Abstract
Recent studies have uncovered an unexpected relationship between factors that are essential for germline development in Drosophila melanogaster: the arginine protein methyltransferase 5 (dPRMT5/Csul/Dart5) and its cofactor Valois, methylate the Piwi family protein Aub, enabling it to bind Tudor. The RNA helicase Vasa is another essential protein in germline development. Here, we report that mouse (mouse Vasa homolog), Xenopus laevis, and D. melanogaster Vasa proteins contain both symmetrical and asymmetrical dimethylarginines. We find that dPRMT5 is required for the production of sDMAs of Vasa in vivo. Furthermore, we find that the mouse Vasa homolog associates with Tudor domain-containing proteins, Tdrd1 and Tdrd6, as well as the Piwi proteins, Mili and Miwi. Arginine methylation is thus emerging as a conserved and pivotal post-translational modification of proteins that is essential for germline development.
Keywords: Drosophila, MicroRNA, Protein Methylation, RNA-Binding Protein, Spermatogenesis, Piwi, Tudor Protein, Vasa Helicase, Drosophila Oogenesis, PiRNA
Introduction
Germline specification is an essential process for all sexually reproducing organisms. In Drosophila melanogaster, germline development is initiated by the formation of the germ cell precursors, pole cells, which are induced through the activity of maternally inherited, cytoplasmic determinants deposited in the pole plasm (also known as germ plasm) at the posterior pole of the oocyte (1–3). Genetic studies have identified maternal genes (often referred to as posterior group or grandchild-less genes) that are required for germ cell specification and the protein or RNA products of these genes are invariably concentrated in the pole plasm (2–4). Among these genes are aub (5); csul/dart5 (the D. melanogaster homolog of protein methyltransferase 5, dPRMT5) (6, 7) and its cofactor valois (8), tudor (9, 10, 11), and vasa (12, 13).
Vasa encodes an ∼65-kDa DEAD-box RNA helicase whose expression is exclusively restricted to the germ cell lineage (12, 13). Vasa binds RNA, and this activity is required for germ cell formation but not for localization to the pole plasm (14). Vasa binds to eukaryotic initiation factor 5B, and this interaction is required for germ cell development (15). A proposed function for Vasa is that it regulates the translation of target mRNAs involved in germ cell establishment such as oskar and nanos, and oogenesis such as gurken (15–19). The first component known to arrive at the pole plasm is oskar mRNA that is locally translated in the pole plasm (20). Oskar protein in turn binds to Vasa, and this interaction appears to be essential for Vasa recruitment to the pole plasm (21). Vasa is widely conserved in invertebrate and vertebrate species including D. melanogaster, Xenopus laevis, Danio rerio, mouse, and human (22). The Xenopus Vasa homolog (XVLG1, Xenopus Vasa-like gene 1) is expressed in oocytes and embryos and is required for the formation of germ cells (23–25). Mouse Vasa homolog (MVH,2 DDX4) expression is also restricted to the germ cell lineage (26) and loss of MVH protein function causes a deficiency in the proliferation and differentiation of spermatocytes, leading to male sterility (27).
Protein arginine methylation is an important post-translational modification that is mediated by two types of protein methyltransferases (PRMTs). Type I enzymes (such as PRMT1) catalyze asymmetric dimethylation of arginines (aDMA) and type II enzymes (such as PRMT5) catalyze symmetric arginine dimethylation (sDMA) (Fig. 1C) (28, 29). sDMA modifications occur in sequence motifs composed of arginines flanked by glycines (GRG) or alanines (GRA or ARG) that are often found as repeats, whereas aDMAs frequently occur in repeating “RGG” sequences (28, 29). Methylated arginines and in particular sDMAs bind to Tudor domains of proteins and regulate protein-protein interaction (8, 28, 29). In mammals, PRMT5 associates with its cofactor MEP50 to form the methylosome (30). Valois is the D. melanogaster homolog of MEP50 (dMEP50) (31, 32). In valois-null mutants, dPRMT5 is destabilized, resulting in absence of sDMA modifications of target proteins, indicating that dMEP50 is required for sDMA production in concert with dPRMT5 (7).
FIGURE 1.
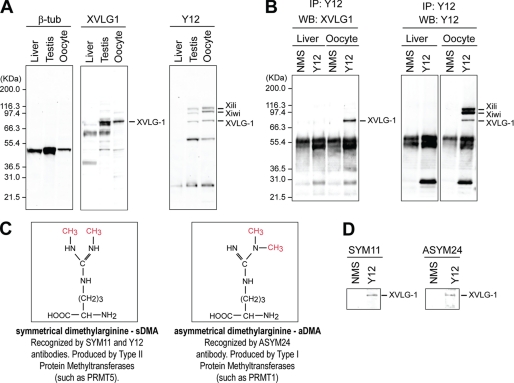
X. laevis Vasa protein XVLG1 is immunoprecipitated by Y12 and contains both sDMA and aDMA. A, Western blots from lysate of indicated X. laevis tissues. B, Y12 immunoprecipitates (IP) from X. laevis liver or oocytes were probed on Western blots (WB) with indicated antibodies (NMS: nonimmune mouse serum; negative control). C, arrangement of methyl groups in dimethylarginines. D, Y12 immunoprecipitates from X. laevis oocytes were probed on Western blots with anti-sDMA (SYM11) or anti-aDMA (ASYM24) antibodies. β-tub, β-tubulin.
