Abstract
The Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) variant BA.2.86 has over 30 mutations in spike compared with BA.2 and XBB.1.5, which raised the possibility that BA.2.86 might evade neutralizing antibodies (NAbs) induced by vaccination or infection. In this study, we show that NAb titers are substantially lower to BA.2.86 compared with BA.2 but are similar or slightly higher than to other current circulating variants, including XBB.1.5, EG.5.1, and FL.1.5.1. Moreover, NAb titers against all these variants were higher in vaccinated individuals with a history of XBB.1.5 infection compared with vaccinated individuals with no history of XBB.1.5 infection, suggesting the potential utility of the monovalent XBB.1.5 mRNA boosters.
Introduction
The continued evolution of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) often leads to increased evasion of vaccine immunity and natural immunity1–3. A highly mutated Omicron variant BA.2.86 has recently been identified with over 30 amino acid changes in Spike compared with BA.2 and XBB.1.5 (Figs. 1–3). As of September 30, 2023, BA.2.86 has been identified in 304 sequences from 24 countries (Fig. 1), which is likely an underestimate due to limited surveillance. The ability of BA.2.86 to evade neutralizing antibodies (NAbs) compared with other currently circulating Omicron variants remains unknown.
Figure 1. Transitions in SARS-CoV-2 variant lineages in 2023.
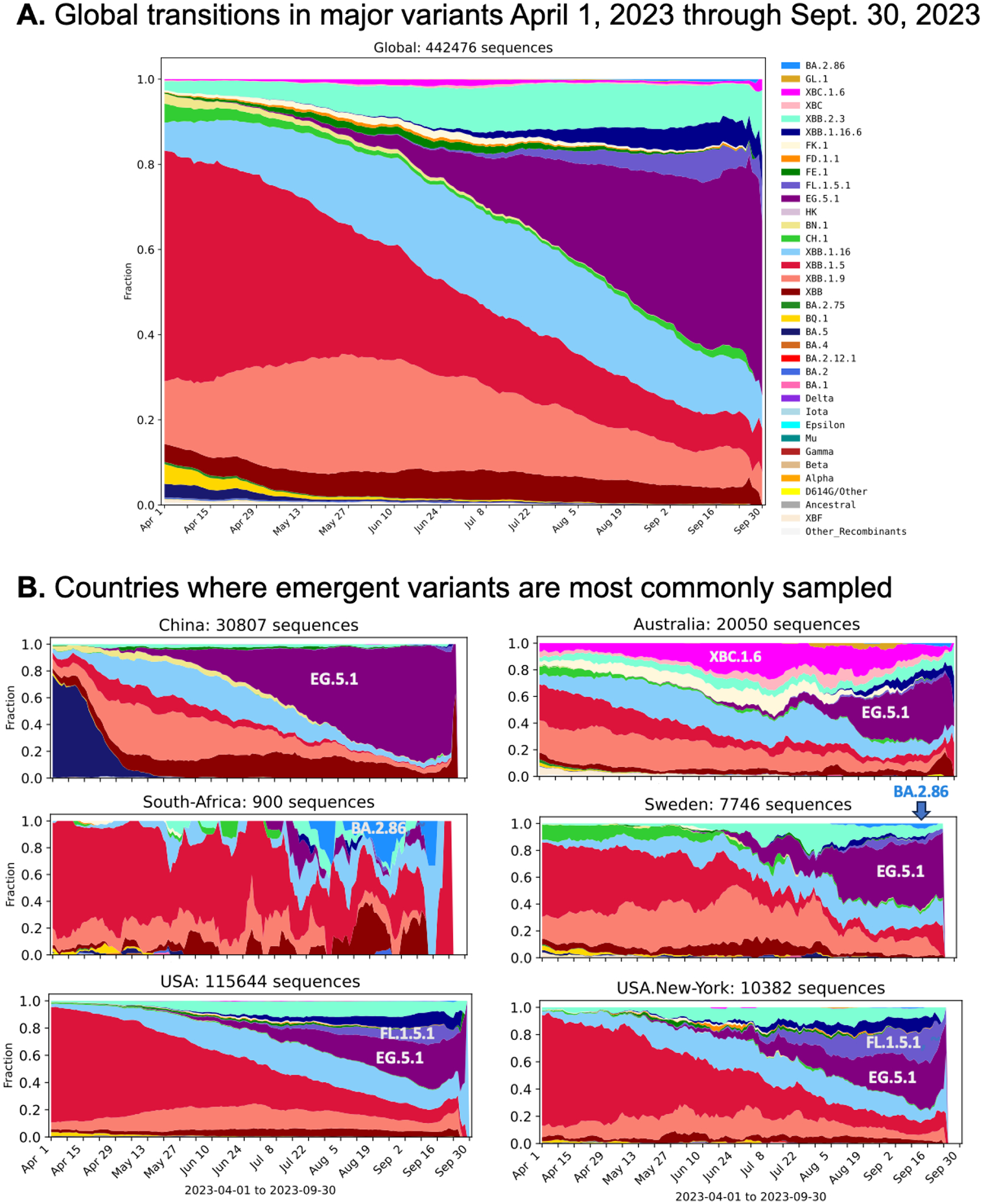
(A) Global frequencies based on GISAID data. Note that recent sampling is biased due to sparse sampling given the inevitable time delays between sampling and sequence submissions to GISAID. EG.5.1 sampling is currently increasing the fastest globally. (B) shows the countries were the emergent lineages are most sampled. EG.5.1 has become the dominant lineage in China, and is increasing everywhere it is sampled. XBC.1 and XBC.1.6 are common in Australia (magenta), but no longer increasing there. The recently detected highly divergent variant BA.2.86 has 66 sequences available in GISAID as of this writing, 2023-09-30; its sublineage BA.2.86.1 and has spread globally more quickly, and has been sampled 238 times. As BA.2.86.1 is distinguished by only two mutations (ORF1a:K1973R, C12815T), both outside of spike, both are grouped here and called BA.2.86. BA.2.86 