Abstract
IR spectroscopy is an excellent method for biological analyses. It enables the nonperturbative, label-free extraction of biochemical information and images toward diagnosis and the assessment of cell functionality. Although not strictly microscopy in the conventional sense, it allows the construction of images of tissue or cell architecture by the passing of spectral data through a variety of computational algorithms. Because such images are constructed from fingerprint spectra, the notion is that they can be an objective reflection of the underlying health status of the analyzed sample. One of the major difficulties in the field has been determining a consensus on spectral pre-processing and data analysis. This manuscript brings together as coauthors some of the leaders in this field to allow the standardization of methods and procedures for adapting a multistage approach to a methodology that can be applied to a variety of cell biological questions or used within a clinical setting for disease screening or diagnosis. We describe a protocol for collecting IR spectra and images from biological samples (e.g., fixed cytology and tissue sections, live cells or biofluids) that assesses the instrumental options available, appropriate sample preparation, different sampling modes as well as important advances in spectral data acquisition. After acquisition, data processing consists of a sequence of steps including quality control, spectral pre-processing, feature extraction and classification of the supervised or unsupervised type. A typical experiment can be completed and analyzed within hours. Example results are presented on the use of IR spectra combined with multivariate data processing.
INTRODUCTION
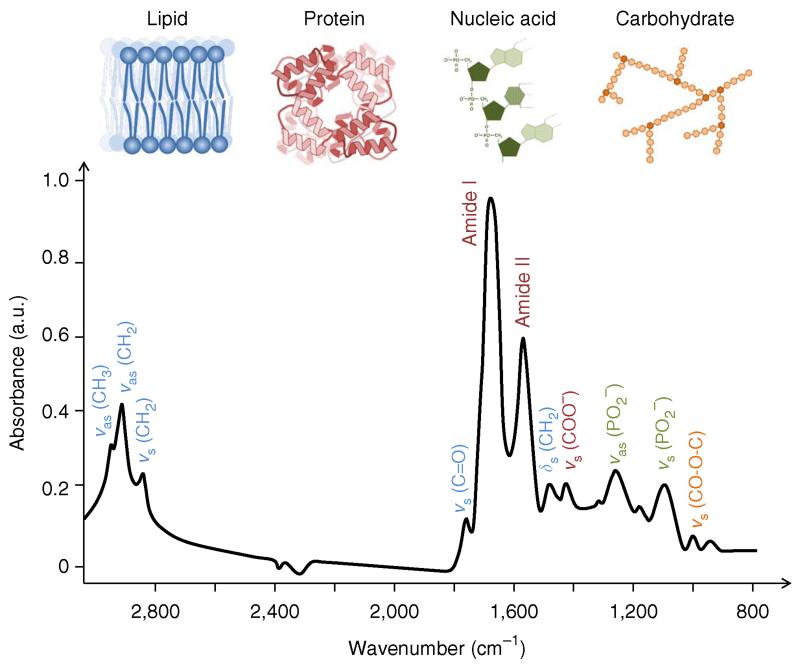
The use of Fourier transform IR (FTIR) spectroscopic techniques for the nondestructive analysis of biological specimens is a rapidly expanding research area, with much focus on its utility in cytological and histological diagnosis through the generation of spectral images1,2. Molecular bonds with an electric dipole moment that can change by atomic displacement owing to natural vibrations are IR active. These vibrational modes are quantitatively measurable by IR spectroscopy3, providing a unique, label-free tool for studying molecular composition and dynamics without perturbing the sample. For interrogating biological materials, the most important spectral regions measured are typically the fingerprint region (600–1,450 cm−1) and the amide I and amide II (amide I/II) region (1,500–1,700 cm−1). The higher-wavenumber region (2,550–3,500 cm−1) is associated with stretching vibrations such as S-H, C-H, N-H and O-H, whereas the lower-wavenumber regions typically correspond to bending and carbon skeleton fingerprint vibrations4. Together, these regions comprise a biochemical fingerprint of the structure and function of interrogated cellular specimens. A typical biological IR spectrum with molecular assignments is shown in Figure 1.
Figure 1.
Typical biological spectrum showing biomolecular peak assignments from 3,000–800 cm−1, where ν = stretching vibrations, δ = bending vibrations, s = symmetric vibrations and as = asymmetric vibrations. The spectrum is a transmission-type micro-spectrum from a human breast carcinoma (ductal carcinoma in situ). The sample was cryosectioned (8 μm thick) and mounted on BaF2 slides (1 mm thick) before IR microspectroscopy. Equipment: Bruker IR scope II, circular diameter of aperture ~60 μm; a.u., arbitrary units.
IR microspectroscopy
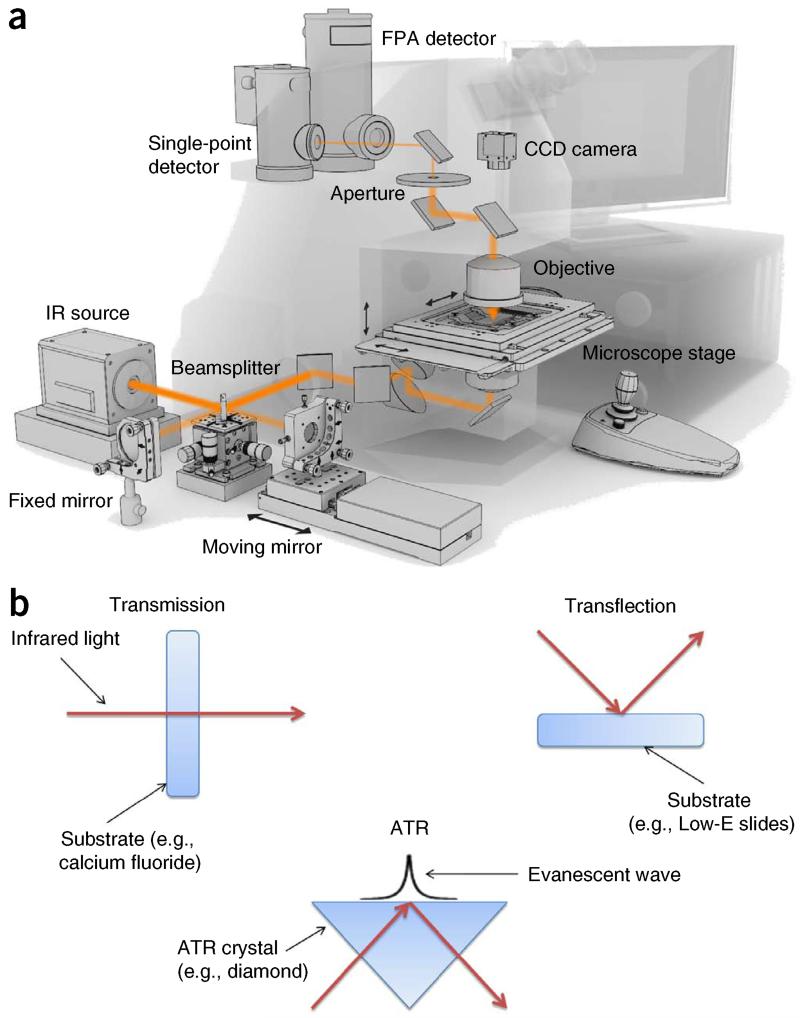
Although the spectral domain allows chemical identification, the combination with microscopy (microspectroscopy) permits the examination of complex tissues and heterogeneous samples5. Detection by microscopy (see schematic of instrumentation in Fig. 2) may be accomplished by raster-scanning a point illuminated on the sample or by using wide-field illumination and focal plane array (FPA) or linear array detectors6. At present, wide-field scanning of a sample is possible in seconds, providing tens of thousands of spectra. A variety of choices are available for the IR source, including globar7, synchrotron8-12 and quantum-cascade lasers (QCLs)13, as well as for the detector (2D FPA, linear array or single element)14. The three major IR-spectroscopic sampling modes (Fig. 2b) are transmission, transflection and attenuated total reflection (ATR). Each mode offers convenience for some samples and challenges for others. In transflection mode, for illustration, the sample is placed on an inexpensive IR-reflecting surface (such as that found on low-emissivity (Low-E) slides) and measurements are generated by a beam passing through the sample and reflecting back from the substrate (i.e., the reflective surface) through the sample. As is clear from both theoretical and experimental studies15,16, the recorded spectral intensities depend on both sample morphology and chemistry. Hence, care should be taken on substrate choice17,18. Recently, topographical features of the sample and its effects have been shown to be minimized by inputting second derivative spectra in the classification model; better segregation of normal versus various disease categories facilitates potential spectral histopathological diagnosis19. Research by Cao et al.20 has demonstrated that if this pre-processing data analysis approach is performed (e.g., after both transflection and transmission measurements on dried cellular monolayers), the resulting classification is the same. This example suggests that irrespective of sampling geometry, mathematical tools can be applied to minimize confounding effects and to interpret their influence. As such, spectral processing may determine the diagnostic efficacy of spectral processing, not only from a biological perspective but also from the ability to control optical or distorting influences.
Figure 2.
The instrumentation underlying the main forms of IR spectroscopic sampling. (a) Schematic of modern FTIR-imaging spectrometer. Reproduced with permission from ref. 6. (b) Schematic representation of the three main sampling modes for FTIR spectroscopy. Reprinted from Trends Biotechnol, 31, Dorling, K.M. and M.J. Baker, Highlighting attenuated total reflection Fourier transform infrared spectroscopy for rapid serum analysis, 327–328, Copyright 2013 with permission from Elsevier (ref. 132).
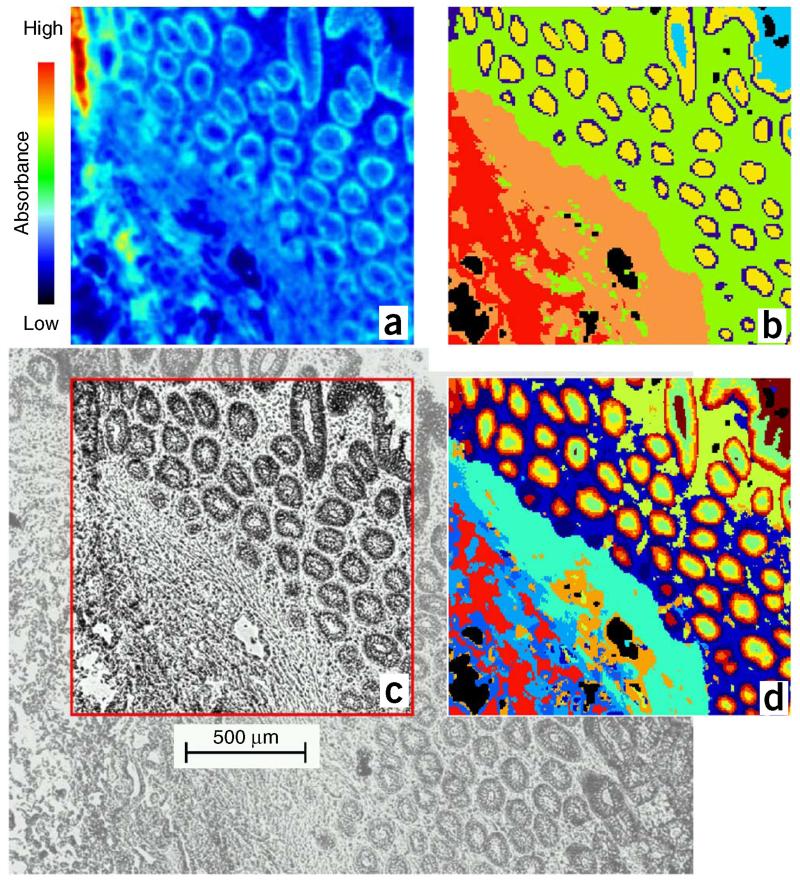
FTIR imaging provides spatially resolved information based on chemically specific IR spectra in the form of an information-rich image of the tissue or cell type being interrogated21-23. Further multivariate data analysis allows potential diagnostic markers to be elucidated, thus providing a fast and label-free technology to be used alongside conventional techniques such as histology2,22. At present24, rapid imaging permits imaging in hours for a whole-organ cross-section, such as that from the prostate; this not only allows one to objectively visualize pathology in situ but the aforementioned classification models could also allow one to grade disease on the basis of the cateogries into which spectra might be aligned. One excellent interpretation application of IR imaging data is to consider it as a metabolomic tool that allows the in situ, nondestructive analysis of biological specimens, (e.g., determining the glycogen levels in cervical cytology)25.
Data can be recorded from a variety of samples, ranging from live cells to formalin-fixed, paraffin-embedded (FFPE) archival tissue typical of a pathology specimen. IR spectra representing distinguishing fingerprints of specific cell types (e.g., stem cells versus transit-amplifying cells versus terminally differentiated cells) within a defined tissue architecture (e.g., crypts of the gastrointestinal tract and cornea)9,26 are now easily recorded. Consequently, spectral analyses delineate cellular hierarchy on the basis of protein, lipid and carbohydrate composition and/or DNA conformational changes27. For biomedical analyses, the major goal today is to derive an image of tissue architecture expressing the underlying biochemistry in a label-free fashion28, a development that can considerably extend our diagnostic potential beyond present capabilities. For example, to distinguish cells committed toward a pathological process (e.g., transformation) that conventional methods (e.g., visual scoring) might identify as normal. The screening of cervical cytology specimens to distinguish normal versus low-grade versus high-grade cells4,29, to grade primary neoplasia30, or to determine whether tissue margins and potential metastatic sites are tumor free31,32 are examples of this concept across many types of tissues. It is this bridge from the technology and potential of IR spectroscopy and imaging to biological, mainly clinical, applications that is the subject of this protocol (Fig. 3).
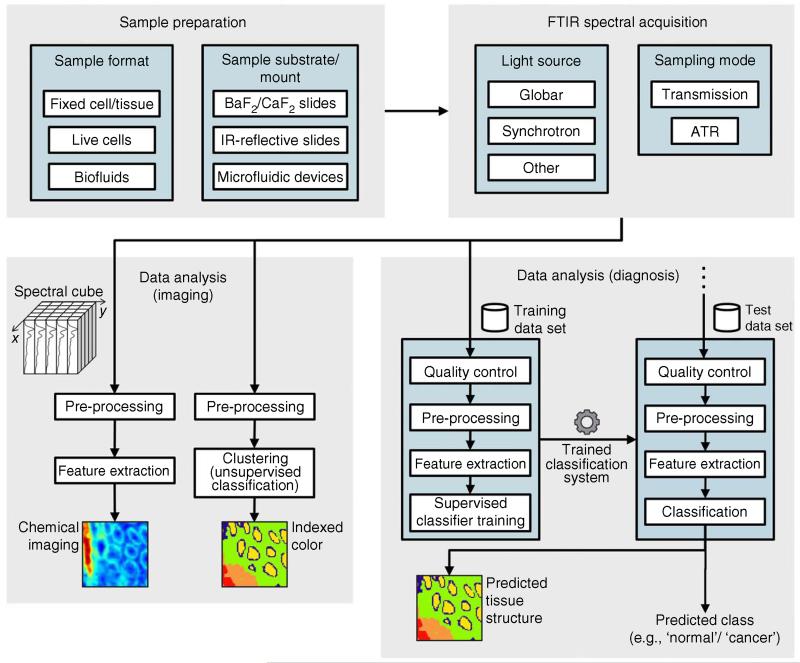
Figure 3.
