Abstract
Objective:
The complex molecular etiology of psychosis in schizophrenia (SZ) and psychotic bipolar disorder (PBP) is not well defined, presumably due to their multifactorial genetic architecture. Neurobiological correlates of psychosis can be identified through genetic associations of intermediate phenotypes such as event-related potential (ERP) from auditory paired stimulus processing (APSP). Various ERP components of APSP are heritable and aberrant in SZ, PBP and their relatives, but their multivariate genetic factors are less explored.
Methods:
We investigated the multivariate polygenic association of ERP from 64-sensor auditory paired stimulus data in 149 SZ, 209 PBP probands, and 99 healthy individuals from the multisite Bipolar-Schizophrenia Network on Intermediate Phenotypes study. Multivariate association of 64-channel APSP waveforms with a subset of 16 999 single nucleotide polymorphisms (SNPs) (reduced from 1 million SNP array) was examined using parallel independent component analysis (Para-ICA). Biological pathways associated with the genes were assessed using enrichment-based analysis tools.
Results:
Para-ICA identified 2 ERP components, of which one was significantly correlated with a genetic network comprising multiple linearly coupled gene variants that explained ~4% of the ERP phenotype variance. Enrichment analysis revealed epidermal growth factor, endocannabinoid signaling, glutamatergic synapse and maltohexaose transport associated with P2 component of the N1-P2 ERP waveform. This ERP component also showed deficits in SZ and PBP.
Conclusions:
Aberrant P2 component in psychosis was associated with gene networks regulating several fundamental biologic functions, either general or specific to nervous system development. The pathways and processes underlying the gene clusters play a crucial role in brain function, plausibly implicated in psychosis.
Key words: schizophrenia, bipolar disorder, psychosis, single nucleotide polymorphism, gene, pathway, event-related potential
Introduction
Growing evidence supports common genetic mechanisms underlying the psychosis dimension in schizophrenia (SZ), schizoaffective (SZA), and psychotic bipolar disorder (PBP).1–3 Numerous genetic markers have been implicated in the etiology of these disorders,4 yet none of them prove to have a high penetrance,5 consistent with a polygenic common disease common variant (CDCV) model comprising multiple interacting genes contributing to a cumulative effect, distinct from the weak individual gene effect in disease risk or association.6 Further, the underlying pathways and mechanisms of the genes mediating the neurophysiological abnormalities and clinical symptoms are not well ascertained,7,8 and their polygenic effects are relatively unstudied. An effective strategy for gene and disease target discovery underlying complex neuropsychiatric diseases is to employ endophenotypes that are heritable state-independent biological measures associated with illness due to shared genetic influences. The concept of endophenotype in psychiatric genetics has elicited increased attention in gene characterization due to the presumed tractable genetic framework and close proximity to the level of gene action,9 as opposed to the complex clinical disease.
One such endophenotype is the event-related potential (ERP) measured via noninvasive paired click auditory sensory processing (auditory paired stimulus processing [APSP]) paradigm. Auditory sensory gating is assessed with the paired stimulus paradigm, involving presentation of 2 (S1, S2) closely separated identical sounds (500ms inter-stimulus interval), with long intervals (6–10 s) between stimulus pairs.10,11 The earliest component of the mid-latency ERPs is the P50, a positive wave peaking at ~50ms (between 35 and 85ms) following presentation of each stimulus.12 Aberrant P50 gating is a key biomarker marker of SZ and PBP13–15 and also abnormal in unaffected relatives.16 Informative measurements in paired-stimuli paradigms are not limited to P50 peaks. ERPs derived from APSP paradigms also comprise complex cascade of mid/late latency including N1, the negative voltage ERP peaking approximately 100ms and P2, the positive ERP peaking approximately 200ms after S1 onset and sustained deflections that carry unique, substantial, and complementary elements depicting SZ and PBP pathophysiology.10,17,18 The N1 component is also compromised in unaffected SZ relatives.19 P50, N1, and P2 components are associated with various attentive stages of information processing.20,21 Less traditional ERP measurements from APSP, such as a pre-S2 negativity, have been shown to capture heritable disease heterogeneity in a large sample of SZ and PBP proband and first-degree relatives.22 That study did not include SZA probands, thus distinctions between SZ and PBP from SZA probands were not investigated. Despite the lack of clear pathophysiological boundary between SZ and SZA, and the clinical heterogeneity manifested in SZA disorder (includes depressive and manic subtypes), an earlier study examined auditory P300 ERP in both these probands and reported that SZA did not exhibit P300 amplitude deficits observed in SZ.23 A recent report24 showed that P300 feature of response inhibition did not differentiate SZA from SZ and controls. Additionally, a previous study employing electrophysiological indices demonstrated lack of diagnostic specificity in comparing SZA to SZ and PBP but revealed that such measures strongly differentiated SZ subtypes.25 A detailed review of electrophysiological studies comparing SZA to other controls and probands can be found in ref.26. The inclusion of SZA probands in either the SZ or PBP group has been debated due to clinical heterogeneity and overlapping symptoms.26 The current study analyzed the entire ERP waveform derived from APSP including all 64 sensors across the entire scalp. Such a strategy offers a comprehensive representation of the biomarkers compared to a single biomarker based genetic studies. In this study, we empirically assessed time-voltage ERP from subset of probands in conjunction with genetic data, different from prior study22 examining both time-voltage and time-frequency activity across probands and relatives. Although significant familiality (h 2 ≅ 0.2–0.5) was noted in that study no unaffected relatives’ effects were present.
Although few reports indicate P50 impairments to be associated with dysfunctions in various neurotransmitter systems,27–29 the neurobiological correlates of other APSP components are known and not well characterized. As an endophenotype, heritability of APSP components has been substantiated by several twin studies.19,30–32 Genetic linkage studies have proposed various risk loci and candidate genes for P50 abnormalities in SZ and PBP including the 15q13-14 (the site of alpha-7 nicotinic acetylcholine receptor gene CHRNA7),33,34 22q11-q12,35 18q21,36 COMT,37 and CACNA1C.38 However, these findings have not been consistently replicated.39 One prior study examined genetic associations of N1 and P2 ERP components but yielded no significant linkage.40 To our knowledge, no prior studies have extensively evaluated the genetic correlates of the N1 and P2 components in psychosis.
Genome-wide association (GWA) of single nucleotide polymorphisms (SNPs) or pedigree-based linkage analysis is the primary tool for identifying complex traits in most genetic studies. The univariate strategy of GWA examines a SNP with single phenotype of interest. Thus, failing to account for the multifactorial model,41 where linear and epistatic interaction between multiple SNPs confers risk for complex psychiatric diseases.42 For instance, because most of the common SNP variants usually confer small increments in risk of the disease, they often do not reach genome-wide significance due to stringent multiple comparison correction for numerous SNP variants in the GWA model. Thus, large study samples are required to achieve genome wide significance. To mitigate these problems and improve the predictive power of complex phenotypes in moderate sized samples, novel statistically efficient data-driven multivariate association approaches based on parallel independent component analysis (Para-ICA)43,44 are employed to model the presumed polygenic architecture of complex psychiatric illnesses.45 Such methods have been successfully used to identify gene correlates implicated in Alzheimer’s disease46 and psychoses.47–49 In multivariate model, the aggregate constituent from linearly coupled gene variants is linked (linear additive model) to complex phenotypes including multiple biological measures (as opposed to single feature in GWA). Under the multifactorial assumption, several gene variants mediate psychotic disorders in concordance with the notion that multiple complex neurobiological mechanisms underlie the disease etiology.
