Abstract
Background
Chemotherapy remains the primary treatment modality for patients with lung cancer; however, substantial inter-patient variability exists in responses to chemotherapeutic agents. Therefore, predicting individual responses is critical for optimizing treatment outcomes and improving patient prognosis.
Methods
This study developed a model to predict chemotherapy response in lung cancer patients by integrating multi-omics and clinical data from the Genomics of Drug Sensitivity in Cancer database, employing 45 machine learning algorithms. Data from the Gene Expression Omnibus database were utilized to validate the model. The impact of key genes on chemotherapy response was assessed in cell lines.
Results
A model combining random forest and support vector machine algorithms exhibited superior performance in both the training and validation sets. Furthermore, patients in the sensitive group demonstrated longer overall survival compared to those in the resistant group. TMED4 and DYNLRB1 genes were identified as pivotal features in the model and exhibited higher expression levels in the chemotherapy-resistant group. SiRNA-mediated knockdown of gene expression enhanced the chemosensitivity of lung cancer cell lines to chemotherapeutic agents.
Conclusions
This study successfully developed a high-performance machine learning model for predicting chemotherapy response in lung cancer and elucidated a strong correlation between TMED4 and DYNLRB1 gene expression and chemotherapy resistance. We further provide a user-friendly web server (available at https://smuonco.shinyapps.io/LC-DrugPortal/) to enable clinical utilization of our model, promoting personalized chemotherapy selection for lung cancer patients.
Keywords: Machine Learning, Chemosensitivity, Cell Line Validation, Lung Cancer, Prediction
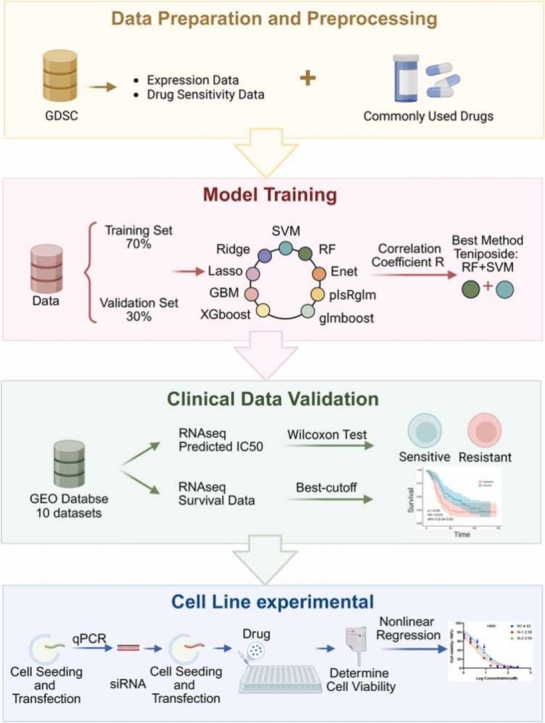
Graphical Abstract
1. Introduction
Lung cancer is a malignant neoplasm originating from the bronchial mucosal epithelium or lung parenchyma. As one of the most prevalent malignancies, lung cancer poses a significant threat to global public health[1]. Lung cancer is primarily classified into two major histological types: non-small cell lung cancer (NSCLC) and small cell lung cancer (SCLC)[2], [3], [4], [5]. NSCLC can be further subclassified into several histological subtypes, including adenocarcinoma, squamous cell carcinoma, and large cell carcinoma[5]. Globally, lung cancer consistently ranks among the malignancies with the highest incidence and mortality rates. According to World Health Organization statistics, in 2020, there were approximately 2.2 million new cases of lung cancer worldwide, with mortality reaching approximately 1.8 million cases[6]. Despite significant advancements in early screening methodologies and diagnostic techniques for lung cancer in recent years, the prognosis for patients with advanced-stage disease remains poor.
The therapeutic landscape for lung cancer patients encompasses multiple modalities, including surgical intervention, radiotherapy, systemic chemotherapy, molecularly targeted therapy, and immunotherapy[7], [8], [9], [10]. Surgical resection remains the preferred treatment for early-stage lung cancer patients, particularly when distant metastasis has not occurred. However, surgical intervention is primarily applicable to patients diagnosed with early-stage disease, which unfortunately constitutes only a minority of cases. Regrettably, the majority of patients are diagnosed at an advanced stage, often with metastatic dissemination already present. While radiotherapy remains a crucial treatment, it may cause a range of severe adverse effects, including fatigue, dermatological complications, and the risk of radiation-induced secondary malignancies. Molecularly targeted therapy depends on comprehensive genetic profiling of tumor cells; however, not all patients harbor actionable genetic alterations targetable by currently available therapeutic agents. Although immunotherapy represents a paradigm-shifting approach in lung cancer treatment, it may induce severe immune-related adverse events that require careful management[7], [11]. Consequently, for patients with advanced-stage lung cancer, therapeutic goals primarily focus on prolonging overall survival, improving progression-free survival, and enhancing quality of life while minimizing treatment-related toxicities.
Chemotherapy remains a cornerstone in lung cancer management, demonstrating not only a significant extension of patient survival but also an effective enhancement of quality of life[12], [13], [14]. Chemotherapeutic agents predominantly exert their antitumor effects by disrupting critical biological processes, including, but not limited to, DNA replication and protein synthesis in neoplastic cells[15], [16], [17], [18]. In contemporary clinical practice, the armamentarium of chemotherapeutic agents commonly employed in lung cancer treatment includes platinum-based compounds (e.g., cisplatin [DDP] and carboplatin), taxanes (e.g., paclitaxel and docetaxel), vinca alkaloids (e.g., vinorelbine), and antimetabolites (e.g., gemcitabine [GEM] and pemetrexed)[14]. Nevertheless, owing to substantial inter-individual biological variability, the therapeutic efficacy and adverse effects of chemotherapeutic agents exhibit considerable heterogeneity among patients. For example, doxorubicin, used in lung cancer treatment, predominantly exerts its antineoplastic effect by inhibiting topoisomerase II. However, in cases of topoisomerase II mutations, neoplastic cells may acquire resistance, thereby evading the cytotoxic effects of the drug[19]. These inter-individual disparities may arise from multiple factors, including genetic polymorphisms, tumor heterogeneity, variations in drug metabolism capacity, and heterogeneity in immune system function[20], [21]. As a result, accurately predicting patient responsiveness to specific chemotherapeutic agents has become a critical challenge in achieving personalized treatment paradigms.
Historically, the selection of chemotherapeutic agents has predominantly depended on the empirical judgment of clinical oncologists. However, this approach has significant limitations, including difficulties in accurately predicting individual patient sensitivity to specific drugs and inefficacy in preventing potentially unnecessary toxic reactions[22]. Given these challenges, accurately predicting patient sensitivity to specific chemotherapeutic agents has become a critical issue in achieving personalized treatment[23], [24], [25], [26], [27], [28]. The identification and application of biomarkers have substantially advanced research on predicting treatment sensitivity in lung cancer patients, establishing a vital foundation for implementing individualized therapy. Biomarkers provide clinicians with essential predictive information, aiding in the assessment of patients' potential responses to specific treatments, thereby enabling the development of optimized individualized treatment plans. In NSCLC, mutations in the epidermal growth factor receptor (EGFR) gene are recognized as among the most clinically relevant biomarkers. Patients harboring EGFR mutations typically exhibit significant therapeutic sensitivity to EGFR tyrosine kinase inhibitors, including gefitinib and erlotinib[29]. Therefore, developing high-precision prediction models and identifying reliable biomarkers not only enhances treatment efficacy and reduces unnecessary toxic reactions but also refines individualized treatment plans, thereby substantially improving patients' quality of life and prognosis. Recent studies have underscored the critical roles of Transmembrane P24 Trafficking Protein 4 (TMED4) and Dynein Light Chain Roadblock-Type 1 (DYNLRB1) in lung cancer pathogenesis. TMED4, a member of the α subfamily of TMED proteins, has been shown to be an important regulator of protein transport between the endoplasmic reticulum and the Golgi apparatus. A comprehensive review by Zhou et al. revealed that TMED family members, particularly TMED4, are involved in essential cellular processes including protein trafficking, cell signaling, and inflammatory responses. The study demonstrated that dysregulation of TMED proteins significantly contributes to malignant tumor development through multiple pathways[30]. DYNLRB1, a key component of the cytoplasmic dynein complex, plays essential roles in cellular transport processes and cell division. In lung cancer, DYNLRB1 is especially notable for its potential involvement in chemotherapy resistance mechanisms, particularly in response to platinum-based agents, through its regulation of intracellular trafficking and protein transport.