FTIR spectroscopy work flow for imaging and diagnosis. The three major steps are sample preparation, FTIR spectral acquisition and data analysis. Sample preparation may differ depending on the sample format, requiring different materials and procedures. At FTIR spectral acquisition, several options have to be considered for light source and sampling mode. Data analysis presents different paths depending on the analysis goal (i.e., imaging or diagnosis). The framework for diagnosis is somewhat more complex, involving training of classification systems and validation of these systems using test data sets. Although not illustrated, the data sets used for testing are also obtained through sample preparation followed by FTIR spectral acquisition.
IR spectroscopy in cancer classification and imaging
By using IR spectroscopy either as an imaging tool or by classifying spectral categories, it has been possible to distinguish between benign and malignant tumors in tissue samples of breast32-35, colon22,23,36, lung37 and prostate8,30,38,39 along with cervical cytology or biopsies4,28,40. IR spectroscopic analysis is also an ideal tool for the study of biofluids such as urine, saliva, serum or whole blood; the use of biofluids is desirable in a clinical setting as samples are obtained rapidly and relatively noninvasively, and minimal sample preparation is required. By using such methods, a spectral fingerprint of the biofluid can be obtained, which allows the subsequent classification of spectra from different categories with computational methods and possibly the identification of biomarkers41-44.
FTIR imaging of tissue and cells
Imaging of live cells is possible using both globar and synchrotron-based light sources, with the latter permitting greater lateral spatial resolution and data quality owing to higher flux21,45-47. Diffraction-limited resolution with ATR-FTIR imaging can also be advantageous as it allows analysis of live cells in aqueous systems21,48. In addition, the spatial resolution of the image can be increased by incorporating optics with a high refractive index21,34.
We describe a protocol that has three components: (i) specimen preparation and removal of possible sample contaminants; (ii) acquisition of spectra with a sufficiently high signal-to-noise ratio (SNR); and (iii) data processing for classification and imaging. As the precise steps in acquisition of spectra and data processing are, respectively, dependent on the instrument and software available, this protocol covers (ii) and (iii) to deliver a general understanding of the steps involved. Supplementary Methods 1–4 correspond to four different examples of standard operating procedures (with troubleshooting) specific to common instruments and acquisition/analysis software. Together, this protocol and the material contained in Supplementary Methods 1–4 are designed to build researchers’ confidence in conducting their studies using their own instrumentation and computational settings.
Application of this protocol to other research areas
The application of this protocol is not limited to the biomedical field. IR spectroscopy has previously been used in the fields of environmental toxicology49-52, consumer safety53,54, taxonomy55-57, and in the food industry58; a non-instrument– and non-software–specific protocol for imaging and classification could be of considerable use to these areas of research.
Experimental design: instrumental options
The main steps required to analyze a sample of interest are sample preparation, instrumental setting, acquisition of spectra and data processing (Fig. 4). Before instrumental options are chosen, it is important for the user to understand the expectations from the intended experiment. These include the desired spectral and spatial resolution and type of study (e.g., diagnostic versus exploratory). In addition, proper consideration must be given to potential sample restrictions such as acquiring appropriate sample thickness for respective modes.
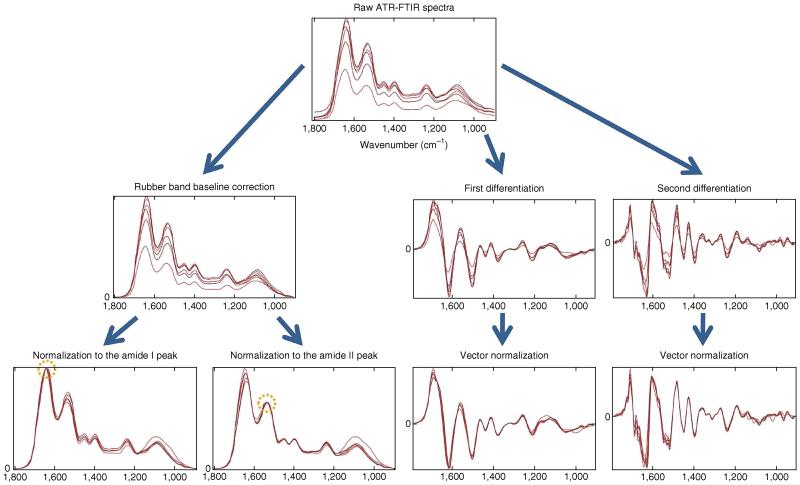
Figure 4.
Visual effect of different pre-processing steps on a set of FTIR spectra. Two common pre-processing sequences are rubber band baseline correction followed by normalization to the amide I/II peak and first or second differentiation followed by vector normalization. Rubber band baseline correction subtracts a rubber band, which is stretched ‘bottom-up’ at each spectrum, eliminating slopes. Amide I/II normalization forces all spectra to have the same absorbance intensity at the amide I/II peak. Differentiation (Savitzki-Golay (SG) method) has the advantage of eliminating slopes while also resolving overlapped bands, but has the drawback of altering the shape of the spectra (the y axis unit is no longer a.u. (arbitrary units), but ‘a.u. per wavenumber’ (first differentiation) or ‘a.u. per wavenumber squared’ (second differentiation)) and enhancing noise (note how second-differentiated spectra are visibly more noisy). Vector normalization is typically applied after differentiation. This normalization technique does not require a reference peak as amide I/II normalization does.
Sampling modes
Figure 2b shows a schematic representation of each sampling mode and details of each can be seen in Table 1; however, it is important to note that different manufacturer systems may vary slightly in some parameters, such as sampling apertures. Transmission and transflection sampling modes have been applied to a variety of biological specimens that can be sectioned into a thin layer allowing for accurate spectral data acquisition59. ATR-FTIR mode differs in that the IR beam is directed through an internal reflection element (IRE) with a high refractive index (e.g., diamond, zinc selenide, germanium or silicon)60. The evanescent wave extends beyond the IRE surface penetrating the sample, which must be in direct contact with the IRE. The penetration depth of this wave typically ranges from 1 to 2 μm within the 1,800–900 cm−1 region, but it should be remembered that there is still ~5% intensity at a depth of 3 μm (refs. 18,61,62). It has been shown that samples with thicknesses of <2 μm may give rise to spectral artifacts with IR-reflective substrates such as MirrIR Low-E slides (Kevley Technologies); therefore, when these substrates are used with ATR-FTIR spectroscopy, a thicker sample is recommended18.
Table 1.
FTIR spectroscopy modes used for the interrogation of cellular materials.
| Mode | Suitable samples |
Substrate | Typical interrogation area (μm) |
Pros | Cons |
|---|---|---|---|---|---|
| ATR | Tissues, cells and biofluids |
Calcium or barium fluoride, zinc selenide, MirrIR Low-E-coated glass |
250 × 250 | High SNR Reduced scattering Analysis of large target area Better for aqueous samples with appropriate substrate Highest spatial resolution (because of the refractive index n, which is 3.5 or even 4 in case of Si or Ge) |
Can be destructive because of pressure Air between sample and IRE will affect spectra Minimum sample thickness is required (~2.3 μm) Interactions of samples with the IRE leading to structural alterations (e.g., secondary protein structure) |
| Transmission | Tissues, individual cells, cellular components and biofluids |
Calcium or barium fluoride and zinc selenide |
5 × 5 to 150 × 150 |
High spatial resolution Nondestructive of prepared sample Automated stage allows for spec- tral acquisition at several different locations of choice with little user interaction |
Lower SNR than ATR Maximum sample thickness is required Sample thickness should be twice as large as for transflection to achieve the same absorbance Longer sample and machine preparation is required |
| Transflection | Tissues, individual cells, cellular components and biofluids |
Calcium or barium fluoride and zinc selenide |
5 × 5 to 150 × 150 |
High spatial resolution Nondestructive of prepared sample Automated stage allows for spec- tral acquisition at several different locations of choice with little user interaction Approximate sample thickness can be 1–4 mm, whereas for transmission it needs to be 2–8 μm |
May give rise to standing wave artifacts Lower SNR than ATR Maximum sample thickness is required Longer sample and machine preparation is required Scattering effects such as RMieSc will be much more intense in transflection type measurements |
A magnification-limited digital camera may be used for visualization in order to guide manual navigation across a given sample so as to locate a region of interest and help identify basic microscopic features such as separation between cancer cells and stromal elements. An alternative setup for ATR involves placing the sample directly onto the IRE aperture of the ATR accessory. This is particularly useful for biofluid analysis as it bypasses any potential contributions from any slide substrate that the sample could be placed on (Supplementary Method 1). This methodology may also help to reduce experimentation time owing to reduced sample preparation.
Light sources
In IR microspectroscopy, the user has the option of several light sources: a conventional thermal (globar) or synchrotron radiation source for FTIR interferometric measurements or alternative sources such as QCLs63 and filters64, which obviate the use of interferometers. The majority of benchtop instruments use conventional thermal light sources often in conjunction with single-element detectors. A globar source is composed of a silicon carbide rod that generates IR radiation, and can typically generate a collimated mean of ~1,000 μm in diameter, providing a uniformly illuminated aperture of 20–100 μm of the diameter at the sample65. It has been shown that single-cell investigations can be conducted using standard globar IR sources to derive subcellular information66.
A synchrotron radiation light source is ~100–1,000-fold brighter than current benchtop thermal ones, but it illuminates a much smaller area. Thus, a synchrotron source has a natural sampling aperture of 10–20 μm in diameter with a high SNR67. It is therefore possible to achieve single-cell and large organelle (e.g., nucleus) lateral spatial resolution with these modern sources, allowing subcellular molecular distribution analysis68,69. There are ~50 synchrotron facilities worldwide, all easily accessible for routine use as they operate on a call-for-projects basis70. Alternatively, other available sources that may be advantageous to individual studies include optic parametric oscillator (OPO) lasers, QCLs and free-electron lasers (FELs); traditionally they have been primarily used for gas sensing because of intrinsically narrow linewidths71,72; however, modern QCLs can cover much broader wavelength regions (hundreds of cm−1).
Mapping versus imaging
Broadly speaking, detectors can be separated into single-element, linear array and FPA detectors; the detector choice will be influenced by the requirement being imaging (i.e., FPA) or point spectra with high SNR (i.e., single element). The use of a single-element detector allows for individual point spectra to be obtained across a whole sample (for instance, useful when analyzing biofluids); a particular application has been to derive single-cell–specific fingerprint spectra across a heterogeneous tissue section. Acquiring large data sets containing point spectra is a method regularly used in biomedical and environmental studies coupled with multivariate data analysis40,73. Although time consuming, point spectra often have a high SNR, resulting in high-quality spectra, as spatial resolution is limited by IR apertures74. Maps can be generated when point spectra are collected in a stepwise manner in a grid from a target area, which is useful for comparing the different cell types from that particular area, e.g., gastrointestinal crypt23. Spectral maps take a much longer time than individual point spectra and, thus, in order to make large maps feasible to run, the acquisition time for each point can be reduced leading to a lower SNR. The absorbance intensity at each spectral point within the map becomes an individual pixel in the resultant pseudocolor images, which can give details of how different biomolecules vary across the target area.
In contrast to aperture-based systems, non-aperture-based instruments such as FPA and linear array detectors provide imaging using spatially arranged detectors. Multielement detectors allow for simultaneous spectral acquisition, which, combined with suitable optics, produce spectral images with good SNR and lateral spatial resolution close to the diffraction limit75. Measurements using an FPA detector (typically 32 × 32, 64 × 64 or 128 × 128) are rapid as such detectors allow for the acquisition of thousands of spectra simultaneously76; for a typical methodology see Supplementary Method 2. The acquired spectral data can be used to generate pseudocolor images of the target area such as shown in the characterization of prostate tissue77 and cervical biopsy samples28. The benefits of using a synchrotron radiation light source with FPAs also mean that much smaller pixel sizes can be used (e.g., 0.54 μm × 0.54 μm at some synchrotron facilities) resulting in higher spatial-resolution images of the target area76.
ATR-FTIR spectroscopy coupled to an array detector can allow for sample imaging down to diffraction-limited resolution for the spectral range of interest78. The spatial resolution of non-aperture-based techniques is determined by the optics chosen, and it has been shown that a germanium optic is preferential, although ZnSe and diamond crystals can also be used34. Although transmission and transflection imaging have been widely implemented in biological tissues, imaging in ATR mode is a versatile option, because little sample preparation is required owing to minimal sample-thickness restrictions, which thus means that it has been implemented in biological fields such as pharmacology and subcellular interrogation59,78,79.
Experimental design: sample preparation
Sample formats
The main sample formats for clinical IR spectroscopy are fixed cell and tissue samples, biofluids and live cells. Spectroscopic approaches can be used to examine tissues of human extraction (all require the appropriate ethical approval before their use). The type of sample used greatly determines which type of IR spectroscopy is appropriate and how it should be prepared for analysis. Table 2 shows the main types of samples and how they should be prepared for analysis.
Table 2.
