Summary
Neonatal acute respiratory distress syndrome (NARDS) is challenging to differentiate from other respiratory conditions, and gestational age (GA) may influence gene expression. This study characterized whole blood transcriptomic profiles of NARDS in a pilot cohort of 48 neonates with respiratory distress, demonstrating a significant GA-dependent modulation of gene expression. Functional analyses revealed prominent involvement of interferon-related pathways in NARDS, with greater suppression in neonates born before 34 weeks. Immune cell infiltration was observed in term or late preterm neonates but was absent in more preterm cases. Machine learning identified three key predictive genes, among which ALOX15 and PTGDR2 were validated in an independent cohort, with area under the curve ranging from 0.68 to 0.83 across different GAs. The gene changes were also confirmed in a neonatal lipopolysaccharide-induced lung injury mouse model. These findings highlight the potential predictive and therapeutic value of ALOX15 and PTGDR2 for NARDS.
Subject areas: Transcriptomics, Machine learning
Graphical abstract
Highlights
-
•
Gestational age significantly influences neonatal peripheral blood gene expression
-
•
Whole blood transcriptomics reveal that interferon pathways are central to NARDS
-
•
Neonates born before 34 weeks exhibit weaker immune responses to ARDS
-
•
Blood ALOX15 and PTGDR2 are potential biomarkers and therapeutic targets for NARDS
Transcriptomics; Machine learning
Introduction
Acute respiratory distress syndrome (ARDS) is a life-threatening condition caused by various etiologies, and was first described in 1967.1 Since then, diagnostic criteria of ARDS have been established for adults2 and children,3 respectively; however, these criteria do not apply to neonates. Neonatal ARDS (NARDS) remained poorly understood for decades until 2017, when the Montreux Criteria4 were specifically proposed to address the unique perinatal factors associated with neonates. Recent international multicenter researches5,6 on NARDS have begun to uncover its basic clinical characteristics. NARDS occurs in 1.44%–1.5% of admissions, with mortality rates ranging from 12.6% to 24%. Disparities in the immaturity of pulmonary and immune system and different perinatal triggering factors may contribute to the heterogeneity of NARDS.6,7 Early identification remains challenging due to the distinct physiological characteristics of neonatal lungs and immune systems. Respiratory distress, a common yet nonspecific manifestation, affecting 7% of deliveries.8 In addition to ARDS, which can occur in neonates born at every gestational age (GA), common causes of respiratory distress primarily include neonatal respiratory distress syndrome (NRDS) due to primary surfactant (PS) deficiency in preterm infants, and transient tachypnea of the neonate (TTN) in later period. The Montreux definition explicitly stated that the diagnosis of NARDS requires the exclusion of NRDS, TTN, or other primary acute respiratory conditions, while suggesting that a response to PS therapy and lung recruitment should be considered in confirming the diagnosis.4 No specific treatments are currently available for NARDS, while patients typically require longer courses of antibiotic therapy, mechanical ventilation and higher mortality compared to those with NRDS or TTN.5,9
Currently, blood transcriptomics has provided valuable insights into the heterogeneity of both adult and pediatric ARDS, facilitating the identification of novel biomarkers and construction prediction model,10 classifying disease sub phenotypes,11,12 and uncovering the underlying pathophysiological processes13 that may lead to more effective therapies.14 However, this field remains blank in NARDS. Our study seeks to bridge this gap by establishing a cohort of respiratory distress across various GA. We will integrate high-throughput sequencing and machine learning techniques to identify novel predictive biomarkers and offer new perspectives and references for the treatment of NARDS.
Results
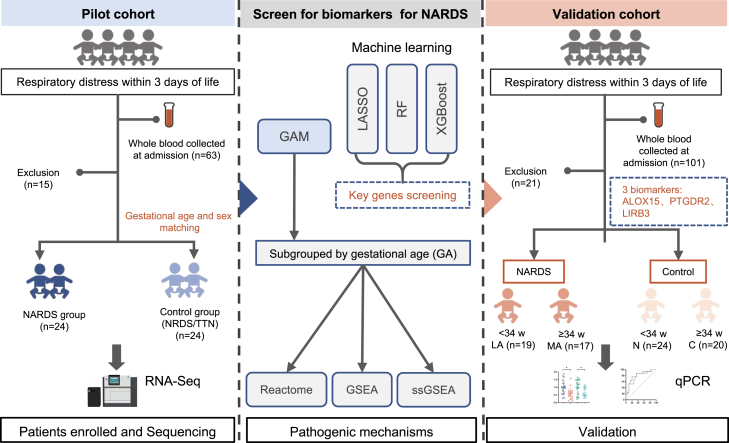
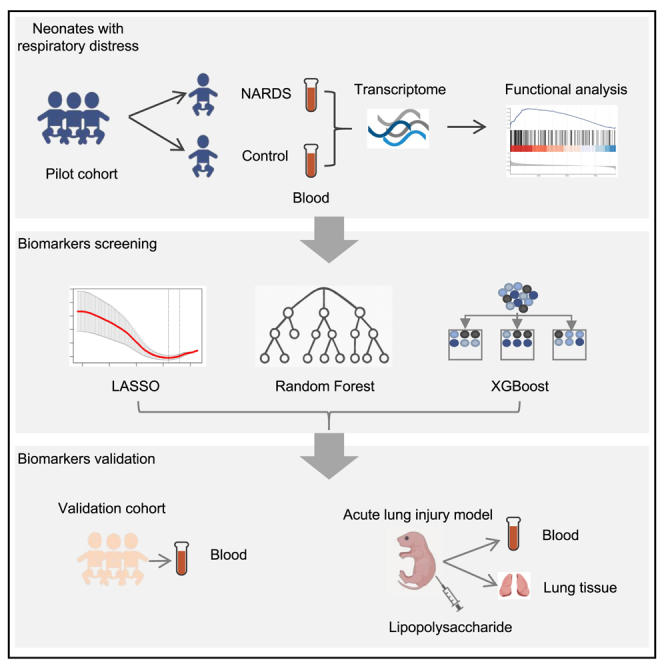
To investigate the blood transcriptomic signatures of NARDS, we prospectively established a clinical cohort of neonates presenting with respiratory distress as the primary manifestation at admission. The pilot cohort comprised 24 NARDS cases [12 in group MA (GA more than 34 weeks), 12 in group LA (GA less than 34 weeks)] and 24 GA/sex-matched controls [12 in group C (GA more than 34 weeks), 12 in group N (GA less than 34 weeks)] selected from an initial pool of 63 neonates, with 15 exclusions based on: congenital anomalies or hereditary metabolic diseases (n = 3), declined to participate (n = 7), poor quality of isolated RNA (n = 5). The group N were all diagnosed with NRDS, while the group C consisted of individuals with NRDS or TTN. For validation, a subsequent cohort was constructed using identical inclusion criteria, ultimately enrolling 80 neonates after excluding 21 ineligible candidates. The validation cohort distribution included 17 MA cases, 19 LA cases, 24 group N controls, and 20 group C controls. Figure 1 illustrates the clinical study workflow, detailing cohort establishment, stratification logic, and analytical pipelines.
Figure 1.
The schematic overview of this study
Demographics and clinical characteristics
Table 1 summarizes the demographic and clinical characteristics of the NARDS and control groups in the pilot cohort, and Table S3 provides details for the validation cohort. The NARDS groups and control groups had no significant differences in birth weight, cesarean delivery, maternal gestational diabetes, antenatal steroid treatment, premature rupture of membranes, or white blood counts. Among the NARDS patients enrolled, about 50%–58.3% were diagnosed with early-onset sepsis (EOS), 83.3%–100% required invasive respiratory support, with a prolonged duration of ventilation (p < 0.05). In the group of neonates born before 34 weeks of GA (group N and LA), NARDS patients had significantly lower 5-min Apgar scores, a higher frequency of receiving PS treatment, and approximately 41.7% of these infants succumbed to the severity of their condition. While in neonates born after 34 weeks, five NARDS patients received surfactant treatment, with three requiring more than two doses. One (8.3%) of these infants did not survive.
Table 1.
