Abstract
Genomic instability underlies many cancers and generates genetic variation that drives cancer initiation, progression, and therapy resistance. In contrast with classical assumptions that mutations occur purely stochastically at constant, gradual rates, microbes, plants, flies, and human cancer cells possess mechanisms of mutagenesis that are upregulated by stress responses. These generate transient, genetic-diversity bursts that can propel evolution, specifically when cells are poorly adapted to their environments—that is, when stressed. We review molecular mechanisms of stress-response-dependent (stress-induced) mutagenesis that occur from bacteria to cancer, and are activated by starvation, drugs, hypoxia, and other stressors. We discuss mutagenic DNA break repair in Escherichia coli as a model for mechanisms in cancers. The temporal regulation of mutagenesis by stress responses and spatial restriction in genomes are common themes across the tree of life. Both can accelerate evolution, including the evolution of cancers. We discuss possible anti-evolvability drugs, aimed at targeting mutagenesis and other variation generators, that could be used to delay the evolution of cancer progression and therapy resistance.
Keywords: chemotherapy, double-strand break repair, evolution, HSP90, hypoxia, kataegis, genome instability, MMBIR, stress response, trinucleotide repeat instability
INTRODUCTION
In the 1930s and 1940s, biologists fused Charles Darwin’s principles of evolution by variation and selection (1859) with Gregor Mendel’s principles of genetic inheritance (1865). They articulated (Huxley 1942) that gene mutations create variation between individuals and the fittest proliferate. Predating molecular biology, evolutionary biologists could not know that genes were DNA (Avery et al. 1944, Hershey & Chase 1952), how DNA replicated (Meselson & Stahl 1958), nor how mutations arise. So they tentatively modeled mutation formation simply (attractively!) as a stochastic, probabilistic process, constant and gradual over time (Mayr 1985): a ticking clock that, with enough time, could be essentially limitless.
Nowell (1976), and then others, modeled cancer development as an evolutionary process (reviewed in Greaves & Maley 2012, Yates & Campbell 2012, Gerlinger et al. 2014). Cells acquire characteristics of cancer by multiple heritable changes (Fearon & Vogelstein 1990, Fishel et al. 1993), many of them genetic. Genomic instability fuels many cancers (reviewed in Lengauer et al. 1998, Lee et al. 2016) by increasing the probability of mutations, which drive every stage of cancer progression. The origin of mutations is, therefore, fundamental to cancer biology. Mutations also underpin the evolutionary arms races between the chemotherapies we develop and the cancer cells and pathogens that out-evolve them.
Stress-induced mutagenesis encompasses molecular mechanisms of abrupt, transient genomic instability that are upregulated by stress responses, as reviewed here. These mechanisms produce mutations nonrandomly, preferentially when cells are maladapted to their environments—when stressed—and can accelerate adaptation in changing environments (Ram & Hadany 2012, 2014). In some mechanisms, the mutations also occur nonrandomly in genomic space: in localized clusters (Lin et al. 2011, Shee et al. 2012), which could speed concerted evolution within genes (Ponder et al. 2005, Ram & Hadany 2014), a major problem in the evolution of new protein functions (Camps et al. 2003). Mutation clusters occur across phylogeny (Drake et al. 2005), including in mutation storms in cancer genomes (Nik-Zainal et al. 2012, Roberts et al. 2012, Alexandrov et al. 2013).
We review molecular mechanisms of stress-induced mutagenesis across the tree of life, induced by starvation, drugs, hypoxia, protein misfolding, or other stressors and upregulated by various stress responses. These illustrate that virtually all types of mutations observed in human cancer genomes could result from stress-induced mutagenesis. We discuss their implications for cancer and drug resistance and consider potential novel anti-evolvability drugs that could target genomic instability and other variation generators that drive cancer evolution.
HISTORICAL PERSPECTIVE
Harrison Echols (1981) discussed the idea of stress-response “inducible evolution” when considering the Escherichia coli SOS DNA damage response. He, and previously Barbara McClintock (summarized in McClintock 1978), thought that cells could sense stress and adapt by activating a burst of upregulated genomic instability, which Echols discussed as a paradigm shift for evolution. Both proposals followed Evelyn Witkin’s (1967; Witkin & George 1973) and Miroslav Radman’s (1974, 1975) articulation of the SOS response, in which cells upregulate DNA repair, mutagenesis, prophage induction, and cell division inhibition. Others still argue that cells induce SOS to repair DNA, and mutagenesis is an unavoidable by-product (i.e., nonmutagenic DNA repair cannot evolve; e.g., Lynch 2010, Andersson & Hughes 2014). Of course, both sides may be right: SOS is needed for DNA repair, but it also could accelerate adaptation (modeled in Ram & Hadany 2012, 2014). During the past 15 years, this argument has been transcended by discoveries of molecular mechanisms of mutagenesis that are upregulated by stress responses not needed for DNA repair.
MUTAGENIC BREAK REPAIR IN ESCHERICHIA COLI MODELS CANCER PROCESSES
Discovered in E. coli (Harris et al. 1994, Rosenberg et al. 1994), mutagenic DNA break repair upregulates mutation mechanisms temporally, during stress, and nonrandomly in genomic space, and its mechanisms are illuminating similar mechanisms in cancers.
Two Mutagenic Break Repair Mechanisms
In starving E. coli (Cairns & Foster 1991, Al Mamun et al. 2012), double-strand break (DSB) repair becomes mutagenic, depending on the general stress response (Ponder et al. 2005, Shee et al. 2011), by either of two mechanisms. Homologous recombination (HR)–based mutagenic break repair (MBR) (Figure 1a–f) generates base substitutions and indels (Foster & Trimarchi 1994, Rosenberg et al. 1994, Ponder et al. 2005, Petrosino et al. 2009, Shee et al. 2011, Al Mamun et al. 2012) (Figure 1). An alternative break-induced microhomologous MBR mechanism produces copy-number alterations (CNAs) (Figure 1h) and other genomic rearrangements (Hastings et al. 2000, Slack et al. 2006, Lin et al. 2011). Both mechanisms are useful models for human cancer (e.g., Mihaylova et al. 2003, Bindra et al. 2005). They mirror the two kinds of genomic instability seen in many cancers (reviewed in Lengauer et al. 1998, Lee et al. 2016), which could result from stress-induced mutagenesis.
Figure 1.
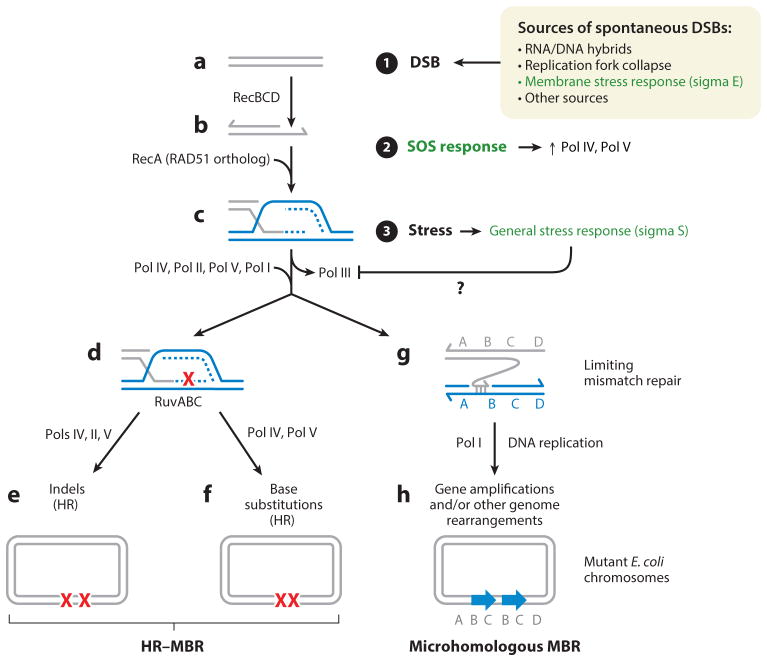
Models of Escherichia coli MBR. (➀) DSBs arise via many routes. (a–c) RecBCD nuclease (an analog of human BRCA2) loads RecA HR protein (an ortholog of human RAD51) onto ssDNA, analogously to human BRCA2 loading RAD51, facilitating base-pairing with a strand of identical duplex DNA (blue) (e.g., a sister chromosome). Parallel lines represent base-paired DNA strands; arrowheads represent 3′ DNA ends. Repair synthesis (dashed lines) is switched to a mutagenic mode by the general stress response (sigma S; ➂). (d–f ) In HR-MBR, a switch to mutagenic HR repair of DSBs occurs when the SOS DNA damage response and general stress response are activated. (➁) The SOS response upregulates error-prone Pols IV, V, and II. (➂) The general stress response licenses the use of, or errors made by, alternative DNA polymerases in repair synthesis by as yet unknown means. DSB repair is otherwise high fidelity and dependent on high-fidelity DNA Pol III. HR-MBR produces (e) indels and (f) base substitutions, both dependent on Pol IV. Pol V contributes to some indels, and base substitutions. Pol II contributes to some indels. The red Xs represent DNA polymerase errors that become mutations. ( g,h) Microhomologous MBR requires DNA Pol I for template switching to regions containing microhomology ( g) and initiates replication, creating genome rearrangements (h) including duplications (blue arrows). Microhomologous MBR requires steps ➀ and ➂, but not the SOS response. Abbreviations: DSB, double-strand break; HR, homologous recombination; MBR, mutagenic break repair; Pol, DNA polymerase; SS, single-stranded.
Three Stress Responses and Three Events Trigger Homologous Recombinational Mutagenic Break Repair
HR-MBR requires three simultaneous events (Ponder et al. 2005, Shee et al. 2011) (Figure 1, steps 1–3): a DSB (some promoted by the sigma E stress response) and its repair using HR (step 1); activation of the SOS DNA damage response (step 2); and activation of the general (sigma S) stress response, which is triggered by many different stressors (step 3). These events occur as follows.