can be visualized on the global plot in (A) as the thin blue line at the top. The county where BA.2.86 has the highest frequency of sampling is South Africa, while the country where BA.2.86.1 has the highest frequency of sampling is Sweden. The FL.1.5.1 sublineage is increasing in the USA, and New York is showing the most rapid increase in FL.1.5.1 sampling frequency at the state level which is highlighted here. FL.1.5.1 can be differentiated from FL.1 by 2 additional mutations in Spike, F456L and T478R. Of note, these are common mutations in currently expanding lineages.
Figure 3. Structural models of BA.2.86 mutations.
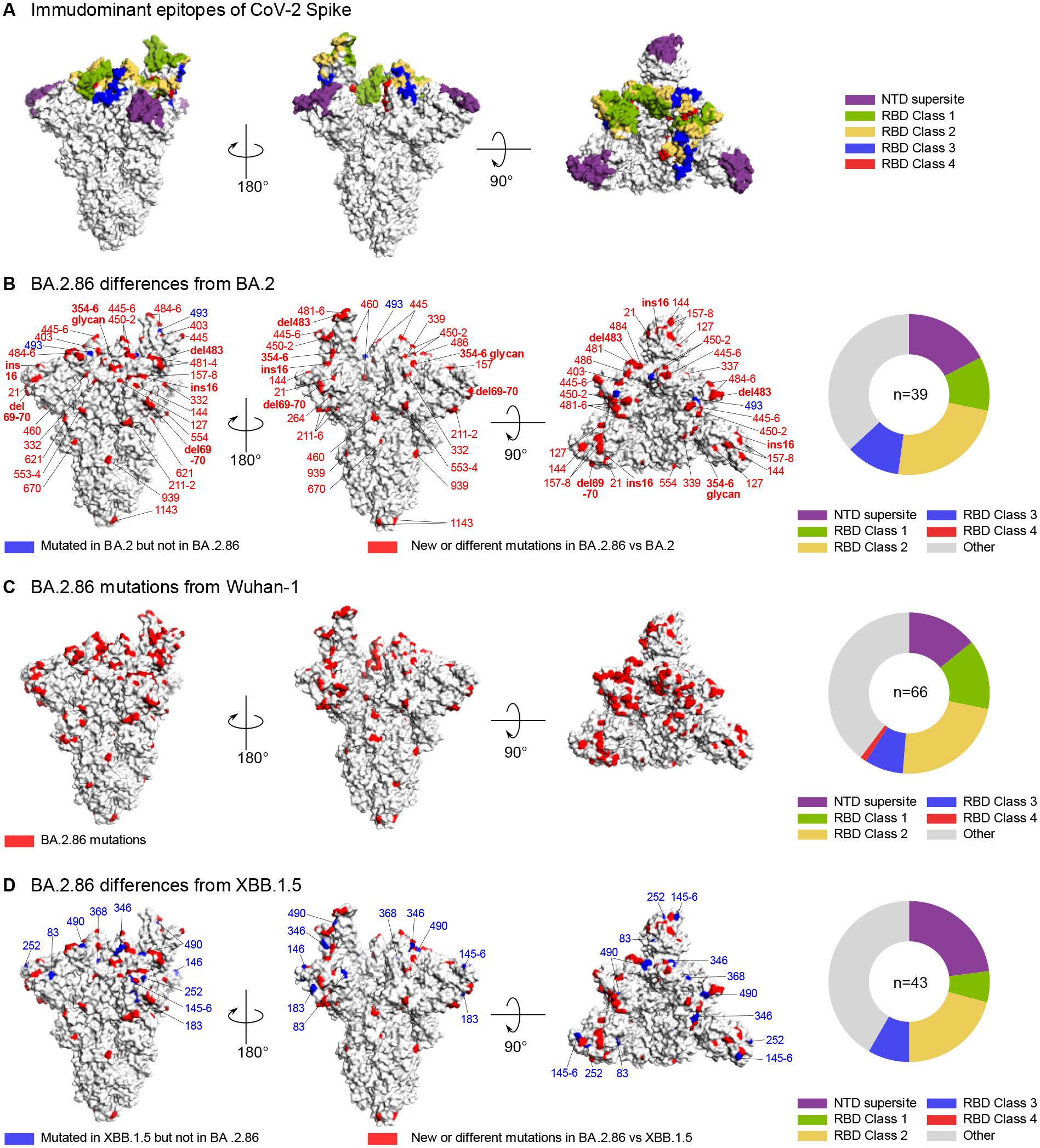
(A) Structural mapping of key neutralizing antibody epitopes. Spike trimer structure in the one-up configuration from Benton et al.10 (PDB: 7A94) is used. RBD class and epitope definitions are from Barnes et al.11, and NTD supersite epitope definitions are from Cerutti et al.12. Epitopes are color-coded per legend. (B-D) Mutations in BA.2.86 versus BA.2, Wuhan-1, and XBB.1.5. Sites with either new or different mutations in BA.2.86 with respect to the comparator variant are shown in red, and sites that have mutations in the comparator strain but not in BA.2.86 are shown in blue. An N-linked glycosylation site introduced in BA.2.86 at Spike N354 (resulting from a K356T change) and insertions and deletions are in bold as they may be particularly impactful. The pie charts on the right show the fraction of sites with amino acid differences between BA.2.86 and the comparator variant that fall in each of the epitopes from (A); grey indicates fraction of sites with sequence differences that did not fall in any of the epitopes from (A). The total number of sites with sequence differences is indicated in the center, and sites that occur in multiple epitopes are counted in each epitope.
Results
We evaluated NAb responses in 44 individuals who received the bivalent mRNA booster in fall 2022 and in 22 individuals who did not receive the bivalent mRNA booster (Tables 1, S1). Participants had a median of 3 Coronavirus Disease-19 (COVID-19) vaccine doses prior to the bivalent mRNA boost, and 70–73% had at least one documented SARS-CoV-2 infection. In participants who received the bivalent mRNA boost, baseline NAb responses against WA1/2020, BA.1, BA.2, BA.5, XBB.1.5, XBB.1.16, EG.5, EG.5.1, FL.1.5.1, XBC.1.6, and BA.2.86 were 16,172, 1287, 1594, 603, 149, 127, 157, 107, 65, 149, and 304, respectively (Fig. 4).