Machine learning, a computational approach that predicts outcomes through large-scale data mining and complex algorithmic analysis[31], has demonstrated significant potential in the precise and personalized selection of chemotherapeutic agents for lung cancer. This approach effectively integrates multidimensional biomedical data, including genomics, transcriptomics, proteomics, drug chemical structures, and drug response data, enabling accurate prediction of lung cancer patients' responses to various chemotherapeutic agents[32], [33], [34]. For instance, Jang et al. employed advanced deep learning models to analyze large-scale gene expression profiles and drug sensitivity data from lung cancer patients, successfully developing a high-performance model capable of accurately predicting NSCLC patients' responses to multiple chemotherapeutic agents[35]. This model demonstrated excellent predictive performance on independent external validation sets, providing robust support for clinical treatment decision-making. Numerous studies suggest that machine learning methods have the potential to significantly accelerate the screening of chemotherapeutic agents for lung cancer and the development of precise and personalized medication strategies, offering innovative approaches to enhance treatment efficacy and reduce adverse reaction rates[31], [36], [37]. However, existing models still exhibit substantial limitations in predictive accuracy, generalizability, and clinical applicability, necessitating further optimization.
This study aims to address this critical research gap by systematically and comprehensively investigating and integrating multi-dimensional omics data alongside rich clinical information to develop highly accurate and interpretable predictive models. Our objective is to devealop a highly reliable clinical decision support system that accelerates the widespread application of machine learning in precision and personalized treatment for lung cancer, ensuring patients receive optimized treatment regimens tailored to their individual biological characteristics. This innovative approach not only assists clinicians in making more precise and personalized treatment decisions but also may reveal potential, underutilized therapeutic opportunities, paving the way for novel research directions and treatment strategies in lung cancer therapy.
2. Methods
2.1. Data sources
RNA-seq data and drug IC50 values for cell lines were obtained from the Genomics of Drug Sensitivity in Cancer (GDSC) database[38], including both the GDSC1 and GDSC2 datasets, for model training purposes. Lung cancer-related cell lines were selected, and based on clinical expertise, commonly prescribed drugs for lung cancer treatment were chosen as training targets for the drug sensitivity model. These drugs included Cyclophosphamide, Dacarbazine, Dactinomycin, Irinotecan, Mitoxantrone, Oxaliplatin, Teniposide, Topotecan, 5-Fluorouracil (5-FU), Bleomycin, DDP, Cytarabine, Docetaxel, Doxorubicin, Etoposide (VP-16), GEM, Methotrexate, Mitomycin, Paclitaxel, Pemetrexed, Temozolomide, Vinblastine, and Vinorelbine. For model validation, 10 datasets were selected from the Gene Expression Omnibus (GEO) that contained both cell line RNA-seq data and drug sensitivity data. These datasets included GSE106609 [39], GSE116192[40], GSE162256 [41], GSE162285 [42], GSE163013 [43], GSE20185 [44], GSE37543 [45], [46], GSE58074 [47], [48], GSE78512 [49], and GSE85183 [50]. Furthermore, five datasets containing patient RNA-seq data, clinical medication information, and survival data were utilized for validating the model's survival prediction. These datasets comprised GSE25055 [51], [52], GSE26193 [53], [54], [55], [56], GSE69795 [57], GSE132342 [58], and GSE151335[59].
2.2. Model training and validation
This study incorporated 45 machine learning models, including Support Vector Machine (SVM), Ridge Regression, Random Forest (RF), Elastic Net (Enet), Least Absolute Shrinkage and Selection Operator (Lasso), Gradient Boosting Machine (GBM), Partial Least Squares Regression (plsRglm), and XGBoost, among others (see Table S1 for comprehensive details), to perform an initial screening of genes significantly correlated with IC50. The dataset was then randomly partitioned into training and validation sets at a 7:3 ratio. IC50 prediction models were constructed based on gene expression data using cross-validation methods, and the contribution of individual genes to model performance was evaluated. The correlation coefficient (R) between predicted and actual IC50 values was calculated for both the training and validation sets. The optimal prediction model for each drug was then selected based on the mean correlation coefficient (R) value.
2.3. Clinical data validation
Using the optimized prediction models for each chemotherapeutic agent, we estimated IC50 values based on cell line RNA-seq data obtained from the GEO database. Next, we applied the Wilcoxon rank-sum test to compare the differences in estimated IC50 values between drug-sensitive and drug-resistant cell groups. Using patient survival data and RNA-seq expression profiles from the GEO database, we applied the optimized prediction models for each chemotherapeutic agent to estimate patient-specific IC50 values. Then, we employed the optimal cutoff method to determine the threshold for stratifying patients into drug-sensitive and drug-resistant groups. Finally, we generated model-predicted Kaplan-Meier survival curves to analyze the survival differences between these groups.
2.4. Cell line experimental validation
This investigation utilized a panel of widely used chemotherapeutic agents, including DDP, VP-16, GEM, epirubicin (EPI), and 5-FU, to assess drug sensitivity across multiple lung cancer cell lines. The study employed three non-small cell lung cancer (NSCLC) cell lines: NCI-H460 (Pricella, cat: CL-0299), PC-9 (Pricella, cat: CL-0668), and NCI-H1299 (Pricella, cat: CL-0165), along with one small cell lung cancer (SCLC) cell line: NCI-H1339 (Beijing Zhongke Quality Inspection Network, cat: ZK1488). All cell lines were maintained in RPMI-1640 medium (KeyGEN BioTECH, cat: KGL-1501–500) supplemented with 10 % fetal bovine serum (Pricella, 164210) and 1 % penicillin/streptomycin. Cells were cultured at 37°C in a humidified atmosphere containing 5 % CO2. We initially seeded these cell lines at a density of 5 × 10^5 cells/well in six-well plates. Subsequently, these cells were transfected with siRNA targeting specific genes using PepMute™ siRNA transfection reagent. At 24 h post-transfection, total RNA was extracted using the EZBioscience RNA extraction kit following the manufacturer's protocol. The extraction process included cell lysis with Lysis Buffer, RNA binding to spin columns at 4000 g for 1 min, washing with 500 μL Wash Buffer at 12,000g for 1 min, and final elution with 20–30 μL Elution Buffer. RNA concentration and purity were measured using a NanoDrop spectrophotometer, with OD260/OD280 ratios between 1.9 and 2.1 indicating high RNA purity. The extracted RNA was then analyzed using real-time quantitative PCR (qPCR) to identify siRNA sequences exhibiting significant interference effects for use in subsequent experiments. The RT-qPCR was performed under the following conditions: pre-denaturation at 95°C for 3 min, followed by 39 cycles of denaturation (95°C for 10 s) and annealing/extension (60°C for 30 s). Following the identification of effective siRNA sequences, cells were reseeded at a density of 5 × 10^5 cells/well, and target gene knockdown was performed using PepMute™ siRNA transfection reagent. At 6 h post-transfection, the culture medium was replaced with complete medium supplemented with 10 % fetal bovine serum. At 12 h post-transfection, cells were transferred to 96-well plates and seeded at a density of 6 × 10^3 cells/well. Following complete cell adhesion (approximately 12–24 h), varying concentrations of chemotherapeutic agents (VP-16, 5-FU, EPI, DDP, and GEM) were applied to each well for a 48-hour drug exposure period. Post-drug exposure, cell viability was assessed using the Cell Counting Kit-8 colorimetric assay. Nonlinear regression analysis was employed to generate drug concentration-cell viability curves and calculate the half-maximal inhibitory concentration (IC50) for each experimental group. This approach allowed us to evaluate the impact of TMED4 gene silencing on chemotherapeutic drug sensitivity across various lung cancer cell lines.
2.5. Statistical analysis
Statistical analysis and data visualization in this study were primarily conducted using R software (version 4.3.0) and the Python programming language (version 3.9.5). We employed the nonparametric Wilcoxon rank-sum test to compare differences in predicted IC50 values between chemosensitive and chemoresistant cell groups. For data visualization, we utilized the ComplexHeatmap package[60] (version 2.8.0) to generate heatmaps, the circlize package[61] (version 0.4.13) for circular plots, and the ggplot2 package[62] (version 3.3.5) for box plots. Survival analysis was performed using the survival package[63] (version 3.2–13) and the survminer package[64] (version 0.4.9) to generate Kaplan-Meier survival curves and conduct log-rank tests. Additionally, we employed the pRRophetic package[65] (version 0.5) to calculate Pearson correlation coefficients between predicted and observed IC50 values. Cell line experimental data analysis was conducted using GraphPad Prism software (version 9.0), fitting drug concentration-cell viability curves through four-parameter logistic nonlinear regression analysis. Statistical significance was defined as a two-sided P value < 0.05, with all tests adjusted for multiple comparisons using the Benjamini-Hochberg procedure.
