Abstract
For over a century, bacterial infections have been studied through the lens of the one-microbe, one-disease paradigm. However, it is now clear that multi-pathogen infections are common, and many infectious diseases are inherently polymicrobial. These complex infections can involve a variety of pathogens, including viruses, bacteria, fungi, and parasites, with polyviral and viral-bacterial interactions being the most extensively studied. In this review, we focus on polybacterial infections, providing an in-depth analysis of the diverse strategies bacteria employ to thrive in co-infection scenarios. We examine the mechanisms of bacterial competition, competition avoidance through spatial or temporal separation, and cooperation. Given the association of polymicrobial infections with more severe clinical outcomes and heightened antibiotic tolerance, we also explore novel therapeutic targets to treat these increasingly common and complex infections. Although our review summarizes current knowledge, the vast scope of this phenomenon suggests that many more mechanisms remain undiscovered and warrant further investigation.
Keywords: bacterial competition, bacterial pathogens, competition avoidance, pathogen cooperation, polymicrobial infection, polymicrobial infection therapy
Introduction
Following Koch's postulates, linking single microbes to specific diseases, infectious disease research has focused on single pathogens growing in isolation (see [1] for a modern take on Koch’s postulates). This approach has proven instrumental in advancing knowledge by determining disease etiology, thereby promoting diagnosis, prevention, and treatment strategies. However, likely due to improved diagnostic techniques, the percentage of infections found to be polymicrobial (involving two or more infectious agents) has been on the rise, reaching up to 40% of pediatric clinical samples [2–7]. Nevertheless, the true prevalence of polymicrobial infections is likely underestimated, as diagnostic protocols are designed mainly for mono-infections and the likelihood of detecting polymicrobial infections varies based on the techniques employed and the clinician’s intent [8]. Although certain diseases (periodontitis, otitis media, urinary tract infections (UTIs)), are well-established as polymicrobial [9, 10], others are traditionally considered monomicrobial, with diagnosis focusing on the dominant pathogen. However, there is a growing recognition that many infectious diseases are multifactorial, involving multiple pathogens.
Theoretical models comparing monomicrobial and polymicrobial infections predict the latter result in greater host damage [11, 12]. Indeed, previous work showed polymicrobial infections can be associated with more severe clinical symptoms, possibly due to increased overall infectious burden or heightened virulence of at least one of the pathogens, and are often more tolerant to antibiotics [4, 13–18]. However, the outcome of polymicrobial infections is shaped by both the infectious agents and host context, making it difficult to predict how these interactions will unfold. Under some conditions, polymicrobial infections may be just as, or even less, severe than mono-infections. Regardless, the observation that, under some circumstances, polymicrobial infections can result in increased virulence emphasizes the need for clear criteria in diagnosing and treating such polymicrobial infections, likely leading to better patient outcomes [19, 20].
Although polymicrobial infections can involve viruses, bacteria, fungi, or parasites, this review focuses specifically on polybacterial infections. We discuss how multiple bacterial pathogens detect one another and modulate their behavior to either collaborate or compete. Also, it was not our goal to dissect the interactions that occur between pathogens and commensal microbes. Due to the complexity of the human microbiome, pathogens are constantly interacting with commensals, with significant implications for infection dynamics. However, this is beyond the scope of this review, and our focus is on the less explored realm of pathogen-pathogen interactions.
Polybacterial infections in humans
Polybacterial infections occur at various body sites, including the respiratory tract, gastrointestinal system, and skin [21–23]. Often, specific bacterial pairs are found together, suggesting these associations are not random. In skin and soft tissue infections (SSTIs), such as chronic wounds, pressure ulcers, and burns, polymicrobial infections are particularly prevalent [21]. Bacterial species associated with polymicrobial SSTIs include Staphylococcus, Enterobacteriaceae, Pseudomonas aeruginosa, Enterococcus, Acinetobacter baumannii, Streptococcus, and strict anaerobes, with Staphylococcus aureus being the most frequent agent [21]. Due to the high incidence of methicillin-resistant S. aureus (MRSA), these infections tend to show more severe clinical manifestations and limited treatment options [24, 25]. Most S. aureus-associated SSTIs are polymicrobial, with P. aeruginosa, Enterococcus faecalis, A. baumannii, Corynebacterium spp., or Escherichia coli being co-isolated [26–29]. In diabetic foot ulcers, S. aureus is most frequently isolated alongside Gram-negative bacteria (P. aeruginosa, Acinetobacter, E. coli, Enterobacter, Citrobacter, Proteus, and Klebsiella), although some Gram-positive bacteria, such as Enterococcus, are also co-isolated [26, 30]. In pressure ulcers, S. aureus is frequently co-isolated with P. aeruginosa, E. coli, P. mirabilis, Enterobacter cloacae, and E. faecalis [31, 32], and, in burn wounds, with P. aeruginosa [33].
In the oral cavity, another known site for polymicrobial infections, Porphyromonas gingivalis, Treponema denticola, Tannerella forsythia, and Fusobacterium nucleatum are usually co-isolated [22, 34, 35]. These infections typically lead to periodontal disease, potentially causing tooth loss. Although bacteria directly harm tissues, damage is mostly by triggering host cell activation, leading to production of tissue-degrading substances. Although most research on periodontal disease uses single-species infection models, recent studies adopted polybacterial models, better mimicking natural disease [22, 34, 35]. In a murine co-infection abscess model with F. nucleatum and P. gingivalis, pathogen ratios and timing of infection were critical determinants of disease severity [34]. Higher proportions of F. nucleatum significantly reduced lesion formation. Other infection protocols, such as simultaneous inoculation with both pathogens at different sites, increased lesion formation by a trypsin-like protease of P. gingivalis [34]. These data suggest intricate interactions between pathogens, also likely involving the host immune system, are at play.
UTIs have historically been associated with single pathogens, like uropathogenic E. coli [36]. However, polybacterial infections are frequent in specific populations, such as individuals using urinary catheters, the elderly, and the immunocompromised [37]. Assessment of polymicrobial UTIs is complicated by sample contamination with periurethral and vaginal microbes [36]. Nevertheless, up to 86% of catheter-associated UTIs are polymicrobial [38]. The pathogens involved include E. coli, Klebsiella pneumoniae, P. aeruginosa, Proteus mirabilis, and Providencia stuartii. Among these, P. mirabilis and P. stuartii are consistently co-isolated from catheter-associated UTIs and increase urolithiasis and bacteremia in mice [38]. A recent analysis of urine and catheter colonization in long-term catheterized individuals revealed that long-term co-isolation was observed with uropathogenic E. coli and E. faecalis [39]. In vitro data, using artificial urine media, showed co-culturing with E.coli increases E. faecalis growth.
Although viruses are the primary cause of respiratory diseases, several bacteria can lead to severe illness, including Streptococcus pneumoniae, Streptococcus pyogenes, Haemophilus influenzae, S. aureus, Neisseria meningitidis, Mycobacterium tuberculosis, and Bordetella pertussis. Polymicrobial respiratory infections are widely reported, but consist mainly of viral-bacterial infections, where bacterial agents are considered secondary to initial damage caused by viruses [40]. Polybacterial lung infections are most commonly observed in patients with severe conditions, such as patients who underwent major surgery, chemotherapy and radiotherapy patients, and patients with the genetic disease cystic fibrosis (CF). In CF, infections often include S. aureus, H. influenzae, P. aeruginosa, and Burkholderia cepacia [41]. The interplay between these species in CF is intricate and dynamic. Some species are associated with poor clinical outcomes, whereas others are not. Ratios between different species vary widely and display longitudinal variation, also being affected by host factors, such as age [42].
Detecting polybacterial infections in the gastrointestinal tract is challenging due to the dense microbiome that populates this environment, making it difficult to evaluate the impact of commensals during infection. Also, several gut commensals can act as pathogens depending on the context (barrier breach, microbial density, acquisition of virulence genes through horizontal gene transfer). Interactions between gut commensals and classic enteric pathogens are common, and play significant roles in microbial virulence and disease severity. A healthy intestinal microbiome is associated with host colonization resistance to pathogens, adding complexity to pathogen-pathogen interactions [43]. A study of two cohorts of infants in Bangladesh found that two pathogen pairs, Enterotoxigenic E. coli (ETEC) with Enteropathogenic E. coli (EPEC) and ETEC with Campylobacter spp., appeared together more frequently than expected at random, suggesting these pairs are associated with gastrointestinal coinfections [44]. Additional studies showed multiple enteric pathogens can be detected in diarrheal samples, many of which carry ETEC and Vibrio cholerae [4, 6, 45]. Patients with these co-infections tend to experience more days with diarrhea, worse dehydration, and a need for higher intravenous fluid intake.
Interactions between bacterial species in complex communities can span from cooperation to competition, and this is likely also the case for pathogen-pathogen interactions [46]. In some cases, bacteria compete for resources and niche dominance, inhibiting each other's growth and colonization. Other scenarios involve cooperation, where different species support each other's survival and virulence through metabolic cross-feeding and biofilm formation [46]. These interactions impact the progression and severity of infections, underscoring the need to understand their underlying dynamics to develop more effective treatment strategies.
Challenges in the diagnosis of polymicrobial infections
Currently, several methods are available for diagnosing monomicrobial infections, with culture-based techniques regarded as the gold standard [47]. Culture-based methods are cost-effective, require minimal specialized equipment and training, and uniquely allow for the detection of viable bacteria, which is essential for phenotypic antimicrobial susceptibility testing. However, they are time-consuming, require specific growth conditions for fastidious strains, and are often ineffective once antibiotic treatment has begun. In polymicrobial infections, traditional culture-based techniques are inherently limited, often selectively detecting the most abundant or the fastest-growing species. This methodological bias can lead to incomplete identification of the causative agents of these infections [48]. Molecular methods involving DNA sequencing, DNA/protein fingerprinting technology, and target-specific testing, such as multiplex PCR, digital PCR, Peptide Nucleic Acid Fluorescence In Situ Hybridization (PNA-FISH), and Molecular Antimicrobial Susceptibility Testing (AST) have significantly enhanced the diagnosis of polymicrobial infections [48–52]. These approaches enable rapid pathogen detection, with high specificity and sensitivity, and support multiplex assays for the simultaneous diagnosis of multiple pathogens. Yet, these approaches are relatively expensive for routine clinical use, requiring specialized equipment and training. Also, they cannot distinguish between viable and non-viable microbes and provide limited antimicrobial susceptibility information.
Another useful approach is the use of immunological-based methods, which include antigen detection techniques, such as ELISA, and serological methods that target host antibody responses. Antigen detection techniques offer relatively rapid results and are suitable for automation; however, they may suffer from reduced sensitivity and specificity and cannot confirm organism viability, as they detect only microbial components [53, 54]. Serological methods are valuable for identifying infections by measuring host immune responses, but they are less effective for early diagnosis and often cannot differentiate between past and current infections [55]. Microscopy and mass-spectrometry methods both rely on prior microbe isolation but offer rapid results, which are based on phenotypic presentation [56, 57]. These methods have been optimized for the detection of a single pathogen and therefore require adaptation to multiplex platforms to effectively identify and characterize polymicrobial infections [58, 59].
In order to be valuable, diagnostic tools must be accurate (high specificity and sensitivity) and optimally deliver rapid results. They should also be cost-effective and accessible in multiple health care settings. For polymicrobial infections, diagnostic tools must enable simultaneous detection of all involved pathogens, regardless of their growth or metabolic requirements, while also determining their antimicrobial profile, allowing for proper treatment of these infections. While significant advancements in bacterial diagnostic methods have occurred in recent years, options for the diagnosis of polymicrobial infections remain limited, creating a critical gap between clinical need and available technology.
Strategies to succeed in polymicrobial environments
Bacterial infections can stem from several contamination sources, including food and water, surfaces and objects, especially in healthcare settings, and environmental sources (soil, air) [60]. These sources may harbor single pathogens, but more commonly contain multiple pathogens that can be introduced simultaneously into the host. The presence of multiple bacteria in contaminated environments can drive diverse strategies that promote successful infections: (i) competition, (ii) competition avoidance, or (iii) cooperation (Fig. 1).
Figure 1.
