Abstract
Super-enhancers (SEs) represent a distinct category of cis-regulatory elements notable for their robust transcriptional activation capabilities. In tumor cells, SEs intricately regulate the expression of oncogenes and pivotal cancer-associated signaling pathways, offering significant potential for cancer treatment. However, few studies have systematically discussed the crucial role of SEs in hepatocellular carcinoma (HCC), which is one of the most common liver cancers with late-stage diagnosis and limited treatment methods for advanced disease. Herein, we first summarize the identification methods and the intricate processes of formation and organization of super-enhancers. Subsequently, we delve into the roles and molecular mechanisms of SEs within the framework of HCC. Finally, we discuss the inhibitors targeting the key SE-components and their potential effects on the treatment of HCC. In conclusion, this review meticulously encapsulates the distinctive characteristics of SEs and underscores their pivotal roles in the context of hepatocellular carcinoma, presenting a novel perspective on the potential of super-enhancers as emerging therapeutic targets for HCC.
Keywords: Hepatocellular carcinoma, Super-enhancer, Therapeutic targets
Introduction
Hepatocellular carcinoma (HCC) is the most prevalent primary liver cancer, which accounts for 90% of all new occurrences of primary liver cancer worldwide [1]. Due to the asymptomatic nature of the disease, most HCC patients are diagnosed at an advanced stage, which contributes to the fact that the prognosis of HCC is very poor (the 5-year OS rate is approximately 18%) [2, 3]. Although clinically beneficial treatment methods for HCC include surgical resection [4], transplantation [5], ablation [6] and transcatheter arterial chemoembolization [7], there is basically no effective anticancer medication. Therefore, it is urgent to fully understand the crucial mechanisms for the development of new therapies.
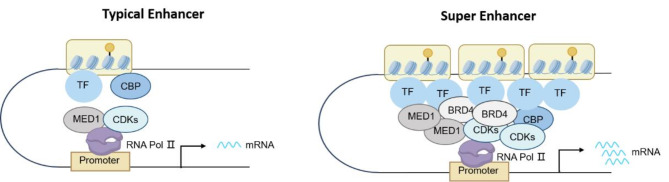
Enhancers and super-enhancers are receiving significant attention for their pivotal roles in gene transcription, which is a highly intricate and meticulously coordinated process in the pathogenesis of complex diseases in humans [8–11]. Enhancers are distant noncoding cis-regulatory elements that play a vital role in controlling cell-type-specific spatiotemporal gene expression programs by attracting specific transcription factors that increase promoter activity to activate target genes, often through long-range chromosomal interactions [12, 13]. Super-enhancers are large clusters of enhancers with abnormally high levels of transcriptional activation ability than typical enhancers (TEs) [14]. Similar to the TEs, SEs can be bound by some factors, including transcription factors (TFs), coactivators, chromatin regulators, and the RNA Pol II complex. However, there are substantial structural and functional differences between super-enhancers and typical enhancers, as shown in Fig. 1. SEs are characterized by a dense occupancy of transcription factors, with an average of 10-fold higher density than TEs [15–17]. Notably, many of the transcription factors that bind to super-enhancers are integral to key oncogenic signaling pathways, such as Wnt and TGF-β [18, 19]. And SEs-related genes are critical for maintaining cancer cell identity and promoting tumorigenic gene transcription [20]. Additionally, multiple studies elucidated that inhibitors targeting SE structures have achieved significant clinical results in cancers, including prostate cancer, colorectal cancer and acute myeloid leukemia [21–23]. Banerji et al. revealed that many types of tumor cells are particularly sensitive to the inhibition of the SEs complexes, and inhibitors of the SEs complex have demonstrated positive anticancer effects [24]. Consequently, SEs are capable of driving the transcription of their target genes at a much higher rate and play a more critical role in the development and progression of cancers, which are potential new targets for cancer therapy.
Fig. 1.
The difference between typical enhancers and super-enhancers
However, there is a scarcity of studies that systematically elucidated the role of SEs in HCC and its new prospects for therapeutic targets. Thus, this review summarizes the identification, formation and organization approaches of SEs, focusing on the mode of mechanism as well as the drug inhibitors and therapeutic effect for targeting SEs, which may provide new insights for future research and optimum treatment strategies in HCC.
Identification of super-enhancers
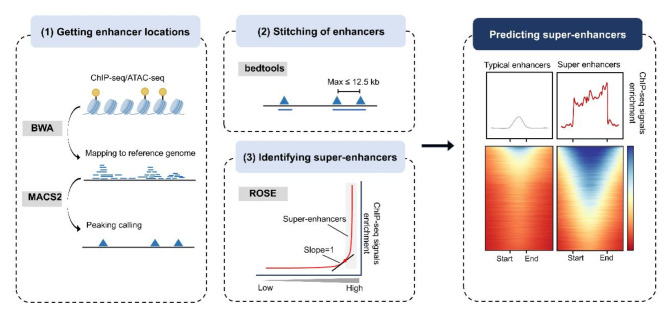
Young et al. were the pioneers in identifying a unique class of enhancers in mouse embryonic stem cells (mESCs) that were unusually dense with transcription factors and the Mediator complex. They designated these extensive clusters of enhancers, which significantly influence the transcriptional expression of genes, as ‘super-enhancers’ [14, 25]. Here we conclude three main steps of identifying SEs (Fig. 2): (1) Achieving the region of active enhancers. First, techniques such as chromatin immunoprecipitation sequencing (ChIP-seq), Assay for Transposase Accessible Chromatin using sequencing (ATAC-seq) are performed to map the sequences of transcription factors (e.g. Oct4, Sox2, and Nanog), Mediator, and histone modifications (H3K27ac and H3K4me1) [14, 26–29] across the genome by using Burrows-Wheeler Aligner (BWA) [30]. Then, run Model-based Analysis for ChIP-Seq version 2 (MASC2) to obtain objective peak files [31]. (2) Stitching enhancer. Bedtools version 2.27.0 is used to merge neighboring enhancers (within 12.5 kb) into one group to capture dense enhancer clusters [32]; (3) Identifying super-enhancers. ROSE algorithm (http://younglab.wi.mit.edu/super_enhancer_code.html) is a tool for finding SEs using gff file (regions of enhancer) and bam file (the factor density of enhancers). After ranking stitched enhancers according to their ChIP-seq signals, SEs are separated from typical enhancers based on a cutoff value [14, 27].
Fig. 2.