3. Results
3.1. Development of machine learning models for predicting lung cancer patients' sensitivity to common chemotherapy drugs
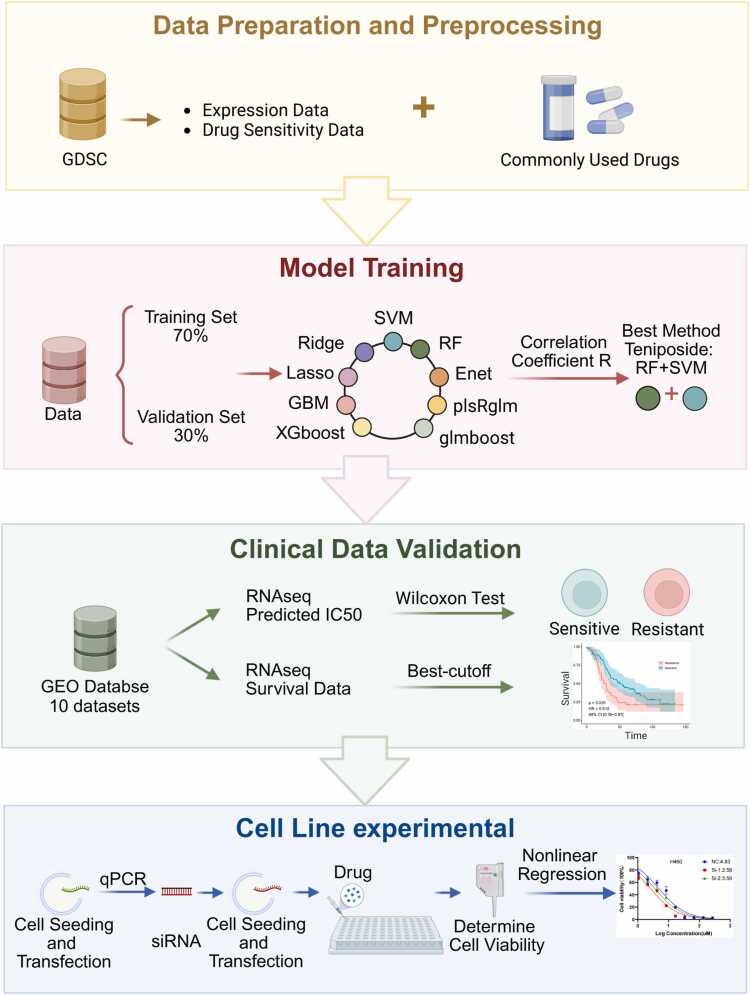
Fig. 1 illustrates the comprehensive experimental design and analytical workflow of this study. To address the challenge of personalized treatment in lung cancer chemotherapy, this study aimed to develop an integrative predictive model based on an ensemble of machine learning algorithms to accurately forecast lung cancer patients' responsiveness to standard-of-care chemotherapeutic agents. By integrating large-scale transcriptomic profiles and clinical data from public repositories, along with in vitro cell line experiments, this study not only validated the predictive performance of the constructed models but also identified key genetic biomarkers capable of robustly predicting chemotherapeutic efficacy.
Fig. 1.
Schematic overview of the development and validation of machine learning models for predicting chemotherapy drug sensitivity in lung cancer patients, and the exploration of the predictive value of TMED4 and DYNLRB1 genes in lung cancer chemotherapy response.
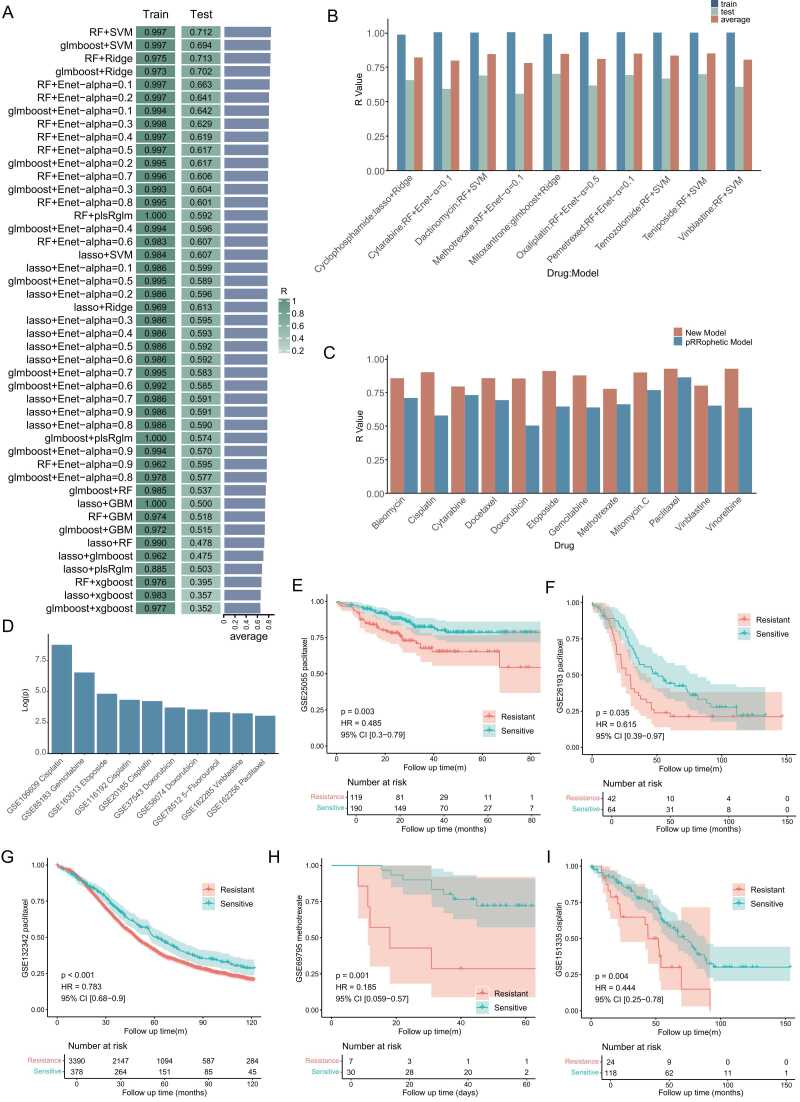
By evaluating Pearson correlation coefficients (R values) in both the training and validation sets, we identified models with optimal predictive performance. Among the 45 machine learning models evaluated, the combination model of RF+SVM exhibited the best predictive performance, with R values of 0.997, 0.972, and 0.854 for the training set, validation set, and their combination, respectively (Fig. 2A). For various chemotherapy drugs, the R values of their respective optimal models accurately reflected predictive performance in both the training and validation sets. The average R values for the top 10 models demonstrating the best predictive performance in both the training and validation sets were as follows: Teniposide: RF+SVM (R=0.846), Pemetrexed: RF+Enet-α= 0.1 (R=0.844), Mitoxantrone: glmboost+Ridge (R=0.842), Dactinomycin: RF+SVM (R=0.841), Temozolomide: RF+SVM (R=0.830), Cyclophosphamide: lasso+Ridge (R=0.817), Oxaliplatin: RF+Enet-α= 0.5 (R=0.806), Vinblastine: RF+SVM (R=0.800), Cytarabine: RF+Enet-α= 0.1 (R=0.794), and Methotrexate: RF+Enet-α= 0.1 (R=0.776) (Fig. 2B). These results indicate that the constructed models can effectively predict sensitivity to various chemotherapy drugs. Furthermore, we compared the predictive performance of our constructed models with that of the pRRophetic software package[65], a widely used tool in this field. The results revealed that for 12 commonly used chemotherapy drugs, including bleomycin, DDP, cytarabine, docetaxel, doxorubicin, VP-16, GEM, methotrexate, mitomycin C, paclitaxel, vinblastine, and vinorelbine, our constructed models exhibited higher correlations in predicting IC50 values (Fig. 2C). These findings provide compelling evidence for the superior predictive performance of the models developed in this study.
Fig. 2.
Evaluation of chemotherapy drug sensitivity prediction models. (A) Pearson correlation coefficients (r values) of the models in the training set, validation set, and their average. (B) R values of prediction models for the top 10 chemotherapy drugs based on the average of training and validation sets. (C) Comparison of r values between the newly developed model and the pRRophetic model for predicting the same drugs. (D) Log10(p) values of IC50 differences between predicted drug-sensitive and drug-resistant groups across multiple datasets, as determined by the Wilcoxon rank-sum test. (E-I) Kaplan-Meier survival curves for paclitaxel, methotrexate, and cisplatin, comparing overall survival between predicted drug-sensitive and drug-resistant groups across various datasets.
3.2. Significant prognostic differences exist between drug-sensitive and drug-resistant groups in lung cancer patients
This study employed the Wilcoxon test to evaluate differences in IC50 values between predicted drug-sensitive and drug-resistant cell groups. Statistically significant differences in IC50 values between predicted drug-sensitive and drug-resistant groups were observed across multiple datasets (p < 0.05). The analyzed datasets and their corresponding -log10(p-values) for IC50 differences were as follows: GSE106609 (8.770), GSE85183 (6.535), GSE163013 (4.818), GSE116192 (4.329), GSE20185 (4.226), GSE37543 (3.707), GSE58074 (3.555), GSE78512 (3.325), GSE162285 (3.239), and GSE162256 (3.043) (Fig. 2D). These findings provide further validation of the model's accuracy in predicting drug sensitivity. Subsequent survival analyses demonstrate that the drug sensitivity prediction model effectively stratifies patient prognosis across multiple independent datasets. Paclitaxel exhibited consistent prognostic trends across three independent datasets: GSE25055 (HR = 0.485, 95 % CI: 0.30–0.79, p = 0.003), GSE26193 (HR = 0.615, 95 % CI: 0.39–0.97, p = 0.035), and GSE132342 (HR = 0.783, 95 % CI: 0.68–0.90, p < 0.001) (Fig. 2E-G). Analysis of the GSE69795 dataset revealed that methotrexate-sensitive patients exhibited significantly higher survival rates compared to the resistant group (HR = 0.185, 95 % CI: 0.059–0.570, p = 0.001) (Fig. 2H). Similarly, analysis of the GSE151335 dataset showed that the DDP-sensitive group demonstrated superior survival outcomes compared to the resistant group (HR = 0.444, 95 % CI: 0.250–0.780, p = 0.004) (Fig. 2I). These results consistently demonstrate that patient cohorts predicted to be drug-sensitive exhibit significantly reduced mortality risk compared to drug-resistant groups across multiple chemotherapeutic agents. These findings not only substantiate the efficacy of drug sensitivity prediction models in identifying potentially responsive patient populations but also provide crucial evidence to support personalized treatment decisions in lung cancer therapy.