Table 1.
Study population.
| No Bivalent mRNA Boost N=22 |
Bivalent mRNA Boost N=44 |
|
|---|---|---|
| Age (years), median (range) | 36 (25–62) | 39 (23–73) |
| Sex at birth, Female | 19 (86) | 35 (80) |
| Race | ||
| White | 14 (64) | 38 (86) |
| Asian | 2 (9) | 3 (7) |
| Black | 1 (5) | 0 |
| More than one race | 3 (14) | 2 (5) |
| Other* | 2 (9) | 1 (2) |
| Ethnicity | ||
| Hispanic or Latino | 3 (14) | 3 (7) |
| Non-Hispanic | 19 (86) | 40 (91) |
| Declined | 0 | 1 (2) |
| Medical condition | ||
| Obesity (BMI ≥ 30 kg/m2) | 6 (27) | 8 (18) |
| Hypertension | 2 (9) | 8 (18) |
| Diabetes | 0 | 1 (2) |
| Pregnant^ | 6 (27) | 9 (20) |
| Asthma | 5 (23) | 2 (5) |
| Bivalent mRNA Boost | ||
| Pfizer | N/A | 15 (34) |
| Moderna | N/A | 29 (66) |
| COVID-19 Vaccines (Excluding Bivalent mRNA Boost) | ||
| BNT (3 doses) | 7 (32) | 8 (18) |
| BNT (3 doses) / 1273 | 0 | 2 (5) |
| BNT (3 doses) / Ad26 | 0 | 1 (2) |
| BNT (2 doses) / 1273 (2 doses) | 0 | 1 (2) |
| BNT (2 doses) / 1273 | 1 (5) | 4 (9) |
| BNT (2 doses) / Ad26 / 1273 | 0 | 1 (2) |
| BNT (2 doses) / Ad26 / BNT | 1 (5) | 2 (5) |
| BNT (2 doses) / Ad26 | 1 (5) | 2 (5) |
| 1273 (4 doses) | 1 (5) | 4 (9) |
| 1273 (3 doses) | 7 (32) | 10 (23) |
| 1273 (2 doses) | 2 (9) | 1 (2) |
| 1273 (2 doses) / Ad26 | 0 | 1 (2) |
| 1273 (2 doses) / BNT | 0 | 1 (2) |
| Ad26 / BNT (1 dose) | 0 | 1 (2) |
| NVX (3 doses) | 2 (9) | 0 |
| NVX (2 doses) / 1273 (3 doses) | 0 | 1 (2) |
| NVX (2 doses) / 1273 | 0 | 1 (2) |
| NVX (2 doses) / BNT | 0 | 1 (2) |
| NVX (2 doses) / Ad26 / BNT | 0 | 1 (2) |
| NVX (2 doses) / Ad26 / 1273 | 0 | 1 (2) |
| Days from bivalent boost to week 3 peak sampling | N/A | N=27 |
| 20 (16–37) | ||
| Days from last vaccine dose to month 6 durability sampling | N=22 | N=30 |
| 482 (388–559) | 179 (173–187) | |
| Known COVID-19 positive | N=16 | N=31 |
| 1 prior infection | 16 (73) | 24 (55) |
| 2 prior infections | 0 | 7 (16) |
| Days from last infection to month 6 durability sampling | N=16 | N=30 |
| 283 (106–331) | 248 (49–338) |
Data displayed as median (range or interquartile range, IQR) and n (%); BMI, body mass index; pregnant designation reflects time of last vaccine dose and/or time of sampling.
