Abstract
The transdifferentiation from cardiac fibroblasts to myofibroblasts is an important event in the initiation of cardiac fibrosis. However, the underlying mechanism is not fully understood. Circ-sh3rf3 (circular RNA SH3 domain containing Ring Finger 3) is a novel circular RNA which was induced in hypertrophied ventricles by isoproterenol hydrochloride, and our work has established that it is a potential regulator in cardiac hypertrophy, but whether circ-sh3rf3 plays a role in cardiac fibrosis remains unclear, especially in the conversion of cardiac fibroblasts into myofibroblasts. Here, we found that circ-sh3rf3 was down-regulated in isoproterenol-treated rat cardiac fibroblasts and cardiomyocytes as well as during fibroblast differentiation into myofibroblasts. We further confirmed that circ-sh3rf3 could interact with GATA-4 proteins and reduce the expression of GATA-4, which in turn abolishes GATA-4 repression of miR-29a expression and thus up-regulates miR-29a expression, thereby inhibiting fibroblast–myofibroblast differentiation and myocardial fibrosis. Our work has established a novel Circ-sh3rf3/GATA-4/miR-29a regulatory cascade in fibroblast–myofibroblast differentiation and myocardial fibrosis, which provides a new therapeutic target for myocardial fibrosis.
Supplementary Information
The online version contains supplementary material available at 10.1007/s00018-023-04699-7.
Keywords: Circular RNA, Fibroblast–myofibroblast differentiation, Myocardial fibrosis, microRNAs
Introduction
Myocardial fibrosis is a common pathological process involved in the development of many different types of heart diseases and is characterized by excessive accumulation of extracellular matrix proteins and myofibroblasts in the heart [3, 20, 40]. In general, when the heart is subjected to stimulation, such as pressure overload and myocardial inflammation [20], cardiac fibroblasts transdifferentiate into myofibroblasts, which promote excess collagen deposition and show stronger contractile activity, thus protecting the myocardium from rupture to initially form fibrotic scars at the injured sites [15]. However, myofibroblast persistence ultimately leads to cardiac fibrosis, cardiac remodeling, and even heart failure [20, 35, 40]. Therefore, it is important to understand the mechanism underlying the formation of myofibroblasts and cardiac fibrosis to determine future therapeutic targets.
Recent studies have demonstrated that several circular RNAs (circRNAs) participate in the regulation of myocardial fibrosis [5, 11, 30, 39]. Given that the differentiation of cardiac fibroblasts to myofibroblasts is an important process during cardiac fibrosis, it is believed that these circRNAs are also involved in the fibroblast-to-myofibroblast transition. Another circular RNA, circ-sh3rf3, which we previously discovered to be derived from the Sh3rf3 (SH3 domain containing ring finger 3) gene, has also been revealed to be a potential regulator in isoproterenol hydrochloride-induced cardiac hypertrophy [48]. Given that persistent myocardial hypertrophy can lead to cardiac fibrosis [43], we speculated that circ-sh3rf3 plays roles in cardiac fibrosis and fibroblast-to-myofibroblast transition.
One of the ways by which circular RNA works is as a microRNA sponge. Indeed, recent studies have revealed that several microRNAs, such as miR-141, miR-433, and miR-29b-3p [12, 25, 30], can regulate cardiac fibrosis via sponge adsorption by circRNAs, which results in altered expression of microRNA target genes, including cardiac fibrosis-associated genes. Among these microRNAs, miR-29a has been revealed to be involved in myocardial fibrosis [34]. In addition, several lines of evidence proved that miR-29a was adsorbed by circRNAs, such as circHIPK3 and cZNF532 in the heart [17, 44], suggesting that miR-29a may play roles in myocardial fibrosis through interaction with circular RNA. In addition to its role in myocardial fibrosis, miR-29a also plays roles in cardiac hypertrophy via the inhibition of PPARδ expression [53], and given that circ-sh3rf3 also exerts effects on cardiac hypertrophy, we speculated that circ-sh3rf3 may function via miR-29a.
In addition to acting as a sponge, circular RNA can also function by interacting with proteins. For instance, circular RNA circFndc3b can bind to fused protein to regulate cardiac repair after myocardial infarction [13]. These proteins include RNA binding proteins and transcription factors, which play roles in synergy with circular RNA by assembling to these circRNAs [9], but for important transcription factors in the heart, specifically for GATA-4 [6], it is unclear whether GATA-4 interacts with circRNAs to play roles in cardiac fibrosis and fibroblast-to-myofibroblast transition.
Here, we provide evidence that circ-sh3rf3 could interact with GATA-4 and inhibit its expression. Inhibiting GATA-4 expression results in elevated miR-29a expression, which in turn attenuates the transition of fibroblast-to-myofibroblast and myocardial fibrosis, thereby underlining the regulatory cascade involving circ-sh3rf3/GATA-4/miR-29a and its clinical relevance for the treatment of myocardial fibrosis.
Results
circ-sh3rf3 is downregulated in isoproterenol (ISO)-induced cardiac fibrosis
Previously, we showed that the circular RNA circ-sh3rf3 was markedly decreased in hypertrophied ventricles induced by isoproterenol [48], which was formed by back-splicing from exon 2 to exon 3 of the Sh3rf3 gene and localized both in the cytoplasm and nucleus (Figure S1A, B). Other than in the ventricle, qPCR assays revealed that both in neonatal rat cardiomyocytes and in cardiac fibroblasts treated with 10 μM isoproterenol for 48 h, the mRNA level of circ-sh3rf3 was significantly reduced (Fig. 1A, B), along with increased mRNA levels of the myocardial hypertrophy markers ANF and BNP (Fig. 1C) and cardiac fibrosis-associated markers, including transforming growth factor-β1 (TGF-β1), connective tissue growth factor (CTGF), matrix metalloproteinase-2 (MMP-2), matrix metalloproteinase-9 (MMP-9), collagen type I alpha 1 (COL1A1) and α-smooth muscle actin (α-SMA) (Fig. 1D), indicating that circ-sh3rf3 is involved not only in cardiomyocyte hypertrophy but also in cardiac fibrosis. To validate the presence of cardiomyocytes, troponin T or α-actinin staining was typically utilized [51]. Therefore, α-actinin staining was used to evaluate the purity of the cardiomyocytes (Fig. S1C), and cardiac fibrosis induced by isoproterenol was verified by Masson trichrome staining in the ventricle (Fig. 1E), suggesting that circ-sh3rf3 may play roles in cardiac fibrosis.
Fig. 1.
Decreased circ-sh3rf3 expression in isoproterenol (ISO)-induced cardiac fibrosis. (A, B) qPCR for circRNA_Sh3rf3 and Sh3rf3 in isoproterenol-treated rat cardiomyocytes (A) and cardiac fibroblasts (B). (C) qPCR assays for atrial natriuretic factor (ANF) and B-type natriuretic peptide (BNP) were performed in isoproterenol-treated rat cardiomyocytes. (D) The expression of transforming growth factor-β1 (TGF-β1), connective tissue growth factor (CTGF), matrix metalloproteinase-2 (MMP-2), matrix metalloproteinase-9 (MMP-9), collagen type I alpha 1 (COL1A1), and α-smooth muscle actin (α-SMA) was determined by qPCR in isoproterenol-treated rat cardiac fibroblasts. (E) Masson trichrome staining of left ventricular sections from mice infused with isoproterenol or saline (control) for 2 weeks (left), and fibrotic areas were quantitated by with Image J software (right). Scale bars: 50 μm. Iso isoproterenol. The data shown are the means ± SEMs from three independent experiments. *P < 0.05, **P < 0.01 compared with the control, and rat cardiomyocytes and cardiac fibroblasts treated with PBS (A–D) served as the control
circ-sh3rf3 expression is decreased during fibroblast differentiation into myofibroblasts
Given that cardiac fibroblast-to-myofibroblast conversion is an inevitable process in the development of myocardial fibrosis [2], we speculate that circ-sh3rf3 will be implicated in the process of cardiac fibroblast–myofibroblast transformation. Since normal culture media containing 10% FBS could lead to automatic differentiation of cardiac fibroblasts into myofibroblasts [7], we cultured primary fibroblasts in medium with 10% serum compared with 1% serum, which maintained the fibroblast phenotype. Typically, vimentin is employed to identify fibroblasts [37]. Here, to determine the differentiation from cardiac fibroblast to myofibroblast, F-actin staining was utilized to assess the change in the presence of stress fibers in cardiac fibroblast and myofibroblast [35]. Indeed, culture of cardiac fibroblasts in 10% FBS resulted in the formation of myofibroblasts, as indicated by the increased presence of stress fibers and the expression of α-SMA, an important marker of myofibroblasts, along with a well-spread myofibroblast-like morphology (Fig. 2A–C). In addition, we found that the expression of circ-sh3rf3 was markedly decreased compared with that in cells exposed to 1% serum medium (Fig. 2D), indicating that circ-sh3rf3 may be involved in the transition of fibroblasts to myofibroblasts. In addition to exposure to 10% FBS, transforming growth factor-beta1 (TGF-β1) is a well-characterized stimulator of the transformation of fibroblasts to myofibroblasts [35]; therefore, we treated cardiac fibroblasts with different concentrations of TGF-β1 (5, 10, 20 ng/mL) for 24 or 48 h, and the mRNA level of α-SMA was analyzed to determine the optimal condition of TGF-β1 treatment. Treatment with 5 ng/ml TGF-β1 for 48 h was sufficient to result in the highest mRNA level of α-SMA (Fig. 2E). Moreover, we observed more obvious changes in cell morphology, including increased cell size, the presence of stress fibers, and the upregulation of α-SMA, CTGF and TGF-β1, all crucial markers of myofibroblasts (Fig. 2F–J). Collectively, these data indicate that the myofibroblast phenotype is induced by TGF-β1. Furthermore, similar to what was observed in 10% FBS exposure, dramatic downregulation of circ-sh3rf3 also occurred in TGF-β1-treated cardiac fibroblasts compared to untreated fibroblasts (Fig. 2K), suggesting a potential role of circ-sh3rf3 in myofibroblast transdifferentiation.