Piwi family proteins bind to 25–31 nucleotides Piwi-interacting RNAs (piRNAs) to form Piwi-ribonucleoproteins and play an essential role in germline development and specification. D. melanogaster expresses three Piwi proteins termed Aub, Piwi, and Ago3, whereas mice express three Piwi proteins known as Mili, Miwi, and Miwi2 (33–35). Piwi family proteins from diverse species contain sDMA modifications (36–38), and in D melanogaster, the sDMAs are catalyzed by dPRMT5 (36). Aub binds to Tudor through interactions between sDMAs of Aub and Tudor domains of Tudor (39, 40), and the sDMA-dependent Aub interaction with Tudor is required for pole plasm assembly (39). Collectively, these findings provide an explanation for the relationship between the protein products of four posterior grandchild-less genes; dPRMT5 and dMEP50 produce sDMAs in Aub, which, in turn, are required for binding to Tudor (39). The binding of sDMA-modified Piwi proteins with Tudor domain proteins extends beyond D. melanogaster and appears to be an evolutionary conserved interaction in germ cells. In mice, sDMA modifications in Mili are required for binding to the Tudor domain-containing protein Tdrd1 (37, 38). Tdrd6, the mouse homolog of Drosophila Tudor, associates predominantly with Miwi and also Mili (38, 39, 41). Tdrd2/Tdrdkh interacts with Miwi (42), and Tdrd9, the mouse homolog of Drosophila Spindle E, binds to Miwi2 (43). X. laevis Xiwi protein associates with Xenopus Tudor (44).
Here, we report that mouse, X. laevis, and D. melanogaster Vasa proteins contain both sDMAs and aDMAs. We identify dPRMT5 as the enzyme that catalyzes sDMAs of the Vasa protein in vivo. Furthermore, we find that MVH is part of Piwi-ribonucleoprotein complexes and interacts with Tudor domain-containing proteins, Tdrd1 and Tdrd6, as well as the Piwi proteins Mili and Miwi.
EXPERIMENTAL PROCEDURES
Antibodies
For Western blots and immunoprecipitations of mouse (MVH), Drosophila (Vasa), and Xenopus (XVLG1) Vasa proteins, anti-MVH (Abcam), anti-Vasa (Developmental Studies Hybridoma Bank), and anti-Vasa-H80 (Santa Cruz Biotechnology) were used, respectively. Anti-Vasa-H80 reacts with XVLG1 because it was raised against a region (amino acids 631–710) of a human Vasa protein whose sequence is almost identical to amino acids 617–662 of XVLG1. Other antibodies used were anti-sDMA (SYM11, Millipore), anti-aDMA (ASYM24, Millipore), anti-β-tubulin (Developmental Studies Hybridoma Bank), anti-Miwi (39), and G82 (Cell Signaling Technology), anti-Mili monoclonal 17-8 (Cell Signaling Technology) (36), anti-FLAG (Sigma), anti-Myc (9E-10), and anti-penta His (Qiagen). Y12 antibody was a kind gift from G. Dreyfuss. Antibodies against Tdrd1 and Tdrd6 were gifts from S. Chuma (45). Drosophila anti-Tudor antibody was a gift from P. Lasko (46).
Western Blots and Immunoprecipitations
Western blots and immunoprecipitations were performed essentially as described previously (36). Cell lysates were prepared from mouse testis (Pel-Freez Biologicals); Xenopus laevis oocytes, testis, and liver; or Drosophila ovaries in a lysis buffer (20 mm Tris-HCL, pH 7.5, 200 mm NaCl, 2.5 mm MgCl2, 0.5% Nonidet P-40, 0.1% Triton X-100 and complete EDTA-free protease inhibitors (Roche Diagnostics). Anti-mouse IgM-agarose (Sigma) was used for immunoprecipitation with anti-Vasa (Developmental Studies Hybridoma Bank), whereas Protein-G-agarose (Invitrogen) was used with other antibodies.
X. laevis
Oocytes were isolated from ovaries and defolliculated as described in Ref. 47. Testis and liver tissues were procured from euthanized animals.
Drosophila Stocks and Immunofluorescence
csul flies (csulRM: w−;csulRM50/CyO) were a gift from J. Anne (6). Immunofluorescence of Drosophila ovaries was performed as described previously (36).
RNA Isolation and Labeling
RNA isolation and labeling were performed as described previously (36). Briefly, RNA was isolated from immunoprecipitates using TRIzol (Invitrogen) and treated with calf intestinal phosphatase (New England Biolabs). The 5′-dephosphorylated RNAs were then subjected to 5′-end labeling using [γ-32P]ATP and T4 polynucleotide kinase (New England Biolabs). The labeled RNAs were resolved by 15% PAGE containing 7 m urea and were visualized by storage phosphor autoradiography using a Storm 860 PhosphorImager and ImageQuant software (GE Healthcare).
Transfections and Co-immunoprecipitation Experiments
Full-length mili and miwi coding sequence were cloned without a tag in pcDNA3 expression vectors. Tdrd1 (C terminus His tag) and Tdrd6 (N terminus FLAG tag) pCAG constructs were kind gifts from S. Chuma. HEK 293T cells were transfected with a total of 12 μg of vector DNA, keeping a molar ratio of 1:1 for each pair of constructs and using Lipofectamine 2000. Cells were treated with dimethyl sulfoxide alone (solvent) or MTA (750 μm) 1 day prior to transfection and were harvested 2 days after transfection. Culture media was replaced every 12 h. Cells were lysed by sonication in a buffer containing 20 mm Tris-HCL, pH 7.5, 150 mm NaCl, 2.5 mm MgCl2, 0.1% Nonidet P-40, and complete EDTA-free protease inhibitors (Roche Diagnostics).