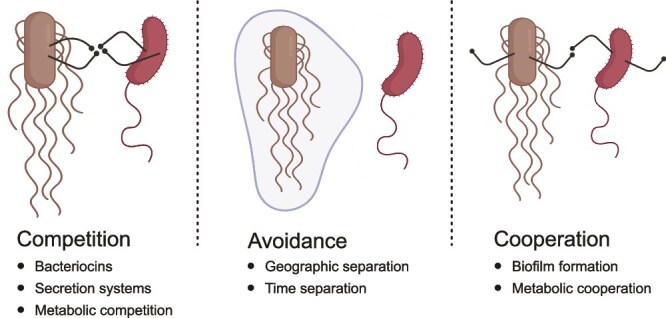
Strategies for survival and growth in polymicrobial environments. The figure illustrates the main strategies employed by bacteria in mixed communities: (1) bacterial competition, where organisms directly antagonize competitors through mechanisms such as production of bacteriocins, delivery of effectors by secretion systems, or metabolic competition for limited resources; (2) competition avoidance, where bacteria minimize direct competition through spatial or temporal separation; and (3) bacterial cooperation, where bacteria engage in mutually beneficial interactions including metabolite sharing, collaborative biofilm formation, and cross-feeding relationships. These strategies are not mutually exclusive, and bacteria often employ multiple approaches simultaneously depending on the environmental conditions and community composition.
Bacterial competition
Bacterial competition relies on the concept that only the most resilient and efficient strains will flourish in competitive environments. To achieve that, bacteria develop mechanisms to kill competitors or compete more effectively for nutrients or space [61]. This natural selection can enhance beneficial traits, like antibiotic resistance and metabolic efficiency. Studying bacterial competition led to the discovery of antimicrobials, as bacteria often produce compounds to inhibit competitors [62, 63]. Although pathogens with effective competitive strategies frequently dominate infections, leading to mono-infection, competitors may display defensive strategies, such as immunity proteins or altered targets to reduce susceptibility, allowing them to persist alongside the dominant pathogen. Furthermore, successful competition strategies by pathogens can affect the microbiome, disrupting colonization resistance and favoring polymicrobial infections.
Bacteriocins
Bacteriocins are small proteinaceous molecules that inhibit bacterial growth, allowing producers to outcompete other strains for resources and space [64]. Bacteriocins are highly diverse in structure, size, mode of killing, and mechanisms of production and secretion. Bacteriocins can be classified according to their target (membrane, DNA, RNA, proteins) or according to their function (pore-forming activity, nuclease activity, inhibition of peptidoglycan production, modulation of enzyme activity, quorum-sensing (QS) interference) [65].
Due to their ability to target specific bacteria and relatively high heat stability, bacteriocins have been investigated as food preservatives [66]. Recently, the potential of bacteriocins in targeting antibiotic-resistant pathogens has been explored. Commensal E. coli was shown to produce small bacteriocins, such as the microcin H47 toxin, that limit Salmonella enterica serovar Typhimurium and E. coli pathogenesis toward host cells [67, 68]. Nevertheless, most bacteriocins have narrow spectrum, typically inhibiting or killing genetically related bacteria, suggesting they are probably not major players in polymicrobial infections. However, recent studies reported that bacteriocins produced by Gram-positive bacteria exhibit broader spectrum, targeting other bacterial species or genera [69]. This suggests bacteriocins may play significant roles in bacterial interactions.
Bacterial secretion systems
Bacteria evolved secretion systems to deliver proteins, nucleic acids, and metabolites to the surrounding medium or directly into cells [70, 71]. Some of these secretion systems are essential for growth and survival in some environments, whereas others are dedicated to secreting components crucial for host infection or competition with nearby bacteria. The effectors of these secretion systems can act on various targets within the bacterial pathogens or the host cells they infect (Fig. 2). Among the various types of secretion systems (I-IX), four are particularly known for their roles in facilitating bacterial competition [71].
Figure 2.
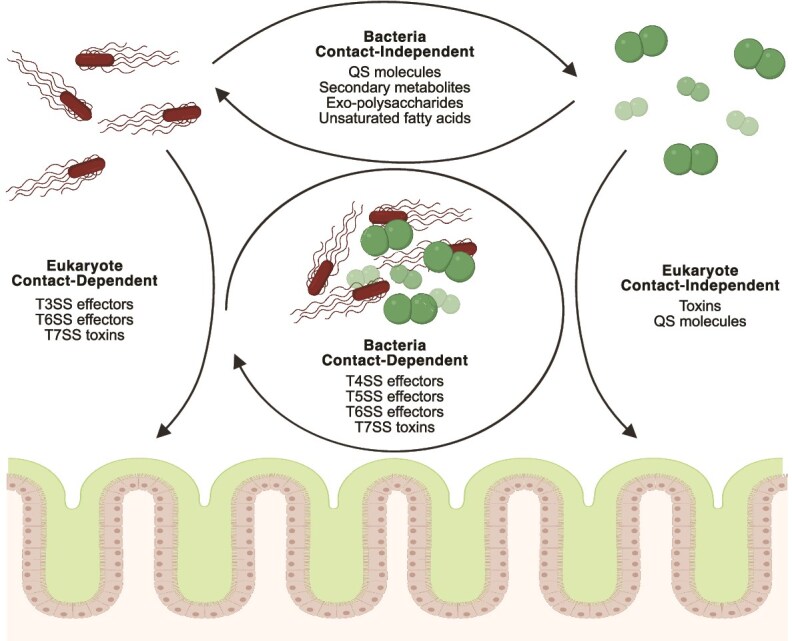
Effectors used by bacteria to target other pathogens or host cells in polymicrobial infections. Bacteria enhance their colonization and replication by manipulating host cells and competing against other microbes through the activity of various effectors. These effectors are delivered via contact-independent secretion, including the release of quorum sensing (QS) molecules, secondary metabolites, exopolysaccharides, and unsaturated fatty acids that influence neighboring bacteria, as well as secreted toxins and QS molecules that act toward eukaryotic cells. In contrast, contact-dependent secretion relies on direct contact and the use of secretion systems such as the T4SS, T5SS, T6SS, and T7SS to target other bacteria and T3SS, T6SS, and T7SS to deliver effectors into eukaryotic cells. These strategies contribute to bacterial competition, cooperation, and virulence in different environments.
Type 4 secretion systems (T4SSs)—T4SSs are large protein complexes predominantly found in Gram-negative bacteria. T4SSs transport substrates directly from the cytoplasm of donor cells into the cytoplasm of recipients. T4SSs can interact with both eukaryotic and prokaryotic cells and can transfer DNA between bacteria or virulence proteins (effectors) into prokaryotic/eukaryotic cells. T4SSs facilitate bacterial competition by delivering toxic effectors into competitors within the same niche. These toxins employ diverse killing mechanisms, including enzymatic degradation of essential cellular components or membrane disruption. For instance, Stenotrophomonas maltophilia, a bacterium associated with medical device-related infections, induces contact-dependent killing of E. coli, K. pneumoniae, and P. aeruginosa by T4SS toxins [72]. A recent study of two soil bacteria, Lysobacter enzymogenes and Pseudomonas fluorescens, identified a mechanism through which T4SS promotes bacterial competition [73]. The translocated effector disrupts the production of QS molecules in P. fluorescens, leading to reduced biofilm formation and increased susceptibility to killing. Moreover, due to their well-known role in horizontal gene transfer, T4SS could potentially increase pathogen competition by horizontally transferring genes that could enhance or reduce bacterial virulence and survival during infection.
Type 5 secretion systems (T5SSs)—Protein secretion by T5SSs takes place in a two-step procedure and is autonomous—the secreted protein is responsible for its transport across the membrane. The first secretion step, across the inner membrane and into periplasmic space, is executed by the general bacterial secretion route (Sec pathway), an essential, ubiquitous, and universal export machinery. Sec transport is mediated by a short, cleavable amino-terminal signal peptide recognized by the Sec machinery [74]. The second step, through the outer membrane to the extracellular environment, is mediated by the beta-barrel domain at the C-terminus of the T5SS protein. This domain folds across the outer membrane to create a pore through which the secreted portion of the protein, the passenger domain, translocates to the bacterial surface. In most cases, the passenger domain is released after translocation, by cleavage between the passenger domain and the beta-barrel domain, which remains anchored in the outer membrane [75]. Passenger domains of T5SSs have a broad spectrum of functions, including adhesion, enzymatic activity, immune modulation, and even toxin production. T5SS substrates can act in their cell-surface adhered form (contact-dependent) or as secreted proteins (contact-independent). T5SSs are involved in bacterial competition by facilitating toxin delivery (nucleases, lipases, proteases) into neighboring bacteria, damaging them by degrading essential biomolecules. One of the most documented T5SS involved in competition is the Contact-Depended growth Inhibition (Cdi) system, CdiAB [76, 77]. CdiA has a C-terminal effector domain that is delivered into competitors by CdiB. Once inside cells, the effector degrades nucleic acids, inhibiting growth and, ultimately, causing death [78]. The specificity of these systems ensures that toxins selectively target competing species/strains, sparing those that express immunity proteins that neutralize effectors [79].
Type 6 secretion systems (T6SSs)—The T6SS is another contact-dependent killing mechanism employed by Gram-negative bacteria. T6SSs are needle-like structures that inject toxic effectors directly into target cells (prokaryotic or eukaryotic), making them versatile tools for bacterial competition and virulence [80–83]. Structurally, T6SSs resemble a phage tail or a contractile injection system with several key components (baseplate, sheath, needle). There are many examples of T6SS involvement in interbacterial interactions, including those that allow pathogens to outcompete commensals in their environment. However, only a few studies explored T6SSs in pathogen-pathogen interactions. A recent study revealed that Aeromonas dhakensis, a seafood bacterium, uses its T6SS to eliminate Vibrio competitors in shrimp [84]. Similarly, another study showed that T6SS effectors of P. syringea possess DNase activity, actively killing E. coli through DNA degradation [85]. Moreover, the T6SS of Shigella sonnei provides a competitive advantage against S. flexneri in mixed cultures [86]. By injecting toxins into competitors, T6SSs disrupt cell membranes, degrade cell walls, and interfere with essential processes, inhibiting growth or killing them [87]. This provides a competitive advantage in microbial communities and aids in establishing infections. Many pathogens, such as V. cholerae and P. aeruginosa, express T6SSs with high levels of antagonism against a broad range of commensals and pathogens, including E. coli and S. enterica.
Type 7 secretion systems (T7SSs)—The T7SS is found mostly in Gram-positive bacteria and has been implicated in nutrient acquisition, interbacterial competition, and virulence [88]. Given their limited similarity, T7SSs are classified into two subtypes; T7SSa in Actinobacteria and T7SSb in Firmicutes. While T7SSs are not as well-studied as other secretion systems, previous studies showed that some pathogens, such as S. aureus and Enterococcus faecalis, can promote the killing of competitors by delivering toxins through their T7SSs [89, 90].
Metabolic competition
One of the most common ways microbes compete in polymicrobial environments is by fighting for nutrients. Because many bacteria share metabolic needs, competition for resources is intense. This leads to the evolution of metabolic traits that allow bacteria to bypass competition, often accessing less common nutrients that competitors cannot utilize for growth. Metabolic competition is widely recognized as a key mechanism by which the microbiome provides colonization resistance against invaders. Multiple pathogens also use this strategy when competing against the microbiome and other pathogens alike.
Metabolic competition in the gut environment
Ethanolamine is a gut nutrient derived from membrane phospholipids. Recently, the opportunistic pathogen Klebsiella pneumoniae was shown to utilize ethanolamine as carbon and nitrogen sources during gut colonization, an ability conferred by genes encoded within two eut loci, which are strongly induced during gut colonization [91]. Also, single and double mutants of eut loci showed defects in gut colonization in the presence of an intact gut microbiome. However, gut microbiome ablation by antibiotic treatment eliminated these differences, suggesting eut loci are involved in the ability of K. pneumoniae to outcompete the microbiome [91].