Demographic and clinical characteristics of the pilot cohort
| N (n = 12) | LA (n = 12) | P1 | C (n = 12) | MA (n = 12) | P2 | |
|---|---|---|---|---|---|---|
| Gestational age (weeks) | 30.5 ± 2.0 | 28.9 ± 1.9 | 0.061 | 38.0 ± 2.0 | 37.3 ± 0.8 | 0.288 |
| Sex (male, %) | 8(66.7) | 8(66.7) | 1.0 | 9(75.0) | 8(66.7) | 1.0 |
| Birth weight (g) | 1391.8 ± 281.4 | 1310.0(800.0,1383.8) | 0.183 | 3165.0 ± 572.3 | 3012.5 ± 341.5 | 0.436 |
| Cesarean (%) | 9(75.0) | 7(58.3) | 0.667 | 6(50.0) | 8(66.7) | 0.680 |
| Gestational diabetes (%) | 5(41.7) | 2(16.7) | 0.37 | 3(25) | 2(16.7) | 1.0 |
| Antenatal corticoids (%) | 11(91.7) | 8(66.7) | 0.32 | 3(25) | 1(8.3) | 0.590 |
| PROM (%) | 6(50.0) | 4(33.3) | 0.68 | 1(8.3) | 2(16.7) | 0.530 |
| Apgar 5 min | 10(8,10) | 7.6 ± 2.0 | 0.011 | 10(10,10) | 10(8.3,10) | 0.435 |
| White blood count (×109/L) | 11.3 ± 4.3 | 10.0(7.3,14.4) | 0.488 | 16.9 ± 5.5 | 15.0 ± 6.9 | 0.454 |
| Procalcitonin (ng/mL) | 6.2 ± 6.0 | 8.1 ± 7.9 | 0.538 | 0.4(0.2,0.6) | 4.6(0.6,12.2) | 0.012 |
| Early onset of sepsis (%) | 0(0) | 7(58.3) | 0.005 | 0(0) | 6(50.0) | 0.014 |
| Pneumonia (%) | 0(0) | 2(16.7) | 0.5 | 0(0) | 3(25) | 0.093 |
| Pulmonary hemorrhage (%) | 0(0) | 2(16.7) | 0.5 | 0(0) | 4(33.3) | 0.093 |
| PPHN (%) | 6(50.0) | 10(83.3) | 0.193 | 1(8.3) | 7(58.3) | 0.030 |
| Surfactant (%) | 11(91.7) | 11(91.7) | 1.0 | 0(0) | 5(41.7) | 0.037 |
| Surfactant ≥2 times (%) | 0(0) | 5(41.7) | 0.037 | 0(0) | 3(25) | 0.093 |
| Invasive ventilation (%) | 3(25.0) | 12(100) | 0.000 | 0(0) | 10(83.3) | 0.000 |
| Duration of ventilation (day) | 7.50(6.25,18.8) | 21.5(8.0,34.5) | 0.040 | 1.83 ± 1.4 | 11.50(7.25,14.75) | 0.000 |
| Death (%) | 0(0) | 5(41.7) | 0.037 | 0(0) | 1(8.3) | 1.0 |
Data are presented as mean ± standard deviation or median [interquartile range (IQR)] according to normality by Shapiro-Wilk test; P1 indicates significance level for N vs. LA, P2 for C vs. MA; “n” represents the number of subjects.
LA: subjects with neonatal acute respiratory distress syndrome (NARDS) and gestational age <34 weeks; N (Control for LA): Subjects without NARDS and gestational age <34weeks; MA: Subjects with NARDS and gestational age≥34 weeks; C (Control for MA): Subjects without NARDS and gestational age≥34 weeks.
PROM, Premature rupture of membranes; PPHN, persistent pulmonary hypertension of newborn.
Evaluation of gene expression variance caused by GA and NARDS signature genes screening
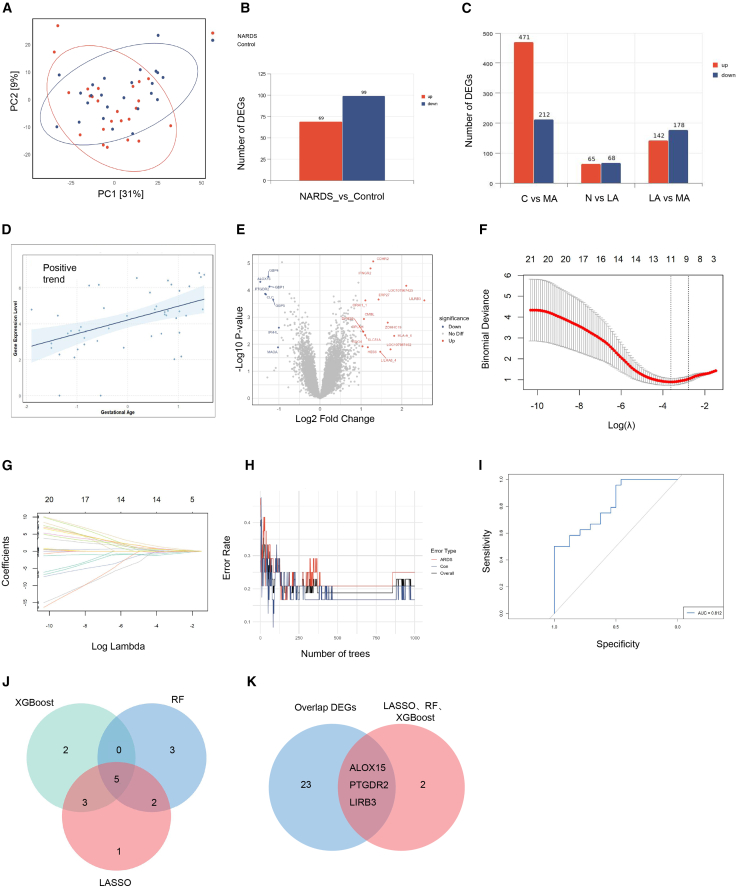
A total of 22,323 genes were identified by RNA sequencing of peripheral blood samples from patients. Principal component analysis (PCA) revealed that the first principal component (PC1) accounted for 31% of the total variance for the differentiation of NARDS and control group (Figure 2A). Notably, a tighter clustering was observed among respiratory distress patients across different GAs (Figure S1A). Further analysis focusing on GA demonstrated considerable heterogeneity in gene expression among different subgroups. Specifically, gene expression displayed more dispersion between subgroups with larger GA (MA and C), while they were less significant between those with smaller GA (LA and N) (Figures S1B and S1C). The primary number of DEGs between NARDS and control was 168. (Figure 2B) While it increased to 837 DEGs between group MA and group C, decreasing to 133 between group LA and group N, and 320 DEGs were found between LA and MA. (Figure 2C) Collectively, both GA and the disease status accounted for the heterogeneity of gene expression.
Figure 2.
Evaluation of gene expression variance caused by GA and NARDS signature genes screening
(A) Principal component analysis (PCA) of variance between NARDS and control. The ellipses represent 95% confidence intervals for each cluster. the first principal component (PC1) explained 31% of the total variance, while the second principal component (PC2) accounted for an additional 9% of the variance.
(B) Number of differentially expresses genes (DEGs) (p < 0.05, |log Foldchange| > 1) in NARDS and control.
(C) Number of DEGs (p < 0.05, |log Foldchange| > 1) in C-MA, N-LA, LA-MA comparisons.
(D) Gene expression of whole blood in relation to gestational age.
(E) Volcano plots of DEGs in NARDS and control after adjustment by gestational age.
(F) LASSO coefficient computation. The vertical dashed line shows the ideal lambda value and 11 was selected in this study.
(G) 10-fold cross-validation for LASSO model parameter adjustment. Each curve represents a gene.
(H) The correlation between the erro rate and the number of trees in the random forest, the error rate stabilized when the number of trees reached 100 to 200.
(I) The area under the curve (AUC) of the XGBoost model was 0.812, indicating a relative strong classification performance.
(J) A Venn diagram illustration of the intersection of genes selected by LASSO, random forest, and XGBoost algorithms.
(K) A Venn diagram illustration of the intersection of genes of three machine algorithms and overlapped DEGs in C-MA and N-LA. NARDS: neonatal acute respiratory distress syndrome.
We therefore included GA as a covariate in a GAM to assess and adjust its influence on gene expression between the NARDS and Control groups. Residual plot and Quantile-Quantile (Q-Q) plot demonstrated that the model had good applicability and robustness (Figure S2). The model revealed that GA positively influenced the expression of neonatal peripheral blood genes (Figure 2D) and a total of 24 DEGs were identified between NARDS and control after adjustment (p < 0.05, |log2FoldChange| > 1) (Figure 2E). Further investigation using three machine learning algorithms —LASSO (Figures 2F and 2G), RF (Figure 2H), and XGBoost (Figure 2I)— identified five key feature genes for NARDS (Figure 2J; Table S4). Of them, ALOX15, PTGDR2, and LILRB3 were significantly different in both MA-C and LA-N group (p < 0.05, |log2FoldChange| > 1, Figure 2K). Especially, the expression of ALOX15 and PTGDR2 showed a positive correlation with GA (Figure S3).