Mass Spectrometry for Analysis of Arginine Methylation
The immunopurified MVH band was excised from a 4–12% NuPAGE followed by trypsin digestion as described elsewhere (48). In brief, proteins were reduced in 10 mm dithiothreitol for 30 min at 37 °C, alkylated in 55 mm iodoacetamide for 20 min at room temperature in the dark, and digested overnight at 37 °C with 12.5 ng/μl trypsin (Proteomics grade, Sigma). The digestion media were then acidified to 0.1% of trifluoroacetic acid and spun onto StageTips as described elsewhere (49). Peptides were eluted in 20 μl of 80% acetonitrile in 0.1% trifluoroacetic acid and were concentrated to 4 μl (Concentrator 5301, Eppendorf AG). They were then diluted to 5 μl by 0.1% trifluoroacetic acid for liquid chromatography-tandem mass spectrometry analysis.
For mass spectrometry analysis, an LTQ-Orbitrap mass spectrometer (Thermo Fisher Scientific) was coupled online to an Agilent 1100 binary nanopump and an HTC PAL autosampler (CTC Analysis). To prepare an analytical column with a self-assembled particle frit (50), C18 material (ReproSil-Pur C18-AQ 3 μm; Dr. Maisch, GmbH) was packed into a spray emitter (75-μm inner diameter, 8-μm opening, and 70-mm length; New Objectives) using an air pressure pump (Proxeon Biosystems). Mobile phase A consisted of water, 5% acetonitrile, and 0.5% acetic acid; mobile phase B consisted of acetonitrile and 0.5% acetic acid. The gradient went from 0 to 20% mobile phase B in 75 min and then to 80% mobile phase B in 13 min at a 300 nanoliters/min flow. The six most intense peaks of the mass spectrometry (MS) scan were selected for tandem mass spectrometry in the ion trap (normal scan, wideband activation, filling 5e5 ions for MS scan, 104 ions for MS2, maximum fill time 100 msec, and dynamic exclusion for 180 s). Raw files were processed using DTAsupercharge 0.62. The generated peak lists were searched against the ipi.MOUSE.v3.56 database using Mascot 2.0 with the parameters: monoisotopic masses, 6 ppm peptide tolerance, and 0.6 Da tandem MS tolerance, ESI TRAP parameters, fully tryptic specificity, cysteine carbamidomethylated as fixed modification, oxidation on methionine, arginine methylation and dimethylation as variable modifications, with two missed cleavage sites allowed. The results were parsed through MSQuant, and a cut-off 5-ppm peptide tolerance was applied to the recalibrated list. Peptides with scores 25 and higher were reported, and all methylations and dimethylations were reported and in individual cases manually validated.
RESULTS
X. laevis Vasa Protein XVLG1 Is Recognized by Y12 Antibody and Contains Both sDMAs and aDMAs
We have previously reported that the widely used Y12 monoclonal antibody (51) recognizes X. laevis Piwi proteins, Xiwi and Xili, and that both Xiwi and Xili contain sDMA modifications (36). In addition to these proteins, we found that on Western blots, Y12 recognized a protein of ∼70 kDa specifically in testis and oocytes, but not in liver of X. laevis, corresponding in size to XVLG1 (Fig. 1A). We performed immunoprecipitations with Y12 from X. laevis-defolliculated oocytes and liver, followed by Western blots probed by anti-XVLG1 or Y12 antibody. As shown in Fig. 1B (left panel), XVLG1 protein was present in the Y12 immunoprecipitates from oocytes, but not from liver. Furthermore, Y12 recognized XVLG1 as well as Xiwi and Xili on Western blot of Y12 immunoprecipitates (Fig. 1B, right panel). These results indicated that Y12 reacts directly with XVLG1.
The Sm proteins of small nuclear ribonucleoproteins constitute the main antigen for Y12, and the epitope consists of sDMAs in the glycine-arginine-rich regions of the proteins (51, 52). Moreover, Y12 recognizes sDMAs of Piwi proteins (36). We thus reasoned that Y12 likely reacted with sDMA-containing epitopes in XVLG1. We tested whether XVLG1 contains arginine methylations by using SYM11 or ASYM24 antibody, which specifically recognize proteins containing sDMA-glycine or aDMA-glycine repeats, respectively (Fig. 1C) (53, 54). As shown in Fig. 1D, Western blots clearly revealed that both SYM11 and ASYM24 antibodies reacted with Y12-immunopurified XVLG1, indicating that XVLG1 contained both sDMAs and aDMAs. We searched for arginine residues that could be symmetrically or asymmetrically methylated in Vasa proteins. Intriguingly, we found that Vasa proteins from diverse species as well as XVLG1 contain putative sDMA and aDMA motifs (Table 1).
TABLE 1.
DMA motifs in Vasa homologs
Putative sDMA (GRG and ARG/GRA) or aDMA (RGG) motifs present in animal Vasa homologs; numbers refer to amino acid positions.