Fig. 2.
circ-sh3rf3 expression decreases during fibroblast differentiation into myofibroblasts. (A) Representative images of cardiac fibroblasts cultured in 1% or 10% serum using immunofluorescence. Cardiac fibroblasts spontaneously differentiated into myofibroblasts in 10% serum with marked changes in cell morphology and the presence of stress fibers by staining with F-actin. (B–C) α-SMA expression was determined by Western blot (B) and qPCR (C). The protein bands were quantitated by densitometry (B, lower). (D) qPCR assay for circ-sh3rf3 in cardiac fibroblasts maintained in 10% serum. (E) Cardiac fibroblasts were starved for 24 h, followed by treatment with different concentrations of TGF-β1 (5, 10, 20 ng/ml) for 24 or 48 h. The mRNA level of α-SMA was analyzed by qPCR. (F) Representative images of fibroblasts treated with 5 ng/ml TGF-β1 for 48 h using immunofluorescence. F-actin staining (upper panel) shows marked changes in cell morphology, and immunofluorescence (lower panel) shows increased expression of α-SMA in cardiac fibroblasts stimulated with TGF-β1. (G–I) qPCR for α-SMA, CTGF, and TGF-β1 in cardiac fibroblasts treated with TGF-β1. (J) Western blotting for α-SMA, CTGF, and TGF-β1 in TGF-β1-treated cardiac fibroblasts (left), and the protein bands were quantitated by densitometry (right). (K) qPCR analysis of circ-sh3rf3 in cardiac fibroblasts treated with TGF-β1. The data shown are the means ± SEMs from three independent experiments, **P < 0.01 compared with the control, and cardiac fibroblasts cultured in 1% serum (B–D) and treated with PBS (E, G–K) served as the control. Scale bars: 50 μm
circ-sh3rf3 inhibits the conversion of fibroblasts to myofibroblasts
To further investigate the role of circ-sh3rf3 in cardiac fibroblast-to-myofibroblast conversion, we tested the effects of exogenous expression of circ-sh3rf3 on the TGF-β1-induced transformation. The expression of circ-sh3rf3 was confirmed in cardiac fibroblasts infected with circ-sh3rf3 adenovirus by green fluorescent protein (GFP) reporter and qPCR (Fig. 3A, B), while the expression of Sh3rf3 was not changed (Fig. 3C). Furthermore, the expression of CTGF and α-SMA was significantly decreased in cardiac fibroblasts overexpressing circ-sh3rf3 and cultured with 10% FBS compared with that in the empty vector control (Fig. 3D, E), indicating that circ-sh3rf3 inhibits the conversion of fibroblasts to myofibroblasts.
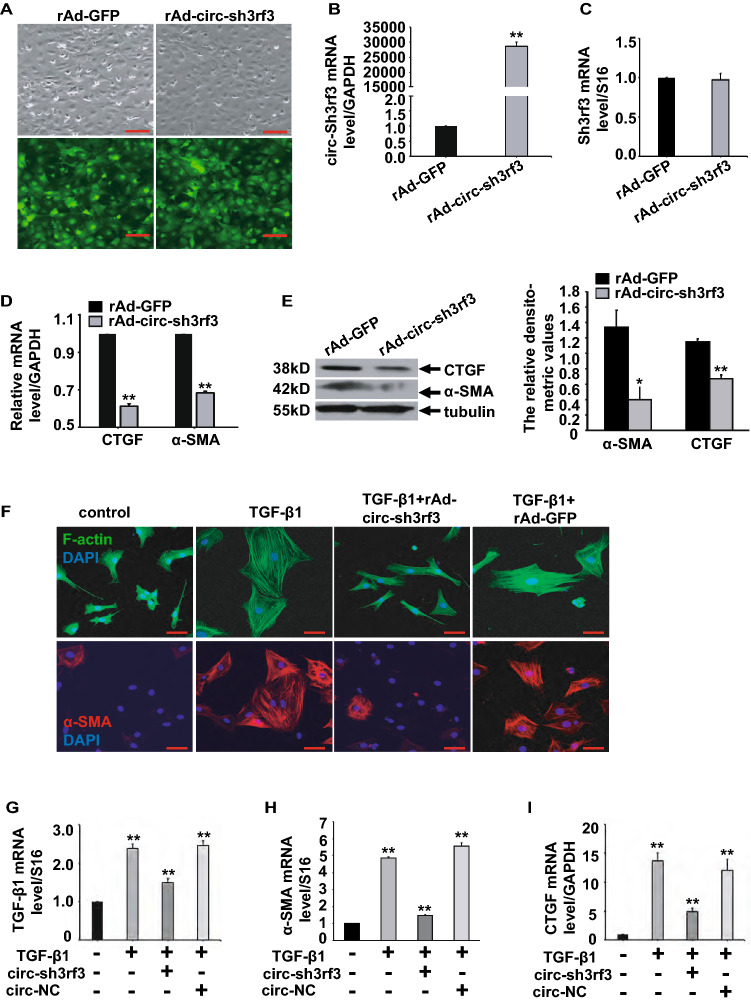
Fig. 3.
circ-sh3rf3 inhibits fibroblast-to-myofibroblast differentiation. (A) Adenovirus-mediated overexpression of circ-sh3rf3 in cardiac fibroblasts cultured in medium with 10% serum. Recombinant circ-sh3rf3 adenovirus or empty adenovirus vector with the coexpression of green fluorescent protein (GFP) was constructed and then used to infect cardiac fibroblasts, and the infection efficiency of recombinant circ-sh3rf3 adenovirus (rAd-circ-sh3rf3) was reflected by the expression of GFP. rAd-GFP, empty adenovirus vector. Scale bars: 100 μm. (B–C) qPCR of circ-sh3rf3 (B) and sh3rf3 (C) in cardiac fibroblasts infected with rAd-circ-sh3rf3 or rAd-GFP (control). (D–E) Expression of CTGF and α-SMA was determined by qPCR (D) and Western blotting (E, left) in adenovirus-infected cardiac fibroblasts, rAd-GFP served as the control. and the protein bands were quantitated by densitometry (E, right). (F) Immunofluorescence analysis of F-actin (upper panel) and α-SMA (lower panel) in cardiac fibroblasts treated with TGF-β1 and/or circ-Sh3rf3. (G–I) Expression of TGF-β1, α-SMA and CTGF in cardiac fibroblasts treated with TGF-β1 and/or circ-sh3rf3 was detected by qPCR, and cardiac fibroblasts treated with PBS served as the control. The data shown are the means ± SEMs from three independent experiments, **P < 0.01 compared with the control. Scale bars: 50 μm. circ-NC: circular RNA negative control
Next, we evaluated the anti-transdifferentiation effect of circ-sh3rf3 in cardiac fibroblasts induced by 5 ng/ml TGF-β1 for 48 h. As shown in Fig. 3F, circ-sh3rf3 overexpression resulted in decreased cell size and stress fibers and α-SMA protein expression in TGF-β1-treated cardiac fibroblasts compared with the empty vector control. Furthermore, the mRNA levels of TGF-β1, CTGF and α-SMA were also downregulated in circ-sh3rf3-overexpressing CFs treated with TGF-β1 (Fig. 3G–I). Taken together, these data indicated that circ-sh3rf3 could inhibit the transdifferentiation of fibroblasts to myofibroblasts.