3.3. Exploring the association between TMED4 and DYNLRB1 genes and drug sensitivity
We conducted a comprehensive analysis of the predictive performance of the TMED4 and DYNLRB1 genes across multiple drug sensitivity prediction models. Our statistical analysis demonstrated that TMED4 and DYNLRB1 were consistently incorporated at the highest frequency across all evaluated prediction models. TMED4 was integrated into 17 machine learning models (Fig. 3A), and this high prevalence strongly indicates its potential as a biomarker for drug sensitivity. Similarly, DYNLRB1 was incorporated into 11 machine learning models (Fig. 4A), further substantiating its potential value in predicting drug sensitivity. Subsequent gene expression analysis elucidated the potential roles of DYNLRB1 and TMED4 in conferring resistance to multiple chemotherapeutic agents. Our comprehensive data analysis revealed that DYNLRB1 exhibited statistically significant expression differences between resistant and sensitive groups across a wide range of drugs. Of particular note, TMED4 demonstrated significant upregulation in resistant groups across an extensive array of drugs, including 5-FU, Bleomycin, DDP, Cyclophosphamide, Docetaxel, VP-16, GEM, Irinotecan, Methotrexate, Mitomycin-C, Mitoxantrone, Oxaliplatin, Paclitaxel, Pemetrexed, Temozolomide, Teniposide, Topotecan, and Vinblastine (Fig. 3A). Analogously, DYNLRB1 exhibited statistically significant upregulation in resistant groups across a substantial number of drugs, including Bleomycin, Cytarabine, Dactinomycin, Docetaxel, Doxorubicin, VP-16, GEM, Irinotecan, Mitomycin-C, Teniposide, Vinblastine, and Vinorelbine (Fig. 4A). Our in silico analysis identified TMED4 and DYNLRB1 as key features in multiple drug sensitivity models, with both genes consistently upregulated in resistant cell lines (e.g., TMED4 in cisplatin-resistant groups, Fig. 3A). Subsequent siRNA knockdown experiments confirmed their functional relevance, as silencing these genes enhanced chemosensitivity across multiple cell lines (Figs. 3C and 4C). While these findings suggest TMED4 and DYNLRB1 as potential biomarkers for chemotherapy resistance, further validation in patient-derived samples is necessary to assess their clinical utility. Detailed information on specific drug brands is provided in Supplementary Table S2.
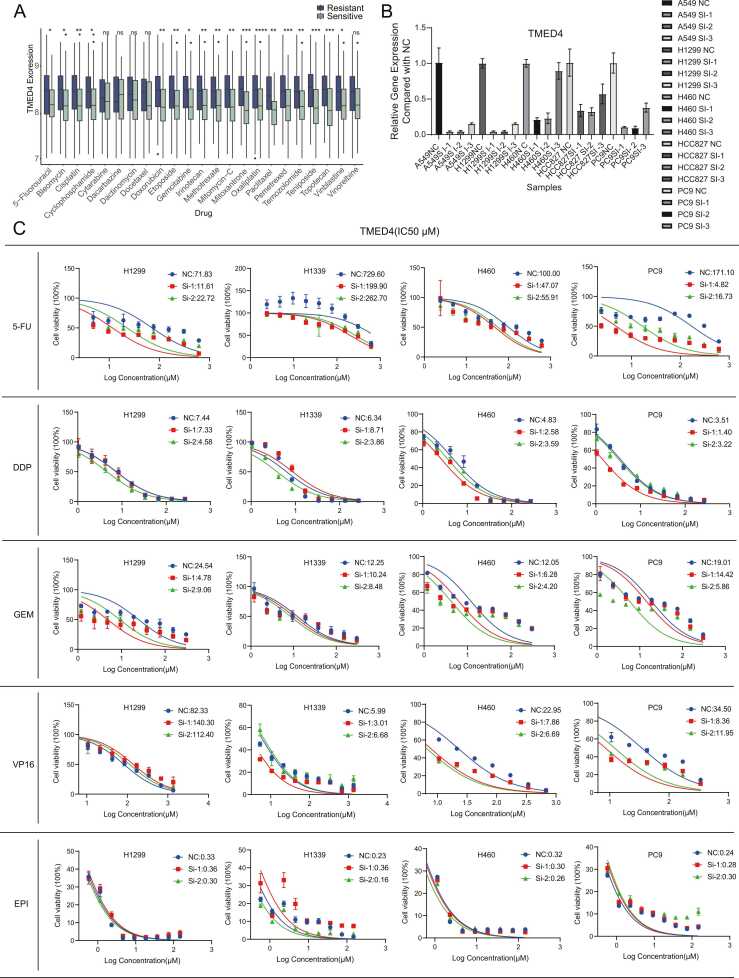
Fig. 3.
TMED4 expression in lung cancer cell lines and its impact on chemotherapy drug sensitivity. (A) TMED4 expression levels in drug-resistant and drug-sensitive cells for various lung cancer chemotherapy drugs. (B) Relative expression levels of TMED4 in five lung cancer cell lines (A549, H1299, H460, HCC827, and PC9) treated with three distinct siRNA sequences. (C) Dose-response curves for five chemotherapeutic agents (5-fluorouracil, cisplatin, gemcitabine, etoposide, and epirubicin) in TMED4 knockdown lung cancer cell lines.
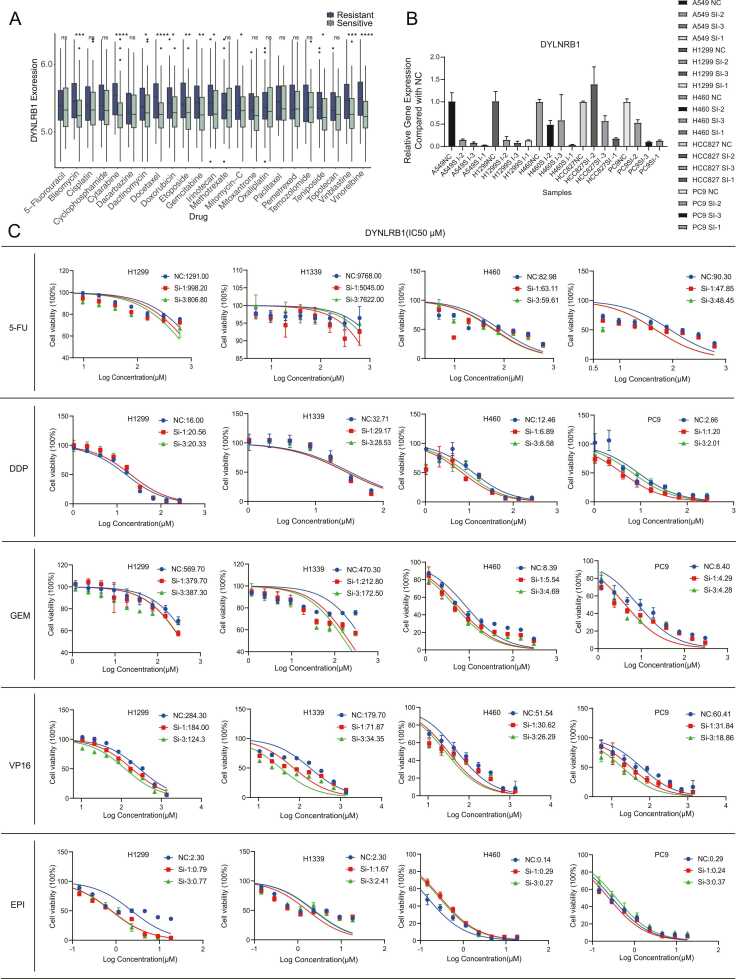
Fig. 4.
DYNLRB1 expression in lung cancer cell lines and its impact on chemotherapy drug sensitivity.cancer cell. (A) DYNLRB1 expression levels in drug-resistant and drug-sensitive cells for various lung cancer chemotherapeutic agents. (B) Relative expression levels of DYNLRB1 in five lung cancer cell lines (A549, H1299, H460, HCC827, and PC9) treated with three distinct siRNA sequences. (C) Dose-response curves for five chemotherapeutic agents (5-fluorouracil, cisplatin, gemcitabine, etoposide, and epirubicin) in DYNLRB1 knockdown lung cancer cell lines.