| Species | Proteins | sDMA or aDMA motifs |
|---|---|---|
| Human (Homo sapiens) | Vasa | 130-RGG, 147-RGGRG, 157-RGG, 208-RGG, 601-ARG, 638-GRA |
| Mouse (Mus musculus) | MVH | 61-GRG, 105-RGG, 122-RGGRG, 132-RGG, 152-RGG, 180-GRGG, 574-ARG, 611-GRA, 662-RGG |
| Frog (X. laevis) | XVLG1 | 53-RGG, 75-GRGRG, 108-GRGG, 126-GRGG, 132-RGG, 175-GRGG, 186-GRGRRGGRGG, 587-ARG |
| Zebrafish (D. rerio) | Vasa | 61-GRGGRGG, 69-RGGRGG, 97-RGRGRGG, 105-RGG, 131-RGRGRGG, 143-RGG, 174-GRGRGRGG, 183-RGG, 200-GRGG, 420-GRG, 590-ARG, 627-GRA |
| Fruit fly (D. melanogaster) | Vasa | 16-ARGG, 64-GRGG, 73-RGG, 94-RGG, 101-RGG, 108-RGG, 115-RGG, 122-RGG, 150-RGG, 163-RGG, 587-GRA, 642-RGRG |
D. melanogaster Vasa Proteins Contain sDMAs and aDMAs, and the sDMAs Are Generated by dPRMT5 in Vivo
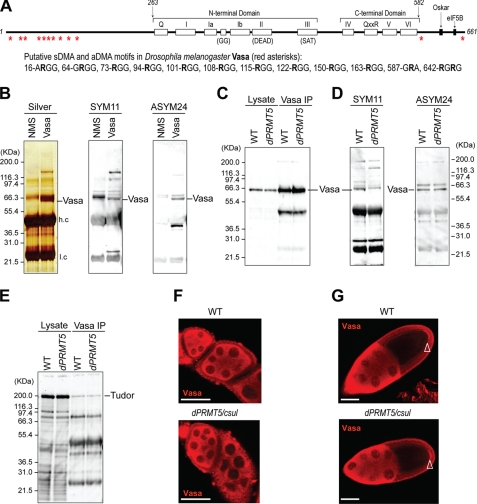
Next, we asked whether Drosophila Vasa protein (schematic shown in Fig. 2A) contained methylated arginines. We immunopurified Vasa from Drosophila ovaries (Fig. 2B) and confirmed the identity of the purified protein by mass spectrometry (supplemental Table 1). A nonspecific protein band of ∼67 kDa, right above Vasa, is present in the Vasa immunoprecipitates and also in the nonimmune mouse serum negative control immunoprecipitates (silver stain panel of Fig. 2B). This band corresponds, at least in part, to undissociated, single heavy and light antibody chains because it is recognized by anti-mouse Ig antibodies.3 As shown in the Western blots in Fig. 2B, both SYM11 and ASYM24 reacted with immunopurified Vasa, indicating that it contained sDMAs and aDMAs.
FIGURE 2.
D. melanogaster Vasa protein contains sDMAs and aDMAs, and Drosophila PRMT5 (dPRMT5/csul/dart5) is required for sDMA modifications. A, a schematic of Vasa and its putative DMAs is shown. The conserved DEAD-box helicase motifs and domains of Vasa that interact with other proteins (Oskar and eukaryotic initiation factor 5B (eIF5B) are also shown. B, Vasa or nonimmune mouse serum (NMS) immunoprecipitates (IP) from D. melanogaster ovaries were resolved by NuPAGE and stained by silver-staining or probed on Western blots with SYM11 or ASYM24 (NMS, nonimmune mouse serum; negative control). Vasa protein is indicated and was also confirmed by mass spectrometry (see supplemental Table 1). H.c, heavy antibody chain; l.c, light chain. Lysate or Vasa immunoprecipitates from WT or dPRMT5/csul ovaries were probed on Western blots with anti-Vasa (C); SYM11 and ASYM24 (D); and anti-Tudor (E). Localization of Vasa in WT and dPRMT5/csul ovaries in early (F) or late egg chambers (G); arrowhead indicates pole (germ) plasm. Scale bar is 40 μm.
dPRMT5/Csul/Dart5 is the enzyme that is responsible for sDMA modifications of Sm proteins (6, 7) and of Piwi family proteins (36). We reasoned that dPRMT5 might be the methyltransferase that produces sDMAs in Vasa as well. To test this, we used ovaries from csulRM50/Df(2R)Jp7 females which give rise to embryos that are genetic nulls for dPRMT5 (6) and w− as a wild-type (WT) control. We immunoprecipitated Vasa proteins from WT and homozygous dPRMT5/csul mutant ovaries (csul) and probed the immunoprecipitates with anti-Vasa, SYM11, and ASYM24 (Fig. 2, C and D). Similar amounts of Vasa protein were detected in the lysate and in Vasa immunoprecipitates from WT and csul ovaries (Fig. 2C), indicating that the protein levels of Vasa are not affected in csul mutants. As shown in Fig. 2D, SYM11 reacted with immunopurified Vasa from WT but not from csul ovaries, indicating that dPRMT5 is essential for production of Vasa sDMAs. ASYM24 reacted with Vasa from WT and csul ovaries (Fig. 2D), indicating that Vasa contains aDMAs in the absence of dPRMT5 activity. D. melanogaster homologs of type I PRMTs, such as dPRMT1 (dart1) and dPRMT4 (dart4) have been identified, and dPRMT1 has been reported to methylate Vasa in vitro (55). dPRMT1 or other Drosophila type I PRMTs might be the enzyme that produces aDMAs in Vasa.
We have recently shown that sDMAs of Aub direct its interaction with Tudor protein (39). We hypothesized that sDMAs of Vasa might also mediate interaction with Tudor domain-containing proteins. However, as shown in Fig. 2E, a similar amount of Tudor protein was found in Vasa immunoprecipitates from WT and csul ovaries. This result suggests that the association of Vasa with Tudor may be independent of sDMA modifications of Vasa.
Next, we analyzed by confocal microscopy the localization of Vasa in WT and csul ovarioles. Representative images are shown in Fig. 2, F and G, and reveal that the localization of Vasa was not altered in csul early egg chambers (Fig. 2F), and Vasa accumulated in the pole plasm of csul oocytes (Fig. 2G). Staining with secondary antibody only (negative control) showed variable nonspecific staining of the outer aspect of anterior follicle cells without staining in nurse cells or in the oocyte (supplemental Fig. 1).