The main goals in this study are to (1) identify prior psychosis candidate genes and novel genes associated with auditory processing abnormalities in psychosis. (2) Examine gene networks and associated molecular and biological attributes collectively influencing distinctive auditory processing features in SZ and PBP. In line with prior evidence, we hypothesized that ERP abnormalities in APSP would be influenced by genes involved in axon guidance and synaptic transmission known to be associated with psychosis.
Materials and Methods
Bipolar-Schizophrenia Network on Intermediate Phenotypes Study
The genotype–phenotype multivariate association was evaluated using multimodal data fusion (integration of data from multiple modalities to leverage the joint or combined information for better characterization of brain activity) of SNP variants and ERP data from APSP pooled across SZ, PBP, and healthy comparison (HC) individuals from the Bipolar-Schizophrenia Network on Intermediate Phenotypes (BSNIP) study.
Subject Recruitment
We examined 457 subjects including 99 HC, 149 SZ, and 209 PBP probands. Study participants in this sample were selected such that each had both genotype data (N = 620 subjects inclusive of SZ, SZA and PBP probands and HC only) and had participated in APSP paradigm (N ≅ 1600 subjects comprising all probands, relatives and HC). A subset of these APSP ERP data (N ≅ 1100) were previously published elsewhere.22 Thus the sample comprised an intersection of both subsets (N = 620 and 1600 subjects), which were part of the overall 2441-person cohort from the BSNIP study (refer to ref.50 for complete details including participant recruitment and exclusion criteria). The subjects were aged 15–65 years and recruited (see supplementary material) from the 5 participating BSNIP sites. Probands were on stable doses of psychotropic medication (supplementary table ST1) for ≥4 weeks. The study was explained to all subjects and institutional review board approved written informed consent was obtained at the respective participating sites. Detailed demographic characteristics and medication information for the study sample are listed in table 1 and supplementary table ST1, respectively.
Table 1.
Demographic and Clinical Characteristics of Study Sample Including Probands and Healthy Comparison Subjects
| Variable | Healthy Comparison Subjects (N = 99) | Schizophrenia (N = 149) | Psychotic Bipolar Disorder (N = 209) | P Value | |||
|---|---|---|---|---|---|---|---|
| Mean | SD | Mean | SD | Mean | SD | ||
| Age (y)a | 36.06 | 11.88 | 32.58 | 11.63 | 35.75 | 12.47 | <.02 |
| PANSS-positive | — | — | 16.2 | 5.36 | 14.3 | 5.25 | |
| PANSS-negative | — | — | 16.91 | 5.84 | 12.95 | 4.43 | |
| PANSS-general | — | — | 32.64 | 9.13 | 30.27 | 8.87 | |
| SBS | — | — | 7.66 | 1.33 | 2.03 | 1.8 | |
| N | % | N | % | N | % | ||
| Sexb | |||||||
| Male | 39 | 39 | 106 | 71 | 80 | 38 | <5e-10 |
| Female | 60 | 60 | 43 | 29 | 129 | 62 | |
| Site | |||||||
| Baltimore | 15 | 15 | 37 | 24.8 | 36 | 17.2 | <.06 |
| Bostonc | 3 | 3 | 4 | 2.7 | 1 | 0.5 | |
| Chicago | 26 | 26 | 31 | 20.8 | 65 | 31.1 | |
| Dallasc | 7 | 7 | 15 | 10.1 | 28 | 13.3 | |
| Detroit | 17 | 17 | 15 | 10.1 | 21 | 10.04 | |
| Hartford | 31 | 31 | 47 | 31.5 | 58 | 27.7 | |
| Schizoaffective disorderd | 0 | 0 | 52 | 34.9 | 136 | 65.1 | |
| Race | |||||||
| Caucasian | 57 | 57.6 | 78 | 52.3 | 137 | 65.5 | |
| African American | 26 | 26.2 | 46 | 30.8 | 42 | 20.1 | |
| Hispanic | 5 | 5.05 | 14 | 9.4 | 19 | 9.1 | |
| Asian | 7 | 7.07 | 3 | 2.01 | 2 | 0.95 | |
| Mixed | 4 | 4.04 | 8 | 5.3 | 9 | 4.3 | |
Note: EEG, electroencephalogram; PANSS, Positive and Negative Syndrome Scale; SBS, Schizo-Bipolar Scale.
aHealthy comparison subjects and bipolar probands > Schizophrenia probands.
bDisproportionate number of males in schizophrenia probands and disproportionate number of females in psychotic bipolar probands.
cThere were only 8 subjects from the Boston site and because identical EEG equipment was used to collect data at both Boston and Detroit sites which were managed by the same investigator (2 sites were merged in the study). Site was used as a factor with 5 levels in the statistical analysis.
d136 schizoaffective (SZA)-manic probands were assigned to the psychotic bipolar group and 52 SZA-depressive probands were assigned to the schizophrenia group.
APSP Paradigm and Electroencephalogram Data Recording
Electroencephalogram (EEG) data were collected at each site from identical Neuroscan equipment equipped with 64 Ag/AgCl electrodes (Quik-Cap, Compumedrics), while subjects listened to 150 binaural broadband auditory stimuli pairs (4ms duration at 75 dB), separated by an average of 9.5 seconds, with 500ms between stimuli in a pair (see supplementary material). Electrode positions were defined according to the International 10-10 EEG system, with the inclusion of mastoids and CP1/2 locations to provide greater signal sampling below the cantho-meatal line. Nose served as a reference with a forehead ground. Electrode impedances were maintained below 5 KΩ throughout the procedure. EEG recordings were amplified with a gain of 12 500 and digitized at a sampling rate of 1000 Hz, using Neuroscan Acquire and SynAmps2 recording systems (Compumedics Neuroscan).
SNP Data Collection
DNA was extracted using blood samples collected from participants. Approximately 1 million (1 140 419) target SNP variants across the whole genome were genotyped using the Illumina Human Omni1-Quad chip and BeadArray platform at Genomas Inc, Hartford Hospital.
ERP Data Processing
Raw EEG data were inspected and adjusted (see supplementary figure SF1) for bad electrodes and artifacts (<5% for any subject), using spherical spline interpolation (BESA 5.3; MEGIS Software). Data were converted to an average reference and digitally band-pass filtered from 0.5–55 Hz (zero phase filter; rolloff: 6 and 48 dB/octave, respectively). The current filter settings are suitable for deriving the N1 and P2 ERP components, while they were suboptimal to extract the P50 component.51 Refer to supplementary material for additional details on ERP data processing.
SNP Data Processing
Genotyped SNP data typically represented as homozygous (AA, BB) and heterozygous (AB or BA) were converted to numerical format by additive coding for the number of minor alleles (AA = 0, AB = 1, and BB = 2, assuming B is minor allele). SNP data were subjected to quality control process outlined in ref.52 using Plink (see supplementary figure SF1 and supplementary text for processing details) and stratification bias (see Q-Q plot in supplementary figure SF2) was corrected using principal component analysis (PCA)53 (using custom Matlab scripts). Data were reduced from a ~106 to 16 999 SNPs by selecting those significant at P < .05 in either of the proband (SZ, PBP) vs control logistic regression.
Para-ICA-Based SNP-ERP Association
Data were constructed as a matrix of (1) subjects by SNPs (457×16 999), (2) subjects by ERP waveforms (457×80 000 [64 channels × 1250 time points of ERP data]) and were reduced to 12 and 2 hidden components or factors, respectively (see supplementary text). ERP and SNP data were simultaneously processed by Para-ICA43,44 (see supplementary figure SF3). The original ERP data,22 were processed using principal component analysis (see supplementary text for PCA vs ICA differences). A leave-one-out cross validation was conducted using the same parameters used in the original run to assess the reliability of SNP and ERP components.