Sample types and preparation.
| Sample type | Preparation | Removal of contaminants | Sample mount | Considerations |
|---|---|---|---|---|
| Biofluids | Biofluids such as blood, urine, saliva and synovial fluid should be collected as per hospital SOPs Samples that are not immediately used should be frozen and stored at −80 °C Samples should be thawed fully before use |
When using blood-based biofluids such as serum and plasma, spectra from erythrocytes may mask that of other biomolecules, so they should be removed if not being directly investigated41 |
Biofluids may be placed onto slides and dried, or dried directly onto the, IRE44 |
Dry film analysis (where the fluid is dried onto the slide) often results in large signals compared with the wet biofluid, but measurements may be impeded by uneven distribution Only small sample volumes are needed, normally in the region of a few nanoliters136 |
| FFPE tissue samples |
FFPE tissue should be de-waxed for a minimum of 5 min in xylene and three washes should be performed Sample thickness should not exceed 8–12 μm (transmission, less for transflection; see Table 1) in order to avoid a nonlinear detector response (at absorbance values > 1.2 (for MCT) or > 1.5 (for deuterated triglycine sulfate)), to even total absorption |
Samples must be de-waxed in order to probe the full wavenumber range, as paraffin is known to have significant peaks at ~2,954 cm−1, 2,920 cm−1, 2,846 cm−1, 1,462 cm−1 and 1,373 cm−1, which may mask solvent-resistant methylene components of native tissue128,129 Samples are then cleared with acetone to remove any final xylene contamination Another recent and emerging alternative is to model the paraffin contribution and numerically de-paraffinize the sample36. In this way, the sample is not affected by chemical de-paraffinization, and intact tissue biochemical information is used for spectral histology |
De-waxed tissue should be floated onto slides |
If using tissue for imaging and extraction of tissue cell type, sample thickness is not just an SNR issue. The thicker the tissue, the greater the chance of probing heterogeneous layers and possibly multiple cell types, rendering cell type signal less pure |
| Cryosectioned tissue samples |
Tissue must be thoroughly thawed before IR analysis Once a sample is thawed, compo- nents may start to degrade, so we suggest imaging sections as soon as possible after thawing and drying, in a dark environment130 However, under dry conditions, cryosections can be stored for months without major problems other than lipid oxidation, as seen by the decrease of the ester carbonyl bands (degrades within 2 weeks; this can be avoided when samples are stored in a N2 atmosphere) |
Serial sections should be carefully isolated from the cryoblock to prevent OCT compound contamination of the final tissue slice |
Snap-frozen tissue should be cut and placed onto slides |
Although snap-freezing negates the use of fixatives such as formalin or the use of paraffin, it may damage the structural integrity of the tissue |
| Fixed cells | Medium contaminants must be removed before cells are placed in fixative such as ethanol or formalin For formalin fixation, cells should be washed twice in PBS before suspension in formalin for at least 30 min Slides should be dipped three times in double-distilled water (this should not be extended beyond quick dips) as formalin fixation can be reversed in the presence of water92 For ethanol fixation, cells should be washed three times in ethanol (min. 70% (vol/vol)) before being left to stand in ethanol for at least 1 h |
After formalin fixation, cells should be washed in HBSS before IR analysis to remove residual phosphate ions After ethanol fixation, slides should be left to dry for 24 h on the benchtop and 24 h in a desiccator so that all residual ethanol evaporates |
Cells can be grown onto IR substrates that have been first sterilized in 70% (vol/vol) ethanol, as growing directly onto the slides can preserve cell morphology They can also be grown in a 3D culture matrix, which can then be fixed or frozen and sectioned; this may provide the most realistic environ- ment in which cells can be studied |
If grown on slides, cells will typically be thin, as they grow and stretch over a 2D surface Cells fixed and then placed onto slides may be uneven in thickness, which may be resolved using cytospinning, which allows cells to be proportionally dispersed over the substrate |
| Live cells | Cells that are to be analyzed in suspension should be detached from the growth substrate using trypsin and then stored at 4 °C to prevent autolysis137 For ATR-FTIR measurements, cells can be seeded and grown directly onto the ATR IRE using a cell chamber59 |
Cells in suspension must be washed with PBS to remove residual medium or trypsin |
Spectra recorded in an aqueous environment show minimal dispersion because the refractive index of aqueous medium for the background single- beam spectrum closely matches that of the cell for the sample spectrum47 Therefore, cell suspension can be placed onto the IR slides as microdroplets Cells can be grown directly onto a detachable IRE such as diamond for ATR-FTIR analysis Live cells can also be analyzed in situ by the use of microfluidic devices21 |
The critical β-DNA conformational marker bands are enhanced in the hydrated state2, and thus can be used to determine the concentration of DNA in simple cells138,139 Single-cell micro- spectroscopy is inherently difficult because of the strong absorptivity of the water molecule, which can swamp the spectrum especially when the sample path length is > 10 μm (ref. 140) A bright source of IR photons is required to achieve a good SNR because the IR beam must usually pass through two IR transparent windows, cell medium and the hydrated cell, causing attenuation of the IR signal Thus, most measurements performed on single living cells with an FTIR microscope configuration use a synchrotron light source |
Sample thickness
Sufficient thickness of material needs to be placed onto the support matrix to allow a sufficiently large absorbance intensity to be recorded. In transmission and transflection modes, the specimen thickness needs to be adjusted appropriately: if it is too thick, the detector response function will be nonlinear so that Beer-Lambert’s law cannot be applied anymore. This has serious consequences for subsequent quantitative and classification analyses. In contrast, to achieve an adequate SNR and to avoid interactions of the evanescent wave with the underlying substrate, samples must also not be too thin. For example, when using ATR-FTIR spectroscopy, it is ideal if the specimen is three- or fourfold thicker than the penetration depth (that said, there is no maximum thickness for ATR-FTIR, and samples that are even a millimeter thick can be analyzed). This is pertinent for internal reflection measurements, which are commonly used for the disease diagnosis of biofluids; such samples can be naturally thinner in composition (especially with regard to cerebrospinal fluid (CSF), although this is not so much the case with blood or serum/plasma; serum, for example, is a solution containing a high protein concentration, ~80 mg ml−1). The effect of substrate interference on spectra, especially in reference to transflection measurements, has now been shown independently in the last year by several groups17,18,80. Given this, we would urge extreme caution regarding the use of Low-E slides with transflection measurements; with ATR-FTIR, it is unlikely that there will be optical effects associated with substrate.
Substrate choice
Proper consideration of the substrate (the slide or matrix) upon which the sample will be placed and any preparation steps associated with this are essential in order to acquire the best and most-reproducible spectra. For transmission measurements, this needs to be an IR-transparent material such as BaF2 or CaF2 (the latter, in particular, for live-cell IR spectroscopy), whereas for reflection or transflection measurements an IR-reflective substrate (e.g., Low-E slides) is required because glass alone absorbs the radiation and has a spectral signature in the mid-IR region81,82. Previously, it had been recommended that biological materials be placed on IR-reflective substrates. However, there now appears to be a shift in the general consensus that suggests that transmission or ATR spectroscopy measurements are more applicable to interrogation of biological material.
Microfluidic devices
Traditionally, aqueous sampling environments were unsuitable for IR spectroscopy because of the contribution of water. Development of microfluidic devices and processing to remove the water contribution has made it possible to achieve real-time, live-cell monitoring with IR spectroscopy. Nondestructive to cells, it better replicates physiological conditions; no labeling is required and the resolution is such that single cells can be studied83. The nondestructive nature of these methods has allowed studies to look at samples over time (e.g., stem cells in situ as they differentiate and chemical reactions in flow systems have been monitored84,85.
The key challenge of IR spectroscopy using microfluidics is associated with the materials’ transparency over the spectral range to be studied, and especially when live-cell monitoring is desirable. Many potential window materials are unsuitable on the basis of their water solubility (e.g., KBr and NaCl), toxicity toward the cells under observation (e.g., CdTe) or spectral dispersion (e.g., ZnS and BaF2)86. A flow chamber is used that combines IR transparency and robustness of diamond as window material. Although manufacture is complicated, the windows must be sufficiently thin (0.4–0.8 μm) to avoid multiple internal reflections86. CaF2 is extensively used as a window material, and a simple flow cell with inlet and outlet flow is constructed by clamping two CaF2 plates together. One of the plates is etched to form a 10-μm well, designed for the IR observation of live cells in aqueous media85. A similar device has been used for synchrotron IR spectroscopy of living cells using a surface micro-etched silicon substrate87. Further advances in the field have led to the development of sandwich devices and entirely polymeric devices.
Experimental design: spectral acquisition
Instrumental and operational settings to maximize spectral quality
When acquiring spectra, it is important to maximize as best as possible the SNR in order to produce high-quality spectral data (Table 3). There are a number of noise-related and signal-related parameters, with an effect on SNR, which can be altered depending on the instrument mode being used (e.g., point mode versus imaging)88-91. The instrumental and operational settings will be specific to the user experimental setup; Table 1 compares properties of different sampling modes for optimized spectral acquisition. An initial noise-related parameter that can be altered is the sampling aperture in point or mapping mode; this will reduce the SNR when the aperture size is reduced92. However, in imaging mode there is no aperture. The interferometer mirror velocity may also have an effect on SNR3. Weighting the interferogram with an apodization function will also contribute to a reduction in SNR, as this smoothing effect can incorporate spectral artifacts while one is attempting to optimize the information contained93. In general, the square root of the number of co-additions is proportional to the SNR, and therefore an increased number will enhance the SNR94.
Table 3.
Typical conditions of the main variables affecting SNR in spectroscopy instruments.
| Instrument options |
||||||
|---|---|---|---|---|---|---|
| FTIR | ATR-FTIR | |||||
|
|
||||||
| Variable | Single-element detector |
FPA |
Single-element detector |
FPA |
||
| Light source | Globara | Synchrotronb | Globarc | Synchrotrond | Globar e | Globarf |
| Sampling aperture | 15 × 15 to 150 × 150 μm |
5 × 5 to 20 × 20 μm |
700 × 700 μm FOV |
50 × 50 to 175 × 175 μm |
250–250 μm | 60 × 60 to 700 × 700 μm |
| No. of co-additions | 512 | 256 | 64 or 128 | 128 | 32 | 32 |
| Spectral resolution | 4 or 8 cm−1 | 4 or 8 cm−1 | 4 or 8 cm−1 | 4 or 8 cm−1 | 8 cm−1 | 4 or 8 cm−1 |
IR spectroscopy has a spatial resolution that is limited by the diffraction limit; hence, as the resolution approaches this value, the SNR is reduced to a point where there is no further gain in image quality95 A synchrotron radiation source (e.g., at the IR Environmental Imaging Facility (IRENI) at the Synchrotron Radiation Centre (SRC)) in the mid-IR region is 1,000 times brighter than a thermal globar source and thus may generate enhanced SNR spectra when using apertures approaching the diffraction limit; however, when using an FPA detector, this cannot be exploited as the brightness is applied over a larger area. By using multiple beams, such as at IRENI, the single-beam disadvantage when using an FPA may be overcome.
It is important to consider that an optimized and well-aligned benchtop instrument is not considered to be inferior with regard to SNR or image quality to a general synchrotron-based machine63. A number of options regarding the detector can also have an effect on the SNR, such as the choice between a thermal detector versus a quantum detector. A mercury cadmium telluride (MCT) quantum detector usually provides a superior SNR than, for example, a thermal detector such as a deuterated triglycine sulfate detector96. An optimized cooling system in the detector, such as thermoelectrical cooling, will also reduce the dark current produced by the detector, which has been shown to have a detrimental effect on SNR97,98. In addition, signal-related parameters can affect the SNR; for instance, an increase in the optical path length can reduce spectral quality, which has been particularly important in the analysis of aqueous samples such as biofluids33. When producing spectral images with the help of multielement detectors, such as an FPA, one must consider optimizing the SNR. The authors point readers to the authoritative reference on FTIR spectroscopy by Griffiths and De Haseth3 for theoretical and instrumental discussions; this book has supported the authors since their undergraduate studies and continues to support them today3.
Water vapor and instrument purging
The presence of water vapor in the instrumentation and sample area can result in reduced transmission of IR light, potentially obscuring important spectral details even at low spectral resolutions often used in biomedical IR spectroscopy. Water vapor interference can be minimized by computational subtraction of a pure water vapor spectrum from the sample spectrum99. The efficacy of this compensation is limited and it is therefore considered crucial before spectral acquisition to purge the instrumentation with dry air or nitrogen and/or desiccants to remove any water vapor that may contaminate spectra between 1,350 and 1,950 cm−1, and between 3,600 and 3,900 cm−1 (ref. 100). By doing so, ambient CO2 is also purged, thereby reducing its contribution to the spectra.
Acquisition of sample and background
Measurements of an FTIR absorption spectrum involve collecting a ‘single-beam’ spectrum. A background single-beam spectrum provides the source intensity, as modified by the instrument; placing a sample in the beam path and measuring the single beam again, theoretically, provides just the additional effect of the sample absorbance. A logarithm (to the base 10) of the ratio of these quantities provides the absorbance, which is directly related to concentration by Beer’s law. With point spectra, a background spectrum is typically retained for recording 5–10 sample spectra and with each different sample to reduce the effects of constantly changing atmospheric conditions. As spectral maps are composed of a large number of point spectra acquired in a stepwise manner, it is necessary to set up background scans to be taken at set intervals (e.g., at the end of every row) to account for the atmospheric variation over the extended acquisition time66. When acquiring spectral images, background spectra should be acquired over a defined time period, depending on the sample acquisition time.
Experimental design: data processing
Data processing is carried out in a sequence of steps (Fig. 3) and the most important factor determining its workflow is the analysis goal; typical spectroscopy software programs used are shown in Table 4. Here we describe two analysis goals: imaging and diagnosis. Other goals not covered here include pattern finding and biomarker identification101,102.
Table 4.
Some existing FTIR spectroscopy data analysis software.
| Software | Website | Description | License |
|---|---|---|---|
| Cytospec | http://www.cytospec.com | Software for hyperspectral imaging (IR and Raman) |
Commercial; free demo available |
| IRootLab | https://code.google.com/p/irootlab/ | MATLAB toolbox for biospectroscopy data analysis |
Open source |
| OPUS | http://www.bruker.com | Spectral acquisition software with data processing capabilities |
Commercial |
| Pirouette | http://www.infometrix.com | Chemometrics modeling software | Commercial |
| Unscrambler X | http://www.camo.com | Multivariate data analysis and design of experiments |
Commercial |
| PLS, MIA, EMSC toolboxes | http://www.eigenvector.com | MATLAB toolboxes for spectroscopy data analysis |
Commercial |
| OMNIC | http://www.thermoscientific.com | Spectral acquisition software with data processing capabilities |
Commercial |
| PyChem | http://pychem.sourceforge.net/ | Package for univariate and multivariate data analysis |
Open source |
| ENVI, IDL | http://www.exelisvis.com | Integrated development, data analysis and image processing suite |
Commercial |
| MCR-ALS toolbox | http://www.cid.csic.es/homes/rtaqam/tmp/WEB_MCR/welcome.htm | MATLAB Toolbox implementing the MCR-ALS algorithm |
Open source |
Imaging is defined as data analysis that uses an unsupervised data processing method to reveal tissue structure on a ‘spectral cube’ acquired by a mapping or imaging technique. Imaging allows for the study of shape and penetration of important histopathological features on the basis of the underlying chemistry28.
In contrast, a diagnosis using IR spectroscopy requires a more complex framework that uses supervised classification methods. A supervised data processing method is one that uses classes assigned a priori to each IR spectrum as teaching information to build models that are used later to predict the classes of a data set that does not have classes associated with its spectra103,104. The modeling process for diagnosis requires separate training and testing stages and respective training and test data sets. The optimal size of a training data set (i.e., one that will maximize classification accuracy at a reasonable cost of data set generation) has been underinvestigated to date, but it has been suggested that it may be problem dependent105. For example, in a study, one could start with ten samples (acquiring 5–10 spectra from each sample), creating a trained model with eight samples and testing the model using the remaining two samples; one could then repeat this procedure four more times, each time using two different samples for testing and the remaining eight samples for training (this is called five-fold cross-validation). The number of times that the classifier correctly guessed the class of the testing sample would be counted to calculate a classification rate (i.e., the number of correct guesses divided by the total number of guesses). Next, one could acquire spectra from an additional five samples and repeat the cross-validation process, comparing the new classification rate with the old one (it is expected to improve). The process of adding samples and repeating cross-validation could continue until the classification rate stops improving.
It is important to note that a diagnostic framework may be set to use either point spectra or image maps; in the latter case, the trained classification system can be used to predict tissue structure.
We describe the following data analysis steps: pre-processing, feature extraction (FE), clustering (unsupervised classification) and supervised classification, and we exemplify some visualization options in the ANTICIPATED RESULTS section. Quality control is another step that is not covered in this protocol, but there are guidelines on this available in the literature105,106.