Besides K. pneumoniae, other pathogens use ethanolamine as a substrate during metabolic competition. An eutC mutant of S. enterica exhibited a significant competitive disadvantage in mouse co-infections with the wildtype strain [92]. S. enterica cannot efficiently utilize ethanolamine as a carbon source during fermentative growth in vitro. However, during anaerobic respiration in the presence of the alternative electron acceptor tetrathionate, S. enterica can grow on ethanolamine as the sole carbon source. The requirement for tetrathionate respiration also occurs in vivo, as an eutC mutant showed no competitive defect against the wildtype when ttrA, the gene encoding the tetrathionate reductase subunit A, was absent. Therefore, S. enterica uses anaerobic respiration in the gut to access a substrate that provides limited benefit to the predominantly fermentative microbial community of the anoxic mammalian gut [92]. Whether K. pneumoniae and S. enterica compete for ethanolamine in vivo remains unknown. However, that both pathogens devised mechanisms to access this nutrient suggests such competition likely occurs. Also, although considered a pathogen, K. pneumoniae can asymptomatically colonize the human gut. Under these conditions, K. pneumoniae may prevent S. enterica colonization by utilizing ethanolamine.
Metabolic competition in the CF lungs
CF is caused by mutations that impair chloride transport across the cell membrane, disturbing osmotic balance at the epithelial surface [93]. This results in dehydrated, thick mucus on the airway surfaces, favoring the attachment and proliferation of bacteria, which resist clearance. Polybacterial infections in CF commonly involve S. aureus and P. aeruginosa, highly proficient biofilm formers. A recent study demonstrated that P. aeruginosa secretes polysaccharides that specifically inhibit S. aureus growth, a finding confirmed using CF clinical isolates [94]. Recently, metabolic competition between these pathogens was studied using a synthetic CF medium [95]. Metabolomics was used to determine the nutrients consumed and metabolites produced by these pathogens. As expected, a vast overlap of metabolites consumed and produced was observed when different strains of the same species were compared. However, a sizable overlap was also observed in the metabolites consumed by both pathogens. Specifically, 15 metabolites, including beta-leucine, tryptophan, homoserine, indole-3-acrylic acid, serine, fructose, lactate, and glutamate, were degraded by all strains tested, indicating they can be reliable sources of carbon and energy for both species. Although these experiments were performed in monocultures, the vast metabolic overlap identified likely has implications during co-infection in vivo.
Aside from the competition between S. aureus and P. aeruginosa for carbon and energy sources, P. aeruginosa can also directly alter S. aureus metabolism, with implications for competition. A co-culture model of S. aureus and P. aeruginosa on CFTR-deficient human bronchial epithelial cells showed that P. aeruginosa shifts S. aureus metabolism from aerobic respiration to fermentation [96]. mRNA sequencing revealed P. aeruginosa strongly upregulated S. aureus fermentation genes (≥10-fold), an effect reproduced with P. aeruginosa cell-free supernatants. In contrast, the effect of S. aureus on P. aeruginosa gene expression was minimal, with only one gene being differentially regulated. Production of 2-heptyl-4-hydroxyquinoline N-oxide (HQNO) and the siderophores pyoverdine and pyochelin by P. aeruginosa was responsible for this phenomenon. These mechanisms of competition between S. aureus and P. aeruginosa have biological relevance. During lung infections of CF patients, although both S. aureus and P. aeruginosa are present, a succession is commonly observed, where S. aureus dominates the CF microbial community earlier in life, with P. aeruginosa taking over later on.
Competition avoidance
Competitive avoidance describes a state where bacterial species that use similar resources coexist by avoiding one another. This can be achieved by maintaining physical or temporal separation, reducing direct competition. In some of the examples below, competition avoidance may seem like an active process, whereas in others it may be a simple consequence of extraneous factors. Whether active or passive, this phenomenon allows for a balance where multiple species can thrive within the same environment without outcompeting each other to extinction.
Geographic separation
One strategy to avoid competition is to geographically divide the niche into smaller areas. Although this strategy was mainly demonstrated among fungal pathogens [97, 98], there are examples for geographic separation between bacterial pathogens and commensals. Bacterial populations can create physical separation by producing compounds, such as extracellular polysaccharides (EPS), polyamides, and extracellular DNA, that define distinct microenvironments forming physical barriers between cells [99]. One example is Cutibacterium acnes, a prevalent member of the skin microbiota that comprises ~92% of bacterial populations of oily skin regions. C. acnes was shown to maintain intraspecies diversity by promoting spatial segregation of different strains across distinct skin pores [100]. Each pore is colonized by a single strain, although multiple strains are present in neighboring pores. No genetic adaptations were found to drive this phenomenon; pore anatomy and physiology seem to induce bottlenecking, avoiding direct competition between different C. acnes strains, and enhancing overall strain diversity. Although C. acnes is generally a commensal, it can cause significant prosthetic joint infections in humans, and it is possible that geographic separation also occurs during polymicrobial infections [101].
Spatial separation between P. aeruginosa and S. aureus through a cell-free physical barrier has also been described, a phenomenon derived from the production of phenol-soluble modulins (PSMs) by S. aureus colonies [102]. PSMs form amyloid fibrils around the colonies, deflecting the surfactant flow created by P. aeruginosa and steering its swarms away. Recently, it was also shown that S. aureus produces extracellular surfactants that affect P. aeruginosa motility, enabling it to spread across semi-solid surfaces that would otherwise restrict its movement [103]. Using another species’ surfactants would, in theory, allow P. aeruginosa to access different environmental niches within the host while avoiding competition. Another study reported that V. cholerae can produce an EPS that functions as an armor that protects it from T6SS-mediated attacks by heterologous bacterial species, enabling coexistence in the same niche [104].
Physical separation between pathogens can also be achieved through their natural preference for intracellular versus extracellular environments. A recent study that examined the prevalence of diarrhoeal pathogens in pediatric samples found high numbers of bacterial poly-infections of intracellular-extracellular pathogens, such as the triplets Shigella, Enteroinvasive E. coli (EIEC), and EPEC; Enteroaggregative E. coli (EAEC), EPEC and Campylobacter; and the pair EAEC and Salmonella [105]. Although we are unaware of evidence that the presence of a competitor in the extracellular environment could drive invasion by an intracellular pathogen, the ability of intracellular pathogens to physically separate themselves from extracellular competitors may allow them to survive and multiply in what would be an otherwise hostile environment due to pathogen competition.
Time separation
Sequential bacterial infections are co-infections that occur at slightly different times. Until recently, these infections were considered to stem from the arrival of distinct infectious agents at different times. The first (primary) infection was considered to facilitate sequential infections (secondary or super-infections) by harming the immune response, altering the native microbiome, or disrupting host tissues. However, recent models suggest these infections might also stem from the simultaneous arrival of multiple pathogens into the host, but with clinical manifestations occurring at different times, as a means to support both infections. In most sequential infections, the primary pathogen is a virus [106, 107]. However, specific pairs of bacterial pathogens were reported to occur in sequential infections. For example, following M. tuberculosis infections, superinfection with H. influenzae or S. aureus is common [108]. A high incidence of P. aeruginosa infection (23.5%) was detected among patients receiving the antibiotic tigecycline, mainly used to treat nosocomial infections by MRSA, penicillin-resistant S. pneumoniae, vancomycin-resistant Enterococcus spp., and Enterobacteriaceae, and P. aeruginosa likely appears as a late-stage co-infecting pathogen in these infections [109].
Time separation between pathogens was recently reported between V. cholerae and EPEC. V. cholerae, the causative agent of cholera, causes an estimated 3-5 million cases annually, with ~100,000 deaths. In endemic countries, about half of the deaths occur in young children [7, 110]. EPEC is a prevalent diarrheal pathogen that leads to pediatric and persistent diarrhea [111]. While EPEC-related illness has not been highly prevalent in developed countries since the 1950’s, it is recently reemerging, with severe disease outcomes in the community and hospital settings [111]. V. cholerae and EPEC share the same route of infection, source of infection (contaminated water or food), and symptoms (gastroenteritis). Additionally, both display tropism toward the small intestine and are often found together in clinical samples. A recent study showed EPEC senses the presence of V. cholerae and adjusts its virulence accordingly to coordinate successful host colonization [112]. By measuring the concentration of the main V. cholerae QS molecule (Cholerae Autoinducer-1 (CAI-1)), EPEC monitors the population size and virulence status of V. cholerae. To optimize its infection for times of minimal competition, EPEC upregulates its virulence when it detects high CAI-1 concentrations, indicating downregulation of V. cholerae virulence and their preparation to leave the host [112]. These results were unexpected, as CAI-1 was previously considered an exclusively intra-genus (Vibrio) signal [113, 114]. A follow-up study demonstrated that production of indole by members of the gut microbiome, like Bacteroides thetaiotaomicron, disrupt EPEC’s ability to sense CAI-1 and time its infection to periods of lower competition with V. cholerae [115]. These findings suggest that bacterial interactions involving members of the gut microbiome may also affect the time separation displayed by V. cholerae and EPEC during polymicrobial infection.
Co-infections of the sexually transmitted bacteria Neisseria gonorrhoeae and Chlamydia trachomatis, which infect the epithelium of the endocervix in women and the urethra in men, also exemplify time separation. Researchers found that, during infection, when C. trachomatis is exposed to extracellular N. gonorrhoeae at mid-cycle, it continues to grow but does not transition into the infectious elementary body state as fast as it does during mono-infections [116]. This slower kinetics of bacterial development results in delayed release of progeny, which can benefit C. trachomatis by timing the release of infectious elementary bodies to periods when the environment is less hostile.
Pathogen cooperation
Microbial cooperation enhances the fitness of all microbes by enabling them to share resources, create protective structures that promote survival under environmental stresses, like antibiotics, and perform complementary metabolic functions. Microbial synergy can occur between pathogens during infection, allowing them to colonize more effectively. For instance, one pathogen might create conditions that favor the growth of another, such as altering the local environment to reduce immune responses or providing essential nutrients. These cooperative interactions can contribute to the persistence and virulence of pathogens, leading to harder-to-treat infections [117, 118].
Biofilm formation
Biofilms have increased resistance to antimicrobial agents and immune responses [119, 120]. Generally, monobacterial biofilms are more commonly associated with acute infections, such as UTIs or uncomplicated skin infections. In contrast, biofilms containing multiple species are more frequently observed in chronic and harder-to-treat infections. Polymicrobial biofilms are frequently found in periodontitis, otitis media, diabetic foot wounds, burn wounds, and infections of CF patients. These polymicrobial biofilms provide an optimal environment for different species to interact antagonistically, additively, or synergistically, impacting their persistence and stability during infection [31, 121]. Consistent with their detrimental effect on treatment, polymicrobial infections in chronic wounds delay closure and promote antibiotic resistance compared to mono-infections [122, 123]. Furthermore, polymicrobial biofilms were shown to initiate and aggravate CF and burn infections [124, 125]. By establishing multi-species biofilms, bacteria gain competitive advantages, including antimicrobial resistance, metabolic cooperation, QS regulation, and higher chances of horizontal gene transfer.
Two common partners in polymicrobial infections, P. aeruginosa and S. aureus, have been shown to cooperate in an in vitro wound model by displaying enhanced antibiotic tolerance [126]. Enhanced resistance is dependent on components of the host matrix and extracellular polymeric substances produced by both species. The increased aggregation and tobramycin resistance exhibited by P. aeruginosa is dependent on the interaction between S. aureus protein A and P. aeruginosa Psl (an exopolysaccharide) [127]. In contrast, S. aureus senses HQNO produced by P. aeruginosa, forming small colony variants, which are aminoglycoside-resistant and associated with persistent infections [127, 128]. It has also been shown that S. aureus displays passive resistance to ampicillin by becoming closely surrounded by a dense layer of ampicillin-resistant P. aeruginosa [129].
The spatial organization of biofilms can modulate virulence and overall progression of infection. Cooperative interactions in biofilms have been classified as derived from physical contact or chemical signaling [130]. One example of contact-dependent interaction that segregates species in biofilms happens between P. aeruginosa and S. aureus. In CF, several variants of P. aeruginosa overproduce the polysaccharide alginate, enhancing biofilm formation with S. aureus and B. cepacia and leading to highly structured biofilms with increased antibiotic resistance [131, 132]. This species organization impacts biofilm stability and persistence, consequently affecting the success of the infection. In another case of contact-dependent interaction, two periodontal pathogens, Treponema denticola and Porphyromonas gingivalis, have been shown to form thicker biofilms when co-cultured [133], a phenomenon dependent on the expression of membrane-associated proteins or adhesins, gingipains, by P. gingivalis.
