Abstract
Konzo is a neglected paralytic neurological disease associated with food (cassava) poisoning that affects the world’s poorest children and women of childbearing ages across regions of sub-Saharan Africa. Despite understanding the dietary factors that lead to konzo, the molecular markers and mechanisms that trigger this disease remain unknown. To identify potential protein biomarkers associated with a disease status, plasma was collected from two independent Congolese cohorts, a discovery cohort (n = 60) and validation cohort (n = 204), sampled 10 years apart and subjected to multiple high-throughput assays. We identified that Glutathione Peroxidase 3 (GPx3), a critical plasma-based antioxidant enzyme, was the sole protein examined that was both significantly and differentially abundant between affected and non-affected participants in both cohorts, with large reductions observed in those affected with konzo. Our findings raise the notion that reductions in key antioxidant mechanisms may be the biological risk factor for the development of konzo, particularly those mediated through pathways involving the glutathione peroxidase family.
Subject terms: Predictive markers, Neurotoxicity syndromes, Motor neuron disease, Diseases of the nervous system
The search for biomarkers associated with konzo disease has been a long-standing question within the research community. Here, the authors have uncovered a potential biomarker for konzo that is associated with the ability to manage oxidative damage.
Introduction
Konzo is a neglected and irreversible paralytic neurological condition that predominately affects the legs of children and women of childbearing age in sub-Saharan Africa. The occurrence of konzo outbreaks has been consistently associated with undernourished populations that have a chronic dietary reliance on cyanogenic cassava as their main source of food1. However, the biological factors that cause some individuals to develop the disease remain elusive. Konzo is characterized by a sudden onset of a non-progressive spastic paraparesis of varying severity, evolving as a mild (stage 1), moderate (stage 2), or severe (stage 3) form of neurodisability1. In addition to the physical manifestations, visual deficits and speech disturbances are commonly reported2. Recent studies have also reported cognition deficits which, however, were also documented in children unaffected with konzo within the same study population3. While epidemiological evidence indicates that this motor system disease is associated with poisoning from cyanogenic glucosides found in improperly processed cassava, not all individuals who rely on a cyanogenic diet are affected during outbreaks of konzo, suggesting additional and yet unknown factors required for onset of disease4,5.
The primary cyanogenic glucoside found in cassava, linamarin, liberates glucose and hydrogen cyanide (HCN) following hydrolysis in the gastrointestinal (GI) tract, presumably by the gut flora which harbors the necessary β-glucosidase enzyme6. Thus, reliance on a monotonous diet rich in cyanogenic glucosides effectively exposes populations to constant doses of cyanide (CN) to degrees that can greatly vary depending on environmental stressors such as drought7. While the molecular consequences of cyanide exposure and its role in arresting the electron transport chain during aerobic respiration have been extensively studied, the effects of long-term and chronic exposure to sub-lethal toxicity as observed in those residing in konzo-prone regions are less understood8. Because of the lethal potential of cyanide toxicity, most organisms have developed effective detoxification mechanisms that classically require sulfur donors, primarily sulfur-containing amino acids, to convert CN into thiocyanate (SCN), a putatively less toxic molecule9. However, the prevailing biological hypothesis for konzo development is that famine-stricken populations rely on diets that are deficient in the key sulfur amino acids methionine and cysteine, likely decreasing their ability to fully detoxify cyanide10,11. In addition, due to the inability to effectively detoxify CN, a deficiency of such amino acids has also been postulated to reduce a key antioxidant, glutathione, thereby hindering responses to the cyanide exposure-related oxidative stress, however, direct evidence of this mechanism has been scarce12,13.
Many studies have clearly demonstrated an increased incidence of oxidative damage either from reactive oxygen species (ROS) or peroxides in other neurological diseases including Parkinson’s14, Huntington’s15, Alzheimer’s16, and Amyotrophic Lateral Sclerosis (ALS)17. Since cyanide is a key trigger of konzo development, oxidative damage has also been suggested as a key mechanism of neuronal dysfunction in the pathogenesis of this disease18,19. This theory has been augmented by several studies that demonstrated increased markers of oxidative stress (by-products of non-enzymatic lipid peroxidation) in children suffering from konzo as well as documentation of deficiencies in micronutrients which are required to maintain effective antioxidant response in konzo-prone populations18,19.
Using multiple high-throughput proteomics assays in two independent cohorts of konzo-affected children compared to unrelated individuals (Discovery) or their unaffected sibling (Validation) respectively, we demonstrate that levels of the critical plasma-based antioxidant enzyme Glutathione Peroxidase 3 (GPx3) are significantly reduced in those affected. This finding is of particular interest, as genomic and proteomic differences in GPx3 have recently been identified in large cohorts of individuals affected with a similar motor neuron disease, ALS20,21. Additionally, we performed quantitative amino acid profiling of individuals from konzo-prone regions which suggests a deficiency in cystine, the precursor for the antioxidant glutathione, which is required by GPx3 to effectively remove ROS from circulation22. Collectively, these data have identified a biological difference between children who are discordant for konzo and emphasize the role of oxidative stress in neglected motor system diseases, effectively opening new venues for research into potential disease mitigation strategies.
Results
Key proteomic differences between children discordant for konzo in pilot study
To determine if protein levels differed between children affected with konzo and unrelated/unaffected individuals, we carried out liquid chromatography-tandem mass spectrometry analyses (LC-MS/MS) of plasma samples collected from 60 individuals (31 affected, 29 unaffected) from the Kahemba region of the DRC in 2011 (Discovery Cohort). In an untargeted analysis, we identified a high degree of homogeneity across all sample groups at the global scale (Fig. 1A) However, when assessing individual protein differences where a specific protein signal had to be present in at least 75% of the participants, we showed that 28 proteins were significantly increased, and 5 proteins significantly reduced in children affected with konzo versus unrelated/unaffected individuals (q < 0.05, Welch t-test) (Supplementary Fig. 1A, B, Supplementary Data 1). Of the 3 proteins that had a significant differential abundance and a log2 transformed fold change (log(FC)) greater than 0.5, Glutathione Peroxidase 3 (GPx3) displayed the largest signal reduction in affected children (q < 0.01 Welch t-test, log(FC) = −0.88) (Fig. 1B). Using the same criteria, we found that the variable region of an Immunoglobulin heavy chain (IGHV3OR16-12) displayed the highest signal increase in affected children; however, the functionality of the protein is not well established (Fig. 1B).
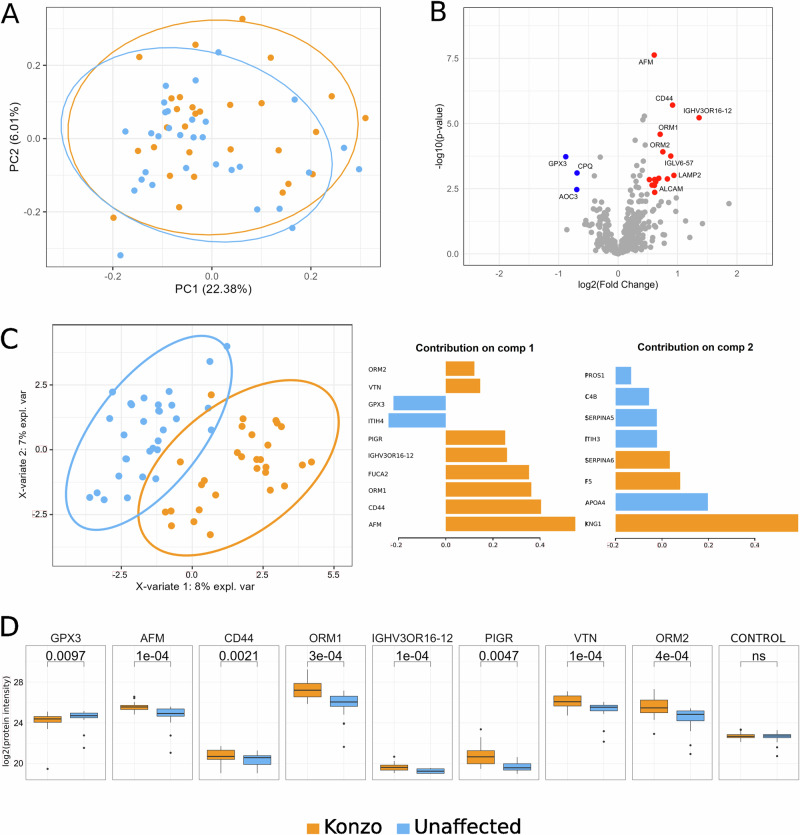
Fig. 1. Identification of potential biomarkers of Konzo disease in the discovery cohort.
Blood plasma proteome from 29 children affected with konzo (orange) and 31 unaffected (blue) were analyzed by untargeted and targeted LC-MS/MS analyses. A Principal component analysis from the untargeted analysis with ellipses drawn at the 95% confidence interval level for each group. B Untargeted analysis representation of log2 of protein intensity ratio versus -log10 of two-sided Welch t-test p-value. Significant proteins (Benjamini–Hochberg adjusted p-value and log2(fold-change) thresholds) are highlighted in blue when they were less expressed in konzo or in red when overexpressed in affected children. C Left: Untargeted analysis sparse partial least square discriminant analysis (sPLS-DA). Right: Top loadings of PC1 and PC2 are highlighted in bar plots where the weight of each variable contribution is represented along the x-axis. D Targeted analysis boxplots represented using log2 transformed intensities of some of the regulated proteins selected based on the results from the untargeted analysis sPLS-DA component 1 (n = 29 konzo individuals, n = 31 unaffected individuals). Two-sided paired Welch t-test p-values are represented above the plots. For each box, data is illustrated as follows: an inner line to represent the median, lower edge and upper edge for the 25th percentile and 75th percentile and whiskers extends from the hinges to either the highest or lowest value within a maximum of 1.5 times the interquartile range (IQR) from the hinge. Data points beyond the IQR are considered as outliers and represented as dots. Within the box plots, orange represents measures obtained from children affected with konzo and blue denotes measures obtained from their siblings who are unaffected with konzo.