circ-sh3rf3 inhibits the conversion of fibroblasts to myofibroblasts via the upregulation of miR-29a
We subsequently investigated the mechanism through which circ-sh3rf3 inhibits the transdifferentiation from fibroblasts to myofibroblasts by searching for downstream targets that might mediate this inhibitory effect. Considering that miR-29a has an inhibitory effect on cardiac fibrosis [26, 34], we examined the expression of miR-29a in cardiac fibroblasts overexpressing circ-sh3rf3. As expected, miR-29a was significantly upregulated in cardiac fibroblasts infected with circ-sh3rf3 (Fig. 4A), suggesting that circ-sh3rf3 suppresses the transdifferentiation of myofibroblasts via miR-29a. Next, we evaluated the roles of miR-29a in cardiac fibrosis induced by isoproterenol. In previous studies, isoproterenol was administered at a dose of 30 mg/kg/day for 13, 14, 15, and 21 days, or 5 weeks [22, 31, 32]. Given 14 days of treatment, mice could successfully develop left ventricular hypertrophy and cardiac fibrosis [54], in the present study, isoproterenol was administered at a dose of 30 mg/kg/day for 14 days. As shown in Fig. 4B, there was decreased collagen deposition in the ventricles of mice treated with miR-29a and isoproterenol compared with only the isoproterenol- and/or miR-29a-negative control-treated group (Fig. 4B and S1D). Moreover, the mRNA levels of CTGF, α-SMA, collagen type III alpha 1 (COL3A1), and MMP-2 were decreased in the left ventricles of mice treated with miR‐29a and isoproterenol compared with those in isoproterenol- and/or miR-29a-negative control-treated ventricles (Fig. 4C–F). On the other hand, the expression of miR-29a was downregulated in the ventricles of mice, and rat cardiomyocytes and cardiac fibroblasts treated with isoproterenol (Figures S1E and 4G). Altogether, the results indicated that miR-29a could protect the myocardium from cardiac fibrosis induced by isoproterenol.
Fig. 4.
circ-sh3rf3 attenuates fibroblast-to-myofibroblast differentiation by upregulating miR-29a. (A) qPCR assays for miR-29a in cardiac fibroblasts infected with rAd-circ-sh3rf3. The empty vector served as the control. (B) Masson trichrome staining of left ventricular sections from mice infused with Iso and/or miR-29a agomir or agomir negative control. (C–F) mRNA levels of CTGF, α-SMA, collagen type III alpha 1 (COL3A1), and MMP-2 were determined by qPCR in the left ventricles of mice treated with Iso and/or miR-29a agomir or agomir negative control, and treatment with saline served as the control. (G) The expression of miR-29a in isoproterenol-treated cardiac myocytes(cMs) and fibroblasts (cFs) was determined by qPCR, and treatment with PBS served as the control. (H) qPCR for miR-29a in cardiac fibroblasts transfected with miR-29a agomir or agomir negative control, and PBS treatment served as the control. (I–L) qPCR for CTGF, TGF-β1, α-SMA, and MMP-2 in rat cardiac fibroblasts treated with Iso and/or miR-29a agomir or agomir negative control, and PBS treatment served as the control. (M) Western blotting for CTGF, TGF-β1, α-SMA, MMP-2, TGFβ-receptor1 (TGFβ-R1), and Smad3 phosphorylation (p-Smad3) in rat cardiac fibroblasts treated with Iso and/or miR-29a agomir or agomir negative control (upper). PBS treatment served as the control. The protein bands were quantitated by densitometry (lower). (N) F-actin and α-SMA staining using immunofluorescence analysis in cardiac fibroblasts treated with 5 ng/ml TGF-β1 for 48 h and/or miR-29a agomir. (O) qPCR for CTGF, α-SMA and TGF-β1 in cardiac fibroblasts overexpressed with circ-Sh3rf3 and/or transfected with miR-29a antagomir (left), and the expression of miR-29a was determined by qPCR (right). Cardiac fibroblasts overexpressed with antagomir negative control (right) and circ-Sh3rf3 (left) served as the control. The data shown are the means ± SEMs from three independent experiments, *P < 0.05, **P < 0.01 compared with the control. Scale bars: 50 μm. Ctr: control, c-sh3rf3: circular sh3rf3. Iso: isoproterenol, 29a: miR-29a agomir, NC: miR-29a agomir negative control, anti-29a: miR-29a antagomir, anti-NC: antagomir negative control
To further investigate the anti-fibrotic role of miR-29 in the ventricle, we tested the fibrotic response of miR-29a in cardiac fibroblasts. Delivery of miR-29a in these isoproterenol-treated cells caused reduced expression of CTGF, TGF-β1, α-SMA, and MMP-2 compared to isoproterenol- and/or miR-29a-negative control-treated cardiac fibroblasts (Fig. 4H–M). Given that TGF-β1/Smad3 signaling is activated during cardiac fibrosis, we detected the expression of TGFβ-receptor1 (TGFβ-R1) and Smad3 phosphorylation (p-Smad3), and the results showed that miR-29a could decrease the expression of TGFβ-R1 and p-Smad3 in isoproterenol-treated cardiac fibroblasts (Fig. 4M). Taken together, these data revealed that miR-29a could attenuate isoproterenol-induced cardiac fibrosis by negatively regulating the TGF-β1/Smad3 signaling pathway.
Next, we examined the roles of miR-29a in the TGF-β1-induced transformation of fibroblasts to myofibroblasts. The results showed that miR-29a inhibited the transdifferentiation of myofibroblasts, as evidenced by decreased cell size and stress fibers and α-SMA protein expression in TGF-β1- and miR-29a-treated cardiac fibroblasts compared with that in TGF-β1- and/or miR-29a-negative control-treated cells (Fig. 4N). Furthermore, if miR-29a expression was inhibited by the addition of miR-29a antagomir, the expression of CTGF, α-SMA, and TGF-β1 in cardiac fibroblasts overexpressing circ-sh3rf3 was upregulated (Fig. 4O), indicating that circ-sh3rf3 inhibits the conversion of fibroblasts to myofibroblasts via miR-29a.
circ-sh3rf3 upregulated miR-29a expression to suppress cardiac fibroblast-to-myofibroblast conversion through inhibition of GATA-4 expression
To further address the mechanism, whereby circ-sh3rf3 exerts its anti-fibrotic function via upregulation of miR-29a expression, transient transfection experiments were performed to determine whether circ-sh3rf3 could enhance the transcription of miR-29a. The results showed that the promoter activity of miR-29a was consistently unchanged with the increase in circ-sh3rf3 (Figure S2A). Given that it has been widely reported that circRNAs can act as miRNA sponges [24, 28, 41], we performed bioinformatic analysis to identify the potential miRNA binding sites within the sequence of circ-sh3rf3 using circMir software. However, no miR-29a binding sites on circ-sh3rf3 were predicted by the two algorithms, i.e., miRanda (Figure S2B) and RNA hybrid (Figure S2C). In addition, an RNA pull-down assay revealed that miR-29a was not captured by the circ-sh3rf3 probe, indicating that miR-29a could not bind to circ-sh3rf3 (Figure S2 D–E). Altogether, these results indicated that circ-sh3rf3 could not directly interact with miR-29a.
Apart from acting as sponges for miRNAs, circRNAs were reported to play important roles in the regulation of biological processes by interacting with proteins [8, 13, 18, 47]. As a key transcription factor, GATA-binding protein 4 (GATA-4) was reported to be involved in cardiac fibrosis induced by dexamethasone [4], so we speculated that circ-Sh3rf might interact with GATA-4 to play roles in cardiac fibrosis. As shown in Fig. 5A, there were many GATA-4 binding motifs on the sequence of circ-sh3rf3 predicted by the bioinformatics software RBPmap [33]. In addition, GATA-4 could be pulled down by circ-sh3rf3 in cardiac fibroblasts by RNA pull-down assays (Fig. 5B). Furthermore, RNA immunoprecipitation assays further validated the binding of circ-sh3rf3 to GATA-4 in H9C2 cells (Fig. 5C) and cardiac fibroblasts (Fig. 5D). Collectively, these data indicate that circ-sh3rf3 could interact with GATA-4.