Within polymicrobial biofilms, different species can detect one another by sensing metabolites. These metabolites include byproducts produced during cross-feeding or dedicated signals, such as QS molecules. Metabolite-based detection has been shown to occur between two opportunistic pathogens commonly associated with otitis media, Moraxella catarrhalis and H. influenzae [10]. Although M. catarrhalis cannot produce QS autoinducer-2 (AI-2), it responds to it. Co-culturing M. catarrhalis with H. influenzae, which produces AI-2, results in more robust biofilms, which are more tolerant to antimicrobials. In an animal model of otitis media, the interaction between these species was shown to support higher levels of M. catarrhalis growth in an AI-2-dependent fashion. A similar pattern was observed for P. aeruginosa; although P. aeruginosa cannot produce AI-2, it enhances virulence factor expression in CF lung infections in response to AI-2 from co-infecting bacteria [134].
Metabolic cooperation
Metabolic cooperation can promote polymicrobial interactions, as seen in E. faecalis and E. coli, which are commonly found in urinary catheters and wound infections. Co-culturing E. faecalis and E. coli enhanced their ability to form biofilms under iron-limiting conditions [135]. The underlying mechanism involves L-ornithine production by E. faecalis, triggering E. coli to produce the siderophore enterobactin, boosting iron acquisition, bacterial growth, and biofilm formation. In a mouse wound infection model, E. coli reached significantly higher colonization levels during co-infections.
Synergistic interactions based on metabolic cross-feeding also occur during periodontitis. Aggregatibacter actinomycetemcomitans is an oral pathogen found in patients with aggressive periodontitis. These infections are often polymicrobial, with Streptocoocus gordonii frequently present. It has been shown that an A. actinomycetemcomitans mutant that cannot metabolize lactose can cause infections in monoculture, but cannot synergize with S. gordonii in polymicrobial infections. Because A. actinomycetemcomitans utilizes lactate as a carbon source and S. gordonii produces large amounts of lactate during carbohydrate metabolism, A. actinomycetemcomitans cross-feeds on lactate to enhance the severity of the disease [136]. In turn, A. actinomycetemcomitans neutralizes S. gordonii-produced hydrogen peroxide via catalase. Hydrogen peroxide also upregulates A. actinomycetemcomitans apiA expression; apiA encodes a protein that binds human factor H, a regulator of the alternative complement pathway [137]. This interaction increases resistance to immune-mediated killing by A. actinomycetemcomitans when co-cultured with S. gordonii.
Targeting pathogen interactions for therapy
Current treatments for infections typically target individual pathogens despite the high prevalence of polymicrobial infections. This narrow focus overlooks the complex dynamics often present in mixed infections. Treatment of polymicrobial infections can be more challenging because different bacterial species often display different antimicrobial susceptibility patterns. Furthermore, synergistic interactions enhance biofilm formation, adding an extra layer of resistance to many treatments [128, 138, 139]. However, the presence of multiple pathogens does not always result in an additive virulence effect. This was demonstrated using a Galleria mellonella model, where the primary determinant of virulence in a polymicrobial infection was found to be the most virulent species [140]. In certain pathogen combinations, the presence of a co-infecting pathogen actually reduced the fitness of the primary pathogen. This was specific to certain bacterial pairs, highlighting the complexity of predicting the outcome of polymicrobial infections. Nevertheless, developing effective treatment strategies for polybacterial infections remains crucial and requires a multi-faceted approach (Fig. 3). Two main antimicrobial strategies can be employed: broad-spectrum antimicrobials targeting multiple bacterial species simultaneously or combination therapies integrating several narrow-spectrum drugs, each selectively targeting one pathogen. Because many polybacterial infections involve biofilms, which are significantly more resistant, anti-biofilm drugs can be deployed to target these resilient infections. Furthermore, targeting bacterial communication through QS inhibitors represents an innovative approach that could prevent bacterial coordination and reduce virulence. Anti-biofilm and anti-QS therapeutics could be deployed either independently or in combination with antibiotics. Disrupting these virulence mechanisms may boost the host's immune response to eradicate the pathogens on its own or make bacteria more susceptible to antibiotics. While these strategies offer promising alternatives to conventional treatments, their clinical potential remains largely underexplored and warrants further investigation.
Figure 3.
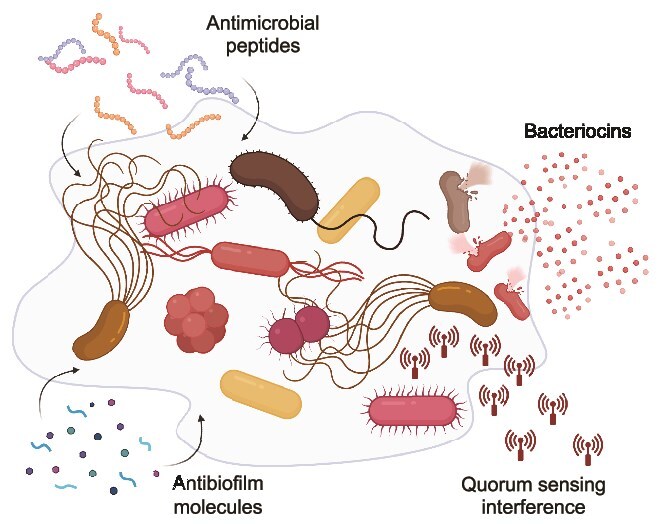
Therapeutic strategies targeting polymicrobial infections. The illustration depicts a polymicrobial biofilm and various therapeutic approaches that can be used alone or in combination with conventional antibiotics to combat these complex bacterial communities. Antibiofilm molecules may target the extracellular matrix, including matrix-degrading enzymes and dispersal agents that compromise biofilm structural integrity. They may also interfere with biofilm maturation processes. Bacterial cells within the biofilm can be targeted by antimicrobial peptides, which disrupt bacterial cell membranes, and bacteriocins, which provide diverse antimicrobial activity. Quorum-sensing inhibitors interfere with bacterial cell-cell communication through multiple mechanisms, including degradation of signaling molecules and receptor antagonism. This multi-modal approach addresses the various aspects of the polymicrobial pathogen community, potentially offering more effective treatment strategies for polymicrobial infections compared to conventional antibiotics alone.
Bacteriocins
Bacteriocins emerged as potent antimicrobials for the treatment of mono-infections. Traditionally, bacteriocins are considered to have a narrow spectrum of activity, primarily targeting genetically related bacteria. As a result, their use is largely confined to the food industry. However, recent studies demonstrated that several bacteriocins are effective against a wide range of pathogens [141–143]. For example, lactobacilli produce bacteriocins that interfere with cell wall biosynthesis and induce the formation of pores in S. aureus membranes [144]. To counter this activity, S. aureus produces bacteriocins (staphylococcin Au-26, Bac1829, BacR1, aureocin A53) that inhibit lactobacilli growth [144]. The combination of multiple bacteriocins or the use of bacteriocins with antibiotics demonstrates the highest antibacterial efficacy [145, 146]. Bacteriocins are also used during competition between S. aureus and Corynebacterium spp., microbes associated with skin, nasopharynx, and catheter-related infections. Several bacteriocins with activity against Corynebacterium spp. (Bac 1829, aureocin A70, aureocin A53, and staphylococcin 188) are secreted by S. aureus [147–149]. However, despite their potential as antimicrobial agents, nisin is the only bacteriocin approved for use by regulatory agencies, and even then, its use is limited to food preservation. It would be valuable to investigate the impact of bacteriocins on polymicrobial infections, as they may influence bacterial dynamics during infection and improve pathogen eradication.
Antibiofilm molecules
Because biofilm formation offers considerable advantage to polymicrobial infections, disrupting biofilms will likely profoundly impact infections. There are two strategies for biofilm disruption: preventing their formation or dismantling them after they have formed. Biofilm prevention is considered easier and can be done by changing materials used in medical devices, coating them with anti-adherence compounds, or interfering with signaling pathways that induce biofilms [150]. The most common strategy is creating materials with reduced adhesion properties, including materials with enhanced smoothness, reduced hydrophobicity, and low surface energy. These properties can be achieved by selecting appropriate materials or applying coatings imparting the necessary characteristics. For example, the extremely hydrophilic 2-methacryloyloxyethyl phosphorylcholine polymers can significantly decrease bacterial pathogen attachment and prevent biofilm formation [151]. Moreover, coating with silicone rubber or surfactants reduces interfacial tension and the ability of bacteria, like S. aureus, to form biofilms [152, 153]. Another strategy to disrupt biofilms involves targeting signaling pathways essential for their development. These include QS molecules and nucleotide second messengers that reduce bacterial motility, promote surface attachment (such as pilus formation), or enhance biofilm matrix production (EPS, cellulose, curli fibers). The regulation of biofilm formation via nucleotide second messengers mainly includes (p)ppGpp, c-di-AMP, and c-di-GTP [154]. Interference with these secondary metabolites can be achieved by reducing their synthesis or enhancing their degradation. For example, small molecules that antagonize the activity of diguanylate cyclases or diadenylyl cyclases, which synthesize c-di-GMP and c-di-AMP, respectively, inhibit biofilm formation [155, 156].
Dismantling mature biofilms is considered more challenging but can be accomplished by physical or biochemical methods [157]. Physical methods, such as ultrasound and magnetic field application, are effective for eradicating biofilms on medical devices and patient-contact surfaces but unsuitable for treating infections directly. Biochemical methods, including phage lysins, degradative enzymes, and microbial metabolites, are more appropriate for treating patients. Several enzymes effectively metabolize components of the biofilm and its matrix. These include EPS hydrolytic enzymes, proteases that cleave biofilm-associated proteins, and DNases that degrade extracellular DNA (eDNAs) [158]. A combination of multiple antibiofilm agents showed enhanced effectiveness against biofilms of mixed pathogenic species (L. monocytogenes with E. coli or P. fluorescens) [159]. Despite significant potential, only a few anti-biofilm agents have been evaluated in clinical trials, and their full therapeutic potential has yet to be explored [160].
Although individual strategies show promise in disturbing biofilms, combining antibiofilm and antibacterial drugs may be more effective by targeting both biofilm structure and the cells within. Often, results demonstrate enhanced activity at lower doses, reducing the risk of resistance and side effects [161]. Acting on biofilm EPS can effectively reduce tolerance and the damage caused by the immune responses. A recent study showed that a combination of trypsin and DNAse I, both of which degrade EPS, effectively eradicates dual-species biofilms of P. aeruginosa and S. aureus [162]. In another study, researchers screened a small molecule library for antagonists of PQS, a quinolone-dependent QS system that controls virulence and biofilm production in P. aeruginosa. One antagonist, quinazolinone (QZN), successfully eradicated P. aeruginosa and S. aureus mixed biofilms in combination with tobramycin. These findings are particularly interesting given that P. aeruginosa is known to protect S. aureus from tobramycin-induced killing by promoting the formation of small colony variants [163]. Recently, a combination of phages and gentamicin was shown to effectively kill P. aeruginosa and S. aureus mixed biofilms using a wound model on artificial dermis [164]. Simultaneous and multiple consecutive doses were required to avoid the emergence of resistance to this treatment.
Antimicrobial peptides
Antimicrobial peptides (AMPs) have been described as potential alternatives to treat biofilm-associated polymicrobial infections (reviewed in [139]). AMPs have several advantages compared to conventional antibiotics, displaying broad-spectrum antimicrobial and antibiofilm activity with a shorter incubation time and at similar or lower concentrations than antibiotics. Furthermore, these molecules often exhibit anti-inflammatory properties, speeding healing. In one study, a short synthetic AMP, DRGN-1, inhibited monomicrobial and polymicrobial biofilms of P. aeruginosa and S. aureus [165]. Using a mouse wound model, the peptide was shown to reduce the colonization of both species and also stimulate keratinocyte migration, accelerating wound closure. More recently, a short synthetic peptide, K6, showed bactericidal activity against P. aeruginosa and S. aureus polymicrobial biofilms in a mouse model of persistent infection [166]. In addition, K6 exhibited higher antibiofilm activity than gentamicin, inducing bacterial membrane permeability by forming nanostructured micelles. A study using a mouse model of pneumonia with multi-drug resistant P. aeruginosa and A. baumannii reported that pre-treatment of mice intravenously with the AMP tachyplesin III effectively prolonged survival and reduced bacterial counts in bronchoalveolar lavage fluid, even compared to meropenem-treated mice [167]. An additional synthetic AMP, Nal-P-113, showed bactericidal activity against polymicrobial biofilms of S. gordonii, F. nucleatum, and P. gingivalis at lower concentrations than penicillin or metronidazole [168]. Although promising, the use of AMPs has limitations, including potential for toxicity, susceptibility to protease degradation, low stability at physiological salt concentrations, sequestration by components of body fluids, and high costs.