BNT=BNT162b2; 1273=mRNA-1273; Ad26=Ad26.COV2.S; NVX=Novavax
Other race: 1 Cape Verdean, 1 Hispanic, 1 declined to report
Pregnant during last vaccine dose and/or during month 6 sampling
Figure 4. Neutralization escape from SARS-CoV-2 variants.
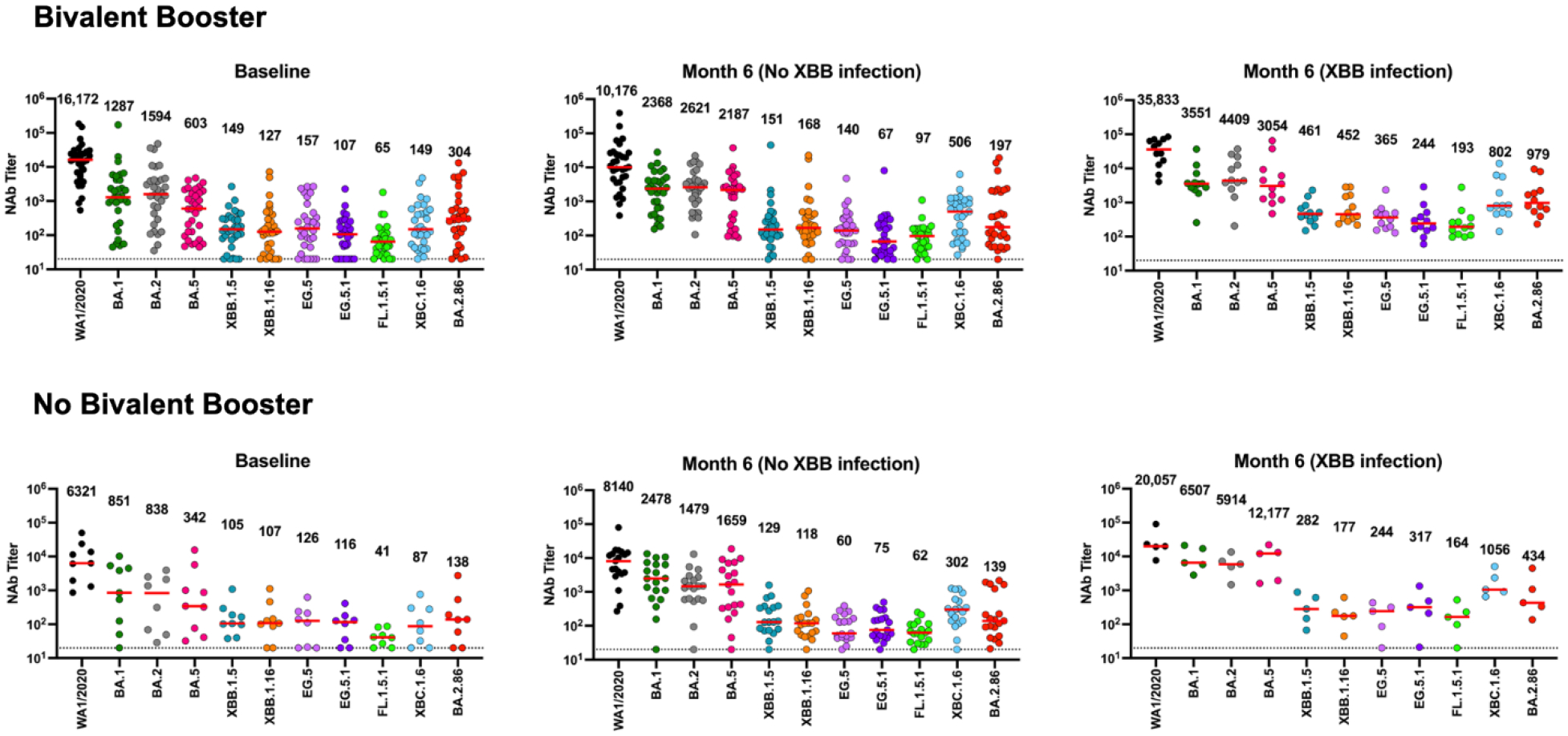
Neutralizing antibody (NAb) against the WA1/2020, BA.1, BA.2, BA.5, XBB.1.5, XBB.1.16, EG.5, EG.5.1, FL.1.5.1, XBC.1.6, and BA.2.86 variants by luciferase-based pseudovirus neutralization assays at baseline and at 6 months in individuals who did or did not receive the bivalent mRNA booster in fall 2022. Also shown are participants at 6 months who were infected during the XBB.1.5 predominant period. The horizontal red bar reflects median values. The bivalent mRNA booster cohort with no XBB infection had n=32 participants, and the bivalent mRNA booster cohort with XBB infections had n=12 participants. The no bivalent booster cohort with no XBB infection had n=9–19 participants, and the no bivalent booster cohort with XBB infection had n=5 participants.