The specific process of SE identification and advanced techniques
The formation and organization of super-enhancers
Super-enhancers gradually recruit biomolecules to form transcription complexes
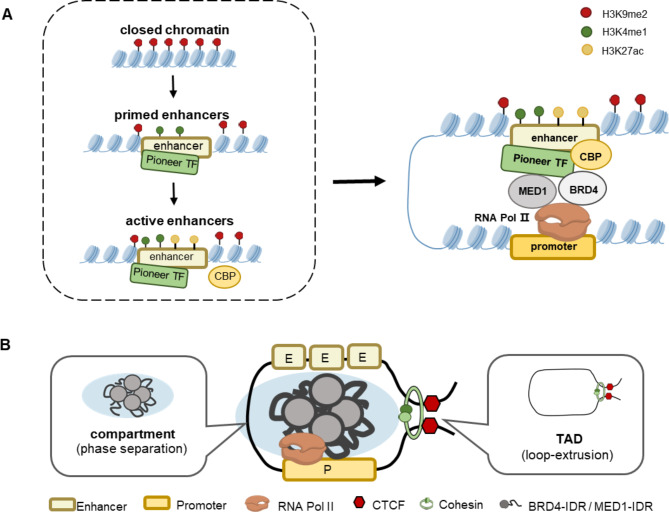
SE-regulated gene expression in mammalian cells is a biological process in which biomolecules act in cooperation with each other. The transcription complexes recruited by SEs contain biomolecules including transcription factors, chromatin regulators (e.g. CBP/P300 and HDAC), transcriptional co-activators (e.g. MED1 and BRD4) and RNA polymerase II (RNA pol II) [33–35]. However, it is unclear how SEs recruit these transcriptional regulators. Here, we briefly describe the process by which SEs form transcription complexes stepwise (Fig. 3A). First, pioneer factors bind to highly folded DNA and establish the accessible chromatin conformations to promote the binding of additional transcriptional elements [36–39]. Pioneer factors such as FOXA and SOX2 are instrumental in opening chromatin by facilitating the separation of terminal nucleosomal DNA from histone octamer [40–43]. Notably, the DNA-binding domain (DBD) of the pioneer factor is crucial for recognizing specific DNA sequence motifs, which determines the precise location of SEs in the genome [44, 45]. Subsequently, the pioneer factor-induced changes of chromatin accessibility transform “closed chromatin” with high levels of H3K9me2 into “primed enhancers” characterized by H3K4me1 [46–49]. Thereafter, the primed enhancers will divide into two directions depending on the type of co-transcription factors they recruit: (1) Becoming active enhancers by recruiting the co-activator CBP/P300 and completing the recruitment of H3K27ac [50, 51]; (2) Becoming inactive enhancers due to the presence of components such as histone deacetylases (HDACs) in the recruitment complex [52, 53]. Finally, BRD4 binds to the active enhancer due to its ability to read highly acetylated histones and serves as a bridge to recruit Med1 and RNA pol II [54, 55]. It is noteworthy that Mediator acts as a coordinator to mediate the interaction between transcription factors and RNA polymerase II machinery, facilitating communication between SEs and the target gene promoters. Together, SEs regulate chromatin accessibility by modulating histone modifications, recruit multiple biomolecules, and ultimately form transcriptional activation complexes, which lays the compositional foundation for the formation of the more complex structures of SEs.
Fig. 3.
Super-enhancers recruit transcription complexes and form 3D structures. (A) Pioneer factors bind to closed chromatin with a high level of H3K9me2 and transform it into “primed enhancers” characterized by H3K4me1. Through the recruitment of CBP/P300, primed enhancers can get H3K27ac at nucleosomes and become active enhancers. Finally, the histone “reader” BRD4 binds to the active enhancers and recruits Med1 and RNA pol II, which form transcription complexes at the SE region. (B) Compartments - Topologically associating domains” model of super-enhancers. The IDR of BRD4 and MED1 mediate the formation of phase-separated and promote compartment formation. The CTCF and cohesion promote the process of loop-extrusion and determine the TAD boundaries
Super-enhancers organize intricate structure to achieve efficient and precise regulation
Compared to typical enhancers, SEs exhibit extended lengths and elevated densities of transcription factors, which enhance transcriptional efficiency and make SEs more effective in regulating gene expression. Furthermore, recent studies have shown that SEs possess more intricate structures at the three-dimensional chromatin level. Herein, we propose the “compartments-Topologically associating domains (TADs)” model (Fig. 3B) to explain the advanced structural features of SEs and reveal how SEs play an efficient and precise role in transcriptional regulation. For its efficiency, the main reason is the intrinsically disordered regions (IDRs) of BRD4 and MED1(two key SEs transcriptional coactivators), which mediate the formation of phase-separated structures and promote compartment formation to facilitate efficient biochemical reactions [56–60]. For its precision, CCCTC-binding factor (CTCF) and cohesion, which were widely detected at the boundaries of SEs, promote the process of loop-extrusion and determine the TAD distinct boundaries to maintain the precise regulation of genes [42, 61–66]. Overall, the organization of the genome is the result of the interaction between phase-separation-driven compartments and loop-extrusion-driven TADs.
The roles and molecular mechanisms of SEs in HCC
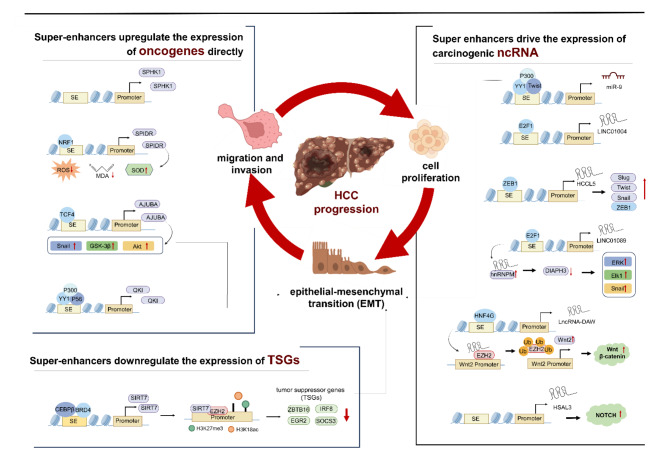
There is mounting evidence suggesting that SEs exert pivotal control over central hallmarks of cancer, including abnormal cell proliferation, migration, invasion, and epithelial-mesenchymal transition (EMT) [67]. In the progression of HCC, SEs not only influence protein-coding genes but also play a crucial role in the transcriptional regulation of non-coding RNAs (ncRNAs), such as microRNAs (miRNAs) and long non-coding RNAs (lncRNAs). Here, we summarize the roles and molecular mechanisms of SEs in the development of HCC, which will provide the potential targets for the treatment of HCC (Table 1; Fig. 4).
Table 1.
Recent studies on SEs in hepatocellular carcinoma
| SE-related-targets | Classification | Enriched TF | Implication | Potential drug |
|---|---|---|---|---|
| SPHK1 | gene | - | Cell proliferation, migration |
SKI-II, CBP30, JQ1 and THZ1 |
| SPIDR | gene | NRF1 |
Cell proliferation, oxidative stress response |
JQ1 |
| AJUBA | gene | TCF4 |
Cell proliferation, migration, invasion, EMT, Akt/GSK-3β/Snail pathway |
- |
| QKI | gene | YY1/p65/p300 complex | Cell proliferation, migration, invasion, EMT | JQ1, Hyperoside |
| SIRT7 | gene | C/EBPβ | Cell proliferation, invasion, EMT | JQ1 |
| miR-9 | microRNA | Twist1-YY1-p300 complex | Cell proliferation, migration, invasion, EMT | Metformin |
| LINC01004 | lncRNA | E2F1 | Cell proliferation, migration | - |
| HCCL5 | lncRNA | ZEB1 | Cell proliferation, migration, invasion, EMT | - |
| LINC01089 | lncRNA | E2F1 |
Cell proliferation, migration, invasion, EMT, ERK/Elk1/Snail axis |
- |
| lncRNA-DAW | lncRNA | HNF4G |
Cell proliferation, migration, invasion, Wnt/β-catenin pathway |
- |
| HSAL3 | lncRNA | HCFC1, HSF1 |
Cell proliferation, migration, NOTCH signaling |
JQ1 |
EMT, epithelial-mesenchymal transition;
Fig. 4.