Correlation and Statistical Analysis
The strength of association was evaluated using correlations between SNP and ERP loading coefficients (LC). The effects of confounding demographic factors including age, sex and site on LCs were accounted using partial-correlation. Correlations for all component pair combinations (2×12 = 24) were evaluated and significance levels adjusted using Bonferroni multiple comparison correction for number of combinations (P < .05/24). Supplementary analyses including partial-correlation accounting for group variations, multiple regression analysis and correlation with symptoms are presented in the supplementary material. ERP and SNP LCs of significant component pairs were examined between groups using pairwise t tests. The SNPs and ERP features in each component contributing to the association were identified using a threshold of |Z| = 2.5.
Pathway Analysis
Genes representing SNPs selected from the genetic network were entered into the GeneGo MetaCore software, version 6.17 (https://portal.genego.com) and the ConsensusPathDB database (CPDB), release 28 (http://consensuspathdb.org)54 (includes KEGG and Reactome resources from CPDB) to examine the biological pathways/mechanisms underlying sensory gating abnormalities (see supplementary material).
Results
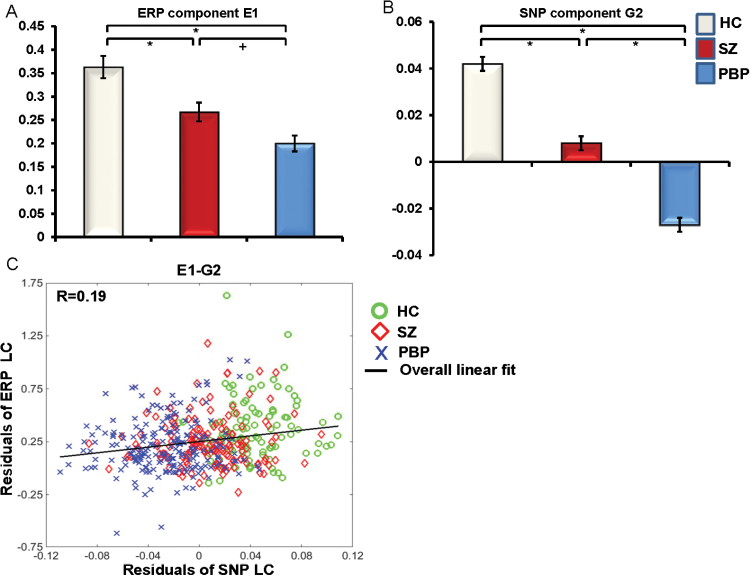
We identified 1 significant gene-ERP linkage. Of the 2 ERP components (E1-E2) derived from Para-ICA (shown in supplementary figure SF4), E1 was significantly correlated with G2 (r = .19, P = .00004, N = 457) (after multiple comparison correction). The linkage between ERP and SNP LC was assessed using partial-correlation after adjusting for the effects of demographic variables (see supplementary table ST2 for their effects on LC). The association between E1-G2 was significant when both proband groups and HC were combined as a single group (figure 1); however no significant association was observed within each group (SZ, PBP, and HC) separately (see supplementary figure SF5), likely due to constrained LC variability (LC range) within each diagnostic group. The component pair E2-G3 did not yield a significant linkage after Bonferroni adjustment (r = −.09, P = .07, N = 457). Refer to supplementary material for results from the partial-correlation after adjusting for group variations (supplementary figure SF6) and multiple regression analysis.
Fig. 1.
Mean loading coefficients (LC) for significantly associated component pair (A) ERP (B) gene network. (C) Scatter plot of LC for all subjects (including schizophrenia, bipolar disorder, and healthy comparison) combined as a single group. The ERP and SNP LC denote each subject’s contribution to the component waveform and the minor allele frequency from the SNP variants, respectively. The LC was adjusted for effects of demographics variables including sex, site, and age. Error bars represent standard errors. Post hoc comparisons included pairwise t tests between groups. The overall association between the LCs across 3 groups is displayed as the linear fit. *Groups significantly differed at P < .001. +trend at P < .1. ERP = event-related potential; SNP = single nucleotide polymorphism.
E1 and E2 comprised N1-P2 wave complex with a dominant P2 and secondary N1 feature, respectively. The ERP components are plotted at peak location Fcz and the topography for E1 and E2 was mapped using the peak amplitude in the P2 (150–250ms) and N1 time range (75–130ms) in response to stimulus S1, respectively. The topographies for both components revealed a fronto-central distribution. E1 was positively correlated with G2 (see figure 1 for scatter plot), reflecting increased linear combination of minor allelic variation related to increased E1, respectively. The genetic network G2 comprised 281 SNPs/170 unique genes. The top 20 genes from the genetic network G2, their functional properties and their known associations with neuropsychiatric diseases are summarized in table 2 and supplementary table ST3, respectively. No significant association was found between ERP LC and symptom measures including Positive and Negative Syndrome Scale (PANSS) positive (r = −.02, P = .6), PANSS negative (r = −.027, P = .6), PANSS general (r = .037, P = .48), MADRS (r = −.015, P = .77), Young mania (r = −.07, P = .18), and SBS ratings (r = .035, P = .51). Group comparisons on ERP and SNP LC revealed significant differences between HC and both SZ and PBP proband groups for E1-G2 pair (see figure 1 and supplementary figure SF7). A P2 (response to S1) deficit was observed in both proband groups (HC > SZ > PBP) compared to HC. Further, pairwise comparison revealed a trend toward greater P2 deficit in PBP compared to SZ probands [t(1.69), P < .08; see figure 1]. Reliability evaluated using leave-one-out cross-validation revealed an average within modality correlation > 0.9, indicating stable ERP and SNP components.
Table 2.