Fig. 5.
circ-sh3rf3 upregulates the expression of miR-29a by inhibiting GATA-4 expression. (A) Bioinformatic analysis of potential GATA-4 binding sites on the circ-sh3rf3 sequence using RBPmap. (B) Cell lysates from cardiac fibroblasts infected with recombinant circ-sh3rf3 adenovirus were mixed with biotinylated circ-sh3rf3 or random probe, incubated with streptavidin beads and finally subjected to Western blotting using anti-GATA-4 antibody. (C–D) Cell lysates from H9C2 cells (C) or cardiac fibroblasts (D) infected with recombinant circ-sh3rf3 adenovirus were subjected to immunoprecipitation with antibodies against rabbit IgG (control) or GATA-4, followed by qPCR. (E) The mRNA level of GATA-4 in cardiac fibroblasts transfected with circ-sh3rf3 was determined by q-PCR (Left), and the protein level of GATA-4 and GATA-6 were detected by Western blotting and quantitated by densitometry (right). (F) Expression of miR-29a (upper) and GATA-4 (lower) in cardiac fibroblasts transfected with RNAi GATA-4 by qPCR. (G) mRNA levels of CTGF, TGF-β1, and α-SMA in cardiac fibroblasts transfected with RNAi GATA-4 by qPCR. The empty vector served as the control (E–G). (H–I) qPCR for CTGF and α-SMA in cardiac fibroblasts treated with TGF-β1 and Ri-GATA4. (J) Cardiac fibroblasts were treated with TGF-β1 and/or transfected with Ri-GATA4, and F-actin (upper panel) and α-SMA (lower panel) staining was determined by immunofluorescence. PBS treatment served as the control (H–J). The data shown are the means ± SEMs from three independent experiments,**P < 0.01 compared with the control. Scale bars: 50 μm. rAd-circ-sh3rf3 Recombinant circ-sh3rf3 adenovirus, CFs cardiac fibroblasts, ctr control, c-sh3rf3 circular sh3rf3, Ri-G4 RNAi-GATA-4, Tβ TGF-β1
Next, we evaluated the interaction effect of circ-sh3rf3 and GATA-4 on miR-29a expression. The results showed that overexpression of circ-sh3rf3 reversed the GATA-4-mediated downregulation of miR-29a (Fig. S2F), which demonstrated that the interaction between circ-sh3rf3 and GATA-4 could relieve the inhibitory miR-29a expression mediated by GATA-4. Furthermore, overexpression of circ-sh3rf3 remarkably reduced the binding ability of GATA-4 to the miR-29a promoter (Fig. S2G). Taken together, these data strongly demonstrated that circ-sh3rf3 interacts with GATA-4 to relieve the suppressed the expression of miR-29a mediated by GATA-4.
Moreover, overexpression of circ-sh3rf3 reduced the expression of GATA-4, especially, the protein levels of GATA-6, a functionally redundant transcription factor with GATA-4, were also down-regulated (Fig. 5E). Furthermore, knockdown of GATA-4 resulted in elevated expression of miR-29a (Fig. 5F), indicating that circ-sh3rf3 activated the expression of miR-29a through inhibition of GATA-4 expression.
To further investigate the role of circ-sh3rf3 in the heart following isoproterenol treatment, we tested the mRNA level of GATA-4, ANF, BNP, circ-sh3rf3, and miR-29a in the left ventricles of mice, rat cardiomyocytes, and cardiac fibroblasts treated with Iso and/or circ-sh3rf3 or circRNA empty vector. As shown in Fig. S1F–H, the result showed that over-expression of circ-sh3rf3 could reduce the expression of GATA-4, ANF, and BNP induced by isoproterenol, but its effect on the expression of miR-29a presented an opposite trend, indicating that circ-sh3rf3 play roles in the development of cardiac hypertrophy induced by isoproterenol via the regulation of miR-29a and GATA-4 expression.
Next, we evaluated the effect of RNAi GATA-4 on the trans-differentiation of myofibroblasts. The RNAi GATA-4 constructs were designed as previously described [49], and silencing of GATA-4 resulted in significant downregulation of CTGF, TGF-β1 and α-SMA in 10% FBS culture medium (Fig. 5G). Furthermore, inhibition of GATA-4 expression in TGF-β1-treated fibroblasts resulted in decreased expression of α-SMA and CTGF and marked changes in cell morphology, including decreased cell size and stress fibers (Fig. 5H–J), suggesting that downregulation of GATA-4 could attenuate the TGF-β1-induced differentiation of fibroblasts to myofibroblasts. Altogether, these data provide strong evidence that Circ-sh3rf3 inhibits fibroblast–myofibroblast differentiation and myocardial fibrosis through the upregulation of miR-29a via the suppression of GATA-4 expression (Fig. 6), thus identifying a potential target for the treatment of myocardial fibrosis.
Fig. 6.
Model depicting the role of the circ-sh3rf3-GATA-4-miR-29a regulatory axis in fibroblast–myofibroblast differentiation and myocardial fibrosis. This regulatory axis involves circ-sh3rf3 interacting with GATA-4 and inhibiting GATA-4 expression, which subsequently elevated miR-29a expression by abolishing the suppression of miR-29a expression mediated by GATA-4, thus further inhibiting fibroblast-myofibroblast differentiation and myocardial fibrosis
Discussion
Our study provides a new circ-sh3rf3/GATA-4/miR-29a regulatory axis in fibroblast–myofibroblast differentiation and myocardial fibrosis. We found that circ-sh3rf3 could suppress GATA-4 expression, which subsequently resulted in the upregulation of miR-29a due to the abolishment of miR-29a expression inhibition by GATA-4 and eventually inhibited the differentiation from fibroblasts to myofibroblasts. Given that few studies have focused on the roles of circular RNA in the differentiation of fibroblasts to myofibroblasts and that the function of circular RNA in this conversion is a very intriguing and crucial question which must be answered to uncover the mechanisms of myocardial fibrosis, our data provide a molecular basis for understanding myocardial fibrosis.
The anti-fibrotic effects of circ-sh3rf3 may be attributed to several underlying mechanisms, including both direct effects of the circ-sh3rf3/GATA-4/miR-29a regulatory cascade and indirect effects mediated by this regulatory cascade. The direct effects indicated that circ-sh3rf3 reduced GATA-4 expression and activated miR-29a expression, while both down-regulated GATA-4 and up-regulated miR-29a resulted in inhibited the differentiation of fibroblasts to myofibroblasts. In addition, the downstream target genes regulated by GATA-4 or miR-29a as well as their interacting signaling molecules may be involved in the antifibrotic effects mediated by the circ-sh3rf3/GATA-4/miR-29a regulatory axis, as evidenced by the finding that atrial and brain natriuretic peptide and antiangiogenic matricellular protein thrombospondin 2, which are regulated by GATA-4 and miR-29a, respectively, all suppress cardiac fibrosis [10, 16, 19, 38]. Therefore, circ-sh3rf3 plays inhibitory roles in myocardial fibrosis via a complicated regulatory network consisting of the circ-sh3rf3/GATA-4/miR-29a regulatory cascade and its functionally related molecules.
In addition, we found that circ-sh3rf3 could interact with GATA-4 and alleviate the inhibited expression of miR-29a mediated by GATA-4. Considering our findings that circ-sh3rf3 could reduce the enrichment of GATA-4 on the promoter regions of miR-29a, we speculated that circ-sh3rf3 could segregate the binding of GATA-4 to the miR-29a promoter. Moreover, circ-sh3rf3 inhibited the expression of GATA-4, which in turn abolished the inhibition of miR-29a expression mediated by GATA-4. A mechanistic link between circ-sh3rf3 and reduced GATA-4 expression suggests that circ-sh3rf3 may be involved in the post-transcriptional regulation and post-translational modification of GATA-4, including mRNA degradation and protein ubiquitination. In particular, back-splicing may compete with canonical splicing during the generation of circ-sh3rf3, so the altered expression level of circ-sh3rf3 in the process of cardiac fibroblast-to-myofibroblast conversion may affect the expression level of sh3rf3, which is an E3 ubiquitin–protein ligase, thereby mediating GATA-4 protein degradation through ubiquitination modification. On the other hand, circ-sh3rf3 may have the potential to encode proteins which may have the activity of ubiquitination modification enzymes, thereby regulating the degradation of GATA-4 protein. However, the exact mechanism remains to be elucidated in future investigations. As GATA-4 is an important transcription factor in physiological and pathological processes in the heart [6], we identified the central role of GATA-4 in the regulatory cascade of cardiac fibrosis is involving circ-sh3rf3 and miR-29a. To our knowledge, it is the first time to explore the roles of GATA-4 as a primary regulator of fibroblast–myofibroblast differentiation. Indeed, we found that silencing of GATA-4 significantly inhibited the expression of α-SMA, CTGF and TGF-β1, all crucial markers of myofibroblasts. This is in accordance with our previous finding that GATA-4 is up-regulated in ISO-induced myocardial hypertrophy in mice [53], further confirming the important role of GATA-4 in myocardial fibrosis. Taken together, the functional consequence of determining its critical roles lies in the benefit of identifying GATA-4 as a potential therapeutic target in cardiac fibrosis.