Pre-processing
Pre-processing essentially aims to improve the robustness and accuracy of subsequent multivariate analyses and to increase the interpretability of the data by correcting issues associated with spectral data acquisition107. Pre-processing methods may be divided into de-noising, spectral correction, normalization and other manipulations; two or three methods are often combined (e.g., de-noising followed by spectral correction and normalization). The choices of pre-processing methods may depend on the analysis goal, the physical state of the sample, and the time and computing power available.
De-noising of IR spectra may be carried out with Savitzki-Golay (SG) smoothing, minimum noise fraction108 or wavelet de-noising (WDN)101. The latter is known to be the best method for eliminating high-frequency noise while still keeping intact high sharp peaks (this is essential in Raman spectra processing, but WDN works well on IR spectra too). Another option is to decompose the spectra by principal component analysis (PCA), and then reconstruct them from only a few of their principal components (PCs), thus discarding those PCs that represent mostly noise85,109.
Measurement characteristics that may require spectral correction include:
Sloped or oscillatory baselines that result from scattering, with resonant Mie scattering in biological materials being the most pronounced effect. The effects of sample (scattering centers, edges and substrates) have often been lumped together and the effects of the same on spectra are termed ‘artifacts’. Although this terminology was initially acceptable, it is now clear that there is a rational explanation for these effects and they arise merely from the coupling of morphology and optics. Hence, we will refer to these as morphological effects on spectra. There are two major efforts in understanding and resolving these effects to recover absorption spectra free from the effects of morphology. The first group of methods is termed ‘physics based’. In this approach, explicit optical image–formation modeling from first principles is used to predict and correct data. Here each sample effect (boundary scattering, scattering centers in the sample and substrate) needs to be explicitly accounted for. The theory has been shown to be generally valid and there are methods now for correcting the same for films, spheres and fibers16,110,111. Extension to more complex samples is still the subject of ongoing research. A second group of methods may be termed ‘model based’. In these methods, a model is assumed to explain all sample effects, typically, Mie scattering. Subsequently, rigorous theory is used to recover spectra, e.g., including extended multiplicative scattering correction (EMSC)112, resonant Mie scattering correction (RMi-eSC)113-115 and rubber band baseline correction116. An indirect way to deal with baseline slope is to apply first or second derivative to spectra using the SG algorithm. This alters the shape of the spectra, but may also resolve overlapped bands. Model-based methods will generally be faster than explicit modeling methods and may prove to be broadly useful but need to be validated in each case. A third approach, which was traditionally used but is now recognized to be of limited value, is to simply correct baselines with a piecewise linear approach. Obviously, this method is the fastest, as it requires the least effort to apply and no modeling. It is as yet unclear which of these methods works best.
Spectral contributions may arise from atmospheric water vapor, carbon dioxide, paraffin or other interfering compounds. Although these artifacts may be compensated mathematically through EMSC117 or other least-squares–based technique118, the most common actions are to remove contaminated spectral bands from the data set, improve the control of atmospheric conditions or take background spectra more often. In this aspect, before pre-processing, it is often useful to implement quality tests to verify SNR and minimize water vapor contribution. By following this approach, ‘bad quality’ spectra are discarded as they can influence subsequent analysis. The threshold values for defining ‘bad’ and ‘good’ spectra can be adjusted according to the biological application.
It is vital to normalize IR spectra to account for confounding factors such as varying thickness of sample. Common normalization methods are amide I/II peak normalization and vector normalization. Amide I/II normalization is often used after baseline correction, whereas vector normalization is often used after differentiation of spectra (after correction by differentiation, there is no longer a consistent amide I/II peak in the spectra to allow for amide I/II peak normalization). For imaging, leaving spectra non-normalized for chemical imaging or unsupervised clustering will reveal tissue structures primarily based on absorbance intensity, whereas normalization will highlight differences in biochemical structure. For diagnosis, some form of spectral normalization is conducted.
The optimal pre-processing method or sequence to apply is a subject of discussion and no universal best approach exists for all samples. Often the choices are based on the problems visually spotted in the spectra; a more objective criterion is to optimize the pre-processing method (e.g., through a genetic algorithm)119. In this protocol, we offer several alternatives based on cues identified by visual expression of raw (non-pre-processed) spectra, although objective validation will probably become more common in the future.
FE
FE methods process the IR spectra to form new variables based on the original variables (which are absorbance intensities). FE has an important or even essential role in both imaging and diagnosis. For imaging, FE is responsible for generating a single value based on the whole of an input IR spectrum. This value can subsequently be used to set the color of a pixel in the image; FE is repeated for all spectra, thus forming the pseudocolor image. Popular FE methods for imaging include calculating the ratios between wavenumber absorbance intensities, area under a subregion of the spectrum, selecting a single wavenumber or an ensemble of wavenumbers, or performing PCA. PCA may be applied to the spectral data set, followed by selection of a single PCA factor for the color gradient.
For diagnosis, FE constitutes an important data reduction step in order to match the complexity of the subsequent supervised classifier with the amount of data available so as to avoid over-fitting or undertraining. PCA is one particular popular form of unsupervised FE that is used for this purpose103. The number of PCA factors to retain may be subject to optimization. One way out is to order the PCA factors from the most to the least discriminant on the basis of their P values as determined by a statistical test. The percentage of explained variance can also be taken into account. Within FE, the subgroup of feature selection (FS) methods is particularly interesting because it can confer biological interpretability (i.e., identify the wavenumbers most important for classification) to the classification system. Popular FS methods include forward FS120 and COVAR121. Variance analyses may also be used to select spectral variables for elimination122. Another approach to FS is to use spectral features that are obtained from a biochemical understanding of the problem123. These cases in which direct spectral interpretation is possible are termed metrics for measures of biochemical activity in the samples. It is important to note that not all metrics may be useful biomarkers. Thus, even FE may be a multistep process, (i.e., one in which metrics are converted to statistically relevant biomarkers).
Clustering (unsupervised classification)
Clustering aims at sorting different objects (i.e., spectra) into categories or clusters on the basis of a so-called distance measure124. Clustering methods such as hierarchical cluster analysis (HCA) and k-means clustering (KMC) are frequently used in IR-imaging studies to identify tissue morphology23,125. HCA groups spectra into mutually exclusive clusters; in IR-imaging studies, HCA-based segmentation is achieved by assigning a distinct color to the spectra in one cluster. Because each spectrum of an IR-imaging experiment has a unique spatial (x,y) position, pseudocolor segmentation maps can be easily generated by plotting specifically colored pixels as a function of the spatial coordinates.
Supervised classification
Supervised or concept-driven classification techniques are machine-learning techniques for creating a classification function from training data. These methods involve a supervised learning procedure in which models are created that map input objects (spectra) to desired outputs (class assignments). Popular supervised techniques are artificial neural networks, support vector machines (Supplementary Method 3), linear discriminant classifier11,103,126 and Bayesian inference-base methods77. Among the many criteria guiding the choice of classifier, the most important is probably the accuracy (related to sensitivity and specificity) when tested on an independent test data set. Other criteria include ease to train, computational time, spatial resolution considerations127 and software availability. Classifiers such as artificial neural networks and support vector machines may require a two-stage training, where the first stage is dedicated to finding optimal tuning parameters or architecture and the second stage fits the classifier model to the training data. Linear discriminant classifier (LDC) is a parameterless classifier that requires only the fitting stage. A general rule of thumb is that if two different classifiers are equally well performing on an independent test data set, the simplest one should be preferred over the more complex one, as simpler classifiers are more likely to be better generalizers103.
MATERIALS
REAGENTS
▲ CRITICAL For sample preparation and analysis, please refer to Tables 1 and 2 and the INTRODUCTION for further information.
FFPE blocks: see Reagent Setup for further information
Sample preparation: advice regarding collection of biofluids, cryosectioned tissue samples, fixed cells and live cells can be found in the Reagent Setup section! CAUTION Human tissues (including biofluids, cytology or FFPE blocks) should be obtained with appropriate local institutional review board (e.g., in the UK, this is a Local Research Ethics Committee (LREC)) approval; generally, ethical permission will be granted for a carefully designed study in which patient participants sign a consent form. Worldwide, studies using human tissues should adhere to the principles of the Declaration of Helsinki. Similarly, for research using animals, appropriate approvals are required; The Animals (Scientific Procedures) Act of 1986 is the legislation that regulates the use of animals in scientific procedures in the United Kingdom and this is enforced by the Home Office, which issues the licenses required.
Other reagents
ThinPrep (PreservCyt Solution, Cytyc)
SurePath (TriPath Care Technologies)
-
Formalin, 10% (vol/vol), neutral buffered (Sigma-Aldrich, cat. no. HT501128)
! CAUTION It is a potential carcinogen, an irritant and an allergenic. Always work in a fume hood while handling it.
Acetone (Fisher Scientific, cat. no. A/0600/17) ! CAUTION Its vapors may cause dizziness. Always work in a fume hood while handling it.
Ethanol, 2.5 liters (Fisher Scientific, cat. no. E/0600DF/17)
Virkon (Antec, DuPont, cat. no. A00960632)! CAUTION It is an irritant.
Paraplast Plus paraffin wax (Thermo Fisher Scientific, cat. no. SKU502004)
Xylene (Sigma-Aldrich, cat. no. 534056)! CAUTION It is a potential carcinogen, an irritant and an allergenic. Always work in fume hood while handling it.
Histoclear (Fisher Scientific, cat. no. HIS-010-010S)! CAUTION It is an irritant.
Isopentane (Fisher Scientific, cat. no. P/1030/08)! CAUTION It is an extremely flammable, irritant, aspiration hazard and toxic reagent. Always work in fume hood while handling it.
Optimal cutting temperature (OCT) compound (Agar Scientific, cat. no. AGR1180)
Liquid nitrogen (BOC, CAS no. 7727-37-9)! CAUTION May cause asphyxiation and contact with skin will cause burns. Wear cryoprotective clothing and use it in a fume hood.
EQUIPMENT
Electronic equipment
For a list of commercial instruments available, please refer to Table 5
Table 5.
Instruments and corresponding data acquisition software.
| Manufacturer | Instruments | Software |
|---|---|---|
| Agilent Technologies |
Agilent 670-IR spectrometer |
Resolutions Pro |
| Cary 600 series FTIR spectrometers |
||
| Agilent 600 series FTIR microscope |
||
| Bruker Optics | Bruker Tensor 27 spectrometer | OPUS |
| ALPHA FT-IR spectrometer | ||
| HYPERION series FT-IR microscope |
||
| LUMOS FT-IR microscope | ||
| JASCO UK | JASCO FTIR-4100 series | Spectra Manager |
| JASCO FTIR-6000 series | ||
| IRT-5000 FTIR microscope | ||
| PerkinElmer | PerkinElmer Frontier | Spectrum 10 |
| Spectrum Two | ||
| Spotlight FTIR microscope system |
||
| Thermo Fisher Scientific |
Thermo Nicolet iS50 spectrometer system |
OMNIC 8 |
| Thermo Nicolet Scientific FTIR 5700 spectrometer with continuum microscope |
||
| Shimadzu | IRTracer-100 spectrometer IRAffinity-1S spectrometer |
Lab Solutions IR |
Substrate
Low-E slides (Kevley Technologies, CFR)
BaF2 slides (Photox Optical Systems)
Silicon multi-well plate (Bruker Optics)
Superfrost slides: these can be obtained from various manufacturers, e.g., Menzel Glazer Superfrost slides (Menzel-Glaser, cat. no. AA00008132E); Thermo Scientific SuperFrost slides (Thermo Fisher Scientific); or Fisherbrand Superfrost slides (Fisher Scientific)
Accessories
Coverslips (Thermo Fisher Scientific, cat. no. 102440)
Specac Golden Gate single-reflection diamond ATR accessory (Specac)
Microtomes: these can be obtained from various manufacturers, e.g., Microtome (Surgipath Medical Industries); Leica rotary microtomes (Leica Microsystems, Davy Avenue Knowlhill); or Bright Cryostat (Bright Instruments)
Microtome blades: these can be obtained from various manufacturers, e.g., Feather disposable microtome blades S35 (VWR, cat. no. SURG08315E), Edge-Rite disposable microtome blades (Thermo Fisher Scientific); or Leica Surgipath DB80 blade (Leica Microsystems)! CAUTION Blades are extremely sharp; handle and dispose of them with care.
Paraffin section mounting bath (40–75 °C; Electrothermal, cat. no. MH8515)
Desiccator: these can be obtained from various manufacturers, e.g., desiccator (Duran Group) or WHEATON Dry-Seal vacuum desiccators (Wheaton Industries)
Labofuge 400e (Heraeus Instruments)
REAGENT SETUP
FFPE blocks
These are prepared according to the standard methods used routinely in all pathology laboratories; the overall steps are: immerse fresh tissue in formalin solution that acts as a chemical fixative; dehydrate the tissue in sequential washes of xylene and ethanol; and embed the tissue in paraffin wax, which creates an airtight barrier. Tissue blocks can then be stored indefinitely at room temperature (20–22 °C).
Biofluids
These are primarily blood plasma or serum, but can also potentially include cerebrospinal fluid, saliva or urine. Typically, after acquisition, such samples should be stored in appropriate tubes at −85 °C until they are thawed before analysis.
Cryosectioned tissue samples
Tissue samples can be snap-frozen and stored at −80 °C before use. Tissue should be coated with optimal cutting temperature (OCT) compound before freezing, and it should be frozen with isopentane cooled with liquid nitrogen.
Fixed cells
Typically, these would originate from cytology specimens placed in a fixative buffer; an ideal example of this is cervical cytology. However, it could be extended to any cell type isolated in the form of a suspension in a preservative buffer solution.
Live cells
This is an emerging area within the field whereby viable cells can be spectrochemically analyzed, primarily in a constructed microfluidic platform (for a typical method, see Supplementary Method 4).
EQUIPMENT SETUP
Software
Two types of software are required: spectral acquisition and data analysis. Spectral acquisition software is normally provided by the instrument manufacturer. Most instrumentation software also provides a number of preprocessing and sometimes more advanced data analysis options. Various data analysis software programs and packages exist, ranging from those for general-purpose use to those targeting specific data analysis tasks (e.g., multiplicative curve resolution–alternating least squares (MCR-ALS)). A popular development environment and programming language is MATLAB (http://www.mathworks.com) in which customized software can be written for specific tasks. Python (http://www.python.org) is another programming language that is becoming increasingly popular in the FTIR spectroscopy field, and it has the advantage of being open source. For a list of commonly used software and packages, please refer to Table 4.
PROCEDURE
Sample preparation
1| Prepare the samples by following the steps listed in one of the options given below. Perform the steps in option A for FFPE tissue samples; option B for cryosectioned tissue samples; option C for cytological specimens; and option D for biofluids.
Live cells may be prepared in three main ways for IR-transmission studies: grown directly onto IR substrates; grown in a 3D culture matrix (and then processed as described in options A and B); or fixed in suspension, e.g., as cervical cytological specimens in fixative obtained from hospital pathology laboratories. Cells that are fixed in suspension should be processed by following the steps in option C.