NAb responses increased at 3 weeks after the boost (Fig. 5) but largely declined back to baseline at 6 months after the boost in individuals who did not have a documented infection during the XBB.1.5 predominant period, with NAb responses against WA1/2020, BA.1, BA.2, BA.5, XBB.1.5, XBB.1.16, EG.5, EG.5.1, FL.1.5.1, XBC.1.6, and BA.2.86 of 10,176, 2368, 2621, 2187, 151, 168, 140, 67, 97, 506, and 197, respectively. At 6 months after the boost in individuals who had a documented infection during the XBB.1.5 predominant period, NAb responses were 35,833, 3551, 4409, 3054, 461, 452, 365, 244, 193, 802, and 979, respectively (Fig. 4). Participants who did not receive the bivalent boost had comparable NAb responses at 6 months. We utilized the initial BA.2.86 variant sequence containing the I670V mutation, but BA.2.86 without this mutation showed similar neutralization titers4.
Figure 5. NAb titers induced by bivalent mRNA vaccines against SARS-CoV-2 variants during peak immunogenicity.
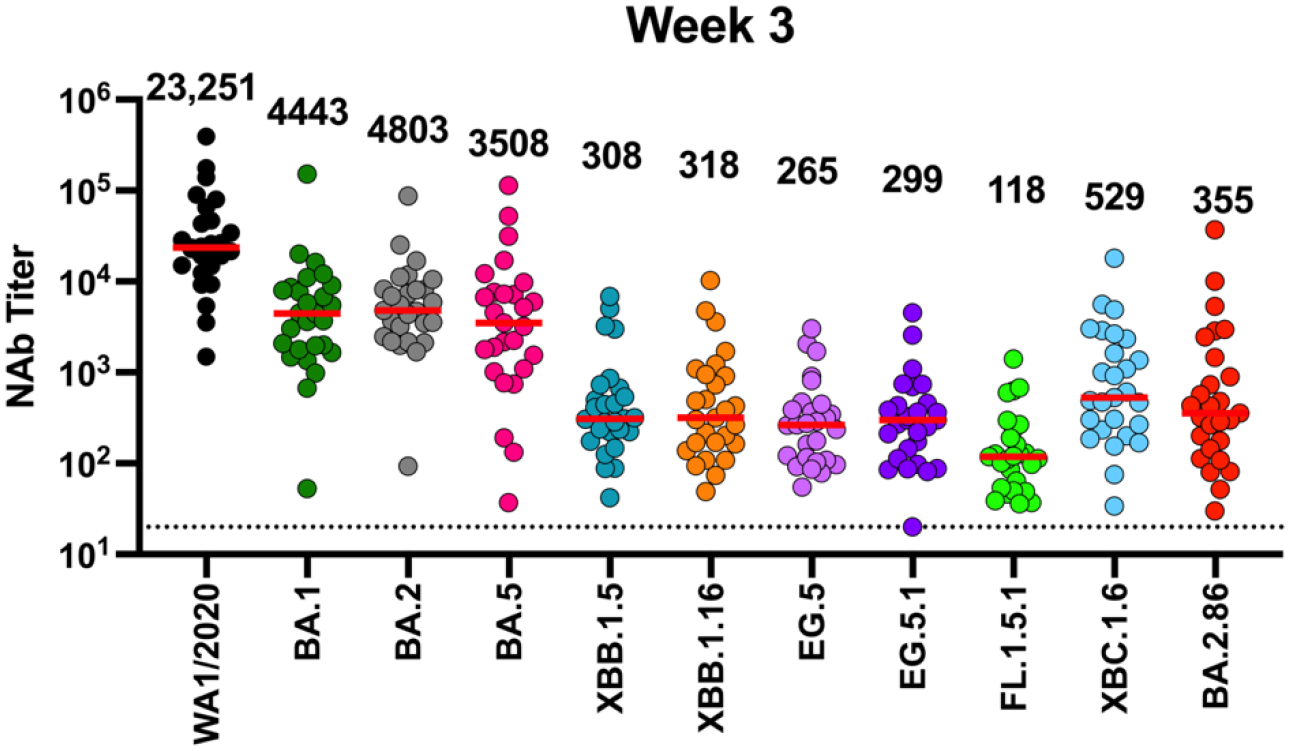
NAb against the WA1/2020, BA.1, BA.2, BA.5, XBB.1.5, XBB.1.16, EG.5, EG.5.1, FL.1.5.1, XBC.1.6, and BA.2.86 variants by luciferase-based pseudovirus neutralization assays at week 3 following the bivalent mRNA booster in fall 2022. The horizontal red bar reflects median values. This cohort had n=27 participants.
