Abstract
Here we describe the design, preparation and characterization of 10 EF-Tu mutants of potential utility for the study of Escherichia coli elongation factor Tu (EF-Tu) interaction with tRNA by a fluorescence resonance energy transfer assay. Each mutant contains a single cysteine residue at positions in EF-Tu that are proximal to tRNA sites within the aminoacyl-tRNA·EF-Tu·GTP ternary complex that have previously been labeled with fluorophores. These positions fall in the 323–326 and 344–348 regions of EF-Tu, and at the C terminus. The EF-Tus were isolated as N-terminal fusions to glutathione S-transferase (GST), which was cleaved to yield intact EF-Tus. The mutant EF-Tus were tested for binding to GDP, binding to tRNA in gel retardation and protection assays, and activity in poly-U translation in vitro. The results indicate that at least three EF-Tu mutants, K324C, G325C and E348C, are suitable for further studies. Remarkably, GST fusions that were not cleaved were also active in the various assays, despite the N-terminal fusion.
Keywords: elongation factor Tu, FRET, site-directed mutagenesis, structure-function, transfer RNA
Introduction
The quest to collect data that provide precise real-time information on complex actions of single molecules often demands advanced approaches to fluorescence labeling. The work described in this paper is part of a larger project funded by a grant from the $1000 Genome Initiative of NIH and is aimed at collecting sequence data for nucleic acids using the ribosome as an engine to facilitate this process. The principle of the method is single-pair fluorescence resonance energy transfer (spFRET) involving two molecules, a specific aminoacyl-tRNA (aa-tRNA) conjugated to a fluorescent dye (donor) and the elongation factor Tu (EF-Tu) conjugated to a quencher (acceptor). The active form of EF-Tu is bound to GTP and catalyzes the binding of aminoacyl-tRNA to the A site of the mRNA-programmed ribosome. Upon donating aa-tRNA to the codon-programmed decoding site of the ribosome, the fluorescent dye and the quencher will become spatially separated as a result of EF-Tu dissociation from the ribosome. Such dissociation follows correct codon–anticodon pairing and GTP hydrolysis. The donor–acceptor separation will be accompanied by a burst of donor fluorescence which can be detected by a photon counting instrument capable of signal acquisition from sub-femtoliter volumes. A fuller description has been published previously (Mandecki et al., 2008).
The success of this approach depends on well-characterized, properly labeled and active components, i.e. tRNA and EF-Tu. Here we describe the construction and properties of several E.coli EF-Tu mutants designed to meet the requirements of the proposed sequencing method described earlier.
One of the challenges in creating a proper EF-Tu construct was to make sure that the fluorescent label is directed, in a conjugation experiment, to the desired site on EF-Tu. The problem was that the native E.coli EF-Tu has three cysteine residues, Cys 81, Cys 137 and Cys 255, which could potentially be labeled, thus complicating greatly the interpretation of the results. Cys 81 is located within domain 1, between β-strand d (residues 75–80) and α-helix B (residues 84–92, switch region II) in a turn region (Fig. 1). Cys 137 is located within domain 1, between β-strand f (residues 130–135) and α-helix D (residues 143–159) in a turn region. Cys 255 is located within domain 2, at the end of β-strand f2 (residues 251–255) (Song et al., 1999).
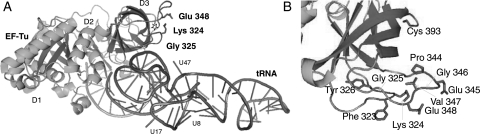
Fig. 1.
Structure of the EF-Tu·GTP·aa-tRNA complex. (A) Highlighted are positions of three key EF-Tu mutations and three nucleotide residues in tRNA commonly used for labeling with fluorescent dyes. The C-terminal residue is marked with a star. Symbols D1, D2 and D3 denote Domains 1, 2 and 3, respectively. Only fragments of Domain 2 can be seen in this drawing. The drawing is based on the coordinates from 1ob2.pdb (R.C. Nielsen, O. Kristensen, M. Kjeldgaard, S. Thirup and P. Nissen, to be published). Phe-tRNAPhe is from baker's yeast, and EF-Tu is from E.coli. The figure was created in PyMol. (B) Close-up view of the mutated regions 323–326, 344–348 and residue 393 showing positions of the amino acid side-chains.
The thiol groups of all three cysteines have been found to be reactive (C.R.K., unpublished results). However, the cysteines at positions 138 and 255 are expected to be the most reactive due to their location on the surface. In order to avoid interferences during labeling with the quencher, the three native cysteines were mutated to Ser, Ala and Val, respectively, based on sequence alignment and homologies to EF-Tus from other species. The activity of the resulting triple-point mutant (named EF-TuSAV) was assessed by monitoring its activity in GDP-binding and translation in vitro. Wild-type behavior was observed in both of these assays (C.R.K., unpublished results).
We used the gene for EF-TuSAV to construct a series of EF-Tu mutants having only one cysteine residue. Positions of the introduced Cys were chosen based on two criteria, the proximity of the residue to a fluorescent dye attached to aa-tRNA and the apparent lack of essential involvement of the mutated amino acid in the function of EF-Tu. Amino acid residues chosen for mutagenesis to Cys are located in the turn regions between β-strands a3 and b3 (Phe 323, Lys 324, Gly 325, Tyr 326), between β-strands c3 and d3 (Pro 344, Glu 345, Gly 346, Val 347, Glu 348) and at the C terminal end of EF-Tu (393Cins) (Fig. 1).
The aim of this study was to characterize EF-Tu mutants with only one Cys residue introduced in 10 various positions and select those that are most active and suitable for further use in the sequencing project. The mutated EF-Tu molecules were expressed as fusions to glutathione S-transferase (GST), a 201 amino acid protein (Mr of 23 kDa), to allow isolation and purification. A linker was placed between the two sequences. The linker, Ile-Glu-Gly-Arg, contains a factor Xa cleavage site, thus EF-Tu can be released from the GST fusion in its native form through a proteolytic cleavage with factor Xa. No additional residues are added via the inserted linker. The first residue of the cleaved EF-Tu corresponds to residue Ser 1.