Mouse Vasa Protein MVH Contains sDMAs and aDMAs and Binds to Tdrd1, Tdrd6, Miwi, and Mili
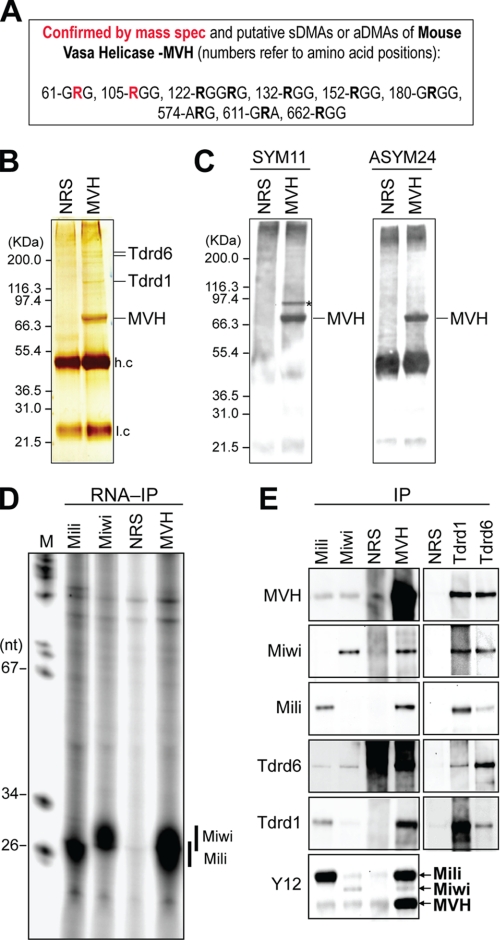
We next asked whether MVH contained methylated arginines (Fig. 3A). We immunoprecipitated MVH from mouse testis lysates and analyzed the immunoprecipitates by NuPAGE and silver staining (Fig. 3B). Mass spectrometry analysis identified 574 peptides of MVH, providing 91% coverage (supplemental Table 2), and indicated with high confidence the presence of two DMAs in endogenous MVH (Fig. 3A and supplemental Table 3). The first is Arg62 and is found in a GRG triplet, which is the classic substrate for sDMA modification by type II PRMTs. The second is Arg105 and is found in an RGG triplet that is the typical substrate for aDMA modification by type I PRMTs. Monomethylated arginines were also identified with high confidence (see supplemental Table 3). Additional proteins co-immunoprecipitated with MVH and were identified by mass spectrometry as Tdrd6 and Tdrd1 (Fig. 3B and supplemental Table 1). To further interrogate the presence of arginine methylation in MVH, we probed Western blots of immunopurified MVH with SYM11 and ASYM24 antibodies. As shown in Fig. 3C, both SYM11 and ASYM24 reacted strongly with MVH, indicating that, similar to X. laevis and D. melanogaster Vasa proteins, MVH contains both sDMAs and aDMAs. SYM11 (but not ASYM24) also reacted with a ∼95-kDa band present in the MVH immunoprecipitates (shown with asterisk in Fig. 3C). This protein likely corresponds to Miwi because Miwi is a ∼95-kDa protein found in MVH immunoprecipitates (see below), and Miwi is recognized by SYM11 but not ASYM24 (36). We also analyzed RNAs from MVH, Mili, and Miwi immunoprecipitates of mouse testis lysates. As shown in Fig. 3D, short (∼26 nucleotides) and long (∼29 nucleotides) piRNAs were present in the Mili and Miwi immunoprecipitates, respectively, consistent with the known association of Mili with short piRNAs and Miwi with long piRNAs (33, 34). MVH immunoprecipitates contained both short and long piRNAs (Fig. 3D), indicating that MVH associates with Mili and Miwi Piwi-ribonucleoproteins. Collectively, these results suggest that MVH interacts and forms complexes with Tdrd1, Tdrd6, Miwi, and Mili. These interactions were confirmed by immunoprecipitations from mouse testis lysates and Western blots against each protein (Fig. 3E). We detected the previously reported Mili-Tdrd1 (37, 56), Miwi-Tdrd6 (38, 41), and Mili-MVH and Miwi-MVH (57) interactions (Fig. 3E). In MVH immunoprecipitates, Miwi, Mili, Tdrd6, and Tdrd1 were detected, whereas MVH was present in Miwi, Mili, Tdrd1, and Tdrd6 immunoprecipitates.
FIGURE 3.
Mouse Vasa protein MVH contains sDMAs and aDMAs and associates with Miwi, Mili, Tdrd1, and Tdrd6. A, confirmed and putative DMAs in MVH. Arg62 and Arg105 (red) were identified as dimethylated by mass spectrometry (mass spec) (see supplemental Table 3). B, MVH immunoprecipitates from mouse testis were resolved by NuPAGE and stained by silver-staining. Co-precipitating proteins (indicated) were identified by mass spectrometry (see supplemental Table 1). NRS, nonimmune rabbit serum (negative control). C, MVH immunoprecipitates were probed on Western blots with SYM11 or ASYM24; an asterisk indicates a protein band that likely corresponds to Miwi. D, RNA-immunoprecipitation with indicated antibodies from mouse testis. Miwi- and Mili-bound piRNAs are indicated. E, immunoprecipitates (IP; NRS, MVH, Miwi, Mili, Tdrd1, and Tdrd6) from mouse testis were probed on Western blots with indicated antibodies. nt, nucleotide; M, marker.