To further expand potential protein targets of interest in this cohort, we performed a supervised analysis (Sparse PLS Discriminant Analysis (sPLS-DA)) to determine if factors other than GPx3 may be relevant to the biology of konzo. The approach showed that groups with and without konzo had sample distributions with minimal overlap in their confidence ellipses as well as high predictive value regarding group prediction (Fig. 1C, Supplementary Fig. 1B), albeit with small contributing percentages along the primary and secondary component axis (Fig. 1C). In addition to GPx3, we found other proteins that are largely involved in inflammation, immune response and metabolic disease such as CD44, Afamin (AFM), Alpha-1-acid glycoprotein 1 (ORM1) and Kininogen-1 (KNG1) were the most influential contributors to the differences between children affected with konzo and those unaffected, identifying additional targets that may warrant investigation for pathogenetic relevance to konzo (Fig. 1C). Considering the important role of GPx3 as the only known plasma-based antioxidant enzyme of the Glutathione Peroxidase family, and the relevance of this pathway for neurodegeneration, we next sought to confirm these finding using a targeted LC-MS/MS acquisition. We also targeted peptides of the 38 other proteins that displayed a statistically significant difference, along with those identified through the sPLS-DA analysis that served as important contributors to distinguish groups. Using this targeted acquisition, we found that of the 39 proteins targeted, 25 displayed significant differences in abundance (Fig. 1D, Supplementary Fig. 1C, Supplementary Data 1) with GPx3 still displaying a significant reduction in protein signal (Welch t-test q = 0.01, log(FC) = −0.35) in the konzo-affected children, adding additional layers of confidence to the observations of the untargeted analysis (Fig. 1D).
GPX3 is significantly reduced in affected siblings who are discordant for konzo disease
To validate our findings from our pilot/discovery study we conducted a new LC-MS/MS untargeted analysis on a larger cohort (n = 204) of siblings discordant for konzo (Validation Cohort). Using a more sensitive mass spectrometer than for the previous study, operating in Data Independent Acquisition (DIA) for high throughput deep proteome coverage, we found that GPx3 was the only protein that showed a large signal reduction in children affected with konzo (Welch t-test q < 0.05, log(FC) > 0.5) (Fig. 2A, Supplementary Data 1). When comparing GPx3 signal intensities between affected children versus their unaffected siblings, we found that while individuals with stage 1 of the disease had a significant reduction of the GPx3 signal intensity when compared to their unaffected siblings (Welch t-test q < 0.05), this reduction was the most modest of the stage comparisons (log(FC) = −0.30) (Fig. 2A). We also found that children with stage 2 or 3 of the disease had more dramatic reductions of GPx3 signal relative to their unaffected siblings (Welch t-test q < 0.05, log(FC) = −0.88 and −0.66, respectively) (Fig. 2A, Supplementary Data 1).
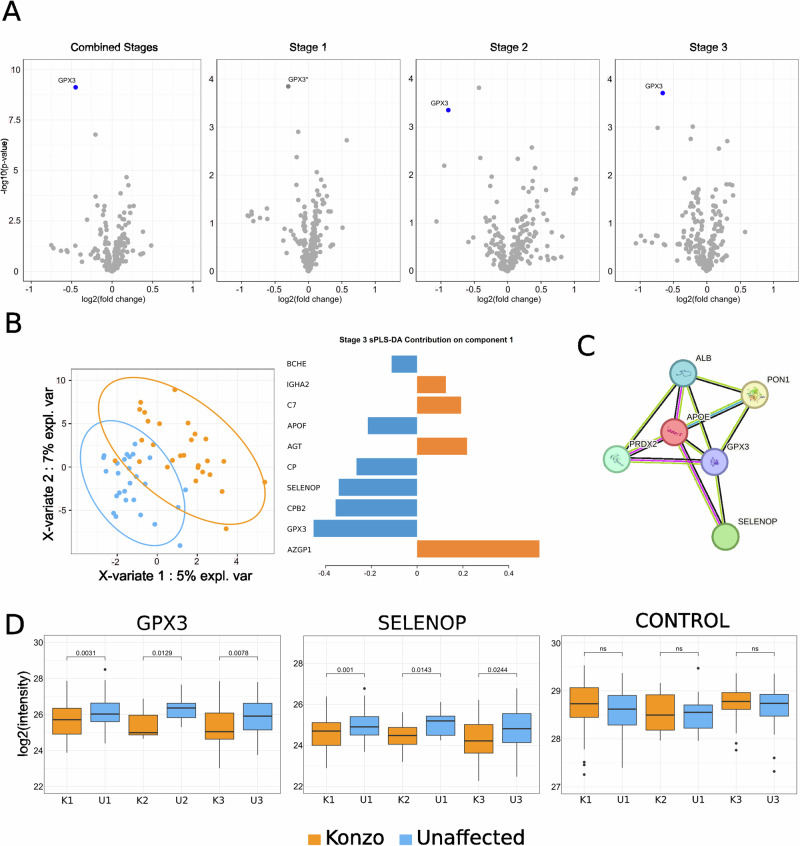
Fig. 2. GPx3 identified as the main konzo biomarker candidate in the validation cohort.
Plasma from unaffected children (blue) and their siblings with different stages of konzo disease (orange). A Untargeted analysis protein volcano plot for all stages combined or for each stage independently. Statistical significance of Glutathione Peroxidase 3 (GPx3) regulation was observed in all comparisons (two-sided paired Welch Benjamini–Hochberg adjusted p-value < 0.05) and a log2Fold change greater than 0.5 was observed for all groups except for Stage 1 comparisons. 204 samples were analyzed with 63 sibling pairs for stage 1, 11 pairs for stage 2 and 28 pairs for stage 3. B Left: Stage 3 sPLS-DA for the untargeted analysis. Right: Top protein contributors on component 1. C StringDB GPx3 interaction network of high confidence experimental interactors identified in the untargeted analysis. D Targeted analysis boxplot representation of log2 transformed protein intensities from the 3 monitored proteins (Glutathione Peroxidase 3 and SelenoProtein1) for each stage (K1/U1 for stage 1 with 62 sibling pairs, K2/U2 for stage 2 with 11 pairs and K3/U3 for stage 3 with 27 pairs). Two-sided paired Welch t-test p-values are represented above the plots. For each box, data is illustrated as follows: an inner line to represent the median, lower edge and upper edge for the 25th percentile and 75th percentile and whiskers extends from the hinges to either the highest or lowest value within a maximum of 1.5 times the interquartile range (IQR) from the hinge. Data points beyond the IQR are considered as outliers and represented as dots. Within the box plots, orange represents measures obtained from children affected with konzo and blue denotes measures obtained from their siblings who are unaffected with konzo.
Like the approach used for the discovery cohort, we next searched for additional proteins that may be indicative of differences in the same cohort using (s)PLS-DA analysis. We found that GPx3 remains one of the most important proteins to discriminate children living with konzo from those not affected, using this supervised model for group clustering, regardless of disease stage severity (Fig. 2B, Supplementary Fig. 2A). When comparing children with stage 1 of konzo with their unaffected siblings, this supervised method identified Ceruloplasmin (CP) and Protein C (PROC) as important classifiers along the component 1 axis (Supplementary Fig. 2A). When comparing stage 2 siblings, it was identified that Ceruloplasmin (CP), Butyrylcholinesterase (BCHE) and Complement component 7 (C7) were the main additional proteins that aided in differentiating between those affected from those unaffected (Supplementary Fig. 2A). Lastly, when comparing children with stage 3 konzo with their unaffected siblings, we found that in addition to GPx3, Zinc-alpha-2-glycoprotein (AZGP1) and Carboxypeptidase B2 (CPB2) were major contributors to group segregation, with CP protein also being identified as being a potential contributor (Fig. 2B).
Among the stage 3 comparisons for the sPLS-DA component 1 proteins, we also observed that one of the top-ranking proteins was Selenoprotein P (SELENOP), one of the few GPx3 interactors detected experimentally (Fig. 2B). Considering that SELENOP is also involved in the defense against oxidative damage and had a q-value that was second (Welch t-test q = 0.08, log(FC) = −0.66) to GPx3 in this group comparisons using an untargeted approach, we sought to monitor peptides of both of these proteins by targeted LC-MS/MS (Supplementary Fig. 2B, Supplementary Data 1). We found that regardless of disease stage, GPx3 was significantly reduced in affected children (Welch t-test p < 0.05) (Fig. 2D), validating the findings obtained using the untargeted LC-MS/MS acquisition. We determined through this target approach that SELENOP was also significantly reduced in children with all three stages of konzo (Welch t-test p < 0.05) (Fig. 2D) and displayed a strong positive correlation with GPx3 signal intensity (Pearson correlation coefficient (r) = 0.9, p = 2.2 × 10−16) (Supplementary Data 1, Supplementary Fig. 2C).