Individual Gene Information and Function for Top 20 Unique Genes in Genetic Network (G2) Obtained From Multivariate Association Analysis
| Gene G2 | SNP | Chr | Position | Z score | RW | Functional Attribute |
|---|---|---|---|---|---|---|
| SCAIa,b | rs394750I | 9q33.3 | 127801065 | −6.8909 | 1 | Transcription corepressor activity, protein binding, cell migration and signal transduction |
| PPP6Cb | rs1048251Utr-3 | 9q33.3 | 127910498 | −6.7805 | 0.983 | Phosphatase/hydrolase activity, protein/metal ion binding and mitotic cell cycle regulation |
| RABEPKb | rs450331I | 9q33.3 | 127985834 | −6.1289 | 0.889 | Protein binding and vesicle-mediated transmembrane transport |
| HSPA5b | rs12009Utr-3 | 9q33.3 | 127997303 | −5.489 | 0.796 | Calcium ion/ATP binding, protein binding/ folding, immune response, and meiosis |
| CSGALNACT1a,b | rs13261732I | 8p21.3 | 19330215 | 5.0214 | 0.728 | Hexosyl transferase activity, metal ion binding and neuronal development |
| GOLGA1a,b | rs1058540Utr-3 | 9q33.3 | 97254160 | −4.9367 | 0.716 | Protein binding and maintaining Golgi structure |
| UGGT2 | rs2062276I | 13q32.1 | 96656144 | 4.6448 | 0.674 | Glycoprotein glucosyltransferase activity and post-translational protein modification/ folding |
| HS6ST3a,b | rs9562025I | 13q32.1 | 96753630 | 4.5491 | 0.660 | Sulfotransferase activity, heparan sulfate/ heparin synthesis and axon guidance |
| PALM2b | rs10759382I | 9q31.3 | 112634640 | −4.4345 | 0.643 | Neuronal anatomical plasticity and interaction with D3 dopamine receptors |
| CSMD1a,b | rs7816319I | 8p23.2 | 3432860 | 4.3756 | 0.634 | Protein phosphatase activity, possible role in neuronal growth cone function and complement activation in developing CNS |
| DNAJC3 | rs9561932I | 13q32.1 | 96347177 | 4.3089 | 0.625 | Protein kinase inhibitor, chaperone/protein binding |
| ASIC2a,b | rs4795769I | 17q12 | 31679192 | −4.1551 | 0.603 | Voltage-gated sodium channel, protein binding, transmembrane/potassium/sodium transport and synaptic transmission |
| ZMIZ1a,b | rs703965I | 10q22.3 | 80955067 | −3.8628 | 0.560 | Zinc ion binding, androgen receptor signaling |
| PLD5 | rs2796135I | 1q43 | 242555949 | −3.8616 | 0.560 | Catalytic activity |
| AJAP1 | rs4654583I | 1p36.32 | 4730452 | −3.8567 | 0.559 | Adherens junction, CA and cell migration |
| EPHA8 | rs649754I | 1p36.12 | 22906247 | −3.8021 | 0.551 | Protein binding, CA, axon guidance, neuronal development/remodeling and synaptic plasticity |
| NCKAP5a | rs17802295I | 2q21.2 | 134157295 | −3.801 | 0.551 | Peripheral clock protein, possible role in nuclear localization signaling |
| SHANK2a | rs10459038I | 11q13.2 | 70846961 | −3.7144 | 0.539 | GKAP, dendritic spine morphogenesis/ synaptogenesis, organizing glutamatergic postsynaptic density, dopamine-glutamate interaction |
| CDH13 | rs11860907I | 16q23.3 | 83271443 | 3.69 | 0.535 | Calcium ion/(lipo) protein/cadherin binding, CA, regulation of neuronal growth |
| PCSK5a,b | rs10781332I | 9q21.3 | 78713300 | −3.6786 | 0.533 | Endopeptidase activity, peptide/protein binding, neurotrophins processing and neuronal plasticity |
Note: ALS, amyotrophic lateral sclerosis; ATP, adenosine triphosphate; CA, cell adhesion; CNS, central nervous system; GKAP, guanylate kinase-associated protein, I, intronic; ICA, independent component analysis; Utr-3, 3 prime untranslated region. Only the single nucleotide polymorphism (SNP) with maximal loading within each gene is listed in the table, (most genes were represented by several SNPs). Relative weights (RW) indicate normalized Z scores (ZS) (absolute Z score divided by the maximum absolute SNP weight). SNP locations are based on GRCh37/hg19 assembly. Functional attributes were obtained from Dbsnp and genecards. A fraction (60%, N = 12) of the top 20 genes from the parallel ICA of the original admixture sample was present in the top 20 genes identified in a separate parallel ICA of Caucasian sample (a subset of the original data with N = 272).
aMultiple SNP occurrences (>2) of the gene within the genetic network.
bGenes\SNPs that were present in the top 20 genes\SNPs identified from the parallel ICA of the Caucasian sample.
Pathway Analysis
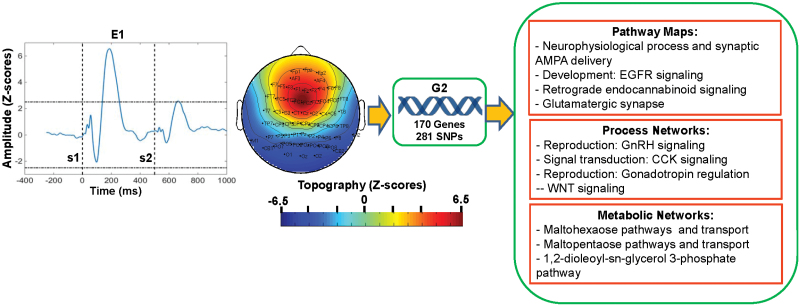
The significant pathways found in GeneGo and consensus pathway data base (CPDB) databases are listed in supplementary tables ST4 and ST5, respectively. G2 network (related to E1 network) was significantly enriched with genes belonging to retrograde endocannabinoid signaling and glutamatergic synapse (see figure 2) in CPDB. Significant pathway maps in GeneGo included epidermal growth factor receptor (EGFR), immune response and development slit-robo signaling. Several process networks associated with G2 included GnRH, cholecystokinin, gonadotropin, WNT signaling and development neurogenesis (axon guidance and synaptogenesis). Prominent metabolic networks included maltohexaose pathway and transport.
Fig. 2.
Gene networks and pathways associated with event-related potential (ERP) components from auditory paired stimulus processing obtained using multivariate data fusion analysis. Of the 2 ERP components, only 1 ERP-gene pair was significantly correlated. E1 represents a fronto-centrally distributed component (at peak location Fcz) capturing the N1-P2 complex with a dominant P2 in response to stimulus S1. The horizontal dotted line denotes the threshold of 2.5 for selecting dominant ERP feature. SNPs within each gene network were selected using a threshold of 2.5 and mapped to genes, which were in turn entered into enrichment analysis to obtain biological functions associated with gene clusters.
Discussion
The complex genetic architecture underlying psychosis in SZ and PBP is largely unknown. Etiology can be examined by probing intermediate phenotypes such as ERPs derived from APSP, a biological measure presumably closer to gene action than clinical phenomenology. Various features from APSP exhibit significant heritability in family-based studies30 and less densely related individuals,22 suggesting genetic control over ERP components. We hypothesized that multiple coordinated genes contribute to ERP abnormalities in APSP in SZ and PBP.
APSP ERP Components
We observed deficits in E1 (dominant P2 response to S1) component in SZ and PBP, consistent with previous studies.17,22 P2 is related to filtering mechanisms associated with attention allocation.20 Although the neural basis of P2 is not well characterized, P2 deficits in patients may index disruptions of sensory registration of information allocated by stimuli and problems with auditory stimulus identification and classification.55
Gene Networks
Para-ICA identified networks of synchronized genes, with each gene associated with a weight reflecting the individual contribution to the ERP-genetic linkage. It is important to note that both the P2 ERP and the related gene network showed pairwise group-differences in a similar ordinal fashion (HC > SZ > PBP), perhaps reflecting a common latent or hidden factor partly mediating the variability in both phenotype and genotype, further contributing to the diagnostic variations. This important finding revealed that a common source of influence may explain the existence of correlation between the ERP-gene networks that was influenced by diagnosis, plausibly providing insights towards search of trans-diagnostic psychosis dimensions. Variance in both ERP and genetic networks may thus influence the disease outcome. The genetic network identified in this study explained ~4% of the ERP phenotype variance, similar to other polygenic studies that have reported slightly higher explained variance than GWAS type studies1 (~2%). Thus, a majority of the phenotypic variability is still unresolved, likely due to the small contributions of each SNP toward the total variance. Additionally, environmental factors may influence the phenotype. We discuss the genetic basis of P2 abnormalities by examining the functionality of the top-ranked and repeatedly identified genes (multiple SNP occurrences), as well as the biological pathways enriched in the gene clusters.