Functional enrichment analysis of genes expressed in subgroup of NARDS and control
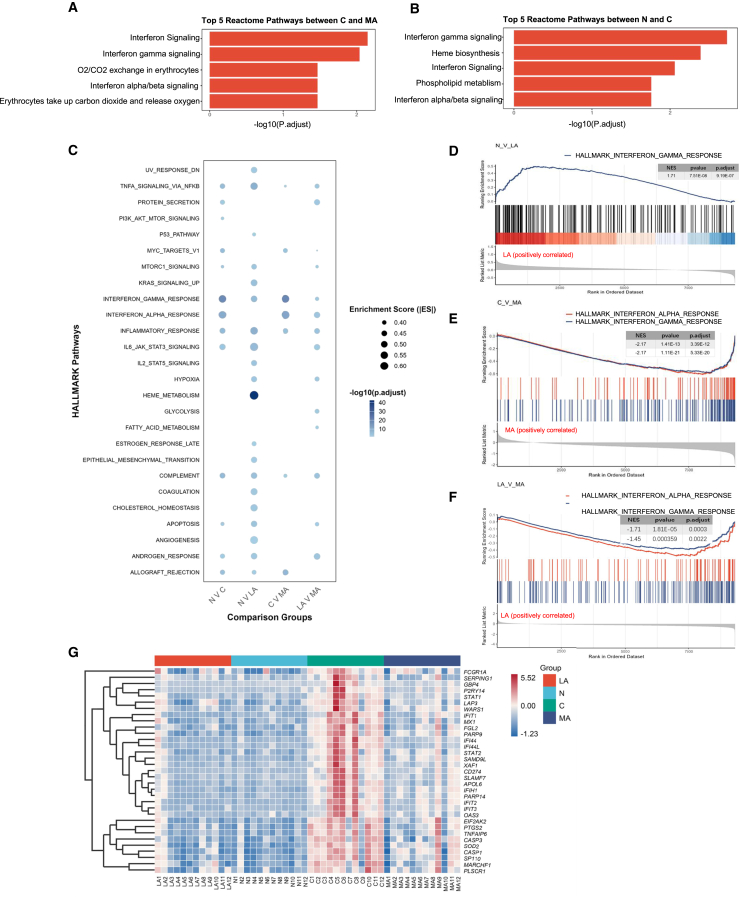
To investigate the potential biological mechanisms of NARDS, we initially performed Reactome pathway enrichment analysis on DEGs across various comparison groups. The results showed a significant enrichment in interferon(IFN)-related signaling pathways, O2/CO2 exchange in erythrocytes and erythrocytes take up carbon dioxide and release oxygen in the MA - C comparison, GPCR ligand binding and nucleotide-like (purinergic) receptors were enriched in the N-LA comparison, while none were enriched in the LA-MA comparison (Figure 3A, p. adjust<0.05). When comparing group N to group C, interferon-related pathways were also enriched, along with heme biosynthesis and phospholipid metabolism. Subsequently, we conducted GSEA against Hallmark Gene Sets, with significantly enriched pathways across the groups summarized in Figure 3C, Table S5. Multiple immune pathways, including TNFα signaling via NF-κB, complement, inflammatory response, and interferon response pathways were consistently enriched across various group comparisons. Notably, the interferon gamma response pathway appeared relatively higher activity in the group LA compared with group N (Figure 3D). The interferon alpha/gamma response in the group MA was suppressed compared with group C, while activated compared with group LA (Figures 3E and 3F). Figure 3G showed a heatmap of the expression levels of interferon-related genes across the groups.
Figure 3.
Functional enrichment analysis of genes expressed in subgroup of NARDS and control
(A) Results of Reactome pathway analysis of DEGs in group C and MA presented in a barplot.
(B) Results of Reactome pathway analysis of DEGs in group N and C presented in a barplot.
(C) Results of GSEA analysis of ranked gene lists of N-C, N-LA, C-MA, LA-MA comparison groups in a bubble plot, with bubble size representing the absolute value of enrichment score (ES) (larger circles indicate larger ES) and color representing the adjusted p-value (darker blue shades indicate smaller p-values).
(D–F) Significant enrichment in interferon-related pathways of GSEA analysis. (D) Interferon-gamma response was activated in group LA compared with group N. (F) Interferon-alpha/gamma response was suppressed in MA compared with group C, while (F) activated when compared with group LA.
(G) Expression pattern of interferon-related genes in different subgroups in a heatmap. NARDS: neonatal acute respiratory distress syndrome, GSEA: gene set enrichment analysis, DEG: differentially expresses genes.
Immune cell infiltration analysis and correlation analysis
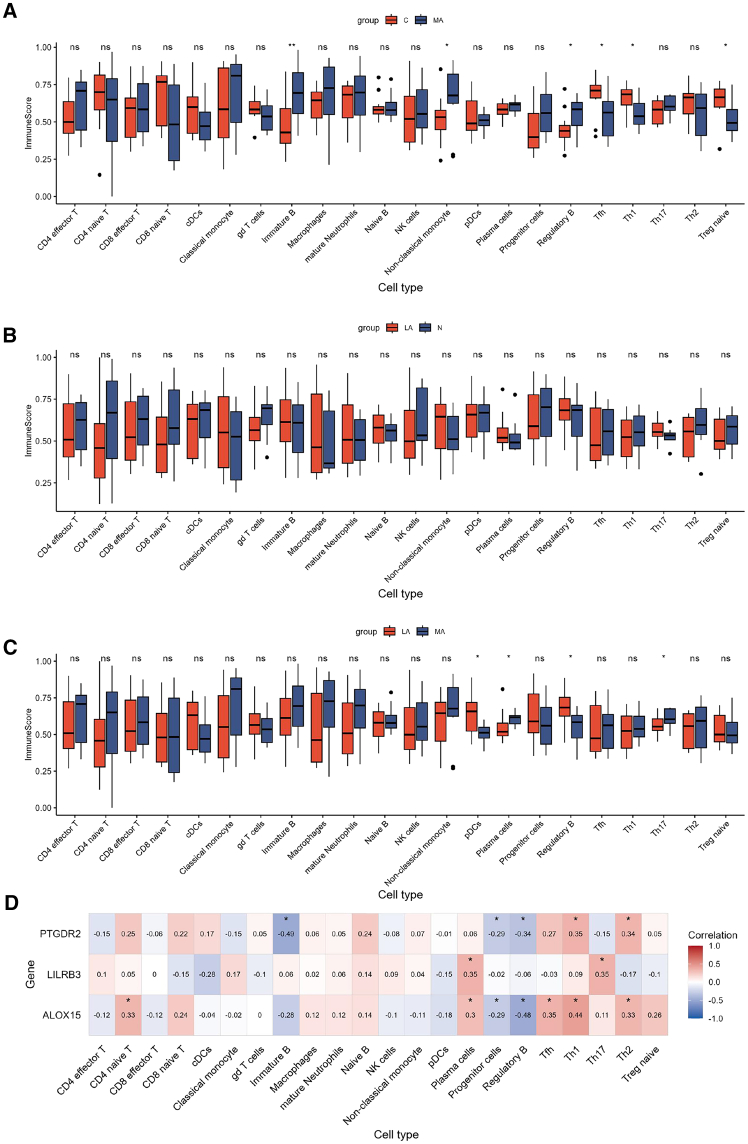
The ssGSEA algorithm was implemented to analyze the infiltration of 22 immune cell types. A significant increase in immature B cells, non-classical monocytes and regulatory B cells (Bregs), along with a significant decrease in naïve-IFN CD8 T cells, Tfh and Treg naive cells, was observed between the MA and C groups (Figure 4A). However, no significant differences were found when comparing the LA and N groups (Figure 4B). pDCs, Bregs, and Th17 cells were found to be changed between LA and MA groups. (Figure 4C). Spearman correlation analysis revealed that PTGDR was negatively correlated with immature B, progenitor cells, Bregs, and positively correlated with Th1 and Th2 cells. LIRB3 was positively correlated with plasma and Th17 cells. ALOX15 was correlated with certain immune cell types like regulatory B, Th1, Tfh, Th2 cells, etc. (Figure 4D).
Figure 4.
Immune cell infiltration analysis and correlation analysis
(A–C) Boxplot showing the difference in immune cell infiltration between C-MA, N-LA, LA-MA. p values were calculated using the non-parametric Mann-Whitney U test. The significance levels are: ∗p < 0.05, ∗∗p < 0.01, ns (not significant).
(D) Spearman correlation analysis of the expression levels of three key genes (ALOX15, LIRB3, PTGDR2) with 22 immune cells. The color intensity reflects the magnitude of the correlation coefficient. Blue indicates a negative correlation and red indicates a positive correlation. ∗ indicates p < 0.05.
Validation and diagnostic efficacy of signature genes
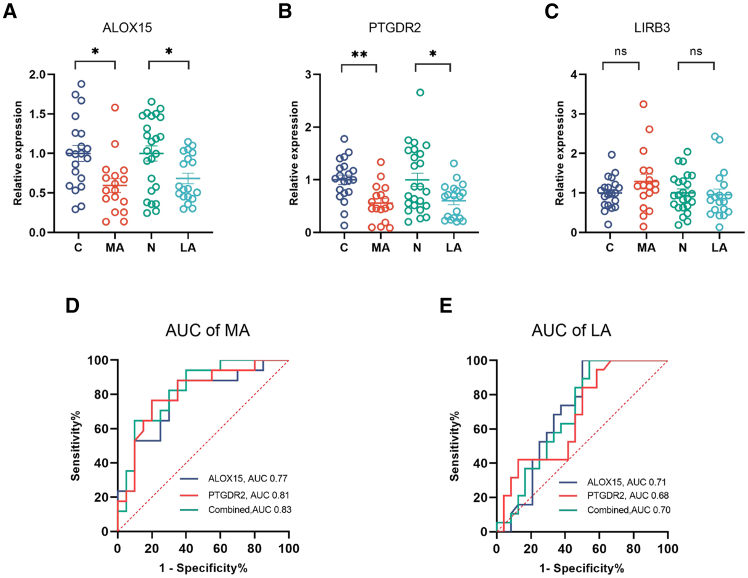
The relative expression levels of three selected signature genes —ALOX15, PTGDR2, and LILRB3— were validated using quantitative reverse transcription polymerase chain reaction (qRT-PCR) in an independent cohort of 80 newborns with respiratory distress. The expression levels of ALOX15 and PTGDR2 were significantly downregulated in both the MA and LA groups compared to the C and N groups. In contrast, LILRB3 showed no statistically significant change (Figures 5A–5C). Receiver operating characteristic (ROC) curve analysis revealed that the AUC for ALOX15 was 0.77 in the MA group and 0.71 in the LA group. PTGDR2 exhibited AUC values of 0.81 and 0.68 in the MA and LA groups, respectively. Combining biomarkers enhanced the diagnostic potential for MA identification, increasing the AUC to 0.83. Whereas this combination showed only limited additional diagnostic value for LA, with an AUC of 0.70. (Figures 5D and 5E).