The roles and molecular mechanisms of super-enhancers in HCC
Super-enhancers upregulate the expression of oncogenes directly
Compared with typical enhancers, super-enhancers have a higher density of histone modifications and can bind a higher density of transcription factors to tremendously promote gene expression [25, 68]. Here, we summarize some oncogenes upregulated by super-enhancers in HCC, such as SPHK1, SPIDR, AJUBA, and QKI. SPHK1 is a kinase that mediates sphingosine phosphorylation to produce sphingosine-1-phosphate (S1P), which plays an important role in sphingolipid metabolism of HCC and regulates a variety of pro-survival functions [69]. After interfering with SE-promoter interactions by CRISPR, the expression level of SPHK1 was significantly reduced, leading to decreased proliferation and migration of hepatocellular carcinoma cells [70]. Besides, SPIDR is an HCC-specific gene that is closely associated with the oxidative stress response. The transcription factor NRF1 activates the transcription of SPIDR by binding to its SE region. Silencing SPIDR or NRF1 results in increased levels of reactive oxygen species (ROS) and malondialdehyde (MDA), and decreased levels of superoxide dismutase (SOD) in HCC cells, indicating an impairment of antioxidant capacity [71]. Additionally, it is noteworthy that numerous SE-driven oncogenes, including AJUBA and QKI, exert a pivotal role in facilitating EMT [72, 73]. Specifically, AJUBA is a SE-associated gene regulated by the transcription factor TCF 4 in HCC, with high expression correlated with aggressive phenotype and poor prognosis in patients [74]. High levels of AJUBA expression induce EMT by recruiting tumor necrosis factor-associated factor 6 (TRAF6), enhancing Akt phosphorylation, and activating the Akt/GSK-3β/Snail pathway. Furthermore, during the transcription of alternative splicing factor Quaking (QKI), the super-enhancer of QKI which was bind by YY1/p65/p300 complex can activate the transcription of QKI gene. This activation promotes EMT and facilitates the migration and invasion in hepatocellular carcinoma cells [75]. Together, SEs accelerate cell proliferation, migration, invasion, and EMT mainly through genes such as SPHK1, AJUBA, QKI and SPIDR, promoting the malignant progression of hepatocellular carcinoma.
Super-enhancers downregulate the expression of tumor suppressor genes through epigenetic modification
The process of carcinogenesis is highly intricate, involving not only alterations in oncogene function but also perturbations in the function of tumor suppressor genes (TSGs). In this context, SEs possess the capability to modulate TSGs expression through the influence on their epigenetic modifications. In HCC cells, SIRT7-SE enhances SIRT7 transcription by promoting co-occupancy of C/EBPβ and BRD4. Notably, Sirtuin 7 (SIRT7) can induce genome-wide deacetylation of H3K18 and physically interact with the H3K27 methyltransferase EZH2 to promote cooperative epigenetic silencing. This ultimately results in the suppression of TSGs such as EGR2, IRF8, SOCS, and ZBTB, which are critical for regulating metabolism and immunity, thereby promoting the tumorigenicity of HCC cells [76]. In conclusion, SE promotes the transcriptional silencing of TSGs through SIRT7-mediated epigenetic modification, indicating that it is particularly important to study the potential regulation of SE on tumor suppressor genes.
Super enhancers drive the expression of carcinogenic ncRNA
Non-coding RNAs (ncRNAs) are crucial in determining cell fate by interacting with proteins or other RNAs [77]. Based on existing research, SE-associated ncRNAs in HCC can be primarily classified into two categories, microRNAs and lncRNAs, which play significant roles in cancer cell proliferation, carcinogenic signaling pathways, and the EMT process (Table 1).
MiR-9 is a SE-related microRNA that can promote the process of EMT in HCC [78]. It is noteworthy that the Twist1/YY1/p300 complex can bind to the SE region of miR-9 and form phase-separated condensates to regulate the expression of miR-9. Interestingly, metformin can dissolve these phase-separated puncta, thereby inhibiting the progression of HCC. Regarding lncRNA-SE, the binding of transcription factor E2F1 to the LINC01004-SE can promote the transcription of LINC01004, thereby facilitating cell proliferation and metastasis [79]. Additionally, SE-associated lncRNAs can regulate EMT to promote HCC cell migration and invasion. For example, lncRNA HCCL5 is transcriptionally driven by ZEB1 binding to its super-enhancer region, which increases the expression of cancer-associated transcription factors including Snail, Slug, ZEB1 and Twist in hepatocellular carcinoma cells, thereby accelerating the EMT phenotype [80]. Moreover, SE-driven lncRNA LINC01089 can promote EMT by regulating DIAPH3 expression. The transcription factor E2F1 binds to the LINC01089-SE and promotes its transcription. LINC01089 interacts with heterogeneous nuclear ribonucleoprotein M (hnRNPM) in the nucleus, thereby enhancing hnRNPM functions and suppressing DIAPH3 protein levels, which in turn promotes the ERK/Elk1/Snail axis and ultimately induces EMT progression [81]. In addition, lncRNA can act by participating in the process of oncogenic signaling pathways. LncRNA-DAW is activated by the liver-specific super-enhancer and physically interacts with EZH2 (negative regulator of Wnt2), resulting in EZH2 phosphorylation and ubiquitination, which leads to the de-repression of Wnt2 and the activation of the Wnt/β-catenin pathway [82]. Furthermore, another SE-associated lncRNA called HSAL3 exerts its oncogenic function through regulating NOTCH signaling to promote HCC cells growth and metastasis [83].
However, despite the crucial role of SE-driven ncRNAs in modulating cancer progression, the precise mechanism of how SE regulates the production of ncRNA remains unclear. Altogether, leveraging SE-related ncRNA to treat HCC is a potential research field that warrants further investigation.
Super-enhancers as new prospects for therapeutic targets and inhibitors in HCC
Super-enhancers (SEs) have emerged as promising therapeutic targets in hepatocellular carcinoma (HCC). Here are three mainly potential therapeutic strategies for targeting SEs in cancer treatment: (1) epigenetic modulators that regulate chromatin accessibility to alter SE activity and restore gene expression [20]; (2) CRISPR-Cas9 genome editing to delete or modulate critical components for precise and targeted disruption of SEs [84, 85]; (3) small molecule inhibitors designed to disrupt interactions among key SE components, especially for three critical SE components including BRD4, CDKs, and CBP/P300 (EP300) (Table 2). Here we concluded the potential function and the inhibitors of three key components of SEs, which may provide new prospects for optimum treatment of HCC by targeting SEs.
Table 2.