G2
Top-Ranked and Repeatedly Identified Genes
The highest ranking gene in G2 was SCAI with 7 SNPs identified within the network. This gene encodes a protein regulating cell migration and function in the RhoA-Dia1 signal transduction pathway. The next ranking gene was PPP6C, encoding the catalytic subunit of protein phosphate regulating cell cycle progression. Although neither gene has no known specific central nervous system functions or neuropsychiatric disease association, cell cycle regulation plays a key postmitotic role in neuronal migration, axon pruning and synapse maturation.56 The CSMD1 gene (12 SNPs) encodes complement control-related protein suggested to inhibit the classical immune c3 activation,57 while also involved in neuronal wiring and altering synaptic strength.58 This gene plays an active role in immune system pathways and is associated with psychosis risk.59–61 ASIC2 encodes the degenerin/epithelial sodium channel associated with response to lithium treatment in PBP patients.62 SHANK2 encodes synaptic proteins that functions as molecular scaffolds in excitatory synapses implicated in autism risk.63 This gene is also involved in neuronal morphogenesis and regulates calcium signaling.64
Biological Pathways, Processes and Metabolic Networks
Pathway enrichment analysis and gene network ontologies are ongoing fields of development.65 Although various gene annotation databases exist, these differ in underlying statistical approaches, thus presenting challenges in interpretation, and requiring refinement. However, such techniques are essential in identifying key biological substrates associated with key endophenotypes in complex psychiatric disorders. Numerous signaling pathways comprising endocannabinoid, EGF receptor, immune response (IL-15), tubby, developmental slit-robo influenced P2 in psychosis. Retrograde endocannabinoid signaling mediates synaptic plasticity in various brain regions.66 In the hippocampus, this pathway is triggered by NMDA receptor-mediated calcium entry into neurons.67 Endocannabinoid signaling regulates hippocampal GABA release68 and glutamatergic neurotransmission through D2 dopamine receptors.69,70 EGF receptor signaling has neurotrophic effects on developing dopaminergic neurons71 playing major role in neuronal proliferation and differentiation.72 One prior study reported abnormal EGF generation in chronic SZ tissues.73 EGFR (ErbB-1) is a member of the ErbB receptor tyrosine kinase proteins, and interacts with neuregulins, including NRG1, associated with SZ susceptibility.74
Additional signaling networks including gonadotropin-releasing hormone (GnRH), cholecystokinin and WNT were enriched in G2. GnRH is highly expressed in hippocampal75 and hypothalamic pyramidal neurons76 and contributes to pathology of SZ.77 Hypothalamic-pituitary-gonadal axis dysfunction is linked to affective arousal deficits in SZ.78 WNT signaling is associated with cortical development,79 hippocampal neurogenesis80 and synaptic plasticity.81 It is implicated in SZ through interaction with DISC182,83 and is found to be upregulated in bipolar disorder.84 The top metabolic networks including maltohexaose and maltopentaose, and 2 other carbohydrate metabolism networks related to glucose and sucrose were enriched in G2. Although the mechanism of metabolism in psychoses is unclear, prior evidence points to role of estrogen based glucose metabolism in SZ.85 Glucose metabolism interacts with dopamine signaling,86,87 a key network implicated in depression, SZ and PBP.
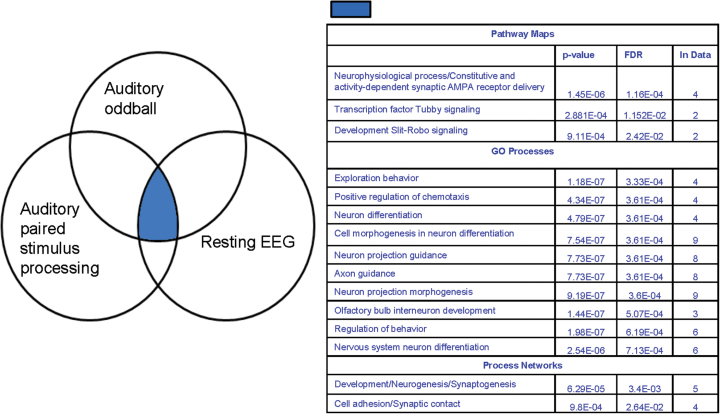
Validation of the current findings can be evaluated as percent overlap between genes\SNPs or pathways88 obtained from an independent sample, which was unavailable in this study. Several candidate genes (N = 13) and mechanisms reported in the current study have previously been implicated in psychosis. In particular, the large scale Psychiatric Genomics Consortium dataset identified immune and neuronal signaling pathways89 associated with SZ. Similarly, the Collaborative Study on Genetics (COGS) of SZ examined association of custom chip SNPs with various (N = 12) endophenotypes including P50 and identified axonal guidance and glutamate signaling90,91 pathways. Additional, pathway analysis studies revealed axon guidance,47,92 netrin signaling, neural cell adhesion process,93,94 immune response signaling,95 and glutamatergic synapse96 underlying psychosis. The individual genes identified using various BSNIP intermediate phenotypes may differ, as these measures assay differential neurophysiological aspects. Several neuronal processes (Venn diagram in figure 3) including neuron projection, differentiation, development, neurogenesis and axon guidance mediating various BSNIP EEG phenotypes comprising auditory oddball ERPs,49 resting EEG frequency activity97,98 and APSP ERPs in psychoses are comparable and similar, validating the current findings and our approach to combine functional and genetic data to unravel the complex etiology underlying these disorders.
Fig. 3.
Venn diagram showing the overlap in gene ontology process, process networks, and pathway maps across 3 different electrophysiological phenotypes. The overlap between the processes and pathway maps is shaded. The intersection of gene clusters obtained from 3 independent multivariate genetic associations of different electroencephalogram (EEG) phenotypes derived from resting EEG, auditory oddball, and auditory paired stimulus processing event-related potentials (ERPs) were entered into GeneGo’s enrichment analysis. The P values denote the significance or the probability of over-representation of the common genes in the data relative to GeneGo’s database by chance.
Limitations
Study limitations include: (1) Demographic variables including age and sex were unbalanced across data collection sites. (2) We grouped the SZA depressive- and manic-type probands into SZ and PBP, respectively. An ongoing debate exists on the inclusion of SZA in psychiatric nosology, perhaps, due to its clinical heterogeneity and lack of sufficient comparisons among SZA vs SZ and PBP. Several prior studies included SZA probands in either SZ or PBP categories, but there is no consensus on a best strategy; however a dimensional approach may be utilized in future studies to circumvent this problem. Thus, the current findings have to be interpreted with caution due to the boundaries we adopted in classifying the SZA probands. (3) Effects of sex and age were regressed through post hoc partial correlation between the loading coefficients of ERP and SNP modalities. (4) Another limitation was possible confounding effects of medications, which could not be controlled for, due to significant variation in the number of medications and duration of use (probands were using multiple medications making it difficult to dissect these into cells). (5) Relatives of probands were not included in the association analyses due to lack of genetic data. (6) Multivariate genetic analyses were confined to only subset of disease associated SNPs, excluding the possibility of identifying other key genetic networks. The current findings may be sensitive to the SNPs selected based on individual logistic regression analysis, comparing each proband group to HC. Future studies can investigate multivariate association of SNPs, selected from univariate analysis comparing all probands to HC, which would clarify the pathways common to psychosis across all probands. (7) Although population stratification effects were corrected using top 3 principal components (associated with race), sample admixture may have influenced the findings. Additionally, missing SNPs were imputed, which may introduce bias in association analysis. Hence we advise caution in interpreting these data. Future studies may employ sophisticated imputation techniques utilizing a reference haplotype to minimize imputation bias. (8) We were unable to verify the current findings due to the lack of replication sample. (9) Samples were too small to examine disorder specific effects.
Conclusions
Several studies have investigated GWA data for SZ and PBP, while some have conducted hypothesis-driven types of pathway analyses. In the current study, we were able to identify data-driven multivariate ERP-genetic associations by probing the auditory sensory system in psychotic probands via a noninvasive paired click paradigm. The genetic network identified in this study explained ~4% of the ERP phenotype variance. The ERP-genetic association is influenced by probands vs controls group difference; specifically PBP-HC group difference was highly prominent. The ERP-genetic association influenced by diagnosis may be of interest to clinical neuroscientists in the identification of trans-diagnostic dimensions of psychosis variance. The most significant pathways and processes that enriched the gene networks were related to several fundamental biologic functions, either general or specific to the nervous system, including neuronal differentiation, neurogenesis, glutamatergic, EGFR, endocannabinoid, and immune signaling. In summary, several genes and multiple biological pathways that have direct impact on brain function and development were found to be associated with P2 abnormalities in psychosis. The pathways identified in this study may be explored for therapeutic targets in future animal model studies. The present study did not examine the P50 ERP component; whose genetic basis will be explored in future studies. Future genetic studies can also examine epistasis among the identified genes.