The properties of both types of mutated EF-Tus, i.e. native and fused to GST, were evaluated by a variety of in vitro assays to examine the proteins' ability to bind GDP and aa-tRNA, as well as their activity in translating a poly(U) message into poly(Phe). The three most active EF-Tu mutants, EF-TuSAVK324C, EF-TuSAVG325C and EF-TuSAVE348C, were chosen for further study.
Materials and methods
Reagents
Reagents for electrophoresis and silver gel staining were purchased from Bio-Rad. Ampicillin, sorbitol, betaine, glutathione-agarose, reduced glutathione, phenylmethanesulfonyl fluoride (PMSF), imidazole, phosphoenolpyruvate (PEP), pyruvate kinase (500 U/mg), DTT, GTP, GDP, ATP, poly U, spermidine, putrescine were from Sigma. The QuikChange Lightning site-directed mutagenesis kit (QCM) was purchased from Stratagene (La Jolla, CA, USA). Factor Xa was obtained from Novagen (concentration 2 U/µl). Factor Xa removal resin was provided by Qiagen. [8,5′-3H] guanosine 5′-diphosphate, trisodium salt (8.2 Ci/mmol) and L-[14C] phenylalanine (496 mCi/mmol) were from PerkinElmer (Boston, MA, USA). IPTG was obtained from Promega. Escherichia coli tRNAPhe was purchased from Chemical Block (Moscow, Russia). DABCYL Plus C2 maleimide was obtained from Anaspec (Fremont, CA, USA).
EF-Tu expression vector
The plasmid pGEX-FX-tufA was constructed as previously described (Knudsen et al., 1992). The tufA gene encodes E.coli EF-Tu. Subsequently, in a series of three mutagenesis steps in which the Stratagene's QCM kit was used, the three native Cys residues at positions 81, 137 and 255 of E.coli EF-Tu were replaced by the Ser, Ala and Val residues, respectively; hence, the name pGEX-FX-tufA-SAV. This plasmid was constructed in the laboratory of one of the coauthors (C.R.K.).
The plasmid expressing Staphylococcus aureus EF-Tu was a gift to B.S.C. from Mandana Sassanfar.
Site-directed mutagenesis
The plasmid pGEX-FX-tufA-SAV was used as the mutagenesis template. Ten primer pairs designed to mutate the tufA gene were synthesized by Biosynthesis, Inc. (Lewisville, TX, USA). The mutagenesis procedure is a modification of methods described by Wang and Malcolm (1999) and Tseng et al. (2008) and consists of two steps. In step one, two extension reactions are performed in different tubes: one containing the sense primer and the other the antisense primer. In a single primer extension reaction, 50–200 ng of the pGEX-FX-tufA-SAV plasmid along with 10 pmol of the primer were used in the 25 µl reaction containing 2.5 µl of 10× reaction buffer, 0.5 µl of QuikChange Lightning enzyme, 0.5 µl of dNTP cocktail, 0.75 µl of QuikSolution reagent and deionized water. The extension reaction was initiated by preheating the reaction mixture to 95°C for 2 min, followed by three cycles of 95°C for 20 s, 60°C for 10 s, 68°C for 3.5 min and 68°C for 5 min. Following the completion of the extension reactions, 25 µl of each reaction was mixed in one tube and subjected to the standard QuikChange Lightning (QCM) procedure (Stratagene, La Jolla, CA, USA). Following the cycling reaction, 2 µl of DpnI was added (as per the QCM protocol), mixed well and incubated at 37°C for 1.5 h before transformation. Two microliters of the final PCR products were transformed into 45 µl of XL10-Gold ultracompetent cells, and 10 µl volumes were spread on LB agar plates containing ampicillin (100 µg/ml). Five clones were isolated and amplified in 3 ml of LB medium containing ampicillin (100 µg/ml). Plasmids were isolated using Wizard Plus SV minipreps DNA purification system and a vacuum adapter kit (Promega, Madison, WI, USA). To confirm the introduction of desired mutations, plasmid DNAs were sequenced.
Expression of EF-Tu wt, SAV, mutants and His-tagged versions
Escherichia coli and S.aureus EF-Tu were expressed as a fusion with GST or a His6 tag, respectively. EF-Tu fused to GST was expressed in E.coli strain XL-Blue10, from the expression vector pGEX-FX-tufA. Cells expressing wild-type EF-Tu were grown in LB medium with ampicillin and protein production was induced with 1 mM IPTG, followed by 3 h incubation at 30°C. Because of their tendency to form inclusion bodies, the EF-Tu mutants were expressed in LB medium enriched with 1 M sorbitol and 2.5 mM betaine, at 25°C for 18–24 h.
Isolation of EF-Tu
Isolation of EF-Tu fused to GST was performed according to Knudsen et al. (1992) with minor modifications. The cell culture was placed on ice and medium removed by centrifugation at 8500g (Beckmann rotor JA 10; 7000 rpm) for 15 min. Cells were washed and resuspended in buffer U (50 mM Tris–HCl, 100 mM NaCl, 10 mM MgCl2, 15 µM GDP, pH 7.6 at 4°C), followed by sonication (6 × 10 s) in the presence of 1% Triton X-100. Cell debris was removed by centrifugation at 7500g (JA 25,50; 8000 rpm) for 15 min. The supernatant was loaded onto an affinity column (glutathione-agarose, 1 ml bed volume) equilibrated with buffer U. Unbound protein was washed off the column with 15–20 ml of buffer U. Fusion protein was then eluted with 10 ml of buffer U containing 5 mM reduced glutathione. Fractions containing GST-EF-Tu were pooled, concentrated and glutathione was washed off with buffer U on Millipore 10 kDa cut-off filters. EF-Tu was recovered from the GST fusion by proteolysis with factor Xa in a reaction performed according to the manufacturer's recommendations. The cleavage was performed overnight at 4°C using one unit of enzyme per 200 µg of substrate, followed by removal of factor Xa using the Xa removal resin, according to the manufacturer's instructions. After cleavage was completed, GST was separated from EF-Tu by affinity chromatography on a glutathione-agarose column. To ensure that any remaining factor Xa present in the preparation of EF-Tu is inactive, 0.5 mM PMSF was added to fractions containing the protein. Finally, EF-Tu was concentrated on Millipore filters and stored in 15% glycerol at −20°C.