Figure 5.
Validation and diagnostic efficacy of signature genes
(A–C) Relative mRNA levels of ALOX15, PTGDR2, LIRB3 in the peripheral blood of validation cohort. Data are presented as the means ± SEM and error bars represent SEM. Statistical differences between groups were analyzed using the non-parametric Mann-Whitney U test. ∗ Indicates a difference between groups. ∗p < 0.05, ∗∗p < 0.01, ns (not significant).
(D and E) ROC curves indicate the predictive performance of ALOX15, PTGDR2 and the combined two genes in NARDS subgroups. NARDS: neonatal acute respiratory distress syndrome, ROC: receiver operating characteristic. SEM: standard error of the mean.
Validation of ALOX15 and PTGDR2 in lipopolysaccharide (LPS)-induced neonatal acute lung injury (ALI) model
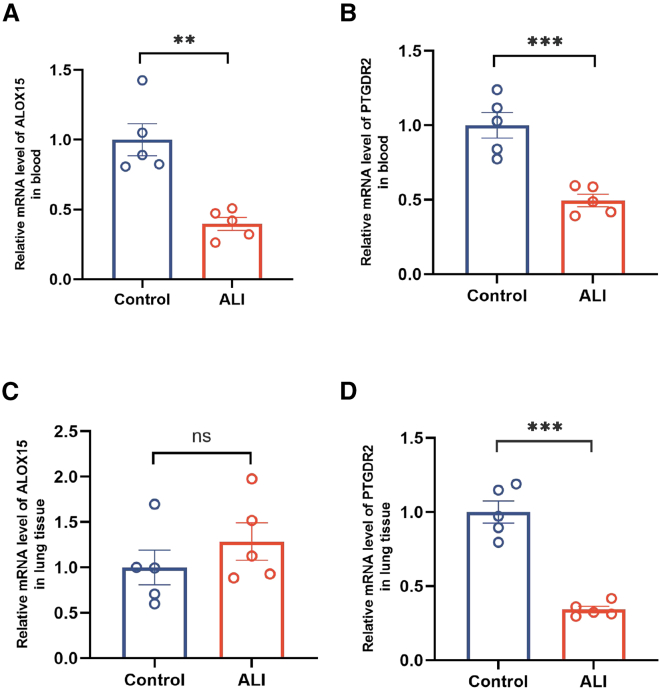
To further investigate the expression patterns of previously validated differential genes in clinical cohorts, we established a neonatal mouse model of ALI through intraperitoneal injection of 25 mg/kg LPS in 6-day-old mice. Compared with the control group, the ALI group exhibited elevated lung injury scores and increased neutrophil infiltration (Ly6G+) (Figure S4), confirming the successful establishment of the ALI neonatal model. qRT-PCR analysis revealed significant reduction of ALOX15 and PTGDR2 mRNA expression in peripheral blood of ALI mice(Figures 6A and 6B). In lung tissue, PTGDR2 expression was similarly reduced, whereas ALOX15 levels showed no statistically significant alteration compared to controls (Figures 6C and 6D).
Figure 6.
Validation of ALOX15 and PTGDR2 in neonatal ALI model
(A and B) ALOX15 and PTGDR2 mRNA levels in peripheral blood.
(C and D) ALOX15 and PTGDR2 mRNA levels in lung tissue. Data are presented as the means ± SEM and error bars represent SEM (n = 5). Statistical differences were analyzed using a two-tailed, unpaired Student's t-test. ∗ Indicates a difference between groups.∗∗p < 0.01,∗∗∗p < 0.001, ns (not significant) versus control group. ALI: acute lung injury. SEM: standard error of the mean.
Discussion
This study provides a comprehensive transcriptomic analysis of NARDS using RNA-seq. Through the analysis of whole blood samples from neonates with respiratory distress, we identified two key genes, ALOX15 and PTGDR2, which demonstrated strong predictive performance in an independent cohort. We further validated the gene changes in a neonatal ALI mouse model induced by LPS, thus supporting their potential as predictive biomarkers.
Early diagnosis of NARDS in neonates remains challenging, especially for preterm infants, as imaging findings often resemble those of ARDS, leading to diagnostic uncertainty in the early stages. This uncertainty can delay treatment and increase the risk of mortality and complications. The Montreux definition4 emphasizes that the diagnosis of NARDS depends on comprehensive clinical evaluation, including the timeline, lung imaging, oxygenation deficits, and the exclusion of other causes of respiratory distress. To address this challenge, we enrolled a cohort of neonates with respiratory distress to identify features that are specific to NARDS in this population. By selecting neonates younger than three days, we minimized gene expression differences due to normal physiological variations, ensuring comparability between groups.
Systemic acute inflammation involving a series of immune cells and networks of cytokines is one of the prominent features of ARDS (especially caused by indirect triggers).15 Transcriptomic analysis on peripheral blood helps to uncover significant alterations in immune characteristics within the circulatory system, including potential dysregulation of innate and adaptive immunity.16 While this approach has been widely applied to ARDS in adults and children, its findings may not be directly applicable to NARDS due to the unique perinatal conditions and the substantial changes in both the lung status and the immune system.17 Our study highlights these differences and provides new insights into the immune mechanisms that contribute to NARDS.
Our study showed that GA has a strong effect on the transcriptomic profile of newborns, which is consistent with prior studies.18,19 PCA analysis presented distinct clustering of samples from the NARDS and control groups driven by GA. The GAM model also suggested a positive trend on gene expression by GA. These results highlighted the major changes in the rapid development of neonate. To further explore the underlying effects, we stratified our cohort with 34 weeks of GA as the cutoff. In the subgroup of neonates born before 34 weeks’ gestation, 50% of NRDS and 33% of ARDS cases were associated with PROM, ARDS patients exhibited significantly lower 5-min Apgar scores and a higher incidence of EOS, suggesting a synergistic interplay among birth asphyxia, systemic inflammatory responses, and immature immune development in shaping the distinct transcriptomic profile of lower GA-associated ARDS. In contrast, among neonates born at ≥34 weeks’ gestation, ARDS cases demonstrated markedly elevated plasma PCT levels, with 50% of these cases secondary to EOS. This observation highlights potential divergence in inflammatory activation mechanisms underlying ARDS pathogenesis at different GAs.20
Reactome pathway and GSEA analysis revealed the enrichment of multiple biological pathways in NARDS patients. In the pilot control group, neonates with smaller GA were all diagnosed with NRDS, and 91.7% of them received PS administration, none of the controls with GA>34 weeks required supplementation. Consistent with a prior study, our functional analysis demonstrated that, compared to group C, smaller NRDS cases shared overlapping pathological responses with ARDS neonates of higher GA, like interferon signaling and several inflammatory pathways. Additionally, NRDS demonstrated distinct abnormalities in phospholipid metabolism— a process critically involved in synthesis, secretion, and functional regulation of PS.21 These findings highlight the clinical necessity of PS supplementation in this population. Moreover, NARDS with larger GA showed significant enrichment in pathways such as “O2/CO2 exchange in erythrocytes" and “erythrocytes’ uptake of carbon dioxide and release of oxygen”, while those with a GA < 34 weeks demonstrated significant enrichment in the “heme biosynthesis” pathway. These results suggest oxygenation dysfunction in NARDS patients, revealing the urgent need for ventilation support. It is noteworthy that while ARDS in more preterm infants showed certain activation in a wide range of immune pathways compared to NRDS, several immune responses remained suppressed relative to ARDS with larger GA(Table S5). Infants with NARDS of smaller GA experience the double “hit” of both inflammation and PS deficient, in our cohort, this group required more frequent PS, longer duration of mechanical ventilation, while exhibited a higher mortality rate. This disparity may suggest a relative deficiency in immunomodulatory function in response to inflammation and more severe condition associated with smaller GA. Intriguingly, IFN-related pathways exhibited differential regulatory patterns in NARDS pathogenesis. Functional analyses demonstrated suppressed type I/II IFN responses in term/late preterm ARDS, contrasting with upregulated type II activity in <34-week cases. Researches indicate that type I IFN (IFN-α/β) responses were enhanced22,23 in older populations while reduced in more severe ARDS in infants, and a reduction of IFN-γ response correlated with increased severity,24,25 which was consistent with our results. Clinical trials have demonstrated that both IFN-α/β26,27 and IFN-γ28 contribute to the recovery of ARDS patients from diverse etiologies, significantly reducing mortality. Additionally, short-term IFN-γ therapy may enhance the pro-inflammatory and phagocytic responses in leukocytes from preterm infants.29 However, the role of interferons in NARDS remains unexplored, targeted modulation of this axis may represent a novel therapeutic strategy, though systematic evaluation through preclinical studies and translational trials remains essential to validate efficacy and safety profiles.
