and 2 peptides were found in bands 12 and 13, respectively. Furthermore, this summary included a protein identification only when two or more peptide matches to the protein were obtained from the same gel slice. The summary program also produces additional spreadsheet cells with the larger number of peptides found in a single gel band, and the percent AA sequence coverage of the sum of the peptide matches, thus facilitating data analysis.
Results and Discussion
cDNA Library Characteristics
A total of 1,483 clones were sequenced and used to assemble a database (Supplemental Table S1) that yielded 698 clusters of related sequences, 561 of which contained only one EST. The consensus sequence of each cluster is named either a contig (deriving from two or more sequences) or a singleton (deriving from a single sequence). For sake of simplicity, this paper uses ‘cluster’ to denote sequences derived from both consensus sequences and singletons. The 698 clusters were compared using the program BlastX, BlastN, or RPSBlast 42 to the nonredundant protein database of the NCBI (NR database), a gene ontology database,43 the conserved domains database of the NCBI,47 and a custom-prepared subset of the NCBI nucleotide database containing either mitochondrial or rRNA sequences.
Because the libraries used are unidirectional, three-frame translations of the dataset were also derived, and open reading frames (ORFs) starting with a methionine and longer than 40 AA residues were submitted to SignalP server49 to help identify putative-secreted proteins. The EST assembly, BLAST, and signal peptide results were loaded into an Excel spreadsheet for manual annotation and are provided in Supplemental Table S1.
Three categories of expressed genes derived from the manual annotation of the contigs were created (Table 1). The putatively secreted (S) category contained 19% of the clusters and 51% of the sequences, with an average number of 5.8 sequences per cluster. The housekeeping (H) category had 26% and 19% of the clusters and sequences, respectively, and an average of 1.6 sequences per cluster. Fifty-six percent of the clusters, containing 30% of all sequences, were classified as unknown (U), because no functional assignment could be made. This category had an average of 1.2 sequences per cluster. A good proportion of these transcripts could derive from 3/ or 5/ untranslated regions of genes of the above two categories, as was recently indicated for a sialotranscriptome of An. gambiae.23
Table 1.
Transcript Abundance According to Functional Class
| Class | Clustersa | Sequencesa | Sequences/Cluster |
|---|---|---|---|
| Secreted | 131 (18.8) | 756 (51.0) | 5.77 |
| Housekeeping | 179 (25.6) | 279 (18.8) | 1.56 |
| Unknown | 388 (55.6) | 448 (30.2) | 1.15 |
| Total | 698 | 1483 |
Number (percent of total).
Housekeeping (H) Genes
The 179 clusters (comprising 279 ESTs) attributed to H genes expressed in the salivary glands of S. vittatum were further divided into 16 subgroups according to function (Table 2). Not surprisingly for an organ specialized for the secretion of polypeptides, the two larger sets were associated with protein synthesis machinery (50 clusters containing 95 ESTs) and energy metabolism (31 clusters containing 46 ESTs), a pattern also observed in other sialotranscriptomes.21, 26, 51. We have arbitrarily included a group of 67 ESTs (33 clusters) in the H category that represent highly conserved proteins of unknown function, presumably associated with cellular function. They are named conserved proteins of unknown function in Supplemental Table S1, immediately preceding the clusters of the unknown class. These sets may help functional identification of the ‘conserved hypothetical’ proteins as previously reviewed by Galperin and Koonin.52 The complete list of all 179 gene clusters, along with further information about each, is given in Supplemental Table S1.
Table 2.
Functional Classification of Housekeeping Transcripts
| Function | Clusters | Sequences | Sequences/Cluster |
|---|---|---|---|
| Protein synthesis | 50 | 95 | 1.90 |
| Unknown conserved | 33 | 67 | 2.03 |
| Metabolism, energy | 31 | 46 | 1.48 |
| Protein modification | 8 | 10 | 1.25 |
| Proteasome machinery | 10 | 10 | 1.00 |
| Protein export | 8 | 9 | 1.13 |
| Transcription machinery | 8 | 9 | 1.13 |
| Transporter/Storage | 6 | 8 | 1.33 |
| Cytoskeletal | 6 | 6 | 1.00 |
| Signal transduction | 6 | 6 | 1.00 |
| Metabolism, amino acid | 3 | 3 | 1.00 |
| Metabolism, carbohydrate | 3 | 3 | 1.00 |
| Transcription factors | 3 | 3 | 1.00 |
| Metabolism, detoxication | 2 | 2 | 1.00 |
| Metabolism, lipid | 1 | 1 | 1.00 |
| Nuclear regulation | 1 | 1 | 1.00 |
| Total | 179 | 279 |
Possibly Secreted (S) Class of Expressed Genes
Inspection of Supplemental Table S1 indicates the expression of several expanded gene families, including those coding for Kunitz-domain containing polypeptides, antigen-5 family members, odorant-binding/D7 protein families, vasodilatory proteins, and mucins (Table 3). Several proteins unique to Simuliidae were found, including the observation that the previously described SVEP 10 actually belongs to an expanded protein family.
Table 3.
Functional classification of transcripts coding for secreted proteins
| Family | Clusters | Sequences | Sequences/Cluster |
|---|---|---|---|
| Proline/Glutamine rich family | 19 | 209 | 11.00 |
| Amylase | 7 | 77 | 11.00 |
| SVEP | 19 | 67 | 3.53 |
| Odorant binding family | 8 | 64 | 8.00 |
| Immunity related peptides | 8 | 45 | 5.63 |
| Collagen-like | 4 | 36 | 9.00 |
| Mucins | 7 | 34 | 4.86 |
| Antigen 5 | 4 | 27 | 6.75 |
| Trypsins | 10 | 20 | 2.00 |
| Conserved secreted | 5 | 13 | 2.60 |
| Aegyptin-like | 2 | 10 | 5.00 |
| Apyrase | 2 | 3 | 1.50 |
| Yellow | 1 | 1 | 1.00 |
| Phospholipase | 1 | 1 | 1.00 |
| TIL domain peptide | 1 | 1 | 1.00 |
| Other putative secreted peptides | 26 | 127 | 4.88 |
| Total | 124 | 735 |
Analysis of the S. vittatum Sialome
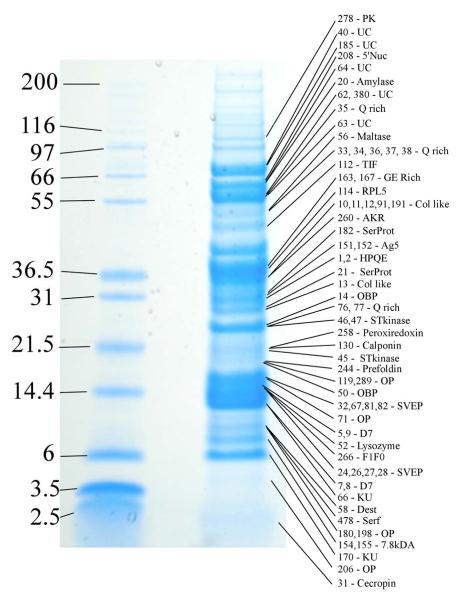
Several clusters of sequences coding for housekeeping and putative secreted polypeptides indicated in Supplemental Table S1 are abundant and complete enough to extract novel consensus sequences. Additionally, we have performed primer extension studies in several clones to obtain full- or near-full-length sequences of products of interest. A total of 117 novel sequences, 72 of which code for putative secreted proteins, are grouped together in Supplemental Table S2. Table 4 has a summary of the secreted subset, with links to GenBank. With this database in hand, we characterized the Simuliidae proteome via analysis of SDS-PAGE separated proteins and MS (Figure 1). The results of this experiment are integrated within the description of the deduced proteins from the transcriptome analysis, as outlined below.
Table 4.