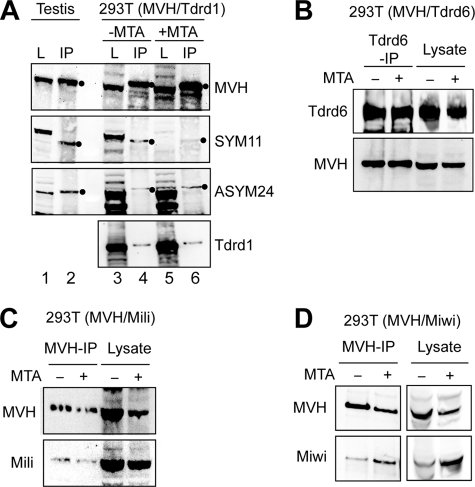
To further investigate the protein associations of MVH, we co-expressed in 293T cells Myc-tagged MVH with either Miwi or Mili (both untagged), Tdrd1-His or FLAG-Tdrd6, and we tested the association between the proteins with co-immunoprecipitation experiments. We also cultured a separate cohort of transfected cells in the presence of the methyltransferase inhibitor 5′-deoxy-5′-(methylthio)-adenosine (MTA), to test the requirement of arginine methylation of MVH for these associations. MVH was immunoprecipitated, and the extent of arginine methylation of the purified protein was examined by SYM11 and ASYM24 Western blots. Immunopurified MVH expressed in 293T cells reacted with SYM11 and ASYM24 (Fig. 4A, lane 4), like endogenous MVH immunoprecipitated from mouse testis (Fig. 4A, lane 2), indicating that MVH expressed in 293T cells contained sDMAs and aDMAs. MTA treatment decreased substantially the SYM11 reactivity of 293T total cell lysates but had small effect in ASYM24 reactivity (Fig. 4A, lane 5; compare with untreated lysates in lane 3; entire blot membranes are shown in supplemental Fig. 2). Similar results were seen for the 293T-expressed MVH (Fig. 4A, lane 6), indicating that the sDMA (but not the aDMA) levels of MVH were reduced with the methyltransferase inhibitor. However, the association of MVH with Tdrd1 was similar between the nontreated and MTA-treated cells (Fig. 4A, lanes 4 and 6). Similarly, Tdrd6-MVH, Mili-MVH, and Miwi-MVH interactions were tested in co-transfection experiments in 293T cells, by probing Tdrd6 immunoprecipitates with anti-MVH antibody (Fig. 4B) or by probing MVH immunoprecipitates with anti-Mili (Fig. 4C) or anti-Miwi (Fig. 4D) antibody. As shown in Fig. 4, B–D, MVH associated with Tdrd6, Mili, and Miwi, and none of these interactions were affected by MTA.
FIGURE 4.
Association of MVH with Tdrd1, Tdrd6, Miwi and Mili in untreated and MTA-treated 293T cells. A, lysate (L) or MVH immunoprecipitates (IP) from mouse testis or 293T cells expressing MVH and Tdrd1, in the presence (+) or absence (−) of the methylation inhibitor MTA, were probed on Western blots with indicated antibodies. All filled circles indicate immunopurified MVH. Many methylated proteins are recognized by SYM11 and ASYM24 in the lysates of untreated 293T cells (lane 3 in A and B). Entire blot membranes for SYM11 and ASYM24 are shown in supplemental Fig. 2. B, Tdrd6 immunoprecipitates from 293T cells expressing MVH and Tdrd6, in the presence or absence of MTA, were probed on Western blots with anti-Tdrd6 or anti-MVH antibodies. MVH immunoprecipitates from 293T cells expressing MVH and either Mili (C) or Miwi (D) and in the presence (+) or absence (−) of MTA, were probed on Western blots with indicated antibodies.
DISCUSSION
sDMA modification of Piwi proteins is essential for their interaction with Tudor domain-containing proteins (Tdrds) (37–39, 40, 42). Recent studies have shown that sDMA-modified Piwi proteins interact with specific Tdrds and form distinct, subcellular bodies that likely reflect the sites of biogenesis and function of piRNAs (41, 43, 58).
In D. melanogaster, sDMA modification of Aub is required for its interaction with Tudor (39, 40) and for accumulation of Aub in the pole plasm (39). The Tudor-Aub and Tudor-Ago3 interactions also appear to play a role in biogenesis of piRNAs (40).
Here, we show that Vasa contains aDMA and sDMAs and like Aub, the sDMAs are catalyzed by dPRMT5. However, unlike Aub, neither the association between Vasa and Tudor nor the localization of Vasa in the pole plasm is dependent on sDMAs of Vasa. In mouse, MVH associates with Miwi, Mili, Tdrd1, and Tdrd6, but the MTA methylation inhibitor did not affect these interactions. These findings suggest that sDMAs of Vasa and MVH may be dispensable for interaction with Tudor and with Tdrd1 and Tdrd6, respectively. However, we note that MTA treatment did not affect appreciably the extent of aDMAs in MVH expressed in 293T cells, and it is also possible that residual sDMA modifications remained in MVH. Thus, it is possible that arginine methylation is still required for association of MVH with Tdrds. Another possibility is that MVH interacts indirectly with Tdrd1 and Tdrd6 via its association with Miwi and Mili. Lastly, MVH may be interacting with as yet unidentified Tdrds in an sDMA-dependent manner. Such association may be important for cytoplasmic compartmentalization of MVH akin to that of Piwi proteins when bound to Tudor proteins (41, 43, 58). It is interesting to note that in mouse male gonocytes, MVH co-localizes with Mili/Tdrd1 in intermitochondrial cement/pi-bodies and also with Miwi2/Tdrd9 in nuage/piP-bodies (43, 58), suggesting that MVH has important roles in piRNA biogenesis and/or function. Future experiments are required to uncover the specific function of the conserved arginine methylation of Vasa proteins.