Discussion
Our data demonstrate that NAb responses to BA.2.86 were 5–13-fold lower than to BA.2 but were comparable or slightly higher than to XBB.1.5, XBB.1.16, EG.5, EG.5.1, and FL.1.5.1. These data suggest that BA.2.86 evolved directly from the less resistant BA.2 variant, rather than from the current highly resistant circulating recombinant variants, which presumably were selected for increased NAb escape following infection with XBB lineage viruses. Thus, BA.2.86 does not show increased antibody escape compared with current circulating variants. Our findings are concordant with other studies from the U.S4,5 but contrast with studies from Asia6,7, which may reflect differences in population immunity due to different vaccine and variant exposures in various regions of the world. XBC.1.6 is another highly mutated variant that is a BA.2/Delta recombinant (Figs. S1, S2) and similarly shows less NAb escape than XBB.1.5.
Our data also show that NAb profiles at 6 months were comparable in participants who did or did not receive the bivalent mRNA boost, consistent with its limited clinical durability8,9. However, NAb titers to multiple variants increased substantially following XBB infection, suggesting that the monovalent XBB.1.5 booster will likely increase these NAb responses. It will also be important to monitor for potential further evolution or recombination of BA.2.86.
Materials and Methods
Study Population
A specimen biorepository at Beth Israel Deaconess Medical Center (BIDMC) obtained samples from individuals who received monovalent SARS-CoV-2 vaccines as well as bivalent COVID-19 mRNA boosters. The BIDMC institutional review board approved this study (2020P000361). All participants provided informed consent. This study included 66 individuals who received the bivalent mRNA booster in September to November 2022 (N=44) or who did not receive the bivalent mRNA booster (N=22). Participants were followed for 6 months and were excluded if they received immunosuppressive medications.
Pseudovirus Neutralizing Antibody Assay
Neutralizing antibody (NAb) titers against SARS-CoV-2 variants were determined using pseudo-typed viruses expressing a luciferase reporter gene. In brief, a luciferase reporter plasmid pLenti-CMV Puro-Luc (Addgene), packaging construct psPAX2 (AIDS Resource and Reagent Program), and Spike protein expressing pcDNA3.1-SARS-CoV-2 SΔCT were co-transfected into human embryonic kidney (HEK)-293T cells (ATCC CRL_3216) with lipofectamine 2000 (ThermoFisher Scientific). Pseudo-typed viruses of SARS-CoV-2 variants were generated using the Spike protein from WA1/2020 (Wuhan/WIV04/2019, GISAID accession ID: EPI_ISL_402124), Omicron BA.1 (GISAID ID: EPI_ISL_7358094.2), BA.2 (GISAID ID: EPI_ISL_6795834.2), BA.5 (GISAID ID: EPI_ISL_12268495.2), XBB.1.5 (GISAID ID: EPI_ISL_16418320), XBB.1.16 (GISAID ID: EPI_ISL_17646715), EG.5 (GISAID ID: EPI_ISL_17976635), EG.5.1 (GISAID ID: EPI_ISL_18125149), FL.1.5.1 (GISAID ID: EPI_ISL_18126515), XBC.1.6 (GISAID ID: EPI_ISL_17851490), and BA.2.86 (GISAID ID: EPI_ISL_18110065). 48 hours post-transfection, the supernatants containing the pseudo-typed viruses were collected and purified by filtration with 0.45-μm filter. To determine NAb titers in human sera, HEK293T-human angiotensin converting enzyme 2 (hACE2) cells were seeded in 96-well tissue culture plates at a density of 2 × 104 cells per well overnight. Three-fold serial dilutions of heat-inactivated serum samples were prepared and mixed with 60 μl of pseudovirus, and incubated at 37 °C for 1 h before adding to HEK293T-hACE2 cells. 48 h later, cells were lysed in Steady-Glo Luciferase Assay (Promega) according to the manufacturer’s instructions. SARS-CoV-2 neutralization titers were defined as the sample dilution at which a 50% reduction (NT50) in relative light units was observed relative to the average of the virus control wells.