Putative Secreted Proteins Deducted from the Salivary Transcriptome Analysis, and Indication of Expression by Proteomic Analysis
| Protein name |
gi number and link to NCBI |
Description | Fraction → number of peptidesa |
% Peptide coverageb |
|---|---|---|---|---|
| SVEP family of vasodilatory peptides | ||||
| SV-81 | 197260890 | Erythema protein SVEP-2 | F23 → 8| F22 → 7| | 63.4 |
| SV-82 | 197260892 | Erythema protein SVEP-3 | F23 → 6| F22 → 4| | 44.4 |
| SV-26 | 197260758 | Erythema protein SVEP-4 | F24 → 8| F23 → 6| | 71.2 |
| SV-24 | 197260750 | Erythema protein SVEP allele | F24 → 5| F23 → 4| | 39.5 |
| SV-27 | 197260764 | Erythema protein SVEP-5 | F24 → 7| F23 → 5| | 58.2 |
| SV-28 | 197260770 | Erythema protein SVEP-6 | F24 → 6| F23 → 4| | 46.4 |
| SV-67 | 197260864 | Erythema protein SVEP-7 | F23 → 5| | 46.1 |
| SV-32 | 197260792 | Erythema protein SVEP-8 | F23 → 7| F24 → 7| | 60.9 |
| Acidic H P Q E-rich proteins of low complexity | ||||
| SV-1 | 197260658 | Hypothetical secreted protein with HG-PEQ repeats | F18 → 2| F19 → 2| | 11.5 |
| SV-2 | 197260726 | Hypothetical secreted protein with HG-PEQ repeats 3 | F18 → 2| F19 → 2| | 10.6 |
| SV-3 | 197260780 | Hypothetical secreted peptide precursor with HG-PEQ repeats variant 2 | ||
| SV-4 | 197260816 | Hypothetical secreted protein with HG-PEQ repeats 4 | ||
| Acidic Q- rich family | ||||
| SV-33 | 197260794 | Hypothetical protein with QQQ repeats—truncated at 5 prime | F13 → 9| F12 → 8| | 80.3 |
| SV-34 | 197260796 | Hypothetical protein with QQQ repeats—truncated at 5 prime | F13 → 9| F12 → 8| | 92.7 |
| SV-35 | 197260798 | Hypothetical protein with QQQ repeats—truncated at 5 prime | F12 → 5| F13 → 5| | 77.0 |
| SV-36 | 197260800 | Hypothetical protein with QQQ repeats—truncated at 5 prime | F13 → 9| F12 → 8| | 100.0 |
| SV-37 | 197260804 | Hypothetical secreted protein with QQQ repeats | F13 → 9| F12 → 8| | 56.5 |
| SV-38 | 197260808 | Hypothetical secreted protein with QQQ repeats | F13 → 9| F12 → 8| | 56.5 |
| SV-76 | 197260883 | GM-2672 hypothetical secreted protein with QE repeats | F20 → 5| F21 → 4| | 50.3 |
| SV-77 | 197260885 | Hypothetical secreted protein with QQ repeats | F20 → 4| F21 → 3| | 44.7 |
| Collagen-like | ||||
| SV-10 | 197260660 | Collagen-like secreted salivary peptide | F17 → 17| F18 → 11| | 87.5 |
| SV-12 | 197260672 | Collagen-like salivary secreted protein | F17 → 22| F16 → 10| | 100.0 |
| SV-13 | 197260678 | Hypothetical secreted protein similar to Aedes mucin | F19 → 7| F18 → 3| | 42.2 |
| SV-11 | 197260664 | Hypothetical secreted with GK and KKKKKK domains protein | F17 → 8| F18 → 7| | 62.0 |
| SV-191 | 197260722 | Hypothetical secreted protein— truncated at 3 prime | F17 → 6| F18 → 6| | 61.6 |
| SV-91 | 197260898 | Hypothetical secreted protein with basic tail | F17 → 7| F18 → 7| | 63.6 |
| GE-rich protein | ||||
| SV-163 | 197260700 | Hypothetical protein—truncated at 5 prime | F16 → 13| F17 → 11| | 89.5 |
| SV-167 | 197260706 | Hypothetical secreted protein | F16 → 10| F17 → 9| | 100.0 |
| Mucins | ||||
| SV-75 | 197260879 | Hypothetical protein with QQQ and TTT repeats—truncated at 5 prime | ||
| SV-74 | 197260877 | Hypothetical protein with QQQ and TTT repeats—truncated at 5 prime | ||
| Odorant-binding protein family | ||||
| SV-14 | 197260684 | Hypothetical secreted protein with two odorant binding domains | F20 → 16| F21 → 8| | 80.3 |
| SV-50 | 197260843 | Salivary D7 secreted protein | F22 → 5| F23 → 5| | 37.1 |
| Simulidin family—Polymorphic 13-kDa protein similar to the D7/SP-15 family of phlebotomines | ||||
| SV-9 | 197260896 | Hypothetical secreted salivary protein | F24 → 2| | 43.8 |
| SV-7 | 197260866 | Salivary OBP/D7 family mamber | F25 → 2| | 34.8 |
| SV-8 | 197260887 | SV short D7 protein-1 | F25 → 2| | 34.8 |
| SV-5 | 197260841 | Short salivary D7 protein | F24 → 2| | 43.8 |
| Antigen-5 family | ||||
| SV-151 | 197260686 | Salivary secreted antigen 5 related protein | F18 → 12| F17 → 8| | 49.5 |
| SV-152 | 197260688 | Salivary secreted antigen 5 related protein | F18 → 12| F17 → 8| | 49.5 |
| Kunitz domain-containing peptides | ||||
| SV-170 | 197260710 | Single Kunitz protease inhibitor | F29 → 3| | 41.0 |
| SV-66 | 197260862 | Single Kunitz protease inhibitor | F25 → 2| F26 → 2| | 24.5 |
| Orphan peptides | ||||
| 5 Cys family | ||||
| SV-72 | 197260873 | Hypothetical 5-Cys secreted protein | ||
| SV-73 | 197260875 | Hhypothetical secreted protein with 5 Cys | ||
| Sv 7.8-kDa family | ||||
| SV-154 | 197260690 | Hypothetical salivary secreted protein | F28 → 4| F30 → 4| | 72.8 |
| SV-155 | 197260692 | Hypothetical secreted peptide precursor | F28 → 4| F30 → 4| | 73.6 |
| Sv 4.8-kDa family | ||||
| SV-158 | 197260694 | Similar to artifact salivary secreted peptide | ||
| SV-159 | 197260696 | Hypothetical secreted peptide precursor | ||
| SV-160 | 197260698 | Hypothetical secreted peptide precursor | ||
| Sv 7.0-kDa family | ||||
| SV-179 | 197260714 | Hypothetical secreted peptide precursor | ||
| SV-178 | 197260712 | Hhypothetical secreted peptide precursor | ||
| Other orphan peptides | ||||
| SV-180 | 197260716 | Hypothetical secreted protein | F27 → 3| F24 → 2| | 46.2 |
| SV-71 | 197260870 | Hhypothetical secreted protein | F23 → 5| F26 → 5| | 40.0 |
| SV-198 | 197260724 | Hypothetical secreted protein | F27 → 4| F30 → 4| | 41.0 |
| SV-206 | 197260734 | Hhypothetical secreted peptide precursor | F30 → 8| F26 → 7| | 85.7 |
| SV-119 | 197260670 | Hhypothetical secreted protein with 7 cys | F22 → 5| F21 → 3| | 27.6 |
| SV-289 | 197260772 | Hypothetical secreted protein | F22 → 3| F21 → 2| | 38.8 |
| SV-123 | 197260674 | Hypothetical secreted protein | ||
| Enzymes | ||||
| 5/-nucleotidase/apyrase | ||||
| SV-208 | 197260736 | Putative apyrase/nucleotidase—truncated at 5 prime | F11 → 18| F12 → 18| | 61.6 |
| Serine proteases | ||||
| SV-21 | 197260738 | Salivary serine protease | F19 → 3| F20 → 3| | 16.5 |
| SV-182 | 197260718 | Salivary serine protease | F18 → 10| F19 → 4| | 44.9 |
| SV-465 | 197260833 | Trypsin—truncated at 3 prime | ||
| SV-164 | 197260702 | Serine protease similar to hypodermin—truncated at 5 prime | ||
| SV-168 | 197260708 | Salivary serine protease similar to hypodermin—truncated at 5 prime | ||
| SV-165 | 197260704 | Salivary serine protease—truncated at 5 prime | ||
| Destabilase | ||||
| SV-58 | 197260851 | Salivary destabilase | F27 → 2| | 15.9 |
| Lysophospholipase | ||||
| SV-383 | 197260812 | Lysophospholipase—truncated at 5′ | ||
| Amylase/maltase | ||||
| SV-20 | 197260728 | Salivary alpha-amylase—truncated at 5 prime | F11 → 24| F12 → 8| | 75.1 |
| SV-56 | 197260849 | Salivary alpha-amylase—truncated at 5 prime | F13 → 7| F12 → 6| | 43.2 |
| Immunity-related proteins | ||||
| Lysozyme | ||||
| SV-52 | 197260845 | Salivary lysozyme | F24 → 4| | 42.6 |
| Cecropin | ||||
| SV-133 | 197260682 | Cecropin precursor | ||
| SV-31 | 197260786 | Salivary expressed cecropin precursor | F32 → 3| F30 → 2| | 50.0 |
| SV-201 | 197260730 | Hypothetical secreted peptide precursor | ||
| Gram-negative binding protein | ||||
| SV-313 | 197260790 | Gram-negative bacteria binding protein—fragment | ||
Indicates band location on Figure 1 gel where matching peptides from tryptic digests were found. Convention for F# → X is for fraction #, X number of peptides were found. Results show only two most abundant fractions. For more details, see Supplemental Table S2.
Indicates percentage coverage of predicted protein with matching peptides in the gel fraction with most fragments.
Figure 1.
1D gel electrophoresis of Simulium vittatum salivary gland homogenates. The left lane shows the protein MW markers (kDa). The right lane shows the fractionated salivary gland proteins, with the proteins identified from various locations within the gel indicated. More detailed information about the proteins can be found in Supplemental Table S2 by appending the number shown to the prefix SV-. Abbreviations for the protein class are, from top to bottom: PK, protein kinase; UC, unknown conserved; 5/Nuc, 5/ nucleotidase/apyrase; Q rich, glutamine-rich protein family; TIF, translation initiation factor; RPL5, ribosomal protein L5; Col like, collagen like; AKR, aldo-keto reductase; SerProt, serine protease; Ag5, antigen 5; HPQE, HPQE-rich family; OBP, odorant-binding protein; STkinase, serine/threonine kinase; OP, orphan protein; SVEP, S. vittatum erythema protein; D7, D7 protein family; KU, Kunitz protein; Dest, destabilase; Serf, similar to SERF-like protein; 7.8 kDa, 7.8-kDa family.
SVEP
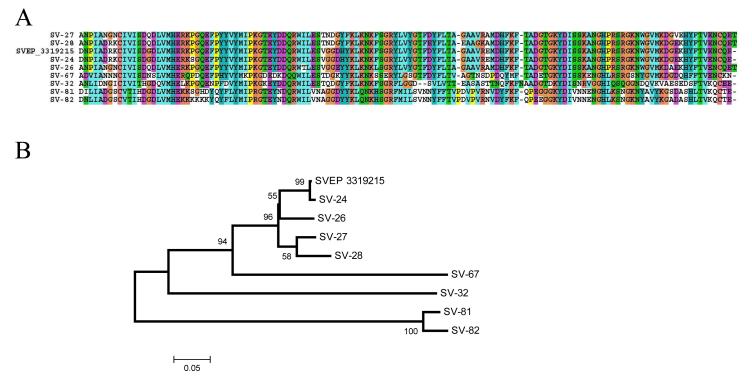
Salivary homogenates from various black fly species were shown to produce a prolonged vasodilatation in rabbit skin. A vasodilatory salivary polypeptide was isolated from S. vittatum,11 leading to the production of a recombinant protein named rSVEP, which had potent vasodilatory activity.10 rSVEP has a unique sequence, yielding no similar matches to any other protein in the NR database. Analysis of the sialotranscriptome of S. vittatum indicates that in addition to the rSVEP sequence, several other similar sequences exist, some of which were 50% identical to rSVEP. The alignment and dendrogram of nine related sequences indicates that possibly five genes exist, some of which are polymorphic based on an AA sequence difference of < 5% (such as SVEP and SV24/SV26, and the pairs SV-27/SV-28 and SV-81/SV-82). It is interesting to note that sv-contig_25, a cluster assembled from two ESTs, matches 100% of the rSVEP AA sequence, while clusters 81 and 82 (leading to the sequences SV-81 and SV-82 in Figure 2) have 17 and 12 ESTs, respectively, indicating these to be more abundantly expressed than rSVEP.