3.4. Enhanced sensitivity to chemotherapeutic agents in multiple lung cancer cell lines following TMED4 and DYNLRB1 gene knockdown
This study investigated the relative expression levels of TMED4 and DYNLRB1 genes in various lung cancer cell lines following transfection with distinct siRNAs. Experimental results revealed that all siRNA treatment groups (SI-1, SI-2, SI-3) induced significantly reduced TMED4 gene expression levels in lung cancer cell lines including A549, H1299, H460, HCC827, and PC9 (Fig. 3B). Simultaneously, DYNLRB1 gene expression demonstrated a marked decrease in A549, H1299, H460, and PC9 cell lines (Fig. 4B).
These findings suggest that siRNA transfection effectively downregulates the expression of TMED4 and DYNLRB1 genes, particularly in A549, H1299, H460, and PC9 cell lines. This establishes a reliable cellular model for subsequent studies investigating the impact of TMED4 and DYNLRB1 gene knockdown on cellular chemosensitivity.
Results from TMED4 gene silencing experiments demonstrated that TMED4 expression inhibition significantly enhanced cellular sensitivity to 5-FU in all four lung cancer cell lines examined. Among all tested cell lines, PC9 and H1299 demonstrated the most pronounced enhancement in 5-FU sensitivity, exhibiting 35.50-fold (from 171.10 to 4.82 µM) and 6.19-fold (from 71.83 to 11.61 µM) reductions in IC50 values respectively, while H460 and H1339 showed 2.12-fold (from 100.00 to 47.07 µM) and 3.65-fold (from 729.60 to 199.90 µM) decreases respectively (Fig. 3C). For DDP sensitivity, PC9 showed the strongest response with a 2.50-fold decrease (from 3.51 to 1.40 µM), followed by H460 and H1299 with 1.87-fold (from 4.83 to 2.58 µM) and 1.62-fold (from 7.44 to 4.58 µM) reductions, respectively. H1339 showed a 1.4-fold increase with Si-1 but a 1.64-fold decrease with Si-2 (from 6.34 to 3.86 µM) (Fig. 3C). In GEM experiments, H1299 exhibited the strongest response with a 5.1-fold reduction (from 24.54 to 4.78 µM), while H460 and PC9 showed 2.98-fold (from 12.05 to 4.20 µM) and 3.24-fold (from 19.01 to 5.86 µM) decreases, respectively. H1339 showed a 1.44-fold reduction (from 12.25 to 10.24 µM) (Fig. 3C). For VP-16, PC9, H460, and H1339 showed 4.13-fold, 3.43-fold, and 1.99-fold decreases in IC50 respectively, while H1299 displayed a 1.37-fold increase (from 82.33 to 112.40 µM) (Fig. 3C). EPI induced varied responses: H460 showed a 1.2-fold decrease (from 0.32 to 0.26 µM), while H1229 and H1339 showed increases of 1.09-fold and 1.56-fold with Si-1, but decreases of 1.09-fold and 1.43-fold with Si-2, respectively (Fig. 3C). Collectively, these results strongly suggest that the TMED4 gene plays a critical role in modulating the sensitivity of various lung cancer cell lines to commonly used chemotherapeutic agents. Inhibition of TMED4 expression predominantly enhanced cellular response to chemotherapeutic agents, although some heterogeneity was observed across different cell lines and drugs.
Analogous to the TMED4 gene, knockdown experiments targeting the DYNLRB1 gene revealed its critical role in modulating chemotherapeutic sensitivity. In all four examined lung cancer cell lines, DYNLRB1 knockdown elicited a statistically significant enhancement in sensitivity to VP16, 5-FU, and GEM. The effect of DYNLRB1 knockdown on DDP sensitivity demonstrated heterogeneity across the cell lines examined: H1339, H460, and PC9 cells exhibited increased sensitivity to DDP following knockdown, while the H1299 cell line displayed a marginal increase in resistance (Fig. 4C). The response to EPI following DYNLRB1 knockdown manifested a cell line-specific pattern: H1339 and H460 cells exhibited enhanced resistance, whereas H1299 and PC9 cells demonstrated increased sensitivity (Fig. 4C). In conclusion, these DYNLRB1 knockdown experiments provide compelling evidence for the pivotal role of this gene in modulating cellular response to chemotherapeutic agents, while also revealing drug- and cell line-specific modulatory effects.
4. Discussion
In the contemporary landscape of lung cancer therapeutics research, the development of personalized treatment strategies has become a paramount and increasingly critical focus. Specifically, in the context of chemotherapeutic agent selection, the ability to predict drug responses based on patient-specific gene expression profiles is essential for optimizing treatment efficacy while minimizing adverse effects. Recent studies have demonstrated that this challenge can be significantly addressed through the integration of multi-source datasets and the application of advanced machine learning algorithms[66], [67], [68]. This study systematically collected and integrated cell line data from the GDSC database and clinical sample data from the GEO database to develop and implement multiple machine learning algorithms for predicting the sensitivity of lung cancer cells to standard-of-care chemotherapeutic agents. Subsequently, we validated the model's performance using survival data and in vitro cell line experiments, further elucidating the impact of gene expression profiles on cellular chemosensitivity. Through this comprehensive investigation, we not only validated the predictive accuracy of our model but also systematically delineated the influence of gene expression patterns on chemotherapeutic responses. The significance of this research lies in its potential to provide clinicians with more precise guidance for chemotherapeutic agent selection, thereby facilitating the development of more personalized treatment regimens that may enhance therapeutic efficacy and mitigate treatment-associated risks. Moreover, the findings of this study identify potential biomarkers for future drug development and clinical trial design, promising to further advance the implementation of precision medicine in lung cancer therapeutics.
This study's findings demonstrate that drug sensitivity prediction models, based on multiple machine learning algorithms, exhibit significant accuracy and substantial clinical application potential in predicting chemotherapeutic drug sensitivity for lung cancer. Among the 45 machine learning models evaluated, the RF+SVM combination model exhibited the highest correlation between predicted and observed IC50 values in both the training (R=0.997) and validation sets (R=0.972). However, the performance in the combined dataset (R=0.854) indicates potential limitations in generalizability, possibly due to dataset heterogeneity or overfitting during the training phase. This high-performance prediction model provides clinicians with a robust decision support tool, potentially significantly enhancing the development of personalized treatment plans for lung cancer patients. Further validation through survival analysis revealed a significant correlation between drug sensitivity prediction results and actual patient survival rates. In validation across multiple independent datasets, patient groups predicted to be drug-sensitive exhibited a significantly reduced mortality risk compared to drug-resistant groups (p < 0.05). For instance, in the GSE69795 dataset, patients in the methotrexate-sensitive group demonstrated significantly higher survival rates than those in the resistant group (hazard ratio [HR] = 0.185, 95 % confidence interval [CI]: 0.059–0.57, p = 0.001). This finding not only further validates the clinical relevance of the constructed model but also provides crucial empirical evidence for developing precise, personalized treatment strategies in lung cancer.