Summary of the inhibitors targeting SEs in HCC
| Targets | Inhibitor | Target model | Methods | Results | References |
|---|---|---|---|---|---|
| BRD4 | JQ1 | HepG 2 and LO2 cells | 0.5 µM, 48 h | JQ1 significantly reduced SIRT7 mRNA levels and a loss of enhancer activity | [76] |
| JQ1 | MHCC97H and HepG2.2.15 cells | 0.5 µM, 48 h | BRD4 exposure significantly reduced ETV4 expression in both MHCC97H and HepG2.2.15 cells | [86] | |
| JQ1 | HepG2 and LM3 cells | 0.5 µM, 6 h | JQ1 preferred to inhibit SE-associated lncRNA transcription in HCC | [83] | |
| JQ1 | HepG2 and Huh7 cells | / | JQ1 reduced the expression of target gene at both the mRNA and protein levels, substantially abrogated the BRD4 occupancy in the SE region | [70] | |
| AZD5153 | HCCLM3, HepG2, and Huh7 cells | 1 to 100 µM, 72 h, | AZD5153 reduced cell proliferation dose-dependently in all HCC cell lines tested. | [87] | |
| AZD5153 | Mouse (NSG mice with subcutaneously transplanted HCCLM3 cells) | 3 mg/kg/day, 3 weeks | AZD5153 significantly reduced tumor weight and volume and the proliferation of tumor cells. | [87] | |
| CDK7 | THZ1 | HepG2 and Huh7 cells | 100 nM, 6 h | Treatment of THZ1 significantly attenuated cell proliferation, colony formation, cell migration, and induced apoptosis in HCC cells. | [70] |
| THZ1 | Mouse (Luciferase-labeled Huh7 cells are injected into the liver of nude mice to establish liver tumor) | 5 mg/kg/day, 14 days | THZ1 significantly reduced the liver tumor size | [70] | |
| CDK9 | Alvocidib | KOB and ST-1 ATL cell lines | 100 nM, 4.5 h | The current literature lacks reports on the impact of CDK9 inhibitors via SE in HCC | [88] |
| EP300 | CBP30 | HepG2 and Huh7 cells | / | CBP300 repressed the expression of the 13 HCC-SE genes in HCC cells | [70] |
BRD4 inhibitors in HCC
BRD4, a member of the bromodomain and extra terminal domain (BET) family of proteins, is known as epigenetic readers of acetylated histones and plays a key role in chromatin remodeling and transcriptional regulation [89–92]. As an important element of super-enhancers, BRD4 can regulate the expression of target genes by recognizing histone acetylation and recruiting different transcriptional regulators such as MED1 and positive transcription elongation factor b (P-TEFb) [93–95]. Lovén et al. showed that the use of bromine domain inhibitors caused preferential loss of BRD4 occupation at SEs [96]. Remarkably, BRD4 is noticeably upregulated in HCC tissues as well as in liver cancer cell lines, and its overexpression in cancerous tissues correlates with an unfavorable prognosis in HCC patients [97]. Given BRD4 proteins contain two N-terminal bromodomains, BD1 and BD2, small-molecule inhibitors of BET proteins can be divided into two categories: monovalent inhibitors (e.g. JQ1) which can bind to either BD1 or BD2, and bivalent inhibitors (e.g. AZD5153) which can bind to both BD1 and BD2 [87, 89, 98]. For BRD4 monovalent inhibitors, JQ1 is recognized as a selective inhibitor of the BD1 domain and plays a pivotal role in BRD4 signaling pathway, which is widely used for preclinical studies of various malignancies including HCC [99–101]. Therefore, JQ1 can significantly reduce the activity of carcinogenic super-enhancers and suppress the activity of various HCC-related genes, including SIRT7, ETV4 and SPHK1, and thereby inhibiting the proliferation of hepatocellular carcinoma cells [70, 76, 86]. However, the pharmacokinetic profile of JQ1 in vivo presents challenges to its broad application. High doses have been correlated with lethal outcomes in experimental mice [102], and its short half-life of approximately 1 h in CD1 mice indicates a non-sustained therapeutic effect [98, 103–105]. These characteristics represent suboptimal properties to the in vivo application of JQ1. For BRD4 bivalent inhibitors, the novel AZD5153 has shown encouraging results [87]. Compared with monovalent inhibitors, AZD5153 shows improved potency and a wider range of applications. At the animal and cellular levels, AZD5153 showed significant anti-HCC activity. At the clinical trial level, Phase 1 clinical trials of AZD5153 have been completed in patients with relapsed or refractory solid tumors (NCT03205176). Altogether, BRD4 inhibitors are of great research value and extensive clinical application prospect, but are still of great challenge that needs to be addressed in future studies.
CDK inhibitors in HCC
The mammalian cyclin-dependent kinases (CDKs) contain subfamilies with specific functions related to cell cycle (CDK1, CDK2, CDK4, CDK6) and transcriptional regulation (CDK7, CDK8, CDK9, CDK12, and CDK19) [106]. Among them, CDK7 plays the role in the oncogenic SE-involved transcription by phosphorylating the subunit of RNA Pol II. As a member of the transcription factor TFIIHb, CDK7 is up-regulated in HCC tissues and its expression is negatively correlated with the survival of HCC patients [107]. Studies have shown that CDK7 promotes the growth and migration of hepatocellular carcinoma cells, and targeting CDK7 induces apoptosis and inhibits hepatocellular carcinoma tumor growth [108]. Among that, THZ1 is the most widely used specific covalent inhibitor of CDK7 which suppresses CDK7 kinase activity based on modification of a unique cysteine residue [109]. After brief treatment with low concentrations of THZ1 (100 nM for 6 h), the transcript levels of SE-related genes were significantly reduced in THZ1-treated HCC cells. THZ1-sensitive genes are significantly enriched in biological processes, including transcription, cellular metabolic processes, regulation of gene expression, cell cycle progression and DNA repair, suggesting that essential genes involved in the sustainability of HCC cells are particularly susceptible to CDK7 inhibition [70]. Notably, another cyclin-dependent kinase CDK9 also plays a pivotal role in transcription process. As the subunit of positive transcription elongation factor b (P-TEFb), CDK9 can also phosphorylate the subunit of RNA Pol II similar to CDK7 (but not the same sites) to stimulate transcription elongation [110]. In addition, targeting CDK9 has shown positive effects in tumor therapy. Sakamoto’s research showed that alvocidib, an inhibitor of CDK9, can inhibit adult T-cell leukemia/lymphoma (ATL) cell proliferation through SE-mediated down-regulation of IRF4 expression [88]. Unfortunately, although CDK9 has been shown to be an important component of SEs, no specific studies have been performed to clarify that CDK9 inhibits hepatocellular carcinoma development by affecting SEs in HCC cells, which is worthy of further exploration [111–114]. Together, considering the significant anticancer effects of CDK7 and CDK9 inhibitors in HCC and the complexity of their regulatory mechanisms, further research on the CDK inhibitors of SEs is of great significance.
CBP/p300 inhibitors in HCC
Recent studies have shown that the CBP/p300 was overexpressed in hepatocellular carcinoma and other various cancer cells which were regulated by SEs and could activate oncogene transcription and induce cancer cell proliferation, survival, tumorigenesis, and metastasis [75, 78]. Among that, the catalytic core of CBP/p300 protein consists of histone acetyltransferase (HAT), bromodomain and ZZ-type zinc finger domain, of which the HAT and the bromodomain components’ inhibitors play more significant roles in suppressing SEs functions [115–117]. The main reason is that the HAT induces acetylation of histone lysine H3 in the super-enhancer region of target genes while the bromodomain region of CBP/p300 can bind to the HAT structural domain of thereby amplifying HAT activity. As a CBP/p300 HAT inhibitor, B029-2 reduces glycolytic function and nucleotide synthesis to inhibit hepatocellular carcinoma cell proliferation, migration and invasion in vitro and tumor progression in vivo [118]. CBP/p300 bromodomain inhibitor CBP30 was able to inhibit cell proliferation and colony formation as well as the expression of SE-associated genes in hepatocellular carcinoma cells HepG 2 and Huh 7 cells through selective bromodomain blockade of CBP [70]. It is worth mentioning that the transcriptional profile of CBP30-treated human T cells showed a much more limited effect on gene expression compared to other bromodomain inhibitors like JQ1, and thus selective targeting of the CBP/p300 bromodomain by CBP30 may result in fewer side effects [119]. For clinical applications, a variety of CBP/p300 inhibitors are currently being used in clinical trials in cancer patients. Although no clinical trials have been reported for HCC, given the sensitivity as well as the safety of CBP/p300 inhibitors, targeted clinical trials are still of great research value.
Conclusion
Currently, it is widely accepted that alterations in cis-regulatory elements represent a major mechanism underlying cancer development, particularly hepatocellular carcinoma, which is one of the most prevalent malignancies with high recurrence rates. Here, we review the process of identification and constitution of SEs, summarize the mechanism how SE plays an oncogenic role in HCC, and discuss current insights into the inhibitors targeting SE components and their potential value for application in HCC therapy. However, the following issues still need to be considered. First, although a large body of literature suggests that both TADs and compartments characterize SEs, the detailed mechanisms of how the two cooperate with each other remain largely unexplored, and the complex composition as well as the more advanced structure of SEs need to be further investigated. Second, despite the encouraging results of SEs inhibitors in animal tests as well as cellular experiments, there are few clinical trials and their dosage should be further evaluated based on individual differences to determine potential adverse reactions. In summary, targeting super-enhancers offers a promising avenue for developing novel therapeutic strategies in HCC, though further research on the specific action machinery and clinical data is needed.