Figure 2.
The S. vittatum erythema protein (SVEP) family. (A) ClustalW alignment of SVEP (gi|3319215) with eight novel proteins of the same family. (B) Dendrogram derived from the alignment. The mature (signal peptide removed) protein sequences were aligned by the Clustal program,39 and the dendrogram was done with the Mega package41 after 10,000 bootstraps with the neighbor-joining (NJ) algorithm. The numbers on the tree bifurcations indicate the percentage bootstrap support above 50%. The bar at the bottom represents 5% AA substitution.
Four SVEP members were identified from a densely stained band excised from the SDS-PAGE gel shown in Figure 1. This band migrated just above the 14.4-kDa marker, consistent with the predicted mature molecular masses of the SVEP proteins. Four additional members were identified in an adjacent intensely stained band indicating a slightly larger molecular weight (MW). The identified peptides covered over 40% of the predicted protein sequence, indicating a high level of expression of these proteins.
Acidic, low complexity proteins rich in His, Pro, or Gln
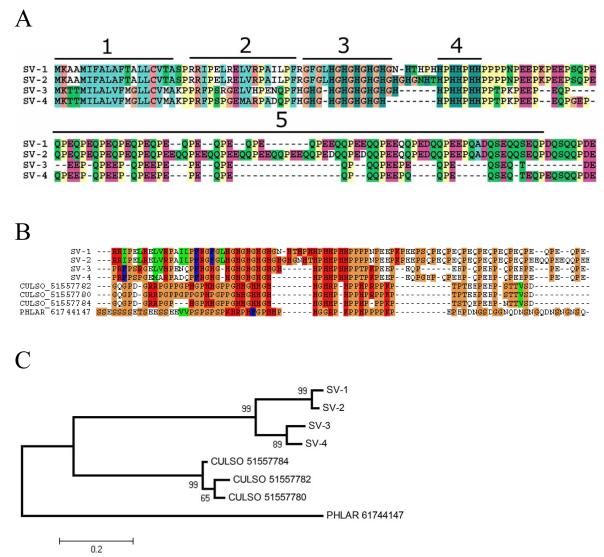
The most abundantly expressed cluster found in the S. vittatum sialotranscriptome, with 51 ESTs, encodes a secreted salivary protein of low complexity that is rich in His and Pro, Gln, and Glu residues (shown in Supplemental Table S2 as SV-1). When the non-redundant database is searched with this protein sequence as query by Blastp (with the filter of low complexity turned off, – F F command line option), bacterial proteins are retrieved but no matches to metazoan proteins are seen. Three other protein sequences are similar to SV-1, with their alignment indicating a similar signal peptide, an Arg rich region, a Gly-His repeat domain, a His-Pro-His repeat domain, a loose Gln-Pro-Glu repeat domain, and a C-terminus ending with the common acidic dyad Asp-Glu (Figure 3A). These four protein sequences account for nearly 10% of all ESTs within the current library. Their repeat nature suggests they may interact with matrix proteins, possibly collagen, and function in a manner analogous to mosquito aegyptins, which inhibit collagen-induced platelet aggregation.53 Of interest, when the nucleotide sequence of these proteins is compared with Culicidae sequences deposited on the NR database using the program blastx, additional protein matches are retrieved. These matches are all from salivary proteins of Nematocera containing a histidine-rich domain followed by an acidic domain, indicating these proteins may derive from a common ancestral gene (Figure 3B). The bootstrapped tree from the alignment of the Nematocera proteins (Figure 3C) shows distinct species clades, including two subclades within S. vittatum, indicating at least two genes exist in the black fly coding for proteins of this family.
Figure 3.
The S. vittatum GHPQ-rich protein family. (A) Clustal alignment of the S. vittatum proteins indicates five domains: 1) signal peptide indicative of secretion, 2) arginine-rich region, 3) HG repeat region, 4) HPH repeat region, and 5) QPE repeat region. (B) Alignment of regions 2, 3, 4, and part of 5 with salivary proteins from Culicoides sonorensis (CULSO) and Phlebotomus argentipes (PHLAR). (C) Bootstrapped dendrogram from the alignment in (B). The number following the non-Simulium protein names refers to the GenBank GI number.
Expression of SV-1 and SV2 was identified by tryptic peptides obtained from a protein band that migrated near the 31-kDa marker, which is about twice the predicted MW of the mature proteins (Figure 1). Only 10% coverage of these proteins was obtained by MS analysis, possibly owing to the limited number of MS-compatible tryptic peptides that are produced by these highly negatively charged proteins.
Acidic Gln-rich proteins
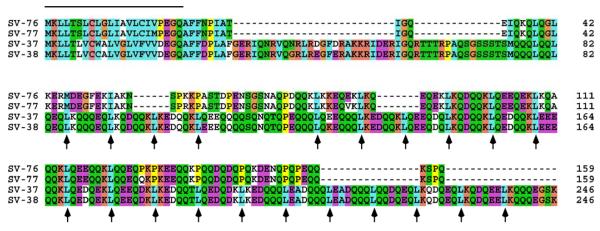
Another unique family of putative salivary secreted proteins is characterized by acidic polypeptides of low complexity and rich in Gln residues. At least two genes may code for proteins of mature MW of either 16 or 27 kDa. Four of these sequences are shown in Figure 4, two for the short and two for the longer proteins. It is possible that these four proteins correspond to alleles of two genes or to a larger number of genes coding for similar products. Other possible alleles are shown in Supplemental Table S2. Notice that this family of proteins has many Leu residues at every seventh position (arrows in Figure 4), characterizing a Leu zipper repeat known to be involved in dimerization of proteins.54, 55 This protein family is well transcribed in S. vittatum salivary glands, as there are 27 ESTs coding for SV-38 alone. These protein sequences do not identify similar proteins in the NR database, except for repeat regions of diverse products. The function of this protein family may be related to the binding to host extracellular matrix proteins or receptors, as is the case with the Aegyptin/GE-rich proteins of mosquitoes, which bind to collagen53.
Figure 4.
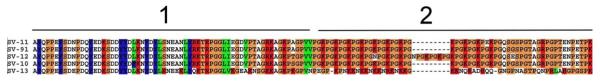
The S. vittatum QE-rich protein family. Clustal alignment indicates at least two genes coding for the shorter proteins, named SV-76 and SV-77, and two larger proteins, named SV-37 and SV-38. The bar at the top of the alignment indicates the signal peptide region, the mature proteins starting with the Phe-Phe doublet. The arrows point to leucine repeats spaced by 6 AA.
Consistent with their abundant EST expression, a large number of tryptic peptides originating from members of this protein family were identified by MS. Tryptic peptides for SV-76 and SV-77, both having a predicted mature MW of 16 kDa, were found in a gel band (Figure 1) between the 21.5- and 31-kDa markers, indicating they may be post-translationally modified. The predicted larger protein members (MW of 27 kDa) encoded by SV-37 and SV-38 were also identified in a location consistent with a larger mass, between the 36.5- and 55-kDa markers, while SV-35, derived from a 5/ truncated clone, was identified in a gel band located at the 55-kDa marker.
Collagen-like basic proteins
Several putative proteins in the sialotranscriptome of S. vittatum were found to be similar to collagen by virtue of their Pro-Gly repeats. These putative mature proteins vary from 12–27 kDa and are all basic in nature, with a minimum pI of 9.4. Alignment of the deduced sequences of five proteins indicates the presence of an acidic amino terminal region followed by a basic region containing Pro-Gly-Lys repeats (Figure 5). It appears that this family contains at least two genes, based on the divergence of the sequences, particularly that of SV-13. The carboxy terminal region of these putative proteins, not shown in Figure 5, is divergent. This result could be either real or artifactual in nature due to polymerase slippage, which is common in DNA repeats.56 These protein sequences do not identify similar proteins in the NR database, except for repeat regions of diverse products. The Pro-Gly repeats of this family, reminiscent of collagen57, suggests it may interact with matrix components or receptors.
Figure 5.
The S. vittatum collagen-like protein family. Clustal alignment indicates a mature amino terminal region of higher complexity and rich in acidic residues (1), then a basic carboxyterminal region with Pro-Gly-Lys repeats. The alignment excludes the signal peptide region and the carboxy termini of the proteins, which are divergent.
This divergent protein family contains members with predicted MW varying from 12-27 kDa. The proteome analysis (Figure 1) identified several protein members having an apparent molecular mass of ~36 kDa, which is larger than their expected masses. SV-13, with a predicted mature MW of 20 kDa, was identified between the 21.5- and 31-kDa markers, indicating a possible post-translation modification of this protein.
GE-rich/30-kDa allergen distant family members
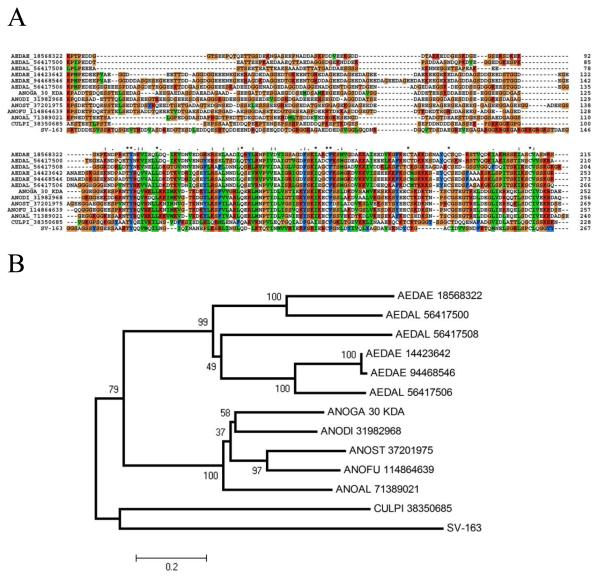
Members of the 30-kDa allergen (also described as GE-rich proteins) are ubiquitously and exclusively found in the salivary glands of adult female mosquitoes. Recombinant members of this family are known to be antigenic to humans.58 Only one member of this family is found in the An. gambiae genome,23 but at least three genes exist in Aedes albopictus,24 and two genes were found in Aedes aegypti.25 Other mosquito transcriptomes have yielded similar proteins, including other anopheline species and Culex pipiens quinquefasciatus. Recently, a family member from both Ae. aegypti and Anopheles stephensi were shown to inhibit collagen-induced platelet aggregation, indicating a conserved function of this protein family53, 59 after 150 million years of evolutionary history, the time estimated from the Anopheline/Culicine divergence.60 The sialotranscriptome of S. vittatum yielded 10 ESTs matching members of this protein family. The deduced protein sequence encoded in SV-163 (derived from a 5/ truncated clone) aligns well with members of the family (Figure 6A) showing an N-terminal domain rich in acidic and Gly residues and a C terminal domain richer in aliphatic and Lys residues; however, sequence identity levels are only about 30% when comparing the black fly sequences with those of mosquitoes. The bootstrapped phylogram shows robust clades for Anopheles and Aedes, with Culex and Simulium constituting individual branches (Figure 6B). The finding of a member of this unique protein in Simulium can be explained by convergent evolution or, alternatively, supports a common blood-feeding ancestor for black flies and mosquitoes, as previously proposed.61
Figure 6.