The uncontrolled inflammation in ARDS may induce the infiltration of various innate and adaptive immune cells (e.g., macrophages, neutrophils, and T cells), along with immune tolerance and suppression.16,29 In this study, we manually collected single-cell data of blood immune cells combined with ssGSEA algorithm to peer into the potential changes of NARDS circulatory immunity. Neonates with larger GA(≥34 weeks) exhibited characteristic immune dysregulation featuring expanded regulatory populations alongside several diminished T helper subtypes – a pathological overlap with neonatal sepsis associated with, and potentially mediated through, Bregs expansion30 and Treg reduction.31 Preterm infants are characterized by diminished immune cell responsiveness and reduced production of multiple pro-inflammatory cytokines,32 for infants born before 34 weeks, no significant change of immune cell infiltration in ARDS were found compared with NRDS, potentially attributable to this inherent immune immaturity. While neonates with larger GA showed decreased pDC and regulatory B cells and increased plasma and Th17 cells, suggesting that the imbalance in immune cell development may contribute to the phenotypic differences observed in NARDS across varying GA. However, the intricate changes of these immune cells in NARDS still need to be comprehensively verified and evaluated by advanced multi-parameter flow cytometry, single-cell transcriptomics and targeted cytokines analysis.
Signature genes of NARDS were screened by GAM and three machine learning algorithms (LASSO, RF, and XGBOOST). Among these candidates, ALOX15 and PTGDR2 were validated as significantly dysregulated through q-PCR in an independent cohort of 80 neonates with respiratory distress, demonstrating consistent diagnostic performance for NARDS across GA subgroups. Further validation in an LPS-induced neonatal mouse ALI model revealed tissue-specific gene expression divergence: ALOX15 mRNA levels were significantly reduced in peripheral blood but remained unchanged in lung tissue, whereas PTGDR2 expression was downregulated in both circulatory and lung tissue compared to controls.
ALOX15(15-lipoxygenase), is a key functional gene in the lipoxygenase (LOX) family, constitutively expressed in various human cells, including reticulocytes, eosinophils, dendritic cells, alveolar macrophages, and immature dendritic cells and so on.33 ALOX15 and its metabolites have been shown to be both pro- and anti-inflammatory, of which metabolite 12-hydroxyeicosatetraenoic acid (12(S)- HETE) act as a potent pro-inflammatory chemoattractant for neutrophils and leukocytes.34 Additionally, ALOX15 is involved in the lipoxygenase pathway to generate eicosanoids, which are essential mediators of homeostatic and inflammatory processes and serve as potent regulators of macrophage function.33 ALOX15 plays a critical role in the pathogenesis of ARDS. In animal models of acute lung injury, the elimination or inhibition of ALOX15 reduces neutrophil recruitment,35 alleviates lung iron deposition, inflammation, and tissue damage.36 Notably, a recent study has highlighted the key role of ALOX15 in programming neutrophil-dependent alveolar macrophage function during lung development.37 PTGDR2 (prostaglandin D2 receptor 2), also known as chemoattractant receptor-homologous molecules expressed on Th2 (CRTH2) cells, is a G protein–coupled receptor for prostaglandin D2 (PGD2). Activation of PTGDR2 has been shown to elicit proinflammatory responses in leukocytes, including the chemotaxis of eosinophils, basophils, and Th2 cells.38 Researches indicate that the activation of PGD2/CRTH2 on macrophages promotes neutrophil migration and the release of pro-inflammatory cytokines, which worsens disease.39 The administration of PTGDR2 antagonists can significantly reduce the infiltration of inflammatory cells, neutrophils and macrophages.40 One study suggests that lymphocytopenia in patients with ARDS may be associated with PGD2/PTGDR2 signaling, which could be addressed with its antagonists.41 However, most of these findings are derived from studies conducted in adults. In our study, the expression of ALOX15 and PTGDR2 in blood was significantly reduced in NARDS patients and neonatal ALI mouse, while their expression in ALI lung tissue showed variability. The observed differential expression of these genes may due to the incomplete maturation of immune defense mechanisms and underdeveloped hematopoietic function of bone marrow during the early postnatal stage, particularly in preterm infants,42,43 as the patients in our study were within first three days after birth. Additionally, NARDS is strongly associated with the IFN pathway. Immune-related analysis revealed that ALOX15 and PTGDR2 are linked to various immune cells, including Th1 and Breg cells, which, through the cross-talk of cytokines such as IFN-γ and IL-10, regulate the dynamic balance between immune activation and homeostasis maintenance within the IFN signaling pathway.44,45 Therefore, we will further explore the immune regulatory roles and molecular mechanisms of ALOX15 and PTGDR2 in NARDS in future research.
Collectively, we used the transcriptome analysis to describe the peripheral blood gene expression profile of NARDS. We highlighted the potential disparities in the pathological response to ARDS that caused by different GA. By integrating bioinformatical analysis with machine learning algorithms, we identified and independently validated two genes (ALOX15, PTGDR2) which show good predictive performance of NARDS at various gestation. The two genes were further validated in an independent clinical cohort and an LPS-induced neonatal mouse model of ALI. These findings lay the groundwork for future research into the early diagnosis and personalized treatment of NARDS.
Limitations of the study
There are some limitations that need to be acknowledged. First, as our purpose is to identify the characteristics of NARDS in patients with respiratory distress after birth, we included children in the first three days of life, which may not represent all cases of NARDS. This may lead to discrepancies in the etiological spectrum of NARDS compared to other publications. The primary trigger of our NARDS was sepsis. Second, due to the small sample size and the unique nature of the included cohort, subgroup analysis for different injury factors could not be conducted. Third, we collected blood samples upon admission, which may be ideal for predicting long-term outcomes. Finally, although we validated key genes in an independent cohort, the study’s single-center design and relatively small sample size may limit the generalizability and statistical power of the findings, so our results still needed to be verified by data from larger, multicenter cohorts. Future multicenter, longitudinal cohorts and well-designed molecular functional experiments may enhance our understanding of the progression and development of NARDS.
Resource availability
Lead contact
Requests for further information and resources should be directed to and will be fulfilled by the lead contact, Yuan Shi (shiyuan@hospital.cqmu.edu.cn).
Materials availability
This study did not generate new unique reagents.
Data and code availability
-
•
Transcriptomic data are available in the SRA database under accession number [SRA: PRJNA1194327] (See https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1194327).
-
•
This paper does not report original code.
-
•
Requests for any additional information are available from the corresponding author on reasonable request.
Acknowledgments
This study was funded by National Key Research and Development Program of China (No. 2022YFC2704803), and the National Natural Science Foundation of China (No. 82001602), Special Funding for Postdoctoral Research Projects of Chongqing (No. 2022CQBSHTB3085), China Postdoctoral Science Foundation (No. 2023MD744152), Natural Science Foundation of Chongqing (No. CSTB2024NSCQ-MSX0491), the special key project of technology innovation and application development of Chongqing Science and Technology Bureau (CSTC2021jscx-gksb-N0015), Key Research and Development Program of Jiangxi (No. 20243BBI91020).
Author contributions
Y.H. and Y.S. conceptualized and directed this study. Q.A., W.J., J.Y., Y.F., L.S., Y.Y., F.C., and Y.L. were responsible for recruitment, sample collection, and acquisition of clinical data. C.L. was responsible for analysis, interpretation of data, and drafting of the manuscript. Y.H. and Y.S. critically revised the manuscript. All authors reviewed the manuscript and approved the final version before submission. All authors had full access to the data in this study.
Declaration of interests
All authors declare no competing interests.
Declaration of generative AI and AI-assisted technologies
During the drafting of this work the authors used ChatGPT for language refinement only. All authors have reviewed this manuscript and take full responsibility for its originality, accuracy, and integrity.
STAR★Methods
Key resources table
| REAGENT or RESOURCE | SOURCE | IDENTIFIER |
|---|---|---|
| Antibodies | ||
| anti-Ly6G antibody | ServiceBio | RRID: AB_2814689 |
| Biological samples | ||
| Whole blood | Children’s Hospital of Chongqing Medical University | |
| Chemicals, peptides, and recombinant proteins | ||
| LPS(Escherichia coli O111:B4) | Sigma | L2630 |
| Critical commercial assays | ||
| PAXgene RNA tubes | BD Biosciences | 762165 |
| PAXgene Blood miRNA Kit | Qiagen | 763134 |
| NovaSeq 6000 platform | Illumina | |
| SteadyPure Blood RNA Extraction Kit | Accurate Biotechnology | AG21025 |
| SteadyPure Quick RNA Extraction Kit | Accurate Biotechnology | AG21023 |
| Evo M-MLV RT Mix Tracking Kit with gDNA Clean for qPCR | Accurate Biotechnology | AG11734 |
| SYBR Green Premix Pro Taq HS qPCR Kit | Accurate Biotechnology | AG11733 |
| Deposited data | ||
| Respiratory distress cohort | SRA database | SRA: PRJNA1194327 |
| Experimental models: organisms/strains | ||
| Mouse: C57BL/6 | Laboratory Animal Center of Chongqing Medical University | |
| Oligonucleotides | ||
| Primers for mRNA expression, see Table S2 | ||
| Software and algorithms | ||
| R | cran.r-project.org | v 4.4.1 |
| HISAT2 | daehwankimlab.github.io/hisat2 | v 2.1.0 |
| HTSeq | Python | v 0.9.1 |
| Fastp | cran.r-project.org | v 0.22.0 |
| DESeq | bioconductor.org | v 1.38.3 |
| DESeq2 | bioconductor.org | v 1.44.0 |
| Mgcv | cran.r-project.org | v 1.9–1 |
| Glmnet | cran.r-project.org | v 4.1-8 |
| randomForest | cran.r-project.org | v 4.7-1.2 |
| Xgboost | cran.r-project.org | v 1.7.8.1 |
| ReactomePA | bioconductor.org | v 1.48.0 |
| ClusterProfiler | bioconductor.org | v 4.12.6 |
| GSVA | bioconductor.org | v 1.52.3 |
| ggplot2 | cran.r-project.org | v 4.12.6 |
| SPSS | IBM | v 26 |
| Prism | GraphPad | v 8.0.2 |
| Other | ||
| NovaSeq | Illumina | 6000 |
Experimental model and study participant details
Human participants
This was a prospective cohort study performed from October 2022 to August 2023 and approved by Children’s Hospital of Chongqing Medical University (Approval No. 2021-414) and was registered on Chinese Clinical Trial Registry (ChiCTR) (Registration No. ChiCTR2400085274), with consent confirmed by guardians before enrollment. Inclusion criteria were (1) admitted within three days after birth, (2) presented with respiratory distress within the first 24 hours after birth, including progressive dyspnea, grunting, intercostal and subcostal retractions, and cyanosis, (3) requirement for non-invasive/invasive respiratory support within 24 hours of birth, and (4) relevant radiographic findings such as air bronchograms, ground-glass opacities, or “white lung”. Exclusion criteria were (1) congenital structural or developmental abnormalities impacting respiratory function, (2) intubation due to surgical intervention, (3) hereditary metabolic diseases, or (4) inability to obtain consent.