Author contributions
X.L.: Conceptualization, writing original draft preparation. M.Z.: Search literature. X.P.: Visualization. J.M.: Investigation. R.W.: Supervision, visualization. Y.W.: Visualization. S.C.: Supervision. Y.Y.: Methodology, writing original draft. Y.Z.:Conceptualization, project administration, supervision, validation. All authors reviewed the manuscript.
Funding
This research was funded by the National Natural Science Foundation of China, grant number 82300661; the Natural Science Foundation of Anhui province, grant number 2308085QH246; Scientific Research of BSKY from Anhui Medical University, grant number XJ201925; the Natural Science Foundation of the Anhui Higher Education Institutions, grant number KJ2021A0205; Basic and Clinical Cooperative Research Promotion Program of Anhui Medical University, grant number 2022xkjT013.
Data availability
No datasets were generated or analysed during the current study.
Declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Contributor Information
Yan Yan, Email: yanyht@163.com.
Yaling Zhu, Email: zhuyaling@ahmu.edu.cn.
References
- 1.Rinaldi L, Guarino M, Perrella A, Pafundi PC, Valente G, Fontanella L, Nevola R, Guerrera B, Iuliano N, Imparato M, et al. Role of Liver Stiffness Measurement in Predicting HCC occurrence in direct-acting antivirals setting: a real-life experience. Dig Dis Sci. 2019;64(10):3013–9. [DOI] [PubMed] [Google Scholar]
- 2.Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391(10127):1301–14. [DOI] [PubMed] [Google Scholar]
- 3.Villanueva A. Hepatocellular Carcinoma. N Engl J Med. 2019;380(15):1450–62. [DOI] [PubMed] [Google Scholar]
- 4.Sugawara Y, Hibi T. Surgical treatment of hepatocellular carcinoma. Biosci Trends. 2021;15(3):138–41. [DOI] [PubMed] [Google Scholar]
- 5.Vibert E, Schwartz M, Olthoff KM. Advances in resection and transplantation for hepatocellular carcinoma. J Hepatol. 2020;72(2):262–76. [DOI] [PubMed] [Google Scholar]
- 6.Wang K, Wang C, Jiang H, Zhang Y, Lin W, Mo J, Jin C. Combination of ablation and immunotherapy for Hepatocellular Carcinoma: where we are and where to go. Front Immunol. 2021;12:792781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ando Y, Kawaoka T, Amioka K, Naruto K, Ogawa Y, Yoshikawa Y, Kikukawa C, Kosaka Y, Uchikawa S, Morio K, et al. Efficacy and safety of Lenvatinib-Transcatheter arterial chemoembolization sequential therapy for patients with Intermediate-Stage Hepatocellular Carcinoma. Oncology. 2021;99(8):507–17. [DOI] [PubMed] [Google Scholar]
- 8.Cramer P. Organization and regulation of gene transcription. Nature. 2019;573(7772):45–54. [DOI] [PubMed] [Google Scholar]
- 9.Field A, Adelman K. Evaluating enhancer function and transcription. Annu Rev Biochem. 2020;89:213–34. [DOI] [PubMed] [Google Scholar]
- 10.Pang B, van Weerd JH, Hamoen FL, Snyder MP. Identification of non-coding silencer elements and their regulation of gene expression. Nat Rev Mol Cell Biol. 2023;24(6):383–95. [DOI] [PubMed] [Google Scholar]
- 11.Zabidi MA, Stark A. Regulatory enhancer-core-promoter communication via transcription factors and cofactors. Trends Genet. 2016;32(12):801–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Banerji J, Rusconi S, Schaffner W. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell. 1981;27(2 Pt 1):299–308. [DOI] [PubMed] [Google Scholar]
- 13.Grosveld F, van Assendelft GB, Greaves DR, Kollias G. Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell. 1987;51(6):975–85. [DOI] [PubMed] [Google Scholar]
- 14.Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, Rahl PB, Lee TI, Young RA. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153(2):307–19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Moorthy SD, Davidson S, Shchuka VM, Singh G, Malek-Gilani N, Langroudi L, Martchenko A, So V, Macpherson NN, Mitchell JA. Enhancers and super-enhancers have an equivalent regulatory role in embryonic stem cells through regulation of single or multiple genes. Genome Res. 2017;27(2):246–58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pott S, Lieb JD. What are super-enhancers? Nat Genet. 2015;47(1):8–12. [DOI] [PubMed] [Google Scholar]
- 17.Tang F, Yang Z, Tan Y, Li Y. Super-enhancer function and its application in cancer targeted therapy. NPJ Precis Oncol. 2020;4:2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Scholz BA, Sumida N, de Lima CDM, Chachoua I, Martino M, Tzelepis I, Nikoshkov A, Zhao H, Mehmood R, Sifakis EG, et al. WNT signaling and AHCTF1 promote oncogenic MYC expression through super-enhancer-mediated gene gating. Nat Genet. 2019;51(12):1723–31. [DOI] [PubMed] [Google Scholar]
- 19.Zhang T, Xia W, Song X, Mao Q, Huang X, Chen B, Liang Y, Wang H, Chen Y, Yu X, et al. Super-enhancer hijacking LINC01977 promotes malignancy of early-stage lung adenocarcinoma addicted to the canonical TGF-β/SMAD3 pathway. J Hematol Oncol. 2022;15(1):114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bacabac M, Xu W. Oncogenic super-enhancers in cancer: mechanisms and therapeutic targets. Cancer Metastasis Rev. 2023;42(2):471–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cao Z, Shu Y, Wang J, Wang C, Feng T, Yang L, Shao J, Zou L. Super enhancers: pathogenic roles and potential therapeutic targets for acute myeloid leukemia (AML). Genes Dis. 2022;9(6):1466–77. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen X, Ma Q, Shang Z, Niu Y. Super-enhancer in prostate cancer: transcriptional disorders and therapeutic targets. NPJ Precis Oncol. 2020;4(1):31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu Q, Guo L, Lou Z, Xiang X, Shao J. Super-enhancers and novel therapeutic targets in colorectal cancer. Cell Death Dis. 2022;13(3):228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Shin HY. Targeting super-enhancers for Disease Treatment and diagnosis. Mol Cells. 2018;41(6):506–14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-André V, Sigova AA, Hoke HA, Young RA. Super-enhancers in the control of cell identity and disease. Cell. 2013;155(4):934–47. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Buenrostro JD, Wu B, Chang HY, Greenleaf WJ. ATAC-seq: A Method for Assaying Chromatin Accessibility Genome-Wide. Curr Protoc Mol Biol 2015, 109:21.29.21–21.29.29. [DOI] [PMC free article] [PubMed]
- 27.Creyghton MP, Cheng AW, Welstead GG, Kooistra T, Carey BW, Steine EJ, Hanna J, Lodato MA, Frampton GM, Sharp PA, et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc Natl Acad Sci U S A. 2010;107(50):21931–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Grandi FC, Modi H, Kampman L, Corces MR. Chromatin accessibility profiling by ATAC-seq. Nat Protoc. 2022;17(6):1518–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kaya-Okur HS, Wu SJ, Codomo CA, Pledger ES, Bryson TD, Henikoff JG, Ahmad K, Henikoff S. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat Commun. 2019;10(1):1930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Abuín JM, Pichel JC, Pena TF, Amigo J. BigBWA: approaching the Burrows-Wheeler aligner to Big Data technologies. Bioinformatics. 2015;31(24):4003–5. [DOI] [PubMed] [Google Scholar]