The GE-rich/30-kDa antigen family of mosquitoes and S. vittatum. (A) Clustal alignment. Notice the first half of the alignment is dominated by Gly, Asp, and Glu, and the second part is dominated by hydrophobic as well as by lysine residues. The signal peptide region is not shown. The S. vittatum protein is represented by SV-163. The remaining proteins are from Aedes aegypti, Ae. albopictus, Anopheles albimanus, An. gambiae, An. stephensi, An. dirus, An. funestus, and Culex pipiens, recognized by the three letters of the genus name followed by the first two letters of the species name, followed by the NCBI GI number, except for the An. gambiae protein, which was obtained from Arca et al.23 The symbols above the alignment indicate identity (*), high similarity (:), and similarity (.) of residues in the indicated alignment position. (B) Phylogram following 10,000 bootstraps; numbers indicate the percentage of bootstrap support.
Members of this family were identified at a gel location coincident with the 36.5-kDa marker. Because our clones were truncated at the 5/ end, we do not have estimates of the predicted mature MW to compare with their observed gel migrations.
Mucins
Mucins are usually low-complexity proteins rich in Thr and/or Ser residues to which galactosyl residues are attached. These proteins are commonly found in sialotranscriptomes of blood-sucking arthropods. They may function in the maintenance of the salivary gland ducts (many such proteins have chitin-binding domains), or their galactosylation may assist some other function (e.g., some double-Kunitz proteins in ticks have a Ser/Thr-rich carboxy terminus62). Two protein fragments were identified from the sialotranscriptome of S. vittatum as having over 100 galactosylation sites, as predicted by the NetOglyc server;63 however, these sequences lack a chitin-binding domain. The protein sequence of SV-75 is also a member of the acidic Gln-rich family described above but uniquely contains a long carboxy terminus rich in Ser/Thr residues. The second mucin described in Supplemental Table S2, SV-74, has more than 50% of its AA residues consisting of Ser and Thr.
Although no mucins were identified by more than a single tryptic fragment from the gel shown in Figure 1, a single fragment, not containing Ser or Thr residues, indicated that SV-75 migrates to a gel location coincident with the 116-kDa marker. It is possible that multiple glycosylations of these proteins may have interfered with the identification of these proteins by the SDS-PAGE and MS/MS strategy used in this study.
Putative D7/odorant-binding protein (OBP) family
The D7 protein from Ae. aegypti was the first mosquito salivary protein to be cloned.64 It was later found to be a member of the OBP superfamily.65 Members of this family have been found in all mosquito transcriptomes so far studied. Single- or double-domain proteins exist, constituting the short and long D7 subfamilies.66 Ae. aegypti and An. gambiae have at least eight genes encoding these proteins, with three genes encoding the long forms and five encoding the short forms. Sand flies from the genera Lutzomyia and Phlebotomus, as well as members of the genus Culicoides also have these salivary proteins. Mosquito proteins have recently been shown to bind biogenic amines,67 thus helping blood feeding by sequestering vasoconstrictory and platelet aggregation agonists. This function, however, may be shared by only some members of the family, as other members have been shown to bind leukotrienes (E. Calvo, personal communication). The crystal structure of a short D7 protein from An. gambiae has been solved, showing that its OBP domain has two C-terminal alpha helices that are not present in canonical OBP proteins.68 In Simulium, we find both short and long D7 proteins, thus extending the presence of this protein family into all families of blood-sucking Nematocera flies. The long D7 protein sequence encoded by SV-14 has two weak OBP domains as indicated by RPSBlast against the Smart database and also shows weak similarity to phlebotomine proteins of the D7 protein family. Ten ESTs were found to code for SV-14, indicating that it is relatively highly expressed; however, we failed to identify a signal peptide indicative of secretion, possibly indicating truncation of the clone. Eighty percent peptide coverage for this protein was found by MS, revealing 16 tryptic peptides originating from a single densely stained gel band (Figure 1) located between 21.5 and 31 kDa, which is consistent with the predicted MW of this protein.
SV-50 represents a protein sequence displaying a signal peptide indicative of secretion and a predicted weight for the mature polypeptide of 16 kDa. It has a single OBP domain and is weakly homologous to mosquito D7 proteins when compared to a database of salivary proteins of Diptera. SV-50 was identified in the SDS-PAGE gel of Figure 1 at its expected location, between the 14.4- and 21.5-kDa MW markers.
Phlebotomine sand flies have an expanded family of short D7 proteins, the first of which, named SP-15, was used as an antigen to protect mice from leishmaniasis.69 The S. vittatum sequences SV-5, SV-7, SV-8, and SV-9 are quite similar to each other but only weakly similar to a protein member of the SP-15 family, to which they may belong, or they may constitute a unique family. Evidence for the expression of these proteins was found by two tryptic peptides originating from a band that migrated to a location consistent with its predicted mature MW (Figure 1). SV-7 is identical to the Simulium anti-thrombin sequence reported by Cupp and Cupp (2000) in a patent application. This corresponds to the protein named simulidin by Abebe et al. in 1995.7 Given the high degree of similarity of the members of this cluster, it is likely that they all encode antithrombins. If these proteins are indeed related to the short D7 proteins, this would be a novel biologic activity for the family, analogous to the evolution of anti-kallikrein activity by a short D7 protein from An. stephensi. 70
Antigen 5 protein family
AG5-related salivary products are members of a group of secreted proteins that belong to the CAP family (Cys-rich secretory proteins; AG5 proteins of insects; pathogenesis-related protein 1 of plants).71 Members of this protein family are found in most blood-sucking insect sialotranscriptomes. The majority of these animal proteins have no known function. The notable exceptions include proteolytic activity in Conus,72 smooth muscle-relaxing activity in snake venoms,73, 74 and salivary neurotoxin activity in the venomous lizard Heloderma horridum.75 Recently, an antigen-5 protein from the saliva of a tabanid fly76 was shown to inhibit platelet aggregation by the unusual acquisition of a typical RGD domain that is known to prevent fibrinogen binding to platelets and its ensuing aggregation.77 No function has been determined for any Nematoceran antigen-5 protein. The S. vittatum sialotranscriptome revealed the presence of at least one member of the antigen-5 family, which is possibly polymorphic, and most homologous to protein sequences retrieved from phlebotomine and mosquito sialotranscriptomes. The MS results (Figure 1) identified SV-151 and SV-152 at a location coincident with the 31-kDa marker, in accordance with their predicted MW of 32 kDa.
Kunitz-domain containing peptides
The Kunitz domain is associated with ubiquitous proteins having serine protease inhibitor activity.78, 79 Many tick salivary anticlotting peptides belong to this protein family.80-82 In blood-sucking Diptera, however, no anticlotting protein with a Kunitz domain has been biochemically characterized. Anopheles mosquitoes have the unique anopheline family of peptides that act as antithrombin agents,34, 83 while Aedes have an anti-Xa serpin.35 No transcripts coding for Kunitz domains have been found in transcriptomes of mosquitoes or sand flies. Characterization of the sialotranscriptome of the biting midge Culicoides sonorensis, however, revealed at least four different Kunitz-domain-containing polypeptides,30 suggesting this family of proteins may function as the salivary Xa-ase inhibitor previously described in Culicoides variipennis.84 The sialotranscriptome of S. vittatum revealed 10 ESTs coding for members of the Kunitz family. Supplemental Table S2 presents two full-length sequences, each having both a signal sequence and a typical Kunitz domain. Both peptides were identified by MS (Figure 1), with SV-170 corresponding to an intensely stained gel band near the 6-kDa marker and SV-66 corresponding to another intensely stained band just below the 14.4-kDa marker, consistent with their predicted mature MW of 9 and 12 kDa, respectively. These peptides may be responsible for the previously described anti-Xa activity of S. vittatum salivary gland homogenates.7, 85
Orphan putative secreted peptides
Mining of the sialotranscriptome of S. vittatum lead to the discovery of 16 additional polypeptides grouped in 11 families without any similarity to other known protein sequences or to conserved domains available in the CDD, KOG, PFAM, SMART, or PROSITE databases. Some of these are highly transcribed, such as the two possible alleles of the 5Cys family (with 27 ESTs), the Sv7.8 family (SV-154 and SV-155) with 21 ESTs, and the Sv7.0 family (SV-179 and SV-178), with 11 ESTs. The remaining seven unique polypeptides are represented from between one and eight ESTs each. Several of these polypeptides were identified in the proteome experiment shown in Figure 1. The function of these polypeptides remains to be identified.