His-tagged EF-Tu was isolated according to a standard procedure. Briefly, cells were washed and resuspended in buffer A (50 mM Tris–HCl, pH 7.6, 60 mM NH4Cl, 7 mM MgCl2, 7 mM 2-mercaptoethanol, 15% glycerol). Sonication, centrifugation, washing and storage were the same as for GST-fused EF-Tu. His-tagged EF-Tu was retained on a Ni-Sepharose column (bed volume of 1 ml). Unbound proteins were washed off with 10 ml of buffer B (buffer A with 300 mM KCl), followed by 10 ml of buffer C (buffer B with 20 mM imidazole). Bound EF-Tu was eluted from the resin with 10 ml of buffer D (buffer B with 300 mM imidazole). Concentrations of all isolated proteins were measured by the Bradford (1976) method. A reducing agent, either DTT or β-mercaptoethanol, was included in all purification steps and assays to ascertain that intermolecular dimers are not formed.
Expression and isolation of PheRS
Escherichia coli strain XL1Blue, overexpressing E.coli phenylalanyl-tRNA synthetase (PheRS), was kindly provided by Dr M. Ibba. The PheRS gene (expressing both α and β subunits) was carried on the pQE31 vector. Subunit α had an N-terminal His6 tag that allowed its isolation together with the co-purifying subunit β using standard His-tagged protein purification procedures. Expression was induced with 1 mM IPTG at 30°C for 24 h. Cells were spun down, resuspended in sonication buffer (20 mM Tris–HCl, pH 8.0, 300 mM NaCl, 5 mM imidazole) and disrupted by sonication. After spinning down cell debris, the lysate was applied onto a Ni-Sepharose column. Unbound proteins were washed off with 4 ml of sonication buffer and 10 ml of wash buffer (20 mM Tris–HCl, pH 8.0, 300 mM NaCl, 20 mM imidazole), followed by elution of bound proteins with elution buffer (20 mM Tris–HCl, pH 8.0, 300 mM NaCl, 250 mM imidazole). Fractions containing PheRS were collected, concentrated, transferred to storage buffer (20 mM Tris–HCl, pH 7.4, 150 mM KCl, 5 mM 2-mercaptoethanol) and stored in 50% glycerol at −20°C. Protein concentration was measured according to Bradford (1976).
GDP-binding assay
GDP-binding activity was measured according to Miller and Weissbach (1974). Reaction mixtures (20 µl) contained 200 pmoles of 3H-GDP (specific activity 8.2 Ci/mmol) and 10 pmoles of EF-Tu in binding buffer (50 mM Tris–HCl, pH 7.4, 50 mM NH4Cl, 10 mM KCl, 10 mM MgCl2, 5 mM DTT). After incubation at 30°C for 30 min, the reaction was stopped by diluting 10-fold with cold washing buffer (10 mM Tris–HCl, pH 7.4, 10 mM MgCl2, 10 mM NH4Cl) and placing the reaction on ice. Two hundred microliter portions were filtered through 0.45 µm nitrocellulose filters (Schleicher & Schuell BA 85 or Millipore HAWP02500) soaked in washing buffer. The filters were then washed with 5–10 ml of washing buffer and the 3H-GDP·EF-Tu complex bound to the filters was counted in a scintillation counter. The background GDP binding obtained in a control without EF-Tu was subtracted from the results in the presence of EF-Tu.
Aminoacylation
The aminoacylation mix contained 80 mM HEPES (pH 7–7.6), 8 mM MgCl2, 24 mM KCl, 0.08 mM EDTA, 2 mM ATP, 200 µM [14C]Phe (specific activity 496 mCi/mmol), 0.2 mg/ml E.coli tRNAPhe and 7 ng/µl of purified PheRS. After 10–30 min of incubation at 37°C, the reaction was stopped by precipitation with 5% TCA on 3 mm Whatman paper, followed by washing in 5% cold TCA (3 × 10 min) and ethanol (10 min) to remove unbound radioactive Phe, drying and scintillation counting.
Non-enzymatic hydrolysis protection assay
The labile aminoacyl bond of aa-tRNA in the ternary complex is protected by EF-Tu (Pingoud and Urbanke, 1979). The formation of ternary complex between EF-Tu:GTP and aminoacyl-tRNA was determined by quantifying the protective effect of the complex formation on non-enzymatic deacylation of aminoacyl-tRNA. The ternary complex was formed by incubating 2.7 or 0.5 µM EF-Tu with 0.5 µM [14C]Phe-tRNAPhe in 50 mM Tris–HCl, pH 7.4, 10 mM MgCl2, 50 mM NH4Cl, 1 mM PEP, 0.05 mg/ml pyruvate kinase, 5 mM DTT and 1 mM GTP at room temperature. At various time points, 10-µl aliquots of the reaction mixture were withdrawn and applied to Whatman 3 mm filters, which were washed three times in 5% cold trichloroacetic acid and once in cold ethanol. The dried filters were assayed for radioactivity using a Beckman Coulter LS 6500 liquid scintillation counter.
Translation in vitro
Reaction mixtures (20 µl) contained 10 mM magnesium acetate, 5 mM potassium phosphate, pH 7.4, 100 mM potassium glutamate, pH 7.7, 95 mM potassium chloride, 5 mM ammonium chloride, 0.5 mM calcium chloride, 1 mM spermidine, 8 mM putrescine, 1 mM DTT, 0.9 mM ATP, 4.5 mM PEP, 1 mM GTP, 91–150 µg/ml pyruvate kinase, 1.8–2.9 µM EF-G, 0.4 µM 70S ribosomes, 36–42 µg/ml [14C]Phe-tRNAPhe and variable amounts of EF-Tu. The reaction was carried out at 30 or 37°C for 30 min, and then incubated with an equal volume of 0.4 M NaOH at 37°C for 10 min. After hydrolysis of unreacted [14C]Phe-tRNAPhe was completed, the sample was applied onto Whatman filter paper, washed in 5% ice cold TCA (3x10 min) and in ethanol (10 min). Subsequently, the filter papers were dried, immersed in solution of 0.4% (w/v) Omnifluor (New England Nuclear) in toluene (4 g/l) and the radioactivity in the precipitates was determined by scintillation counting.