In step 1, spontaneous DSBs occur in ≤1% of growing E. coli and are estimated to form similarly per base pair in human genomes (Pennington & Rosenberg 2007, Shee et al. 2013). The sigma E membrane stress response is required for MBR at some loci (Figure 1, step 1), can be substituted by the delivery of a DSB near the mutation reporter gene assayed, and so promotes spontaneous DSBs, by unknown means (Gibson et al. 2010; discussed in Rogers et al. 2016). DSB repair (reviewed in Kuzminov 2013) uses human cancer protein analogs and orthologs (Figure 1) (Aravind et al. 1999). RecBCD (an analog of human BRCA2) loads RecA (an ortholog of human RAD51) onto single-stranded DNA at DSBs (Figure 1b); identical strands are exchanged (with, e.g., a sister chromosome), and repair synthesis or replication (Figure 1c) replaces eroded DNA. In unstressed cells, HR-DSB repair is relatively nonmutagenic (Ponder et al. 2005, Shee et al. 2011), using the high-fidelity major replicative DNA polymerase (Pol) III (Motamedi et al. 1999).
In step 2, the SOS DNA damage response is induced in about 25% of cells with a reparable DSB (Pennington & Rosenberg 2007), and so follows DNA breakage (Figure 1, steps 1 and 2). SOS promotes HR-MBR (Cairns & Foster 1991, McKenzie et al. 2000) by its 10-fold transcriptional upregulation of the Pol IV (DinB) error-prone DNA polymerase (Galhardo et al. 2009). Pol IV is required for most HR-MBR-generated base substitutions and indels (McKenzie et al. 2001, Ponder et al. 2005, Shee et al. 2011) (Figure 1e, f). Artificial production of Pol IV to SOS-induced levels substitutes for a functional SOS response (Galhardo et al. 2009). SOS also allows production of error-prone DNA Pol V, which is used in MBR at some sites (Petrosino et al. 2009, Shee et al. 2011), and increases Pol II, which promotes some MBR indels (Frisch et al. 2010). Although SOS-induced cells have 10 times more Pol IV and produce Pol V, the repair of DSBs is still relatively nonmutagenic unless the general stress response is also induced (Ponder et al. 2005, Shee et al. 2011, Al Mamun et al. 2012). Whereas it has long been argued that nonmutagenic DNA repair could not evolve (reviewed in the section Historical Perspective, above), the ability to remove the mutagenic component from MBR without reducing repair demonstrates that this is not the case (see the sidebar High-Fidelity Break Repair Has Evolved—Mutagenesis Not Needed).
HIGH-FIDELITY BREAK REPAIR HAS EVOLVED—MUTAGENESIS NOT NEEDED.
Mutants that lack functional Pol IV survive endonuclease-induced DSBs as well as, or better than, isogenic Pol IV–proficient cells (Ponder et al. 2005, Shee et al. 2011). This counters arguments that nonmutagenic repair could not evolve (see the section Historical Perspective) (e.g., Lynch 2010, Andersson & Hughes 2014). High-fidelity DSB repair has evolved; E. coli has it. It is efficient and Pol IV appears to be used for reasons other than DSB survival or repair. Instead, E. coli might use Pol IV in DSB repair because the generation of variation during stress is itself selected (Ram & Hadany 2012, 2014).
In step 3, the general stress response throws a switch that allows mutagenic DSB repair. The general (sigma S) stress response is activated by starvation, cold, acid, and oxidative and osmotic stresses, and it directly or indirectly up- or downregulates the transcription of about 10% of E. coli genes (hundreds of genes) (reviewed in Battesti et al. 2011). Sigma S is a master regulator of stress responses that may be analogous to human mTOR (mammalian target of rapamycin). When induced by either stress or artificial means, sigma S licenses the use of error-prone DNA Pols IV, II, and V, and/or errors made by, these Pols in the HR repair of site-specific endonuclease-induced and spontaneous DSBs in living cells (Figure 1, step 3) (Ponder et al. 2005, Shee et al. 2011). This demonstrates a sigma S–controlled switch to mutagenic HR-DSB repair. Base substitutions and indels result.
Purified Pol IV in solution can load at a D loop HR-intermediate-like structure (Figure 1c), catalyze DNA synthesis, and produce errors such as those seen in HR-MBR (Figure 1d–f) (Pomerantz et al. 2013). These data provide biochemical support for the HR-MBR mechanism.
Sigma S also allows DNA Pol I–dependent microhomologous MBR at endonuclease-induced (Ponder et al. 2005) and spontaneous DSBs (see the section Microhomologous Mutagenic Break Repair, below) (Figure 1c,g,h). It is unknown which sigma S–controlled gene or genes allow either MBR mechanism and how this occurs. One hypothesis is that the sigma S response biases DNA polymerase competition toward DNA polymerases other than Pol III at the DSB repair replisome (Figure 1) (Frisch et al. 2010). Other mechanisms are possible.
Mismatch Repair Becomes Limiting During Homologous Recombinational Mutagenic Break Repair
DNA mismatch repair (MMR) is conserved from bacteria to humans, acts after replication to correct DNA polymerase errors, and also prevents genomic rearrangements by preventing HR between imperfectly identical DNA sequences (reviewed in Li 2008). Humans with mutations in the orthologs of bacterial MMR genes display high mutation rates and Lynch syndrome colon and ovarian cancers (Fishel et al. 1993).
Transient downregulation or limitation of MMR was suggested (Stahl 1988) and then discovered in E. coli MBR. First, HR-MBR mutation sequences (Foster & Trimarchi 1994, Harris et al. 1994, Rosenberg et al. 1994) mimic mutations in unstressed MMR-defective mutant cells (Longerich et al. 1995), even though most MBR-generated mutants are MMR proficient once formed (Longerich et al. 1995, Torkelson et al. 1997, Rosenberg et al. 1998). This implies that there is transient MMR deficiency. Second, the overproduction of the MMR protein MutL inhibits HR-MBR, as if MutL were limiting for MMR during HR-MBR, but MutL protein levels are not decreased, whereas MutS and MutH are (Harris et al. 1997). The general stress response downregulates MutS and MutH protein levels (Tsui et al. 1997). How MMR becomes limiting transiently during MBR remains unsolved, although the finding of this transient limitation led to the discovery of the transcriptional downregulation of human MutL and MutS orthologs by hypoxic stress responses, which causes genomic instability in human cancer cells (e.g., Mihaylova et al. 2003; see the section Hypoxic Stress Responses Downregulate Mismatch Repair, below), and the discovery of MMR downregulation in other stress-induced mutation mechanisms (Brégeon et al. 1999, Bjedov et al. 2003, Pedraza-Reyes & Yasbin 2004, Gutierrez et al. 2013).
NONRANDOM MUTATIONS IN GENOMIC SPACE
Homologous Recombinational Mutagenic Break Repair Generates Mutation Clusters at Double-Strand Breaks Similar to Kataegis in Cancers
HR-MBR-generated mutations promoted by site-specific endonuclease-generated DSBs occur mostly in the first kilobase (kb) pair of the DSB, in hot spots that fall off to approximately 60 kb on either side (Shee et al. 2012), and occur in clusters (Bull et al. 2000). Similarly, roughly 100 kb showers of multiple mutations were found concurrently in human cancer genomes and inferred to result from MBR (Nik-Zainal et al. 2012, Roberts et al. 2012, Alexandrov et al. 2013). Mutation clusters in cancers are called kataegis (Greek for storms). Mutation clusters in yeast HR-MBR (Sakofsky et al. 2014) further support HR-MBR causing kataegis in cancers. Moreover, HR-MBR is a mechanism that E. coli normally and frequently uses (see the sidebar HR-MBR: A Mechanism Normally Used).
Mutation clusters could promote concerted evolution via multiple simultaneous mutations within genes or linked gene families (Ponder et al. 2005, Ram & Hadany 2014). The need for multiple mutations in the evolution of new protein functions is a major problem (Camps et al. 2003), potentially addressed by the observation that coincident nearby mutations occur across phylogeny (Drake et al. 2005).
Are Spontaneous Double-Strand Breaks Nonrandom in Space?
The genomic locations of spontaneous DSBs in somatic cells are unknown, and they might or might not be random. Few mechanisms of spontaneous DNA breakage are known (reviewed in Merrikh et al. 2012, Shee et al. 2013). Some spontaneous DSBs in E. coli result from RNA-DNA hybrids generated by transcription (R loops) (Figure 1a, step 1) (Wimberly et al. 2013) and the sigma E membrane stress response (Gibson et al. 2010). Transcriptional R loops also promote DNA damage in yeast and other organisms (reviewed in Santos-Pereira & Aguilera 2015). The connection of DSBs with transcription raises the possibility that MBR hot spots might fall in or near active genes, potentially biasing mutations to genes currently under selection, which might additionally accelerate evolution. This hypothesis remains to be tested. Caporale (2006) reviews many biological systems in which genomes appear to possess intrinsic mutation-targeting devices.
HR-MBR: A MECHANISM NORMALLY USED.
When no engineered DSBs are given to plasmid-free E. coli starved in liquid, half of the base substitutions and indels detected in two chromosomal mutation reporter gene assays required the proteins for HR-MBR: DSB repair proteins, a functional SOS response regulator, sigma S, and DNA Pols IV and V (Shee et al. 2011). Both reporter genes assayed drug-resistance mutations that were induced, but not selected, by starvation. Dependence on the DSB-specific RecB protein implies that spontaneous DSBs promote the mutagenesis. The data indicate that HR-MBR is a mechanism that E. coli normally uses. Further, the mutation signatures of HR-MBR are overrepresented in bacterial genomes, suggesting its widespread use (Maharjan & Ferenci 2015).