In addition to GATA-4, circ-sh3rf3 also regulates miR-29a expression. The fact that circ-sh3rf3 acts as an upstream regulator of miR-29a may be due to several underlying mechanisms, including both direct regulation via GATA-4 and indirect regulation via the GATA-4 partner. These partners may include MEF2C, TBX5, and SWI/SNF chromatin-remodeling complexes [1, 14, 29]. However, the exact role of the GATA-4 partner in the regulation of miR-29a expression by circ-sh3rf3 needs to be further investigated in the future. Impressively, a mechanistic link between circ-sh3rf3 and inhibited conversion from cardiac fibroblasts to myofibroblasts is suggested by the increased miR-29a expression, which is consistent with previous findings that miR-29a inhibits cardiac fibrosis [34, 42]. As a regulator of cardiac fibrosis, the mechanism of miR-29a remains unclear. Here, we revealed that miR-29a is a novel inhibitor of fibroblast–myofibroblast differentiation, which further supports our previous finding that miR-29a exerts beneficial effects on cardiac hypertrophy [53]. Since there are no binding sites of miR-29a in the circ-sh3rf3 sequence and the RNA pull-down assay revealed that miR-29a could not bind circ-sh3rf3, unlike most circular RNAs, circ-sh3rf3 cannot act as a sponge of miR-29a but instead regulates the expression of miR-29a, which may contribute to opening up a new mode of action for circular RNAs.
The difficulty of precisely constructing gain and loss circular RNA mice has hindered studies of the function of circular RNAs. However, recent studies have used the adenovirus infection technique in cells to characterize the roles of circular RNAs in cardiac fibrosis [50]. Impressively, several circular RNAs play roles in myocardial fibrosis [30, 50]. Indeed, our data further supported that circ-sh3rf3 acts as a cardiac fibrosis suppressor. Moreover, consistent with our experimental findings that circ-sh3rf3 acts as a sponge of GATA-4, many circRNAs function by acting as protein sponges [21, 36]. In fact, until now, the potential mechanism and function of circular RNA have not been fully explored. The function of circRNAs has been proposed as protein interactors, microRNAs, and protein sponges [13, 24, 36]. Some circRNAs have coding functions or function through their parent genes [23, 46]. For circ-sh3rf3, which is derived from exons 2 and 3 of the Sh3rf3 gene, how it is produced and whether it could encode a protein or play roles via the Sh3rf3 gene need to be further explored in the future. Moreover, the detailed mechanism, whereby circ-sh3rf3 regulates the conversion from fibroblasts to myofibroblasts, especially for the altered expression of markers of myofibroblasts such as CTGF and α-SMA, remains to be elucidated.
In conclusion, we herein identified the regulatory cascade involving circ-sh3rf3/GATA-4/miR-29a as a novel regulator of fibroblast–myofibroblast differentiation and myocardial fibrosis. Circ-sh3rf3 inhibits the differentiation from fibroblasts to myofibroblasts through the inhibition of GATA-4 and the activation of miR-29a. These findings pave the way to unveil clinical relevance for the treatment of myocardial fibrosis.
Materials and methods
Animals
ICR mice (8–10 weeks) were intraperitoneally injected with isoproterenol hydrochloride (ISO, 30 mg/kg/day; Sigma, St. Louis, MO, USA) for 14 days, and the vehicle group was treated with saline. Following treatment with isoproterenol, ICR mice received infusion of miR-29a agomir (6 µg/g/day) or agomir negative control (NC) for 3 days by intravenous injection. At the end of the infusion, the mice were anesthetized with 2.5% Avertin (2 mg/0.01 kg) by intraperitoneal injection, once unresponsive to toe pinch, mice were euthanized by heart collection or perfusion fixation with neutral formaldehyde for histological analysis. All animal experiments were approved by the Animal Ethics Committee of Zhengzhou University and carried out in accordance with the Guide for the Care and Use of Laboratory Animals (US NIH, 2011).
Isolation of rat cardiomyocytes and cardiac fibroblasts
Rat cardiomyocytes and cardiac fibroblasts were isolated from neonatal 1–2-day-old Sprague–Dawley rats by enzymatic digestion. Briefly, hearts were isolated after decapitation. Next, the dissected ventricles were finely minced and digested with 200 U/ml collagenase II (Worthington) solution, and the digested cells were harvested by centrifugation and resuspended in Dulbecco’s modified Eagle’s medium (DMEM, Gibco) with 5% fetal bovine serum and 1% penicillin/streptomycin solution. For separation of cardiomyocytes and cardiac fibroblasts, two cycles of 30 min pre-plating were carried out at 37 °C in an incubator with 5% CO2, and cardiac fibroblasts were first adherent to the culture plates. After serum starvation for 24 h, cardiac fibroblasts at passages 2–3 were treated with TGF-β1 (5 ng/ml; PeproTech, Rocky Hill, NJ, USA) or isoproterenol (10 μM; Sigma) for 48 h.
Infection with recombinant adenoviruses and transfection
Cardiac fibroblasts were infected with rAd-GFP or rAd-circ-sh3rf3 adenovirus (Genechem, Shanghai, China) at a multiplicity of infection (MOI) of 100 for 48 h in culture medium without antibiotics. For the transfection experiments, cardiac fibroblasts were transfected with miR-29a agomir, antagomir or their negative control (50 nM; GenePharma, Shanghai, China) for 24 h using Lipofectamine 2000 (Invitrogen). After transfection, the cells were starved and then treated with 5 ng/ml TGF-β1 for 48 h.
Total RNA isolation and quantitative polymerase chain reaction (qPCR)
Total RNA was extracted from cultured cardiac fibroblasts or left ventricles of mice with TRIzol reagent (Invitrogen). Next, reverse transcription was performed using the PrimeScript™ RT reagent Kit (Takara, Tokyo, Japan), and the expression levels of mRNA, miRNA and circRNA were determined by SYBR Green Master Mix (Takara). Notably, the reverse transcription of circRNA was specifically performed with random primers (Thermo Fisher), and its expression level was determined by qPCR using divergent primers. A list of the sequences of the primers is shown in Table 1.
Table 1.
Primer sequences
| Gene | Primer | Sequence |
|---|---|---|
| MMP-2 | Forward | GCACCACCGAGGATTATGAC |
| Reverse | CACCCACAGTGGACATAGCA | |
| MMP-9 | Forward | CCTCTGCATGAAGACGACATAA |
| Reverse | GGTCAGGTTTAGAGCCACGA | |
| COL1A1 | Forward | CATGTTCAGCTTTGTGGACCT |
| Reverse | GCAGCTGACTTCAGGGATGT | |
| COL3A1 | Forward | TCCCCTGGAATCTGTGAATC |
| Reverse | TGAGTCGAATTGGGGAGAAT | |
| TGFβ1 | Forward | CCTGGAAAGGGCTCAACAC |
| Reverse | CAGTTCTTCTCTGTGGAGCTGA | |
| α-SMA | Forward | CTGTGCTATGTCGCTCTGGA |
| Reverse | ATAGGTGGTTTCGTGGATGC | |
| CTGF | Forward | CTGTGAGGAGTGGGTGTG |
| Reverse | ATGTGTCTTCCAGTCGGTAGG | |
| miR-29a | Forward | GCGGTAGCACCATCTGAAAT |
| Reverse | GTGCAGGGTCCGAGGT | |
| U6 | Forward | CTCGCTTCGGCAGCACA |
| Reverse | AACGCTTCACGAATTTGCGT | |
| circ-sh3rf3 (divergent) | Forward | AACTGTGATCCGGAGAGTGG |
| Reverse | GGATGATGATGTCCCCTTTG | |
| circ-sh3rf3 (convergent) | Forward | AGGGGAAAGAACCTGGTGAC |
| Reverse | CATAAAGTGCTTTGCCTTGG | |
| miR-29a distal promoter | Forward | CAAGTCCTGGTGTCCCTAAC |
| Reverse | CGGTCTGTTCTTGCGTGAG | |
| miR-29a proximal promoter | Forward | CTGCTTACCTCGGTGTTGTG |
| Reverse | GGGCCTTCTGTCTGTTGTAC |
The subcellular localization analysis of circular RNA
Cardiac fibroblasts were harvested and resuspended in hypotonic buffer (20 mM HEPES pH 7.9, 20 mM sodium fluoride, 1 mM sodium pyrophosphate, 1 mM sodium orthovanadate, 1 mM EDTA, 1 mM EGTA, 0.25 mM sodium molybdate, 10 μg/ml leupeptin, 10 μg/ml aprotinin, 10 μg/ml pepstatin, 2 mM DTT, 0.5 mM PMSF and 100 nM okadaic acid) for 15 min. Next, 10% NP-40 were added, and the cytoplasm and nucleus were separated by centrifugation at 7000 r.p.m. at 4 °C, subsequently, RNA was extracted from the nuclear and cytoplasmic fractions.