Secondary analysis using an ELISA based approach confirms GPx3 is substantially reduced in children affected with konzo
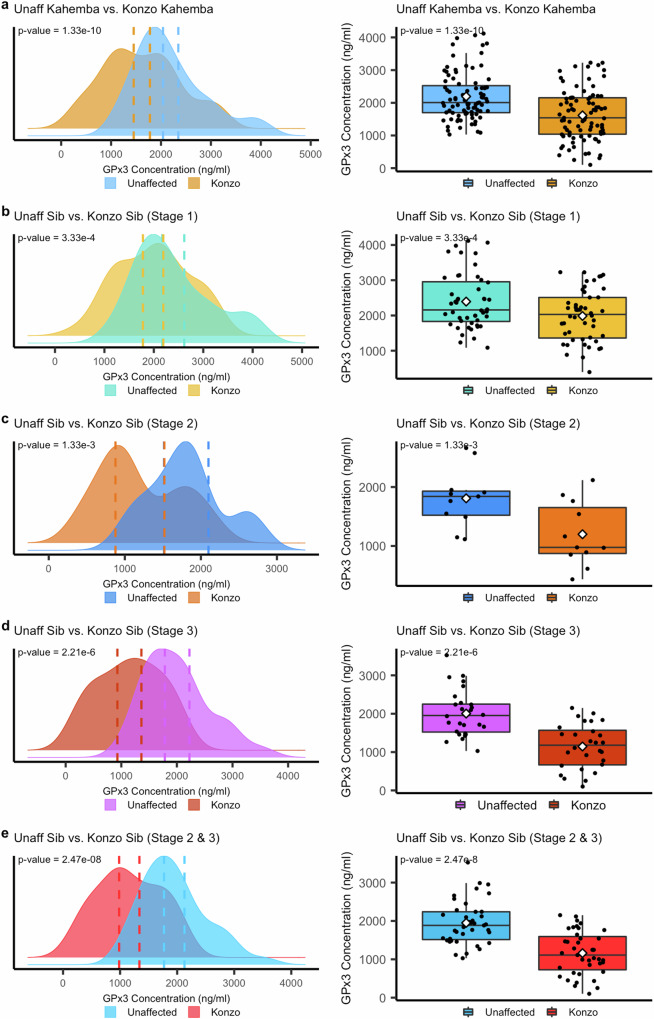
To validate the findings obtained using proteomic-based applications, we sought to determine if the reduced abundance of GPx3 protein in konzo affected children as compared to their unaffected siblings could also be detected using an orthogonal ELISA-based approach. When assessing a large subset of konzo affected children (n = 87) regardless of stage versus their unaffected siblings (n = 87), we detected a significant reduction of GPx3 in those affected (p < 0.0001, Paired t-test) with non-overlapping 95% confident intervals (95% CI Konzo [1.45 µg/ml, 1.78 µg/ml], 95% CI Unaffected [2.04 µg/ml, 2.35 µg/ml]) (Fig. 3a, Supplementary Data 1). These findings appear to be a result of disease and not other variables such as age or biological sex, as no significant difference in biological sex distribution within the cohort was found (Student’s t Test, p = 0.98) (Supplementary Data 1). When GPx3 was correlated to age, we also found a poor correlation exists between these two variables (Pearson correlation coefficient (r) = 0.17, p = 0.02), indicating that age is not strongly influencing the differences of GPx3 abundance in this sibling cohort (Supplementary Fig. 3D). When separating by disease severity, we found that children with stage 1 of the disease had a significant reduction in GPx3 as compared to their unaffected siblings (p < 0.001, Paired t-test), with very minimal overlap in the 95% confidence intervals (95% CI Stage 1 [1.78 µg/ml, 2.19 µg/ml], 95% CI Unaffected [2.17 µg/ml, 2.62 µg/ml]) (Fig. 3b). For those with stage 2 of konzo, the difference between siblings was again significant (p < 0.01, Paired t-test), with minimal overlapping 95% confidence intervals of [0.87 µg/ml, 1.52 µg/ml] for those affected with konzo as compared to [1.52 µg/ml, 2.10 µg/ml] for their unaffected siblings (Fig. 3c). While measuring GPx3 levels in children with the most severe form of konzo, stage 3, we again observed a significant difference in protein levels (p < 0.0001, Paired t-test), however these differences were the most dramatic between the sibling comparisons. We find that those affected with stage 3 have the lowest overall levels of GPx3 and display non-overlapping 95% confidence intervals as compared to their unaffected siblings (95% CI Stage 3 [0.93 µg/ml, 1.36 µg/ml], 95% CI Unaffected [1.78 µg/ml, 2.23 µg/ml]) (Fig. 3d). Given the smaller sample size of children with Stage 2, we next combined the most severe stages (2 and 3) and assessed differences in GPx3 concentration, to increase statistical confidence. Our findings, mirror the differences of Stage 2 or Stage 3 siblings when independently measured and show dramatic and statistically significant reductions of GPx3 in the most severely affect children (p = 2.47 × 10−8, Paired t-test) again with non-overlapping 95% confidence intervals (95% CI Unaffected Stage 2/3 [1.7 ug/ml, 2.1 ug/ml], 95% CI Konzo Stage 2/3 [0.98 ug/ml, 1.3 ug/ml]) (Fig. 3e).
Fig. 3. Validating a decrease of GPx3 concentration in konzo affected children using ELISA.
A general distribution visualization and box plot representations of Glutathione Peroxidase 3 (GPx3) plasma concentration (ng/ml) as measured by human GPx3 ELISA for a Unaffected controls (Unaff Sib) by all konzo (Konzo Sib) stages combined (88 Unaffected, 88 Affected), b Sibling comparisons for Stage 1 konzo (48 Unaffected, 48 Affected), c Sibling comparisons for stage 2 konzo (11 Unaffected and 11 Affected), d Sibling comparisons for Stage 3 konzo (28 Unaffected, 28 Affected) and e Sibling comparisons for Stage 2 and 3 combined (39 Unaffected and 39 Affected). 95% confidence intervals are highlights in dashed lines with corresponding colors to distinguish the unaffected group or those affected with konzo. Statistical differences were calculated using a two-sided paired-t test of the average GPx3 concentration (ng/ml) of the duplicate runs for each sample for Stage 1, 2, and 3 and a T-test of the mean of each sample for unaffected by affected comparisons and for the comparison assessing stages 2 and 3 combined (A and e respectively). For each box, data is illustrated as follows: an inner line to represent the median, lower edge and upper edge for the 25th percentile and 75th percentile and whiskers extends from the hinges to either the highest or lowest value within a maximum of 1.5 times the interquartile range (IQR) from the hinge. Data points beyond the IQR are considered as outliers. The diamond within the box plots represents the data mean for that measure and data point measures can be found in the Data Source File.
When comparing the GPx3 intensity values obtained through mass spectrometry to the quantified values of GPx3 as measured by ELISA for the same individuals, we observe a moderate to strong correlation (Pearson correlation coefficient (r) = 0.64, p < 2.2 × 10−16), indicating that both approaches yield similar conclusions of differential protein abundance (Supplementary Fig. 3A). We next sought to determine how well the ratio of GPx3 abundance/signal between matched discordant siblings (unaffected/affected sibling) correlated between both approaches. We find that a moderate correlation exists between ELISA ratios and Mass Spectrometry intensity ratios for sibling pair differences (Pearson correlation coefficient (r) = 0.55, p < 4.9 × 10−8) (Supplementary Fig. 3B). However, if 2 sibling pairs that display differences in the same direction just to larger degrees depending on method used are removed from analysis, the correlation of ratios between methods substantially increases (Pearson correlation coefficient (r) = 0.75, p < 2.7 × 10−16) (Supplementary Fig. 3C), indicating a good agreement between mass spectrometry and ELISA measures, particularly regarding sibling pair differences.
Because a risk factor for severe konzo disease could cluster in families, we asked whether GPx3 levels might vary between families with different stages of konzo disease, even for unaffected siblings. When comparing unaffected children with a stage 1 sibling versus unaffected children with a stage 2,3, or combined (2 and 3) siblings, we observe a significant difference for all three (p < 0.05, t-test), but for each comparison of unaffected children there are larger overlaps in the 95% confidence intervals (Supplementary Figs. 4A, 5A, 4B, 5B and 4D, 5D respectively). However, when assessing GPx3 protein levels of unaffected children with a stage 2 sibling compared to unaffected children with a stage 3 sibling, we find no significant difference (p = 0.3, t-test) (Supplementary Figs. 4C, 5C). Thus, GPx3 levels trend lower in families affected by more severe konzo disease, even for unaffected siblings.
When measuring the GPx3 levels for children with stage 1 of konzo compared to stage 2, we found a significant decrease in protein abundance for those affected with the moderate stage of disease as compared to the less severe stage (p < 0.01), with stage 1 having a higher lower limit of the 95% confidence interval (1.78 µg/ml) than the upper 95% CI (1.52 µg/ml) of stage 2 (Supplementary Figs. 4e, 5e). The most dramatic difference is observed when measuring protein levels between stage 1 and stage 3 konzo, where the most significant differences are observed with the lower limit of the 95% CI for unaffected siblings being 1.78 µg/ml being as compared to the upper limit of 1.36 µg/ml for the 95% CI for those with stage 3 (Supplementary Fig. 4F, 5F). Like what we observed for the unaffected children, we found that overall GPx3 levels between stage 2 and stage 3 are not significantly different (p = 0.79) (Supplementary Figs. 4G, 5G). Overall, these findings indicate that reductions in GPx3 abundance trends with disease severity with stage 1 having the highest mean levels of GPx3 (1.99 µg/ml ± 0.73 µg/ml) and the highest K1/U1 ratio (0.86), followed by the stage 2 with a mean of 1.2 µg/ml ± 0.54 µg/ml and average K2/U2 ratio of 0.67, with the most severe form of konzo, stage 3, displaying the largest reduction in average protein abundance (mean: 1.15 µg/ml ± 0.58 µg/ml) and the lowest K3/U3 ratio of 0.59 (Supplementary Data 1). When combining the most severe forms of konzo (Stages 2/3) as compared to the milder form (Stage 1) we again see a strong statistical difference (p < 0.0001) with non-overlapping 95% confidence intervals (Stage 1 95% CI [1.78ug/ml, 2.1ug/ml], Stages 2/3 95% CI [0.98 ug/ml, 1.3 ug/ml) (Supplementary Figs. 4H, 5H).