A LARGE MUTAGENIC BREAK REPAIR GENE NETWORK COUPLES MUTAGENESIS TO STRESS, PREDICTS DRUG TARGETS
A screen for MBR-defective mutants discovered a network of more than 93 genes that promote MBR, of which only 16 had been identified previously (Al Mamun et al. 2012). Using mechanism-directed functional tests, more than half the genes were assigned or implicated in MBR roles upstream of the activation of the three key stress-response regulators: sigma S, sigma E, and the SOS response. The data show that sensing and signaling stress are central to mutagenesis. Electron transfer, the largest class of MBR network genes discovered, upregulates sigma S, indicating that cells sense starvation and transduce the signal via electron transfer. The three stress-response regulators are nonredundant network hubs (Al Mamun et al. 2012) and so are potentially vulnerable targets for proposed new drugs aimed at slowing the evolution of pathogens and stress-induced evolution in cancer (see the section Anti-Evolvability Drugs, below) (Al Mamun et al. 2012, Rosenberg & Queitsch 2014).
MICROHOMOLOGOUS MUTAGENIC BREAK REPAIR CAUSES COPY-NUMBER ALTERATIONS AND OTHER GENOMIC REARRANGEMENTS VIA MICROHOMOLOGY-MEDIATED BREAK-INDUCED REPLICATION
Microhomologous MBR, discovered in E. coli assays for gene amplification (Tlsty et al. 1984, Hastings et al. 2000), produces CNAs and other genome rearrangements (Figure 1h) that model human cancer–driving rearrangements (Hastings et al. 2000, Slack et al. 2006, Lin et al. 2011). Microhomologous MBR is stress induced, requiring the sigma S response (Lombardo et al. 2004, Ponder et al. 2005), and is initiated by and requires DSBs (Ponder et al. 2005, Wimberly et al. 2013), HR-DSB repair proteins, and DNA Pol I (Slack et al. 2006), which is used in E. coli lagging-strand synthesis. A microhomology-mediated break-induced replication (MMBIR) mechanism (Hastings et al. 2009) allowed by the sigma S response (Figure 1g,h) is supported for microhomologous MBR. In MMBIR, DSBs undergo nonhomologous repair via repeated rounds of DNA Pol I–promoted replication template switching to regions of less than 20 bp of sequence identity (Figure 1g) (Slack et al. 2006). Single-stranded 3′ DNA ends (Figure 1b), proposed to prime the repair replication (Figure 1g), are demonstrable intermediates in that the overproduction of a 3′ single-strand-dependent exonuclease reduced sigma S-dependent, break-dependent amplifications (Slack et al. 2006). Intramolecular direct tandem repeat amplifications (Hastings et al. 2000) are likely to result from additional HR-mediated expansions of the initial MMBIR-generated duplications (Figure 1h) (Slack et al. 2006). The MMBIR model developed for E. coli is illuminating human genome rearrangements in cancer and other diseases (Hastings et al. 2009, Liu et al. 2012).
MUTAGENIC BREAK REPAIR IN SUBPOPULATIONS, IN EVOLUTION, AND IN OTHER ORGANISMS
MBR is nonrandom not just in time and space but also in cell subpopulations (see the sidebar A Subpopulation Phenomenon). MBR is predicted to promote evolution and to be selected on that basis (see the sidebar Stress-Induced Mutagenesis Can Increase the Ability to Evolve). Mutation mechanisms like HR-MBR are apparent in multiple organisms and circumstances (see the sidebar Mutagenesis Mechanisms Similar to Escherichia coli Homologous Recombinational Mutagenic Break Repair and the rest of this review).
A SUBPOPULATION PHENOMENON.
MBR occurs in cell subpopulations (Torkelson et al. 1997, Gonzalez et al. 2008, Lin et al. 2011). For HR-MBR, (a) ≤1% of E. coli possess a spontaneous DSB (Pennington & Rosenberg 2007, Shee et al. 2013); (b) 10–40% of stationary-phase cells have a second chromosome copy (Åkerlund et al. 1995) and so could repair with a sister; and (c) 25% of reparable DSBs activate the SOS response (Pennington & Rosenberg 2007). The ≤10−3 of cells with all three conditions could be limited further by other heterogeneities. Mutagenesis in small cell subpopulations might be a bet-hedging strategy (discussed in Torkelson et al. 1997, Veening et al. 2008) that mitigates risk in the larger population, but allows some individuals to experiment or explore the fitness landscape. Tumors (Berman et al. 2010) and bacterial colonies (Saint-Ruf et al. 2014) show heterogeneous stress-response induction and genomic instability; bet-hedging might also promote tumor evolution.
STRESS-INDUCED MUTAGENESIS CAN INCREASE THE ABILITY TO EVOLVE.
Mathematical modeling shows that stress-inducible mutagenesis accelerates adaptation in changing environments and is selectable on that basis (Tenaillon et al. 2004; Denamur & Matic 2006; Ram & Hadany 2012, 2014). In asexual tumor populations, constitutive mutators can be counter selected because the mutator phenotype is disadvantageous after adaptation to a particular environment (Funchain et al. 2000). Tumor microenvironments fluctuate, suggesting that inducible mutability will be favored.
MUTAGENESIS MECHANISMS SIMILAR TO ESCHERICHIA COLI HOMOLOGOUS RECOMBINATIONAL MUTAGENIC BREAK REPAIR.
HR-MBR in yeast (Strathern et al. 1995) requires error-prone DNA Pol zeta (see, for example, Holbeck & Strathern 1997), which has some similarities to E. coli Pol IV. Whether stress-response regulators permit yeast HR-MBR is unknown (untested). By contrast, nonhomologous end-joining protein Ku promotes drug-induced mutagenesis in yeast that requires the MSN environmental stress response (Shor et al. 2013). Ku also promotes mutations during starvation (Heidenreich et al. 2003) and mutations in flies (Yu & McVey 2010), none of which has been tested for stress-response dependence. MSN stress-induced mutagenesis also requires error-prone DNA polymerases Rev1 and Pol zeta, similar to the E. coli HR-MBR DNA polymerases. The same yeast error-prone DNA polymerases were required for spontaneous mutagenesis in yeast (Quah et al. 1980). Perhaps yeast HR-MBR and most yeast spontaneous mutations arise in occasional stressed cells, inducing MSN or other stress responses.
ANTIBIOTIC-INDUCED MUTAGENESIS IS SIMILAR TO ESCHERICHIA COLI HOMOLOGOUS RECOMBINATIONAL MUTAGENIC BREAK REPAIR AND MODELS CANCER CHEMOTHERAPY–INDUCED MUTAGENESIS AND RESISTANCE
The anticancer drugs etoposide and doxorubicin, and quinolone antibiotics, are type II topo-isomerase inhibitors that cause DSBs (reviewed in Pommier et al. 2010). HR-MBR is implicated in mutagenesis induced by the quinolone ciprofloxacin. Ciprofloxacin-treated E. coli produced ciprofloxacin-resistant mutants rapidly (Zhang et al. 2011), depending on DSB repair proteins, SOS, and DNA Pols IV, V, and II, similar to E. coli HR-MBR (Cirz et al. 2005). This appears to occur in multigenome cells, possibly by recombining DNA (Bos et al. 2015). However, distinguishing selection from the induction of mutations is complicated (Cirz et al. 2005). Diverse antibiotics, including other quinolones, induce the sigma S response (Gutierrez et al. 2013), and some induce cross-resistance mutagenesis (Kohanski et al. 2010). A penicillin-like antibiotic induced cross-resistance mutations, requiring sigma S, Pol IV, and MutS-limited MMR in diverse pathogens (Gutierrez et al. 2013), showing antibiotic-driven stress-induced mutagenesis with some, but not all, HR-MBR-like features. Bile, a host antibacterial defense, also induced Salmonella bile–resistance mutations dependent on DSB repair proteins, SOS, and DNA Pol IV, also similar to E. coli HR-MBR (Prieto et al. 2006). Antibiotics also induce DSBs, so they might both generate DSBs (Belenky et al. 2015) and trigger sigma S–dependent mutagenesis (Gutierrez et al. 2013). Cancer therapies, such as etoposide and doxorubicin, are likely to provoke stress response–dependent error-prone repair mechanisms, as antibiotics do in bacteria (Gutierrez et al. 2013, Belenky et al. 2015) (see the sidebar Convergent Evolution of Stress-Induced Mutagenesis).
CONVERGENT EVOLUTION OF STRESS-INDUCED MUTAGENESIS.
Similar to E. coli MBR, a Bacillus subtilis mutagenesis mechanism requires starvation stress responses (Rudner et al. 1999, Sung & Yasbin 2002), a DNA Pol IV (DinB) ortholog (Sung et al. 2003), and transient MMR deficiency (Pedraza-Reyes & Yasbin 2004), and is promoted by transcription (Martin et al. 2011). HR proteins are not required (Sung & Yasbin 2002), implying convergent evolution, which is the apparent rule (see also Bjedov et al. 2003). Across species and kingdoms, the most common feature is stress-response upregulation (of different stress responses), with DSBs or error-prone DNA polymerases, or both, in some, but not all, mechanisms.
HSP90 AND OTHER STRESS RESPONSES DESTABILIZE EUKARYOTIC GENOMES
The HSP90 protein chaperone is an evolutionary capacitor that massages mutant proteins into functional conformations, but abandons them during heat stress, revealing diverse new phenotypes of the mutations that have accumulated (reviewed in Lindquist 2009). HSP90 depletion also promotes genomic instability.