- 31.Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9(9):R137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26(6):841–2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Allen BL, Taatjes DJ. The Mediator complex: a central integrator of transcription. Nat Rev Mol Cell Biol. 2015;16(3):155–66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Chernukhin I, Shamsuddin S, Kang SY, Bergström R, Kwon YW, Yu W, Whitehead J, Mukhopadhyay R, Docquier F, Farrar D, et al. CTCF interacts with and recruits the largest subunit of RNA polymerase II to CTCF target sites genome-wide. Mol Cell Biol. 2007;27(5):1631–48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, Chen X, Taipale J, Hughes TR, Weirauch MT. The human transcription factors. Cell. 2018;172(4):650–65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Iwafuchi-Doi M, Zaret KS. Cell fate control by pioneer transcription factors. Development. 2016;143(11):1833–7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jerkovic I, Cavalli G. Understanding 3D genome organization by multidisciplinary methods. Nat Rev Mol Cell Biol. 2021;22(8):511–28. [DOI] [PubMed] [Google Scholar]
- 38.Mayran A, Drouin J. Pioneer transcription factors shape the epigenetic landscape. J Biol Chem. 2018;293(36):13795–804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Zaret KS. Pioneer Transcription Factors Initiating Gene Network Changes. Annu Rev Genet. 2020;54:367–85. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cirillo LA, Zaret KS. An early developmental transcription factor complex that is more stable on nucleosome core particles than on free DNA. Mol Cell. 1999;4(6):961–9. [DOI] [PubMed] [Google Scholar]
- 41.Dodonova SO, Zhu F, Dienemann C, Taipale J, Cramer P. Nucleosome-bound SOX2 and SOX11 structures elucidate pioneer factor function. Nature. 2020;580(7805):669–72. [DOI] [PubMed] [Google Scholar]
- 42.Gong Y, Lazaris C, Sakellaropoulos T, Lozano A, Kambadur P, Ntziachristos P, Aifantis I, Tsirigos A. Stratification of TAD boundaries reveals preferential insulation of super-enhancers by strong boundaries. Nat Commun. 2018;9(1):542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Michael AK, Grand RS, Isbel L, Cavadini S, Kozicka Z, Kempf G, Bunker RD, Schenk AD, Graff-Meyer A, Pathare GR, et al. Mechanisms of OCT4-SOX2 motif readout on nucleosomes. Science. 2020;368(6498):1460–5. [DOI] [PubMed] [Google Scholar]
- 44.Adam RC, Yang H, Rockowitz S, Larsen SB, Nikolova M, Oristian DS, Polak L, Kadaja M, Asare A, Zheng D, et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature. 2015;521(7552):366–70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Pelletier A, Mayran A, Gouhier A, Omichinski JG, Balsalobre A, Drouin J. Pax7 pioneer factor action requires both paired and homeo DNA binding domains. Nucleic Acids Res. 2021;49(13):7424–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Balsalobre A, Drouin J. Pioneer factors as master regulators of the epigenome and cell fate. Nat Rev Mol Cell Biol. 2022;23(7):449–64. [DOI] [PubMed] [Google Scholar]
- 47.Kang Y, Kang J, Kim A. Histone H3K4me1 strongly activates the DNase I hypersensitive sites in super-enhancers than those in typical enhancers. Biosci Rep 2021, 41(7). [DOI] [PMC free article] [PubMed]
- 48.Mayran A, Khetchoumian K, Hariri F, Pastinen T, Gauthier Y, Balsalobre A, Drouin J. Pioneer factor Pax7 deploys a stable enhancer repertoire for specification of cell fate. Nat Genet. 2018;50(2):259–69. [DOI] [PubMed] [Google Scholar]
- 49.Wang C, Lee JE, Lai B, Macfarlan TS, Xu S, Zhuang L, Liu C, Peng W, Ge K. Enhancer priming by H3K4 methyltransferase MLL4 controls cell fate transition. Proc Natl Acad Sci U S A. 2016;113(42):11871–6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Dorighi KM, Swigut T, Henriques T, Bhanu NV, Scruggs BS, Nady N, Still CD 2nd, Garcia BA, Adelman K, Wysocka J. Mll3 and Mll4 facilitate enhancer RNA synthesis and transcription from promoters independently of H3K4 Monomethylation. Mol Cell. 2017;66(4):568–e576564. [DOI] [PMC free article] [PubMed]
- 51.Lai B, Lee JE, Jang Y, Wang L, Peng W, Ge K. MLL3/MLL4 are required for CBP/p300 binding on enhancers and super-enhancer formation in brown adipogenesis. Nucleic Acids Res. 2017;45(11):6388–403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Haberland M, Montgomery RL, Olson EN. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat Rev Genet. 2009;10(1):32–42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Nguyen TTT, Zhang Y, Shang E, Shu C, Torrini C, Zhao J, Bianchetti E, Mela A, Humala N, Mahajan A, et al. HDAC inhibitors elicit metabolic reprogramming by targeting super-enhancers in glioblastoma models. J Clin Invest. 2020;130(7):3699–716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Donati B, Lorenzini E, Ciarrocchi A. BRD4 and Cancer: going beyond transcriptional regulation. Mol Cancer. 2018;17(1):164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Lee JE, Park YK, Park S, Jang Y, Waring N, Dey A, Ozato K, Lai B, Peng W, Ge K. Brd4 binds to active enhancers to control cell identity gene induction in adipogenesis and myogenesis. Nat Commun. 2017;8(1):2217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hyman AA, Weber CA, Jülicher F. Liquid-liquid phase separation in biology. Annu Rev Cell Dev Biol. 2014;30:39–58. [DOI] [PubMed] [Google Scholar]
- 57.Mirny LA, Imakaev M, Abdennur N. Two major mechanisms of chromosome organization. Curr Opin Cell Biol. 2019;58:142–52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sabari BR, Dall’Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K, Abraham BJ, Hannett NM, Zamudio AV, Manteiga JC et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science 2018, 361(6400). [DOI] [PMC free article] [PubMed]
- 59.Shin Y, Chang YC, Lee DSW, Berry J, Sanders DW, Ronceray P, Wingreen NS, Haataja M, Brangwynne CP. Liquid Nuclear condensates mechanically sense and restructure the genome. Cell. 2018;175(6):1481–e14911413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Wei MT, Elbaum-Garfinkle S, Holehouse AS, Chen CC, Feric M, Arnold CB, Priestley RD, Pappu RV, Brangwynne CP. Phase behaviour of disordered proteins underlying low density and high permeability of liquid organelles. Nat Chem. 2017;9(11):1118–25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Dixon JR, Selvaraj S, Yue F, Kim A, Li Y, Shen Y, Hu M, Liu JS, Ren B. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature. 2012;485(7398):376–80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Dowen JM, Fan ZP, Hnisz D, Ren G, Abraham BJ, Zhang LN, Weintraub AS, Schujiers J, Lee TI, Zhao K, et al. Control of cell identity genes occurs in insulated neighborhoods in mammalian chromosomes. Cell. 2014;159(2):374–87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Fudenberg G, Imakaev M, Lu C, Goloborodko A, Abdennur N, Mirny LA. Formation of chromosomal domains by Loop Extrusion. Cell Rep. 2016;15(9):2038–49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Hansen AS, Pustova I, Cattoglio C, Tjian R, Darzacq X. CTCF and cohesin regulate chromatin loop stability with distinct dynamics. Elife 2017, 6. [DOI] [PMC free article] [PubMed]