To grow cells on IR substrates, sterilize the IR substrate for 1 h in 70% (vol/vol) ethanol before growing cells directly onto the chosen IR substrate.
| Cells grown onto IR substrates | Sterilize the IR substrate for 1 h in 70% (vol/vol) ethanol before growing cells directly onto the chosen IR substrate. Generally, cellular materials are then fixed in order to preserve their architectural integrity, and the samples are stored in a desiccator prior to spectral acquisition (Step 2). |
| Cells grown in 3D culture matrix | Cells may be grown on 3D culture matrices (a tissue culture environment or device in which live cells can grow or interact with their surroundings in three dimensions), and subsequently fixed or snap-frozen and sectioned as described for tissue samples in Step 1A and Step 1B |
(A) FFPE tissue • TIMING 50 min
-
(i)
Obtain FFPE tissue blocks of interest from a pathology laboratory.
-
(ii)
Place a FFPE block onto an ice block for 10 min. Use a microtome to trim into the block to expose the entire tissue sample to the face of the block. This will ensure that a full tissue section is cut for analysis. Place trimmed blocks back on ice for 10 min.
▲ CRITICAL STEP Make sure that the blocks are cold before cutting sections. This hardens the wax, reducing the friction between the block surface and blade allowing a much smoother cut.
-
(iii)
Cut a ribbon of 10-μm sections and float it onto a heated water bath (40–44 °C). Separate the individual sections with forceps.
▲ CRITICAL STEP Optimal tissue thickness for the maximum SNR should be determined in-house by applying variable thicknesses of sections (depending on the tissue type) to slides for IR interrogation, e.g., ~3 μm (e.g., for bone), 5 or 10 μm (e.g., for prostate tissue), and 15-μm serial sections to BaF2, CaF2 or Low-E slides. SNR is judged on the quality of the raw spectra; in particular, the presence of many narrow, sharp peaks indicates high noise. If using tissue for imaging and extraction of tissue cell type, sample thickness is not just an SNR issue. The thicker the tissue, the greater the chance of probing heterogeneous layers and perhaps multiple cell types, rendering the cell type signal less pure.
▲ CRITICAL STEP Depending on the melting point of the paraffin wax used for embedding tissue samples, the temperature of the water bath will need to be adjusted to prevent melting of the wax.
-
(iv)
Prepare tissue slides by re-floating a single 10-μm-thick tissue section onto a BaF2, CaF2 or Low-E slide for FTIR microspectroscopy or ATR-FTIR spectroscopy. In our experience, a 5–10-μm section is the optimal thickness for maximum SNR.
▲ CRITICAL STEP As BaF2 slides can be 1 cm × 1 cm in size to fit common slide holders, a H&E-stained parallel section may be required to identify an area of interest for analysis. Once a section is floated onto the water bath, sections can be picked up on normal microscope slides, dissected using a scalpel for the area of interest and floated back onto water for application to BaF2 slide.
-
(v)
Place the tissue slide in a 60 °C oven for 10 min.
-
(vi)
De-wax the tissue slide by immersing it in xylene for 5 min at room temperature. Repeat this step twice with fresh xylene. For small, round slides that are difficult to handle during solvent immersion, slides can be encased into plastic histology cassettes that can be threaded round a large metal clip. The same procedure can be conducted using hexane.
▲ CRITICAL STEP For IR analysis, it is necessary to de-wax the tissue in order to probe unhindered the full wavenumber range. This is paramount as paraffin is known to have significant peaks at ~2,954, 2,920, 2,846, 1,462 and 1,373 cm−1. If there is uncertainty about paraffin removal, these regions of the spectrum can be removed from subsequent analysis. However, this comes at the cost of probing many solvent-resistant methylene components of the native tissue128,129.
-
(vii)
Sequentially, wash and clear the tissue slide by immersing it in acetone or 100% ethanol for 5 min at room temperature.
-
(viii)
Allow the tissue slide to air-dry before placing it into an adequate-sized Petri dish for storage in a desiccator.
■ PAUSE POINT Slides can be stored in a desiccator before IR interrogation; in our experience, storage should be <1 year.
(B) Snap-frozen and cryosectioned tissue samples • TIMING 120 min + drying time (3 h)
! CAUTION Snap-freezing should be carried out in a fume hood while you are wearing cryoprotective gloves, clothing and a facemask.
-
(i)
The fresh tissue should be no more than 2 cm in any one dimension; gently blot away any fluids from the surface, place a cryomold and fill the mold with OCT compound.
-
(ii)
Fill a plastic cryobucket with 3–4 cm of liquid nitrogen. Pour isopentane into the stainless steel beaker until it is about 1–2 cm deep. Place the stainless steel beaker into the liquid nitrogen and allow temperatures to equilibrate (3–5 min).
-
(iii)
Take the cryomold containing the tissue sample in OCT compound and use long forceps to lower it into the isopentane; hold until the OCT compound freezes (60–90 s).
-
(iv)
Remove the cryomold and transfer it to the bucket of dry ice. Wrap snap-frozen tissue in aluminum foil and label it before storing it in −80 °C freezer.
■ PAUSE POINT Snap-frozen tissue can be stored in a −80 °C freezer for several months.
-
(v)
Retrieve previously prepared snap-frozen tissue blocks from the −80 °C freezer and transfer them to the cryostat in dry ice to prevent thawing.
-
(vi)
Unwrap the frozen block from its protective foil covering and mount it into the cryostat. Allow the block to equilibrate to the cryostat temperature for 30 min. The optimum cryostat cutting temperature will depend on the sample, but −20 °C will be suitable for most tissues.
-
(vii)
Cut sections with a cryostat until the region of interest is reached. Next, take serial sections of the tissue sample at the desired thickness for your study.
-
(viii)
Carefully mount the sections onto the substrate window. Immediately upon acquiring the cryosection, transfer the slides to a slide box on dry ice, wrap them in foil and store them at −80 °C to preserve the biochemical content.
-
(ix)
Before imaging, bring slides to room temperature in a dark slide box with desiccant for several hours (minimum 3 h) until they are dry.
▲ CRITICAL STEP The tissue needs to be adequately thawed before IR analysis (freezing and thawing may also damage the structural integrity of the tissue). During the thawing process, store the sample under dark, dry conditions at room temperature. Light exposure is only advised during the short time required for the instrumental setup; this maintains the stability of spectral acquisition. Room lights and bright-field microscope illumination should be switched off during measurement collection130.
(C) Cytological specimens • TIMING 30 min + desiccation time (24 h)
-
(i)
For formalin fixation, cellular pellets should be washed twice in PBS to remove culture medium before resuspension in formalin solution (in which they should remain for at least 30 min). Before IR analysis, cells should be washed with HBSS to wash out the residual phosphate ions.
▲ CRITICAL STEP SurePath and ThinPrep fixative solutions, used in hospital pathology laboratories, have IR signatures in the biochemical cell–fingerprint region and should therefore be removed from the sample by sequential washes before analysis. Alcohol-based fixatives may remove some lipids from the sample.
-
(ii)
Resuspend the remaining cell pellet in 0.5 ml of distilled water, transfer the cells to the appropriate IR slide and allow them to air-dry before storing in a desiccator. Cells may be transferred to a slide as microdroplets, or they can be cytospun.
-
(iii)
For cytospinning, take a maximum volume of 200 μl of cells in suspension (spin-fixed cells at 800g (g force = 0.0000118 × radius of rotation (mm) × r.p.m.2) for 5 min). After spinning, leave the slide to air-dry for 24 h; the centrifugal force will have squashed the cells onto the slide, but if you try to wash the slides with water straight away you might lose them. After this time, wash the slide with 1-ml aliquots of deionized water three times (at around 5–10 s per wash, more water can be used if found necessary and cells stick adequately). The cells will remain on the slide and can always be washed further if traces of salts remain.
▲ CRITICAL STEP When transferring the cellular material to the slide, ensure that an even deposit of cells is placed on the slide. Cytospinning allows the cells to be proportionally dispersed over the substrate. If the cells are particularly small, they may ‘bounce’ off the slide during the spinning instead of getting stuck down. In this case, do a 5-min spin at 400g, then another 5-min spin at 800g to ensure firm plating.
(D) Biofluids • TIMING 10 min
-
(i)
Biofluids (i.e., urine, serum, plasma and saliva) should be immediately stored at −80 °C in cryovials after collection from pathology laboratories and thawed at room temperature before use.
-
(ii)
Samples of biofluids are painted directly onto the aperture (e.g., for ATR analysis) or a standard amount is pipetted onto suitable IR substrates (50–250 μl would be typical, but depending on the biofluid, preliminary analysis would be needed).
-
(iii)
Samples are allowed to dry before analysis.
▲ CRITICAL STEP Contact of the sample with the crystal is a very important parameter for ATR-FTIR analysis. If you are using an aperture ATR-FTIR accessory, 1 μl of sample has been shown to be dry within 8 min44.
Acquisition of spectra
2| Acquire spectra by ATR-FTIR spectroscopy (option A) or transmission FTIR microspectroscopy (option B) or FPA. A standard operating procedure for direct-drop ATR-FTIR for biofluid analysis is included in Supplementary Method 1; this would primarily be used when very small aliquots of the sample are available. A standard operating procedure for FTIR-FPA imaging with an Agilent 670-IR spectrometer coupled with an Agilent 620-IR microscope and FPA detector is included in Supplementary Method 2 (this file also contains a troubleshooting section).
(A) ATR-FTIR spectroscopy • TIMING 20 min (10 spectra)
-
(i)
Open the instrument-operating software.
-
(ii)
Apply instrumental settings (guidelines are described in ‘Experimental design: spectral acquisition’).
-
(iii)
Check the path where files are to be saved; set the file name according to a previously devised file-naming convention.
-
(iv)
Visualize the sample through the instrument digital camera to locate the region of interest from which you wish to acquire the spectrum.
▲ CRITICAL STEP If the instrument has been switched off, make sure you check the interferogram signal for the correct location and amplitude. The system may need to be re-aligned if it has been moved or if components have been changed.
-
(v)
Clean the ATR IRE with distilled water and dry it with tissue.
▲ CRITICAL STEP Make sure that the crystal is thoroughly cleaned and dried before a background acquisition.
-
(vi)
To acquire a background spectrum, the IRE should not be in contact with the sample or slide, and it should be open to the surrounding environment. Record a background spectrum.
▲ CRITICAL STEP It is very important that a background spectrum is taken before every sample. Also, a background spectrum should be taken if atmospheric changes occur (e.g., if a door has been suddenly opened).
-
(vii)
Place the slide in contact with the IRE.
▲ CRITICAL STEP Ensure that the ATR IRE is completely covered by the sample and that the minimum sample thickness is 3–4 times the depth of penetration to ensure that there is no interference from the substrate.
-
(viii)
Acquire a spectrum.
(B) FTIR microspectroscopy • TIMING 1 h per sample (~12 spectra) or 6 h per sample (image map, ~72 spectra)
-
(i)
Switch on the microscope and instrument.
-
(ii)
Fill the detector with liquid N2.
▲ CRITICAL STEP If you are using a MCT detector, filling it with liquid N2 is essential; allow the detector (and therefore the signal) to stabilize (~10 min) to an optimal peak-to-peak value. Top up with N2 every 9 h (depending on the instrument).
-
(iii)
Open the instrument-operating software.
-
(iv)
Apply settings.
-
(v)
Use the software to get a view of the slide as seen through the microscope.
-
(vi)
Load the sample onto the stage and focus the microscope.
-
(vii)
To check the signal quality, move to a sample-free area of the slide and adjust the position to bring the surface of the blank area of the substrate into focus.
-
(viii)
In our experience, the optimal sample aperture for a benchtop FTIR spectrometer with a globar source is 20 μm × 20–100 μm × 100 μm (dependent on sample quality and instrumental limitations). Apply the aperture size.
▲ CRITICAL STEP Optimization of the aperture size should be performed to confirm the smallest possible aperture that can be used to acquire spectra with a high SNR.
? TROUBLESHOOTING
-
(ix)
Use the joystick to move the sample around the microscope stage to identify points or areas to interrogate.
-
(x)
Select a clean, sample-free point on the slide and acquire a background spectrum according to your device.
▲ CRITICAL STEP Acquire a background spectrum each time the detector is filled with liquid N2 and at regular intervals (or before each sample) to account for atmospheric changes.
-
(xi)
Acquire a sample measurement either as a point map or as an image map.
A point map Select a number and location of points of interest Image map Use automatic allocation of adjacent points in a grid ▲ CRITICAL STEP Be sure to define a number of points (or map size) that does not exceed the scheduled time frame of the liquid N2 top-up.
▲ CRITICAL STEP The integration time is essentially a measure of the time for which the shutter is open to collect the incoming photons. The aim is to optimize the SNR without saturating the detector. If the integration time is too high, the user will observe saturation effects in the FTIR images; if it is too low, the data quality and SNR will be reduced as the FPA has not been fully illuminated. This calibration is a nonuniformity correction, and results are shown with measures of high and low flux (in counts) and the number of out-of-range pixels.
-
(xii)
Acquire spectra.
■ PAUSE POINT Once the spectra are saved they can be stored in a database until data processing.
Data pre-processing • TIMING 15 min–4 h (depending on the size of the data set)
▲ CRITICAL Steps 3–7 below all contain different options at each step; however, there are combinations of these steps that may be more or less appropriate than others, depending on the sample type, instrumentation setup, noise level, need for visualization of spectra, personal preference and classification performance among other factors (Table 4). Although they are usually carried out in the sequence presented, none of the steps from 3 to 7 are mandatory. For guidelines on choosing specific preprocessing steps and options, please refer to the ‘Experimental design data processing’ section. The reader may also refer to the Supplementary Method 3 for an illustrated example of a pre-processing sequence applied to a real-world data set using specific software.
3| De-noise the spectra (optional, depending on the SNR of the spectra). Consider using one of the following de-noising algorithms: Savitzky-Golay de-noising, WDN (not commonly used, but is a nonlinear method with its own advantages), PCA noise reduction or minimum noise fraction.
4| Perform spectral correction, which can be carried out using physical theory–based methods such as RMieSC or rubber band baseline correction49,53,113-115.
5| Perform SG differentiation (first differentiation is most used; second differentiation is also common).
6| Perform data normalization. This can be done using min-max normalization (e.g., normalization to the amide I/II peak) or vector normalization.
7| Scale the variables: this could be done by standardization (normalization of variables to zero mean and unit s.d.) or by normalization to a 0–1 range.
Data analysis • TIMING 1 h–2 d (depending on file size)
8| Choose a data analysis procedure appropriate to your analysis goal; here we cover diagnosis (supervised classification; option A below) and imaging (options B–D). For option A, data sets obtained with a single-element IR detector are normally used and the physical location of one spectrum within its sample is used in the data analysis. The procedures for creating an image use a data set acquired by point or array mapping, or an FPA detector. Each spectrum has its Cartesian (x, y) location. There are three different options for achieving this step (options B–D). Options B and C are suitable for chemical imaging using the FE method and unsupervised classification, respectively. Option D (supervised classification) is suitable for spatial diagnosis of tissues. It uses a training set of images to build a model to subsequently apply this model to unknown images. After performing options B, C or D, follow the instructions in option E.