Aneuploidy
Chen et al. (2012) found that in yeast, exposure to heat shock or the drug radicicol, which inhibits HSP90, induced aneuploidy and populations with karyotypic and phenotypic variation. Radicicol disrupted kinetochore assembly, a probable explanation for the aneuploidy, and increased the emergence of cells resistant to radicicol and unrelated drugs. Stress-associated aneuploidy causing resistance to antifungal drugs occurs widely (reviewed in Forche 2014) and might result, similarly, from HSP90 titration, but possibly other stress responses too.
Transposition in Flies
Genetic inhibition of HSP90 in Drosophila melanogaster disrupted piwi-interacting RNA-mediated silencing of transposition in germline cells and induced the transposition of multiple DNA-based and retrotransposable elements (Specchia et al. 2010). The transpositions produced morphological variation in subsequent generations. These heat-shock response-induced transpositions suggest at least one explanation for the widely reported stress-associated transpositions in flies and plants (reviewed in Belyayev 2014, Piacentini et al. 2014). Other stress responses upregulate transpositions and the genomic rearrangements likely to drive micro- and macroevolutionary events across phylogeny (see the sidebar Stress-Induced Transposition Mechanisms) (reviewed in Casacuberta & Gonzalez 2013).
STRESS-INDUCED TRANSPOSITION MECHANISMS.
Both the excision of an E. coli transposable prophage and a Pseudomonas transposon required the sigma S general stress response, indicating stress inducibility (Lamrani et al. 1999, Ilves et al. 2001). A different starvation stress response promoted the retrotransposition of a Lactobacillus lactis group II intron (Coros et al. 2009). A fascinating transposition mechanism is promoted directly by oxidative stress in a bacteriophage homing intron, which inserts into a particular genomic site (Robbins et al. 2011). During oxidative stress, direct oxidation of the homing endonuclease caused an alternative protein conformation that allowed promiscuous insertions into other sites—that is, transposon mutagenesis was induced directly. Thus, there can be stress-induced genomic instability without a stress response: by direct action on a protein.
HSP90, Other Stress Responses, and Triplet Repeat Instability in Human Cells and Plants
Trinucleotide (triplet) repeat instability alters gene expression and protein function, and repeat expansions promote various inherited neurological diseases (reviewed in Fonville et al. 2012). The inhibition of HSP90 destabilized triplet repeats in cultured human fibroblasts (Mittelman et al. 2010). The authors hypothesized that HSP90 inhibition decreased the activity of RAD51, an HSP90 client, and showed that RAD51 knockdown destabilized triplet repeats similarly. In plants, HSP90 inhibition also increased somatic alterations of triplet repeat microsatellite sequences (Queitsch et al. 2012). These data imply that microsatellites are destabilized when proteins denature, such as during heat shock, and HSP90 chaperoning activity becomes limiting.
Chatterjee et al. (2015) found that cold, heat, hypoxic, and oxidative stresses all induced triplet repeat instability in human cells. Each stressor required different stress-response regulator proteins to destabilize the repeats: cold, the CIRP cold shock response regulator; heat, the heat-shock response factor HSF1; and hypoxia and oxidative stresses, HSF1 and the hypoxic- or oxidative stress-response regulators Hif1 and NRF2. All required the DNA replication–related proteins FEN1 endonuclease and PIF1 helicase, suggesting a common, replication-based mechanism. The data demonstrate that triplet repeat mutagenesis is broadly stress-response upregulated in human cells. The stress inducibility of the triplet repeat instability might help explain the phenomenon of genetic anticipation in triplet repeat–associated neurological diseases (see the sidebar Genetic Anticipation in Neurological Diseases: Hypothesis). The existence of stress-induced metagenesis in multicellular organisms is at first glance a perplexing problem (see the sidebar Why Do Multicellular Organisms Have Stress-Induced Mutagenesis?).
RETURN TO LIFE AFTER APOPTOSIS: TRANSIENT GENOMIC INSTABILITY AND TRANSFORMATION
Cells of multicellular organisms initiate a process of programmed cell death, or apoptosis, in response to various stressors. Apoptosis was previously thought to be irreversible, especially once the protease Caspase3 was activated and genome fragmentation had occurred. However, Tang et al. (2012) demonstrated that many human cell lines recover from late-stage apoptosis and begin dividing again upon removal of the inducing stressor, even after extensive DNA damage and genome fragmentation have occurred. Recovered primary liver cells and NIH 3T3 cells showed karyotypic abnormalities indicative of incomplete and inaccurate repair of apoptotic DNA damage. Recovered NIH 3T3 cells also displayed hallmarks of oncogenic transformation, such as anchorage-independent growth and loss of contact inhibition. Recovery from late-stage apoptosis occurs frequently in flies (Ding et al. 2016) and is likely to occur in other organisms. Therefore, transient stress can trigger incomplete apoptosis and recovery, resulting in transient genome instability and potentially initiating tumorigenesis.
GENETIC ANTICIPATION IN NEUROLOGICAL DISEASES: HYPOTHESIS.
Stress response–induced triplet repeat instability (Mittelman et al. 2010, Queitsch et al. 2012, Chatterjee et al. 2015) might underlie anticipation in neurological diseases—in which children and grandchildren of individuals with expanded repeats acquire successively larger repeat expansions and more severe pathologies (reviewed in Fonville et al. 2012). In a triplet repeat disease caused by an altered poly-glutamate-containing protein, overproduction of the HSP70 protein chaperone ameliorated pathological phenotypes (Cummings et al. 1998), implying that cellular pathology induced by repeat expansion, in turn, induced a heat-shock response that depleted a chaperone, reducing protein chaperoning. We suggest that anticipation could result from stress-response activation by triplet repeat–induced protein or gene expression changes, and that the stress responses then provoke greater repeat instability.
WHY DO MULTICELLULAR ORGANISMS HAVE STRESS-INDUCED MUTAGENESIS?
Stress-induced mutation mechanisms in plants, flies, human cells, and human cancer cells are startling because, unlike microbes, multicellular creatures can get cancer. Understanding why stress-induced mutation mechanisms exist in macrobes is a problem, yet they do. Regulatable mutagenesis might (a) be retained for another required role of the mutagenic proteins, (b) promote evolution via the germline, or (c) promote somatic cell evolution, which might be needed for multicellular life. All three probably occur. For example, immune responses require more genetic diversity than a genome encodes. Three of the four human homologs of E. coli Pol IV (DinB) promote somatic hypermutation of immunoglobulin genes (reviewed in Goodman & Woodgate 2013), supporting this idea, although not Pol κ (DINB1), the ortholog of E. coli Pol IV (DinB). Pol κ (DINB1) is produced in lymphoid, adrenal gland, and testis cells (suggestive of species evolution?) (Schenten et al. 2002, Velasco-Miguel et al. 2003). All of the Pol IV homologs also promote cell survival via translesion synthesis. Biological processes other than immune diversification might require regulatable evolution in continuously renewing somatic cells or tissues (such as liver adaptation to toxins or colon or epithelium adaptation to microbes?). For whatever reason, stress-induced mutation mechanisms exist in humans and so must be considered in cancer biology and therapy resistance.
HYPOXIC STRESS–INDUCED MUTAGENESIS IN CANCER CELLS
Hypoxia in tumor microenvironments predicts aggressive phenotypes, therapy resistance, and poor clinical outcomes (reviewed in Luoto et al. 2013), and it also upregulates mutagenesis.
Hypoxic Stress Responses Downregulate Mismatch Repair
Similar to transient MMR deficiency during E. coli MBR (discussed above), multiple cancer cell lines show hypoxic stress–response downregulation of MMR protein orthologs of E. coli MutL and MutS: MLH (MutL homologous proteins) and MSH (MutS homologous proteins). The hypoxia-activated transcriptional repressor Mad1–Max downregulates MLH1 and MSH2, causing increased mutagenesis via transient MMR deficiency (Mihaylova et al. 2003, Bindra & Glazer 2007a). The Hif1 hypoxic stress response transcriptionally downregulates MSH2 and MSH6, also increasing mutagenesis, as does the DNA damage response regulator p53 (Koshiji et al. 2005). These data demonstrate that the downregulation of MMR is a stress-induced mutation mechanism in cancer cells and reveal the underlying stress-responsive signal-transduction pathways. Malignancy and therapy resistance in hypoxic tumors (reviewed in Luoto et al. 2013) could be fueled by hypoxia-induced genomic instability.
A Hypoxia-Induced Switch to Genome-Destabilizing Double-Strand Break Repair
The general stress response switches DSB repair from accurate to mutagenic in E. coli, causing small mutations and genomic rearrangements (Figure 1). A switch to MBR is also implicated in hypoxic cancer cells and tumors. Activated by hypoxia, the E2F4–p130 transcriptional repressor downregulates the HR-DSB repair proteins BRCA1, RAD51, and FANCD2 (Bindra et al. 2005, Bindra & Glazer 2007b, Scanlon & Glazer 2014), which are also silenced translationally (Chan et al. 2008). Nonhomologous break repair remains active in hypoxic BRCA1- and RAD51-repressed cells (Bindra et al. 2005). Thus, downregulated HR is expected to switch DSB repair to nonhomologous mechanisms, which produce CNAs, translocations, and loss of heterozygosity (e.g., Figure 1g,h). The following findings correlate the functional deficiency of HR-DSB repair with hypoxia. First, poly(ADP-ribose) polymerase-1 (PARP1) inhibition kills HR-DSB repair-defective BRCA-mutant tumor cells and also kills cultured hypoxic cancer cells (Chan et al. 2010). Second, hypoxic regions of PARP inhibitor–treated xenograft tumors display increased DNA damage and death after radiotherapy (Chan et al. 2010), both of which are expected in DSB repair deficiency. Third, downregulated HR-DSB repair appears generally relevant to tumors in that cervical and colon tumor xenografts show RAD51 protein levels that are inversely correlated with hypoxia-marker staining (Bindra et al. 2004). These data suggest that, analogously to E. coli MBR, hypoxic stress responses switch DSB repair from high fidelity to chromosome destabilizing.