In the current landscape of lung cancer treatment research, the identification and application of reliable biomarkers are of paramount importance for optimizing therapeutic regimens[69]. In particular, within the realms of chemotherapeutic agent selection and personalized treatment strategy formulation, the ability to predict drug responses based on gene expression profiles has emerged as a critical factor in enhancing therapeutic efficacy and mitigating adverse reactions[70], [71]. Recent research trends have focused on developing high-precision predictive models by integrating preclinical and clinical data[72], [73]. Compared to existing approaches like pRRophetic[65], our study provides three key advancements: (1) integration of 45 machine learning algorithms instead of single-model approaches, (2) experimental validation of TMED4/DYNLRB1 roles in chemoresistance, and (3) development of a clinical decision support web server (available at https://smuonco.shinyapps.io/LC-DrugPortal/). This multi-faceted strategy addresses the critical gap between computational prediction and clinical implementation highlighted in recent reviews[66]. Our study contributes novel insights and potential applications to this evolving field, particularly in understanding the role of TMED family genes that have been widely implicated in lung cancer development and therapeutic responses[30]. Our investigation centers on two genes: TMED4 and DYNLRB1. TMED4 was incorporated into 17 machine learning models, while DYNLRB1 was included in 11 models, indicating their potential as robust biomarkers for personalized lung cancer treatment. These two genes exhibit a significant modulatory effect on the sensitivity of lung cancer cells to various common chemotherapeutic agents, particularly in the expression of drug-resistant phenotypes. Notably, TMED4 exhibited significant upregulation in drug-resistant groups for most agents, including 5-FU, DDP, and Docetaxel. Similarly, DYNLRB1 demonstrated significant upregulation in drug-resistant groups for various agents, including Bleomycin, Cytarabine, and Docetaxel. Upon individual knockdown of these two genes in cell lines, the cells exhibited enhanced sensitivity to chemotherapeutic agents compared to the control group, underscoring the pivotal roles of TMED4 and DYNLRB1 in modulating the chemotherapeutic response of lung cancer cells. From a clinical perspective, novel diagnostic methods can be developed based on the expression profiles of these genes to predict patient responses to specific chemotherapeutic agents. This approach can not only aid clinicians in selecting optimal treatment regimens prior to therapy but may also reduce time and resource expenditure resulting from ineffective treatments. For instance, by quantifying the expression levels of TMED4 and DYNLRB1 in tumor biopsy specimens, clinicians may more accurately evaluate patient resistance to specific chemotherapeutic agents and subsequently select more efficacious alternative therapies. Future investigations should focus on elucidating the molecular mechanisms by which these genes modulate the effects of chemotherapeutic agents, as well as characterizing their differential expression across various lung cancer subtypes and diverse patient populations. Moreover, studies should explore the existence of additional genes with comparable predictive potential and investigate how interactions among these genes orchestrate drug responses. Through these comprehensive investigations, we can enhance our understanding of the intricate landscape of lung cancer treatment and develop more precise and personalized therapeutic strategies for patients. To facilitate the clinical translation of our predictive model, we have developed an open-access web server (available at https://smuonco.shinyapps.io/LC-DrugPortal/). This comprehensive interactive platform offers several key advantages: it provides seamless access to curated drug sensitivity data for over 300 chemotherapy agents across lung cancer subtypes without requiring programming expertise; features an intuitive interface with real-time filtering capabilities that enable rapid identification of relevant drug-cancer combinations; delivers standardized sensitivity metrics in downloadable formats for further analysis; and ensures reproducible research through consistent data presentation and query methodologies. By providing this robust, user-friendly resource, we aim to bridge the gap between computational predictions and real-world clinical decision-making, ultimately accelerating the translation of pharmacogenomic insights into personalized lung cancer treatment strategies.
Despite significant advancements in predicting chemotherapy drug sensitivity for lung cancer, this study acknowledges several notable limitations. The primary challenge arises from the inherent characteristics of the data. Although this study utilized large-scale datasets from public databases such as GDSC and GEO, these datasets may inherently be limited by issues such as insufficient sample size and heterogeneity. These dataset characteristics may affect the model's ability to generalize its predictive results, thereby limiting its clinical applicability to a broader population. Consequently, future research should focus on increasing sample sizes and validating the model's practicality and reliability through multicenter clinical trials, which are essential for the model's further development. Moreover, although the model developed in this study demonstrates excellent predictive performance, it may face the challenge of the "black box" effect, a common issue in many machine learning models. The lack of transparency in the model's decision-making process impedes the interpretation of how the model derives specific predictive outcomes. This opacity may affect clinicians' trust and willingness to adopt the model; therefore, improving model interpretability will be a critical focus for future research. Finally, extrapolating from cell line data to actual patient populations presents a potential challenge. Although the model developed in this study demonstrates robust performance on cell line data, clinical scenarios in actual patients often exhibit greater complexity. Factors such as the tumor microenvironment and individual patient heterogeneity may significantly influence prediction accuracy. Therefore, more comprehensive and in-depth validation and optimization are required before translating research findings into clinical practice[74], [75]. While these limitations impose constraints on the current study to some extent, they also provide valuable guidance for future research directions. By systematically addressing these issues, we anticipate substantially enhancing the model's accuracy and clinical utility, ultimately enabling the development of more precise personalized treatment strategies for lung cancer patients.
5. Conclusions
This comprehensive study integrated large-scale datasets from the GDSC and GEO databases and applied state-of-the-art machine learning algorithms. Through rigorous evaluation using the Pearson correlation coefficient (R-value), we identified the optimal combination model comprising RF and SVM algorithms from a pool of 45 machine learning models to accurately predict lung cancer patients' sensitivity to chemotherapeutic agents. Subsequently, we conducted stringent validation of the top 10 performing models using independent datasets sourced from the GEO database. Our analysis revealed that specific genes, notably TMED4 and DYNLRB1, exhibit significant predictive value for lung cancer chemotherapy response. The expression levels of these genes demonstrated robust correlations with chemotherapy drug sensitivity, with particularly pronounced associations observed in resistant phenotypes. In summary, our study demonstrates that machine learning models integrating multi-omics data can predict chemotherapy sensitivity, with TMED4 and DYNLRB1 emerging as promising resistance biomarkers. Despite these promising findings, several limitations should be acknowledged. Our study primarily relied on cell line data, which may not fully capture the complexity of clinical scenarios. The interpretability of machine learning models remains a challenge for clinical implementation. Future studies should focus on prospective clinical validation in larger patient cohorts, further investigation of the molecular mechanisms underlying TMED4- and DYNLRB1-mediated chemoresistance, and the development of more interpretable prediction models incorporating additional clinical parameters. Moving forward, the ongoing refinement of predictive models for lung cancer chemotherapy drug sensitivity, coupled with comprehensive validation and mechanistic studies of pertinent biomarkers, will provide critical support for the development of precision treatment strategies and innovative drug discovery for lung cancer patients, ultimately enhancing patient survival rates and quality of life.
List of Abbreviations
- NSCLC
non-small cell lung cancer
- SCLC
small cell lung cancer
- EGFR
epidermal growth factor receptor
- GDSC
Genomics of Drug Sensitivity in Cancer
- GEO
Gene Expression Omnibus
- DDP
cisplatin
- VP-16
etoposide
- GEM
gemcitabine
- EPI
epirubicin
- 5-FU
5-fluorouracil
- IC50
half-maximal inhibitory concentration
- SVM
support vector machine
- RF
random forest
- Enet
elastic net
- Lasso
least absolute shrinkage and selection operator
- GBM
gradient boosting machine
- plsRglm
partial least squares regression
- TMED4
transmembrane P24 trafficking protein 4
- DYNLRB1
dynein light chain roadblock-type 1
CRediT authorship contribution statement
Peng Luo: Writing – review & editing, Supervision, Conceptualization, Methodology. Ting Wei: Writing – review & editing, Supervision, Conceptualization, Methodology. Jinghong Chen: Writing – original draft, Visualization, Data curation. Yonglin Yi: Visualization, Data curation, Writing – original draft. Haoxuan Ying: Writing – review & editing. Chunqian Yang: Writing – original draft, Visualization, Data curation. Anqi Lin: Writing – review & editing, Supervision, Conceptualization, Methodology. Jian Zhang: Writing – review & editing.
Authors' contributions
Peng Luo, Ting Wei, and Anqi Lin contributed to the conception, manuscript design, and supervision. Jinghong Chen, Yonglin Yi, and Chunqian Yang curated data, drafted the main manuscript, and prepared the visualization. Jinghong Chen built the web server. Jinghong Chen, Haoxuan Ying, Jian Zhang, Anqi Lin, Ting Wei, and Peng Luo participated in the revision of the manuscript. All authors read and approved the final manuscript.
Consent for publication
Not applicable.
Ethical approval
This study was approved by the Ethics Committee of Zhujiang Hospital, Southern Medical University, and all experimental procedures were reviewed and approved, with experiments conducted in strict compliance with established ethical guidelines.
Funding
Not applicable.
Declaration of Competing Interest
All authors declare no competing financial or non-financial interests.
Acknowledgments
Not applicable.
Footnotes
Supplementary data associated with this article can be found in the online version at doi:10.1016/j.csbj.2025.07.043.
Contributor Information
Anqi Lin, Email: smulinanqi0206@i.smu.edu.cn.
Ting Wei, Email: weitingyouyou@qq.com.
Peng Luo, Email: luopeng@smu.edu.cn, luopeng@hku.hk.
Appendix A. Supplementary material
Supplementary material
Table S2. Concentrations and sources of chemotherapeutic agents used in cell line studies
Data Availability
The data used in this study are available from the following public repositories: Genomics of Drug Sensitivity in Cancer (GDSC) database (https://www.cancerrxgene.org/), Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). All other data generated and analyzed in this study are available from the corresponding authors upon reasonable request.