QS interference
QS is a key bacterial communication strategy that regulates population behaviors, host-microbe interactions, and virulence, prompting the evolution of mechanisms of QS interference as mechanisms of competition [169]. Indeed, the ability of microbes to interfere with QS of competitors, termed quorum quenching, is extensively described [170]. This can occur, for instance, through the degradation of QS signals, typically performed by acylases and lactonases. P. aeruginosa, including human clinical isolates, can grow in 3-oxododecanoyl-homoserine lactone (3OC12HSL), the primary QS signal of this species, as the sole source of carbon and energy [171]. The acylase enzyme involved in this activity, PvdQ, is also required to mature pyoverdine, an important virulence factor of P. aeruginosa. Indeed, inhibition of PvdQ is protective against P. aeruginosa infection in a G. mellonella model [172]. Therefore, QS enzymes have essential roles in P. aeruginosa virulence. Although, to our knowledge, the role of this enzyme in microbial competition has not been directly demonstrated, its ability to interfere with QS could have consequences during co-infections, such as those occurring in the CF lungs. In this complex microbial community, other AHL-producing pathogens, such as Burkholderia cenocepacia, are commonly found and use QS in their pathogenesis [173]. Two acylases of P. aeruginosa, PvdQ and QuiP, allow the degradation of multiple types of AHLs [171, 174]. In addition, AhlD, a lactonase from Arthrobacter sp. with orthologs in several bacteria, including the human pathogen K. pneumoniae, was previously described [175]. Also, lab-engineered variants of the SsoPox lactonase showed anti-virulence activity against multiple human pathogens, including A. baumannii, P. aeruginosa, and B. cepacia. Biofilm formation by these pathogens and toxicity towards macrophages by P. aeruginosa were significantly dampened by lactonase treatment [176]. Besides acylases and lactonases, a third class of enzymes, the oxidoreductases, interfere with AHL signaling. Although oxidoreductases do not effectively degrade AHLs, they chemically modify them by oxidizing or reducing acyl chains. This can alter signal specificity and, therefore, activity. Although this has been investigated mostly in environmental and plant-associated microorganisms, these enzymes were found in Burkholderia, a genus encompassing several human pathogens [177].
Conclusions
Historical focus on single-pathogen infections has underestimated the clinical relevance of polymicrobial infections until recently. It is now evident that bacterial pathogens frequently exist within complex communities, both within the host and in environmental reservoirs. This has driven the evolution of sophisticated interactions, encompassing sensing, communication, competition, and evasion strategies, often contributing to increased infection persistence and treatment resistance. Understanding the intricate balance between antagonistic and synergistic interactions among pathogens can revolutionize therapeutic and preventative strategies. Investigations of pathogen interactions within the context of polymicrobial infections should be prioritized to fully realize this potential. This includes improving microbial isolation procedures for different clinical specimens and applying diagnostic tools to identify co-infections correctly. Once pathogens involved in polymicrobial infections are identified, the mechanisms by which these interactions facilitate pathogen establishment, persistence, and disease severity can be elucidated. Specifically, studies should focus on defining the molecular basis of interspecies communication, as understanding how pathogens signal and respond to each other is crucial to identifying potential targets for intervention. Additionally, characterizing the dynamics of competitive and cooperative interactions will assist in determining the factors that influence the outcome of these interactions, providing insights into the stability and resilience of polymicrobial communities. Studies should also focus on the role of the host environment, as understanding how host factors influence pathogen interactions and contribute to disease pathogenesis is essential to develop therapies. Finally, future studies that explore the impact of polymicrobial interactions on antimicrobial resistance are needed. Assessing the contribution of interspecies interactions to the evolution and dissemination of resistance is critical for combating the growing threat of antibiotic resistance. The development and validation of in vitro and in vivo models are essential to accurately replicate the complex relationships observed in polymicrobial infections. These models should incorporate relevant environmental and host factors to ensure translational relevance. Ultimately, the accumulated knowledge of pathogen interaction strategies will enable the development of targeted interventions. Strategies such as disrupting cooperative mechanisms or exploiting competitive interactions could provide novel avenues for combating polymicrobial infections. This paradigm shift, from targeting single pathogens to modulating interspecies interactions, holds immense promise for improving patient outcomes and addressing the challenges posed by complex infectious diseases.
Acknowledgements
RBRF and LCMA are supported by startup funds provided by the University of Kansas and grant funds provided by the National Institute of General Medical Sciences (NIGMS) of the National Institutes of Health (P20GM113117). NS is supported by the Israel Science Foundation (988/19), and the Israel Ministry of Science and Technology (0007155).
Contributor Information
Rosana Barreto Rocha Ferreira, Department of Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, KS 66045, United States.
Luis Caetano Martha Antunes, Department of Molecular Biosciences, University of Kansas, 1200 Sunnyside Avenue, Lawrence, KS 66045, United States.
Neta Sal-Man, The Shraga Segal Department of Microbiology and Immunology, Ben-Gurion University of the Negev, Beer Sheva, Israel.
Conflict of interest
None to declare
Funding
None declared.
Data availability
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.
References
- 1. Falkow S. Molecular Koch's postulates applied to bacterial pathogenicity—a personal recollection 15 years later. Nat Rev Microbiol. 2004;2:67–72. 10.1038/nrmicro799 [DOI] [PubMed] [Google Scholar]
- 2. Liu J, Platts-Mills JA, Juma J. et al. Use of quantitative molecular diagnostic methods to identify causes of diarrhoea in children: a reanalysis of the GEMS case-control study. Lancet. 2016;388:1291–301. 10.1016/S0140-6736(16)31529-X [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Peng D, Zhao D, Liu J. et al. Multipathogen infections in hospitalized children with acute respiratory infections. Virol J 2009;6:155. 10.1186/1743-422X-6-155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Chowdhury F, Begum YA, Alam MM. et al. Concomitant enterotoxigenic Escherichia coli infection induces increased immune responses to vibrio cholerae O1 antigens in patients with cholera in Bangladesh. Infect Immun 2010;78:2117–24. 10.1128/IAI.01426-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Faruque AS, Mahalanabis D, Islam A. et al. Severity of cholera during concurrent infections with other enteric pathogens. J Diarrhoeal Dis Res 1994;12:214–8. [PubMed] [Google Scholar]
- 6. Paniagua GL, Monroy E, Garcia-Gonzalez O. et al. Two or more enteropathogens are associated with diarrhoea in Mexican children. Ann Clin Microbiol Antimicrob 2007;6:17. 10.1186/1476-0711-6-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Shrivastava AK, Kumar S, Mohakud NK. et al. Multiple etiologies of infectious diarrhea and concurrent infections in a pediatric outpatient-based screening study in Odisha. India Gut Pathog 2017;9:16. 10.1186/s13099-017-0166-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Balmer O, Tanner M. Prevalence and implications of multiple-strain infections. Lancet Infect Dis 2011;11:868–78. 10.1016/S1473-3099(11)70241-9 [DOI] [PubMed] [Google Scholar]
- 9. Roberts FA, Darveau RP. Microbial protection and virulence in periodontal tissue as a function of polymicrobial communities: symbiosis and dysbiosis. Periodontol 2000;69:18–27. 10.1111/prd.12087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Armbruster CE, Hong W, Pang B. et al. Indirect pathogenicity of haemophilus influenzae and Moraxella catarrhalis in polymicrobial otitis media occurs via interspecies quorum signaling. mBio. 2010;1:e00102-10. 10.1128/mBio.00102-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. West SA, Buckling A. Cooperation, virulence and siderophore production in bacterial parasites. Proc Biol Sci 2003;270:37–44. 10.1098/rspb.2002.2209 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Buckling A, Brockhurst MA. Kin selection and the evolution of virulence. Heredity. 2008;100:484–8. 10.1038/sj.hdy.6801093 [DOI] [PubMed] [Google Scholar]
- 13. Gharibi O, Mirzaei K, Karimi A. et al. Mixed infections of vibrio cholerae O1 Ogawa EL tor with Shigella dysenteriae. Pak J Biol Sci 2010;13:1110–2. 10.3923/pjbs.2010.1110.1112 [DOI] [PubMed] [Google Scholar]
- 14. Brogden KA, Guthmiller JM, Taylor CE. Human polymicrobial infections. Lancet. 2005;365:253–5. 10.1016/S0140-6736(05)70155-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Cigana C, Bianconi I, Baldan R. et al. Staphylococcus aureus impacts Pseudomonas aeruginosa chronic respiratory disease in murine models. J Infect Dis 2018;217:933–42. 10.1093/infdis/jix621 [DOI] [PubMed] [Google Scholar]
- 16. Maliniak ML, Stecenko AA, McCarty NA. A longitudinal analysis of chronic MRSA and Pseudomonas aeruginosa co-infection in cystic fibrosis: a single-center study. J Cyst Fibros 2016;15:350–6. 10.1016/j.jcf.2015.10.014 [DOI] [PubMed] [Google Scholar]
- 17. Limoli DH, Yang J, Khansaheb MK. et al. Staphylococcus aureus and Pseudomonas aeruginosa co-infection is associated with cystic fibrosis-related diabetes and poor clinical outcomes. Eur J Clin Microbiol Infect Dis 2016;35:947–53. 10.1007/s10096-016-2621-0 [DOI] [PubMed] [Google Scholar]
- 18. Yang W, Zhou Y, Wu C. et al. Enterohemorrhagic Escherichia coli promotes the invasion and tissue damage of enterocytes infected with Candida albicans in vitro. Sci Rep 2016;6:37485. 10.1038/srep37485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Verghese A, Prabhu K, Diamond RD. et al. Synchronous bacterial and fungal septicemia. A marker for the critically ill surgical patient. Am Surg 1988;54:276–83. [PubMed] [Google Scholar]
- 20. Pulimood S, Ganesan L, Alangaden G. et al. Polymicrobial candidemia. Diagn Microbiol Infect Dis 2002;44:353–7. 10.1016/S0732-8893(02)00460-1 [DOI] [PubMed] [Google Scholar]
- 21. Esposito S, Ascione T, Pagliano P. Management of bacterial skin and skin structure infections with polymicrobial etiology. Expert Rev Anti-Infect Ther 2019;17:17–25. 10.1080/14787210.2019.1552518 [DOI] [PubMed] [Google Scholar]
- 22. Chukkapalli SS, Velsko IM, Rivera-Kweh MF. et al. Polymicrobial oral infection with four periodontal bacteria orchestrates a distinct inflammatory response and atherosclerosis in ApoE null mice. PLoS One 2015;10:e0143291. 10.1371/journal.pone.0143291 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Jean-Pierre F, Vyas A, Hampton TH. et al. One versus many: polymicrobial communities and the cystic fibrosis airway. mBio. 2021;12:e00006-21. 10.1128/mBio.00006-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Ray GT, Suaya JA, Baxter R. Incidence, microbiology, and patient characteristics of skin and soft-tissue infections in a U.S. population: a retrospective population-based study. BMC Infect Dis 2013;13:252. 10.1186/1471-2334-13-252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Sartelli M, Coccolini F, Kluger Y. et al. WSES/GAIS/WSIS/SIS-E/AAST global clinical pathways for patients with skin and soft tissue infections. World J Emerg Surg 2022;17:3. 10.1186/s13017-022-00406-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Shettigar K, Jain S, Bhat DV. et al. Virulence determinants in clinical Staphylococcus aureus from monomicrobial and polymicrobial infections of diabetic foot ulcers. J Med Microbiol 2016;65:1392–404. 10.1099/jmm.0.000370 [DOI] [PubMed] [Google Scholar]