More Mutagenesis?
Hypoxia also induces transcription of the error-prone DNA Pol iota, a homolog of E. coli Pol IV, although it is unknown whether this promotes mutagenesis (Ito et al. 2006). The rapid evolution of malignancy and therapy resistance in hypoxic tumors (reviewed in Luoto et al. 2013) might result from any or all of the above mechanisms of hypoxia-induced genomic instability and suggests that inhibition of these multiple stress responses might work therapeutically.
MUTATIONS AND THERAPIES IN CANCER
Mutation Storms in Tumors: APOBEC3B and Stress-Induced Mutagenesis?
Above, we reviewed kataegis, the approximately 100 kb clusters of multiple mutations in many cancers (Nik-Zainal et al. 2012, Roberts et al. 2012, Alexandrov et al. 2013), and its probable origin in MBR. Critical to kataegis is APOBEC3B, a cytidine deaminase that promotes C to T mutations via C to U deamination in single-stranded DNA, including at DSB ends (reviewed in Swanton et al. 2015). APOBEC3B levels have been correlated with tumor mutation storms and total mutations, particularly in breast cancer and human papillomavirus–associated cancers (Burns et al. 2013a, b; Roberts et al. 2013). APOBEC3B, a viral defense, is upregulated by the human papillomavirus (Vieira et al. 2014), and also in many other cancers, partly via PKC/NF-κB signaling (Leonard et al. 2015). It seems likely that APOBEC3B overproduction, and thus mutagenesis, is stress inducible. Like E. coli MBR mutation hot spot clusters (Shee et al. 2012, and reviewed above), cancer mutation storms potentially drive rapid, concerted evolution (Ponder et al. 2005, Shee et al. 2012, Swanton et al. 2015).
Human Ortholog of Escherichia coli Error-Prone DNA Polymerase in Lung Cancer Mutagenesis
Pol κ (DINB1), the human ortholog of the major E. coli HR-MBR DNA polymerase, is upregulated in small-cell lung cancers, which are highly mutated, with evidence of DSBs (O-Wang et al. 2001, Bavoux et al. 2005). It is unknown whether this is HR-MBR- or stress-induced, but this ortholog is implicated in driving tumor evolution.
HSP90, Cancer Evolution, and Therapeutic Inhibition
Overproduction of HSP90 protein chaperone characterizes many cancers and indicates poor prognosis (reviewed in Jarosz 2016). HSP90 inhibitors aim to block the HSP90 chaperone support of mutant proteins (reviewed above) to reveal deleterious phenotypes of highly mutated cancer cells (reviewed in Jarosz 2016). HSP90 inhibitors attack cancer cells highly specifically, probably because HSP90 exists in different protein complexes in cancer and normal cells (Kamal et al. 2003). Although we reviewed the many ways that HSP90 depletion promotes mutagenesis above, therapeutic inhibition, nevertheless, appears promising because of its specificity. Potentially, stress-induced titration of HSP90 may drive key, early genetic oncogenic events, whereas HSP90 overproduction promotes the survival and malignancy of heavily mutated cancer cells later in cancer evolution, when these drugs are needed.
Mutational Big Bangs in Tumors
Studies of tumor genome heterogeneity, using single-cell whole-genome sequencing (Navin et al. 2011, Wang et al. 2014) or mathematical reconstruction (Baca et al. 2013, Sottoriva et al. 2015), imply a picture of punctuated mutations. Early in tumor development, cells appear to undergo an explosion of cell division and mutagenesis, then remain relatively stable, sometimes with multiple different cell clones coexisting after the big bang (Baca et al. 2013, Sottoriva et al. 2015). The implication that mutagenesis turns up and then turns down supports the idea of stress-induced mutation mechanisms; if poorly adapted, stressed cells are mutable, then mutated or adapted cells are stable. The mechanisms of mutagenesis in these big bangs are important to understand in the contexts of cancer therapy and prevention.
Probabilistic Cancer-Causing Mutations Could Be Stress Induced
Both the seminal Luria & Delbruck (1943) mutation experiments in E. coli and the recent correlation of cancer incidence with stem cell divisions (Tomasetti & Vogelstein 2015) imply probabilistic relationships between cell generations and mutations (but see Wu et al. 2016). A common conflation is to interpret probabilistic events as random (as opposed to regulated or otherwise induced), as if, for example, the one cell or cell division per billion undergoing a spontaneous mutation is like every other. Such spontaneous mutations could occur in cells that have experienced special internal or external environmental conditions that induced stress responses that upregulated mutagenesis. Regardless of a correlation with generations, probabilistic mutations can arise via stress-induced mutagenesis. The evidence of tumor mutational big bangs, kataegis, and mutagenesis mechanisms in cancer cells and other models, discussed above, supports this view.
ANTI-EVOLVABILITY DRUGS
Most cancer treatments target one or a few of the many cell biological properties of cancer (Figure 2). These are the products of cancer evolution. Turning off the tap of the evolution processes that generate those products could be a fundamentally different and synergistic therapeutic strategy. We suggest targeting the mechanisms that create the variation that drives cancer evolution with anti-evolvability drugs (Figure 2) (Al Mamun et al. 2012, Rosenberg & Queitsch 2014). Targets could include, but not be limited to, activators of stress responses that drive stress-inducible mutagenesis, which are nonredundant network hubs (Al Mamun et al. 2012). APOBEC3B inhibition could also be promising (suggested in Swanton et al. 2015). Anti-evolvability drugs could slow the evolution of malignancy and resistance to chemotherapies, particularly in small populations with fewer already resistant mutant cells, for example, postsurgically. Most chemotherapies kill cells or inhibit growth and probably activate stress responses in the survivors, as various bacterial antibiotics do (Gutierrez et al. 2013). Standard chemotherapies are, therefore, likely to induce mutation mechanisms that provoke the evolution of resistance and malignancy. Anti-evolvability drugs might succeed in early-stage cancers without the use of standard chemotherapies. Alternatively, their use with other drugs might slow the evolution of resistance. Mild HSP90 inhibition shows promise in an anti-evolvability resistance-blocking capacity (Whitesell et al. 2014). Anti-evolvability antimicrobials (Al Mamun et al. 2012, Rosenberg & Queitsch 2014) could block pathogen evolution and allow immune system clearance, potentially with or without standard antibiotics, and could benefit cancer patients postsurgically and during immunosuppressive chemotherapy. Some of the anticancer efficacy of current inhibitors of the HIF response and mTOR (Ortolani et al. 2015) might result from the demonstrated (see the section Hypoxic Stress-Induced Mutagenesis in Cancer Cells), or possible, antimutagenic consequences of blocking those stress responses.
Figure 2.
Proposed anti-evolvability drugs for targeting the processes of cancer evolution. Current anticancer and antipathogen chemotherapies kill cells or inhibit growth and target the phenotypes of cancer—that is, the products of cancer evolution (right). Proposed anti-evolvability drugs would target the evolutionary processes that generate the variation that drives cancer (and pathogen) evolution, including the stress-response regulators that propel mutagenesis and the generators of nongenetic variation. Anti-evolvability drugs could slow evolution to allow other drugs and the immune system to work without inducing resistance. Multiple inputs into, and outputs from, one knot, referred to as bow ties, are universal organizational structures in biological and technological networks that confer evolvability and robustness and predict fragilities in disease (Csete & Doyle 2004).
SUMMARY POINTS.
Stress responses transiently and abruptly upregulate mutation mechanisms, and potentially the ability to evolve, when cells are poorly adapted to their environments—that is, when they are stressed.
Stress-induced mutation mechanisms are similar from bacteria to cancer cells, and they cause base substitutions and indels, transposon hops, copy-number alterations, and other genome rearrangements. Bacterial models led to the discovery of cancer cell mechanisms.
Mutations often occur nonrandomly in genomic space, in clusters, and are limited to small cell subpopulations, and both characteristics are predicted to drive evolution.
Mutation storms and punctuated big bangs of mutagenesis in tumors suggest stress-induced mutation mechanisms that are similar to mutagenic break repair in E. coli.
Cancer cells encounter stress-laden microenvironments. Hypoxic stress responses, and probably others, upregulate multiple mutation mechanisms.
Anti-evolvability drugs proposed to target the generation of variation that drives cancer evolution might slow malignancy and resistance to standard antiproliferative chemotherapies, which induce stress responses and, potentially, mutagenesis. This strategy is promising against pathogens as well.
Some of the efficacy of current anticancer stress-response inhibitors might result from anti-evolvability activity.
FUTURE ISSUES.
Do stress responses underlie most spontaneous mutations? Which responses?
How do stress responses flip the switch to mutagenesis in E. coli mutagenic break repair and in cancer cells?
Are spontaneous double-strand breaks, and the mutation showers they cause, random in genomic space or regulated, biased, or directed? By which mechanisms? Responsively to environments?
Do hypoxia-induced mutation mechanisms occur in tumors and drive tumor evolution?
Do punctuated mutational big bangs and kataegis in tumors result from stress-induced mutagenesis? Which mechanism(s)? Which stress responses?
Can proposed anti-evolvability drugs succeed as cancer cotherapies with current antiproliferative cancer therapies? In cancer prevention? As antipathogen cotherapies?
Could anti-evolvability drugs succeed alone, without standard antiproliferative drugs, in cancer or in infectious disease, or both, by favoring immune clearance?
Acknowledgments
We apologize to colleagues whose work we could not cite due to space limitations. We thank C. Queitsch, R.S. Harris, and I. Matic for helpful discussions; E. Rogers and J. Xia for creative contributions to the figures; and L. Garcia-Villada, C. Maley, J.P. Pribis, D. Shibata, and T. Tlsty for comments on the manuscript. D.M.F. is supported by the Cancer Prevention and Research Institute of Texas, Baylor College of Medicine Comprehensive Cancer Training Program Postdoctoral Fellowship (RP160283). P.J.H. is supported by the NASA Astrobiology Institute under cooperative agreement NN13AA91A, issued though the Science Mission Directorate. This work was supported in part by a gift from the W.M. Keck Foundation and the US National Institutes of Health to S.M.R.