Amino acid profiling siblings discordant for konzo identifies deficiencies in key sulfur amino acids and suggests mitochondrial distress
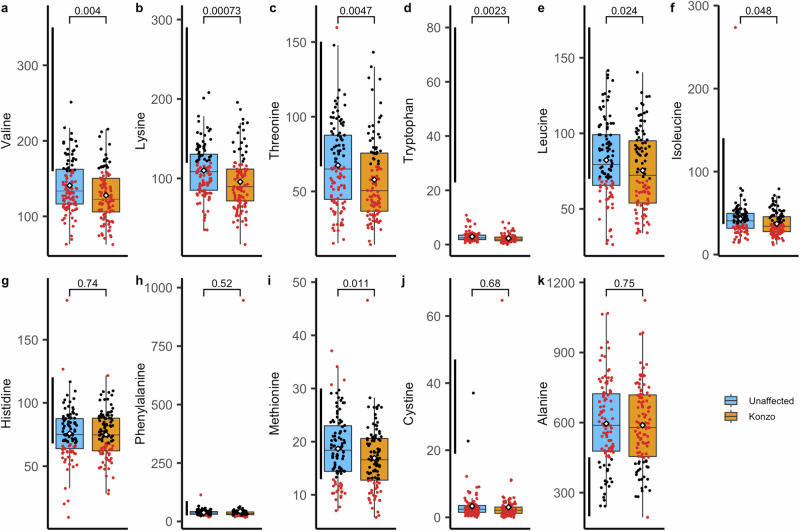
To determine if siblings discordant for konzo had major differences in amino acid profiles which may contribute to changes in GPx3 abundance, we used an HPLC-based approach to quantify plasma amino acids in samples from study participants. Regardless of disease status, the plasma levels of the 9 essential amino acids are suggestive of a protein deficiency (Fig. 4). Particularly, we found that mean plasma levels of Valine (U:140.9 µmol/L, K:127.8 µmol/L), Lysine (U:109.9 µmol/L, K:95.7 µmol/L), Threonine (U:67.5 µmol/L, K:57.9 µmol/L) and Tryptophan (U:2.9 µmol/L, K:2.3 µmol/L), fell below the expected ranges for healthy adolescents (UCSF Clinical Core Accepted Reference Ranges) for both those affected with konzo and their unaffected siblings. Average plasma levels of other essential amino acids such as Leucine (U:82.3 µmol/L, K:75.5 µmol/L), Isoleucine (U:43.0 µmol/L, K:37.1 µmol/L) and Histidine (U:75.8 µmol/L, K:74.9 µmol/L) were also considered at the very low limits for healthy adolescents, except for Phenylalanine (U:37.6 µmol/L, K:43.3 µmol/L) and Methionine (U:18.7 µmol/L, K:16.9 µmol/L), both of which fall within normal expected values, on average, regardless of disease status (Fig. 4, Supplementary Data 1).
Fig. 4. HPLC plasma amino acid profiling of the validation sibling cohort.
Boxplot representations of unaffected (n = 102) and konzo affected (n = 102) children’s amino acids (µmol/L) that were most relevant to the study. Panel a displays Valine levels, b Lysine levels, c Threonine levels, d Tryptophan levels, e Leucine levels, f Isoleucine levels, g Histidine levels, h Phenylalanine levels, i Methionine levels, j Cystine levels and k Alanine levels. Black dots represent samples that fell within the normal range of the particular AA based on UCSF clinical core standards and red dots, represents samples that fell outside of the accepted normal values. Blue shaded box plots represent measures obtained from unaffected siblings while orange shaded box plots represents measures obtained from children affected with konzo. Statistical differences were based on a two-sided t-test outcomes of amino acid means for each sample and compared unaffected groups to affected individuals. For each box, data is illustrated as follows: an inner line to represent the median, lower edge and upper edge for the 25th percentile and 75th percentile and whiskers extends from the hinges to either the highest or lowest value within a maximum of 1.5 times the interquartile range (IQR) from the hinge. Data points beyond the IQR are considered as outliers. Data presented in Fig. 4 can be found in the Data Source File.
When assessing non-essential amino acid profiles, we found that on average, the vast majority fell within healthy reference ranges for adolescents (Supplementary Fig. 6, Supplementary Data 1). However, we found that both Alanine and Cystine substantially fell outside of healthy reference ranges, with Cystine being dramatically reduced in both affected and unaffected siblings with average plasma concentrations of 3.0 µmol/L and 3.3 µmol/L respectively, while Alanine was found to be elevated regardless of disease status (U:595.5 µmol/L, K:589.0 µmol/L). When determining the Alanine to Lysine ratio as an indirect measure of mitochondrial distress we found that unaffected children had an average ratio of 5.8 and those with konzo have an average ratio of 6.9, both of which are substantially higher than the expected ratio of less than 3 for healthy adolescents (Fig. 4). Collectively, regardless of disease status, this cohort of children from a konzo-affected region show signs of protein malnourishment, harbor measures of mitochondrial distress, and deficiencies in a key sulfur-containing amino acid (Cystine).
Discussion
Although konzo was first described in 1938, the biological mechanisms that contribute to the development of this disease have remained elusive23. Studies have continually demonstrated a strong link between a dietary reliance on cyanogenic cassava coupled with general malnutrition, however, these risk factors are present in many regions of sub-Saharan Africa where konzo does not occur5,24,25. In konzo-prone regions, particularly the Democratic Republic of the Congo, this disease appears to affect children and women of childbearing ages at a much higher frequency than adults1. While the exact location of the neurological injury has yet to be identified since MRI (0.5 T) assessments conducted in 1993 on 2 konzo-affected individuals reported no pathological findings, the phenotypic presentation of various severities (Stages 1–3) of hyperreflexia and non-spastic paraparesis suggests damage of motor neurons and corresponding brain regions; conclusions supported by cranial magnetic stimulation procedures26–28. Investigations have sought to identify biological differences between affected and non-affected children residing in prone regions, however, the vast majority of said studies have found that little differences exist between groups regardless of disease status. When searching for potential biomarkers of disease status, it was determined that individuals with konzo did have a higher incidence of albumin carbamoylation, a post-translational modification likely induced by increased exposure to a toxic byproduct of linamarin degradation, cyanate29. While markers of cyanide and cyanate exposure can serve as reliable indicators of acute toxin exposure, there remains a lack of mechanistic understanding of physiological differences that contribute to of disease susceptibility.
To expand the search for biological influences of disease, we sought to investigate if major protein differences were present in children affected with konzo as compared to unaffected individuals. In an initial pilot study using high throughput proteomics applications on plasma samples collected from 29 affected children and 31 unrelated and unaffected children in 2011, we identified several proteins of interest that showed statistical differences between the groups. However, using a more stringent inclusion criteria, we identified one well-characterized protein of interest that was both significantly altered in abundance and displayed a large reduction in signal for those affected with konzo, Glutathione Peroxidase 3 (GPx3). While this protein was an interesting target warranting further investigation, given the smaller sample size and duration post sample collection, we sought to determine if these findings were replicable in a larger and more recent study cohort. Accordingly, in 2021 we acquired plasma from one hundred children affected with all three stages of konzo along with their discordant sex-matched siblings and utilized the most up-to-date mass spectrometry analysis to assess potential protein differences. Interestingly, GPx3 was identified as the sole protein with the most significant reduction in signal for children affected with konzo, a finding that was further validated using ELISA as a secondary method of confirmation.
Glutathione peroxidase 3 belongs to the conserved family of Glutathione Peroxidases and is the only member that is plasma-based, with the remainder of the 7 known GPx enzymes being localized intracellularly30. In humans, this enzyme family serves as one of the critical antioxidant mechanisms, readily scavenging reactive oxides and peroxides that are naturally or exogenously produced, protecting cells and structures from oxidative damage31. GPx3 is also one of the 25 known selenoproteins in humans, which harbor a unique selenocysteine found in the active site that utilizes glutathione as the primary reducing agent to remove oxidative compounds31,32. Therefore, the functionality of this class of enzymes along with other selenium dependent enzymes could be influenced by dietary selenium and adequate levels of glutathione production. In fact, a prior study demonstrated that children residing in konzo-prone regions of the DRC had a collective micronutrient deficiency that trended with disease severity, but such deficiencies are commonplace in populations across sub-Saharan Africa including those with low incidence of konzo18. While our study design did not allow for a direct selenium measure to be accurately determined, we sought to utilize a targeted acquisition to determine if other selenium-containing proteins were also reduced in abundance as observed for GPx3. When focusing on Selenoprotein P (SELENOP), a primary selenium transport protein we found that individuals affected with konzo as compared to their unaffected sibling yielded no significant difference in protein signal in an untargeted analysis. However, when SELENOP was directly targeted, we found that children with all stages of the disease did show a statistically significant reduction in signal as compared to their unaffected sibling. Interestingly, these two plasma-based selenoproteins exhibited a strong positive correlation in regards to protein signal acquired via mass spectrometry, findings that are similar to studies assessing the same measures in both Chinese33 and Swedish cohorts34, where a strong correlation of SELENOP and GPx3 existed. Collectively, this raises the notion that the observed differences here are perhaps related to long-term selenium intake, a micronutrient that has been speculated to play a role in konzo pathogenesis, due to its essential requirement for maintaining antioxidant machinery18. However, since these trends are observed in the sibling study cohort who share a household, overall lifestyle, and likely comparable access to dietary selenium this finding warrants a deeper investigation into the actual causes of these protein differences between siblings discordant for konzo. Particularly, since at time of collection, this cohort appear to harbor overall similar levels of plasma-based amino acids, regardless of disease status or disease severity, indicating minimal differences in nutrition status. An important notion regardless of these and others findings is that all food-stricken populations including those in konzo-prone regions, are likely to have insufficient micronutrient levels that would certainly affect their inherent antioxidant capacities, putting large percentages of children living in developing countries at heightened risk for oxidative damage35,36.