References
- 1.Travis W.D., Brambilla E., Nicholson A.G., et al. The 2015 World Health Organization classification of lung tumors: impact of genetic, clinical and radiologic advances since the 2004 classification. J Thorac Oncol. 2015;10:1243–1260. doi: 10.1097/JTO.0000000000000630. [DOI] [PubMed] [Google Scholar]
- 2.Li Y., Yi Y., Lin A., et al. A comparison of the efficacy of antiangiogenic agents combined with chemotherapy for the treatment of non-small cell lung cancer: a network meta-analysis. Cancer Cell Int. 2020;20:1–18. doi: 10.1186/s12935-020-01639-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Yi Y., Qiu Z., Yao Z., et al. CAMSAP1 mutation correlates with improved prognosis in small cell lung cancer patients treated with platinum-based chemotherapy. Front Cell Dev Biol. 2022;9 doi: 10.3389/fcell.2021.770811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Lin A., Zhou N., Zhu W., et al. Genomic and immunological profiles of small-cell lung cancer between East Asians and Caucasian. Cancer Cell Int. 2022;22:173. doi: 10.1186/s12935-022-02588-w. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zappa C., Mousa S.A. Non-small cell lung cancer: current treatment and future advances. Transl lung Cancer Res. 2016;5:288. doi: 10.21037/tlcr.2016.06.07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sung H., Ferlay J., Siegel R.L., et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA a cancer journal clinicians. 2021;71:209–249. doi: 10.3322/caac.21660. [DOI] [PubMed] [Google Scholar]
- 7.Ramalingam S.S., Owonikoko T.K., Khuri F.R. Lung cancer: New biological insights and recent therapeutic advances. CA a cancer journal clinicians. 2011;61:91–112. doi: 10.3322/caac.20102. [DOI] [PubMed] [Google Scholar]
- 8.Yang S.-R., Schultheis A.M., Yu H., et al. Seminars in cancer biology. Elsevier; 2022. Precision medicine in non-small cell lung cancer: Current applications and future directions; pp. 184–198. [DOI] [PubMed] [Google Scholar]
- 9.Li G., Wan Y., Jiao A., et al. Breaking Boundaries: Chronic Diseases and the Frontiers of Immune Microenvironments. Med Res. 2025;1:62–102. [Google Scholar]
- 10.Fang Y., Kong Y., Rong G., et al. Systematic investigation of tumor microenvironment and antitumor immunity with IOBR. Med Res. 2025;1:136–140. [Google Scholar]
- 11.Gao Z., Jiang A., Li Z., et al. Heterogeneity of intratumoral microbiota within the tumor microenvironment and relationship to tumor development. Med Res. 2025;1:32–61. [Google Scholar]
- 12.Lin A., Zhu L., Jiang A., et al. Activation of the TGF-β pathway enhances the efficacy of platinum-based chemotherapy in small cell lung cancer patients. Dis Markers. 2022;2022 doi: 10.1155/2022/8766448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Lin A., Yang H., Zhang J., et al. CD209 signaling pathway as a biomarker for cisplatin chemotherapy response in small cell lung cancer. Genes Dis. 2024;11 doi: 10.1016/j.gendis.2023.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Li Z., Lin A., Gao Z., et al. B-cell performance in chemotherapy: unravelling the mystery of B-cell therapeutic potential. Clin Transl Med. 2024;14 doi: 10.1002/ctm2.1761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Ye T., Lin A., Qiu Z., et al. Microsatellite instability states serve as predictive biomarkers for tumors chemotherapy sensitivity. Iscience. 2023;26 doi: 10.1016/j.isci.2023.107045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Shi C., Qin K., Lin A., et al. The role of DNA damage repair (DDR) system in response to immune checkpoint inhibitor (ICI) therapy. J Exp Clin Cancer Res. 2022;41:268. doi: 10.1186/s13046-022-02469-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Patnaik S.K., Swaroop A.K., Nagarjuna P., et al. Peptides for dual targeting of ErbB1 and ErbB2: Blocking EGFR cell signaling transduction pathways for Cancer chemotherapy. Curr Mol Pharmacol. 2024;17 doi: 10.2174/1874467216666230224104950. [DOI] [PubMed] [Google Scholar]
- 18.Mirzaei S., Gholami M.H., Zabolian A., et al. Resveratrol augments doxorubicin and cisplatin chemotherapy: a novel therapeutic strategy. Curr Mol Pharmacol. 2023;16:280–306. doi: 10.2174/1874467215666220415131344. [DOI] [PubMed] [Google Scholar]
- 19.Chen A.Y., Liu L.F. In: Anticancer Drug Resistance: Advances in Molecular and Clinical Research. Goldstein L.J., Ozols R.F., editors. Springer US; Boston, MA: 1994. Mechanisms of resistance to topoisomerase inhibitors; pp. 263–281. [Google Scholar]
- 20.Gridelli C., Rossi A., Carbone D.P., et al. Non-small-cell lung cancer. Nat Rev Dis Prim. 2015;1:1–16. doi: 10.1038/nrdp.2015.9. [DOI] [PubMed] [Google Scholar]
- 21.Mansoori B., Mohammadi A., Davudian S., et al. The different mechanisms of cancer drug resistance: a brief review. Adv Pharm Bull. 2017;7:339. doi: 10.15171/apb.2017.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Schiller J.H., Harrington D., Belani C.P., et al. Comparison of four chemotherapy regimens for advanced non–small-cell lung cancer. N Engl J Med. 2002;346:92–98. doi: 10.1056/NEJMoa011954. [DOI] [PubMed] [Google Scholar]
- 23.Lin A., Mou W., Zhu L., et al. Mutations in the DNA polymerase binding pathway affect the immune microenvironment of patients with small-cell lung cancer and enhance the efficacy of platinum-based chemotherapy. Cancer Innov. 2023;2:500–512. doi: 10.1002/cai2.84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Luo P., Lin A., Li K., et al. DDR pathway alteration, tumor mutation burden, and cisplatin sensitivity in small cell lung cancer: difference detected by whole exome and targeted gene sequencing. J Thorac Oncol. 2019;14:e276–e279. doi: 10.1016/j.jtho.2019.08.2509. [DOI] [PubMed] [Google Scholar]
- 25.Liu Y., Zhang S., Liu K., et al. Advances in drug discovery based on network pharmacology and omics technology. Curr Pharm Anal. 2024;21:33–43. [Google Scholar]
- 26.Wang Z., Zhao Y., Zhang L. Emerging trends and hot topics in the application of multi-omics in drug discovery: a bibliometric and visualized study. Curr Pharm Anal. 2024;21:20–32. [Google Scholar]
- 27.Chen J., Lin A., Luo P. Advancing pharmaceutical research: a comprehensive review of cutting-edge tools and technologies. Curr Pharm Anal. 2024;21:1–19. [Google Scholar]
- 28.Liu L., Liang L., Luo Y., et al. Unveiling the power of gut microbiome in predicting neoadjuvant immunochemotherapy responses in esophageal squamous cell carcinoma. Research. 2024;7 doi: 10.34133/research.0529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kobayashi S., Boggon T.J., Dayaram T., et al. EGFR mutation and resistance of non–small-cell lung cancer to gefitinib. N Engl J Med. 2005;352:786–792. doi: 10.1056/NEJMoa044238. [DOI] [PubMed] [Google Scholar]
- 30.Zhou L., Li H., Yao H., et al. TMED family genes and their roles in human diseases. Int J Med Sci. 2023;20:1732. doi: 10.7150/ijms.87272. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhang N., Zhang H., Liu Z., et al. An artificial intelligence network-guided signature for predicting outcome and immunotherapy response in lung adenocarcinoma patients based on 26 machine learning algorithms. Cell Prolif. 2023;56 doi: 10.1111/cpr.13409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hassane M. Artificial intelligence-driven precision medicine in cancer treatment. Science. 2024;1 [Google Scholar]
- 33.Qureshi R., Basit S.A., Shamsi J.A., et al. Machine learning based personalized drug response prediction for lung cancer patients. Sci Rep. 2022;12 doi: 10.1038/s41598-022-23649-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kourou K., Exarchos T.P., Exarchos K.P., et al. Machine learning applications in cancer prognosis and prediction. Comput Struct Biotechnol J. 2015;13:8–17. doi: 10.1016/j.csbj.2014.11.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Jang I.S., Neto E.C., Guinney J., et al. Biocomputing 2014. World Scientific; 2014. Systematic assessment of analytical methods for drug sensitivity prediction from cancer cell line data; pp. 63–74. [PMC free article] [PubMed] [Google Scholar]
- 36.Zhou C., Peng S., Lin A., et al. Psychiatric disorders associated with immune checkpoint inhibitors: a pharmacovigilance analysis of the FDA Adverse Event Reporting System (FAERS) database. eClinicalMedicine. 2023;59 doi: 10.1016/j.eclinm.2023.101967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Gu T., Jiang A., Zhou C., et al. Adverse reactions associated with immune checkpoint inhibitors and bevacizumab: a pharmacovigilance analysis. Int J Cancer. 2023;152:480–495. doi: 10.1002/ijc.34332. [DOI] [PubMed] [Google Scholar]
- 38.Yang W., Soares J., Greninger P., et al. Genomics of Drug Sensitivity in Cancer (GDSC): a resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Res. 2013;41:D955–D961. doi: 10.1093/nar/gks1111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Musiani D., Giambruno R., Massignani E., et al. PRMT1 is recruited via DNA-PK to chromatin where it sustains the senescence-associated secretory phenotype in response to cisplatin. Cell Rep. 2020;30:1208–1222. doi: 10.1016/j.celrep.2019.12.061. . e1209. [DOI] [PubMed] [Google Scholar]