Patients meeting all five Montreux diagnostic criteria4 were included in the NARDS group. To address heterogeneity arising from organ immaturity and immune system development in preterm infants, the NARDS cohort was stratified by GA: group LA(GA less than 34 weeks) and group MA (GA more than 34 weeks). The control groups were 1) diagnosed with NRDS or TTN; 2) rapid resolution of respiratory distress and radiographic pulmonary infiltrates following PS administration, lung recruitment maneuvers, or combined therapy;3) individuals matched to NARDS subgroups by GA and sex ratio. The control cohort was further stratified into group N (GA <34 weeks, matched to group LA) and group C (GA ≥34 weeks, matched to group MA). NRDS was defined as progressive respiratory distress appearing within the first 24 hours of life, caused by insufficient PS due to lung immaturity in newborns.46 TTN was characterized by self-limiting tachypnea developing within 24 hours of birth and resolving spontaneously within 72 hours, resulted from delayed reabsorption and clearance of alveolar fluid.8
For the sample collection, 0.5-1 ml whole blood of each patient was collected using PAXgene RNA tubes (BD Biosciences, San Jose, CA) at admission, thoroughly mixed and then transferred to freezer (−80°C) within 24 hours.
Animal models
6-to-8-week-old specific pathogen-free (SPF) C57BL/6 mice were housed in an SPF-certified animal facility under controlled environmental conditions, maintaining a temperature of 22 ± 2°C, humidity of 55 ± 5%, and a 12-hour light/dark cycle. Pregnant dams were individually housed after confirmed conception, with neonatal offspring at postnatal day 6 (P6) selected as study subjects. All experimental protocols were reviewed and approved by the Experimental Animal Ethics Committee of Children's Hospital Affiliated to Chongqing Medical University (Approval No. CHCMU-IACUC20240628011). P6 neonates were randomly assigned to two experimental groups: the ALI group received intraperitoneal injections of 25 mg/kg LPS (Escherichia coli O111:B4; Sigma, St. Louis, MO, USA) diluted in phosphate-buffered saline (PBS), while the control group was administered equal volumes of PBS. All mice were euthanized by anesthetic overdose 24 hours post-injection, and lung tissue and blood samples were collected for downstream analysis.
Method details
RNA-seq and data processing
Total RNA was isolated using PAXgene Blood miRNA Kit (Qiagen, Germany), mRNA was then purified using poly-T oligo-attached magnetic beads and subsequently fragmented and reverse-transcribed into cDNA. Sequencing libraries were prepared and sequenced on the NovaSeq 6000 platform (Illumina). Quality control of the sequencing data was conducted with fastp (v 0.22.0) to obtain high-quality reads (Clean Data), which were then aligned to the reference genome using HISAT2 (v 2.1.0). Gene counts were obtained with HTSeq (v 0.9.1), then difference expression of genes was analyzed by DESeq (v 1.38.3) with screened conditions as follows: expression difference multiple |log2FoldChange| > 1, significant p-value < 0.05. PlotPCA in DESeq2 (v 1.44.0) was used to implement principal component analysis (PCA) for the high-dimensional data reduction and visualization of variance between groups.
Analysis on gene expression variance and gene screening
The generalized additive model (GAM) was used to assess and adjust for the effect of GA on gene profile of NARDS to identify reliable predictive markers. Subsequently, we employed the least absolute shrinkage and selection operator (LASSO), Random Forest (RF), and Extreme Gradient Boosting (XGBoost) algorithms to pinpoint key biomarkers among the DEGs identified by GAM. For LASSO, we employed a binomial family distribution with pure L1 regularization (α=1). The optimal regularization parameter (λ) was determined through 10-fold cross-validation, with genes exhibiting non-zero coefficients retained as candidate biomarkers. A fixed random seed was implemented throughout the analytical workflow to ensure reproducibility. The Random Forest model was optimized through systematic adjustment of mtry (variables per split) and nodesize (minimum node size) by minimizing out-of-bag (OOB) error to achieve stable model performance. In the XGBoost model, the maximum tree depth was constrained to 6, with 5-fold cross-validation and early stopping to determine optimal iterations, while the area under the AUC served as the primary performance evaluation metric.Significance of the overlapped genes identified by these three machine learning techniques were assessed in subgroup comparisons. These analyses were conducted using the mgcv (v 1.9-1), glmnet (v 4.1-8), randomForest (v 4.7-1.2), and xgboost (v 1.7.8.1) packages in R.
Enrichment analysis and immune cell infiltration analysis
Functional enrichment analysis was applied in the subgroups. The R packages ReactomePA (v 1.48.0) and ClusterProfiler (v 4.12.6) were used to perform enrichment analysis of differentially expressed genes (DEGs) in the Reactome database, significance was confirmed based on an adjusted p-value (p. adjust) threshold of < 0.05. Gene set enrichment analysis (GSEA) of all genes was conducted by ClusterProfiler targeting the MSigDB H gene set (h.all.v2024.1.Hs.symbols.gmt) and the filtering criteria were: p. adjust (< 0.05), FDR q-value (< 0.25), and normalized enrichment score (|NES|> 1). Additionally, the single-sample GSEA (ssGSEA) algorithm was introduced to evaluate 22 immune cell types in the peripheral blood using GSVA (v 1.52.3). Signature panels for each immune cell type were manually curated from several publications on single-cell analysis of blood cells47,48,49,50 and summarized in Table S1. The relative abundance of each immune cell type was represented by an enrichment score in ssGSEA analysis, normalized to a 0-1 scale for visualization. The results were visualized using the ggplot2 (v 4.12.6) package.
Lung histology
Lung tissues were fixed in 4% paraformaldehyde and paraffin-embedded for histological processing. Sequential 4-μm sections were stained with hematoxylin and eosin (H&E) using standardized protocols. Blinded histological assessment was performed by quantifying pathological parameters: alveolar edema, hemorrhage, leukocyte infiltration, and septal wall thickening. Five randomly selected fields per mouse were systematically evaluated, with the mean value of these assessments calculated as the composite lung injury score.51
Immunohistochemistry (IHC) staining
Ly6G IHC was performed to evaluate neutrophil infiltration in lung tissues. Paraffin-embedded lung sections were incubated with an anti-Ly6G antibody (ServiceBio, Wuhan, China), followed by a horseradish peroxidase (HRP)-labeled secondary antibody (AiFang Biological, Hunan, China) for 1 hour at room temperature. Images were captured using a Nikon DS-U microscope (Tokyo, Japan). Ly6G-positive areas were quantified using percentage area coverage (Area%).
Signature genes validated by qRT-PCR in clinical cohort and animal model
The expression levels of three key genes (ALOX15, PTGDR2, and LILRB3) were validated in a clinical cohort using qRT-PCR, and the identified differentially expressed genes were further examined in a neonatal mouse model of ALI. RNA from pediatric blood samples was extracted using the PAXgene Blood miRNA Kit (Qiagen, Germany). For mouse specimens, RNA from blood and lung tissue was isolated using the SteadyPure Blood RNA Extraction Kit (Accurate Biotechnology, Hunan, China) and SteadyPure Quick RNA Extraction Kit (Accurate Biotechnology, Hunan, China), respectively. Reverse transcription was performed with the Evo M-MLV RT Mix Tracking Kit with gDNA Clean for qPCR (Accurate Biotechnology, Hunan, China). qRT-PCR assays were conducted using SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biotechnology, Hunan, China), with primer sequences listed in Table S2. Human blood samples were normalized to ACTB, murine blood to HPRT, and murine lung tissue to Actin, with relative quantification calculated via the 2−ΔΔCq method. Clinical validation results were further analyzed using ROC curves, and the AUC was determined to evaluate the predictive value of these genes for NARDS.