GISAID Data
All genome sequences and associated metadata in this dataset are published in GISAID’s EpiCoV. We gratefully acknowledge the many data contributors to GISAID, the authors and originating laboratories, who share their data, as well as the GISAID staff who facilitate data dissemination. Their work enables an informed response to newly emerging SARS-CoV-2 variants. In particular, we thank the groups that deposited BA.2.86 data which enabled a rapid response to the new distinctive variant. The following acknowledgement links were provided by GISAID for the BA.2.86 data, for the global data sampled after April 1, 2023, and for the Australian data, and data from the Philippines sampled in 2022 and 2023. All sequences in this dataset are compared relative to hCoV-19/Wuhan/WIV04/2019 (WIV04), the official reference sequence employed by GISAID (EPI_ISL_402124).
BA.2.86:
The GISAID EPI_SET_231001ex is composed of 66 individual BA.2.86 genome sequences, with collection dates ranging from 2023-07-24 to 2023-09-22. The data were collected in 12 countries and territories. To view the source and associated sequence data see https://epicov.org/epi3/epi_set/231001ex?main=true.
BA.2.86.1:
The GISAID EPI_SET_231001as is composed of 238 individual BA.2.86 genome sequences, with collection dates ranging from 2023-07-29 to 2023-09-24. The data were collected in 22 countries and territories. To view the source and associated sequence data see https://epicov.org/epi3/epi_set/231001as?main=true.
Global data after April 1, 2023:
The GISAID EPI_SET_231001gp is composed of 423,408 individual genome sequences, with collection dates ranging from 2023-04-01 to 2023-09-27. The data were collected in 137 countries and territories. To view the source and associated sequence data see https://epicov.org/epi3/epi_set/231001as?main=true.
Exploring the origin of XBC.1.16:
The GISAID EPI_SET_231001mn is composed of 14,192 individual genome sequences, with collection dates ranging from 2021-01-01 to 2023-09-27, collected in the Philippines. GISAID EPI_SET_231001fg is composed of 159,253 individual genome sequences, with collection dates ranging from 2021-01-01 to 2023-09-25, collected in Australia. To view the source and associated sequence data see https://epicov.org/epi3/epi_set/231001as?main=true and https://epicov.org/epi3/epi_set/231001fg?main=true.
Supplementary Material
Figure 2. Spike mutations in current circulating SARS-CoV-2 variants.
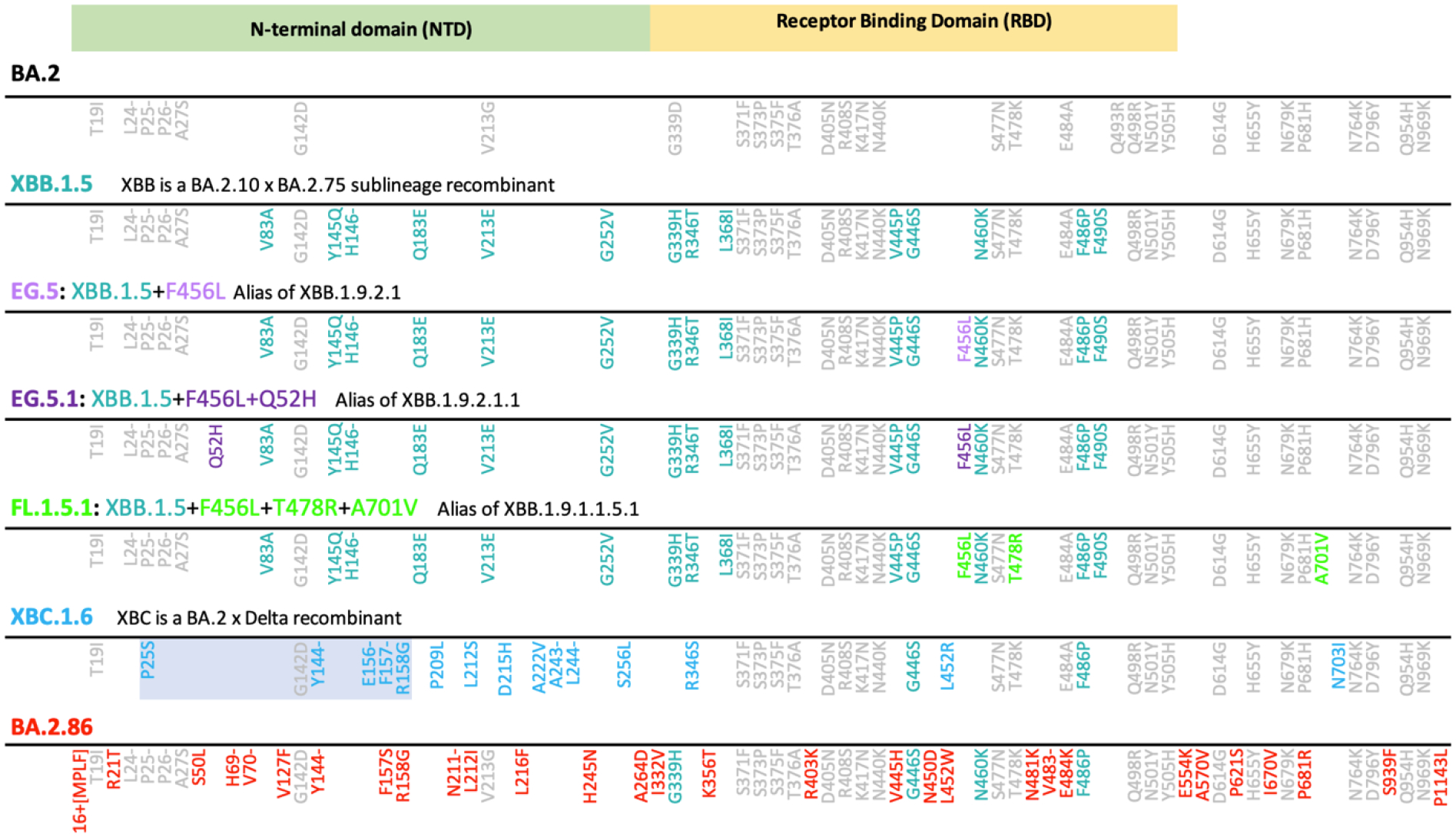

Substitutions in the ancestral BA.2 Omicron lineage relative to the Wuhan/WIV04/ reference strain (https://gisaid.org/WIV04/) are shown in grey. Additional substitutions found in XBB.1.5 are highlighted in teal. Additional substitutions in all other study variants relative to these two patterns are indicated for each variants studied. The mutational pattern used matched the consensus form of the expanding lineage, although in BA.2.86 the I670V mutation was present in the two earliest sampled intact sequences. The highlighted region in XBC.1.6 is a recombinant fragment from a Delta variant.