Glossary
- SOS response
bacterial DNA damage response; the first such response discovered in any organism; it is prototypic for DNA damage responses in humans and other organisms
- DSB
double-strand break
- HR
homologous recombination
- MBR
mutagenic break repair
- Indel
1 to a few bp insertions or deletions; a common error of DinB subfamily error-prone DNA polymerases, such as E. coli Pol IV (DinB) and the human ortholog Pol κ(DINB1)
- Copy-number alteration (CNA)
large deletions, duplications, or amplifications of segments of the genome, thousands to millions of base pairs in length
- HR-MBR
homologous recombinational mutagenic break repair
- Sigma E
a bacterial membrane-protein stress-response activator; upregulates protein orthologs of human chaperones and proteases upregulated by the analogous human unfolded-protein response
- Sigma S
a bacterial general stress-response activator; induced by starvation, antibiotics, and cold, pH, osmotic, oxidative, and other shocks; a possible analog of human mTOR
- RecBCD
E. coli RecA loader at double-strand breaks; an analog of human BRCA2
- RecA
E. coli ortholog of human RAD51 homologous recombination protein
- Pol IV (DinB)
E. coli error-prone Y-family DNA polymerase; an ortholog of human Pol κ (DINB1); a homolog of human DNA Pols iota, eta, and REV1
- Mismatch repair (MMR)
a post-DNA synthesis error correction mechanism conserved from bacteria to humans, inactivation of which increases mutation rate and promotes cancers
- Kataegis
mutation storms (from Greek) of multiple, isolated, approximately 100 kb clusters or showers of mutations seen in cancer genome sequences
- R loop
RNA–DNA hybrid molecule with an RNA strand displacing a like strand of duplex DNA
- MMBIR
microhomology-mediated break-induced replication
- HSP90
a protein chaperone, during heat shock, abandons client proteins to ameliorate others, releasing stored phenotypic diversity in clients that have accumulated mutations
Footnotes
DISCLOSURE STATEMENT
The authors are not aware of any affiliations, memberships, funding, or financial holdings that might be perceived as affecting the objectivity of this review.
LITERATURE CITED
- Åkerlund T, Nordstrom K, Bernander R. Analysis of cell size and DNA content in exponentially growing and stationary-phase batch cultures of Escherichia coli. J Bacteriol. 1995;177:6791–97. doi: 10.1128/jb.177.23.6791-6797.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Al Mamun AA, Lombardo MJ, Shee C, Lisewski AM, Gonzalez C, et al. Identity and function of a large gene network underlying mutagenic repair of DNA breaks. Science. 2012;338:1344–48. doi: 10.1126/science.1226683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alexandrov LB, Nik-Zainal S, Wedge DC, Campbell PJ, Stratton MR. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 2013;3:246–59. doi: 10.1016/j.celrep.2012.12.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andersson DI, Hughes D. Microbiological effects of sublethal levels of antibiotics. Nat Rev Microbiol. 2014;12:465–78. doi: 10.1038/nrmicro3270. [DOI] [PubMed] [Google Scholar]
- Aravind L, Walker AG, Koonin E. Conserved domains in DNA repair proteins and evolution of repair systems. Nucleic Acids Res. 1999;27:1223–42. doi: 10.1093/nar/27.5.1223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Avery O, MacLeod C, McCarty M. Studies on the chemical nature of the substance inducing transformation of pneumococcal types. J Exp Med. 1944;79:137–58. doi: 10.1084/jem.79.2.137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baca SC, Prandi D, Lawrence MS, Mosquera JM, Romanel A, et al. Punctuated evolution of prostate cancer genomes. Cell. 2013;153:666–77. doi: 10.1016/j.cell.2013.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Battesti A, Majdalani N, Gottesman S. The RpoS-mediated general stress response in Escherichia coli. Annu Rev Microbiol. 2011;65:189–213. doi: 10.1146/annurev-micro-090110-102946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bavoux C, Leopoldino AM, Bergoglio V, O-Wang J, Ogi T, et al. Up-regulation of the error-prone DNA polymerase κ promotes pleiotropic genetic alterations and tumorigenesis. Cancer Res. 2005;65:325–30. [PubMed] [Google Scholar]
- Belenky P, Ye JD, Porter CB, Cohen NR, Lobritz MA, et al. Bactericidal antibiotics induce toxic metabolic perturbations that lead to cellular damage. Cell Rep. 2015;13:968–80. doi: 10.1016/j.celrep.2015.09.059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belyayev A. Bursts of transposable elements as an evolutionary driving force. J Evol Biol. 2014;27:2573–84. doi: 10.1111/jeb.12513. [DOI] [PubMed] [Google Scholar]
- Berman HK, Gauthier ML, Tlsty TD. Premalignant breast neoplasia: a paradigm of interlesional and intralesional molecular heterogeneity and its biological and clinical ramifications. Cancer Prev Res. 2010;3:579–87. doi: 10.1158/1940-6207.CAPR-10-0073. [DOI] [PubMed] [Google Scholar]
- Bindra RS, Gibson SL, Meng A, Westermark U, Jasin M, et al. Hypoxia-induced down-regulation of BRCA1 expression by E2Fs. Cancer Res. 2005;65:11597–604. doi: 10.1158/0008-5472.CAN-05-2119. [DOI] [PubMed] [Google Scholar]
- Bindra RS, Glazer PM. Co-repression of mismatch repair gene expression by hypoxia in cancer cells: role of the Myc/Max network. Cancer Lett. 2007a;252:93–103. doi: 10.1016/j.canlet.2006.12.011. [DOI] [PubMed] [Google Scholar]
- Bindra RS, Glazer PM. Repression of RAD51 gene expression by E2F4/p130 complexes in hypoxia. Oncogene. 2007b;26:2048–57. doi: 10.1038/sj.onc.1210001. [DOI] [PubMed] [Google Scholar]
- Bindra RS, Schaffer PJ, Meng A, Woo J, Maseide K, et al. Down-regulation of Rad51 and decreased homologous recombination in hypoxic cancer cells. Mol Cell Biol. 2004;24:8504–18. doi: 10.1128/MCB.24.19.8504-8518.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bjedov I, Tenaillon O, Gérard B, Souza V, Denamur E, et al. Stress-induced mutagenesis in bacteria. Science. 2003;300:1404–9. doi: 10.1126/science.1082240. [DOI] [PubMed] [Google Scholar]
- Bos J, Zhang Q, Vyawahare S, Rogers E, Rosenberg SM, Austin RH. Emergence of antibiotic resistance from multinucleated bacterial filaments. PNAS. 2015;112:178–83. doi: 10.1073/pnas.1420702111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brégeon D, Matic I, Radman M, Taddei F. Inefficient mismatch repair: genetic defects and down regulation. J Genet. 1999;78:21–28. [Google Scholar]
- Bull HJ, McKenzie GJ, Hastings PJ, Rosenberg SM. Evidence that stationary-phase hypermutation in the Escherichia coli chromosome is promoted by recombination. Genetics. 2000;154:1427–37. doi: 10.1093/genetics/154.4.1427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns MB, Lackey L, Carpenter MA, Rathore A, Land AM, et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature. 2013a;494:366–70. doi: 10.1038/nature11881. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burns MB, Temiz NA, Harris RS. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat Genet. 2013b;45:977–83. doi: 10.1038/ng.2701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairns J, Foster PL. Adaptive reversion of a frameshift mutation in Escherichia coli. Genetics. 1991;128:695–701. doi: 10.1093/genetics/128.4.695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camps M, Naukkarinen J, Johnson BP, Loeb LA. Targeted gene evolution in Escherichia coli using a highly error-prone DNA polymerase I. PNAS. 2003;100:9727–32. doi: 10.1073/pnas.1333928100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caporale LH. The Implicit Genome. Oxford, UK: Oxford Univ. Press; 2006. [Google Scholar]
- Casacuberta E, Gonzalez J. The impact of transposable elements in environmental adaptation. Mol Ecol. 2013;22:1503–17. doi: 10.1111/mec.12170. [DOI] [PubMed] [Google Scholar]
- Chan N, Koritzinsky M, Zhao H, Bindra R, Glazer PM, et al. Chronic hypoxia decreases synthesis of homologous recombination proteins to offset chemoresistance and radioresistance. Cancer Res. 2008;68:605–14. doi: 10.1158/0008-5472.CAN-07-5472. [DOI] [PubMed] [Google Scholar]
- Chan N, Pires IM, Bencokova Z, Coackley C, Luoto KR, et al. Contextual synthetic lethality of cancer cell kill based on the tumor microenvironment. Cancer Res. 2010;70:8045–54. doi: 10.1158/0008-5472.CAN-10-2352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chatterjee N, Lin Y, Santillan BA, Yotnda P, Wilson JH. Environmental stress induces trinucleotide repeat mutagenesis in human cells. PNAS. 2015;112:3764–69. doi: 10.1073/pnas.1421917112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen G, Bradford WD, Seidel CW, Li R. Hsp90 stress potentiates rapid cellular adaptation through induction of aneuploidy. Nature. 2012;482:246–50. doi: 10.1038/nature10795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cirz RT, Chin JK, Andes DR, de Crecy-Lagard V, Craig WA, Romesberg FE. Inhibition of mutation and combating the evolution of antibiotic resistance. PLOS Biol. 2005;3:e176. doi: 10.1371/journal.pbio.0030176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coros CJ, Piazza CL, Chalamcharla VR, Smith D, Belfort M. Global regulators orchestrate group II intron retromobility. Mol Cell. 2009;34:250–56. doi: 10.1016/j.molcel.2009.03.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Csete M, Doyle J. Bow ties, metabolism and disease. Trends Biotechnol. 2004;22:446–50. doi: 10.1016/j.tibtech.2004.07.007. [DOI] [PubMed] [Google Scholar]