Quantification and statistical analysis
Spearman correlation analysis was performed using R software (version 4.4.1). Statistical analyses were performed with SPSS (IBM SPSS Statistics, version 26) and graphs were generated using GraphPad Prism (version 8.0.2). Quantitative variables were presented as mean ± SEM unless otherwise indicated. Qualitative variables were presented as numbers and percentages. Differences in quantitative variables were assessed with 2-tailed student’s t test or Mann–Whitney test as appropriate, and qualitative variables were analyzed using with the χ2 test or the Fisher’s test. A two-sided p-value of <0.05 was considered statistically significant.
Additional resources
The prospective cohort study was registered in the Chinese Clinical Trial Registry (Registration number: ChiCTR2400085274; chictr.org.cn) and is available at: https://www.chictr.org.cn/showproj.html?pid=219009.
Published: June 25, 2025
Footnotes
Supplemental information can be found online at https://doi.org/10.1016/j.isci.2025.113007.
Contributor Information
Yu He, Email: heyu@hospital.cqmu.edu.cn.
Yuan Shi, Email: shiyuan@hospital.cqmu.edu.cn.
Supplemental information
References
- 1.Ashbaugh D.G., Bigelow D.B., Petty T.L., Levine B.E. Acute respiratory distress in adults. Lancet. 1967;2:319–323. doi: 10.1016/s0140-6736(67)90168-7. [DOI] [PubMed] [Google Scholar]
- 2.ARDS Definition Task Force. Ranieri V.M., Rubenfeld G.D., Thompson B.T., Ferguson N.D., Caldwell E., Fan E., Camporota L., Slutsky A.S. Acute respiratory distress syndrome: the Berlin Definition. JAMA. 2012;307:2526–2533. doi: 10.1001/jama.2012.5669. [DOI] [PubMed] [Google Scholar]
- 3.Khemani R.G., Smith L.S., Zimmerman J.J., Erickson S., Pediatric Acute Lung Injury Consensus Conference Group Pediatric acute respiratory distress syndrome: definition, incidence, and epidemiology: proceedings from the Pediatric Acute Lung Injury Consensus Conference. Pediatr. Crit. Care Med. 2015;16:S23–S40. doi: 10.1097/PCC.0000000000000432. [DOI] [PubMed] [Google Scholar]
- 4.De Luca D., van Kaam A.H., Tingay D.G., Courtney S.E., Danhaive O., Carnielli V.P., Zimmermann L.J., Kneyber M.C.J., Tissieres P., Brierley J., et al. The Montreux definition of neonatal ARDS: biological and clinical background behind the description of a new entity. Lancet Respir. Med. 2017;5:657–666. doi: 10.1016/s2213-2600(17)30214-X. [DOI] [PubMed] [Google Scholar]
- 5.Chen L., Li J., Shi Y., Chinese Neonatal ARDS ChiNARDS study group Clinical characteristics and outcomes in neonates with perinatal acute respiratory distress syndrome in China: a national, multicentre, cross-sectional study. eClinicalMedicine. 2023;55 doi: 10.1016/j.eclinm.2022.101739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.De Luca D., Tingay D.G., van Kaam A.H., Courtney S.E., Kneyber M.C.J., Tissieres P., Tridente A., Rimensberger P.C., Pillow J.J., Neonatal ARDS Project Collaboration Group Epidemiology of Neonatal Acute Respiratory Distress Syndrome: Prospective, Multicenter, International Cohort Study. Pediatr. Crit. Care Med. 2022;23:524–534. doi: 10.1097/pcc.0000000000002961. [DOI] [PubMed] [Google Scholar]
- 7.Mahoney A.D., Jain L. Respiratory Disorders in Moderately Preterm, Late Preterm, and Early Term Infants. Clin. Perinatol. 2013;40:665–678. doi: 10.1016/j.clp.2013.07.004. [DOI] [PubMed] [Google Scholar]
- 8.Hermansen C.L., Mahajan A. Newborn Respiratory Distress. Am. Fam. Physician. 2015;92:994–1002. [PubMed] [Google Scholar]
- 9.Donda K., Vijayakanthi N., Dapaah-Siakwan F., Bhatt P., Rastogi D., Rastogi S. Trends in epidemiology and outcomes of respiratory distress syndrome in the United States. Pediatr. Pulmonol. 2019;54:405–414. doi: 10.1002/ppul.24241. [DOI] [PubMed] [Google Scholar]
- 10.Cao S., Li H., Xin J., Jin Z., Zhang Z., Li J., Zhu Y., Su L., Huang P., Jiang L., et al. Identification of genetic profile and biomarkers involved in acute respiratory distress syndrome. Intensive Care Med. 2024;50:46–55. doi: 10.1007/s00134-023-07248-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bos L.D.J., Scicluna B.P., Ong D.S.Y., Cremer O., van der Poll T., Schultz M.J. Understanding Heterogeneity in Biologic Phenotypes of Acute Respiratory Distress Syndrome by Leukocyte Expression Profiles. Am. J. Respir. Crit. Care Med. 2019;200:42–50. doi: 10.1164/rccm.201809-1808OC. [DOI] [PubMed] [Google Scholar]
- 12.Yehya N., Varisco B.M., Thomas N.J., Wong H.R., Christie J.D., Feng R. Peripheral blood transcriptomic sub-phenotypes of pediatric acute respiratory distress syndrome. Crit. Care. 2020;24:681. doi: 10.1186/s13054-020-03410-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Kangelaris K.N., Prakash A., Liu K.D., Aouizerat B., Woodruff P.G., Erle D.J., Rogers A., Seeley E.J., Chu J., Liu T., et al. Increased expression of neutrophil-related genes in patients with early sepsis-induced ARDS. Am. J. Physiol. Lung Cell. Mol. Physiol. 2015;308:L1102–L1113. doi: 10.1152/ajplung.00380.2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Englert J.A., Cho M.H., Lamb A.E., Shumyatcher M., Barragan-Bradford D., Basil M.C., Higuera A., Isabelle C., Vera M.P., Dieffenbach P.B., et al. Whole blood RNA sequencing reveals a unique transcriptomic profile in patients with ARDS following hematopoietic stem cell transplantation. Respir. Res. 2019;20 doi: 10.1186/s12931-019-0981-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Bos L.D.J., Ware L.B. Acute respiratory distress syndrome: causes, pathophysiology, and phenotypes. Lancet. 2022;400:1145–1156. doi: 10.1016/S0140-6736(22)01485-4. [DOI] [PubMed] [Google Scholar]
- 16.Kang Z.Y., Huang Q.Y., Zhen N.X., Xuan N.X., Zhou Q.C., Zhao J., Cui W., Zhang Z.C., Tian B.P. Heterogeneity of immune cells and their communications unveiled by transcriptome profiling in acute inflammatory lung injury. Front. Immunol. 2024;15 doi: 10.3389/fimmu.2024.1382449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Khemani R.G., Wilson D.F., Esteban A., Ferguson N.D. Evaluating the Berlin Definition in pediatric ARDS. Intensive Care Med. 2013;39:2213–2216. doi: 10.1007/s00134-013-3094-6. [DOI] [PubMed] [Google Scholar]
- 18.Kwinta P., Bokiniec R., Bik-Multanowski M., Gunther C.-C., Grabowska A., Książek T., Madetko-Talowska A., Szewczyk K., Szwarc-Duma M., Borszewska-Kornacka M.K., et al. Comparison of whole genome expression profile between preterm and full-term newborns. Ginekol. Pol. 2017;88:434–441. doi: 10.5603/GP.a2017.0080. [DOI] [PubMed] [Google Scholar]
- 19.Olin A., Henckel E., Chen Y., Lakshmikanth T., Pou C., Mikes J., Gustafsson A., Bernhardsson A.K., Zhang C., Bohlin K., Brodin P. Stereotypic Immune System Development in Newborn Children. Cell. 2018;174:1277–1292.e14. doi: 10.1016/j.cell.2018.06.045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Humberg A., Fortmann I., Siller B., Kopp M.V., Herting E., Göpel W., Härtel C., German Neonatal Network German Center for Lung Research and Priming Immunity at the beginning of life PRIMAL Consortium Preterm birth and sustained inflammation: consequences for the neonate. Semin. Immunopathol. 2020;42:451–468. doi: 10.1007/s00281-020-00803-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Numata M., Kandasamy P., Voelker D.R. The anti-inflammatory and antiviral properties of anionic pulmonary surfactant phospholipids. Immunol. Rev. 2023;317:166–186. doi: 10.1111/imr.13207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jiang Y., Rosborough B.R., Chen J., Das S., Kitsios G.D., McVerry B.J., Mallampalli R.K., Lee J.S., Ray A., Chen W., Ray P. Single cell RNA sequencing identifies an early monocyte gene signature in acute respiratory distress syndrome. JCI Insight. 2020;5 doi: 10.1172/jci.insight.135678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Grunwell J.R., Stephenson S.T., Mohammad A.F., Jones K., Mason C., Opolka C., Fitzpatrick A.M. Differential type I interferon response and primary airway neutrophil extracellular trap release in children with acute respiratory distress syndrome. Sci. Rep. 2020;10 doi: 10.1038/s41598-020-76122-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Palacios-Gutierrez J.J., Rodriguez-Guardado A., Arias-Guillen M., Alonso-Arias R., Palacios-Penedo S., Garcia-Garcia J.M., Balbin M., Perez-Hernandez D., Sandoval-Torrientes M., Torreblanca-Gil A., et al. Clinical and Epidemiological Correlates of Low IFN-Gamma Responses in Mitogen Tube of QuantiFERON Assay in Tuberculosis Infection Screening During the COVID-19 Pandemic: A Population-Based Marker of COVID-19 Mortality? Arch. Bronconeumol. 