- 65.Sanborn AL, Rao SS, Huang SC, Durand NC, Huntley MH, Jewett AI, Bochkov ID, Chinnappan D, Cutkosky A, Li J, et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc Natl Acad Sci U S A. 2015;112(47):E6456–6465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Willi M, Yoo KH, Reinisch F, Kuhns TM, Lee HK, Wang C, Hennighausen L. Facultative CTCF sites moderate mammary super-enhancer activity and regulate juxtaposed gene in non-mammary cells. Nat Commun. 2017;8:16069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Vogel A, Meyer T, Sapisochin G, Salem R, Saborowski A. Hepatocellular carcinoma. Lancet. 2022;400(10360):1345–62. [DOI] [PubMed] [Google Scholar]
- 68.Witte S, Bradley A, Enright AJ, Muljo SA. High-density P300 enhancers control cell state transitions. BMC Genomics. 2015;16:903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Shida D, Takabe K, Kapitonov D, Milstien S, Spiegel S. Targeting SphK1 as a new strategy against cancer. Curr Drug Targets. 2008;9(8):662–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Tsang FH, Law CT, Tang TC, Cheng CL, Chin DW, Tam WV, Wei L, Wong CC, Ng IO, Wong CM. Aberrant Super-enhancer Landscape in Human Hepatocellular Carcinoma. Hepatology. 2019;69(6):2502–17. [DOI] [PubMed] [Google Scholar]
- 71.Liu B, Dou J, Cao J. Nuclear respiratory factor 1 regulates super enhancer-controlled SPIDR to protect hepatocellular carcinoma cells from oxidative stress. BMC Gastroenterol. 2024;24(1):97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Moreno-Bueno G, Portillo F, Cano A. Transcriptional regulation of cell polarity in EMT and cancer. Oncogene. 2008;27(55):6958–69. [DOI] [PubMed] [Google Scholar]
- 73.Thiery JP, Acloque H, Huang RY, Nieto MA. Epithelial-mesenchymal transitions in development and disease. Cell. 2009;139(5):871–90. [DOI] [PubMed] [Google Scholar]
- 74.Zhang C, Wei S, Sun WP, Teng K, Dai MM, Wang FW, Chen JW, Ling H, Ma XD, Feng ZH, et al. Super-enhancer-driven AJUBA is activated by TCF4 and involved in epithelial-mesenchymal transition in the progression of Hepatocellular Carcinoma. Theranostics. 2020;10(20):9066–82. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Han J, Meng J, Chen S, Wang X, Yin S, Zhang Q, Liu H, Qin R, Li Z, Zhong W, et al. YY1 Complex promotes quaking expression via Super-enhancer binding during EMT of Hepatocellular Carcinoma. Cancer Res. 2019;79(7):1451–64. [DOI] [PubMed] [Google Scholar]
- 76.Wu F, Xu L, Tu Y, Cheung OK, Szeto LL, Mok MT, Yang W, Kang W, Cao Q, Lai PB, et al. Sirtuin 7 super-enhancer drives epigenomic reprogramming in hepatocarcinogenesis. Cancer Lett. 2022;525:115–30. [DOI] [PubMed] [Google Scholar]
- 77.Hombach S, Kretz M. Non-coding RNAs: classification, Biology and Functioning. Adv Exp Med Biol. 2016;937:3–17. [DOI] [PubMed] [Google Scholar]
- 78.Meng J, Han J, Wang X, Wu T, Zhang H, An H, Qin L, Sun Y, Zhong W, Yang C, et al. Twist1-YY1-p300 complex promotes the malignant progression of HCC through activation of miR-9 by forming phase-separated condensates at super-enhancers and relieved by metformin. Pharmacol Res. 2023;188:106661. [DOI] [PubMed] [Google Scholar]
- 79.Li J, Wang J, Wang Y, Zhao X, Su T. E2F1 combined with LINC01004 super-enhancer to promote hepatocellular carcinoma cell proliferation and metastasis. Clin Epigenetics. 2023;15(1):17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Peng L, Jiang B, Yuan X, Qiu Y, Peng J, Huang Y, Zhang C, Zhang Y, Lin Z, Li J, et al. Super-enhancer-associated long noncoding RNA HCCL5 is activated by ZEB1 and promotes the malignancy of Hepatocellular Carcinoma. Cancer Res. 2019;79(3):572–84. [DOI] [PubMed] [Google Scholar]
- 81.Su T, Zhang N, Wang T, Zeng J, Li W, Han L, Yang M. Super enhancer-regulated LncRNA LINC01089 induces alternative splicing of DIAPH3 to Drive Hepatocellular Carcinoma Metastasis. Cancer Res. 2023;83(24):4080–94. [DOI] [PubMed] [Google Scholar]
- 82.Liang W, Shi C, Hong W, Li P, Zhou X, Fu W, Lin L, Zhang J. Super-enhancer-driven lncRNA-DAW promotes liver cancer cell proliferation through activation of Wnt/β-catenin pathway. Mol Ther Nucleic Acids. 2021;26:1351–63. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Yuan XQ, Zhou N, Wang JP, Yang XZ, Wang S, Zhang CY, Li GC, Peng L. Anchoring super-enhancer-driven oncogenic lncRNAs for anti-tumor therapy in hepatocellular carcinoma. Mol Ther. 2023;31(6):1756–74. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Jiang YY, Jiang Y, Li CQ, Zhang Y, Dakle P, Kaur H, Deng JW, Lin RY, Han L, Xie JJ, et al. TP63, SOX2, and KLF5 establish a Core Regulatory Circuitry that controls epigenetic and transcription patterns in esophageal squamous cell Carcinoma Cell lines. Gastroenterology. 2020;159(4):1311–e13271319. [DOI] [PubMed] [Google Scholar]
- 85.Wang SW, Gao C, Zheng YM, Yi L, Lu JC, Huang XY, Cai JB, Zhang PF, Cui YH, Ke AW. Current applications and future perspective of CRISPR/Cas9 gene editing in cancer. Mol Cancer. 2022;21(1):57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Zheng C, Liu M, Ge Y, Qian Y, Fan H. HBx increases chromatin accessibility and ETV4 expression to regulate dishevelled-2 and promote HCC progression. Cell Death Dis. 2022;13(2):116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Lin CH, Kuo JC, Li D, Koenig AB, Pan A, Yan P, Bai XF, Lee RJ, Ghoshal K. AZD5153, a bivalent BRD4 inhibitor, suppresses Hepatocarcinogenesis by Altering BRD4 Chromosomal Landscape and modulating the transcriptome of HCC cells. Front Cell Dev Biol. 2022;10:853652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Sakamoto H, Ando K, Imaizumi Y, Mishima H, Kinoshita A, Kobayashi Y, Kitanosono H, Kato T, Sawayama Y, Sato S, et al. Alvocidib inhibits IRF4 expression via super-enhancer suppression and adult T-cell leukemia/lymphoma cell growth. Cancer Sci. 2022;113(12):4092–103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Dhalluin C, Carlson JE, Zeng L, He C, Aggarwal AK, Zhou MM. Structure and ligand of a histone acetyltransferase bromodomain. Nature. 1999;399(6735):491–6. [DOI] [PubMed] [Google Scholar]
- 90.Filippakopoulos P, Picaud S, Mangos M, Keates T, Lambert JP, Barsyte-Lovejoy D, Felletar I, Volkmer R, Müller S, Pawson T, et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell. 2012;149(1):214–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Morinière J, Rousseaux S, Steuerwald U, Soler-López M, Curtet S, Vitte AL, Govin J, Gaucher J, Sadoul K, Hart DJ, et al. Cooperative binding of two acetylation marks on a histone tail by a single bromodomain. Nature. 2009;461(7264):664–8. [DOI] [PubMed] [Google Scholar]