- Cummings C, Mancini M, Antalffy B, DeFranco D, Orr H, Zoghbi H. Chaperone suppression of aggregation and altered subcellular proteasome localization imply protein misfolding in SCA1. Nat Genet. 1998;19:148–54. doi: 10.1038/502. [DOI] [PubMed] [Google Scholar]
- Denamur E, Matic I. Evolution of mutation rates in bacteria. Mol Microbiol. 2006;60:820–27. doi: 10.1111/j.1365-2958.2006.05150.x. [DOI] [PubMed] [Google Scholar]
- Ding A, Sun G, Argraw Y, Wong J, Easwaran S, Montell D. CasExpress reveals widespread and diverse patterns of cell survival of caspase-3 activation during development in vivo. eLife. 2016;5:e10936. doi: 10.7554/eLife.10936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drake JW, Bebenek A, Kissling GE, Peddada S. Clusters of mutations from transient hypermutability. PNAS. 2005;102:12849–54. doi: 10.1073/pnas.0503009102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Echols H. SOS functions, cancer, and inducible evolution. Cell. 1981;25:1–2. doi: 10.1016/0092-8674(81)90223-3. [DOI] [PubMed] [Google Scholar]
- Fearon E, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61:759–67. doi: 10.1016/0092-8674(90)90186-i. [DOI] [PubMed] [Google Scholar]
- Fishel R, Lescoe M, Rao M, Copeland N, Jenkins N, et al. The human mutator gene homolog MSH2 and its association with hereditary nonpolyposis colon cancer. Cell. 1993;75:1027–38. doi: 10.1016/0092-8674(93)90546-3. [DOI] [PubMed] [Google Scholar]
- Fonville NC, Ward RM, Mittelman D. Stress-induced modulators of repeat instability and genome evolution. J Mol Microbiol Biotechnol. 2012;21:36–44. doi: 10.1159/000332748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forche A. Large-scale chromosomal changes and associated fitness consequences in pathogenic fungi. Curr Fungal Infect Rep. 2014;8:163–70. doi: 10.1007/s12281-014-0181-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foster PL, Trimarchi JM. Adaptive reversion of a frameshift mutation in Escherichia coli by simple base deletions in homopolymeric runs. Science. 1994;265:407–9. doi: 10.1126/science.8023164. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frisch RL, Su Y, Thornton PC, Gibson JL, Rosenberg SM, Hastings PJ. Separate DNA Pol II- and Pol IV-dependent pathways of stress-induced mutation during double-strand-break repair in Escherichia coli are controlled by RpoS. J Bacteriol. 2010;192:4694–700. doi: 10.1128/JB.00570-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Funchain P, Yeung A, Stewart J, Lin R, Slupska M, Miller J. The consequences of growth of a mutator strain of Escherichia coli as measured by loss of function among multiple gene targets and loss of fitness. Genetics. 2000;154:959–70. doi: 10.1093/genetics/154.3.959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galhardo RS, Do R, Yamada M, Friedberg EC, Hastings PJ, et al. DinB upregulation is the sole role of the SOS response in stress-induced mutagenesis in Escherichia coli. Genetics. 2009;182:55–68. doi: 10.1534/genetics.109.100735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerlinger M, McGranahan N, Dewhurst SM, Burrell RA, Tomlinson I, Swanton C. Cancer: evolution within a lifetime. Annu Rev Genet. 2014;48:215–36. doi: 10.1146/annurev-genet-120213-092314. [DOI] [PubMed] [Google Scholar]
- Gibson JL, Lombardo MJ, Thornton PC, Hu KH, Galhardo RS, et al. The σE stress response is required for stress-induced mutation and amplification in Escherichia coli. Mol Microbiol. 2010;77:415–30. doi: 10.1111/j.1365-2958.2010.07213.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez C, Hadany L, Ponder RG, Price M, Hastings PJ, Rosenberg SM. Mutability and importance of a hypermutable cell subpopulation that produces stress-induced mutants in Escherichia coli. PLOS Genet. 2008;4:e1000208. doi: 10.1371/journal.pgen.1000208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman MF, Woodgate R. Translesion DNA polymerases. Cold Spring Harb Perspect Biol. 2013;5:a010363. doi: 10.1101/cshperspect.a010363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greaves M, Maley CC. Clonal evolution in cancer. Nature. 2012;481:306–13. doi: 10.1038/nature10762. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gutierrez A, Laureti L, Crussard S, Abida H, Rodriguez-Rojas A, et al. β-Lactam antibiotics promote bacterial mutagenesis via an RpoS-mediated reduction in replication fidelity. Nat Commun. 2013;4:1610. doi: 10.1038/ncomms2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris RS, Feng G, Ross K, Sidhu R, Thulin C, et al. Mismatch repair protein MutL becomes limiting during stationary-phase mutation. Genes Dev. 1997;11:2416–37. doi: 10.1101/gad.11.18.2426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris RS, Longerich S, Rosenberg SM. Recombination in adaptive mutation. Science. 1994;264:258–60. doi: 10.1126/science.8146657. [DOI] [PubMed] [Google Scholar]
- Hastings PJ, Bull HJ, Klump JR, Rosenberg SM. Adaptive amplification: an inducible chromosomal instability mechanism. Cell. 2000;103:723–31. doi: 10.1016/s0092-8674(00)00176-8. [DOI] [PubMed] [Google Scholar]
- Hastings PJ, Ira G, Lupski JR. A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLOS Genet. 2009;5:e1000327. doi: 10.1371/journal.pgen.1000327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heidenreich E, Novotny R, Kneidinger B, Holzmann V, Wintersberger U. Non-homologous end joining as an important mutagenic process in cell cycle–arrested cells. EMBO J. 2003;22:2274–83. doi: 10.1093/emboj/cdg203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hershey A, Chase M. Independent functions of viral protein and nucleic acid in growth of bacteriophage. J Gen Physiol. 1952;36:39–56. doi: 10.1085/jgp.36.1.39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holbeck S, Strathern J. A role for REV3 in mutagenesis during double-strand break repair in Saccharomyces cerevisiae. Genetics. 1997;147:1017–24. doi: 10.1093/genetics/147.3.1017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huxley J. Evolution of the Modern Synthesis. London: Allen & Unwin; 1942. [Google Scholar]
- Ilves H, Horak R, Kivisaar M. Involvement of σS in starvation-induced transposition of Pseudomonas putida transposon Tn4652. J Bacteriol. 2001;183:5445–48. doi: 10.1128/JB.183.18.5445-5448.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito A, Koshikawa N, Mochizuki S, Omura K, Takenaga K. Hypoxia-inducible factor-1 mediates the expression of DNA polymerase ı in human tumor cells. Biochem Biophys Res Commun. 2006;351:306–11. doi: 10.1016/j.bbrc.2006.10.048. [DOI] [PubMed] [Google Scholar]
- Jarosz D. Hsp90: a global regulator of the genotype-to-phenotype map in cancers. Adv Cancer Res. 2016;129:225–47. doi: 10.1016/bs.acr.2015.11.001. [DOI] [PubMed] [Google Scholar]
- Kamal A, Thao L, Sensintaffer J, Zhang L, Boehm M, et al. A high-affinity conformation of Hsp90 confers tumour selectivity on Hsp90 inhibitors. Nature. 2003;435:407–10. doi: 10.1038/nature01913. [DOI] [PubMed] [Google Scholar]
- Kohanski MA, DePristo MA, Collins JJ. Sublethal antibiotic treatment leads to multidrug resistance via radical-induced mutagenesis. Mol Cell. 2010;37:311–20. doi: 10.1016/j.molcel.2010.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koshiji M, To KK, Hammer S, Kumamoto K, Harris AL, et al. HIF-1αinduces genetic instability by transcriptionally downregulating MutSα expression. Mol Cell. 2005;17:793–803. doi: 10.1016/j.molcel.2005.02.015. [DOI] [PubMed] [Google Scholar]
- Kuzminov A. Homologous recombination—experimental systems, analysis, and significance. EcoSal Plus. 2013;4 doi: 10.1128/ecosalplus.7.2.6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamrani A, Ranquet C, Gama M, Nakai H, Shapiro J, et al. Starvation-induced Mu cts62-mediated coding sequence fusion: a role for ClpXP, Lon, RpoS and Crp. Mol Microbiol. 1999;32:327–43. doi: 10.1046/j.1365-2958.1999.01352.x. [DOI] [PubMed] [Google Scholar]
- Lee JK, Choi YL, Kwon M, Park PJ. Mechanisms and consequences of cancer genome instability: lessons from genome sequencing studies. Annu Rev Pathol. 2016;11:283–312. doi: 10.1146/annurev-pathol-012615-044446. [DOI] [PubMed] [Google Scholar]
- Lengauer C, Kinzler K, Vogelstein B. Genetic instabilities in human cancers. Nature. 1998;396:643–49. doi: 10.1038/25292. [DOI] [PubMed] [Google Scholar]
- Leonard B, McCann JL, Starrett GJ, Kosyakovsky L, Luengas EM, et al. The PKC/NF-κB signaling pathway induces APOBEC3B expression in multiple human cancers. Cancer Res. 2015;75:4538–47. doi: 10.1158/0008-5472.CAN-15-2171-T. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li GM. Mechanisms and functions of DNA mismatch repair. Cell Res. 2008;18:85–98. doi: 10.1038/cr.2007.115. [DOI] [PubMed] [Google Scholar]
- Lin D, Gibson IB, Moore JM, Thornton PC, Leal SM, Hastings PJ. Global chromosomal structural instability in a subpopulation of starving Escherichia coli cells. PLOS Genet. 2011;7:e1002223. doi: 10.1371/journal.pgen.1002223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindquist S. Protein folding sculpting evolutionary change. Cold Spring Harb Symp Quant Biol. 2009;74:103–8. doi: 10.1101/sqb.2009.74.043. [DOI] [PubMed] [Google Scholar]