- 40.Statello L., Ali M.M., Reischl S., et al. The DNA damage inducible lncRNA SCAT7 regulates genomic integrity and topoisomerase 1 turnover in lung adenocarcinoma. NAR Cancer. 2021;3 doi: 10.1093/narcan/zcab002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wang X., Frederick J., Wang H., et al. Spike-in normalization for single-cell RNA-seq reveals dynamic global transcriptional activity mediating anticancer drug response. NAR Genom Bioinforma. 2021;3 doi: 10.1093/nargab/lqab054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Dhimolea E., de Matos Simoes R., Kansara D., et al. An embryonic diapause-like adaptation with suppressed Myc activity enables tumor treatment persistence. Cancer Cell. 2021;39:240–256. doi: 10.1016/j.ccell.2020.12.002. . e211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Hernandez V.A., Carvajal-Moreno J., Papa J.L., et al. CRISPR/Cas9 Genome Editing of the Human Topoisomerase IIα Intron 19 5′ Splice Site Circumvents Etoposide Resistance in Human Leukemia K562 Cells. Mol Pharmacol. 2021;99:226–241. doi: 10.1124/molpharm.120.000173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Poth K.J., Guminski A.D., Thomas G.P., et al. Cisplatin treatment induces a transient increase in tumorigenic potential associated with high interleukin-6 expression in head and neck squamous cell carcinoma. Mol Cancer Ther. 2010;9:2430–2439. doi: 10.1158/1535-7163.MCT-10-0258. [DOI] [PubMed] [Google Scholar]
- 45.Grinde M.T., Hilmarsdottir B., Tunset H.M., et al. Glutamine to proline conversion is associated with response to glutaminase inhibition in breast cancer. Breast Cancer Res. 2019;21:1–13. doi: 10.1186/s13058-019-1141-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Borgan E., Lindholm E.M., Moestue S., et al. Subtype-specific response to bevacizumab is reflected in the metabolome and transcriptome of breast cancer xenografts. Mol Oncol. 2013;7:130–142. doi: 10.1016/j.molonc.2012.10.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Hazar-Rethinam M., de Long L.M., Gannon O.M., et al. A novel E2F/sphingosine kinase 1 axis regulates anthracycline response in squamous cell carcinoma. Clin Cancer Res. 2015;21:417–427. doi: 10.1158/1078-0432.CCR-14-1962. [DOI] [PubMed] [Google Scholar]
- 48.Hazar-Rethinam M., de Long L.M., Gannon O.M., et al. RacGAP1 is a novel downstream effector of E2F7-dependent resistance to doxorubicin and is prognostic for overall survival in squamous cell carcinoma. Mol Cancer Ther. 2015;14:1939–1950. doi: 10.1158/1535-7163.MCT-15-0076. [DOI] [PubMed] [Google Scholar]
- 49.Qu Z., Cui J., Harata-Lee Y., et al. Identification of candidate anti-cancer molecular mechanisms of Compound Kushen Injection using functional genomics. Oncotarget. 2016;7:66003. doi: 10.18632/oncotarget.11788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Welch J.D., Williams L.A., DiSalvo M., et al. Selective single cell isolation for genomics using microraft arrays. Nucleic Acids Res. 2016;44:8292–8301. doi: 10.1093/nar/gkw700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hatzis C., Pusztai L., Valero V., et al. A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA. 2011;305:1873–1881. doi: 10.1001/jama.2011.593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Baldasici O., Balacescu L., Cruceriu D., et al. Circulating small EVs miRNAs as predictors of pathological response to neo-adjuvant therapy in breast cancer patients. Int J Mol Sci. 2022;23:12625. doi: 10.3390/ijms232012625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Gentric G., Kieffer Y., Mieulet V., et al. PML-regulated mitochondrial metabolism enhances chemosensitivity in human ovarian cancers. Cell Metab. 2019;29:156–173. doi: 10.1016/j.cmet.2018.09.002. . e110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Mateescu B., Batista L., Cardon M., et al. miR-141 and miR-200a act on ovarian tumorigenesis by controlling oxidative stress response. Nat Med. 2011;17:1627–1635. doi: 10.1038/nm.2512. [DOI] [PubMed] [Google Scholar]
- 55.Kieffer Y., Bonneau C., Popova T., et al. Clinical interest of combining transcriptomic and genomic signatures in high-grade serous ovarian cancer. Front Genet. 2020;11:219. doi: 10.3389/fgene.2020.00219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Mieulet V., Garnier C., Kieffer Y., et al. Stiffness increases with myofibroblast content and collagen density in mesenchymal high grade serous ovarian cancer. Sci Rep. 2021;11:4219. doi: 10.1038/s41598-021-83685-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.McConkey D.J., Choi W., Shen Y., et al. A prognostic gene expression signature in the molecular classification of chemotherapy-naive urothelial cancer is predictive of clinical outcomes from neoadjuvant chemotherapy: a phase 2 trial of dose-dense methotrexate, vinblastine, doxorubicin, and cisplatin with bevacizumab in urothelial cancer. Eur Urol. 2016;69:855–862. doi: 10.1016/j.eururo.2015.08.034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Chakraborty S., Hosen M.I., Ahmed M., et al. Onco-multi-OMICS approach: a new frontier in cancer research. Biomed Res Int. 2018;2018 doi: 10.1155/2018/9836256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Hu Z., Cunnea P., Zhong Z., et al. The Oxford classic links epithelial-to-mesenchymal transition to immunosuppression in poor prognosis ovarian cancers. Clin Cancer Res. 2021;27:1570–1579. doi: 10.1158/1078-0432.CCR-20-2782. [DOI] [PubMed] [Google Scholar]
- 60.Gu Z., Eils R., Schlesner M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics. 2016;32:2847–2849. doi: 10.1093/bioinformatics/btw313. [DOI] [PubMed] [Google Scholar]
- 61.Gu Z., Gu L., Eils R., et al. circlize implements and enhances circular visualization in R. Bioinformatics. 2014;30:2811–2812. doi: 10.1093/bioinformatics/btu393. [DOI] [PubMed] [Google Scholar]
- 62.Wickham H. A layered grammar of graphics. J Comput Graph Stat. 2010;19:3–28. [Google Scholar]
- 63.Lin H., Zelterman D. Modeling survival data: extending the cox model. Technometrics. 2002;44:85–86. [Google Scholar]
- 64.Kassambara A., Kosinski M., Biecek P. Survminer: drawing survival curves using'ggplot2'. CRAN Contrib Packages. 2016 [Google Scholar]
- 65.Geeleher P., Cox N., Huang R.S. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS One. 2014;9 doi: 10.1371/journal.pone.0107468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Li Y., Wu X., Yang P., et al. Machine learning for lung cancer diagnosis, treatment, and prognosis. Genom Proteom Bioinforma. 2022;20:850–866. doi: 10.1016/j.gpb.2022.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhang J., Li H., Tao W., et al. GseaVis: an R package for enhanced visualization of gene set enrichment analysis in biomedicine. Med Res. 2025;1:131–135. [Google Scholar]
- 68.Liu L., Xie Y., Yang H., et al. HPVTIMER: a shiny web application for tumor immune estimation in human papillomavirus-associated cancers. Imeta. 2023;2 doi: 10.1002/imt2.130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Fang B., Mehran R.J., Heymach J.V., et al. Predictive biomarkers in precision medicine and drug development against lung cancer. Chin J Cancer. 2015;34:1–15. doi: 10.1186/s40880-015-0028-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Zhou J.-B., Tang D., He L., et al. Machine learning model for anti-cancer drug combinations: Analysis, prediction, and validation. Pharmacol Res. 2023;194 doi: 10.1016/j.phrs.2023.106830. [DOI] [PubMed] [Google Scholar]
- 71.Du M., Qu Y., Qin L., et al. The cell death-related genes machine learning model for precise therapy and clinical drug selection in hepatocellular carcinoma. J Cell Mol Med. 2024;28 doi: 10.1111/jcmm.18168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Chen X., Zhang H., Guo D., et al. Risk of intraoperative hemorrhage during cesarean scar ectopic pregnancy surgery: development and validation of an interpretable machine learning prediction model. eClinicalMedicine. 2024;78 doi: 10.1016/j.eclinm.2024.102969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Lin A., Qi C., Wei T., et al. CAMOIP: a web server for comprehensive analysis on multi-omics of immunotherapy in pan-cancer. Brief Bioinforma. 2022;23 doi: 10.1093/bib/bbac129. [DOI] [PubMed] [Google Scholar]
- 74.Lou N., Cui X., Lin X., et al. Development and validation of a deep learning-based model to predict response and survival of T790M mutant non-small cell lung cancer patients in early clinical phase trials using electronic medical record and pharmacokinetic data. Transl lung Cancer Res. 2024;13:706. doi: 10.21037/tlcr-23-737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Bertsimas D., Wiberg H. Machine learning in oncology: methods, applications, and challenges. JCO Clin Cancer Inform. 2020;4 doi: 10.1200/CCI.20.00072. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary material
Table S2. Concentrations and sources of chemotherapeutic agents used in cell line studies
Data Availability Statement
The data used in this study are available from the following public repositories: Genomics of Drug Sensitivity in Cancer (GDSC) database (https://www.cancerrxgene.org/), Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). All other data generated and analyzed in this study are available from the corresponding authors upon reasonable request.