- 27. Esposito S, De Simone G, Pan A. et al. Epidemiology and microbiology of skin and soft tissue infections: preliminary results of a national registry. J Chemother 2019;31:9–14. 10.1080/1120009X.2018.1536320 [DOI] [PubMed] [Google Scholar]
- 28. Nair N, Biswas R, Gotz F. et al. Impact of Staphylococcus aureus on pathogenesis in polymicrobial infections. Infect Immun 2014;82:2162–9. 10.1128/IAI.00059-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Mariani F, Galvan EM. Staphylococcus aureus in polymicrobial skin and soft tissue infections: impact of inter-species interactions in disease outcome. Antibiotics. 2023;12:1164. 10.3390/antibiotics12071164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Sloan TJ, Turton JC, Tyson J. et al. Examining diabetic heel ulcers through an ecological lens: microbial community dynamics associated with healing and infection. J Med Microbiol 2019;68:230–40. 10.1099/jmm.0.000907 [DOI] [PubMed] [Google Scholar]
- 31. Gomes F, Furtado GE, Henriques M. et al. The skin microbiome of infected pressure ulcers: a review and implications for health professionals. Eur J Clin Investig 2022;52:e13688. 10.1111/eci.13688 [DOI] [PubMed] [Google Scholar]
- 32. Fayolle M, Morsli M, Gelis A. et al. The persistence of Staphylococcus aureus in pressure ulcers: a colonising role. Genes. 2021;12:1883. 10.3390/genes12121883 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Pastar I, Nusbaum AG, Gil J. et al. Interactions of methicillin resistant Staphylococcus aureus USA300 and Pseudomonas aeruginosa in polymicrobial wound infection. PLoS One 2013;8:e56846. 10.1371/journal.pone.0056846 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Feuille F, Ebersole JL, Kesavalu L. et al. Mixed infection with Porphyromonas gingivalis and Fusobacterium nucleatum in a murine lesion model: potential synergistic effects on virulence. Infect Immun 1996;64:2094–100. 10.1128/iai.64.6.2094-2100.1996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bakthavatchalu V, Meka A, Mans JJ. et al. Polymicrobial periodontal pathogen transcriptomes in calvarial bone and soft tissue. Mol Oral Microbiol 2011;26:303–20. 10.1111/j.2041-1014.2011.00619.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Gaston JR, Johnson AO, Bair KL. et al. Polymicrobial interactions in the urinary tract: is the enemy of my enemy my friend? Infect Immun 2021;89:10.1128/iai.00652-20. 10.1128/IAI.00652-20 [DOI] [PubMed] [Google Scholar]
- 37. Kline KA, Lewis AL. Gram-positive uropathogens, polymicrobial urinary tract infection, and the emerging microbiota of the urinary tract. Microbiol Spectr. 2016;4:10.1128/microbiolspec.uti-0012-2012. 10.1128/microbiolspec.UTI-0012-2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Armbruster CE, Smith SN, Yep A. et al. Increased incidence of urolithiasis and bacteremia during Proteus mirabilis and Providencia stuartii coinfection due to synergistic induction of urease activity. J Infect Dis 2014;209:1524–32. 10.1093/infdis/jit663 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Nye TM, Zou Z, Obernuefemann CLP. et al. Microbial co-occurrences on catheters from long-term catheterized patients. Nat Commun 2024;15:61. 10.1038/s41467-023-44095-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Bakaletz LO. Viral-bacterial co-infections in the respiratory tract. Curr Opin Microbiol 2017;35:30–5. 10.1016/j.mib.2016.11.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Filkins LM, O'Toole GA. Cystic fibrosis lung infections: polymicrobial, complex, and hard to treat. PLoS Pathog 2015;11:e1005258. 10.1371/journal.ppat.1005258 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Hauser AR, Jain M, Bar-Meir M. et al. Clinical significance of microbial infection and adaptation in cystic fibrosis. Clin Microbiol Rev 2011;24:29–70. 10.1128/CMR.00036-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Nhu NTQ, Young VB. The relationship between the microbiome and antimicrobial resistance. Clin Infect Dis 2023;77:S479–86. 10.1093/cid/ciad641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Colgate ER, Klopfer C, Dickson DM. et al. Network analysis of patterns and relevance of enteric pathogen co-infections among infants in a diarrhea-endemic setting. PLoS Comput Biol 2023;19:e1011624. 10.1371/journal.pcbi.1011624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Black RE, Merson MH, Rahman AS. et al. A two-year study of bacterial, viral, and parasitic agents associated with diarrhea in rural Bangladesh. J Infect Dis 1980;142:660–4. 10.1093/infdis/142.5.660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Freilich S, Zarecki R, Eilam O. et al. Competitive and cooperative metabolic interactions in bacterial communities. Nat Commun 2011;2:589. 10.1038/ncomms1597 [DOI] [PubMed] [Google Scholar]
- 47. Peri AM, Stewart A, Hume A. et al. New microbiological techniques for the diagnosis of bacterial infections and sepsis in ICU including point of care. Curr Infect Dis Rep 2021;23:12. 10.1007/s11908-021-00755-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Cummings LA, Kurosawa K, Hoogestraat DR. et al. Clinical next generation sequencing outperforms standard microbiological culture for characterizing polymicrobial samples. Clin Chem 2016;62:1465–73. 10.1373/clinchem.2016.258806 [DOI] [PubMed] [Google Scholar]
- 49. Cummings LA, Hoogestraat DR, Rassoulian-Barrett SL. et al. Comprehensive evaluation of complex polymicrobial specimens using next generation sequencing and standard microbiological culture. Sci Rep 2020;10:5446. 10.1038/s41598-020-62424-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Zhao Z, Wang Y, Kang Y. et al. A retrospective study of the detection of sepsis pathogens comparing blood culture and culture-independent digital PCR. Heliyon. 2024;10:e27523. 10.1016/j.heliyon.2024.e27523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51. Martin-Loeches I, Pereira JG, Teoh TK. et al. Molecular antimicrobial susceptibility testing in sepsis. Future Microbiol 2024;19:61–72. 10.2217/fmb-2023-0128 [DOI] [PubMed] [Google Scholar]
- 52. Zhang Y, Hu A, Andini N. et al. A 'culture' shift: application of molecular techniques for diagnosing polymicrobial infections. Biotechnol Adv 2019;37:476–90. 10.1016/j.biotechadv.2019.02.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Panwar S, Duggirala KS, Yadav P. et al. Advanced diagnostic methods for identification of bacterial foodborne pathogens: contemporary and upcoming challenges. Crit Rev Biotechnol 2023;43:982–1000. 10.1080/07388551.2022.2095253 [DOI] [PubMed] [Google Scholar]
- 54. Guven E, Azizoglu RO. The recent original perspectives on nonculture-based bacteria detection methods: a comprehensive review. Foodborne Pathog Dis 2022;19:425–40. 10.1089/fpd.2021.0078 [DOI] [PubMed] [Google Scholar]
- 55. Haselbeck AH, Im J, Prifti K. et al. Serology as a tool to assess infectious disease landscapes and guide public health policy. Pathogens. 2022;11:732. 10.3390/pathogens11070732 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Sauer S, Kliem M. Mass spectrometry tools for the classification and identification of bacteria. Nat Rev Microbiol. 2010;8:74–82. 10.1038/nrmicro2243 [DOI] [PubMed] [Google Scholar]
- 57. Riley PA. Principles of microscopy, culture and serology-based diagnostics. Medicine 45:639–44. [Google Scholar]
- 58. Gunasekaran D, Rostovsky I, Taussig D. et al. A dual-channel electrochemical biosensor enables concurrent detection of pathogens and antibiotic resistance. Biosens Bioelectron 2024;257:116314. 10.1016/j.bios.2024.116314 [DOI] [PubMed] [Google Scholar]
- 59. Gehring AG, Albin DM, Reed SA. et al. An antibody microarray, in multiwell plate format, for multiplex screening of foodborne pathogenic bacteria and biomolecules. Anal Bioanal Chem 2008;391:497–506. 10.1007/s00216-008-2044-6 [DOI] [PubMed] [Google Scholar]
- 60. Doron S, Gorbach SL. Bacterial infections: overview. International Encyclopedia of Public Health 2008;273–82. [Google Scholar]
- 61. Garcia-Bayona L, Comstock LE. Bacterial antagonism in host-associated microbial communities. Science. 2018;361:eaat2456. 10.1126/science.aat2456 [DOI] [PubMed] [Google Scholar]
- 62. Hibbing ME, Fuqua C, Parsek MR. et al. Bacterial competition: surviving and thriving in the microbial jungle. Nat Rev Microbiol. 2010;8:15–25. 10.1038/nrmicro2259 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Davies J. Specialized microbial metabolites: functions and origins. J Antibiot 2013;66:361–4. 10.1038/ja.2013.61 [DOI] [PubMed] [Google Scholar]
- 64. Meade E, Slattery MA, Garvey M. Bacteriocins, potent antimicrobial peptides and the fight against multi drug resistant species: resistance is futile? Antibiotics. 2020;9:32. 10.3390/antibiotics9010032 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65. Sugrue I, Ross RP, Hill C. Bacteriocin diversity, function, discovery and application as antimicrobials. Nat Rev Microbiol. 2024;22:556–71. 10.1038/s41579-024-01045-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Arbulu S, Kjos M. Revisiting the multifaceted roles of bacteriocins: the multifaceted roles of bacteriocins. Microb Ecol 2024;87:41. 10.1007/s00248-024-02357-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Cherrak Y, Salazar MA, Yilmaz K. et al. Commensal E. Coli limits salmonella gut invasion during inflammation by producing toxin-bound siderophores in a tonB-dependent manner. PLoS Biol 2024;22:e3002616. 10.1371/journal.pbio.3002616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68. Sassone-Corsi M, Nuccio SP, Liu H. et al. Microcins mediate competition among Enterobacteriaceae in the inflamed gut. Nature. 2016;540:280–3. 10.1038/nature20557 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. De Vuyst L, Leroy F. Bacteriocins from lactic acid bacteria: production, purification, and food applications. J Mol Microbiol Biotechnol 2007;13:194–9. [DOI] [PubMed] [Google Scholar]
- 70. Maphosa S, Moleleki LN, Motaung TE. Bacterial secretion system functions: evidence of interactions and downstream implications. Microbiology. 2023;169:001326. 10.1099/mic.0.001326 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71. Green ER, Mecsas J. Bacterial secretion systems: an overview. Microbiol Spectr 2016;4:10.1128/microbiolspec.VMBF-0012-2015. 10.1128/microbiolspec.VMBF-0012-2015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Cobe BL, Dey S, Minasov G. et al. Bactericidal effectors of the Stenotrophomonas maltophilia type IV secretion system: functional definition of the nuclease TfdA and structural determination of TfcB. mBio. 2024;15:e0119824. 10.1128/mbio.01198-24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Liao J, Li Z, Xiong D. et al. Quorum quenching by a type IVA secretion system effector. ISME J 2023;17:1564–77. 10.1038/s41396-023-01457-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74. Kaushik S, He H, Dalbey RE. Bacterial signal peptides—navigating the journey of proteins. Front Physiol 2022;13:933153. 10.3389/fphys.2022.933153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75. Bernstein HD. Type V secretion in gram-negative bacteria. EcoSal Plus 2019;8:10.1128/ecosalplus.ESP-0031-2018. 10.1128/ecosalplus.esp-0031-2018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76. Ruhe ZC, Low DA, Hayes CS. Bacterial contact-dependent growth inhibition. Trends Microbiol 2013;21:230–7. 10.1016/j.tim.2013.02.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77. Aoki SK, Pamma R, Hernday AD. et al. Contact-dependent inhibition of growth in Escherichia coli. Science. 2005;309:1245–8. 10.1126/science.1115109 [DOI] [PubMed] [Google Scholar]