Western blotting analysis
Western blotting experiments were performed as previously described [27]. Briefly, proteins were separated by SDS–PAGE, transferred onto PVDF membranes (Millipore) and incubated with primary antibodies specific for α-SMA (ab7817, Abcam), CTGF (ab6992, Abcam), MMP-2 (ab86607, Abcam), TGF-β1 (ab92486, Abcam), and α-tubulin (sc-32293, Santa Cruz). Relative expression levels of proteins were normalized to the α-tubulin expression level.
Immunofluorescence and F-actin staining assay
Cardiac fibroblasts were plated in 35-mm confocal dishes. Following TGF-β1 treatment or transfection as described above, cells were washed twice with PBS and fixed with 4% paraformaldehyde (Sigma) for 10 min. For the immunofluorescence assay, fixed cells were permeabilized with 0.1% Triton X-100 for 5 min, blocked with 5% BSA for 1 h and incubated with primary antibody (1:500) at 4 °C overnight. The next day, the cells were incubated with fluorescent-labeled secondary antibody (1:500) at room temperature for 2 h in the dark. The nuclei were stained with 4′,6‐diamidino‐2‐phenylindole (DAPI) for 15 min. For F-actin staining, fixed cells were treated with 0.1% Triton X-100 (containing 3 mg/mL BSA) for 30 min at 37 °C and incubated in a 1 µM solution of FITC-conjugated phalloidin (Sigma) in PBS at 4 °C overnight. Twenty-four h later, the nuclei were stained with DAPI and observed under a laser confocal microscope (OLYMPUS, Tokyo, Japan).
RNA immunoprecipitation
RNA immunoprecipitation was carried out to detect the binding of circRNA and protein as described [8]. Briefly, cultured cardiac fibroblasts (2 × 107 cells) were harvested and resuspended in 700 μl coimmunoprecipitation buffer (20 mM Tris–HCL, pH 7.5, 150 mM NaCl, 1 mM EDTA, 0.5% NP-40, protease inhibitors and RNase inhibitor) on ice for 30 min. After centrifugation, the supernatant was collected and incubated with 5 μg primary antibodies (IgG, anti-GATA4) at 4 °C overnight. Then, 80 μl proteinA/G PLUS-Agarose (sc-2003, Santa Cruz) was added to each sample, and the complexes were incubated at 4 °C overnight. After being washed, the pellets were resuspended in TRIzol-LS Reagent (Invitrogen). The coprecipitated RNA was extracted and subjected to RT–qPCR analysis as described above.
RNA pull-down
The RNA pull-down experiment was performed as described [45]. Briefly, 2 × 107 cardiac fibroblasts were washed and lysed in coimmunoprecipitation buffer. Three micrograms of biotinylated DNA oligo probes against circ-sh3rf3 or random sequences were diluted in 500 μl washing/binding buffer (20 mM Tris–HCl, pH 7.5, 500 mM NaCl, 1 mM EDTA) and incubated with 100 μl of Dynabeads M-280 Streptavidin (Invitrogen) at room temperature for 2 h. Then, the cell lysates were added to each bead-probe pellet and incubated at 4 °C overnight. The next day, the beads were washed briefly with coimmunoprecipitation buffer five times, and the bound proteins were analyzed by Western blotting.
Chromatin immunoprecipitation (ChIP)
ChIP assays were performed as previously described [27, 52]. Briefly, 1 × 107 cardiac fibroblasts transfected with circ-sh3rf3, circRNA negative control and/or GATA-4 were cross-linked in 1% formaldehyde for 15 min at 4 °C and quenched with 125 mM glycine. Then, the cells were collected, washed and sonicated to fragments of approximately 500 bp. Next, the DNA fragments in the sonicated chromatin solution were immunoprecipitated with antibodies against GATA-4 (ab134057, Abcam) or IgG (2729 s, Cell Signaling) overnight at 4 °C, followed by incubation with protein A/G plus agarose overnight at 4 °C. The next day, the chromatin–protein–antibody–bead complexes were washed and eluted. Then, the protein/DNA complex was reversely cross-linked, purified with the QIAquick PCR purification kit (Qiagen) and analyzed by qPCR. Specific primer sets designed to amplify target regions within the rat miR-29a promoter are listed in Table 1.
Statistical analysis
All data are presented as the mean ± SEM. Unpaired Student’s t-tests were used to compare the differences between two groups. One-way analysis of variance was used for multiple comparisons. In all cases, P < 0.05 was considered statistically significant.
Supplementary Information
Below is the link to the electronic supplementary material.
Acknowledgements
We are grateful to members of the Zang’s lab for helpful discussions. We thank Xiu-Hua Ren for histological sections.
Author contributions
YQ and M-XZ conceived, designed, and supervised the study. C-XM, Zh-RW, K-LK, Y-QS, M-HY, J-CS, M-JZ, TS, and Z-WW performed the experiments. JC, WL, T-QW, S-FZ, LX, and MZ provided the technical support and contributed to the discussion of the project and article. YQ and M-XZ analysed the data and wrote the article.
Funding
This study was supported by the National Natural Science Foundation of China (Nos. 82072975, 81771631, and 32171179), the High-Level Talents of Henan Province, particularly the support for the Central Plains Thousand Talents Program, which are the leading talents of Central Plains Basic Research (ZYQR201810120); the key project of discipline construction of Zhengzhou University in 2020 (XKZDQY202002), 2022 Henan Province Science and Technology R&D Program Joint Fund (Cultivation of Superior Disciplines, 222301420094), and Henan Province Science and Technology Projects (202102310057).
Data availability
All data generated during this study are included in this published article and its supplementary information files.
Declarations
Competing interest
The authors declare no competing interest.
Ethics approval and consent to participate
All animal experiments were approved by the Animal Ethics Committee of Zhengzhou University and carried out in accordance with the Guide for the Care and Use of Laboratory Animals (US NIH, 2011).
Consent for publication
The author’s consent to publication.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Qianqian Yin, Email: yinqianqian@bjmu.edu.cn.
Ming-Xi Zang, Email: mzang@zzu.edu.cn.