Enzymes
Apyrase/5/-nucleotidase
Enzymes that destroy ADP and ATP are ubiquitously found in saliva of blood-sucking arthropods, perhaps because of the importance of these nucleotides in activating platelet and neutrophil aggregation.1 Different families of enzymes have been recruited for this function by different arthropod genera in their convergent evolution to blood feeding. Mosquitoes and Triatoma have recruited the 5/-nucleotidase family,86-88 while sand flies and bed bugs have recruited the Cimex family of apyrases.89, 90 Fleas may have recruited a third family, the CD-39 family of ectonucleotidases.91 All nucleotidases require divalent cations to function, with the Cimex family being unique in showing dependence on Ca2+ ions, while the other two families can function with either Ca2+ or Mg2+ ions. A salivary apyrase previously described in S. vittatum and other black flies is activated by either Ca2+ or Mg2+ ions,12 suggesting that it is different from the enzyme found in phlebotomines. The sialotranscriptome of S. vittatum produced three ESTs from which a 417-AA residue protein sequence could be derived that matches different 5/ nucleotidases previously found in blood-sucking Diptera salivary glands. SV-208 matches the salivary apyrases of Anopheles, Aedes, and Chrysops including the protein annotated as chrysoptin precursor, described as an inhibitor of collagen-induced platelet aggregation found in the salivary glands of a deer fly.92 Eighteen SV-208 peptide fragments were identified in an intensely stained gel band by MS (Figure 1) migrating to its expected location near the 66-kDa marker. These findings suggest that S. vittatum apyrase may belong to the 5/-nucleotidase family, as do mosquito apyrases.86, 87
Serine proteases
Protein sequences with similarity to serine proteases have been commonly detected in the sialotranscriptomes of blood-sucking arthropods, where they may function in the immune system, acting as prophenoloxidase activators, or enhance the flow of blood by digesting fibrin or fibrinogen, as occurs in ticks.93 The sialotranscriptome of S. vittatum reveals a variety of transcripts encoding this type of enzyme, with at least four gene products having a serine protease PFAM signature represented. SV-21 encodes a secreted serine protease best matching vertebrate proteases, some of which have been annotated as elastases. This protein was identified by MS (Figure 1) from a gel band that migrated just below the 31-kDa marker. SV-182 also encodes a secreted serine protease that matches invertebrate and vertebrate enzymes, but its best match has only 22% identity, indicating the black fly enzyme to be very divergent. SV-182 was identified by MS from a gel band that migrated near the 36.5-kDa marker. SV-465 contains only the 3/ portion of the gene and is best matched by a protein annotated as Aedes elastase. SV-164, SV-165 ,and SV-168, all 3/ truncated products, appear to result from the same gene by alternative splicing or gene duplication events. These protein sequences produce a weak match to hypodermin, a collagenase expressed in the larval instars of the cattle grub Hypoderma lineatum.94, 95 Because black flies are pool feeders and their salivary glands have hyaluronidase activity,14 it is reasonable to infer that other enzymes may exist in S. vittatum saliva to help create a larger feeding cavity, such as collagenases and elastases. Except for the horse fly Tabanus yao, where two serine proteases were biochemically characterized to have fibrinogenolytic and anticlotting activities,76 no other salivary serine protease from blood-sucking Diptera has been biochemically characterized.
Destabilase
Three ESTs in the S. vittatum sialotranscriptome code for an enzyme having the Destabilase motif. Destabilase is an endo-ξ-(γ-Glu)-Lys isopeptidase, which cleaves isopeptide bonds formed by transglutaminase (Factor XIIIa) between Gln γ-carboxamide and the ξ-amino group of lysine. This enzyme activity leads to dissolution of stabilized fibrin. Destabilase was first described in the salivary glands of the leech Hirudo medicinalis,96 later shown to be the product of a multigene family that is related to the lysozyme superfamily.97, 98 Two tryptic fragments matching SV-58 were identified from a gel band that migrated between the 6- and 14.4-kDa markers, consistent with SV-58 predicted MW of 12 kDa. This result is the first time a member of this protein family has been described in sialotranscriptomes of blood-sucking arthropods.
Amylase/maltase
Enzymes associated with digestion of carbohydrates have regularly been found in the salivary glands of adult mosquitoes and sand flies.19, 28, 99-101 In the present work, we found evidence for the expression of two salivary glycosidases in S. vittatum, in the form of two mRNA fragments. One is similar to a salivary amylase of Aedes, while the other is more similar to a Drosophila enzyme. Both enzymes were identified in the proteome experiment, one each from gel bands migrating near the 66- and 55-kDa markers, consistent with the typical size of these enzymes.
Immunity-related polypeptides
Probably with a function to prevent microbial growth in both sugar and blood meals, the salivary glands of blood-sucking arthropods produce both antimicrobial peptides (AMP) and proteins associated with pathogen pattern recognition motifs, which may be important for opsonization of the microorganism for further recognition by complement-like thioester proteins102 or by the phenoloxidase cascade.1, 103 Evidence of such proteins was observed in the sialotranscriptome of S. vittatum in the form of a typical lysozyme protein that had highest similarity to Lepidoptera peptides, three cecropins that were only recognized through a weak match of one of the members to a previously annotated Culex cecropin, and a 5/ truncated member of the Gram-negative binding protein family that is similar to mosquito proteins. Both the lysozyme and cecropin proteins were recognized in the proteome experiment by at least two tryptic fragments from a single gel band (Figure 1), while the Gram-negative binding protein had a single tryptic fragment identified from a gel band that migrated to a position consistent with the typical size of this protein, between the 31- and 36.5-kDa markers (data not shown).
Housekeeping Proteins
Supplemental Table S2 presents sequence information on 45 proteins classified as housekeeping, several of which were identified in the proteome experiment (Figure 1).
Conclusion
Analysis of the sialome of S. vittatum, the first done for this family of blood-feeding flies, uncovers both the common and divergent evolutionary pathways taken in producing today’s salivary ‘magic potion’ of such arthropods. Blood feeding has been proposed as an ancestral (plesiotypic) feeding mode in the infraorder Culicomorpha, which includes the families Culicidae (mosquitoes), Ceratopogonidae (biting midges), and Simuliidae (black flies);61, 104 however, Pawloswki et al.105 concluded that nectar feeding was plesiotypic in the Culicomorpha, with independent evolution of hematophagy in mosquitoes, black flies, and ceratopogonids. If these dipteran families share a common blood-feeding ancestor, we could expect significant overlap in the presence of protein families, with novel features in each insect family representing evolutionary innovations developed subsequent to their divergence. This pattern is consistent with our observations. In common with mosquitoes, for example, are the findings in Simulium of the antigen 5 family, the immune related polypeptides, the OBP/D7 family, and the putative secreted enzymes found in this work, with the exception of destabilase. Intriguingly, members of the uniquely mosquito protein family known as 30-kDa antigen/aegyptin that inhibit platelet aggregation by collagen53, 59 were also found in Simulium. The antigen 5 and OBP/D7 families are also shared with Culicoides.30 Although it is possible that these diverse protein families have each independently been recruited to a role in hematophagy from some now-unrecognizable ancestral gene, it is more likely that they represent elements of the salivary potion present in a common ancestor of the hematophagous Culicomorpha.
The finding of Kunitz-domain peptides in Simulium has its only parallel in Culicoides,30 the remaining Nematocera having opted for different protein families to act as anticlotting inhibitors [e.g., serpin in culicine mosquitoes35], the novel anophelin family of peptides in anopheline mosquitoes,34, 83 and simulidin in simulids. No Kunitz-domain peptide has so far been biochemically characterized from hematophagous Diptera.
The similarities above contrast with the large number of unique protein families found only in Simulium, including the marydilan/SVEP family (one member of which is a potent vasodilator10), the Q-rich and HPQE acidic protein families, the collagen-like family, mucins, and many other proteins indicated in Supplemental Table S2, several of which appear to derive from multigene families.
The finding of multigenic families in both ubiquitous families (such as the antigen 5 family) and genus or insect family-specific genes indicates the importance of gene duplication events in evolution in general and in the evolution of blood feeding in particular.106 The initial effect of gene duplication is to increase transcript abundance, which may lead to a beneficial increase in protein expression. The newly acquired paralogous gene is free to evolve in divergent ways from its origin. Perhaps the immune pressure imposed by the vertebrate hosts may accelerate the evolutionary pace of these genes, explaining the divergence of sequences even in closely related species20, 22, 24 and of novel families in more distantly related species, as is the case for black flies and mosquitoes.
Supplementary Material
Acknowledgment
This work was supported by the Intramural Research Program of the Division of Intramural Research, National Institute of Allergy and Infectious Diseases, National Institutes of Health, and funded in part with federal funds from the National Cancer Institute, National Institutes of Health, under Contract NO1-CO-12400. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organization imply endorsement by the government of the United States of America. We are grateful to Elmer Gray and Dr. Ray Noblet, University of Georgia, for allowing generous access to the black fly colony, and to NIAID intramural editor Brenda Rae Marshall for assistance.
Because J.F.A., V.M.P., Z.M., and J.M.C.R. are government employees and this is a government work, the work is in the public domain in the United States. Notwithstanding any other agreements, the NIH reserves the right to provide the work to PubMedCentral for display and use by the public, and PubMedCentral may tag or modify the work consistent with its customary practices. You can establish rights outside of the U.S. subject to a government use license.
Footnotes
Supporting Information Available: Supplemental Tables S1 and S2 can be obtained from http://exon.niaid.nih.gov/transcriptome/S_vittatum/T1/SV-tb1.zip and http://exon.niaid.nih.gov/transcriptome/S_vittatum/T2/SV-tb2.zip.