Supplementary Material
Acknowledgments
We are grateful to P. Lasko (McGill University) for Tudor antibody; to J. Anne (Deutsches Krebsforschungszentrum) for csul flies; to S. Chuma (Kyoto University) for Tdrd1 and Tdrd6 antibodies and cDNA constructs; to S. Kuramochi-Miyagawa, T. Nakano (Osaka University), and T. Noce (Mitsubishi Kagaku Institute of Life Sciences) for MVH cDNA construct; and to members of the Mourelatos lab for discussions.
This work was supported, in whole or in part, by National Institutes of Health Grants GM76621 (to P. S. K.), MH086705 (to T. A. J.), GM0720777, and NS056070 (to Z. M.). This work was also supported by a Human Frontier Science Program Long Term fellowship (to Y. K.) and in part by The Institute for the Translational Medicine and Therapeutics grants (to Z. M.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Tables 1–3 and Figs. 1 and 2.
Y. Kirino, A. Vourekas, N. Kim, F. de Lima Alves, J. Rappsilber, P. S. Klein, T. A. Jongens, and Z. Mourelatos, unpublished data.
- MVH
- mouse Vasa homolog
- WT
- wild-type
- piRNA
- Piwi-interacting RNA
- PRMT
- protein methyltransferase
- MTA
- 5′-deoxy-5′-(methylthio)-adenosine
- MS
- mass spectrometry.
REFERENCES
- 1.Williamson A., Lehmann R. (1996) Annu. Rev. Cell Dev. Biol. 12, 365–391 [DOI] [PubMed] [Google Scholar]
- 2.Strome S., Lehmann R. (2007) Science 316, 392–393 [DOI] [PubMed] [Google Scholar]
- 3.Dansereau D. A., Lasko P. (2008) Methods Mol. Biol. 450, 3–26 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Rongo C., Lehmann R. (1996) Trends Genet. 12, 102–109 [DOI] [PubMed] [Google Scholar]
- 5.Harris A. N., Macdonald P. M. (2001) Development 128, 2823–2832 [DOI] [PubMed] [Google Scholar]
- 6.Anne J., Ollo R., Ephrussi A., Mechler B. M. (2007) Development 134, 137–146 [DOI] [PubMed] [Google Scholar]
- 7.Gonsalvez G. B., Rajendra T. K., Tian L., Matera A. G. (2006) Curr. Biol. 16, 1077–1089 [DOI] [PubMed] [Google Scholar]
- 8.Cote J., Richard S. (2005) J. Biol. Chem. 280, 28476–28483 [DOI] [PubMed] [Google Scholar]
- 9.Boswell R. E., Mahowald A. P. (1985) Cell 43, 97–104 [DOI] [PubMed] [Google Scholar]
- 10.Thomson T., Lasko P. (2005) Cell Res. 15, 281–291 [DOI] [PubMed] [Google Scholar]
- 11.Arkov A. L., Wang J. Y., Ramos A., Lehmann R. (2006) Development 133, 4053–4062 [DOI] [PubMed] [Google Scholar]
- 12.Lasko P. F., Ashburner M. (1988) Nature 335, 611–617 [DOI] [PubMed] [Google Scholar]
- 13.Hay B., Jan L. Y., Jan Y. N. (1988) Cell 55, 577–587 [DOI] [PubMed] [Google Scholar]
- 14.Liang L., Diehl-Jones W., Lasko P. (1994) Development 120, 1201–1211 [DOI] [PubMed] [Google Scholar]
- 15.Johnstone O., Lasko P. (2004) Development 131, 4167–4178 [DOI] [PubMed] [Google Scholar]
- 16.Tomancak P., Guichet A., Zavorszky P., Ephrussi A. (1998) Development 125, 1723–1732 [DOI] [PubMed] [Google Scholar]
- 17.Tinker R., Silver D., Montell D. J. (1998) Dev. Biol. 199, 1–10 [DOI] [PubMed] [Google Scholar]
- 18.Liu N., Dansereau D. A., Lasko P. (2003) Curr. Biol. 13, 1905–1909 [DOI] [PubMed] [Google Scholar]
- 19.Styhler S., Nakamura A., Lasko P. (2002) Dev. Cell 3, 865–876 [DOI] [PubMed] [Google Scholar]
- 20.Ephrussi A., Lehmann R. (1992) Nature 358, 387–392 [DOI] [PubMed] [Google Scholar]
- 21.Breitwieser W., Markussen F. H., Horstmann H., Ephrussi A. (1996) Genes Dev. 10, 2179–2188 [DOI] [PubMed] [Google Scholar]
- 22.Raz E. (2000) Genome Biol. 1, REVIEWS1017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Komiya T., Itoh K., Ikenishi K., Furusawa M. (1994) Dev. Biol. 162, 354–363 [DOI] [PubMed] [Google Scholar]
- 24.Ikenishi K., Tanaka T. S. (2000) Dev. Growth Differ. 42, 95–103 [DOI] [PubMed] [Google Scholar]
- 25.Ikenishi K., Tanaka T. S. (1997) Dev. Growth Differ. 39, 625–633 [DOI] [PubMed] [Google Scholar]
- 26.Leroy P., Alzari P., Sassoon D., Wolgemuth D., Fellous M. (1989) Cell 57, 549–559 [DOI] [PubMed] [Google Scholar]
- 27.Tanaka S. S., Toyooka Y., Akasu R., Katoh-Fukui Y., Nakahara Y., Suzuki R., Yokoyama M., Noce T. (2000) Genes Dev. 14, 841–853 [PMC free article] [PubMed] [Google Scholar]
- 28.Bedford M. T., Richard S. (2005) Mol. Cell 18, 263–272 [DOI] [PubMed] [Google Scholar]
- 29.Krause C. D., Yang Z. H., Kim Y. S., Lee J. H., Cook J. R., Pestka S. (2007) Pharmacol. Ther. 113, 50–87 [DOI] [PubMed] [Google Scholar]
- 30.Friesen W. J., Wyce A., Paushkin S., Abel L., Rappsilber J., Mann M., Dreyfuss G. (2002) J. Biol. Chem. 277, 8243–8247 [DOI] [PubMed] [Google Scholar]