2022;58:649–659. doi: 10.1016/j.arbres.2022.01.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Flerlage T., Crawford J.C., Allen E.K., Severns D., Tan S., Surman S., Ridout G., Novak T., Randolph A., West A.N., Thomas P.G. Single cell transcriptomics identifies distinct profiles in pediatric acute respiratory distress syndrome. Nat. Commun. 2023;14:3870. doi: 10.1038/s41467-023-39593-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ranieri V.M., Pettilä V., Karvonen M.K., Jalkanen J., Nightingale P., Brealey D., Mancebo J., Ferrer R., Mercat A., Patroniti N., et al. Effect of Intravenous Interferon beta-1a on Death and Days Free From Mechanical Ventilation Among Patients With Moderate to Severe Acute Respiratory Distress Syndrome: A Randomized Clinical Trial. JAMA. 2020;323:725–733. doi: 10.1001/jama.2019.22525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhou Q., Chen V., Shannon C.P., Wei X.S., Xiang X., Wang X., Wang Z.H., Tebbutt S.J., Kollmann T.R., Fish E.N. Interferon-alpha2b Treatment for COVID-19. Front. Immunol. 2020;11:1061. doi: 10.3389/fimmu.2020.01061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Nguyen L.S., Ait Hamou Z., Gastli N., Chapuis N., Pène F. Potential role for interferon gamma in the treatment of recurrent ventilator-acquired pneumonia in patients with COVID-19: a hypothesis. Intensive Care Med. 2021;47:619–621. doi: 10.1007/s00134-021-06377-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.He D., Yu Q., Zeng X., Feng J., Yang R., Wan H., Zhong Y., Yang Y., Zhao R., Lu J., Zhang J. Single-Cell RNA Sequencing and Transcriptome Analysis Revealed the Immune Microenvironment and Gene Markers of Acute Respiratory Distress Syndrome. J. Inflamm. Res. 2023;16:3205–3217. doi: 10.2147/JIR.S419576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Pan X., Ji Z., Xue J. Percentage of Peripheral CD19+CD24hiCD38hi Regulatory B Cells in Neonatal Sepsis Patients and Its Functional Implication. Med. Sci. Monit. 2016;22:2374–2378. doi: 10.12659/msm.895421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Fortmann I., Dammann M.-T., Siller B., Humberg A., Demmert M., Tüshaus L., Lindert J., van Zandbergen V., Pagel J., Rupp J., et al. Infants Younger Than 90 Days Admitted for Late-Onset Sepsis Display a Reduced Abundance of Regulatory T Cells. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.666447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.van Duuren I.C., van Hengel O.R.J., Penders J., Duijts L., Smits H.H., Tramper-Stranders G.A. The developing immune system in preterm born infants: From contributor to potential solution for respiratory tract infections and wheezing. Allergy. 2024;79:2924–2942. doi: 10.1111/all.16342. [DOI] [PubMed] [Google Scholar]
- 33.Singh N.K., Rao G.N. Emerging role of 12/15-Lipoxygenase (ALOX15) in human pathologies. Prog. Lipid Res. 2019;73:28–45. doi: 10.1016/j.plipres.2018.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kayama Y., Minamino T., Toko H., Sakamoto M., Shimizu I., Takahashi H., Okada S., Tateno K., Moriya J., Yokoyama M., et al. Cardiac 12/15 lipoxygenase-induced inflammation is involved in heart failure. J. Exp. Med. 2009;206:1565–1574. doi: 10.1084/jem.20082596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rossaint J., Nadler J.L., Ley K., Zarbock A. Eliminating or blocking 12/15-lipoxygenase reduces neutrophil recruitment in mouse models of acute lung injury. Crit. Care. 2012;16 doi: 10.1186/cc11518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fang X., Gao F., Zheng L., Xue F.S., Zhu T., Zheng X. Reduced microRNA-744 expression in mast cell-derived exosomes triggers epithelial cell ferroptosis in acute respiratory distress syndrome. Redox Biol. 2024;77 doi: 10.1016/j.redox.2024.103387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pernet E., Sun S., Sarden N., Gona S., Nguyen A., Khan N., Mawhinney M., Tran K.A., Chronopoulos J., Amberkar D., et al. Neonatal imprinting of alveolar macrophages via neutrophil-derived 12-HETE. Nature. 2023;614:530–538. doi: 10.1038/s41586-022-05660-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Kostenis E., Ulven T. Emerging roles of DP and CRTH2 in allergic inflammation. Trends Mol. Med. 2006;12:148–158. doi: 10.1016/j.molmed.2006.02.005. [DOI] [PubMed] [Google Scholar]
- 39.Jandl K., Stacher E., Bálint Z., Sturm E.M., Maric J., Peinhaupt M., Luschnig P., Aringer I., Fauland A., Konya V., et al. Activated prostaglandin D2 receptors on macrophages enhance neutrophil recruitment into the lung. J. Allergy Clin. Immunol. 2016;137:833–843. doi: 10.1016/j.jaci.2015.11.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hussain M., Xu C., Yao M., Zhang Q., Wu J., Wu X., Lu M., Tang L., Wu F., Wu X. CRTH2 antagonist, CT-133, effectively alleviates cigarette smoke-induced acute lung injury. Life Sci. 2019;216:156–167. doi: 10.1016/j.lfs.2018.11.039. [DOI] [PubMed] [Google Scholar]
- 41.Gupta A., Chander Chiang K. Prostaglandin D(2) as a mediator of lymphopenia and a therapeutic target in COVID-19 disease. Med. Hypotheses. 2020;143 doi: 10.1016/j.mehy.2020.110122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kan B., Razzaghian H.R., Lavoie P.M. An Immunological Perspective on Neonatal Sepsis. Trends Mol. Med. 2016;22:290–302. doi: 10.1016/j.molmed.2016.02.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zhang Y., Xie X., Huang Y., Liu M., Li Q., Luo J., He Y., Yin X., Ma S., Cao W., et al. Temporal molecular program of human hematopoietic stem and progenitor cells after birth. Dev. Cell. 2022;57:2745–2760.e6. doi: 10.1016/j.devcel.2022.11.013. [DOI] [PubMed] [Google Scholar]
- 44.Yu F., Sharma S., Jankovic D., Gurram R.K., Su P., Hu G., Li R., Rieder S., Zhao K., Sun B., Zhu J. The transcription factor Bhlhe40 is a switch of inflammatory versus antiinflammatory Th1 cell fate determination. J. Exp. Med. 2018;215:1813–1821. doi: 10.1084/jem.20170155. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zhu Q., Rui K., Wang S., Tian J. Advances of Regulatory B Cells in Autoimmune Diseases. Front. Immunol. 2021;12 doi: 10.3389/fimmu.2021.592914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sweet D.G., Carnielli V.P., Greisen G., Hallman M., Klebermass-Schrehof K., Ozek E., Te Pas A., Plavka R., Roehr C.C., Saugstad O.D., et al. European Consensus Guidelines on the Management of Respiratory Distress Syndrome: 2022 Update. Neonatology. 2023;120:3–23. doi: 10.1159/000528914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Terekhova M., Swain A., Bohacova P., Aladyeva E., Arthur L., Laha A., Mogilenko D.A., Burdess S., Sukhov V., Kleverov D., et al. Single-cell atlas of healthy human blood unveils age-related loss of NKG2C(+)GZMB(-)CD8(+) memory T cells and accumulation of type 2 memory T cells. Immunity. 2023;56:2836–2854.e9. doi: 10.1016/j.immuni.2023.10.013. [DOI] [PubMed] [Google Scholar]
- 48.Xie X., Liu M., Zhang Y., Wang B., Zhu C., Wang C., Li Q., Huo Y., Guo J., Xu C., et al. Single-cell transcriptomic landscape of human blood cells. Natl. Sci. Rev. 2021;8 doi: 10.1093/nsr/nwaa180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Yao M., Cao Y., He J., Dong R., Liu G., Chen Y., Wang J., Zhou J. Single-cell transcriptomic analysis reveals heterogeneous features of myeloid-derived suppressor cells in newborns. Front. Immunol. 2024;15 doi: 10.3389/fimmu.2024.1367230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Leon-Lara X., Fichtner A.S., Willers M., Yang T., Schaper K., Riemann L., Schoning J., Harms A., Almeida V., Schimrock A., et al. gammadelta T cell profiling in a cohort of preterm infants reveals elevated frequencies of CD83+ gammadelta T cells in sepsis. J. Exp. Med. 2024;221 doi: 10.1084/jem.20231987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Amatullah H., Maron-Gutierrez T., Shan Y., Gupta S., Tsoporis J.N., Varkouhi A.K., Teixeira Monteiro A.P., He X., Yin J., Marshall J.C., et al. Protective function of DJ-1/PARK7 in lipopolysaccharide and ventilator-induced acute lung injury. Redox Biol. 2021;38 doi: 10.1016/j.redox.2020.101796. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
-
•
Transcriptomic data are available in the SRA database under accession number [SRA: PRJNA1194327] (See https://www.ncbi.nlm.nih.gov/sra/?term=PRJNA1194327).
-
•
This paper does not report original code.
-
•
Requests for any additional information are available from the corresponding author on reasonable request.