References
- 1.Bevilacqua A, Willis MS, Bultman SJ. SWI/SNF chromatin-remodeling complexes in cardiovascular development and disease. Cardiovasc Pathol. 2014;23:85–91. doi: 10.1016/j.carpath.2013.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Caja L, Dituri F, Mancarella S, Caballero-Diaz D, Moustakas A, Giannelli G, Fabregat I. TGF-beta and the tissue microenvironment: relevance in fibrosis and cancer. Int J Mol Sci. 2018;19:1294. doi: 10.3390/ijms19051294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chothani S, Schafer S, Adami E, Viswanathan S, Widjaja AA, Langley SR, Tan J, Wang M, Quaife NM, Jian Pua C, D'Agostino G, Guna Shekeran S, George BL, Lim S, Yiqun Cao E, van Heesch S, Witte F, Felkin LE, Christodoulou EG, Dong J, Blachut S, Patone G, Barton PJR, Hubner N, Cook SA, Rackham OJL. Widespread translational control of fibrosis in the human heart by RNA-binding proteins. Circulation. 2019;140:937–951. doi: 10.1161/CIRCULATIONAHA.119.039596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.de Salvi GF, de Moraes WM, Bozi LH, Souza PR, Antonio EL, Bocalini DS, Tucci PJ, Ribeiro DA, Brum PC, Medeiros A. Dexamethasone-induced cardiac deterioration is associated with both calcium handling abnormalities and calcineurin signaling pathway activation. Mol Cell Biochem. 2017;424:87–98. doi: 10.1007/s11010-016-2846-3. [DOI] [PubMed] [Google Scholar]
- 5.Dey S, Kwon JJ, Liu S, Hodge GA, Taleb S, Zimmers TA, Wan J, Kota J. miR-29a is repressed by MYC in pancreatic cancer and its restoration drives tumor-suppressive effects via downregulation of LOXL2. Mol Cancer Res MCR. 2020;18:311–323. doi: 10.1158/1541-7786.MCR-19-0594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Dobrzycki T, Lalwani M, Telfer C, Monteiro R, Patient R. The roles and controls of GATA factors in blood and cardiac development. IUBMB Life. 2020;72:39–44. doi: 10.1002/iub.2178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Driesen RB, Nagaraju CK, Abi-Char J, Coenen T, Lijnen PJ, Fagard RH, Sipido KR, Petrov VV. Reversible and irreversible differentiation of cardiac fibroblasts. Cardiovasc Res. 2014;101:411–422. doi: 10.1093/cvr/cvt338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Du WW, Yang W, Chen Y, Wu ZK, Foster FS, Yang Z, Li X, Yang BB. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur Heart J. 2017;38:1402–1412. doi: 10.1093/eurheartj/ehw001. [DOI] [PubMed] [Google Scholar]
- 9.Du WW, Zhang C, Yang W, Yong T, Awan FM, Yang BB. Identifying and characterizing circRNA-protein interaction. Theranostics. 2017;7:4183–4191. doi: 10.7150/thno.21299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Ellmers LJ, Scott NJ, Piuhola J, Maeda N, Smithies O, Frampton CM, Richards AM, Cameron VA. Npr1-regulated gene pathways contributing to cardiac hypertrophy and fibrosis. J Mol Endocrinol. 2007;38:245–257. doi: 10.1677/jme.1.02138. [DOI] [PubMed] [Google Scholar]
- 11.Eyholzer M, Schmid S, Wilkens L, Mueller BU, Pabst T. The tumour-suppressive miR-29a/b1 cluster is regulated by CEBPA and blocked in human AML. Br J Cancer. 2010;103:275–284. doi: 10.1038/sj.bjc.6605751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fu DJ, Song J, Zhu T, Pang XJ, Wang SH, Zhang YB, Wu BW, Wang JW, Zi X, Zhang SY, Liu HM. Discovery of novel tertiary amide derivatives as NEDDylation pathway activators to inhibit the tumor progression in vitro and in vivo. Eur J Med Chem. 2020;192:112153. doi: 10.1016/j.ejmech.2020.112153. [DOI] [PubMed] [Google Scholar]
- 13.Garikipati VNS, Verma SK, Cheng Z, Liang D, Truongcao MM, Cimini M, Yue Y, Huang G, Wang C, Benedict C, Tang Y, Mallaredy V, Ibetti J, Grisanti L, Schumacher SM, Gao E, Rajan S, Wilusz JE, Goukassian D, Houser SR, Koch WJ, Kishore R. Circular RNA CircFndc3b modulates cardiac repair after myocardial infarction via FUS/VEGF-A axis. Nat Commun. 2019;10:4317. doi: 10.1038/s41467-019-11777-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Georges R, Nemer G, Morin M, Lefebvre C, Nemer M. Distinct expression and function of alternatively spliced Tbx5 isoforms in cell growth and differentiation. Mol Cell Biol. 2008;28:4052–4067. doi: 10.1128/MCB.02100-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gurtner GC, Werner S, Barrandon Y, Longaker MT. Wound repair and regeneration. Nature. 2008;453:314–321. doi: 10.1038/nature07039. [DOI] [PubMed] [Google Scholar]
- 16.Hsu CH, Liu IF, Kuo HF, Li CY, Lian WS, Chang CY, Chen YH, Liu WL, Lu CY, Liu YR, Lin TC, Lee TY, Huang CY, Hsieh CC, Liu PL. miR-29a-3p/THBS2 axis regulates PAH-induced cardiac fibrosis. Int J Mol Sci. 2021 doi: 10.3390/ijms221910574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jiang Q, Liu C, Li CP, Xu SS, Yao MD, Ge HM, Sun YN, Li XM, Zhang SJ, Shan K, Liu BH, Yao J, Zhao C, Yan B. Circular RNA-ZNF532 regulates diabetes-induced retinal pericyte degeneration and vascular dysfunction. J Clin Investig. 2020;130:3833–3847. doi: 10.1172/JCI123353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Jie M, Wu Y, Gao M, Li X, Liu C, Ouyang Q, Tang Q, Shan C, Lv Y, Zhang K, Dai Q, Chen Y, Zeng S, Li C, Wang L, He F, Hu C, Yang S. CircMRPS35 suppresses gastric cancer progression via recruiting KAT7 to govern histone modification. Mol Cancer. 2020;19:56. doi: 10.1186/s12943-020-01160-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Koivisto E, Kaikkonen L, Tokola H, Pikkarainen S, Aro J, Pennanen H, Karvonen T, Rysa J, Kerkela R, Ruskoaho H. Distinct regulation of B-type natriuretic peptide transcription by p38 MAPK isoforms. Mol Cell Endocrinol. 2011;338:18–27. doi: 10.1016/j.mce.2011.02.015. [DOI] [PubMed] [Google Scholar]
- 20.Kong P, Christia P, Frangogiannis NG. The pathogenesis of cardiac fibrosis. Cell Mol Life Sci. 2013;71:549–574. doi: 10.1007/s00018-013-1349-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–691. doi: 10.1038/s41576-019-0158-7. [DOI] [PubMed] [Google Scholar]
- 22.Kudej RK, Iwase M, Uechi M, Vatner DE, Oka N, Ishikawa Y, Shannon RP, Bishop SP, Vatner SF. Effects of chronic beta-adrenergic receptor stimulation in mice. J Mol Cell Cardiol. 1997;29:2735–2746. doi: 10.1006/jmcc.1997.0508. [DOI] [PubMed] [Google Scholar]
- 23.Lei M, Zheng G, Ning Q, Zheng J, Dong D. Translation and functional roles of circular RNAs in human cancer. Mol Cancer. 2020;19:30. doi: 10.1186/s12943-020-1135-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Li M, Ding W, Tariq MA, Chang W, Zhang X, Xu W, Hou L, Wang Y, Wang J. A circular transcript of ncx1 gene mediates ischemic myocardial injury by targeting miR-133a-3p. Theranostics. 2018;8:5855–5869. doi: 10.7150/thno.27285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu D, Li S, Cui Y, Tong H, Li S, Yan Y. Podocan affects C2C12 myogenic differentiation by enhancing Wnt/beta-catenin signaling. J Cell Physiol. 2019;234:11130–11139. doi: 10.1002/jcp.27763. [DOI] [PubMed] [Google Scholar]
- 26.Liu Y, Afzal J, Vakrou S, Greenland GV, Talbot CC, Jr, Hebl VB, Guan Y, Karmali R, Tardiff JC, Leinwand LA, Olgin JE, Das S, Fukunaga R, Abraham MR. Differences in microRNA-29 and pro-fibrotic gene expression in mouse and human hypertrophic cardiomyopathy. Front Cardiovasc Med. 2019;6:170. doi: 10.3389/fcvm.2019.00170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ma CX, Song YL, Xiao L, Xue LX, Li WJ, Laforest B, Komati H, Wang WP, Jia ZQ, Zhou CY, Zou Y, Nemer M, Zhang SF, Bai X, Wu H, Zang MX. EGF is required for cardiac differentiation of P19CL6 cells through interaction with GATA-4 in a time- and dose-dependent manner. Cell Mol Life Sci CMLS. 2015;72:2005–2022. doi: 10.1007/s00018-014-1795-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Memczak S, Jens M, Elefsinioti A, Torti F, Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer M, Loewer A, Ziebold U, Landthaler M, Kocks C, le Noble F, Rajewsky N. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495:333–338. doi: 10.1038/nature11928. [DOI] [PubMed] [Google Scholar]