References
- 1.Ribeiro JM, Francischetti IM. Role of arthropod saliva in blood feeding: sialome and post-sialome perspectives. Annu. Rev. Entomol. 2003;48:73–88. doi: 10.1146/annurev.ento.48.060402.102812. [DOI] [PubMed] [Google Scholar]
- 2.Cupp EW, Mare CJ, Cupp MS, Ramberg FB. Biological transmission of vesicular stomatitis virus (New Jersey) by Simulium vittatum (Diptera: Simuliidae) J Med Entomol. 1992;29(2):137–40. doi: 10.1093/jmedent/29.2.137. [DOI] [PubMed] [Google Scholar]
- 3.Mead DG, Mare CJ, Cupp EW. Vector competence of select black fly species for vesicular stomatitis virus (New Jersey serotype) Am J Trop Med Hyg. 1997;57(1):42–8. doi: 10.4269/ajtmh.1997.57.42. [DOI] [PubMed] [Google Scholar]
- 4.Freedman D. Onchocerciasis. 2 ed Churchill Livingstone; Philadelphia: 2005. Chapter 100; p. 1176. [Google Scholar]
- 5.Bernardo MJ, Cupp EW, Kiszewski AE. Rearing black flies (Diptera: Simuliidae) in the laboratory: bionomics and life table statistics for Simulium pictipes. J Med Entomol. 1986;23(6):680–4. doi: 10.1093/jmedent/23.6.680. [DOI] [PubMed] [Google Scholar]
- 6.Brockhouse CL, Adler PH. Cytogenetics of laboratory colonies of Simulium vittatum cytospecies IS-7 (Diptera: Simuliidae) J Med Entomol. 2002;39(2):293–7. doi: 10.1603/0022-2585-39.2.293. [DOI] [PubMed] [Google Scholar]
- 7.Abebe M, Cupp MS, Champagne D, Cupp EW. Simulidin: a black fly (Simulium vittatum) salivary gland protein with anti-thrombin activity. J. Insect Physiol. 1995;41:1001–1006. [Google Scholar]
- 8.Abebe M, Ribeiro JM, Cupp MS, Cupp EW. Novel anticoagulant from salivary glands of Simulium vittatum (Diptera: Simuliidae) inhibits activity of coagulation factor V. J Med Entomol. 1996;33(1):173–6. doi: 10.1093/jmedent/33.1.173. [DOI] [PubMed] [Google Scholar]
- 9.Jacobs JW, Cupp EW, Sardana M, Friedman PA. Isolation and characterization of a coagulation factor Xa inhibitor from black fly salivary glands. Thromb Haemost. 1990;64(2):235–8. [PubMed] [Google Scholar]
- 10.Cupp MS, Ribeiro JM, Champagne DE, Cupp EW. Analyses of cDNA and recombinant protein for a potent vasoactive protein in saliva of a blood-feeding black fly, Simulium vittatum. J Exp Biol. 1998;201(Pt 10):1553–61. doi: 10.1242/jeb.201.10.1553. [DOI] [PubMed] [Google Scholar]
- 11.Cupp MS, Ribeiro JM, Cupp EW. Vasodilative activity in black fly salivary glands. Am J Trop Med Hyg. 1994;50(2):241–6. doi: 10.4269/ajtmh.1994.50.241. [DOI] [PubMed] [Google Scholar]
- 12.Cupp MS, Cupp EW, Ochoa AJ, Moulton JK. Salivary apyrase in New World blackflies (Diptera: Simuliidae) and its relationship to onchocerciasis vector status. Med Vet Entomol. 1995;9(3):325–30. doi: 10.1111/j.1365-2915.1995.tb00141.x. [DOI] [PubMed] [Google Scholar]
- 13.Ribeiro JMC. Blood-feeding arthropods: Live syringes or invertebrate pharmacologists? Infect. Agents Dis. 1995;4:143–152. [PubMed] [Google Scholar]
- 14.Ribeiro JM, Charlab R, Rowton ED, Cupp EW. Simulium vittatum (Diptera: Simuliidae) and Lutzomyia longipalpis (Diptera: Psychodidae) salivary gland hyaluronidase activity. J. Med. Entomol. 2000;37(5):743–7. doi: 10.1603/0022-2585-37.5.743. [DOI] [PubMed] [Google Scholar]
- 15.Hutcheon DE, Chivers-Wilson VS. The histaminic and anticoagulant activity of extracts of the black fly; Simulium vittatum and Simulium venustum. Rev Can Biol. 1953;12(1):77–85. [PubMed] [Google Scholar]
- 16.Wirtz HP. Bioamines and proteins in the saliva and salivary glands of Palaearctic blackflies (Diptera: Simuliidae) Trop Med Parasitol. 1990;41(1):59–64. [PubMed] [Google Scholar]
- 17.Wirtz HP. Quantitating histamine in the saliva and salivary glands of two Palaearctic blackfly species (Diptera: Simuliidae) Trop Med Parasitol. 1988;39(4):309–12. [PubMed] [Google Scholar]
- 18.Cupp MS, Cupp EW. Antithrombin protein and DNA sequences from black fly. 2000. 6,077,825, March 6, 1998.
- 19.Calvo E, Andersen J, Francischetti IM, de LCM, deBianchi AG, James AA, Ribeiro JM, Marinotti O. The transcriptome of adult female Anopheles darlingi salivary glands. Insect Mol. Biol. 2004;13(1):73–88. doi: 10.1111/j.1365-2583.2004.00463.x. [DOI] [PubMed] [Google Scholar]
- 20.Calvo E, Dao A, Pham VM, Ribeiro JM. An insight into the sialome of Anopheles funestus reveals an emerging pattern in anopheline salivary protein families. Insect Biochem Mol Biol. 2007;37(2):164–75. doi: 10.1016/j.ibmb.2006.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Francischetti IM, Valenzuela JG, Pham VM, Garfield MK, Ribeiro JM. Toward a catalog for the transcripts and proteins (sialome) from the salivary gland of the malaria vector Anopheles gambiae. J. Exp. Biol. 2002;205(Pt 16):2429–51. doi: 10.1242/jeb.205.16.2429. [DOI] [PubMed] [Google Scholar]
- 22.Valenzuela JG, Francischetti IM, Pham VM, Garfield MK, Ribeiro JM. Exploring the salivary gland transcriptome and proteome of the Anopheles stephensi mosquito. Insect Biochem. Mol. Biol. 2003;33(7):717–32. doi: 10.1016/s0965-1748(03)00067-5. [DOI] [PubMed] [Google Scholar]
- 23.Arca B, Lombardo F, Valenzuela JG, Francischetti IM, Marinotti O, Coluzzi M, Ribeiro JM. An updated catalogue of salivary gland transcripts in the adult female mosquito, Anopheles gambiae. J. Exp. Biol. 2005;208(Pt 20):3971–86. doi: 10.1242/jeb.01849. [DOI] [PubMed] [Google Scholar]
- 24.Arca B, Lombardo F, Francischetti IM, Pham VM, Mestres-Simon M, Andersen JF, Ribeiro JM. An insight into the sialome of the adult female mosquito Aedes albopictus. Insect Biochem Mol Biol. 2007;37(2):107–27. doi: 10.1016/j.ibmb.2006.10.007. [DOI] [PubMed] [Google Scholar]
- 25.Ribeiro JM, Arca B, Lombardo F, Calvo E, Phan VM, Chandra PK, Wikel SK. An annotated catalogue of salivary gland transcripts in the adult female mosquito, Aedes aegypti. BMC Genomics. 2007;8(1):6. doi: 10.1186/1471-2164-8-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ribeiro JM, Charlab R, Pham VM, Garfield M, Valenzuela JG. An insight into the salivary transcriptome and proteome of the adult female mosquito Culex pipiens quinquefasciatus. Insect Biochem. Mol. Biol. 2004;34(6):543–63. doi: 10.1016/j.ibmb.2004.02.008. [DOI] [PubMed] [Google Scholar]
- 27.Anderson JM, Oliveira F, Kamhawi S, Mans BJ, Reynoso D, Seitz AE, Lawyer P, Garfield M, Pham M, Valenzuela JG. Comparative salivary gland transcriptomics of sandfly vectors of visceral leishmaniasis. BMC Genomics. 2006;7:52. doi: 10.1186/1471-2164-7-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Charlab R, Valenzuela JG, Rowton ED, Ribeiro JM. Toward an understanding of the biochemical and pharmacological complexity of the saliva of a hematophagous sand fly Lutzomyia longipalpis. Proc. Natl. Acad. Sci. U S A. 1999;96(26):15155–60. doi: 10.1073/pnas.96.26.15155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Oliveira F, Kamhawi S, Seitz AE, Pham VM, Guigal PM, Fischer L, Ward J, Valenzuela JG. From transcriptome to immunome: identification of DTH inducing proteins from a Phlebotomus ariasi salivary gland cDNA library. Vaccine. 2006;24(3):374–90. doi: 10.1016/j.vaccine.2005.07.085. [DOI] [PubMed] [Google Scholar]
- 30.Campbell CL, Vandyke KA, Letchworth GJ, Drolet BS, Hanekamp T, Wilson WC. Midgut and salivary gland transcriptomes of the arbovirus vector Culicoides sonorensis (Diptera: Ceratopogonidae) Insect Mol. Biol. 2005;14(2):121–36. doi: 10.1111/j.1365-2583.2004.00537.x. [DOI] [PubMed] [Google Scholar]
- 31.Lerner EA, Ribeiro JMC, Nelson RJ, Lerner MR. Isolation of maxadilan, a potent vasodilatory peptide from the salivary glands of the sand fly Lutzomyia longipalpis. J. Biol. Chem. 1991;266:11234–11236. [PubMed] [Google Scholar]
- 32.Ribeiro JM, Katz O, Pannell LK, Waitumbi J, Warburg A. Salivary glands of the sand fly Phlebotomus papatasi contain pharmacologically active amounts of adenosine and 5′-AMP. J. Exp. Biol. 1999;202(Pt 11):1551–9. doi: 10.1242/jeb.202.11.1551. [DOI] [PubMed] [Google Scholar]
- 33.Ribeiro JM, Modi G. The salivary adenosine/AMP content of Phlebotomus argentipes Annandale and Brunetti, the main vector of human kala-azar. J. Parasitol. 2001;87(4):915–7. doi: 10.1645/0022-3395(2001)087[0915:TSAACO]2.0.CO;2. [DOI] [PubMed] [Google Scholar]
- 34.Valenzuela JG, Francischetti IM, Ribeiro JM. Purification, cloning, and synthesis of a novel salivary anti-thrombin from the mosquito Anopheles albimanus. Biochemistry. 1999;38(34):11209–11215. doi: 10.1021/bi990761i. [DOI] [PubMed] [Google Scholar]
- 35.Stark KR, James AA. Isolation and characterization of the gene encoding a novel factor Xa-directed anticoagulant from the yellow fever mosquito, Aedes aegypti. J. Biol. Chem. 1998;273(33):20802–9. doi: 10.1074/jbc.273.33.20802. [DOI] [PubMed] [Google Scholar]