- 31.Anne J., Mechler B. M. (2005) Development 132, 2167–2177 [DOI] [PubMed] [Google Scholar]
- 32.Cavey M., Hijal S., Zhang X., Suter B. (2005) Development 132, 459–468 [DOI] [PubMed] [Google Scholar]
- 33.Girard A., Hannon G. J. (2008) Trends Cell Biol. 18, 136–148 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Klattenhoff C., Theurkauf W. (2008) Development 135, 3–9 [DOI] [PubMed] [Google Scholar]
- 35.Siomi M. C., Saito K., Siomi H. (2008) FEBS Lett. 582, 2473–2478 [DOI] [PubMed] [Google Scholar]
- 36.Kirino Y., Kim N., de Planell-Saguer M., Khandros E., Chiorean S., Klein P. S., Rigoutsos I., Jongens T. A., Mourelatos Z. (2009) Nat. Cell Biol. 11, 652–658 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Reuter M., Chuma S., Tanaka T., Franz T., Stark A., Pillai R. S. (2009) Nat. Struct. Mol. Biol. 16, 639–646 [DOI] [PubMed] [Google Scholar]
- 38.Vagin V. V., Wohlschlegel J., Qu J., Jonsson Z., Huang X., Chuma S., Girard A., Sachidanandam R., Hannon G. J., Aravin A. A. (2009) Genes Dev. 23, 1749–1762 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Kirino Y., Vourekas A., Sayed N., Lima Alves F., Thomson T., Lasko P., Rappsilber J., Jongens T. A., Mourelatos Z. (2010) RNA 16, 70–78 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Nishida K. M., Okada T. N., Kawamura T., Mituyama T., Kawamura Y., Inagaki S., Huang H., Chen D., Kodama T., Siomi H., Siomi M. C. (2009) EMBO J. 28, 3820–3831 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Vasileva A., Tiedau D., Firooznia A., Müller-Reichert T., Jessberger R. (2009) Curr. Biol. 19, 630–639 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Chen C., Jin J., James D. A., Adams-Cioaba M. A., Park J. G., Guo Y., Tenaglia E., Xu C., Gish G., Min J., Pawson T. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 20336–20341 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Shoji M., Tanaka T., Hosokawa M., Reuter M., Stark A., Kato Y., Kondoh G., Okawa K., Chujo T., Suzuki T., Hata K., Martin S. L., Noce T., Kuramochi-Miyagawa S., Nakano T., Sasaki H., Pillai R. S., Nakatsuji N., Chuma S. (2009) Dev. Cell 17, 775–787 [DOI] [PubMed] [Google Scholar]
- 44.Lau N. C., Ohsumi T., Borowsky M., Kingston R. E., Blower M. D. (2009) EMBO J. 28, 2945–2958 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Hosokawa M., Shoji M., Kitamura K., Tanaka T., Noce T., Chuma S., Nakatsuji N. (2007) Dev. Biol. 301, 38–52 [DOI] [PubMed] [Google Scholar]
- 46.Thomson T., Liu N., Arkov A., Lehmann R., Lasko P. (2008) Mech. Dev. 125, 865–873 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bossi E., Fabbrini M. S., Ceriotti A. (2007) Methods Mol. Biol. 375, 107–131 [DOI] [PubMed] [Google Scholar]
- 48.Shevchenko A., Wilm M., Vorm O., Mann M. (1996) Anal. Chem. 68, 850–858 [DOI] [PubMed] [Google Scholar]
- 49.Rappsilber J., Ishihama Y., Mann M. (2003) Anal. Chem. 75, 663–670 [DOI] [PubMed] [Google Scholar]
- 50.Ishihama Y., Rappsilber J., Andersen J. S., Mann M. (2002) J. Chromatogr. A 979, 233–239 [DOI] [PubMed] [Google Scholar]
- 51.Lerner E. A., Lerner M. R., Janeway C. A., Jr., Steitz J. A. (1981) Proc. Natl. Acad. Sci. U.S.A. 78, 2737–2741 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Brahms H., Raymackers J., Union A., de Keyser F., Meheus L., Lührmann R. (2000) J. Biol. Chem. 275, 17122–17129 [DOI] [PubMed] [Google Scholar]
- 53.Boisvert F. M., Cote J., Boulanger M. C., Cleroux P., Bachand F., Autexier C., Richard S. (2002) J. Cell Biol. 159, 957–969 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Boisvert F. M., Côté J., Boulanger M. C., Richard S. (2003) Mol. Cell Proteomics 2, 1319–1330 [DOI] [PubMed] [Google Scholar]
- 55.Boulanger M. C., Miranda T. B., Clarke S., Di Fruscio M., Suter B., Lasko P., Richard S. (2004) Biochem. J. 379, 283–289 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wang J., Saxe J. P., Tanaka T., Chuma S., Lin H. (2009) Curr. Biol. 19, 640–644 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Kuramochi-Miyagawa S., Kimura T., Ijiri T. W., Isobe T., Asada N., Fujita Y., Ikawa M., Iwai N., Okabe M., Deng W., Lin H., Matsuda Y., Nakano T. (2004) Development 131, 839–849 [DOI] [PubMed] [Google Scholar]
- 58.Aravin A. A., van der Heijden G. W., Castañeda J., Vagin V. V., Hannon G. J., Bortvin A. (2009) PLoS. Genet 5, e1000764. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.