(A) Diagnosis (supervised classification)
▲ CRITICAL Training and test data sets are required. In rare cases, one single training and test data set pair could be enough to obtain a meaningful estimation of real-world classification; however, most of the time such estimation is obtained through cross-validation, in which the procedure below (training and testing) is repeated multiple times to get an average performance (or error) estimation.
-
(i)
FE training. Input pre-processed training data set into the FE algorithm of your choice (see ‘Experimental design: data processing’) in training mode. Generate a model that will be able to subsequently extract features from a test data set.
-
(ii)
Classifier training. Apply the FE model obtained in a previous step to the training data set. Next, input the FE-processed data set to the classification algorithm of your choice (see ‘Experimental design: data processing’) in training mode. Generate a model to be subsequently applied to test data set.
-
(iii)
Testing. Apply the trained FE model to the test data set to obtain an FE-processed output data set; input the FE-processed data set into classification model to obtain one-class estimation per spectrum. If there are several spectra per sample, conduct a ‘majority vote’ procedure to obtain one class estimation per sample.
▲ CRITICAL STEP Training and testing should be repeated through a cross-validation procedure, depending on the sample size.
(B) Chemical imaging using an FE method
-
(i)
Map each spectrum into a single scalar value. Choose an FE technique to obtain a scalar value for each spectrum (this value will be subsequently used to address a particular color within a gradient color map). Refer to ‘Experimental design: data processing’ for guidelines on choosing the FE method.
-
(ii)
Continue to Step 8E.
(C) Clustering (unsupervised classification)
-
(i)
Apply a clustering algorithm (e.g., HCA or k-means) to organize the spectra into clusters.
-
(ii)
Assign a different integer number to each cluster and continue to Step 8E.
(D) Supervised classification for imaging
-
(i)
Conduct histological assessment of the training set to identify different regions within the training set images; this will be used as teaching information for the supervised learning algorithms.
-
(ii)
Apply Step 8A(i,ii), using the training data set obtained in the previous step to obtain a classification model.
-
(iii)
Apply the model to the test data to obtain one class estimation per spectrum in the image.
-
(iv)
Assign a different integer number to each class and continue to Step 8E.
(E) Mapping different scalar values to different color tones
-
(i)
Map different scalar values obtained in previous steps into different color tones. For chemical imaging (Step 8B), a gradient color map is used (e.g., red to yellow, rainbow and so on), whereas for Step 8C and Step 8D, an indexed color map in which each cluster or class is represented by a color of choice is suitable. Although the idea is presented here for understanding, this step is normally carried out by imaging software.
? TROUBLESHOOTING
Optimizing the sample aperture
To overcome the problem of over- and undersampling for fine imaging, the spatial sampling area should be at least two times larger than the (spatial) frequency of the feature under study. The step size should be equal or smaller than the aperture size divided by 2.
For Troubleshooting for FTIR imaging with an Agilent 620 IR microscope coupled with an Agilent 670/680 IR spectrometer, see Supplementary Method 2.
• TIMING
Step 1A, FFPE tissue: 50 min
Step 1B, snap-frozen and cryosectioned tissue samples: 120 min + drying time (3 h)
Step 1C, cytological specimens: 30 min + desiccation time (24 h)
Step 1D, biofluids: 10 min
Step 2A, ATR-FTIR spectroscopy: 20 min (10 spectra)
Step 2B, FTIR microspectroscopy: 1 h per sample (~12 spectra) or 6 h per sample (Image map, ~72 spectra)
Steps 3–7, data pre-processing: 15 min–4 h (depending on the size of the data set)
Step 8, data analysis: 1 h–2 d (depending on the file size)
ANTICIPATED RESULTS
Preprocessing options
Figure 4 is a basic example that shows a set of ATR-FTIR raw spectra (cut to the 1,800–900 cm−1 region) and their appearance after being pre-processed by different methods. Rubber band baseline correction is one of the options to remove sloped baselines. Normalization to the amide I/II peak shifts and scales all the spectra so that their vertical minimum is at zero and the amide I/II peak of all spectra match at the same height. To resolve overlapping bands, mathematical derivatives are used to narrow their full width at half height value (FWHH). Narrower bandwidths (i.e., higher resolution of differential spectra) potentially allow for subtle differences between spectra to be more easily resolved. However, each differentiation amplifies noise and therefore the SG differentiation algorithm (with implicit de-noising) is often used. Vector normalization is applied after differentiation to normalize the Euclidean norm of each spectrum to unity.
Classification of blood plasma
This example shows a comparison of supervised classification performance between different combinations of pre-processing, FE and supervised classification methodologies (Fig. 5). This data set consisted of blood serum and plasma samples of patients with ovarian cancer or endometrial cancer (n = 30 for both) and control patients without ovarian cancer (n = 30) analyzed with ATR-FTIR spectroscopy (7 spectra per sample)41,131. The classification rate, defined as the average between sensitivity and specificity, was used as a classification performance measure to class patients on the basis of their disease status (i.e., ‘normal’ versus ‘cancer’). The example illustrates that no single pre-processing, FE or supervised classification methodology is the absolute best, but a combination of these may be the best solution to the problem posed. The counterpoint to this is that different data sets may require different pre-processing, FE and/or supervised classifier methodologies, as pointed to in the machine-learning literature103. This is evidence that different combinations of methodologies should be attempted and compared in any diagnostic study.
Figure 5.
Classification rates (% classification ± s.d.) of all possible combinations between three different pre-processing, three different feature extraction and two different supervised classifier options. Pre-processing options: rubber band baseline correction followed by normalization to the amide I peak; first Savitzky-Golay (SG) differentiation (7 points; second order) followed by vector normalization; and second SG differentiation followed by vector normalization. FE options: PCA (optimization of number of PCs); forward feature selection (FFS) using multivariate analysis of variance (MANOVA) P values as a criterion to including the next variable (this is similar to the COVAR method for optimization of number of selected features); and ‘Identity’ (FE skipped). Supervised classifier options: linear discriminant classifier (LDC); and support vector machine (SVM; using Gaussian kernel; optimization of the C and γ parameters133,134). The figure’s cells are gradient-colored according to their respective classification rate inside (yellow to red). RBBC, rubber band baseline correction.
Imaging of human colon mucosa in Cytospec using agglomerative HC and KMC
An example of imaging of human colon mucosa sections by using agglomerative hierarchical clustering and KMC is shown in Figure 6. The image was produced using the Cytospec software. Figure 6c shows the original histological image from which FTIR spectra were recorded at a spatial resolution of 232 × 233 pixels. Some images (Fig. 6b,d) have been reconstructed using the multivariate methods of agglomerative hierarchical clustering (AHC) and KMC, respectively, with both demonstrating clear differentiation of the histological structures of the sample analyzed.
Figure 6.
IR image reconstruction of a section of human colon mucosa. (a) Chemical map based on the integrated absorbance of the amide I band (1,620–1,680 cm−1). (b) IR imaging using agglomerative HCA (six clusters). (c) Standard histological preparation of the colonic mucosa. (d) IR map generated on the basis of k-means clustering (15 clusters). Adapted with permission from ref. 135.
Supplementary Material
ACKNOWLEDGMENTS
Over several years, work in F.L.M.’s laboratories has been funded by the UK Engineering and Physical Sciences Research Council (EPSRC), the Rosemere Cancer Foundation and the UK Biotechnology and Biological Sciences Research Council (BBSRC). Work in R.B.’s laboratories has been funded by the US National Institutes of Health (grants R01CA138882 and R01EB009745). M.J.W. acknowledges the Department of Pathology, University of Illinois at Chicago for funding. P.R.F. and P.G. acknowledge the EPSRC. M.J.B. acknowledges the Rosemere Cancer Foundation, the EPSRC, Brain Tumour North West, the Sydney Driscoll Neuroscience Foundation and the Defence Science and Technology (Dstl).
Footnotes
Note: Any Supplementary Information and Source Data files are available in the online version of the paper.
AUTHOR CONTRIBUTIONS
F.L.M. is the principal investigator who conceived the idea for the manuscript; M.J.B. and K.M.D. provided information regarding FTIR FPA imaging; and P.R.F. provided information regarding microfluidic devices; J.T. wrote sections regarding Data processing, ANTICIPATED RESULTS and Figures, as well as maintaining a working manuscript; H.J.B. wrote the Instrumentation and Spectral acquisition sections; K.A.H. and B.O. wrote the Sample preparation and MATERIALS sections; R.J.S. wrote the INTRODUCTION and PROCEDURE, and contributed to the ANTICIPATED RESULTS section; C.H. provided material for Sample preparation, PCA–k-means clustering and cross-validation; P.L. provided figure suggestions and information regarding water vapor; B.R.W. provided information regarding live-cell imaging; R.B. provided significant revisions to the final manuscript; M.J.B., P.B., K.M.D., N.J.F., C.H., P.L., P.L.M.-H., G.D.S., J.S.-S., M.J.W., S.W.F., B.R.W. and P.G. all provided feedback on the manuscript; and F.L.M. brought together the text and finalized the manuscript.
COMPETING FINANCIAL INTERESTS
The authors declare competing financial interests: details are available in the online version of the paper.
References
- 1.Bellisola G, Sorio C. Infrared spectroscopy and microscopy in cancer research and diagnosis. Am. J. Cancer Res. 2012;2:1–21. [PMC free article] [PubMed] [Google Scholar]
- 2.Diem M, Romeo M, Boydston-White S, Miljkovic M, Matthaus C. A decade of vibrational micro-spectroscopy of human cells and tissue (1994-2004) Analyst. 2004;129:880–885. doi: 10.1039/b408952a. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Griffiths P, De Haseth JA. Fourier Transform Infrared Spectrometry. 2nd edn. John Wiley & Sons; 2007. [Google Scholar]
- 4.Walsh MJ, et al. FTIR microspectroscopy coupled with two-class discrimination segregates markers responsible for inter-and intra-category variance in exfoliative cervical cytology. Biomark. Insights. 2008;3:179–189. doi: 10.4137/bmi.s592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bhargava R, Wall BG, Koenig JL. Comparison of the FT-IR mapping and imaging techniques applied to polymeric systems. Appl. Spectrosc. 2000;54:470–479. [Google Scholar]
- 6.Bhargava R. Infrared spectroscopic imaging: the next generation. Appl. Spectrosc. 2012;66:1091–1120. doi: 10.1366/12-06801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Colarusso P, et al. Infrared spectroscopic imaging: from planetary to cellular systems. Appl. Spectrosc. 1998;52:106–120. [Google Scholar]
- 8.German MJ, et al. Infrared spectroscopy with multivariate analysis potentially facilitates the segregation of different types of prostate cell. Biophys. J. 2006;90:3783–3795. doi: 10.1529/biophysj.105.077255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Walsh MJ, et al. Fourier transform infrared microspectroscopy identifies symmetric PO2− modifications as a marker of the putative stem cell region of human intestinal crypts. Stem Cells. 2008;26:108–118. doi: 10.1634/stemcells.2007-0196. [DOI] [PubMed] [Google Scholar]
- 10.Kelly JG, et al. Derivation of a subtype-specific biochemical signature of endometrial carcinoma using synchrotron-based Fourier-transform infrared microspectroscopy. Cancer Lett. 2009;274:208–217. doi: 10.1016/j.canlet.2008.09.018. [DOI] [PubMed] [Google Scholar]
- 11.Romeo MJ, et al. In: Vibrational Spectroscopy for Medical Diagnosis. Diem M, Lasch P, Chalmers J, editors. John Wiley & Sons; 2008. [Google Scholar]
- 12.Diem M, et al. Comparison of Fourier transform infrared (FTIR) spectra of individual cells acquired using synchrotron and conventional sources. Infrared Phys. Technol. 2004;45:331–338. [Google Scholar]
- 13.Kole MR, Reddy RK, Schulmerich MV, Gelber MK, Bhargava R. Discrete frequency infrared microspectroscopy and imaging with a tunable quantum cascade laser. Anal. Chem. 2012;84:10366–10372. doi: 10.1021/ac302513f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bhargava R, Levin IW. Spectrochemical Analysis Using Infrared Multichannel Detectors. Wiley-Blackwell; 2005. [Google Scholar]
- 15.Davis BJ, Carney PS, Bhargava R. Theory of mid-infrared absorption microspectroscopy: II. Heterogeneous samples. Anal. Chem. 2010;82:3487–3499. doi: 10.1021/ac902068e. [DOI] [PubMed] [Google Scholar]
- 16.Davis BJ, Carney PS, Bhargava R. Theory of mid-infrared absorption microspectroscopy: I. Homogeneous samples. Anal. Chem. 2010;82:3474–3486. doi: 10.1021/ac902067p. [DOI] [PubMed] [Google Scholar]
- 17.Filik J, Frogley MD, Pijanka JK, Wehbe K, Cinque G. Electric field standing wave artefacts in FTIR micro-spectroscopy of biological materials. Analyst. 2012;137:853–861. doi: 10.1039/c2an15995c. [DOI] [PubMed] [Google Scholar]