Ternary complex formation and stability: gel-shift assay
The method was performed according to Doi et al. (2007), with minor modifications. EF-Tu (2, 4 or 8 pmol) was preincubated with 7 mM GTP at 37°C for 15 min in 5 µl total volume containing 70 mM HEPES-KOH (pH 7.6), 52 mM NH4OAc, 8 mM Mg(OAc)2, 30 mM KCl, 1.4 mM DTT, 3 µM GDP, 2.6% glycerol, 7.2 mM PEP and 0.8 U/µl pyruvate kinase. To the preincubated EF-Tu solution, 3 µl of aa-tRNA (diluted in 6 mM KOAc) and 2 µl of the ternary complex buffer, containing 150 mM HEPES-KOH (pH 7.6), 195 mM NH4OAc and 30 mM Mg(OAc)2, were added. The mixture of EF-Tu and aa-tRNA was incubated at 37°C for 10 min. Electrophoresis of the samples was performed on 8% polyacrylamide gels at 4°C in a buffer containing 10 mM MES (pH 6), 65 mM NH4OAc, 10 mM Mg(OAc)2 and 10 µM GTP. Gels were stained with silver using Silver Stain Plus kit (Bio-Rad).
Labeling of EF-Tu with quencher
Labeling conditions were as follows (100 µl): 10 nmol EF-Tu or GST-EF-Tu in 50 mM Tris, pH 7.2 incubated with a 10- and 20-fold molar excess of DABCYL Plus in a reaction with EF-Tu and GST-EF-Tu, respectively, at 25°C for 2 h in darkness. Then, the unreacted dye was separated from labeled EF-Tu on a Sephadex G-25 column (NAP-5, GE Healthcare). Fractions containing protein were concentrated on a Millipore filter (10 kDa cut-off), and absorbances at 280 nm and 450 nm were measured using a Nano-Drop 2000 (Thermo Fisher Scientific) UV-Vis spectrophotometer.
Results
Strategy and choice of amino acid residues to be mutated
The mutagenesis strategy aimed at creating an EF-Tu construct having one cysteine residue as close as possible to three or four nucleotide bases in tRNA that can be readily labeled with fluorescent dyes (positions 8, 17/20 and 47), and much closer than the typical Förster distance for many FRET pairs (∼50 ± 10 Å). Additional constraints were: (i) a preferred surface exposure of the amino acid residues to be mutated; and (ii) no significant involvement of the residue in the binding to tRNA or to other molecules participating in ribosomal translation. We hypothesized that the region around the EF-Tu C-terminus should tolerate mutations, since large protein fragments (270 aa) can be fused to the C-terminus without a significant effect on the activity of EF-Tu (Schnell et al., 2003). We were also aware that a significant size nanoparticle, undecagold (Mr of 6200 Da, and particle diameter of 8 Å) can be conjugated to base 47 of tRNA (Blechschmidt et al., 1994); thus, placing a cysteine residue in a location close to nucleotide 47 in tRNA was deemed especially significant. Because the complexity of the EF-Tu-tRNA system and its interactions with other components of the translational machinery adds an element of uncertainty to simple structural analysis, we constructed and functionally screened 10 EF-TuSAV mutants to increase the likelihood of finding a mutant that was suitable for use in a FRET assay. The mutated residues are located in the turn regions between β-strands a3 and b3 (Phe 323, Lys 324, Gly 325, Tyr 326), or between β-strands c3 and d3 (Pro 344, Glu 345, Gly 346, Val 347, Glu 348) (Fig. 1). In addition, we created a mutant having a new C-terminal residue, C394 (the native S393 residue was retained). All of the introduced Cys residues were designed to lie on the surface of domain 3 of EF-Tu, in positions expected not to be involved in the function of EF-Tu. The only exceptions are residues Phe 323 and Glu 348, which interact with EF-Ts (Kawashima et al., 1996).
Positions of three of the mutated residues in the EF-Tu·GTP·aa-tRNA complex are highlighted in Fig. 1.
Purification of EF-Tu
Expression and purification of wt and mutated EF-Tus are summarized in Table I. The yield of purified protein was on the order of 0.01–0.4 mg/g and 0.8 mg/g wet weight for EF-Tu expressed in the GST and His-tag systems, respectively. The former yield refers to EF-Tu mutants released from the GST-tag, while no attempts were made to remove the His-tag from the latter preparation. Bacterial growth conditions to express the EF-Tu mutants had to be modified to minimize formation of inclusion bodies. The solubility of mutated proteins was improved by including sorbitol and betaine, which are known to induce osmotic stress in growing bacteria, and by lowering the temperature of incubation after induction with IPTG. The protein yield of EF-TuSAVE348C expressed under the modified conditions was three times higher than when expressed in standard medium.
Table I.
Summary of EF-Tu isolation from 0.5 l E.coli culture
| EF-Tu species | Fusion form | Fusion protein, mg | Recovered amount of EF-Tu, mg | Purity, % |
|---|---|---|---|---|
| EF-Tu wt | GST | 2.6 | 1 | 87 |
| EF-Tu wta | His tag | N.A. | 7 | 79 |
| EF-TuSAV | GST | 1.5 | 0.7 | 68 |
| EF-TuSAV F323C | GST | 0.9 | 0.2 | 45 |
| EF-TuSAV K324C | GST | 2.2 | 0.9 | 79 |
| EF-TuSAV G325C | GST | 1.7 | 0.4 | 90 |
| EF-TuSAV Y326C | GST | 0.6 | 0.1 | 55 |
| EF-TuSAV P344C | GST | 0.5 | 0.094 | 36 |
| EF-TuSAV E345C | GST | 0.3 | 0.044 | 20 |
| EF-TuSAV G346C | GST | 0.16 | 0.04 | – |
| EF-TuSAV V347C | GST | 0.23 | 0.05 | 17 |
| EF-TuSAV E348C | GST | 1.7 | 0.5 | 84 |
| EF-TuSAV C393ins | GST | 0.29 | 0.06 | 17 |
Percent purity is calculated for EF-Tu, not the fusion protein. N.A., not applicable.
aStaphylococcus aureus EF-Tu.