The notion that oxidative damage likely influences konzo susceptibility has been long theorized since this disease is associated with the consumption of cyanogenic compounds that are well known to increase radicals either through direct inhibition of mitochondrial respiration or through toxicity of the downstream metabolites of cyanide12,18. Despite such a likely connection, only one study to date has assessed differences in markers of oxidative damage in children affected with konzo, where increased levels of 8,12-iso-iPF2α-VI isoprostane in affected children was associated with deficits in neurocognition, but not deficits in motor impairment3. Despite scarce data in relation to oxidative damage in children suffering from konzo, the role of oxidative damage in many other neurologic diseases has been well established37. Particularly, it was recently demonstrated in a large multiethnic study that the GPx3 loci was strongly associated with a similar motor neuron disease, ALS, perhaps identifying a risk locus for neuronal disease susceptibility20,38. Other groups found similar outcomes that linked the GPx3 locus to ALS but were also able to demonstrate that lower GPx3 levels strongly correlated with more advanced disease, in addition to showing significant motor deficits in a knockdown zebra fish model21. Interestingly, in our sibling cohort we also observe that children with the most severe form of konzo (stage 3), have on average the lowest GPx3 levels as well as the largest difference when compared to unaffected siblings. Other groups have also raised the possibility that genetic variants of GPx3 can alter gene expression and have associated certain variants with numerous diseases, including but not limited to gastric cancer39, schizophrenia40 and arterial ischemic stroke41. This raises the notion that perhaps within konzo-prone regions of sub-Saharan Africa there may be genetic variants of GPx3, or genes related to associated pathways that circulate which could alter gene expression and subsequently cause a reduction of protein abundances in these populations. Given the decrease in GPx3 that we observe in konzo affected children, a deeper investigation of possible genetic predispositions is thus warranted, as this has also been a long-standing theory within the research community.
Collectively, these findings represent the first biomarker/molecular difference that has been found to be different between children affected with konzo as compared to unaffected individuals and may serve as a biomarker for not only disease, but to identify children that harbor increased risk for disease susceptibility and severity. Considering the notion has been raised that sub-clinical presentation of konzo exists, these measures may allow for the identification of individuals who show similar reduced antioxidant capacity as those with konzo, but who do not show a visible physical disability, enabling the overdue refinement of the clinical features associated with this disease3,42. As this is the first study to assess such measures, the predictive value of these markers will become even more solidified as future studies are completed and similar protein concentrations can be compared on new and even larger populations from the DRC, as well as konzo affected communities in other countries of sub-Saharan Africa. However, many lines of evidence indicate neuronal sensitivity to oxidative damage; thus it seems highly likely that these processes occur in konzo, a motor neuron disease triggered in part by the consumption of toxic compounds that are well established to increase oxidative stress43. Given that we have established selenoproteins such as GPx3 and SELENOP to be substantially reduced in those affected with disease using multiple applications and cohorts, perhaps a reduction in the ability to effectively manage oxidative damage is a leading factor for disease susceptibility and severity, when all other risk factors have been met, which are typically present even in those without konzo. Findings from the discovery cohort also identified numerous proteins that are associated with acute inflammation, redox balance and immune responses such as ORM1, ORM2 and Afamin (AFM) that were elevated in konzo affected children, perhaps identify additional future targets of investigation44–46. Considering an inability to manage food-induced oxidative stress would damage tissue and cells, an elevation of immune response that altered inflammation markers also seems highly likely47. While these specific proteins were not significantly elevated in the sibling validation cohort, computational predictions based on (s)PLS-DA analysis did identify proteins associated with immunity and inflammatory responses such as Zinc-alpha-2-glycoprotein (AZGP1) and Carboxypeptidase B2 (CPB2) as being important contributors to group separation48–50. So, additional investigation is necessary to link these potential associations to konzo pathogenesis, particularly in relation to time of disease onset, as such markers may only be present during early phases of konzo onset when inflammation or neural damage is likely highest.
In addition to a reduction of GPx3 levels, we also observed in our sibling cohort a large deficiency in cystine, the limiting factor to produce glutathione, which is required as a reducing agent for GPx3 and the entire glutathione peroxidase family30,51. While a sulfur containing amino acid deficiency in konzo-prone regions has been speculated for several decades11, this is the first quantitative evidence that demonstrates a cystine deficiency within these populations, regardless of disease status. Given a reduction in both GPx3 and a likely reduction of glutathione production, as has been theorized by others in the field of konzo13, the role of oxidative damage and mitigation as a contributing factor of disease susceptibility or severity is only further bolstered by the findings of this study. Theoretically, deficiencies in GPx3 and cystine reduce the ability of the body to counter oxidative stress by virtue of the glutathione pathway and associated enzymes, which may lead to increased ROS in the circulation. Given that cyanide exposure from linamarin is strongly associated with the development of konzo, an increase in plasma oxidative stress may also affect cyanide action or metabolism, potentially amplifying its capacity to induce mitochondrial stress or giving rise to its oxidized form cyanate, which is a strong neurotoxin in various experimental models that can further decrease glutathione levels in the CNS52,53. Therefore, based on these and other findings, it should become a priority to begin offering nutritional intervention with a focus on cystine/sulfur containing amino acids and micronutrient supplementation to konzo-prone communities. While changing food related behaviors has always proven difficult, particularly in impoverished communities, a precision nutritional approach focused on increasing the inherent ability to mitigate oxidative stress may be a practical solution—as the global reliance on cassava is poised to only increase in the coming decades, particularly in light of climate change.
While this finding is of great interest to the field of konzo as well as other neurodegenerative diseases, we cannot determine if the reduction in GPx3 is a result of disease or an inherent contributing factor for disease that is triggered by a micronutrient deficiency or underlying genetic variability. Furthermore, the increased reduction in the abundance of GPx3 in unaffected siblings of children with stage 3 as compared to children with a stage 1 sibling with the disease suggests that GPx3 levels may serve as an indicator for individuals within one household most at risk for developing the severe form of disease. As such, it is apparent that additional experimentation focusing on genomics, environmental/nutritional components and their crosstalk is warranted to determine the causal factor(s) that reduce both cystine and selenoprotein levels in children affected with konzo to unravel the biological mechanism behind this disease.
Methods
Ethics statement
Prior to any specimen collection, community consent was first obtained from village leaders. Informed and written consent was then obtained from the Chef de zone/Médecin de zone, who represents the interests of the ministry of health and individuals in the study population. Upon approval and consent by the representatives, verbal and/or written consent was obtained from the parent and/or guardian of the children who participated in the study. Verbal consent was obtained when there were limitations with literacy and the individual expressed a general disinclination to signing written documents that cannot be read and fully comprehended by them. Participants were compensated for their travel expenses to the study site and small monetary and food donations were also provided to the families that participated in this research. The study posed no harm to participants, and participants could chose to not donate samples. The study was approved by the IRB review board at the Oregon Health & Science University (OSHU) (IRB FWA00000161) and by the Ministry of Health of the Democratic Republic of the Congo (DRC).
Inclusion and ethics statement
The study presented here, has been approved and annually reviewed by the ethics board of the Ministry of Health of the Democratic Republic of the Congo (DRC) and the IRB board of the Oregon Health and Sciences University. Additionally, this research was also invited and approved from the regional administration of the Kahemba health zone, where the individual’s assessed in this study reside. As the research presented in this manuscript is highly relevant to the local communities of the DRC, input from the local physicians, scientists and students who are familiar with konzo have been instrumental to the successful outcome of this study. Experts on Konzo who are based at the University of Kinshasa and Institute National de Research Biomedical (INRB) helped design and execute the research activities presented in this manuscript, in collaboration with the large group of co-authors who aided in research activities related to this work. Projects related to unraveling the biological mechanisms of konzo susceptibility have been ongoing and led in the DRC by Dr. Desire Tshala-Katumbay for over 20 years and as such, capacity building and long-term solutions have been at the forefront of all research related activities in the DRC. In addition, any biological material that were collected from DRC-based individuals remains bio banked at the INRB in Kinshasa and managed under the discretion of that local institute, to insure research related to konzo is held at the highest standard when foreign counterparts are involved. The research teams who are authors on this manuscript have been working closely together for over 7 years, and research roles, responsibility and authorship topics are clearly discussed early in research planning, and we strive to be as inclusive as possible, to attribute credit where it is due. As this team is very familiar with conducting research in limited-resource settings that deal with vulnerable populations, safety to both the participants and researchers were and continue to also be a top priority. As such, we strive to prevent any stigmatization or discrimination of individuals affected with konzo during our research studies. As the community of Kahemba is very familiar with the ongoing research efforts to prevent konzo, these studies may even help de-stigmatize individuals with konzo through determining the biology behind this disease all while attempting to draw global attention to this neglected disease. Considering Konzo is an Afro-centric disease that affects very isolated communities, global collaborations are necessary to utilize the most advanced scientific technologies to conduct high-throughput measures as presented in this work. As such, the majority of citations referenced in this manuscript originate from local researchers in the DRC and other sub-Saharan African countries as this is where the current expertise related to konzo is most prevalent. Collectively, this study and others that are conducted with this global team of experts focused on unraveling the biological mechanisms of konzo operates at the highest standards of inclusion, ethical considerations, participant safety and authorship, with the goal of bringing global attention to konzo and other neglected diseases that burden the world’s most impoverished populations.
Sample collection
In October of 2011, our research group comprised of Democratic Republic of the Congo (DRC)-based physicians and experts on konzo along with research scientists collected 60 plasma samples from children affected with konzo and those who were unrelated and unaffected in Kahemba, DRC. In April of 2021, the DRC-based Konzo research group traveled to Kahemba DRC and collected plasma samples from 102 pairs of sex-matched sibling pairs who were discordant for konzo disease. Prior to collection, the Ministry of Health for the DRC and the institutional review board at the Oregon Health and Sciences University provided ethical approval for this study. All participants and parents were consented prior to collection in either French or the appropriate language for the region of collection (Kikongo). Roughly 3 ml of blood was collected from each participant in EDTA-coated tubes and spun at 3000 RPM for 10 min to separate plasma from red blood cells. Plasma was then collected from all participants and then transferred to cryovials and stored in liquid nitrogen within 1 h of sample collection. Upon arrival to the laboratory in Kinshasa, plasma samples were transferred to long-term storage at −80 °C and stored until proteomics applications were ready to be conducted. During sample collection an assessment as to whether an individual was affected with konzo was conducted following the WHO’s 3 main criteria for diagnosis including evidence of (1) a visible symmetric spastic abnormality of gait while walking or running, (2) a history of onset of less than 1 week followed by a nonprogressive course in a formerly healthy person, and (3) bilaterally exaggerated knee or ankle jerks without signs of disease of the spine.