- 18.Bassan P, Sachdeva A, Lee J, Gardner P. Substrate contributions in micro ATR of thin samples: Implications for analysis of cells, tissue and biological fluids. Analyst. 2013;38:4139–4146. doi: 10.1039/c3an00363a. [DOI] [PubMed] [Google Scholar]
- 19.Miljkovi M, Bird B, Lenau K, Mazur AI, Diem M. Spectral cytopathology: new aspects of data collection, manipulation and confounding effects. Analyst. 2013;138:3975–3982. doi: 10.1039/c3an00185g. [DOI] [PubMed] [Google Scholar]
- 20.Cao J, et al. Fourier transform infrared microspectroscopy reveals that tissue culture conditions affect the macromolecular phenotype of human embryonic stem cells. Analyst. 2013;38:4147–4160. doi: 10.1039/c3an00321c. [DOI] [PubMed] [Google Scholar]
- 21.Chan KLA, Kazarian SG. Aberration-free FTIR spectroscopic imaging of live cells in microfluidic devices. Analyst. 2013;138:4040–4047. doi: 10.1039/c3an00327b. [DOI] [PubMed] [Google Scholar]
- 22.Kallenbach-Thieltges A, et al. Immunohistochemistry, histopathology and infrared spectral histopathology of colon cancer tissue sections. J. Biophotonics. 2013;6:88–100. doi: 10.1002/jbio.201200132. [DOI] [PubMed] [Google Scholar]
- 23.Lasch P, Haensch W, Naumann D, Diem M. Imaging of colorectal adenocarcinoma using FT-IR microspectroscopy and cluster analysis. Biochim. Biophys. Acta. 2004;1688:176–186. doi: 10.1016/j.bbadis.2003.12.006. [DOI] [PubMed] [Google Scholar]
- 24.Bassan P, et al. Whole organ cross-section chemical imaging using label-free mega-mosaic FTIR microscopy. Analyst. 2013;138:7066–7069. doi: 10.1039/c3an01674a. [DOI] [PubMed] [Google Scholar]
- 25.Nakamura T, et al. Microspectroscopy of spectral biomarkers associated with human corneal stem cells. Mol. Vis. 2010;16:359–368. [PMC free article] [PubMed] [Google Scholar]
- 26.Hammiche A, German MJ, Hewitt R, Pollock HM, Martin FL. Monitoring cell cycle distributions in MCF-7 cells using near-field photothermal microspectroscopy. Biophys. J. 2005;88:3699–3706. doi: 10.1529/biophysj.104.053926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Walsh MJ, et al. Tracking the cell hierarchy in the human intestine using biochemical signatures derived by mid-infrared microspectroscopy. Stem Cell Res. 2009;3:15–27. doi: 10.1016/j.scr.2009.02.003. [DOI] [PubMed] [Google Scholar]
- 28.Wood BR, Bambery KR, Evans CJ, Quinn MA, McNaughton D. A three-dimensional multivariate image processing technique for the analysis of FTIR spectroscopic images of multiple tissue sections. BMC Med. Imaging. 2006;6:12. doi: 10.1186/1471-2342-6-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Bird B, et al. Cytology by infrared micro-spectroscopy: Automatic distinction of cell types in urinary cytology. Vib. Spectrosc. 2008;48:101–106. doi: 10.1016/j.vibspec.2008.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Baker MJ, et al. FTIR-based spectroscopic analysis in the identification of clinically aggressive prostate cancer. Br. J. Cancer. 2008;99:1859–1866. doi: 10.1038/sj.bjc.6604753. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Bird B, et al. Infrared micro-spectral imaging: distinction of tissue types in axillary lymph node histology. BMC Clin. Pathol. 2008;8:8. doi: 10.1186/1472-6890-8-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Bird B, et al. Detection of breast micro-metastases in axillary lymph nodes by infrared micro-spectral imaging. Analyst. 2009;134:1067–1076. doi: 10.1039/b821166c. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Naumann D, et al. Cells and biofluids analyzed in aqueous environment by infrared spectroscopy. Biomed. Opt. 2005;10:609301. doi: 10.1117/1.1917844. [DOI] [PubMed] [Google Scholar]
- 34.Walsh MJ, Holton SE, Kajdacsy-Balla A, Bhargava R. Attenuated total reflectance Fourier-transform infrared spectroscopic imaging for breast histopathology. Vib. Spectrosc. 2012;60:23–28. doi: 10.1016/j.vibspec.2012.01.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Ooi GJ, et al. Fourier transform infrared imaging and small angle X-ray scattering as a combined biomolecular approach to diagnosis of breast cancer. Med. Phys. 2008;35:2151–2161. doi: 10.1118/1.2890391. [DOI] [PubMed] [Google Scholar]
- 36.Nallala J, et al. Infrared imaging as a cancer diagnostic tool: Introducing a new concept of spectral barcodes for identifying molecular changes in colon tumors. Cytometry Part A. 2013;83:294–300. doi: 10.1002/cyto.a.22249. [DOI] [PubMed] [Google Scholar]
- 37.Bird B, et al. Infrared spectral histopathology (SHP): a novel diagnostic tool for the accurate classification of lung cancer. Lab. Invest. 2012;92:1358–1373. doi: 10.1038/labinvest.2012.101. [DOI] [PubMed] [Google Scholar]
- 38.Baker MJ, et al. Investigating FTIR-based histopathology for the diagnosis of prostate cancer. J. Biophotonics. 2009;2:104–113. doi: 10.1002/jbio.200810062. [DOI] [PubMed] [Google Scholar]
- 39.Gazi E, et al. A correlation of FTIR spectra derived from prostate cancer biopsies with Gleason grade and tumour stage. Eur. Urol. 2006;50:750–761. doi: 10.1016/j.eururo.2006.03.031. [DOI] [PubMed] [Google Scholar]
- 40.Walsh MJ, et al. IR microspectroscopy: potential applications in cervical cancer screening. Cancer Lett. 2007;246:1–11. doi: 10.1016/j.canlet.2006.03.019. [DOI] [PubMed] [Google Scholar]
- 41.Gajjar K, et al. Fourier-transform infrared spectroscopy coupled with a classification machine for the analysis of blood plasma or serum: a novel diagnostic approach for ovarian cancer. Analyst. 2013;138:3917–3926. doi: 10.1039/c3an36654e. [DOI] [PubMed] [Google Scholar]
- 42.Ollesch J, et al. FTIR spectroscopy of biofluids revisited: an automated approach to spectral biomarker identification. Analyst. 2013;138:4092–4102. doi: 10.1039/c3an00337j. [DOI] [PubMed] [Google Scholar]
- 43.Scaglia E, et al. Noninvasive assessment of hepatic fibrosis in patients with chronic hepatitis C using serum Fourier transform infrared spectroscopy. Anal. Bioanal. Chem. 2011;401:2919–2925. doi: 10.1007/s00216-011-5402-8. [DOI] [PubMed] [Google Scholar]
- 44.Hands J, et al. Investigating the rapid diagnosis of gliomas from serum samples using infrared spectroscopy and cytokine and angiogenesis factors. Anal. Bioanal. Chem. 2013;405:7347–7355. doi: 10.1007/s00216-013-7163-z. [DOI] [PubMed] [Google Scholar]
- 45.Munro KL, et al. Synchrotron radiation infrared microspectroscopy of arsenic-induced changes to intracellular biomolecules in live leukemia cells. Vib. Spectrosc. 2010;53:39–44. [Google Scholar]
- 46.Tobin MJ, et al. FTIR spectroscopy of single live cells in aqueous media by synchrotron IR microscopy using microfabricated sample holders. Vib. Spectrosc. 2010;53:34–38. [Google Scholar]
- 47.Whelan DR, Bambery KR, Puskar L, McNaughton D, Wood BR. Synchrotron Fourier transform infrared (FTIR) analysis of single living cells progressing through the cell cycle. Analyst. 2013;138:3891–3899. doi: 10.1039/c3an00316g. [DOI] [PubMed] [Google Scholar]
- 48.Kuimova MK, Chan KLA, Kazarian SG. Chemical imaging of live cancer cells in the natural aqueous environment. Appl. Spectrosc. 2009;63:164–171. doi: 10.1366/000370209787391969. [DOI] [PubMed] [Google Scholar]
- 49.Llabjani V, et al. Polybrominated diphenyl ether-associated alterations in cell biochemistry as determined by attenuated total reflection Fourier-transform infrared spectroscopy: a comparison with DNA-reactive and/or endocrine-disrupting agents. Environ. Sci. Technol. 2009;43:3356–3364. doi: 10.1021/es8036127. [DOI] [PubMed] [Google Scholar]
- 50.Malins DC, et al. Biomarkers signal contaminant effects on the organs of English sole (Parophrys vetulus) from Puget Sound. Environ. Health Perspect. 2006;114:823–829. doi: 10.1289/ehp.8544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Cakmak G, Togan I, Severcan F. 17β-Estradiol induced compositional, structural and functional changes in rainbow trout liver, revealed by FT-IR spectroscopy: a comparative study with nonylphenol. Aquat. Toxicol. 2006;77:53–63. doi: 10.1016/j.aquatox.2005.10.015. [DOI] [PubMed] [Google Scholar]
- 52.Llabjani V, et al. Alterations in the infrared spectral signature of avian feathers reflect potential chemical exposure: A pilot study comparing two sites in Pakistan. Environ. Int. 2012;48:39–46. doi: 10.1016/j.envint.2012.06.019. [DOI] [PubMed] [Google Scholar]
- 53.Trevisan J, et al. Syrian hamster embryo (SHE) assay (pH 6.7) coupled with infrared spectroscopy and chemometrics towards toxicological assessment. Analyst. 2010;135:3266–3272. doi: 10.1039/c0an00586j. [DOI] [PubMed] [Google Scholar]
- 54.Ahmadzai AA, et al. The Syrian hamster embryo (SHE) assay (pH 6.7): mechanisms of cell transformation and application of vibrational spectroscopy to objectively score endpoint alterations. Mutagenesis. 2012;27:257–266. doi: 10.1093/mutage/ger086. [DOI] [PubMed] [Google Scholar]
- 55.Ami D, Natalello A, Zullini A, Doglia SM. Fourier transform infrared microspectroscopy as a new tool for nematode studies. FEBS Lett. 2004;576:297–300. doi: 10.1016/j.febslet.2004.09.022. [DOI] [PubMed] [Google Scholar]
- 56.Hobro AJ, Lendl B. Fourier-transform mid-infrared FPA imaging of a complex multicellular nematode. Vib. Spectrosc. 2011;57:213–219. [Google Scholar]
- 57.Mariey L, Signolle JP, Amiel C, Travert J. Discrimination, classification, identification of microorganisms using FTIR spectroscopy and chemometrics. Vib. Spectrosc. 2001;26:151–159. [Google Scholar]
- 58.Gómez-De-Anda F, et al. Determination of Trichinella spiralis in pig muscles using mid-Fourier transform infrared spectroscopy (MID-FTIR) with attenuated total reflectance (ATR) and soft independent modeling of class analogy (SIMCA) Meat Sci. 2012;91:240–246. doi: 10.1016/j.meatsci.2012.01.019. [DOI] [PubMed] [Google Scholar]
- 59.Kazarian SG, Chan KA. ATR-FTIR spectroscopic imaging: recent advances and applications to biological systems. Analyst. 2013;138:1940–1951. doi: 10.1039/c3an36865c. [DOI] [PubMed] [Google Scholar]
- 60.Glassford SE, Byrne B, Kazarian SG. Recent applications of ATR FTIR spectroscopy and imaging to proteins. Biochim. Biophys. Acta. 2013;1834:2849–2858. doi: 10.1016/j.bbapap.2013.07.015. [DOI] [PubMed] [Google Scholar]
- 61.Stuart B. Infrared Spectroscopy: Fundamentals and Applications. John Wiley and Sons; 2005. [Google Scholar]
- 62.Mantsch HH, Chapman D. Infrared Spectroscopy of Biomolecules. Wiley-Liss; 1996. [Google Scholar]
- 63.Miller LM, Dumas P. Chemical imaging of biological tissue with synchrotron infrared light. Biochim. Biophys. Acta. 2006;1758:846–857. doi: 10.1016/j.bbamem.2006.04.010. [DOI] [PubMed] [Google Scholar]
- 64.Liu J-N, Schulmerich MV, Bhargava R, Cunningham BT. Optimally designed narrowband guided-mode resonance reflectance filters for mid-infrared spectroscopy. Opt. Express. 2011;19:24182–24197. doi: 10.1364/OE.19.024182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Miller LM, Smith RJ. Synchrotrons versus globars, point-detectors versus focal plane arrays: Selecting the best source and detector for specific infrared microspectroscopy and imaging applications. Vib. Spectrosc. 2005;38:237–240. [Google Scholar]
- 66.Lasch P, Boese M, Pacifico A, Diem M. FT-IR spectroscopic investigations of single cells on the subcellular level. Vib. Spectrosc. 2002;28:147–157. [Google Scholar]
- 67.Duncan W, Williams GP. Infrared synchrotron radiation from electron storage rings. Appl. Optics. 1983;22:2914–2923. doi: 10.1364/ao.22.002914. [DOI] [PubMed] [Google Scholar]
- 68.Pijanka JK, et al. Spectroscopic signatures of single, isolated cancer cell nuclei using synchrotron infrared microscopy. Analyst. 2009;134:1176–1181. doi: 10.1039/b821112d. [DOI] [PubMed] [Google Scholar]
- 69.Dumas P, Sockalingum GD, Sule-Suso J. Adding synchrotron radiation to infrared microspectroscopy: what’s new in biomedical applications? Trends Biotechnol. 2007;25:40–44. doi: 10.1016/j.tibtech.2006.11.002. [DOI] [PubMed] [Google Scholar]
- 70.Martin FL. Shining a new light into molecular workings. Nat. Methods. 2011;8:385–387. doi: 10.1038/nmeth.1594. [DOI] [PubMed] [Google Scholar]
- 71.Menzel L, et al. Spectroscopic detection of biological NO with a quantum cascade laser. Appl. Phys. B. 2001;72:859–863. doi: 10.1007/s003400100562. [DOI] [PubMed] [Google Scholar]
- 72.Valle JJ, et al. Free electron laser-Fourier transform ion cyclotron resonance mass spectrometry facility for obtaining infrared multiphoton dissociation spectra of gaseous ions. Rev. Sci. Instrum. 2005;76:023103. [Google Scholar]
- 73.Llabjani V, et al. Differential effects in mammalian cells induced by chemical mixtures in environmental biota as profiled using infrared spectroscopy. Environ. Sci. Technol. 2011;45:10706–10712. doi: 10.1021/es202574b. [DOI] [PubMed] [Google Scholar]
- 74.Schubert JM, Mazur AI, Bird B, Miljkovi M, Diem M. Single point vs. mapping approach for spectral cytopathology (SCP) J. Biophotonics. 2010;3:588–596. doi: 10.1002/jbio.201000023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Carter EA, Tam KK, Armstrong RS, Lay PA. Vibrational spectroscopic mapping and imaging of tissues and cells. Biophys. Rev. 2009;1:95–103. doi: 10.1007/s12551-009-0012-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Nasse MJ, et al. High-resolution Fourier-transform infrared chemical imaging with multiple synchrotron beams. Nat. Methods. 2011;8:413–416. doi: 10.1038/nmeth.1585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Fernandez DC, Bhargava R, Hewitt SM, Levin IW. Infrared spectroscopic imaging for histopathologic recognition. Nat. Biotechnol. 2005;23:469–474. doi: 10.1038/nbt1080. [DOI] [PubMed] [Google Scholar]
- 78.Chan K, Kazarian S. New opportunities in micro-and macro-attenuated total reflection infrared spectroscopic imaging: spatial resolution and sampling versatility. Appl. Spectrosc. 2003;57:381–389. doi: 10.1366/00037020360625907. [DOI] [PubMed] [Google Scholar]