The EF-Tu proteins, prepared from 0.5 l of culture, fall into three groups with respect to yield and purity (Table I). Group 1, including proteins EF-Tu wt, EF-TuSAV, EF-TuSAVK324C, EF-TuSAVG325C and EF-TuSAVE348C, were obtained in higher yield (1.5–2.6 mg GST-fusion protein; and 0.4–1 mg EF-Tu) and higher purity (68–90% as determined by SDS–PAGE analysis). Group 2, including proteins EF-TuSAVF323C, EF-TuSAVY326C, were obtained in intermediate yield (0.5–0.9 mg fusion protein; 0.1–0.2 mg EF-Tu) and purity (36–55%). The third group consisted of four mutants, EF-TuSAVE345C, EF-TuSAVG346C, EF-TuSAVV347C and EF-TuSAVC393ins, which were obtained in low yield and purity.
Functional analysis
The functional analysis of 10 EF-TuSAV mutants as well as the wild-type and EF-TuSAV proteins was performed using four assays: GDP binding, inhibition of aminoacyl-tRNA deacylation, aminoacyl-tRNA-induced gel shift and poly(U)-directed poly(Phe) synthesis. Additionally, for the most active forms of EF-Tu, properties of their GST-fused counterparts were assayed. The results are summarized in Table II.
Table II.
Functional assays of EF-Tu mutants
| EF-Tu species | Fusion form | Deacylation rate constant, k−1×105, s−1 |
Relative GDP binding | Relative poly(U)-dependent poly(Phe) synthesis |
||
|---|---|---|---|---|---|---|
| EF-Tu, 2.7 µM | EF-Tu, 0.5 µM | EF-Tu, 10 pmol | EF-Tu, 1 pmol | |||
| None | 20.4 ± 2.1 | |||||
| EF-Tu wta | His tag | 1 | 2.6 ± 0.3 | |||
| EF-Tu wt | 1–393 | 2 ± 1.2 | 1.0 | 1.0 | 1.0 | |
| EF-TuSAV | 1–393 | 7.9 | 0.8 | 1.7 | ||
| EF-TuSAVF323C | 1–393 | 17.1 | 0.8 | 1.4 | ||
| EF-TuSAVK324C | 1–393 | 9.5 ± 2.5 | 0.9 ± 0.1 | 1.5 | ||
| EF-TuSAVG325C | 1–393 | 22.3 ± 0.6 | 0.3 ± 0.1 | 1.2 | ||
| EF-TuSAVY326C | 1–393 | 24.5 | 1.1 | 0.5 | ||
| EF-TuSAVP344C | 1–393 | 0.8 | ||||
| EF-TuSAVE345C | 1–393 | 0.2 | ||||
| EF-TuSAVV347C | 1–393 | 23.5 | 0.1 | 0.2 | ||
| EF-TuSAVE348C | 1–393 | 18.3 | 0.6 | 1.9 | ||
| EF-TuSAVC393ins | 1–393 | 0.05 | 0.5 | |||
| EF-Tu wt | GST | 0.9 ± 0.3 | 1.3 ± 0.3b | |||
| EF-TuSAVK324C | GST | 4.5 | 0.9 ± 0.5b | |||
| EF-TuSAVG325C | GST | 21.4 ± 3.3 | 0.1b | |||
| EF-TuSAVE348C | GST | 10.1 ± 1.3 | 0.2b | |||
The results for GST-EF-Tu fusions are in the last four rows in the table. Experimental details are presented in Materials and methods. EF-Tu amounts were calculated taking into account percent purities in Table I. Standard deviations (SD) are given for n = 2–5 repetitions. All data presented without the numerical value for SD are only single determinations. Background was subtracted from all poly(Phe) assay data. Blank cells indicate that data were not obtained. 1–393 denotes the native EF-Tu with no peptides fused prepared by Factor Xa cleavage.
aStaphylococcus aureus EF-Tu.
bIndicates that the translation reaction was conducted for 1–2 min at 32°C.
GDP binding
Under saturating conditions, GDP forms an equimolar complex with an intact EF-Tu, which can be captured in a filter-binding assay. The GDP-binding activities of EF-TuSAV, EF-TuSAVF323C, EF-TuSAVK324C and EF-TuSAVY326C were found to be similar to that of wild-type EF-Tu (Table II). Two other mutants, EF-TuSAVG325C and EF-TuSAVE348C, showed reduced GDP binding, while such activity was negligible for the EF-TuSAVV347C and EF-TuSAVC393ins mutants.
Non-enzymatic hydrolysis protection assay
Ternary complex formation can be detected by monitoring EF-Tu-dependent inhibition of [14C]Phe-tRNAPhe deacylation (Pingoud et al., 1977), which we measured for both EF-Tu mutants and for wt His-tagged EF-Tu, giving the results summarized in Table II. The wild-type forms of EF-Tu, as well as their fusions to GST, exhibited the highest protection against hydrolysis of Phe-tRNAPhe. The protection was somewhat reduced for the EF-TuSAV, EF-TuSAVK324C and for the GST fusions of EF-TuSAVK324C and EF-TuSAVE348C mutants. Mutants EF-TuSAVF323C, EF-TuSAVG325C, EF-TuSAVY326C, EF-TuSAVV347C, EF-TuSAVC393ins, as well as fusion protein GST-EF-TuSAVG325C, did not protect against deacylation of Phe-tRNAPhe.