Population characteristics
For this study we utilized data from two independent study cohorts that were consented and sampled 10 years apart. In 2011, plasma samples from the discovery cohort were collected from 60 unrelated individuals (31 unaffected, 29 konzo affected) in the Kahemba region of the Democratic Republic of the Congo, which is approximately 850 km southeast of the capital of Kinshasa. The average age of the unaffected children was 9.5 ± 2.6 years, with samples taken from 17 Males and 14 Females. The average age of those affected with konzo was 9.4 ± 2.6 years, with samples taken from 14 Males and 15 Females. Within the affected group 17 were diagnosed with Stage 1 Konzo, 7 with Stage 2 konzo and 5 with Stage 3 konzo, however proteomic analysis on the discovery cohort were not segregated based on stage of the disease. In 2021, plasma samples from the validation cohort were collected from 102 sibling pairs (102 unaffected and 102 konzo affected, 98 sex-matched, 4 pairs who are discordant for sex) in the Kahemba region of the Democratic Republic of the Congo. The average age of unaffected siblings in this cohort was 8.7 ± 2.6 years with samples taken from 56 unaffected Male siblings and 46 from unaffected Female siblings. The average age of konzo affected children was 9.25 ± 2.2 years with samples taken from 59 konzo affected Male siblings and 43 konzo affected Female Siblings. Within the affected sibling group, 63 were affected with stage 1 konzo, 11 with stage 2 konzo and 28 with stage 3 konzo. Recruitment was based on prior study populations that were familiar with ongoing research related to konzo, therefore the stage groups were generated based simply on those households who chose to participate in this study, which harbored a higher a frequency of children affected with Stage 1 and 3 than Stage 2 konzo.
Plasma protein extraction and digestion
For the discovery cohort (2011), two separate preparations were made with either 1 or 2 µL of plasma added to new tubes containing 24–23 µL of SDC buffer consisting of sodium deoxycholate 1% (DOC), tris(2-carboxyethyl) phosphine 10 mM (TCEP), chloroacetamide 40 mM and Tris 100 mM pH 8.5. Samples were heated at 95 °C for 10 min then cooled off at room temperature. For the untargeted experiment, proteins were digested with 0.7 µg of trypsin (sequencing grade, Promega, Madison, WI) and 0.7 µg of Lys-C (New England Biolabs) for 1 h at 37 °C. For the targeted analysis, proteins were digested with 0.4 µg of trypsin and 0.4 µgLys-C for overnight incubation at 37 °C (aprox. 18 h). After acidification with trifluoroacetic acid 1% (TFA) and centrifugation at 16,000 × g for 5 min, resulting peptides were purified on StageTips according to Rappsilber et al.54 using C18 Empore solid phase (CDS) and vacuum dried. Samples were resuspended in 2% acetonitrile (ACN), 0.05%TFA and peptide concentrations were estimated with 205 nm absorbance readings (Nanodrop, Thermo Fischer). Prior to targeted analysis injections, 1uL of 1X standard indexed retention time peptides (Biognosys AG) was added to each sample to control for the instrumental variability.
For the validation cohort (2021), a similar protocol than above was used in 96 well plates with the following modifications: Digestion was made with 0.25 µg of trypsin only, acidification was performed with formic acid 1% (FA). Plates were then centrifuged at 800 g for 30 min, and the supernatant transferred into new plates. After vacuum evaporation, samples were resuspended in FA 0.1% and peptide concentration was estimated as described above. For the targeted analysis 8 fmol of Cytochrome C digest was added to the samples before injection in LC-MS/MS (Thermo Scientific).
LC-MS/MS acquisitions
For the discovery cohort, untargeted analysis samples were analyzed by nanoLC/MSMS using a Dionex UltiMate 3000 nanoRSLC liquid chromatography system (Thermo Fisher Scientific) interfaced to an Orbitrap Fusion Tribrid mass spectrometer (Thermo Fisher Scientific, San Jose, CA,USA) equipped with a nanoelectrospray ion source. 1 µg of peptides were separated on a 50 cm length × 75 µm internal diameter (ID) C18 Pepmap Acclaim column (Thermo Fisher) using a linear 90-minute acetonitrile gradient. Mass spectra were acquired using a Data Dependent Acquisition mode (DDA) with full scan spectra done in the Orbitrap while fragments ions were generated by Higher energy Collision-induced Dissociation (HCD) and measured in the linear ion trap.
Targeted analysis was carried out on Dionex UltiMate 3000 nanoRSLC coupled to a TSQ Altis mass spectrometer (Thermo Fisher Scientific, San Jose, CA, USA) equipped with a nanoelectrospray ion source. 1 µg of peptides were separated on a 15 cm length × 75 µm ID C18 separation column (Pepmap Acclaim column, ThermoFisher) using a linear 45-minute acetonitrile gradient. 556 parent/fragment transitions corresponding to 95 peptides of 38 proteins were monitored by Selected Reaction Monitoring method (SRM) in a scheduled manner (Supplementary Data 1) using isolation windows of 0.7 m/z in Q1(parent) and 1.2 m/z in Q3 (fragment).
The validation cohort was analyzed on an Evosep One (Evosep Inc., Odense, Denmark) liquid chromatography system coupled to an Orbitrap Exploris 480 mass spectrometer (Thermo Fisher Scientific, Bremen, Germany). 500 ng of peptides loaded on Evotip pure devices (Evosep) according to manufacturer protocol were separated using the pre-programmed 60 samples per day gradient on an 8 cm length ×100 µm ID capillary column (Evosep). The untargeted analysis was carried with a data independent acquisition (DIA) mode in which the precursors were fragmented using HCD on a 350–875 m/z precursor mass range using 35 DIA windows of 15 m/z with a 0.1 m/z overlap. A gas phase fractionation (GPF) library was also acquired by injecting a pool of all samples 8 times in order to cover a 350–910 m/z precursor mass range using 35 DIA windows of 2 m/z with a 0.1 m/z overlap. Targeted analysis was performed using the same chromatographic conditions but with the instrument operating in a parallel reaction monitoring (PRM) method in which targeted HCD MS2 scan were acquired on 12 precursor ions, corresponding to 3 proteins, using an isolation windows of 0.7 m/z (Supplementary Data 1).
Protein identification and data analysis
Mass spectra for the discovery cohort DDA analysis were searched against a Uniprot Homo sapiens sequence database (UniProt Reference Proteome – Proteome ID UP000005640– 93634 entries – 2018.04) using the search engine Andromeda integrated into the MaxQuant software (version 1.6.3.4)55. Trypsin was set as the digestion enzyme. For protein validation, a false discovery rate (FDR) of 1% was allowed at peptide and protein levels based on a target/decoy search. For data processing, the intensity column from the proteinGroups text file were imported in R software. A protein was considered for quantification if it presented at least 2 identified peptides and if there was an intensity value in at least 75% of the replicates in one of the two groups. Mediane-based normalization was performed using protein intensities in each sample and missing values were imputed from the 1st percentile intensity value for each sample independently. A list of candidate biomarkers was determined from the proteins that presented a Welch t-test q-value less than 0.05 (after Benjamini–Hochberg correction for multiple testing) and from the top contributors in component 1 and 2 of a sparse partial least square discriminant analysis (sPLS-Da) computed with the Mixomics R package56.
The data acquired for the targeted analysis of the discovery cohort was imported into the Skyline software57. Peaks were manually validated based on signal intensity, transition chromatogram superposition and retention time. The total area of the 98 peptides were imported into R software for data processing, t-test were done at both the peptide and protein levels. Peptides and proteins were considered regulated between Konzo and unaffected groups if they presented a Welch t-test q-value less than 0.05.
For the DIA analysis of the validation cohort, spectra were analyzed with DIA-NN (version 1.8). First a GPF library was generated using GPF acquisition files and an in silico digested Homo sapiens sequence database (UniProt Reference Proteome – Proteome ID UP000005640– 79052 entries –2022.04) to perform a library-free search. Trypsin/P was set as the enzyme parameter, a maximum of 2 missed cleavage was allowed, carbamidomethylation was set as a fixed modification and methionine N-terminal excision and oxidation were set as variable modifications with a maximum of 2 allowed per peptide. Only 2+ to 4+ precursors were considered along a 350–875 m/z mass range and fragments on a 200–2000 m/z range. Quantification of the sample was done using the same parameters as the GPF library except that the match between run option was enabled. DIANN main report file was processed into R using the diann package and the MaxLFQ normalization algorithm while applying a 1% q-value filter on both precursors and protein groups. Filtering of quantifiable proteins and missing values imputation were done as in the discovery untargeted analysis. Candidate proteins were selected based on a paired Welch’s t-test and z-score thresholds.
Analysis of the PRM data was done in Skyline software and chromatographic peaks were manually validated in a similar fashion as the SRM data which led to keep 4 GPX3 peptides and 2 SEPP1 peptides for quantification. Peptide total peak area from the 6 best fragment ion intensities were exported from Skyline into R software. For each peptide a paired Welch t-test p-value threshold of 0.05 was used to evaluate the significance of abundance variation
GPX3 ELISA
Following plasma thawing, a large subset of samples collected in 2021 from the discordant sibling cohort (n = 174) were vortexed briefly and diluted 1:100 in ELISA sample buffer (NOVUS Biologicals). Methodology was followed as outlined in the manufacture’s protocol for the Human GPX3 ELISA Detection Kit (NOVUS Biologicals). All samples were run in duplicate and the absorbance at 450 nm was averaged prior to statistical analysis. The significance of differences in protein abundance was determined with a Paired T-test for sibling cohort comparisons and a T-test for comparisons of unrelated groups such as Stage 1 versus Stage 3.