Supplementary Material
Supplementary material is available at http://schizophreniabulletin.oxfordjournals.org.
Funding
National Institute of Mental Health (R01 MH077851 to G.D.P.; MH078113 to C.A.T.; MH077945 to M.S.K.; MH077862 to J.A.S.; MH077852 to Dr Gunavant Thaker). Additional support (2R44 MH075481 to G.R.; P20GM1034672 and R01EB006841 to V.D.C.).
Supplementary Material
Acknowledgments
We thank the study participants for their contributions.
J.A.S. has received support from Takeda, BMS, Lilly, Roche and Janssen. M.S.K. has received support from Sunovion. C.A.T. has received funding from Astellas, Eli Lilly, Intracellular Therapies, Lundback, and Pure Tech Ventures. Other authors declare no financial interest in relation to the work described in this manuscript other than the grant funding.
References
- 1. Purcell SM, Wray NR, Stone JL, et al. ; International Schizophrenia C. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Craddock N, O’Donovan MC, Owen MJ. Psychosis genetics: modeling the relationship between schizophrenia, bipolar disorder, and mixed (or “schizoaffective”) psychoses. Schizophr Bull. 2009;35:482–490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Lee SH, Ripke S, Neale BM, et al. ; Cross-Disorder Group of the Psychiatric Genomics C. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet. 2013;45:984–994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Craddock N, Sklar P. Genetics of bipolar disorder. Lancet. 2013;381:1654–1662. [DOI] [PubMed] [Google Scholar]
- 5. Allen NC, Bagade S, McQueen MB, et al. Systematic meta-analyses and field synopsis of genetic association studies in schizophrenia: the SzGene database. Nat Genet. 2008;40:827–834. [DOI] [PubMed] [Google Scholar]
- 6. Kennedy JL, Farrer LA, Andreasen NC, Mayeux R, St George-Hyslop P. The genetics of adult-onset neuropsychiatric disease: complexities and conundra? Science. 2003;302:822–826. [DOI] [PubMed] [Google Scholar]
- 7. Ross CA, Margolis RL, Reading SA, Pletnikov M, Coyle JT. Neurobiology of schizophrenia. Neuron. 2006;52:139–153. [DOI] [PubMed] [Google Scholar]
- 8. Keshavan MS, Tandon R, Boutros NN, Nasrallah HA. Schizophrenia, “just the facts”: what we know in 2008 Part 3: neurobiology. Schizophr Res. 2008;106:89–107. [DOI] [PubMed] [Google Scholar]
- 9. Gottesman II, Gould TD. The endophenotype concept in psychiatry: etymology and strategic intentions. Am J Psychiatry. 2003;160:636–645. [DOI] [PubMed] [Google Scholar]
- 10. Hamm JP, Ethridge LE, Shapiro JR, et al. Spatiotemporal and frequency domain analysis of auditory paired stimuli processing in schizophrenia and bipolar disorder with psychosis. Psychophysiology. 2012;49:522–530. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Light GA, Braff DL. Human and animal studies of schizophrenia-related gating deficits. Curr Psychiatry Rep. 1999;1:31–40. [DOI] [PubMed] [Google Scholar]
- 12. Boutros NN, Belger A. Midlatency evoked potentials attenuation and augmentation reflect different aspects of sensory gating. Biol Psychiatry. 1999;45:917–922. [DOI] [PubMed] [Google Scholar]
- 13. Cabranes JA, Ancin I, Santos JL, et al. P50 sensory gating is a trait marker of the bipolar spectrum. Eur Neuropsychopharmacol. 2013;23:721–727. [DOI] [PubMed] [Google Scholar]
- 14. de Wilde OM, Bour LJ, Dingemans PM, Koelman JH, Linszen DH. A meta-analysis of P50 studies in patients with schizophrenia and relatives: differences in methodology between research groups. Schizophr Res. 2007;97:137–151. [DOI] [PubMed] [Google Scholar]
- 15. Sanchez-Morla EM, Garcia-Jimenez MA, Barabash A, et al. P50 sensory gating deficit is a common marker of vulnerability to bipolar disorder and schizophrenia. Acta Psychiatr Scand. 2008;117:313–318. [DOI] [PubMed] [Google Scholar]
- 16. Schulze KK, Hall MH, McDonald C, et al. P50 auditory evoked potential suppression in bipolar disorder patients with psychotic features and their unaffected relatives. Biol Psychiatry. 2007;62:121–128. [DOI] [PubMed] [Google Scholar]
- 17. Boutros NN, Korzyuko O, Oliwa G, et al. Morphological and latency abnormalities of the mid-latency auditory evoked responses in schizophrenia: a preliminary report. Schizophr Res. 2004;70:303–313. [DOI] [PubMed] [Google Scholar]
- 18. Lijffijt M, Moeller FG, Boutros NN, et al. Diminished P50, N100 and P200 auditory sensory gating in bipolar I disorder. Psychiatry Res. 2009;167:191–201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Turetsky BI, Greenwood TA, Olincy A, et al. Abnormal auditory N100 amplitude: a heritable endophenotype in first-degree relatives of schizophrenia probands. Biol Psychiatry. 2008;64:1051–1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Näätänen R. Attention and Brain Function. Hillsdale, NJ: Erlbaum; 1992. [Google Scholar]
- 21. Lijffijt M, Lane SD, Meier SL, et al. P50, N100, and P200 sensory gating: relationships with behavioral inhibition, attention, and working memory. Psychophysiology. 2009;46:1059–1068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Hamm JP, Ethridge LE, Boutros NN, et al. Diagnostic specificity and familiality of early versus late evoked potentials to auditory paired stimuli across the schizophrenia-bipolar psychosis spectrum. Psychophysiology. 2014;51:348–357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Mathalon DH, Hoffman RE, Watson TD, Miller RM, Roach BJ, Ford JM. Neurophysiological distinction between schizophrenia and schizoaffective disorder. Front Hum Neurosci. 2010;3:70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Chun J, Karam ZN, Marzinzik F, et al. Can P300 distinguish among schizophrenia, schizoaffective and bipolar I disorders? An ERP study of response inhibition. Schizophr Res. 2013;151:175–184. [DOI] [PubMed] [Google Scholar]
- 25. Boutros N, Nasrallah H, Leighty R, Torello M, Tueting P, Olson S. Auditory evoked potentials, clinical vs. research applications. Psychiatry Res. 1997;69:183–195. [DOI] [PubMed] [Google Scholar]
- 26. Abrams DJ, Rojas DC, Arciniegas DB. Is schizoaffective disorder a distinct categorical diagnosis? A critical review of the literature. Neuropsychiatr Dis Treat. 2008;4:20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Mann C, Croft RJ, Scholes KE, et al. Differential effects of acute serotonin and dopamine depletion on prepulse inhibition and p50 suppression measures of sensorimotor and sensory gating in humans. Neuropsychopharmacology. 2008;33:1653–1666. [DOI] [PubMed] [Google Scholar]
- 28. Klinkenberg I, Blokland A, Riedel WJ, Sambeth A. Cholinergic modulation of auditory processing, sensory gating and novelty detection in human participants. Psychopharmacology. 2013;225:903–921. [DOI] [PubMed] [Google Scholar]
- 29. Freedman R, Adler LE, Gerhardt GA, et al. Neurobiological studies of sensory gating in schizophrenia. Schizophr Bull. 1987;13:669–678. [DOI] [PubMed] [Google Scholar]
- 30. Hall MH, Schulze K, Sham P, et al. Further evidence for shared genetic effects between psychotic bipolar disorder and P50 suppression: a combined twin and family study. Am J Med Genet B Neuropsychiatr Genet. 2008;147B:619–627. [DOI] [PubMed] [Google Scholar]
- 31. Young DA, Waldo M, Rutledge JH, III, Freedman R. Heritability of inhibitory gating of the P50 auditory-evoked potential in monozygotic and dizygotic twins. Neuropsychobiology. 1996;33:113–117. [DOI] [PubMed] [Google Scholar]