Gel shift assay
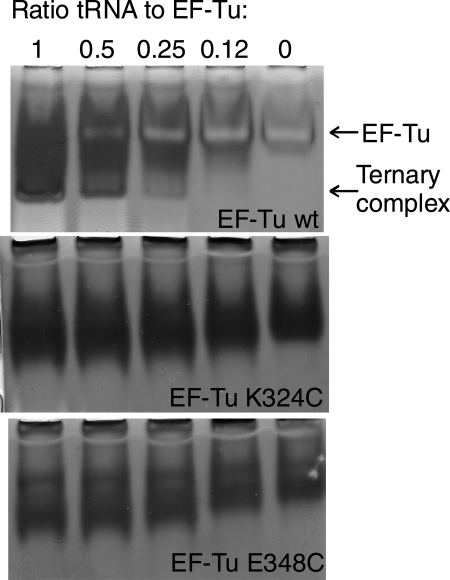
The activities of EF-Tu mutants in forming ternary complex with Phe-tRNAPhe were also monitored by non-denaturing PAGE analysis. Phe-tRNAPhe was titrated from 0.1 to 1.6 µM while keeping the EF-Tu concentration constant at 0.2 or 0.4 µM. For wt EF-Tus (both cleaved from GST and with the His6 tag), a clear band of ternary complex was seen even at low tRNA concentrations (Fig. 2). All wild-type EF-Tu versions participated in ternary complex formation at equimolar concentrations of charged tRNA. The EF-TuSAV mutant showed only a slightly reduced affinity for aa-tRNA compared with wild-type. The affinities of other mutants were reduced more significantly. The EF-TuSAVF323C, EF-TuSAVK324C and EF-TuSAVE348C mutants did form a ternary complex with aa-tRNA, but the relevant bands on the gel only became clear at higher tRNA concentrations. This suggests that the ternary complex had been formed, but was probably unstable under the electrophoresis conditions. The gel shift assay did not show any ternary complex formation by the EF-TuSAVG325C, EF-TuSAVY326C, EF-TuSAVV347C and EF-TuSAVC393ins mutants, even when a 4-fold excess of Phe-tRNAPhe over EF-Tu was used. Control reactions with uncharged tRNA showed no ternary complex formation.
Fig. 2.
Native PAGE of the ternary complex formed by wt EF-Tu with a His6 tag, EF-TuSAVK324C and EF-TuSAVE348C. The molar ratio of Phe-tRNAPhe to EF-Tu is given above each lane. The upper band is EF-Tu·GTP, while the lower band is the ternary complex, EF-Tu·GTP·Phe-tRNAPhe.
Poly(Phe) synthesis
Mutant EF-Tus were tested for their abilities to support poly(U)-dependent poly(Phe) synthesis (Table II). The extent of Phe incorporation into a polypeptide was similar for all tested EF-Tus, except for the EF-TuSAVY326C, EF-TuSAVE345C, EF-TuSAVV347C and EF-TuSAVC393ins mutants, which showed reduced activity. GST-fused EF-Tu wt and EF-TuSAVK324C were fully active, while EF-TuSAVE348C had reduced activity. Mutant EF-TuSAVG325C was the least active among the tested GST-fusions (Table II).
Labeling of EF-Tu with quencher
The reactivities of the introduced Cys residues were tested by labeling of EF-Tu with the quencher DABCYL Plus. Both the native and GST-fused wild-type proteins, as well as three selected EF-Tu mutants (K324C, G325C and E348C) were labeled. The ratio EF-TuSAV:DABCYL Plus was 7:1. Among the native EF-Tus, the highest labeling efficiency was 1.5 for wt EF-Tu (i.e. the number of Cys residues labeled per one EF-Tu molecule). The mutated EF-Tus were labeled to a smaller extent: the labeling level was 0.40, 0.32 and 0.37 for the K324C, G325C and E348C mutants, respectively. It should be noted, however, that the mutants carry only one Cys residue (compared with three cysteine residues in the wild-type EF-Tu). In a control mock labeling experiment with the native EF-TuSAV mutant, the efficiency of labeling due to side reactions was 0.14.
The fusion proteins were labeled to a higher extent, with more than one molecule of DABCYL Plus per protein. The average (n = 3) labeling levels were as follows: 1.9 (SD 0.3), 1.7 (SD 0.8), 1.4 (SD 0.3) and 2.1 (SD 0.7) for GST-EF-Tu wt, K324C, G325C and E348C, respectively. It should be noted that GST has two cysteine residues, which could have been labeled, leading to a higher labeling efficiency. We did not determine which of the five or three Cys residues present in GST-tagged EF-Tu wt or mutant, respectively, were labeled.
Discussion
The major goal of this study was to express, purify and evaluate the properties of E.coli EF-Tu mutants containing one Cys residue introduced at 10 various positions located within domain 3 of the factor (Fig. 1). Analysis of functionally important regions indicates that most of the residues involved in EF-Tu activities are situated within domain 1, though some residues from domains 2 and 3 are also involved in interactions with EF-Ts and in binding aa-tRNAs.
Among residues subjected to mutagenesis in the current study, only residue 348 of E.coli EF-Tu has been previously analyzed by mutagenesis. Zhang et al. (1998) mutated Glu 348 to Ala, and demonstrated that the E348A mutant had unchanged GDP binding activity and a higher binding constant for EF-Ts.
To evaluate conservation of mutated residues among species, we analyzed alignment of EF-Tu domain 3 amino acid sequences from several prokaryotes and eukaryotes to the E.coli sequence. The alignment, originally generated by the BLAST server (Altschul et al., 1997), is shown for both mutated regions (Phe 323–Tyr 326 and Pro 344–Glu 348) in Fig. 3. Sequences within the two regions are quite variable. The most conserved residues are Pro 344 and Tyr 326, and also, to a degree, Glu 348. This relatively high variability suggests that mutagenesis of most of the nine residues in the region should not have a pronounced effect on EF-Tu function.
Fig. 3.
Sequence alignment of the domain 3 fragment from E.coli EF-Tu subjected to labeling with the corresponding fragments from selected prokaryotes and eukaryotes (mitochondrial EF-Tus). The computational analysis was performed using BLAST network service (Altschul et al., 1997). The amino acid residues are numbered as in E.coli EF-Tu. The residues mutated in this study are in the 323–326 and 344–348 regions, as indicated in the figure.
The first difference between analyzed EF-Tu mutants was at the level of expression and purity after isolation. Only EF-Tu wt, EF-TuSAV, EF-TuSAVK324C, EF-TuSAVG325C and EF-TuSAVE348C could be expressed and purified at fairly high levels, while all other forms of EF-Tu gave much lower expression levels and very low purity of the final preparation (Table I). This may be due to lower solubility and/or stability of the mutants.