- 29.Morin S, Charron F, Robitaille L, Nemer M. GATA-dependent recruitment of MEF2 proteins to target promoters. EMBO J. 2000;19:2046–2055. doi: 10.1093/emboj/19.9.2046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ni H, Li W, Zhuge Y, Xu S, Wang Y, Chen Y, Shen G, Wang F. Inhibition of circHIPK3 prevents angiotensin II-induced cardiac fibrosis by sponging miR-29b-3p. Int J Cardiol. 2019;292:188–196. doi: 10.1016/j.ijcard.2019.04.006. [DOI] [PubMed] [Google Scholar]
- 31.Park S, Ranjbarvaziri S, Lay FD, Zhao P, Miller MJ, Dhaliwal JS, Huertas-Vazquez A, Wu X, Qiao R, Soffer JM, Rau C, Wang Y, Mikkola HKA, Lusis AJ, Ardehali R. Genetic regulation of fibroblast activation and proliferation in cardiac fibrosis. Circulation. 2018;138:1224–1235. doi: 10.1161/CIRCULATIONAHA.118.035420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Park SW, Persaud SD, Ogokeh S, Meyers TA, Townsend D, Wei LN. CRABP1 protects the heart from isoproterenol-induced acute and chronic remodeling. J Endocrinol. 2018;236:151–165. doi: 10.1530/JOE-17-0613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Paz I, Kosti I, Ares M, Jr, Cline M, Mandel-Gutfreund Y. RBPmap: a web server for mapping binding sites of RNA-binding proteins. Nucleic Acids Res. 2014;42:W361–367. doi: 10.1093/nar/gku406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Qin RH, Tao H, Ni SH, Shi P, Dai C, Shi KH. microRNA-29a inhibits cardiac fibrosis in Sprague–Dawley rats by downregulating the expression of DNMT3A. Anatol J Cardiol. 2018;20:198–205. doi: 10.14744/AnatolJCardiol.2018.98511. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rodriguez P, Sassi Y, Troncone L, Benard L, Ishikawa K, Gordon RE, Lamas S, Laborda J, Hajjar RJ, Lebeche D. Deletion of delta-like 1 homologue accelerates fibroblast-myofibroblast differentiation and induces myocardial fibrosis. Eur Heart J. 2019;40:967–978. doi: 10.1093/eurheartj/ehy188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Schreiner S, Didio A, Hung LH, Bindereif A. Design and application of circular RNAs with protein-sponge function. Nucleic Acids Res. 2020;48:12326–12335. doi: 10.1093/nar/gkaa1085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tarbit E, Singh I, Peart JN, Rose'Meyer RB. Biomarkers for the identification of cardiac fibroblast and myofibroblast cells. Heart Fail Rev. 2019;24:1–15. doi: 10.1007/s10741-018-9720-1. [DOI] [PubMed] [Google Scholar]
- 38.Temsah R, Nemer M. GATA factors and transcriptional regulation of cardiac natriuretic peptide genes. Regul Pept. 2005;128:177–185. doi: 10.1016/j.regpep.2004.12.026. [DOI] [PubMed] [Google Scholar]
- 39.Tian R, Guan X, Qian H, Wang L, Shen Z, Fang L, Liu Z. Restoration of NRF2 attenuates myocardial ischemia reperfusion injury through mediating microRNA-29a-3p/CCNT2 axis. BioFactors. 2021 doi: 10.1002/biof.1712. [DOI] [PubMed] [Google Scholar]
- 40.Travers JG, Kamal FA, Robbins J, Yutzey KE, Blaxall BC. Cardiac fibrosis: the fibroblast awakens. Circ Res. 2016;118:1021–1040. doi: 10.1161/CIRCRESAHA.115.306565. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wang K, Long B, Liu F, Wang JX, Liu CY, Zhao B, Zhou LY, Sun T, Wang M, Yu T, Gong Y, Liu J, Dong YH, Li N, Li PF. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J. 2016;37:2602–2611. doi: 10.1093/eurheartj/ehv713. [DOI] [PubMed] [Google Scholar]
- 42.Wang R, Peng L, Lv D, Shang F, Yan J, Li G, Li D, Ouyang J, Yang J. Leonurine attenuates myocardial fibrosis through upregulation of miR-29a-3p in mice post-myocardial infarction. J Cardiovasc Pharmacol. 2021;77:189–199. doi: 10.1097/FJC.0000000000000957. [DOI] [PubMed] [Google Scholar]
- 43.Wang Y, Li C, Zhao R, Qiu Z, Shen C, Wang Z, Liu W, Zhang W, Ge J, Shi B. CircUbe3a from M2 macrophage-derived small extracellular vesicles mediates myocardial fibrosis after acute myocardial infarction. Theranostics. 2021;11:6315–6333. doi: 10.7150/thno.52843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wang Y, Zhao R, Liu W, Wang Z, Rong J, Long X, Liu Z, Ge J, Shi B. Exosomal circHIPK3 released from hypoxia-pretreated cardiomyocytes regulates oxidative damage in cardiac microvascular endothelial cells via the miR-29a/IGF-1 pathway. Oxid Med Cell Longev. 2019;2019:7954657. doi: 10.1155/2019/7954657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wu N, Yuan Z, Du KY, Fang L, Lyu J, Zhang C, He A, Eshaghi E, Zeng K, Ma J, Du WW, Yang BB. Translation of yes-associated protein (YAP) was antagonized by its circular RNA via suppressing the assembly of the translation initiation machinery. Cell Death Differ. 2019;26:2758–2773. doi: 10.1038/s41418-019-0337-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Xu X, Zhang J, Tian Y, Gao Y, Dong X, Chen W, Yuan X, Yin W, Xu J, Chen K, He C, Wei L. CircRNA inhibits DNA damage repair by interacting with host gene. Mol Cancer. 2020;19:128. doi: 10.1186/s12943-020-01246-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Yang F, Hu A, Li D, Wang J, Guo Y, Liu Y, Li H, Chen Y, Wang X, Huang K, Zheng L, Tong Q. Circ-HuR suppresses HuR expression and gastric cancer progression by inhibiting CNBP transactivation. Mol Cancer. 2019;18:158. doi: 10.1186/s12943-019-1094-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Yang MH, Wang H, Han SN, Jia X, Zhang S, Dai FF, Zhou MJ, Yin Z, Wang TQ, Zang MX, Xue LX. Circular RNA expression in isoproterenol hydrochloride-induced cardiac hypertrophy. Aging (Albany NY) 2020;12:2530–2544. doi: 10.18632/aging.102761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yao CX, Wei QX, Zhang YY, Wang WP, Xue LX, Yang F, Zhang SF, Xiong CJ, Li WY, Wei ZR, Zou Y, Zang MX. miR-200b targets GATA-4 during cell growth and differentiation. RNA Biol. 2013;10:465–480. doi: 10.4161/rna.24370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Yuan S, Liang J, Zhang M, Zhu J, Pan R, Li H, Zeng N, Wen Y, Yi Z, Shan Z. CircRNA_005647 inhibits expressions of fibrosis-related genes in mouse cardiac fibroblasts via sponging miR-27b-3p. Nan Fang Yi Ke Da Xue Xue Bao. 2019;39:1312–1319. doi: 10.12122/j.issn.1673-4254.2019.11.08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yücel D, Solinsky J, van Berlo JH. Isolation of cardiomyocytes from fixed hearts for immunocytochemistry and ploidy analysis. J Vis Exp JoVE. 2020 doi: 10.3791/60938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Zhang S-B, Lin S-Y, Liu M, Liu C-C, Ding H-H, Sun Y, Ma C, Guo R-X, Lv Y-Y, Wu S-L, Xu T, Xin W-J. CircAnks1a in the spinal cord regulates hypersensitivity in a rodent model of neuropathic pain. Nat Commun. 2019;10:4119. doi: 10.1038/s41467-019-12049-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Zhang S, Yin Z, Dai FF, Wang H, Zhou MJ, Yang MH, Zhang SF, Fu ZF, Mei YW, Zang MX, Xue L. miR-29a attenuates cardiac hypertrophy through inhibition of PPARδ expression. J Cell Physiol. 2019;234:13252–13262. doi: 10.1002/jcp.27997. [DOI] [PubMed] [Google Scholar]
- 54.Zhou L, Miao K, Yin B, Li H, Fan J, Zhu Y, Ba H, Zhang Z, Chen F, Wang J, Zhao C, Li Z, Wang DW. Cardioprotective role of myeloid-derived suppressor cells in heart failure. Circulation. 2018;138:181–197. doi: 10.1161/CIRCULATIONAHA.117.030811. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All data generated during this study are included in this published article and its supplementary information files.