- 92.Wu SY, Chiang CM. The double bromodomain-containing chromatin adaptor Brd4 and transcriptional regulation. J Biol Chem. 2007;282(18):13141–5. [DOI] [PubMed] [Google Scholar]
- 93.Jang MK, Mochizuki K, Zhou M, Jeong HS, Brady JN, Ozato K. The bromodomain protein Brd4 is a positive regulatory component of P-TEFb and stimulates RNA polymerase II-dependent transcription. Mol Cell. 2005;19(4):523–34. [DOI] [PubMed] [Google Scholar]
- 94.Jiang YW, Veschambre P, Erdjument-Bromage H, Tempst P, Conaway JW, Conaway RC, Kornberg RD. Mammalian mediator of transcriptional regulation and its possible role as an end-point of signal transduction pathways. Proc Natl Acad Sci U S A. 1998;95(15):8538–43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Yang Z, Yik JH, Chen R, He N, Jang MK, Ozato K, Zhou Q. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein Brd4. Mol Cell. 2005;19(4):535–45. [DOI] [PubMed] [Google Scholar]
- 96.Lovén J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, Bradner JE, Lee TI, Young RA. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell. 2013;153(2):320–34. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Zhang P, Dong Z, Cai J, Zhang C, Shen Z, Ke A, Gao D, Fan J, Shi G. BRD4 promotes tumor growth and epithelial-mesenchymal transition in hepatocellular carcinoma. Int J Immunopathol Pharmacol. 2015;28(1):36–44. [DOI] [PubMed] [Google Scholar]
- 98.Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, Morse EM, Keates T, Hickman TT, Felletar I, et al. Selective inhibition of BET bromodomains. Nature. 2010;468(7327):1067–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Choi HI, An GY, Baek M, Yoo E, Chai JC, Lee YS, Jung KH, Chai YG. BET inhibitor suppresses migration of human hepatocellular carcinoma by inhibiting SMARCA4. Sci Rep. 2021;11(1):11799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Delmore JE, Issa GC, Lemieux ME, Rahl PB, Shi J, Jacobs HM, Kastritis E, Gilpatrick T, Paranal RM, Qi J, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146(6):904–17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Qi J. Bromodomain and extraterminal domain inhibitors (BETi) for cancer therapy: chemical modulation of chromatin structure. Cold Spring Harb Perspect Biol. 2014;6(12):a018663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Shi X, Liu C, Liu B, Chen J, Wu X, Gong W. JQ1: a novel potential therapeutic target. Pharmazie. 2018;73(9):491–3. [DOI] [PubMed] [Google Scholar]
- 103.Trabucco SE, Gerstein RM, Evens AM, Bradner JE, Shultz LD, Greiner DL, Zhang H. Inhibition of bromodomain proteins for the treatment of human diffuse large B-cell lymphoma. Clin Cancer Res. 2015;21(1):113–22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Moyer MW. First drugs found to inhibit elusive cancer target. Nat Med. 2011;17(11):1325. [DOI] [PubMed] [Google Scholar]
- 105.Matzuk MM, McKeown MR, Filippakopoulos P, Li Q, Ma L, Agno JE, Lemieux ME, Picaud S, Yu RN, Qi J, et al. Small-molecule inhibition of BRDT for male contraception. Cell. 2012;150(4):673–84. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Malumbres M. Cyclin-dependent kinases. Genome Biol. 2014;15(6):122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Duan J, He Y, Fu X, Deng Y, Zheng M, Lu D. CDK7 activated beta-catenin/TCF signaling in hepatocellular carcinoma. Exp Cell Res. 2018;370(2):461–7. [DOI] [PubMed] [Google Scholar]
- 108.Shen S, Dean DC, Yu Z, Duan Z. Role of cyclin-dependent kinases (CDKs) in hepatocellular carcinoma: therapeutic potential of targeting the CDK signaling pathway. Hepatol Res. 2019;49(10):1097–108. [DOI] [PubMed] [Google Scholar]
- 109.Kwiatkowski N, Zhang T, Rahl PB, Abraham BJ, Reddy J, Ficarro SB, Dastur A, Amzallag A, Ramaswamy S, Tesar B, et al. Targeting transcription regulation in cancer with a covalent CDK7 inhibitor. Nature. 2014;511(7511):616–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 110.Brès V, Yoh SM, Jones KA. The multi-tasking P-TEFb complex. Curr Opin Cell Biol. 2008;20(3):334–40. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Gerlach D, Tontsch-Grunt U, Baum A, Popow J, Scharn D, Hofmann MH, Engelhardt H, Kaya O, Beck J, Schweifer N, et al. The novel BET bromodomain inhibitor BI 894999 represses super-enhancer-associated transcription and synergizes with CDK9 inhibition in AML. Oncogene. 2018;37(20):2687–701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Thieme E, Bruss N, Sun D, Dominguez EC, Coleman D, Liu T, Roleder C, Martinez M, Garcia-Mansfield K, Ball B, et al. CDK9 inhibition induces epigenetic reprogramming revealing strategies to circumvent resistance in lymphoma. Mol Cancer. 2023;22(1):64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Yao J, Wang J, Xu Y, Guo Q, Sun Y, Liu J, Li S, Guo Y, Wei L. CDK9 inhibition blocks the initiation of PINK1-PRKN-mediated mitophagy by regulating the SIRT1-FOXO3-BNIP3 axis and enhances the therapeutic effects involving mitochondrial dysfunction in hepatocellular carcinoma. Autophagy. 2022;18(8):1879–97. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Yao JY, Xu S, Sun YN, Xu Y, Guo QL, Wei LB. Novel CDK9 inhibitor oroxylin a promotes wild-type P53 stability and prevents hepatocellular carcinoma progression by disrupting both MDM2 and SIRT1 signaling. Acta Pharmacol Sin. 2022;43(4):1033–45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Delvecchio M, Gaucher J, Aguilar-Gurrieri C, Ortega E, Panne D. Structure of the p300 catalytic core and implications for chromatin targeting and HAT regulation. Nat Struct Mol Biol. 2013;20(9):1040–6. [DOI] [PubMed] [Google Scholar]
- 116.Ogryzko VV, Schiltz RL, Russanova V, Howard BH, Nakatani Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell. 1996;87(5):953–9. [DOI] [PubMed] [Google Scholar]
- 117.Zhang Y, Xue Y, Shi J, Ahn J, Mi W, Ali M, Wang X, Klein BJ, Wen H, Li W, et al. The ZZ domain of p300 mediates specificity of the adjacent HAT domain for histone H3. Nat Struct Mol Biol. 2018;25(9):841–9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 118.Cai LY, Chen SJ, Xiao SH, Sun QJ, Ding CH, Zheng BN, Zhu XY, Liu SQ, Yang F, Yang YX, et al. Targeting p300/CBP Attenuates Hepatocellular Carcinoma Progression through Epigenetic Regulation of Metabolism. Cancer Res. 2021;81(4):860–72. [DOI] [PubMed] [Google Scholar]
- 119.Hammitzsch A, Tallant C, Fedorov O, O’Mahony A, Brennan PE, Hay DA, Martinez FO, Al-Mossawi MH, de Wit J, Vecellio M, et al. CBP30, a selective CBP/p300 bromodomain inhibitor, suppresses human Th17 responses. Proc Natl Acad Sci U S A. 2015;112(34):10768–73. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Data Availability Statement
No datasets were generated or analysed during the current study.