Functional analysis of altered EF-Tu was performed to evaluate the influence of the introduced mutations on guanine-nucleotide binding, ternary-complex formation and activity in translation in vitro (Table II). The results obtained for EF-Tu wt and EF-TuSAV confirmed earlier observations (C.R.K., unpublished results), and demonstrated that mutating the three native Cys residues to Ser, Val and Ala (as in the EF-TuSAV mutant) has no or only slight apparent effect on the GDP-binding assay, translation in vitro and gel shift assay, though a significant, 8-fold effect was seen in the deacylation assay. Much bigger changes were observed after introducing an additional Cys residue into the tested positions within domain 3. This resulted either in problems with expression (EF-TuSAVF323C, EF-TuSAVY326C), lower activity (EF-TuSAVK324C, EF-TuSAVG325C, EF-TuSAVE348C) or both (EF-TuSAVP344C, EF-TuSAVE345C, EF-TuSAVG346C, EF-TuSAVV347C, EF-TuSAVC393ins). Most of the mutants had unchanged (EF-TuSAVF323C, EF-TuSAVK324C, EF-TuSAVY326C) or slightly reduced (EF-TuSAVG325C, EF-TuSAVE348C) ability to bind GDP when compared with the wt EF-Tu. Cys residues introduced at position 347 and at the C-terminus resulted in almost complete loss of GDP binding activity. The loss of activity of the EF-TuSAVC393ins mutant was surprising as fusions to the C-terminus typically do not change EF-Tu's activity (e.g. fusion to a His tag; Boon et al., 1992). It is conceivable that folding of the mutant EF-Tu was affected due to the presence of the GST protein in the expression product.
Residues involved in ternary complex interactions with aa-tRNA or GTP are dispersed in all three domains of EF-Tu [reviewed in (Clark and Nyborg, 1997)]. We employed two methods to evaluate the influence of Cys mutations on aa-tRNA binding, namely the protection assay and the gel shift assay. Both assays gave generally complementary results. Native EF-Tu wt was shown to bind Phe-tRNAPhe well in both assays, while the GST-EF-Tu fusion showed good binding in the protection assay. EF-TuSAV, EF-TuSAV K324C, GST- EF-TuSAVK324C and GST- EF-TuSAVE348C showed partial losses of activity. Other mutants were found to be inactive in the protection assay. The lower activity of EF-TuSAV in protecting aa-tRNA against spontaneous deacylation is in agreement with a similar observation for a Cys 81 mutant (Anborgh et al., 1992).
Other functional tests gave more diverse results. In a number of cases, the tRNA binding activity of EF-Tu was seemingly not correlated to its translation activity in vitro and the ability to bind GDP. For example, mutants EF-TuSAVG325C and EF-TuSAVY326C are active in GDP binding assay and translation in vitro, while they are unable in a binding assay to stably bind to Phe-tRNAPhe. F323C, G325C and Y326C mutations appear not to protect aa-tRNA from hydrolysis, but function relatively well in protein synthesis.
Interestingly, Andersen and Wiborg (1994) also observed a lower protection level of the aminoacyl bond against non-enzymatic hydrolysis of mutants H66A and H118A, while they could not identify any significant differences between wt and the mutants in poly(Phe) synthesis. The authors suggested that the activity of the ribosomes was a limiting factor in their experiments. It is also likely that in the present experiments, the limiting factor in translation is an unidentified step different from the binding activity of EF-Tu to tRNA or of the ternary complex to the ribosomes.
Facing conflicting results, choosing the best mutants for the FRET experiments was challenging. We had to take into account the results of the three assays (GDP binding, protection assay and translation in vitro), and in addition, make sure that sufficient amount of protein can be readily made (expression level). The choice of three mutations for further studies was based on the functional criteria, protein synthesis activity and good expression level. Thus, we chose K324C, G325C and E348 mutants.
The ability to conjugate a dye to the thiols of the three mutants was experimentally verified. Though the labeling efficiency was not very high and ranged from 0.3 to 0.4 DABCYL Plus dye molecules per one EF-Tu molecule (see Results), further enrichment is possible through chromatography approaches.
Seven of the mutants (EF-TuSAVG325C, EF-TuSAVY326C, EF-TuSAVP344C, EF-TuSAVE345C, EF-TuSAVG346C, EF-TuSAVV347C, EF-TuSAVC393ins) were completely defective in Phe-tRNAPhe binding under the conditions tested. This may be due to a significant structural change in the tRNA binding cleft rather than to the loss of a contact crucial for aa-tRNA binding, because none of the listed positions has been so far reported to interact directly with aa-tRNA. This is in agreement with the poor yields obtained for most of the problematic mutants, which may indicate a more general destabilization affecting folding.
We evaluated some of the properties of four GST-tagged EF-Tus due to recovery problems after removing the GST-tag. The GST-fusions of EF-Tu wt and the EF-TuSAVK324C mutant were as effective as the full-length EF-Tu counterparts in protecting aa-tRNA against non-enzymatic deacylation and in translation in vitro. Further, the GST fusion to the EF-TuSAVE348C mutant performed better in the protection assay than the full-length EF-Tu, but had a reduced activity in translation in vitro (Table II). These results indicate that in some cases, particularly when cleavage by factor Xa gives unsatisfactory results, GST-fused EF-Tu could be used in place of EF-Tu wt. Here it is pertinent that cryo-electron microscopy results indicate that EF-Tu within the ternary complex contacts the ribosome through the P loop, Arg 58 and His 84 (Li et al., 2008; Villa et al., 2009), which are located on the opposite side of domain 1 from the N-terminus residue to which the GST moiety is attached.