- 32. Anokhin AP, Vedeniapin AB, Heath AC, Korzyukov O, Boutros NN. Genetic and environmental influences on sensory gating of mid-latency auditory evoked responses: a twin study. Schizophr Res. 2007;89:312–319. [DOI] [PubMed] [Google Scholar]
- 33. Freedman R, Coon H, Myles-Worsley M, et al. Linkage of a neurophysiological deficit in schizophrenia to a chromosome 15 locus. Proc Natl Acad Sci U S A. 1997;94:587–592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Stephens SH, Logel J, Barton A, et al. Association of the 5’-upstream regulatory region of the alpha7 nicotinic acetylcholine receptor subunit gene (CHRNA7) with schizophrenia. Schizophr Res. 2009;109:102–112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Myles-Worsley M, Coon H, McDowell J, et al. Linkage of a composite inhibitory phenotype to a chromosome 22q locus in eight Utah families. Am J Med Genet. 1999;88:544–550. [PubMed] [Google Scholar]
- 36. Greenwood TA, Swerdlow NR, Gur RE, et al. Genome-wide linkage analyses of 12 endophenotypes for schizophrenia from the consortium on the genetics of schizophrenia. Am J Psychiatry. 2013;170:521–532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Majic T, Rentzsch J, Gudlowski Y, et al. COMT Val108/158Met genotype modulates human sensory gating. Neuroimage. 2011;55:818–824. [DOI] [PubMed] [Google Scholar]
- 38. Hall MH, Levy DL, Salisbury DF, et al. Neurophysiologic effect of GWAS derived schizophrenia and bipolar risk variants. Am J Med Genet B Neuropsychiatr Genet. 2014;165:9–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Shaikh M, Hall MH, Schulze K, et al. Do COMT, BDNF and NRG1 polymorphisms influence P50 sensory gating in psychosis? Psychol Med. 2011;41:263–276. [DOI] [PubMed] [Google Scholar]
- 40. Hall MH, Levy DL, Salisbury DF, et al. Neurophysiologic effect of GWAS derived schizophrenia and bipolar risk variants. Am J Med Genet B Neuropsychiatr Genet. 2014;165B:9–18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Wray NR, Yang J, Hayes BJ, Price AL, Goddard ME, Visscher PM. Pitfalls of predicting complex traits from SNPs. Nat Rev Genet. 2013;14:507–515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. McCarthy MI, Abecasis GR, Cardon LR, et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet. 2008;9:356–369. [DOI] [PubMed] [Google Scholar]
- 43. Liu J, Demirci O, Calhoun VD. A parallel independent component analysis approach to investigate genomic influence on brain function. IEEE Signal Process Lett. 2008;15:413–416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Liu J, Kiehl KA, Pearlson G, Perrone-Bizzozero NI, Eichele T, Calhoun VD. Genetic determinants of target and novelty-related event-related potentials in the auditory oddball response. Neuroimage. 2009;46:809–816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Liu J, Pearlson G, Windemuth A, Ruano G, Perrone-Bizzozero NI, Calhoun V. Combining fMRI and SNP data to investigate connections between brain function and genetics using parallel ICA. Hum Brain Mapp. 2009;30:241–255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Meda SA, Narayanan B, Liu J, et al. A large scale multivariate parallel ICA method reveals novel imaging-genetic relationships for Alzheimer’s disease in the ADNI cohort. Neuroimage. 2012;60:1608–1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Meda SA, Ruano G, Windemuth A, et al. Multivariate analysis reveals genetic associations of the resting default mode network in psychotic bipolar disorder and schizophrenia. Proc Natl Acad Sci U S A. 2014;111:E2066–E2075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Chen J, Calhoun VD, Pearlson GD, et al. Multifaceted genomic risk for brain function in schizophrenia. Neuroimage. 2012;61:866–875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Narayanan B, Ethridge LE, O’Neil K, et al. Genetic sources of subcomponents of event-related potential in the dimension of psychosis analyzed from the B-SNIP study. Am J Psychiatry. 2015;172:466–478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Tamminga CA, Ivleva EI, Keshavan MS, et al. Clinical phenotypes of psychosis in the Bipolar-Schizophrenia Network on Intermediate Phenotypes (B-SNIP). Am J Psychiatry. 2013;170:1263–1274. [DOI] [PubMed] [Google Scholar]
- 51. Jerger K, Biggins C, Fein G. P50 suppression is not affected by attentional manipulations. Biol Psychiatry. 1992;31:365–377. [DOI] [PubMed] [Google Scholar]
- 52. Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT. Data quality control in genetic case-control association studies. Nat Protoc. 2010;5:1564–1573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nat Genet. 2006;38:904–909. [DOI] [PubMed] [Google Scholar]
- 54. Kamburov A, Wierling C, Lehrach H, Herwig R. ConsensusPathDB—a database for integrating human functional interaction networks. Nucleic Acids Res. 2009;37:D623–D628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55. Näätänen R, Picton T. The N1 wave of the human electric and magnetic response to sound: a review and an analysis of the component structure. Psychophysiology. 1987;24:375–425. [DOI] [PubMed] [Google Scholar]
- 56. Frank CL, Tsai LH. Alternative functions of core cell cycle regulators in neuronal migration, neuronal maturation, and synaptic plasticity. Neuron. 2009;62:312–326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Kraus DM, Elliott GS, Chute H, et al. CSMD1 is a novel multiple domain complement-regulatory protein highly expressed in the central nervous system and epithelial tissues. J Immunol. 2006;176:4419–4430. [DOI] [PubMed] [Google Scholar]
- 58. Stephan AH, Barres BA, Stevens B. The complement system: an unexpected role in synaptic pruning during development and disease. Annu Rev Neurosci. 2012;35:369–389. [DOI] [PubMed] [Google Scholar]
- 59. Havik B, Le Hellard S, Rietschel M, et al. The complement control-related genes CSMD1 and CSMD2 associate to schizophrenia. Biol Psychiatry. 2011;70:35–42. [DOI] [PubMed] [Google Scholar]
- 60. Xu W, Cohen-Woods S, Chen Q, et al. Genome-wide association study of bipolar disorder in Canadian and UK populations corroborates disease loci including SYNE1 and CSMD1. BMC Med Genet. 2014;15:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature. 2014;511:421–427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Squassina A, Manchia M, Borg J, et al. Evidence for association of an ACCN1 gene variant with response to lithium treatment in Sardinian patients with bipolar disorder. Pharmacogenomics. 2011;12:1559–1569. [DOI] [PubMed] [Google Scholar]
- 63. Tse MT. Neurodevelopmental disorders: exploring the links between SHANK2 and autism. Nat Rev Drug Discov. 2012;11:518. [DOI] [PubMed] [Google Scholar]
- 64. Hwang JI, Kim HS, Lee JR, Kim E, Ryu SH, Suh PG. The interaction of phospholipase C-beta3 with Shank2 regulates mGluR-mediated calcium signal. J Biol Chem. 2005;280:12467–12473. [DOI] [PubMed] [Google Scholar]
- 65. Khatri P, Sirota M, Butte AJ. Ten years of pathway analysis: current approaches and outstanding challenges. PLoS Comput Biol. 2012;8:e1002375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Hashimotodani Y, Ohno-Shosaku T, Kano M. Endocannabinoids and synaptic function in the CNS. Neuroscientist. 2007;13:127–137. [DOI] [PubMed] [Google Scholar]