- 78. Ikryannikova LN, Kurbatov LK, Gorokhovets NV. et al. Contact-dependent growth inhibition in bacteria: do not get too close! Int J Mol Sci 2020;21:7990. 10.3390/ijms21217990 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79. Peterson SB, Bertolli SK, Mougous JD. The central role of interbacterial antagonism in bacterial life. Curr Biol 2020;30:R1203–14. 10.1016/j.cub.2020.06.103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80. Hernandez RE, Gallegos-Monterrosa R, Coulthurst SJ. Type VI secretion system effector proteins: effective weapons for bacterial competitiveness. Cell Microbiol 2020;22:e13241. 10.1111/cmi.13241 [DOI] [PubMed] [Google Scholar]
- 81. Boardman ER, Palmer T, Alcock F. Interbacterial competition mediated by the type VIIb secretion system. Microbiology 2023;169:001420. 10.1099/mic.0.001420 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82. Allsopp LP, Bernal P. Killing in the name of: T6SS structure and effector diversity. Microbiology. 2023;169:001367. 10.1099/mic.0.001367 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83. Ho BT, Dong TG, Mekalanos JJ. A view to a kill: the bacterial type VI secretion system. Cell Host Microbe 2014;15:9–21. 10.1016/j.chom.2013.11.008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84. Bier SB, Toska J, Zhao W. et al. A coordinated attack by a bacterial secretion system and a small molecule drives prey specificity. Commun Biol 2024;7:958. 10.1038/s42003-024-06637-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85. Yu X, Yan Y, Zeng J. et al. T6SS nuclease effectors in pseudomonas syringae act as potent antimicrobials in interbacterial competition. J Bacteriol 2024;206:e0027323. 10.1128/jb.00273-23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86. Anderson MC, Vonaesch P, Saffarian A. et al. Shigella sonnei encodes a functional T6SS used for interbacterial competition and niche occupancy. Cell Host Microbe 2017;21:769–776.e3. 10.1016/j.chom.2017.05.004 [DOI] [PubMed] [Google Scholar]
- 87. Hood RD, Singh P, Hsu F. et al. A type VI secretion system of Pseudomonas aeruginosa targets a toxin to bacteria. Cell Host Microbe 2010;7:25–37. 10.1016/j.chom.2009.12.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88. Unnikrishnan M, Constantinidou C, Palmer T. et al. The enigmatic Esx proteins: looking beyond mycobacteria. Trends Microbiol 2017;25:192–204. 10.1016/j.tim.2016.11.004 [DOI] [PubMed] [Google Scholar]
- 89. Spencer BL, Doran KS. Evolving understanding of the type VII secretion system in gram-positive bacteria. PLoS Pathog 2022;18:e1010680. 10.1371/journal.ppat.1010680 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90. Whitney JC, Peterson SB, Kim J. et al. A broadly distributed toxin family mediates contact-dependent antagonism between gram-positive bacteria. Elife 2017;6:e26938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91. Barnes AJ, Bennett EF, Vezina B. et al. Ethanolamine metabolism through two genetically distinct loci enables Klebsiella pneumoniae to bypass nutritional competition in the gut. PLoS Pathog 2024;20:e1012189. 10.1371/journal.ppat.1012189 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92. Thiennimitr P, Winter SE, Winter MG. et al. Intestinal inflammation allows salmonella to use ethanolamine to compete with the microbiota. Proc Natl Acad Sci USA 2011;108:17480–5. 10.1073/pnas.1107857108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93. Riordan JR, Rommens JM, Kerem B. et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science. 1989;245:1066–73. 10.1126/science.2475911 [DOI] [PubMed] [Google Scholar]
- 94. Liu Y, Gloag ES, Hill PJ. et al. Interbacterial antagonism mediated by a released polysaccharide. J Bacteriol 2022;204:e0007622. 10.1128/jb.00076-22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95. Dunphy LJ, Grimes KL, Wase N. et al. Untargeted metabolomics reveals species-specific metabolite production and shared nutrient consumption by Pseudomonas aeruginosa and Staphylococcus aureus. mSystems. 2021;6:e0048021. 10.1128/mSystems.00480-21 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Filkins LM, Graber JA, Olson DG. et al. Coculture of Staphylococcus aureus with Pseudomonas aeruginosa drives S. Aureus towards fermentative metabolism and reduced viability in a cystic fibrosis model. J Bacteriol 2015;197:2252–64. 10.1128/JB.00059-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97. Al-Naimi FA, Garrett KA, Bockus WW. Competition, facilitation, and niche differentiation in two foliar pathogens. Oecologia. 2005;143:449–57. 10.1007/s00442-004-1814-x [DOI] [PubMed] [Google Scholar]
- 98. Garin G, Pradal C, Fournier C. et al. Modelling interaction dynamics between two foliar pathogens in wheat: a multi-scale approach. Ann Bot 2018;121:927–40. 10.1093/aob/mcx186 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99. Moradali MF, Rehm BHA. Bacterial biopolymers: from pathogenesis to advanced materials. Nat Rev Microbiol. 2020;18:195–210. 10.1038/s41579-019-0313-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100. Conwill A, Kuan AC, Damerla R. et al. Anatomy promotes neutral coexistence of strains in the human skin microbiome. Cell Host Microbe 2022;30:171–182.e7. 10.1016/j.chom.2021.12.007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101. Thoraval L, Varin-Simon J, Ohl X. et al. Cutibacterium acnes and its complex host interaction in prosthetic joint infection: current insights and future directions. Res Microbiol 2025;176:104265. [DOI] [PubMed] [Google Scholar]
- 102. Bru JL, Kasallis SJ, Chang R. et al. The great divide: rhamnolipids mediate separation between P. Aeruginosa and S. Aureus. Front Cell Infect Microbiol 2023;13:1245874. 10.3389/fcimb.2023.1245874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103. Warrell DL, Zarrella TM, Machalek C. et al. Interspecies surfactants serve as public goods enabling surface motility in Pseudomonas aeruginosa. J Bacteriol 2024;206:e0028124. 10.1128/jb.00281-24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104. Toska J, Ho BT, Mekalanos JJ. Exopolysaccharide protects vibrio cholerae from exogenous attacks by the type 6 secretion system. Proc Natl Acad Sci USA 2018;115:7997–8002. 10.1073/pnas.1808469115 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105. Potgieter N, Heine L, Ngandu JPK. et al. High burden of co-infection with multiple enteric pathogens in children suffering with diarrhoea from rural and peri-urban communities in South Africa. Pathogens. 2023;12:315. 10.3390/pathogens12020315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106. Kukoc A, Mihelcic A, Miko I. et al. Clinical and laboratory predictors at ICU admission affecting course of illness and mortality rates in a tertiary COVID-19 center. Heart Lung 2022;53:1–10. 10.1016/j.hrtlng.2022.01.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107. Yoon SM, Lee J, Lee SM. et al. Incidence and clinical outcomes of bacterial superinfections in critically ill patients with COVID-19. Front Med 2023;10:1079721. 10.3389/fmed.2023.1079721 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108. Pilacik N, Kaminska T, Augustynowicz-Kopec E. et al. Etiology of bacterial infections and incidence of comorbidities in patients with tuberculosis, treated in Mazovian treatment Centre of Tuberculosis and Lung Diseases during years 2012-2014. Wiad Lek 2015;68:347–53. [PubMed] [Google Scholar]
- 109. Garcia-Cabrera E, Jimenez-Mejias ME, Gil Navarro MV. et al. Superinfection during treatment of nosocomial infections with tigecycline. Eur J Clin Microbiol Infect Dis 2010;29:867–71. 10.1007/s10096-010-0942-y [DOI] [PubMed] [Google Scholar]
- 110. Zuckerman JN, Rombo L, Fisch A. The true burden and risk of cholera: implications for prevention and control. Lancet Infect Dis 2007;7:521–30. 10.1016/S1473-3099(07)70138-X [DOI] [PubMed] [Google Scholar]
- 111. Croxen MA, Law RJ, Scholz R. et al. Recent advances in understanding enteric pathogenic Escherichia coli. Clin Microbiol Rev 2013;26:822–80. 10.1128/CMR.00022-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112. Gorelik O, Levy N, Shaulov L. et al. Vibrio cholerae autoinducer-1 enhances the virulence of enteropathogenic Escherichia coli. Sci Rep 2019;9:4122. 10.1038/s41598-019-40859-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113. Higgins DA, Pomianek ME, Kraml CM. et al. The major vibrio cholerae autoinducer and its role in virulence factor production. Nature. 2007;450:883–6. 10.1038/nature06284 [DOI] [PubMed] [Google Scholar]
- 114. Ng WL, Perez LJ, Wei Y. et al. Signal production and detection specificity in vibrio CqsA/CqsS quorum-sensing systems. Mol Microbiol 2011;79:1407–17. 10.1111/j.1365-2958.2011.07548.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115. Gorelik O, Rogad A, Holoidovsky L. et al. Indole intercepts the communication between enteropathogenic E. Coli and vibrio cholerae. Gut Microbes 2022;14:2138677. 10.1080/19490976.2022.2138677 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116. Ball LM, Bronstein E, Liechti GW. et al. Neisseria gonorrhoeae drives chlamydia trachomatis into a persistence-like state during in vitro co-infection. Infect Immun 2024;92:e0017923. 10.1128/iai.00179-23 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117. Murray JL, Connell JL, Stacy A. et al. Mechanisms of synergy in polymicrobial infections. J Microbiol 2014;52:188–99. 10.1007/s12275-014-4067-3 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118. Wang S, Mu L, Yu C. et al. Microbial collaborations and conflicts: unraveling interactions in the gut ecosystem. Gut Microbes 2024;16:2296603. 10.1080/19490976.2023.2296603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119. Donlan RM, Costerton JW. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin Microbiol Rev 2002;15:167–93. 10.1128/CMR.15.2.167-193.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120. Lazar V, Oprea E, Ditu LM. Resistance, tolerance, virulence and bacterial pathogen fitness—current state and envisioned solutions for the near future. Pathogens. 2023;12:746. 10.3390/pathogens12050746 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121. Anju VT, Busi S, Imchen M. et al. Polymicrobial infections and biofilms: clinical significance and eradication strategies. Antibiotics. 2022;11:1731. 10.3390/antibiotics11121731 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122. Metcalf DG, Bowler PG. Biofilm delays wound healing: a review of the evidence. Burns Trauma 2013;1:5–12. 10.4103/2321-3868.113329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 123. Bessa LJ, Fazii P, Di Giulio M. et al. Bacterial isolates from infected wounds and their antibiotic susceptibility pattern: some remarks about wound infection. Int Wound J 2015;12:47–52. 10.1111/iwj.12049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 124. Korgaonkar A, Trivedi U, Rumbaugh KP. et al. Community surveillance enhances Pseudomonas aeruginosa virulence during polymicrobial infection. Proc Natl Acad Sci USA 2013;110:1059–64. 10.1073/pnas.1214550110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 125. Peters BM, Jabra-Rizk MA, O'May GA. et al. Polymicrobial interactions: impact on pathogenesis and human disease. Clin Microbiol Rev 2012;25:193–213. 10.1128/CMR.00013-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 126. DeLeon S, Clinton A, Fowler H. et al. Synergistic interactions of Pseudomonas aeruginosa and Staphylococcus aureus in an in vitro wound model. Infect Immun 2014;82:4718–28. 10.1128/IAI.02198-14 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 127. Beaudoin T, Yau YCW, Stapleton PJ. et al. Staphylococcus aureus interaction with Pseudomonas aeruginosa biofilm enhances tobramycin resistance. NPJ Biofilms Microbiomes. 2017;3:25. 10.1038/s41522-017-0035-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 128. Orazi G, O'Toole GA. Pseudomonas aeruginosa alters Staphylococcus aureus sensitivity to vancomycin in a biofilm model of cystic fibrosis infection. mBio. 2017;8:e00873-17. 10.1128/mBio.00873-17 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 129. Connell JL, Ritschdorff ET, Whiteley M. et al. 3D printing of microscopic bacterial communities. Proc Natl Acad Sci USA 2013;110:18380–5. 10.1073/pnas.1309729110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 130. Stacy A, McNally L, Darch SE. et al. The biogeography of polymicrobial infection. Nat Rev Microbiol 2016;14:93–105. 10.1038/nrmicro.2015.8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 131. Bragonzi A, Farulla I, Paroni M. et al. Modelling co-infection of the cystic fibrosis lung by Pseudomonas aeruginosa and Burkholderia cenocepacia reveals influences on biofilm formation and host response. PLoS One 2012;7:e52330. 10.1371/journal.pone.0052330 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 132. Huse HK, Kwon T, Zlosnik JE. et al. Pseudomonas aeruginosa enhances production of a non-alginate exopolysaccharide during long-term colonization of the cystic fibrosis lung. PLoS One 2013;8:e82621. 10.1371/journal.pone.0082621 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 133. Zhu Y, Dashper SG, Chen YY. et al. Porphyromonas gingivalis and Treponema denticola synergistic polymicrobial biofilm development. PLoS One 2013;8:e71727. 10.1371/journal.pone.0071727 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 134. Duan K, Dammel C, Stein J. et al. Modulation of Pseudomonas aeruginosa gene expression by host microflora through interspecies communication. Mol Microbiol 2003;50:1477–91. 10.1046/j.1365-2958.2003.03803.x [DOI] [PubMed] [Google Scholar]