Funding
The authors acknowledge NIH grant CA260476, the Massachusetts Consortium for Pathogen Readiness, and the Ragon Institute (D.H.B.).
Declaration of interests
Dan Barouch reports financial support was provided by National Cancer Institute. Dan Barouch reports financial support was provided by Massachusetts Consortium on Pathogen Readiness. Dan Barouch reports financial support was provided by Ragon Institute.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflicts of Interest
All authors report no conflicts of interest.
References
- 1.Hachmann NP, Miller J, Collier AY, et al. Neutralization Escape by SARS-CoV-2 Omicron Subvariants BA.2.12.1, BA.4, and BA.5. N Engl J Med 2022;387:86–8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Miller J, Hachmann NP, Collier AY, et al. Substantial Neutralization Escape by SARS-CoV-2 Omicron Variants BQ.1.1 and XBB.1. N Engl J Med 2023;388:662–4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Iketani S, Liu L, Guo Y, et al. Antibody evasion properties of SARS-CoV-2 Omicron sublineages. Nature 2022;604:553–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Wang Q, Guo Y, Liu L, et al. Antigenicity and receptor affinity of SARS-CoV-2 BA.2.86 spike. bioRxiv 2023:2023.09.24.559214. [DOI] [PubMed] [Google Scholar]
- 5.Hu Y, Zou J, Kurhade C, et al. Less neutralization evasion of SARS-CoV-2 BA.2.86 than XBB sublineages and CH.1.1. bioRxiv 2023:2023.09.10.557047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Uriu K, Ito J, Kosugi Y, et al. Transmissibility, infectivity, and immune evasion of the SARS-CoV-2 BA.2.86 variant. The Lancet Infectious Diseases. [DOI] [PubMed] [Google Scholar]
- 7.Yang S, Yu Y, Jian F, et al. Antigenicity and infectivity characterisation of SARS-CoV-2 BA.2.86. The Lancet Infectious Diseases. [DOI] [PubMed] [Google Scholar]
- 8.Lin DY, Xu Y, Gu Y, Zeng D, Sunny SK, Moore Z. Durability of Bivalent Boosters against Omicron Subvariants. N Engl J Med 2023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Becerra X, Jha A. Project NextGen - Defeating SARS-CoV-2 and Preparing for the Next Pandemic. N Engl J Med 2023;389:773–5. [DOI] [PubMed] [Google Scholar]
- 10.Benton DJ, Wrobel AG, Xu P, et al. Receptor binding and priming of the spike protein of SARS-CoV-2 for membrane fusion. Nature 2020;588:327–30. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Barnes CO, Jette CA, Abernathy ME, et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 2020;588:682–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cerutti G, Guo Y, Zhou T, et al. Potent SARS-CoV-2 neutralizing antibodies directed against spike N-terminal domain target a single supersite. Cell Host Microbe 2021;29:819–33 e7. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