Positions of the mutated regions were analyzed with regard to their proximity to the 30S subunit and the AT-site tRNA. We used a recently published cryo-electron microscopy structure of the 70S Thermus thermophilus ribosome in which the ternary complex was trapped on the ribosome using kiromycin (Schuette et al., 2009; PDB entries 3FIC and 3FIN), and analyzed the positions of the T.thermophilus EF-Tu residues which were homologous to the mutated residues of the E.coli EF-Tu. The two mutated loops were more than 22 Å away (Cα to Cα) from the nearest ribosomal structure, protein S12 of the 30S subunit and more than 18 Å away from the nearest structure of the 50S subunit, protein L11. The C-terminus of EF-Tu was approximately 10 Å away from protein S12, and 40 Å from protein L11. The positions of the two loops and the C-terminal residue with respect to the AT-site tRNA in the cryo structure was analogous to that found in the structure examined in detail in this paper (1OB2, see Fig. 1). The analysis indicates that no steric hindrance is expected between the labeled forms of EF-Tu and tRNA, which is a key to the success of the series of experiments envisioned in this project, i.e. acquiring of fluorescent signal from a single translating ribosome. The most favorable experimental conditions are expected in a situation where a uridine residue at position 47 of tRNA is labeled with a fluorescent dye, and one of the amino acid residues in loop 323–326 of EF-Tu is labeled with a quencher. The distance between the two sites is in the range of 8–12 Å, potentially suitable for highly efficient contact quenching.
In summary, this study provides detailed information about properties of several E.coli EF-Tu mutants from the perspective of configuring a FRET assay in which the fluorescence of a labeled tRNA molecule is quenched by a quencher-labeled EF-Tu. We have identified three mutations, EF-TuSAVK324C, EF-TuSAVG325C and EF-TuSAVE348C, which will be used in further studies aimed at detailed quantification of energy transfer (quenching) within the ternary complex, leading to a method to collect nucleic acid sequence information from translating ribosomes at a single molecule level.
Funding
This work was supported by a grant from the National Institutes of Health (HG004364 to W.M.).
Acknowledgements
We thank Dr Yale E. Goldman for help in the early phases of the project and for useful discussions. We also thank Dr M. Ibba for providing an E.coli strain expressing the phenylalanyl-tRNA synthetase and Dr Mandana Sassanfar for the S.aureus EF-Tu-expressing plasmid. Karen Margrethe Nielsen is acknowledged for skilled technical assistance.
Conflict of interest: B.S.C. is a consultant to Anima Cell Metrology and owns shares in the company.
Footnotes
Edited by Dieter Soll
References
- Altschul S.F., Madden T.L., Schaffer A.A., Zhang J., Zhang Z., Miller W., Lipman D.J. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anborgh P.H., Parmeggiani A., Jonak J. Eur. J. Biochem. FEBS. 1992;208:251–257. doi: 10.1111/j.1432-1033.1992.tb17180.x. [DOI] [PubMed] [Google Scholar]
- Andersen C., Wiborg O. Eur. J. Biochem. FEBS. 1994;220:739–744. doi: 10.1111/j.1432-1033.1994.tb18674.x. [DOI] [PubMed] [Google Scholar]
- Blechschmidt B., Shirokov V., Sprinzl M. Eur. J. Biochem. FEBS. 1994;219:65–71. doi: 10.1111/j.1432-1033.1994.tb19915.x. [DOI] [PubMed] [Google Scholar]
- Boon K., Vijgenboom E., Madsen L.V., Talens A., Kraal B., Bosch L. Eur. J. Biochem. FEBS. 1992;210:177–183. doi: 10.1111/j.1432-1033.1992.tb17406.x. [DOI] [PubMed] [Google Scholar]
- Bradford M.M. Anal. Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Clark B.F., Nyborg J. Curr. Opin. Struct. Biol. 1997;7:110–116. doi: 10.1016/s0959-440x(97)80014-0. [DOI] [PubMed] [Google Scholar]
- Doi Y., Ohtsuki T., Shimizu Y., Ueda T., Sisido M. J. Am. Chem. Soc. 2007;129:14458–14462. doi: 10.1021/ja075557u. [DOI] [PubMed] [Google Scholar]
- Kawashima T., Berthet-Colominas C., Wulff M., Cusack S., Leberman R. Nature. 1996;379:511–518. doi: 10.1038/379511a0. [DOI] [PubMed] [Google Scholar]
- Knudsen C.R., Clark B.F., Degn B., Wiborg O. Biochem. Int. 1992;28:353–362. [PubMed] [Google Scholar]
- Li W., Agirrezabala X., Lei J., Bouakaz L., Brunelle J.L., Ortiz-Meoz R.F., Green R., Sanyal S., Ehrenberg M., Frank J. EMBO J. 2008;27:3322–3331. doi: 10.1038/emboj.2008.243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandecki W., et al. Proc. SPIE. 2008;6862:68620T. [Google Scholar]
- Miller D.L., Weissbach H. Methods Enzymol. 1974;30:219–232. doi: 10.1016/0076-6879(74)30024-9. [DOI] [PubMed] [Google Scholar]
- Pingoud A., Urbanke C. Anal. Biochem. 1979;92:123–127. doi: 10.1016/0003-2697(79)90632-8. [DOI] [PubMed] [Google Scholar]
- Pingoud A., Urbanke C., Krauss G., Peters F., Maass G. Eur. J. Biochem. FEBS. 1977;78:403–409. doi: 10.1111/j.1432-1033.1977.tb11752.x. [DOI] [PubMed] [Google Scholar]
- Schnell R., Abdulkarim F., Kalman M., Isaksson L.A. FEBS Lett. 2003;538:139–144. doi: 10.1016/s0014-5793(03)00159-5. [DOI] [PubMed] [Google Scholar]
- Song H., Parsons M.R., Rowsell S., Leonard G., Phillips S.E. J. Mol. Biol. 1999;285:1245–1256. doi: 10.1006/jmbi.1998.2387. [DOI] [PubMed] [Google Scholar]
- Schuette J.C., et al. EMBO J. 2009;28:755–765. doi: 10.1038/emboj.2009.26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tseng W.C., Lin J.W., Wei T.Y., Fang T.Y. Anal. Biochem. 2008;375:376–378. doi: 10.1016/j.ab.2007.12.013. [DOI] [PubMed] [Google Scholar]
- Villa E., et al. Proc. Natl Acad. Sci. USA. 2009;106:1063–1068. doi: 10.1073/pnas.0811370106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang W., Malcolm B.A. Biotechniques. 1999;26:680–682. doi: 10.2144/99264st03. [DOI] [PubMed] [Google Scholar]
- Zhang Y., Yu N.J., Spremulli L.L. J. Biol. Chem. 1998;273:4556–4562. doi: 10.1074/jbc.273.8.4556. [DOI] [PubMed] [Google Scholar]