- 67. Ohno-Shosaku T, Hashimotodani Y, Ano M, Takeda S, Tsubokawa H, Kano M. Endocannabinoid signalling triggered by NMDA receptor-mediated calcium entry into rat hippocampal neurons. J Physiol. 2007;584:407–418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Lee SH, Foldy C, Soltesz I. Distinct endocannabinoid control of GABA release at perisomatic and dendritic synapses in the hippocampus. J Neurosci. 2010;30:7993–8000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Yin HH, Lovinger DM. Frequency-specific and D2 receptor-mediated inhibition of glutamate release by retrograde endocannabinoid signaling. Proc Natl Acad Sci U S A. 2006;103:8251–8256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Melis M, Pistis M. Endocannabinoid signaling in midbrain dopamine neurons: more than physiology? Curr Neuropharmacol. 2007;5:268–277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Ferrari G, Toffano G, Skaper SD. Epidermal growth factor exerts neuronotrophic effects on dopaminergic and GABAergic CNS neurons: comparison with basic fibroblast growth factor. J Neurosci Res. 1991;30:493–497. [DOI] [PubMed] [Google Scholar]
- 72. Ayuso-Sacido A, Moliterno JA, Kratovac S, et al. Activated EGFR signaling increases proliferation, survival, and migration and blocks neuronal differentiation in post-natal neural stem cells. J Neurooncol. 2010;97:323–337. [DOI] [PubMed] [Google Scholar]
- 73. Futamura T, Toyooka K, Iritani S, et al. Abnormal expression of epidermal growth factor and its receptor in the forebrain and serum of schizophrenic patients. Mol Psychiatry. 2002;7:673–682. [DOI] [PubMed] [Google Scholar]
- 74. Norton N, Moskvina V, Morris DW, et al. Evidence that interaction between neuregulin 1 and its receptor erbB4 increases susceptibility to schizophrenia. Am J Med Genet B Neuropsychiatr Genet. 2006;141B:96–101. [DOI] [PubMed] [Google Scholar]
- 75. Schang AL, Ngô-Muller V, Bleux C, et al. GnRH receptor gene expression in the developing rat hippocampus: transcriptional regulation and potential roles in neuronal plasticity. Endocrinology. 2011;152:568–580. [DOI] [PubMed] [Google Scholar]
- 76. Wierman ME, Bruder JM, Kepa JK. Regulation of gonadotropin-releasing hormone (GnRH) gene expression in hypothalamic neuronal cells. Cell Mol Neurobiol. 1995;15:79–88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Cantalamessa L, Catania A, Silva A, Orsatti A, Motta P, Cazzullo CL. Gonadotropin releasing hormone elicits abnormal hormone responses in schizophrenia. Psychoneuroendocrinology. 1985;10:481–484. [DOI] [PubMed] [Google Scholar]
- 78. Goldstein JM. Sex, hormones and affective arousal circuitry dysfunction in schizophrenia. Horm Behav. 2006;50:612–622. [DOI] [PubMed] [Google Scholar]
- 79. Chenn A. Wnt/beta-catenin signaling in cerebral cortical development. Organogenesis. 2008;4:76–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Varela-Nallar L, Inestrosa NC. Wnt signaling in the regulation of adult hippocampal neurogenesis. Front Cell Neurosci. 2013;7:100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81. Budnik V, Salinas PC. Wnt signaling during synaptic development and plasticity. Curr Opin Neurobiol. 2011;21:151–159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Inestrosa NC, Montecinos-Oliva C, Fuenzalida M. Wnt signaling: role in Alzheimer disease and schizophrenia. J Neuroimmune Pharmacol. 2012;7:788–807. [DOI] [PubMed] [Google Scholar]
- 83. Singh KK, Ge X, Mao Y, et al. Dixdc1 is a critical regulator of DISC1 and embryonic cortical development. Neuron. 2010;67:33–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Matigian N, Windus L, Smith H, et al. Expression profiling in monozygotic twins discordant for bipolar disorder reveals dysregulation of the WNT signalling pathway. Mol Psychiatry. 2007;12:815–825. [DOI] [PubMed] [Google Scholar]
- 85. Olsen L, Hansen T, Jakobsen KD, et al. The estrogen hypothesis of schizophrenia implicates glucose metabolism: association study in three independent samples. BMC Med Genet. 2008;9:39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Tamminga CA, Burrows GH, Chase TN, Alphs LD, Thaker GK. Dopamine neuronal tracts in schizophrenia: their pharmacology and in vivo glucose metabolism. Ann N Y Acad Sci. 1988;537:443–450. [DOI] [PubMed] [Google Scholar]
- 87. Siuta MA, Robertson SD, Kocalis H, et al. Dysregulation of the norepinephrine transporter sustains cortical hypodopaminergia and schizophrenia-like behaviors in neuronal rictor null mice. PLoS Biol. 2010;8:e1000393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Pearlson GD, Calhoun VD, Liu J. An Introductory review of parallel independent component analysis (p-ICA) and a guide to applying p-ICA to genetic data and imaging phenotypes to identify disease-associated biological pathways and systems in common complex disorders. Front Genet. 2015;6:276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89. Network, Pathway Analysis Subgroup of the Psychiatric Genomics C, International Inflammatory Bowel Disease Genetics C, International Inflammatory Bowel Disease Genetics Consortium I. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat Neurosci. 2015;18:199–209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Greenwood TA, Light GA, Swerdlow NR, Radant AD, Braff DL. Association analysis of 94 candidate genes and schizophrenia-related endophenotypes. PLoS One. 2012;7:e29630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Greenwood TA, Lazzeroni LC, Murray SS, et al. Analysis of 94 candidate genes and 12 endophenotypes for schizophrenia from the Consortium on the Genetics of Schizophrenia. Am J Psychiatry. 2011;168:930–946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Gilman SR, Chang J, Xu B, et al. Diverse types of genetic variation converge on functional gene networks involved in schizophrenia. Nat Neurosci. 2012;15:1723–1728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. O’Dushlaine C, Kenny E, Heron E, et al. Molecular pathways involved in neuronal cell adhesion and membrane scaffolding contribute to schizophrenia and bipolar disorder susceptibility. Mol Psychiatry. 2011;16:286–292. [DOI] [PubMed] [Google Scholar]
- 94. Juraeva D, Haenisch B, Zapatka M, et al. Integrated pathway-based approach identifies association between genomic regions at CTCF and CACNB2 and schizophrenia. PLoS Genet. 2014;10:e1004345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Jia P, Wang L, Meltzer HY, Zhao Z. Common variants conferring risk of schizophrenia: a pathway analysis of GWAS data. Schizophr Res. 2010;122:38–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Nurnberger JI, Jr, Koller DL, Jung J, et al. Identification of pathways for bipolar disorder: a meta-analysis. JAMA Psychiatry. 2014;71:657–664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Narayanan B, O’Neil K, Berwise C, et al. Resting state electroencephalogram oscillatory abnormalities in schizophrenia and psychotic bipolar patients and their relatives from the bipolar and schizophrenia network on intermediate phenotypes study. Biol Psychiatry. 2014;76:456–465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98. Narayanan B, O’Neil K, Calhoun VD, et al. Multivariate genetic determinants of EEG oscillations in schizophrenia and psychotic bipolar disorder from the BSNIP study. Transl Psychiatry. 2015;5:e588. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.