HPLC amino acid quantification
The analysis was conducted using our previously established chromatographic conditions (5) on an Agilent 1260 Infinity II LC System (5). Briefly, a C18 column with dimensions of 3.0 × 100 mm, and a particle size of 2.7 µm was employed. The injection volume was 20 µL, and detection was executed at a wavelength of 338 nm. The binary mobile phase composition and gradient elution program are detailed in the referenced publication. Sample preparation was done by using of an economical 3 K centrifuge filter instead of an acid or alcohol precipitation. This filtration step removes the larger proteins like acid precipitation, which can clog the analytical column. The filters have an added advantage of not diluting the sample, which improves the detection limits of the assay and allows less sample to be used. Peak quantification for corresponding amino acids followed the procedure as directly outlined in the referenced protocol.
Statistics and reproducibility
This study is highly synergistic with the goals of the long-term projects and funding secured by Drs. Tshala, Boivin and Bramble for investigating the molecular mechanisms associated with cassava induced neurotoxicity in the Kahmeba region of the DRC. As such, individuals from the Kahemba region were recruited based on prior diagnosis of konzo by expert phenotyping of this disease, who were participants in the long-term clinical trials aimed at altering cassava processing to reduce konzo. For the 2011 discovery cohort, these samples were obtained during the initial start of the long-term clinical trials conducted by Drs. Tshala and Boivin and individuals recruited for the 2021 validation sibling cohort were also familiar and participants in long-term studies conducted in Kahemba by Drs. Tshala, Boivin and Bramble. Unaffected individuals from both cohorts obtained from Kahemba were not considered to have konzo at the time of sampling following the 3 main criteria for diagnosis outlined by the World Health Organization. Prior to study design that involved proteomic analysis, samples sizes were not considered prior to sampling. The 2011 discovery cohort was sampled years prior to initial proteomics experimentation, however the sample size (n = 60) was sufficient to detect moderate to large differences between groups. However, for the 2021 validation cohort expanded the sample size for more statistical confidence while attempting to control for unknown variables. As such, we recruited 102 sex-matched sibling pairs who were discordant for konzo from the Kahemba region of DRC. This number was more than three times the size of the discovery cohort, while enabling paired statistical analysis due to the cohort being siblings. Details regarding sex and age and cohort size of the recruited participants can be found in the population characteristics section of this manuscript. For this study, all participants that samples were taken from were included. This focus of this study was to determine if children affected with konzo had different plasma-based protein differences that may shed light on the mechanisms of the development of the disease. As such, randomization or blinding was not possible in this study. All statistical tests associated with the outcomes presented in this work are detailed in their corresponding methods sections, presented figures and the results portion of this manuscript.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Supplementary information
Description of Additional Supplementary Files
Source data
Acknowledgements
We would like to thank all the Congolese participants of this study for kindly donating specimens for proteomics analysis. We would also like to thank the funding sources for this project, with support from NIH grant NIEHS/FIC R01ES019841 (D.T.K.), and support by the Fogarty International Center of the National Institutes of Health (NIH) under Award Number 5K01TW011772 (M.S.B.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author contributions
M.S.B., N.V., D.T.K. and A.D. designed and carried out the experiments related to this manuscript and prepared the manuscript for this study. V.F. and F.R-D. performed all proteomic analysis and interpretation and aided in manuscript preparation. G.B., M.S.B., M.L.K., N.V., H.H.-H. and V.K.M. participated in sample collection and preparation for this study. Y.Z., A.M., G.C. and M.S. designed and carried out the experiments related to amino acid quantifications. R.M. and D.S. participated in manuscript and figure preparation and ELISA validation of GPX3. M.B., E.V., D.M.N., L.C., D.R.M., A.D. and D.T.K. provided the senior oversight of this work and aided in manuscript preparation, data finalization and interpretation.
Peer review
Peer review information
Nature Communications thanks Cristina Al-Khalili Szigyarto and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Data availability
The mass spectrometry data generated in this study have been deposited in the Proteomexchange database under accession code PXD049973 and are freely accessible. The processed/cleaned mass spectrometry data associated with this manuscript are available in Supplementary Data 1. The additional ELISA and HPLC data generated in this study are also provided in the Supplementary Information/Source Data file. Source data are provided with this paper.
Code availability
Proteomics data were processed using open-source software: MaxQuant v 1.6 3.4 for DDA experiment (https://maxquant.org/), DIA-NN v1.8 for DIA experiment (https://github.com/vdemichev/DiaNN), Skyline v.23.1.0.380 (https://skyline.ms/project/home/software/Skyline/begin.view) for SRM and PRM experiments. Statistics and visualization of proteomics data were done using R software v.4.1.1 (https://www.r-project.org/) including the R packages Mixomics (http://mixomics.org/) and diann (https://github.com/vdemichev/diann-rpackage).
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors contributed equally: Matthew S. Bramble, Victor Fourcassié.
Contributor Information
Matthew S. Bramble, Email: mbramble@childrensnational.org
Desire Tshala-Katumbay, Email: tshalad@OHSU.edu.
Arnaud Droit, Email: arnaud.droit@crchudequebec.ulaval.ca.
Supplementary information
The online version contains supplementary material available at 10.1038/s41467-024-52136-5.
References
- 1.Kashala-Abotnes, E. et al. Konzo: a distinct neurological disease associated with food (cassava) cyanogenic poisoning. Brain Res. Bull.145, 87–91 (2019). 10.1016/j.brainresbull.2018.07.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Mwanza, J. C., Tshala-Katumbay, D. & Tylleskär, T. Neuro-ophthalmologic manifestations of konzo. Environ. Toxicol. Pharm.19, 491–496 (2005). 10.1016/j.etap.2004.12.012 [DOI] [PubMed] [Google Scholar]
- 3.Boivin, M. J. et al. Neuropsychological effects of konzo: a neuromotor disease associated with poorly processed cassava. Pediatrics131, e1231–e1239 (2013). 10.1542/peds.2012-3011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Tylleskär, T. et al. Epidemiological evidence from Zaire for a dietary etiology of konzo, an upper motor neuron disease. Bull. World Health Organ.69, 581–589 (1991). [PMC free article] [PubMed] [Google Scholar]
- 5.Cliff, J. et al. Konzo and continuing cyanide intoxication from cassava in Mozambique. Food Chem. Toxicol.49, 631–635 (2011). 10.1016/j.fct.2010.06.056 [DOI] [PubMed] [Google Scholar]
- 6.Bramble, M. S. et al. The gut microbiome in konzo. Nat. Commun.12, 5371 (2021). 10.1038/s41467-021-25694-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Oluwole, O. S. Cyclical konzo epidemics and climate variability. Ann. Neurol.77, 371–380 (2015). 10.1002/ana.24334 [DOI] [PubMed] [Google Scholar]
- 8.Hendry-Hofer, T. B. et al. A Review on Ingested Cyanide: Risks, Clinical Presentation, Diagnostics, and Treatment Challenges. J. Med Toxicol.15, 128–133 (2019). 10.1007/s13181-018-0688-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Swenne, I. et al. Cyanide detoxification in rats exposed to acetonitrile and fed a low protein diet. Fundam. Appl Toxicol.32, 66–71 (1996). 10.1006/faat.1996.0107 [DOI] [PubMed] [Google Scholar]
- 10.Baguma, M. et al. Konzo risk factors, determinants and etiopathogenesis: What is new? A systematic review. Neurotoxicology85, 54–67 (2021). 10.1016/j.neuro.2021.05.001 [DOI] [PubMed] [Google Scholar]
- 11.Cliff, J. et al. Association of high cyanide and low sulphur intake in cassava-induced spastic paraparesis. Lancet2, 1211–1213 (1985). 10.1016/S0140-6736(85)90742-1 [DOI] [PubMed] [Google Scholar]
- 12.Nzwalo, H. & Cliff, J. Konzo: from poverty, cassava, and cyanogen intake to toxico-nutritional neurological disease. PLoS Negl. Trop. Dis.5, e1051 (2011). 10.1371/journal.pntd.0001051 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nunn, P. B., Lyddiard, J. R. & Christopher Perera, K. P. Brain glutathione as a target for aetiological factors in neurolathyrism and konzo. Food Chem. Toxicol.49, 662–667 (2011). 10.1016/j.fct.2010.08.037 [DOI] [PubMed] [Google Scholar]
- 14.Dias, V., Junn, E. & Mouradian, M. M. The role of oxidative stress in Parkinson’s disease. J. Parkinsons Dis.3, 461–491 (2013). 10.3233/JPD-130230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kumar, A. & Ratan, R. R. Oxidative Stress and Huntington’s Disease: The Good, The Bad, and The Ugly. J. Huntingt. Dis.5, 217–237 (2016). 10.3233/JHD-160205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Huang, W. J., Zhang, X. & Chen, W. W. Role of oxidative stress in Alzheimer’s disease. Biomed. Rep.4, 519–522 (2016). 10.3892/br.2016.630 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Hemerková, P. & Vališ, M. Role of Oxidative Stress in the Pathogenesis of Amyotrophic Lateral Sclerosis: Antioxidant Metalloenzymes and Therapeutic Strategies. Biomolecules11, 437 (2021). [DOI] [PMC free article] [PubMed]
- 18.Bumoko, G. M. et al. Lower serum levels of selenium, copper, and zinc are related to neuromotor impairments in children with konzo. J. Neurol. Sci.349, 149–153 (2015). 10.1016/j.jns.2015.01.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Makila-Mabe, B. G. et al. Serum 8,12-iso-iPF2α-VI isoprostane marker of oxidative damage and cognition deficits in children with konzo. PLoS One9, e107191 (2014). 10.1371/journal.pone.0107191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tanaka, H. et al. ITIH4 and Gpx3 are potential biomarkers for amyotrophic lateral sclerosis. J. Neurol.260, 1782–1797 (2013). 10.1007/s00415-013-6877-3 [DOI] [PubMed] [Google Scholar]
- 21.Restuadi, R. et al. Functional characterisation of the amyotrophic lateral sclerosis risk locus GPX3/TNIP1. Genome Med14, 7 (2022). 10.1186/s13073-021-01006-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chang, C. et al. Extracellular Glutathione Peroxidase GPx3 and Its Role in Cancer. Cancers, 12, 2197 (2020). [DOI] [PMC free article] [PubMed]
- 23.Trolli, G., Résumé des observations réunies, au Kwango, au sujet de deux affections d’origine indéterminée. (1938).