- 135. Keogh D, Tay WH, Ho YY. et al. Enterococcal metabolite cues facilitate interspecies niche modulation and polymicrobial infection. Cell Host Microbe 2016;20:493–503. 10.1016/j.chom.2016.09.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 136. Ramsey MM, Rumbaugh KP, Whiteley M. Metabolite cross-feeding enhances virulence in a model polymicrobial infection. PLoS Pathog 2011;7:e1002012. 10.1371/journal.ppat.1002012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 137. Ramsey MM, Whiteley M. Polymicrobial interactions stimulate resistance to host innate immunity through metabolite perception. Proc Natl Acad Sci USA 2009;106:1578–83. 10.1073/pnas.0809533106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 138. Ryan RP, Fouhy Y, Garcia BF. et al. Interspecies signalling via the Stenotrophomonas maltophilia diffusible signal factor influences biofilm formation and polymyxin tolerance in Pseudomonas aeruginosa. Mol Microbiol 2008;68:75–86. 10.1111/j.1365-2958.2008.06132.x [DOI] [PubMed] [Google Scholar]
- 139. Batoni G, Maisetta G, Esin S. Therapeutic potential of antimicrobial peptides in polymicrobial biofilm-associated infections. Int J Mol Sci 2021;22:482. 10.3390/ijms22020482 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 140. Schmitz DA, Allen RC, Kummerli R. Negative interactions and virulence differences drive the dynamics in multispecies bacterial infections. Proc Biol Sci 2003;2023:20231119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 141. Hanchi H, Hammami R, Fernandez B. et al. Simultaneous production of formylated and nonformylated enterocins L50A and L50B as well as 61A, a new glycosylated durancin, by enterococcus durans 61A, a strain isolated from artisanal fermented milk in Tunisia. J Agric Food Chem 2016;64:3584–90. 10.1021/acs.jafc.6b00700 [DOI] [PubMed] [Google Scholar]
- 142. Oman TJ, van der Donk WA. Insights into the mode of action of the two-peptide lantibiotic haloduracin. ACS Chem Biol 2009;4:865–74. 10.1021/cb900194x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 143. Ovchinnikov KV, Chi H, Mehmeti I. et al. Novel group of leaderless multipeptide bacteriocins from gram-positive bacteria. Appl Environ Microbiol 2016;82:5216–24. 10.1128/AEM.01094-16 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 144. Charlier C, Cretenet M, Even S. et al. Interactions between Staphylococcus aureus and lactic acid bacteria: an old story with new perspectives. Int J Food Microbiol 2009;131:30–9. 10.1016/j.ijfoodmicro.2008.06.032 [DOI] [PubMed] [Google Scholar]
- 145. Ovchinnikov KV, Kranjec C, Thorstensen T. et al. Successful development of bacteriocins into therapeutic formulation for treatment of MRSA skin infection in a murine model. Antimicrob Agents Chemother 2020;64:e00829-20. 10.1128/AAC.00829-20 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 146. Hanchi H, Hammami R, Gingras H. et al. Inhibition of MRSA and of Clostridium difficile by durancin 61A: synergy with bacteriocins and antibiotics. Future Microbiol 2017;12:205–12. 10.2217/fmb-2016-0113 [DOI] [PubMed] [Google Scholar]
- 147. Saeed S, Ahmad S, Rasool SA. Antimicrobial spectrum, production and mode of action of staphylococcin 188 produced by Staphylococcus aureus 188. Pak J Pharm Sci 2004;17:1–8. [PubMed] [Google Scholar]
- 148. dos Santos NJ, dos Santos KR. et al. Phenotypic and genetic characterisation of bacteriocin-producing strains of Staphylococcus aureus involved in bovine mastitis. Vet Microbiol 2002;85:133–44. 10.1016/S0378-1135(01)00476-X [DOI] [PubMed] [Google Scholar]
- 149. Crupper SS, Iandolo JJ. Purification and partial characterization of a novel antibacterial agent (Bac1829) produced by Staphylococcus aureus KSI1829. Appl Environ Microbiol 1996;62:3171–5. 10.1128/aem.62.9.3171-3175.1996 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 150. Yin W, Xu S, Wang Y. et al. Ways to control harmful biofilms: prevention, inhibition, and eradication. Crit Rev Microbiol 2021;47:57–78. 10.1080/1040841X.2020.1842325 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 151. Hirota K, Yumoto H, Miyamoto K. et al. MPC-polymer reduces adherence and biofilm formation by oral bacteria. J Dent Res 2011;90:900–5. 10.1177/0022034511402996 [DOI] [PubMed] [Google Scholar]
- 152. Li XH, Lee JH. Antibiofilm agents: a new perspective for antimicrobial strategy. J Microbiol 2017;55:753–66. 10.1007/s12275-017-7274-x [DOI] [PubMed] [Google Scholar]
- 153. Xu Y, Jones JE, Yu H. et al. Nanoscale plasma coating inhibits formation of Staphylococcus aureus biofilm. Antimicrob Agents Chemother 2015;59:7308–15. 10.1128/AAC.01944-15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 154. Hengge R, Pruteanu M, Stulke J. et al. Recent advances and perspectives in nucleotide second messenger signaling in bacteria. Microlife. 2023;4:uqad015. 10.1093/femsml/uqad015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 155. Cho KH, Tryon RG, Kim JH. Screening for diguanylate cyclase (DGC) inhibitors mitigating bacterial biofilm formation. Front Chem 2020;8:264. 10.3389/fchem.2020.00264 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 156. Opoku-Temeng C, Sintim HO. Inhibition of cyclic diadenylate cyclase, DisA, by polyphenols. Sci Rep 2016;6:25445. 10.1038/srep25445 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 157. Shrestha L, Fan HM, Tao HR. et al. Recent strategies to combat biofilms using antimicrobial agents and therapeutic approaches. Pathogens. 2022;11:292. 10.3390/pathogens11030292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 158. Kalia VC, Patel SKS, Lee JK. Bacterial biofilm inhibitors: an overview. Ecotoxicol Environ Saf 2023;264:115389. 10.1016/j.ecoenv.2023.115389 [DOI] [PubMed] [Google Scholar]
- 159. Rodriguez-Lopez P, Carballo-Justo A, Draper LA. et al. Removal of listeria monocytogenes dual-species biofilms using combined enzyme-benzalkonium chloride treatments. Biofouling. 2017;33:45–58. 10.1080/08927014.2016.1261847 [DOI] [PubMed] [Google Scholar]
- 160. Lu L, Li M, Yi G. et al. Screening strategies for quorum sensing inhibitors in combating bacterial infections. J Pharm Anal 2022;12:1–14. 10.1016/j.jpha.2021.03.009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 161. Hawas S, Verderosa AD, Totsika M. Combination therapies for biofilm inhibition and eradication: a comparative review of laboratory and preclinical studies. Front Cell Infect Microbiol 2022;12:850030. 10.3389/fcimb.2022.850030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 162. Fanaei Pirlar R, Emaneini M, Beigverdi R. et al. Combinatorial effects of antibiotics and enzymes against dual-species Staphylococcus aureus and Pseudomonas aeruginosa biofilms in the wound-like medium. PLoS One 2020;15:e0235093. 10.1371/journal.pone.0235093 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 163. Orazi G, Ruoff KL, O'Toole GA. Pseudomonas aeruginosa increases the sensitivity of biofilm-grown Staphylococcus aureus to membrane-targeting antiseptics and antibiotics. mBio. 2019;10:e01501-19. 10.1128/mBio.01501-19 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 164. Akturk E, Melo LDR, Oliveira H. et al. Combining phages and antibiotic to enhance antibiofilm efficacy against an in vitro dual species wound biofilm. Biofilm. 2023;6:100147. 10.1016/j.bioflm.2023.100147 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 165. Chung EMC, Dean SN, Propst CN. et al. Komodo dragon-inspired synthetic peptide DRGN-1 promotes wound-healing of a mixed-biofilm infected wound. NPJ Biofilms Microbiomes 2017;3:9. 10.1038/s41522-017-0017-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 166. Chou S, Guo H, Zingl FG. et al. Synthetic peptides that form nanostructured micelles have potent antibiotic and antibiofilm activity against polymicrobial infections. Proc Natl Acad Sci USA 2023;120:e2219679120. 10.1073/pnas.2219679120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 167. Qi J, Gao R, Liu C. et al. Potential role of the antimicrobial peptide Tachyplesin III against multidrug-resistant P. Aeruginosa and A. Baumannii coinfection in an animal model. Infect Drug Resist 2019;Volume 12:2865–74. 10.2147/IDR.S217020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 168. Wang HY, Cheng JW, Yu HY. et al. Efficacy of a novel antimicrobial peptide against periodontal pathogens in both planktonic and polymicrobial biofilm states. Acta Biomater 2015;25:150–61. 10.1016/j.actbio.2015.07.031 [DOI] [PubMed] [Google Scholar]
- 169. Antunes LCM, Ferreira RBR, Buckner MMC. et al. Quorum sensing in bacterial virulence. Microbiology. 2010;156:2271–82. 10.1099/mic.0.038794-0 [DOI] [PubMed] [Google Scholar]
- 170. Sikdar R, Elias M. Quorum quenching enzymes and their effects on virulence, biofilm, and microbiomes: a review of recent advances. Expert Rev Anti-Infect Ther 2020;18:1221–33. 10.1080/14787210.2020.1794815 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 171. Huang JJ, Han JI, Zhang LH. et al. Utilization of acyl-homoserine lactone quorum signals for growth by a soil pseudomonad and Pseudomonas aeruginosa PAO1. Appl Environ Microbiol 2003;69:5941–9. 10.1128/AEM.69.10.5941-5949.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 172. Vogel JGT, Wibowo JP, Fan H. et al. Discovery of chromene compounds as inhibitors of PvdQ acylase of Pseudomonas aeruginosa. Microbes Infect 2022;24:105017. 10.1016/j.micinf.2022.105017 [DOI] [PubMed] [Google Scholar]
- 173. McKeon SA, Nguyen DT, Viteri DF. et al. Functional quorum sensing systems are maintained during chronic Burkholderia cepacia complex infections in patients with cystic fibrosis. J Infect Dis 2011;203:383–92. 10.1093/infdis/jiq054 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 174. Huang JJ, Petersen A, Whiteley M. et al. Identification of QuiP, the product of gene PA1032, as the second acyl-homoserine lactone acylase of Pseudomonas aeruginosa PAO1. Appl Environ Microbiol 2006;72:1190–7. 10.1128/AEM.72.2.1190-1197.2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 175. Park SY, Lee SJ, Oh TK. et al. AhlD, an N-acylhomoserine lactonase in Arthrobacter sp., and predicted homologues in other bacteria. Microbiology. 2003;149:1541–50. 10.1099/mic.0.26269-0 [DOI] [PubMed] [Google Scholar]
- 176. Gonzales M, Kergaravat B, Jacquet P. et al. Disrupting quorum sensing as a strategy to inhibit bacterial virulence in human, animal, and plant pathogens. Pathog Dis 2024;82:ftae009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 177. Chan KG, Atkinson S, Mathee K. et al. Characterization of N-acylhomoserine lactone-degrading bacteria associated with the Zingiber officinale (ginger) rhizosphere: co-existence of quorum quenching and quorum sensing in Acinetobacter and Burkholderia. BMC Microbiol 2011;11:51. 10.1186/1471-2180-11-51 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
Data sharing is not applicable to this article as no datasets were generated or analyzed during the current study.