- 36.Cupp EW, Cupp MS. Black fly (Diptera:Simuliidae) salivary secretions: importance in vector competence and disease. J Med Entomol. 1997;34(2):87–94. doi: 10.1093/jmedent/34.2.87. [DOI] [PubMed] [Google Scholar]
- 37.Huang X, Madan A. CAP3: A DNA sequence assembly program. Genome Res. 1999;9(9):868–77. doi: 10.1101/gr.9.9.868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- 39.Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997;25(24):4876–82. doi: 10.1093/nar/25.24.4876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Page RD. TreeView: an application to display phylogenetic trees on personal computers. Comput. Appl. Biosci. 1996;12(4):357–8. doi: 10.1093/bioinformatics/12.4.357. [DOI] [PubMed] [Google Scholar]
- 41.Kumar S, Tamura K, Nei M. MEGA3: Integrated software for Molecular Evolutionary Genetics Analysis and sequence alignment. Brief Bioinform. 2004;5(2):150–63. doi: 10.1093/bib/5.2.150. [DOI] [PubMed] [Google Scholar]
- 42.Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25(17):3389–402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, Harris MA, Hill DP, Issel-Tarver L, Kasarskis A, Lewis S, Matese JC, Richardson JE, Ringwald M, Rubin GM, Sherlock G. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000;25(1):25–9. doi: 10.1038/75556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Bateman A, Birney E, Durbin R, Eddy SR, Howe KL, Sonnhammer EL. The Pfam protein families database. Nucleic Acids Res. 2000;28(1):263–6. doi: 10.1093/nar/28.1.263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Schultz J, Copley RR, Doerks T, Ponting CP, Bork P. SMART: a web-based tool for the study of genetically mobile domains. Nucleic Acids Res. 2000;28(1):231–4. doi: 10.1093/nar/28.1.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV, Krylov DM, Mazumder R, Mekhedov SL, Nikolskaya AN, Rao BS, Smirnov S, Sverdlov AV, Vasudevan S, Wolf YI, Yin JJ, Natale DA. The COG database: an updated version includes eukaryotes. BMC Bioinformatics. 2003;4(1):41. doi: 10.1186/1471-2105-4-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Marchler-Bauer A, Panchenko AR, Shoemaker BA, Thiessen PA, Geer LY, Bryant SH. CDD: a database of conserved domain alignments with links to domain three-dimensional structure. Nucleic Acids Res. 2002;30(1):281–3. doi: 10.1093/nar/30.1.281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hulo N, Bairoch A, Bulliard V, Cerutti L, De Castro E, Langendijk-Genevaux PS, Pagni M, Sigrist CJ. The PROSITE database. Nucleic Acids Res. 2006;34:D227–30. doi: 10.1093/nar/gkj063. (Database issue) [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Nielsen H, Engelbrecht J, Brunak S, von Heijne G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 1997;10(1):1–6. doi: 10.1093/protein/10.1.1. [DOI] [PubMed] [Google Scholar]
- 50.Hansen JE, Lund O, Tolstrup N, Gooley AA, Williams KL, Brunak S. NetOglyc: prediction of mucin type O-glycosylation sites based on sequence context and surface accessibility. Glycoconj J. 1998;15(2):115–30. doi: 10.1023/a:1006960004440. [DOI] [PubMed] [Google Scholar]
- 51.Ribeiro JM, Andersen J, Silva-Neto MA, Pham VM, Garfield MK, Valenzuela JG. Exploring the sialome of the blood-sucking bug Rhodnius prolixus. Insect Biochem. Mol. Biol. 2004;34(1):61–79. doi: 10.1016/j.ibmb.2003.09.004. [DOI] [PubMed] [Google Scholar]
- 52.Galperin MY, Koonin EV. ‘Conserved hypothetical‘ proteins: prioritization of targets for experimental study. Nucleic Acids Res. 2004;32(18):5452–63. doi: 10.1093/nar/gkh885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Calvo E, Tokumasu F, Marinotti O, Villeval JL, Ribeiro JM, Francischetti IM. Aegyptin, a novel mosquito salivary gland protein, specifically binds to collagen and prevents its interaction with platelet glycoprotein VI, integrin alpha2beta1, and von Willebrand factor. J Biol Chem. 2007;282(37):26928–38. doi: 10.1074/jbc.M705669200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Landschulz WH, Johnson PF, McKnight SL. The leucine zipper: a hypothetical structure common to a new class of DNA binding proteins. Science. 1988;240(4860):1759–64. doi: 10.1126/science.3289117. [DOI] [PubMed] [Google Scholar]
- 55.Alber T. Structure of the leucine zipper. Curr Opin Genet Dev. 1992;2(2):205–10. doi: 10.1016/s0959-437x(05)80275-8. [DOI] [PubMed] [Google Scholar]
- 56.Klintschar M, Wiegand P. Polymerase slippage in relation to the uniformity of tetrameric repeat stretches. Forensic Sci Int. 2003;135(2):163–6. doi: 10.1016/s0379-0738(03)00201-9. [DOI] [PubMed] [Google Scholar]
- 57.Heino J. The collagen family members as cell adhesion proteins. Bioessays. 2007;29(10):1001–10. doi: 10.1002/bies.20636. [DOI] [PubMed] [Google Scholar]
- 58.Simons FE, Peng Z. Mosquito allergy: recombinant mosquito salivary antigens for new diagnostic tests. Int. Arch. Allergy Immunol. 2001;124(13):403–5. doi: 10.1159/000053771. [DOI] [PubMed] [Google Scholar]
- 59.Yoshida S, Sudo T, Niimi M, Tao L, Sun B, Kambayashi J, Watanabe H, Luo E, Matsuoka H. Inhibition of collagen-induced platelet aggregation by anopheline antiplatelet protein, a saliva protein from a malaria vector mosquito. Blood. 2008;111(4):2007–14. doi: 10.1182/blood-2007-06-097824. [DOI] [PubMed] [Google Scholar]
- 60.Krzywinski J, Grushko OG, Besansky NJ. Analysis of the complete mitochondrial DNA from Anopheles funestus: an improved dipteran mitochondrial genome annotation and a temporal dimension of mosquito evolution. Mol Phylogenet Evol. 2006;39(2):417–23. doi: 10.1016/j.ympev.2006.01.006. [DOI] [PubMed] [Google Scholar]
- 61.Grimaldi D, Engel M. Evolution of the insects. Cambridge University Press; New York: 2005. p. 772. [Google Scholar]
- 62.Ribeiro JM, Alarcon-Chaidez F, Francischetti IM, Mans BJ, Mather TN, Valenzuela JG, Wikel SK. An annotated catalog of salivary gland transcripts from Ixodes scapularis ticks. Insect Biochem. Mol. Biol. 2006;36(2):111–29. doi: 10.1016/j.ibmb.2005.11.005. [DOI] [PubMed] [Google Scholar]
- 63.Julenius K, Molgaard A, Gupta R, Brunak S. Prediction, conservation analysis, and structural characterization of mammalian mucin-type O-glycosylation sites. Glycobiology. 2005;15(2):153–64. doi: 10.1093/glycob/cwh151. [DOI] [PubMed] [Google Scholar]
- 64.James AA, Blackmer K, Racioppi JV. A salivary gland-specific, maltase-like gene of the vector mosquito, Aedes aegypti. Gene. 1989;75:73–83. doi: 10.1016/0378-1119(89)90384-3. [DOI] [PubMed] [Google Scholar]
- 65.Hekmat-Scafe DS, Dorit RL, Carlson JR. Molecular evolution of odorant-binding protein genes OS-E and OS-F in Drosophila. Genetics. 2000;155(1):117–27. doi: 10.1093/genetics/155.1.117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Valenzuela JG, Charlab R, Gonzalez EC, Miranda-Santos IKF, Marinotti O, Francischetti IM, Ribeiro JMC. The D7 family of salivary proteins in blood sucking Diptera. Insect Mol. Biol. 2002;11(2):149–55. doi: 10.1046/j.1365-2583.2002.00319.x. [DOI] [PubMed] [Google Scholar]
- 67.Calvo E, Mans BJ, Andersen JF, Ribeiro JM. Function and evolution of a mosquito salivary protein family. J. Biol. Chem. 2006;281(4):1935–42. doi: 10.1074/jbc.M510359200. [DOI] [PubMed] [Google Scholar]
- 68.Mans BJ, Calvo E, Ribeiro JM, Andersen JF. The crystal structure of D7r4, a salivary biogenic amine-binding protein from the malaria mosquito Anopheles gambiae. J Biol Chem. 2007;282(50):36626–33. doi: 10.1074/jbc.M706410200. [DOI] [PubMed] [Google Scholar]
- 69.Valenzuela JG, Belkaid Y, Garfield MK, Mendez S, Kamhawi S, Rowton ED, Sacks DL, Ribeiro JM. Toward a defined anti-Leishmania vaccine targeting vector antigens: characterization of a protective salivary protein. J. Exp. Med. 2001;194(3):331–42. doi: 10.1084/jem.194.3.331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Isawa H, Yuda M, Orito Y, Chinzei Y. A mosquito salivary protein inhibits activation of the plasma contact system by binding to factor XII and high molecular weight kininogen. J. Biol. Chem. 2002;13:13. doi: 10.1074/jbc.M203505200. [DOI] [PubMed] [Google Scholar]
- 71.Megraw T, Kaufman TC, Kovalick GE. Sequence and expression of Drosophila Antigen 5-related 2, a new member of the CAP gene family. Gene. 1998;222(2):297–304. doi: 10.1016/s0378-1119(98)00489-2. [DOI] [PubMed] [Google Scholar]
- 72.Milne TJ, Abbenante G, Tyndall JD, Halliday J, Lewis RJ. Isolation and characterization of a cone snail protease with homology to CRISP proteins of the pathogenesis-related protein superfamily. J Biol. Chem. 2003;278(33):31105–10. doi: 10.1074/jbc.M304843200. [DOI] [PubMed] [Google Scholar]
- 73.Yamazaki Y, Hyodo F, Morita T. Wide distribution of cysteine-rich secretory proteins in snake venoms: isolation and cloning of novel snake venom cysteine-rich secretory proteins. Arch Biochem Biophys. 2003;412(1):133–41. doi: 10.1016/s0003-9861(03)00028-6. [DOI] [PubMed] [Google Scholar]