- 24.Banea, M. et al. [High prevalence of konzo associated with a food shortage crisis in the Bandundu region of zaire]. Ann. Soc. Belg. Med Trop.72, 295–309 (1992). [PubMed] [Google Scholar]
- 25.Tshala-Katumbay, D. et al. Cassava food toxins, konzo disease, and neurodegeneration in sub-Sahara Africans. Neurology80, 949–951 (2013). 10.1212/WNL.0b013e3182840b81 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tshala Katumbay, D., Lukusa, V. M. & Eeg-Olofsson, K. E. EEG findings in Konzo: a spastic para/tetraparesis of acute onset. Clin. Electroencephalogr.31, 196–200 (2000). 10.1177/155005940003100408 [DOI] [PubMed] [Google Scholar]
- 27.Tshala-Katumbay, D. et al. Analysis of motor pathway involvement in konzo using transcranial electrical and magnetic stimulation. Muscle Nerve25, 230–235 (2002). 10.1002/mus.10029 [DOI] [PubMed] [Google Scholar]
- 28.Tylleskär, T. et al. Konzo: a distinct disease entity with selective upper motor neuron damage. J. Neurol. Neurosurg. Psychiatry56, 638–643 (1993). 10.1136/jnnp.56.6.638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rwatambuga, F. A. et al. Motor control and cognition deficits associated with protein carbamoylation in food (cassava) cyanogenic poisoning: Neurodegeneration and genomic perspectives. Food Chem. Toxicol.148, 111917 (2021). 10.1016/j.fct.2020.111917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Brigelius-Flohé, R. & Maiorino, M. Glutathione peroxidases. Biochim Biophys. Acta1830, 3289–3303 (2013). 10.1016/j.bbagen.2012.11.020 [DOI] [PubMed] [Google Scholar]
- 31.Nirgude, S. & Choudhary, B. Insights into the role of GPX3, a highly efficient plasma antioxidant, in cancer. Biochem Pharm.184, 114365 (2021). 10.1016/j.bcp.2020.114365 [DOI] [PubMed] [Google Scholar]
- 32.Moghadaszadeh, B. & Beggs, A. H. Selenoproteins and their impact on human health through diverse physiological pathways. Physiology21, 307–315 (2006). 10.1152/physiol.00021.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hill, K. E. et al. Selenoprotein P concentration in plasma is an index of selenium status in selenium-deficient and selenium-supplemented Chinese subjects. J. Nutr.126, 138–145 (1996). 10.1093/jn/126.1.138 [DOI] [PubMed] [Google Scholar]
- 34.Huang, W. et al. Selenoprotein P and glutathione peroxidase (EC 1.11.1.9) in plasma as indices of selenium status in relation to the intake of fish. Br. J. Nutr.73, 455–461 (1995). 10.1079/BJN19950047 [DOI] [PubMed] [Google Scholar]
- 35.Lopes, S. O. et al. Food Insecurity and Micronutrient Deficiency in Adults: A Systematic Review and Meta-Analysis. Nutrients, 15, 1074 (2023). [DOI] [PMC free article] [PubMed]
- 36.Haug, A. et al. How to use the world’s scarce selenium resources efficiently to increase the selenium concentration in food. Micro. Ecol. Health Dis.19, 209–228 (2007). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Singh, A. et al. Oxidative Stress: A Key Modulator in Neurodegenerative Diseases. Molecules, 24, 1583 (2019). [DOI] [PMC free article] [PubMed]
- 38.Benyamin, B. et al. Cross-ethnic meta-analysis identifies association of the GPX3-TNIP1 locus with amyotrophic lateral sclerosis. Nat. Commun.8, 611 (2017). 10.1038/s41467-017-00471-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang, J. Y. et al. Functional glutathione peroxidase 3 polymorphisms associated with increased risk of Taiwanese patients with gastric cancer. Clin. Chim. Acta411, 1432–1436 (2010). 10.1016/j.cca.2010.05.026 [DOI] [PubMed] [Google Scholar]
- 40.Liu, C. et al. Effects of GSTA1 and GPX3 Polymorphisms on the Risk of Schizophrenia in Chinese Han Population. Neuropsychiatr. Dis. Treat.16, 113–118 (2020). 10.2147/NDT.S236298 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Grond-Ginsbach, C. et al. GPx-3 gene promoter variation and the risk of arterial ischemic stroke. Stroke38, e23 (2007). author reply e24. 10.1161/STROKEAHA.106.479444 [DOI] [PubMed] [Google Scholar]
- 42.Tshala-Katumbay, D. et al. Impairments, disabilities and handicap pattern in konzo–a non-progressive spastic para/tetraparesis of acute onset. Disabil. Rehabil.23, 731–736 (2001). 10.1080/09638280110055075 [DOI] [PubMed] [Google Scholar]
- 43.Salim, S. Oxidative Stress and the Central Nervous System. J. Pharm. Exp. Ther.360, 201–205 (2017). 10.1124/jpet.116.237503 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Seeber, B. E. et al. The vitamin E-binding protein afamin is altered significantly in the peritoneal fluid of women with endometriosis. Fertil. Steril.94, 2923–2926 (2010). 10.1016/j.fertnstert.2010.05.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Jo, M. et al. Astrocytic Orosomucoid-2 Modulates Microglial Activation and Neuroinflammation. J. Neurosci.37, 2878–2894 (2017). 10.1523/JNEUROSCI.2534-16.2017 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Luo, Z. et al. Orosomucoid, an acute response protein with multiple modulating activities. J. Physiol. Biochem71, 329–340 (2015). 10.1007/s13105-015-0389-9 [DOI] [PubMed] [Google Scholar]
- 47.Biswas, S. K. Does the Interdependence between Oxidative Stress and Inflammation Explain the Antioxidant Paradox? Oxid. Med Cell Longev.2016, 5698931 (2016). 10.1155/2016/5698931 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Leung, L. L. K., J. Morser, J. Carboxypeptidase B2 and carboxypeptidase N in the crosstalk between coagulation, thrombosis, inflammation, and innate immunity. J. Thromb. Haemost., 16, 1474–1486 (2018). [DOI] [PubMed]
- 49.Zhou, Q. et al. Both plasma basic carboxypeptidases, carboxypeptidase B2 and carboxypeptidase N, regulate vascular leakage activity in mice. J. Thromb. Haemost.20, 238–244 (2022). 10.1111/jth.15551 [DOI] [PubMed] [Google Scholar]
- 50.Liu, Y. et al. Overexpression of zinc-α2-glycoprotein suppressed seizures and seizure-related neuroflammation in pentylenetetrazol-kindled rats. J. Neuroinflammation15, 92 (2018). 10.1186/s12974-018-1132-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Stipanuk, M. H. et al. Mammalian cysteine metabolism: new insights into regulation of cysteine metabolism. J. Nutr.136, 1652s–1659s (2006). 10.1093/jn/136.6.1652S [DOI] [PubMed] [Google Scholar]
- 52.Kimani, S. et al. Carbamoylation correlates of cyanate neuropathy and cyanide poisoning: relevance to the biomarkers of cassava cyanogenesis and motor system toxicity. Springerplus2, 647 (2013). 10.1186/2193-1801-2-647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Tor-Agbidye, J. et al. Sodium cyanate alters glutathione homeostasis in rodent brain: relationship to neurodegenerative diseases in protein-deficient malnourished populations in Africa. Brain Res.820, 12–19 (1999). 10.1016/S0006-8993(98)01343-2 [DOI] [PubMed] [Google Scholar]
- 54.Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem.75, 663–670 (2003). 10.1021/ac026117i [DOI] [PubMed] [Google Scholar]
- 55.Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol.26, 1367–1372 (2008). 10.1038/nbt.1511 [DOI] [PubMed] [Google Scholar]
- 56.Rohart, F. et al. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol.13, e1005752 (2017). 10.1371/journal.pcbi.1005752 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Pino, L. K. et al. The Skyline ecosystem: Informatics for quantitative mass spectrometry proteomics. Mass Spectrom. Rev.39, 229–244 (2020). [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of Additional Supplementary Files
Data Availability Statement
The mass spectrometry data generated in this study have been deposited in the Proteomexchange database under accession code PXD049973 and are freely accessible. The processed/cleaned mass spectrometry data associated with this manuscript are available in Supplementary Data 1. The additional ELISA and HPLC data generated in this study are also provided in the Supplementary Information/Source Data file. Source data are provided with this paper.
Proteomics data were processed using open-source software: MaxQuant v 1.6 3.4 for DDA experiment (https://maxquant.org/), DIA-NN v1.8 for DIA experiment (https://github.com/vdemichev/DiaNN), Skyline v.23.1.0.380 (https://skyline.ms/project/home/software/Skyline/begin.view) for SRM and PRM experiments. Statistics and visualization of proteomics data were done using R software v.4.1.1 (https://www.r-project.org/) including the R packages Mixomics (http://mixomics.org/) and diann (https://github.com/vdemichev/diann-rpackage).