- 79.Holton SE, Walsh MJ, Bhargava R. Subcellular localization of early biochemical transformations in cancer-activated fibroblasts using infrared spectroscopic imaging. Analyst. 2011;136:2953–2958. doi: 10.1039/c1an15112f. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Bassan P, et al. The inherent problem of transflection-mode infrared spectroscopic microscopy and the ramifications for biomedical single point and imaging applications. Analyst. 2013;138:144–157. doi: 10.1039/c2an36090j. [DOI] [PubMed] [Google Scholar]
- 81.Goormaghtigh E, Raussens V, Ruysschaert JM. Attenuated total reflection infrared spectroscopy of proteins and lipids in biological membranes. Biochim. Biophys. Acta. 1999;1422:105–185. doi: 10.1016/s0304-4157(99)00004-0. [DOI] [PubMed] [Google Scholar]
- 82.Wehbe K, Filik J, Frogley MD, Cinque G. The effect of optical substrates on micro-FTIR analysis of single mammalian cells. Anal. Bioanal. Chem. 2013;405:1311–1324. doi: 10.1007/s00216-012-6521-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Vaccari L, et al. Synchrotron radiation infrared microspectroscopy of single living cells in microfluidic devices: advantages, disadvantages and future perspectives. J. Phys.: Conf. Ser. 2012;359:012007. [Google Scholar]
- 84.Chan KLA, Kazarian SG. FT-IR spectroscopic imaging of reactions in multiphase flow in microfluidic channels. Anal. Chem. 2012;84:4052–4056. doi: 10.1021/ac300019m. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Marcsisin EJ, Uttero CM, Miljkovic M, Diem M. Infrared microspectroscopy of live cells in aqueous media. Analyst. 2010;135:3227–3232. doi: 10.1039/c0an00548g. [DOI] [PubMed] [Google Scholar]
- 86.Nasse M, Ratti S, Giordano M, Hirschmugl C. Demountable liquid/flow cell for in vivo infrared microspectroscopy of biological specimens. Appl. Spectrosc. 2009;63:1181–1186. doi: 10.1366/000370209789553101. [DOI] [PubMed] [Google Scholar]
- 87.Holman H-YN, Bechtel HA, Hao Z, Martin MC. Synchrotron IR spectromicroscopy: chemistry of living cells. Anal. Chem. 2010;82:8757–8765. doi: 10.1021/ac100991d. [DOI] [PubMed] [Google Scholar]
- 88.Bhargava R, Levin IW. Fourier transform infrared imaging: theory and practice. Anal. Chem. 2001;73:5157–5167. doi: 10.1021/ac010380m. [DOI] [PubMed] [Google Scholar]
- 89.Bhargava R, Levin IW. Effective time averaging of multiplexed measurements: A critical analysis. Anal. Chem. 2002;74:1429–1435. doi: 10.1021/ac011153n. [DOI] [PubMed] [Google Scholar]
- 90.Bhargava R, Ribar T, Koenig JL. Towards faster FT-IR imaging by reducing noise. Appl. Spectrosc. 1999;53:1313–1322. [Google Scholar]
- 91.Bhargava R, Schaeberle MD, Fernandez DC, Levin IW. Novel route to faster Fourier transform infrared spectroscopic imaging. Appl. Spectrosc. 2001;55:1079–1084. [Google Scholar]
- 92.Gazi E, et al. Fixation protocols for subcellular imaging by synchrotron-based Fourier transform infrared microspectroscopy. Biopolymers. 2005;77:18–30. doi: 10.1002/bip.20167. [DOI] [PubMed] [Google Scholar]
- 93.Bretzlaff R, Bahder T. Apodization effects in Fourier transform infrared difference spectra. Rev. Phys. Appl. 1986;21:833–844. [Google Scholar]
- 94.Tahtouh M, Despland P, Shimmon R, Kalman JR, Reedy BJ. The application of infrared chemical imaging to the detection and enhancement of latent fingerprints: method optimization and further findings. J. Forensic Sci. 2007;52:1089–1096. doi: 10.1111/j.1556-4029.2007.00517.x. [DOI] [PubMed] [Google Scholar]
- 95.Lasch P, Naumann D. Spatial resolution in infrared microspectroscopic imaging of tissues. Biochim. Biophys. Acta. 2006;1758:814–829. doi: 10.1016/j.bbamem.2006.06.008. [DOI] [PubMed] [Google Scholar]
- 96.Sun D-W. Infrared Spectroscopy for Food Quality Analysis and Control. Academic Press; 2009. [Google Scholar]
- 97.Snook RD, Harvey TJ, Faria EC, Gardner P. Raman tweezers and their application to the study of singly trapped eukaryotic cells. Integr. Biol. 2009;1:43–52. doi: 10.1039/b815253e. [DOI] [PubMed] [Google Scholar]
- 98.Bhargava R, Fernandez DC, Schaeberle MD, Levin IW. Effect of focal plane array cold shield aperture size on Fourier transform infrared micro-imaging spectrometer performance. Appl. Spectrosc. 2000;54:1743–1750. [Google Scholar]
- 99.Bruun SW, et al. Correcting attenuated total reflection-Fourier transform infrared spectra for water vapor and carbon dioxide. Appl. Spectrosc. 2006;60:1029–1039. doi: 10.1366/000370206778397371. [DOI] [PubMed] [Google Scholar]
- 100.Lasch P, Petrich W. Data acquisition and analysis in biomedical vibrational spectroscopy. Biomed. Appl. Sync. Infrared Microspec. 2011;11:192–225. [Google Scholar]
- 101.Trevisan J, Angelov PP, Carmichael PL, Scott AD, Martin FL. Extracting biological information with computational analysis of Fourier-transform infrared (FTIR) biospectroscopy datasets: current practices to future perspectives. Analyst. 2012;137:3202–3215. doi: 10.1039/c2an16300d. [DOI] [PubMed] [Google Scholar]
- 102.Martin FL, et al. Distinguishing cell types or populations based on the computational analysis of their infrared spectra. Nat. Protoc. 2010;5:1748–1760. doi: 10.1038/nprot.2010.133. [DOI] [PubMed] [Google Scholar]
- 103.Duda RO, Hart PE, Stork DG. Pattern Classification. 2nd edn. Wiley Interscience; 2001. [Google Scholar]
- 104.Goodacre R. Explanatory analysis of spectroscopic data using machine learning of simple, interpretable rules. Vib. Spectrosc. 2003;32:33–45. [Google Scholar]
- 105.Naumann D. FTIR spectroscopy of microorganisms at the Robert Koch Institute: experiences gained during successful project; Proc. SPIE; 2008; p. 6853. Biomedical Optical Spectroscopy 68530G. [Google Scholar]
- 106.Lasch P, Petrich W. In: Biomedical Applications of Synchrotron Infrared Microspectroscopy: a Practical Approach, RSC Analytical Spectroscopy Series. Moss D, editor. Vol. 11. 2011. pp. 192–225. RSC Analytical Spectroscopy Series. [Google Scholar]
- 107.Lasch P. Spectral pre-processing for biomedicalvibrational spectroscopy and microspectroscopicimaging. Chemom. Intell. Lab. Syst. 2013;117:100–114. [Google Scholar]
- 108.Bhargava R, Wang S-Q, Koenig JL. Route to higher fidelity FT-IR imaging. Appl. Spectrosc. 2000;54:486–495. [Google Scholar]
- 109.Reddy RK, Bhargava R. Accurate histopathology from low signal-to-noise ratio spectroscopic imaging data. Analyst. 2010;135:2818–2825. doi: 10.1039/c0an00350f. [DOI] [PubMed] [Google Scholar]
- 110.van Dijk T, Mayerich D, Bhargava R, Carney PS. Rapid spectral-domain localization. Opt. Express. 2013;21:12822–12830. doi: 10.1364/OE.21.012822. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.van Dijk T, Mayerich D, Carney PS, Bhargava R. Recovery of absorption spectra from Fourier transform infrared (FT-IR) microspectroscopic measurements of intact spheres. Appl. Spectrosc. 2013;67:546–552. doi: 10.1366/12-06847. [DOI] [PubMed] [Google Scholar]
- 112.Martens H, Stark E. Extended multiplicative signal correction and spectral interference subtraction - new pre-processing methods for near-infrared spectroscopy. J. Pharmaceut. Biomed. 1991;9:625–635. doi: 10.1016/0731-7085(91)80188-f. [DOI] [PubMed] [Google Scholar]
- 113.Bassan P, et al. Resonant Mie scattering in infrared spectroscopy of biological materials - understanding the ‘dispersion artefact’. Analyst. 2009;134:1586–1593. doi: 10.1039/b904808a. [DOI] [PubMed] [Google Scholar]
- 114.Bassan P, et al. Resonant Mie scattering (RMieS) correction of infrared spectra from highly scattering biological samples. Analyst. 2010;135:268–277. doi: 10.1039/b921056c. [DOI] [PubMed] [Google Scholar]
- 115.Bassan P, et al. RMieS-EMSC correction for infrared spectra of biological cells: extension using full Mie theory and GPU computing. J. Biophotonics. 2010;3:609–620. doi: 10.1002/jbio.201000036. [DOI] [PubMed] [Google Scholar]
- 116.Opus 5 Reference Manual. Bruker Optik; 2004. [Google Scholar]
- 117.Ly E, et al. Combination of FTIR spectral imaging and chemometrics for tumour detection from paraffin-embedded biopsies. Analyst. 2008;133:197–205. doi: 10.1039/b715924b. [DOI] [PubMed] [Google Scholar]
- 118.Beier BD, Berger AJ. Method for automated background subtraction from Raman spectra containing known contaminants. Analyst. 2009;134:1198–1202. doi: 10.1039/b821856k. [DOI] [PubMed] [Google Scholar]
- 119.Baker M, et al. An investigation of the RWPE prostate derived family of cell lines using FTIR spectroscopy. Analyst. 2010;135:887–894. doi: 10.1039/b920385k. [DOI] [PubMed] [Google Scholar]
- 120.Guyon I, Gunn S, Nikravesh M, Zadeh L. Feature Extraction, Foundations and Applications. Springer; 2006. [Google Scholar]
- 121.Udelhoven T, Novozhilov M, Schmitt J. The NeuroDeveloper (R): a tool for modular neural classification of spectroscopic data. Chemometr. Intell. Lab. 2003;66:219–226. [Google Scholar]
- 122.Kwak JT, Reddy R, Sinha S, Bhargava R. Analysis of variance in spectroscopic imaging data from human tissues. Anal. Chem. 2011;84:1063–1069. doi: 10.1021/ac2026496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123.Bhargava R, Fernandez DC, Hewitt SM, Levin IW. High throughput assessment of cells and tissues: Bayesian classification of spectral metrics from infrared vibrational spectroscopic imaging data. Biochim. Biophys. Acta. 2006;1758:830–845. doi: 10.1016/j.bbamem.2006.05.007. [DOI] [PubMed] [Google Scholar]
- 124.Naumann D. Encyclopedia of Analytical Chemistry. John Wiley & Sons; 2000. [Google Scholar]
- 125.Hughes C, et al. FTIR microspectroscopy of selected rare diverse sub-variants of carcinoma of the urinary bladder. J. Biophotonics. 2013;6:73–87. doi: 10.1002/jbio.201200126. [DOI] [PubMed] [Google Scholar]
- 126.Hastie T, Tibshirani R, Friedman J. The Elements of Statistical Learning: Data Mining, Inference and Prediction. 2nd edn. Springer; 2009. [Google Scholar]
- 127.Bhargava R. Towards a practical Fourier transform infrared chemical imaging protocol for cancer histopathology. Anal. Bioanal. Chem. 2007;389:1155–1169. doi: 10.1007/s00216-007-1511-9. [DOI] [PubMed] [Google Scholar]
- 128.Berenbaum MC. The histochemistry of bound lipids. Q. J. Microsc. Sci. 1958;s3-99:231–242. [Google Scholar]
- 129.Wigglesworth VB. Bound lipid in the tissues of mammal and insect: a new histochemical method. J. Cell Sci. 1971;8:709–725. doi: 10.1242/jcs.8.3.709. [DOI] [PubMed] [Google Scholar]
- 130.Stitt DM, et al. Tissue acquisition and storage associated oxidation considerations for FTIR microspectroscopic imaging of polyunsaturated fatty acids. Vib. Spectrosc. 2012;60:16–22. [Google Scholar]
- 131.Owens GL, et al. Vibrational biospectroscopy coupled with multivariate analysis extracts potentially diagnostic features in blood plasma/serum of ovarian cancer patients. J. Biophotonics. 2014;7:200–209. doi: 10.1002/jbio.201300157. [DOI] [PubMed] [Google Scholar]
- 132.Dorling KM, Baker MJ. Highlighting attenuated total reflection Fourier transform infrared spectroscopy for rapid serum analysis. Trends Biotechnol. 2013;31:327–328. doi: 10.1016/j.tibtech.2013.03.010. [DOI] [PubMed] [Google Scholar]
- 133.Chang C-C, Lin C-J. LIBSVM: a library for support vector machines. ACM TIST. 2011;2:27. [Google Scholar]
- 134.Trevisan J, et al. Measuring similarity and improving stability in biomarker identification methods applied to Fourier-transform infrared (FTIR) spectroscopy. J. Biophotonics. 2014;7:254–265. doi: 10.1002/jbio.201300190. [DOI] [PubMed] [Google Scholar]
- 135.Lasch P, Fabian H, Thi NAN, Naumann D. Infrarot-bildgebung für die pathohistologische diagnostik. Laborwelt. 2004;2:8–12. [Google Scholar]
- 136.Lasch P, Kneipp J. Biomedical Vibrational Spectroscopy. Wiley-Blackwell; 2008. [Google Scholar]
- 137.Lyng F, Gazi E, Gardner P. In: Preparation of Tissues and Cells for Infrared and Raman Spectroscopy. Moss D, editor. Royal Society of Chemistry; 2011. pp. 147–185. RSC Analytical Spectroscopy Monographs, No. 11. [Google Scholar]
- 138.Whelan DR, Bambery KR, Puskar L, McNaughton D, Wood BR. Quantification of DNA in simple eukaryotic cells using Fourier transform infrared spectroscopy. J. Biophotonics. 2013;6:775–784. doi: 10.1002/jbio.201200112. [DOI] [PubMed] [Google Scholar]
- 139.Whelan DR, et al. Monitoring the reversible B- to A-like transition of DNA in eukaryotic cells using Fourier transform infrared spectroscopy. Nucleic Acids Res. 2011;39:5439–5448. doi: 10.1093/nar/gkr175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140.Rahmelow K, Hubner W. Phase correction in Fourier transform spectroscopy: subsequent displacement correction and error limit. Appl. Optics. 1997;36:6678–6686. doi: 10.1364/ao.36.006678. [DOI] [PubMed] [Google Scholar]
- 141.Gazi E, et al. The combined application of FTIR microspectroscopy and ToF-SIMS imaging in the study of prostate cancer. Faraday Discuss. 2004;126:41–59. doi: 10.1039/b304883g. [DOI] [PubMed] [Google Scholar]
- 142.Patel II, et al. Isolating stem cells in the inter-follicular epidermis employing synchrotron radiation-based Fourier-transform infrared microspectroscopy and focal plane array imaging. Anal. Bioanal. Chem. 2012;404:1745–1758. doi: 10.1007/s00216-012-6314-y. [DOI] [PubMed] [Google Scholar]
- 143.Bassan P, et al. FTIR microscopy of biological cells and tissue: data analysis using resonant Mie scattering (RMieS) EMSC algorithm. Analyst. 2012;137:1370–1377. doi: 10.1039/c2an16088a. [DOI] [PubMed] [Google Scholar]
- 144.Kastyak-Ibrahim M, et al. Biochemical label-free tissue imaging with subcellular-resolution synchrotron FTIR with focal plane array detector. NeuroImage. 2012;60:376–383. doi: 10.1016/j.neuroimage.2011.11.069. [DOI] [PubMed] [Google Scholar]
- 145.Gajjar K, et al. Diagnostic segregation of human brain tumours using Fourier-transform infrared and/or Raman spectroscopy coupled with discriminant analysis. Anal. Methods. 2013;5:89–102. doi: 10.1039/C2AY25544H. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146.Kelly JG, Martin-Hirsch PL, Martin FL. Discrimination of base differences in oligonucleotides using mid-infrared spectroscopy and multivariate analysis. Anal. Chem. 2009;81:5314–5319. doi: 10.1021/ac900546m. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.