- 74.Yamazaki Y, Morita T. Structure and function of snake venom cysteine-rich secretory proteins. Toxicon. 2004;44(3):227–31. doi: 10.1016/j.toxicon.2004.05.023. [DOI] [PubMed] [Google Scholar]
- 75.Mochca-Morales J, Martin BM, Possani LD. Isolation and characterization of helothermine, a novel toxin from Heloderma horridum horridum (Mexican beaded lizard) venom. Toxicon. 1990;28(3):299–309. doi: 10.1016/0041-0101(90)90065-f. [DOI] [PubMed] [Google Scholar]
- 76.Xu X, Yang H, Ma D, Wu J, Wang Y, Song Y, Wang X, Lu Y, Yang J, Lai R. Toward an understanding of the molecular mechanism for successful blood feeding by coupling proteomics analysis with pharmacological testing of horsefly salivary glands. Mol Cell Proteomics. 2008;7(3):582–90. doi: 10.1074/mcp.M700497-MCP200. [DOI] [PubMed] [Google Scholar]
- 77.Scarborough RM, Naughton MA, Teng W, Rose JW, Phillips DR, Nannizzi L, Arfsten A, Campbell AM, Charo IF. Design of potent and specific integrin antagonists. Peptide antagonists with high specificity for glycoprotein IIb-IIIa. J Biol. Chem. 1993;268(2):1066–73. [PubMed] [Google Scholar]
- 78.Ascenzi P, Bocedi A, Bolognesi M, Spallarossa A, Coletta M, De Cristofaro R, Menegatti E. The bovine basic pancreatic trypsin inhibitor (Kunitz inhibitor): a milestone protein. Curr Protein Pept Sci. 2003;4(3):231–51. doi: 10.2174/1389203033487180. [DOI] [PubMed] [Google Scholar]
- 79.Salier JP. Inter-alpha-trypsin inhibitor: emergence of a family within the Kunitz-type protease inhibitor superfamily. Trends Biochem Sci. 1990;15(11):435–9. doi: 10.1016/0968-0004(90)90282-g. [DOI] [PubMed] [Google Scholar]
- 80.Mans BJ, Neitz AW. Adaptation of ticks to a blood-feeding environment: evolution from a functional perspective. Insect Biochem Mol Biol. 2004;34(1):1–17. doi: 10.1016/j.ibmb.2003.09.002. [DOI] [PubMed] [Google Scholar]
- 81.Steen NA, Barker SC, Alewood PF. Proteins in the saliva of the Ixodida (ticks): pharmacological features and biological significance. Toxicon. 2006;47(1):1–20. doi: 10.1016/j.toxicon.2005.09.010. [DOI] [PubMed] [Google Scholar]
- 82.Francischetti IM, Valenzuela JG, Andersen JF, Mather TN, Ribeiro JM. Ixolaris, a novel recombinant tissue factor pathway inhibitor (TFPI) from the salivary gland of the tick, Ixodes scapularis: identification of factor X and factor Xa as scaffolds for the inhibition of factor VIIa/tissue factor complex. Blood. 2002;99(10):3602–12. doi: 10.1182/blood-2001-12-0237. [DOI] [PubMed] [Google Scholar]
- 83.Francischetti IM, Valenzuela JG, Ribeiro JM. Anophelin: kinetics and mechanism of thrombin inhibition. Biochemistry. 1999;38(50):16678–85. doi: 10.1021/bi991231p. [DOI] [PubMed] [Google Scholar]
- 84.de Leon A. A. Perez, Valenzuela JG, Tabachnick WJ. Anticoagulant activity in salivary glands of the insect vector Culicoides variipennis sonorensis by an inhibitor of factor Xa. Exp Parasitol. 1998;88(2):121–30. doi: 10.1006/expr.1998.4210. [DOI] [PubMed] [Google Scholar]
- 85.Abebe M, Cupp MS, Ramberg FB, Cupp EW. Anticoagulant activity in salivary gland extracts of black flies (Diptera: Simuliidae) J Med Entomol. 1994;31(6):908–11. doi: 10.1093/jmedent/31.6.908. [DOI] [PubMed] [Google Scholar]
- 86.Champagne DE, Smartt CT, Ribeiro JM, James AA. The salivary gland-specific apyrase of the mosquito Aedes aegypti is a member of the 5′-nucleotidase family. Proc. Natl. Acad. Sci. U S A. 1995;92(3):694–8. doi: 10.1073/pnas.92.3.694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Sun D, McNicol A, James AA, Peng Z. Expression of functional recombinant mosquito salivary apyrase: A potential therapeutic platelet aggregation inhibitor. Platelets. 2006;17(3):178–84. doi: 10.1080/09537100500460234. [DOI] [PubMed] [Google Scholar]
- 88.Faudry E, Lozzi SP, Santana JM, D’Souza-Ault M, Kieffer S, Felix CR, Ricart CA, Sousa MV, Vernet T, Teixeira AR. Triatoma infestans apyrases belong to the 5′-nucleotidase family. J. Biol. Chem. 2004;279(19):19607–13. doi: 10.1074/jbc.M401681200. [DOI] [PubMed] [Google Scholar]
- 89.Valenzuela JG, Belkaid Y, Rowton E, Ribeiro JM. The salivary apyrase of the blood-sucking sand fly Phlebotomus papatasi belongs to the novel Cimex family of apyrases. J. Exp. Biol. 2001;204(Pt 2):229–37. doi: 10.1242/jeb.204.2.229. [DOI] [PubMed] [Google Scholar]
- 90.Valenzuela JG, Charlab R, Galperin MY, Ribeiro JM. Purification, cloning, and expression of an apyrase from the bed bug Cimex lectularius. A new type of nucleotide-binding enzyme. J. Biol. Chem. 1998;273(46):30583–90. doi: 10.1074/jbc.273.46.30583. [DOI] [PubMed] [Google Scholar]
- 91.Andersen JF, Hinnebusch BJ, Lucas DA, Conrads TP, Veenstra TD, Pham VM, Ribeiro JM. An insight into the sialome of the oriental rat flea, Xenopsylla cheopis (Rots) BMC Genomics. 2007;8:102. doi: 10.1186/1471-2164-8-102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Reddy VB, Kounga K, Mariano F, Lerner EA. Chrysoptin is a potent glycoprotein IIb/IIIa fibrinogen receptor antagonist present in salivary gland extracts of the deerfly. J Biol. Chem. 2000;275(21):15861–7. doi: 10.1074/jbc.275.21.15861. [DOI] [PubMed] [Google Scholar]
- 93.Francischetti IM, Mather TN, Ribeiro JM. Cloning of a salivary gland metalloprotease and characterization of gelatinase and fibrin(ogen)lytic activities in the saliva of the Lyme disease tick vector Ixodes scapularis. Biochem Biophys Res Commun. 2003;305(4):869–75. doi: 10.1016/s0006-291x(03)00857-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Broutin I, Arnoux B, Riche C, Lecroisey A, Keil B, Pascard C, Ducruix A. 1.8 A structure of Hypoderma lineatum collagenase: a member of the serine proteinase family. Acta Crystallogr D Biol Crystallogr. 1996;52(Pt 2):380–92. doi: 10.1107/S090744499501184X. [DOI] [PubMed] [Google Scholar]
- 95.Moire N, Bigot Y, Periquet G, Boulard C. Sequencing and gene expression of hypodermins A, B, C in larval stages of Hypoderma lineatum. Mol Biochem Parasitol. 1994;66(2):233–40. doi: 10.1016/0166-6851(94)90150-3. [DOI] [PubMed] [Google Scholar]
- 96.Baskova IP, Nikonov GI. Destabilase, the novel epsilon-(gamma-Glu)-Lys isopeptidase with thrombolytic activity. Blood Coagul Fibrinolysis. 1991;2(1):167–72. doi: 10.1097/00001721-199102000-00025. [DOI] [PubMed] [Google Scholar]
- 97.Zavalova LL, Artamonova II, Berezhnoy SN, Tagaev AA, Baskova IP, Andersen J, Roepstorff P, Ts A. Egorov. Multiple forms of medicinal leech destabilase-lysozyme. Biochem Biophys Res Commun. 2003;306(1):318–23. doi: 10.1016/s0006-291x(03)00896-9. [DOI] [PubMed] [Google Scholar]
- 98.Zavalova LL, Baskova IP, Lukyanov SA, Sass AV, Snezhkov EV, Akopov SB, Artamonova II, Archipova VS, Nesmeyanov VA, Kozlov DG, Benevolensky SV, Kiseleva VI, Poverenny AM, Sverdlov ED. Destabilase from the medicinal leech is a representative of a novel family of lysozymes. Biochim Biophys Acta. 2000;1478(1):69–77. doi: 10.1016/s0167-4838(00)00006-6. [DOI] [PubMed] [Google Scholar]
- 99.Ribeiro JMC, Rowton ED, Charlab R. Salivary amylase activity of the phlebotomine sand fly, Lutzomyia longipalpis. Insect Biochem. Mol. Biol. 1999 doi: 10.1016/s0965-1748(99)00119-8. Submitted. [DOI] [PubMed] [Google Scholar]
- 100.Grossman GL, Campos Y, Severson DW, James AA. Evidence for two distinct members of the amylase gene family in the yellow fever mosquito, Aedes aegypti. Insect Biochem. Mol. Biol. 1997;27(89):769–81. doi: 10.1016/s0965-1748(97)00063-5. [DOI] [PubMed] [Google Scholar]
- 101.Grossman GL, James AA. The salivary glands of the vector mosquito, Aedes aegypti, express a novel member of the amylase gene family. Insect Mol. Biol. 1993;1(4):223–32. doi: 10.1111/j.1365-2583.1993.tb00095.x. [DOI] [PubMed] [Google Scholar]
- 102.Blandin S, Levashina EA. Thioester-containing proteins and insect immunity. Mol Immunol. 2004;40(12):903–8. doi: 10.1016/j.molimm.2003.10.010. [DOI] [PubMed] [Google Scholar]
- 103.Kanost MR, Jiang H, Yu XQ. Innate immune responses of a lepidopteran insect, Manduca sexta. Immunol Rev. 2004;198:97–105. doi: 10.1111/j.0105-2896.2004.0121.x. [DOI] [PubMed] [Google Scholar]
- 104.Grogan WL, Szadziewski R. A new biting midge from Upper Cretaceous (Cenomanian) amber of New Jersey (Diptera: Ceratopogonidae) J. Paleont. 1988;62:808–812. [Google Scholar]
- 105.Pawloswski J, Szadziewski R, Kmieciak D, Fahrni J, Bittar G. Phylogeny of the infraorder Culicomorpha (Diptera: Nematocera) based on 28S RNA gene sequences. Syst Entomol. 1996;21(2):167–178. [Google Scholar]
- 106.Kondrashov FA, Koonin EV. A common framework for understanding the origin of genetic dominance and evolutionary fates of gene duplications. Trends Genet. 2004;20(7):287–90. doi: 10.1016/j.tig.2004.05.001. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.