- Liu P, Carvalho CM, Hastings PJ, Lupski JR. Mechanisms for recurrent and complex human genomic rearrangements. Curr Opin Genet Dev. 2012;22:211–20. doi: 10.1016/j.gde.2012.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lombardo M-J, Aponyi I, Rosenberg SM. General stress response regulator RpoS in adaptive mutation and amplification in Escherichia coli. Genetics. 2004;166:669–80. doi: 10.1093/genetics/166.2.669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Longerich S, Galloway A, Harris RS, Wong C, Rosenberg SM. Adaptive mutation sequences reproduced by mismatch repair deficiency. PNAS. 1995;92:12017–20. doi: 10.1073/pnas.92.26.12017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luoto KR, Kumareswaran R, Bristow RG. Tumor hypoxia as a driving force in genetic instability. Genome Integr. 2013;4:5. doi: 10.1186/2041-9414-4-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luria S, Delbruck M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics. 1943;28:491–511. doi: 10.1093/genetics/28.6.491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lynch M. Evolution of the mutation rate. Trends Genet. 2010;26:345–52. doi: 10.1016/j.tig.2010.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maharjan R, Ferenci T. Mutational signatures indicative of environmental stress in bacteria. Mol Biol Evol. 2015;32:380–91. doi: 10.1093/molbev/msu306. [DOI] [PubMed] [Google Scholar]
- Martin HA, Pedraza-Reyes M, Yasbin RE, Robleto EA. Transcriptional derepression and Mfd are mutagenic in stressed Bacillus subtilis cells. J Mol Microbiol Biotechnol. 2011;21:45–58. doi: 10.1159/000332751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayr E. The Growth of Biological Thought: Diversity, Evolution, and Inheritance. Cambridge, MA: Harvard Univ. Press; 1985. [Google Scholar]
- McClintock B. Mechanisms that rapidly reorganize the genome. Stadler Genet Symp. 1978;10:25–48. [Google Scholar]
- McKenzie GJ, Harris RS, Lee PL, Rosenberg SM. The SOS response regulates adaptive mutation. PNAS. 2000;97:6646–51. doi: 10.1073/pnas.120161797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKenzie GJ, Lee PL, Lombardo MJ, Hastings PJ, Rosenberg SM. SOS mutator DNA polymerase IV functions in adaptive mutation and not adaptive amplification. Mol Cell. 2001;7:571–79. doi: 10.1016/s1097-2765(01)00204-0. [DOI] [PubMed] [Google Scholar]
- Merrikh H, Zhang Y, Grossman AD, Wang JD. Replication–transcription conflicts in bacteria. Nat Rev Microbiol. 2012;10:449–58. doi: 10.1038/nrmicro2800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meselson M, Stahl F. The replication of DNA in Escherichia coli. PNAS. 1958;44:671–82. doi: 10.1073/pnas.44.7.671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mihaylova VT, Bindra RS, Yuan J, Campisi D, Narayanan L, et al. Decreased expression of the DNA mismatch repair gene Mlh1 under hypoxic stress in mammalian cells. Mol Cell Biol. 2003;23:3265–73. doi: 10.1128/MCB.23.9.3265-3273.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mittelman D, Sykoudis K, Hersh M, Lin Y, Wilson JH. Hsp90 modulates CAG repeat instability in human cells. Cell Stress Chaperones. 2010;15:753–59. doi: 10.1007/s12192-010-0191-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motamedi M, Szigety S, Rosenberg SM. Double-strand-break repair recombination in Escherichia coli: physical evidence for a DNA replication mechanism in vivo. Genes Dev. 1999;13:2889–903. doi: 10.1101/gad.13.21.2889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Navin N, Kendall J, Troge J, Andrews P, Rodgers L, et al. Tumour evolution inferred by single-cell sequencing. Nature. 2011;472:90–94. doi: 10.1038/nature09807. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nik-Zainal S, Alexandrov LB, Wedge DC, Van Loo P, Greenman CD, et al. Mutational processes molding the genomes of 21 breast cancers. Cell. 2012;149:979–93. doi: 10.1016/j.cell.2012.04.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nowell PC. The clonal evolution of tumor cell populations. Science. 1976;194:23–28. doi: 10.1126/science.959840. [DOI] [PubMed] [Google Scholar]
- O-Wang J, Kawamura K, Tada Y, Ohmori H, Kimura H, et al. DNA polymerase κ, implicated in spontaneous and DNA damage-induced mutagenesis, is overexpressed in lung cancer. Cancer Res. 2001;61:5366–69. [PubMed] [Google Scholar]
- Ortolani S, Ciccarese C, Cingarlini S, Tortora G, Massari F. Suppression of mTOR pathway in solid tumors: lessons learned from clinical experience in renal cell carcinoma and neuroendocrine tumors and new perspectives. Future Oncol. 2015;11:1809–28. doi: 10.2217/fon.15.81. [DOI] [PubMed] [Google Scholar]
- Pedraza-Reyes M, Yasbin RE. Contribution of the mismatch DNA repair system to the generation of stationary-phase-induced mutants of Bacillus subtilis. J Bacteriol. 2004;186:6485–91. doi: 10.1128/JB.186.19.6485-6491.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pennington JM, Rosenberg SM. Spontaneous DNA breakage in single living Escherichia coli cells. Nat Genet. 2007;39:797–802. doi: 10.1038/ng2051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petrosino JF, Galhardo RS, Morales LD, Rosenberg SM. Stress-induced β-lactam antibiotic resistance mutation and sequences of stationary-phase mutations in the Escherichia coli chromosome. J Bacteriol. 2009;191:5881–89. doi: 10.1128/JB.00732-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piacentini L, Fanti L, Specchia V, Bozzetti MP, Berloco M, et al. Transposons, environmental changes, and heritable induced phenotypic variability. Chromosoma. 2014;123:345–54. doi: 10.1007/s00412-014-0464-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pomerantz RT, Goodman MF, O’Donnell ME. DNA polymerases are error-prone at RecA-mediated recombination intermediates. Cell Cycle. 2013;12:2558–63. doi: 10.4161/cc.25691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pommier Y, Leo E, Zhang H, Marchand C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem Biol. 2010;17:421–33. doi: 10.1016/j.chembiol.2010.04.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ponder RG, Fonville NC, Rosenberg SM. A switch from high-fidelity to error-prone DNA double-strand break repair underlies stress-induced mutation. Mol Cell. 2005;19:791–804. doi: 10.1016/j.molcel.2005.07.025. [DOI] [PubMed] [Google Scholar]
- Prieto AI, Ramos-Morales F, Casadesus J. Repair of DNA damage induced by bile salts in Salmonella enterica. Genetics. 2006;174:575–84. doi: 10.1534/genetics.106.060889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quah S-K, von Borstel R, Hastings P. The origin of spontaneous mutation in Saccharomyces cerevisiae. Genetics. 1980;96:819–39. doi: 10.1093/genetics/96.4.819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Queitsch C, Carlson K, Girirajan S. Lessons from model organisms: phenotypic robustness and missing heritability in complex disease. PLOS Genet. 2012;8:e1003041. doi: 10.1371/journal.pgen.1003041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radman M. Phenomenology of an inducible mutagenic DNA repair pathway in Escherichia coli: SOS hypothesis. In: Prokash L, Sherman F, Miller M, Lawrence C, Tabor H, editors. Molecular and Environmental Aspects of Mutagenesis. Springfield, IL: Charles C. Thomas; 1974. pp. 128–42. [Google Scholar]
- Radman M. SOS repair hypothesis: phenomenology of an inducible DNA repair which is accompanied by mutagenesis. In: Hanawalt P, Setlow RB, editors. Molecular Mechanisms for Repair of DNA. New York: Plenum; 1975. pp. 355–67. [DOI] [PubMed] [Google Scholar]
- Ram Y, Hadany L. The evolution of stress-induced hypermutation in asexual populations. Evolution. 2012;66:2315–28. doi: 10.1111/j.1558-5646.2012.01576.x. [DOI] [PubMed] [Google Scholar]
- Ram Y, Hadany L. Stress-induced mutagenesis and complex adaptation. Proc R Soc Lond B. 2014;281:20141025. doi: 10.1098/rspb.2014.1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robbins JB, Smith D, Belfort M. Redox-responsive zinc finger fidelity switch in homing endonuclease and intron promiscuity in oxidative stress. Curr Biol. 2011;21:243–48. doi: 10.1016/j.cub.2011.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts SA, Lawrence MS, Klimczak LJ, Grimm SA, Fargo D, et al. An APOBEC cytidine deaminase mutagenesis pattern is widespread in human cancers. Nat Genet. 2013;45:970–76. doi: 10.1038/ng.2702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts SA, Sterling J, Thompson C, Harris S, Mav D, et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol Cell. 2012;46:424–35. doi: 10.1016/j.molcel.2012.03.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers E, Correa R, Barreto B, Bravo Nunez MA, Minnick PJ, et al. Double-strand-break repair, mutagenesis and stress. In: de Bruijn FJ, editor. Stress and Environmental Control of Gene Expression and Adaptation in Bacteria. New York: Wiley; 2016. pp. 185–95. [Google Scholar]
- Rosenberg SM, Longerich S, Gee P, Harris RS. Adaptive mutation by deletions in small mononucleotide repeats. Science. 1994;265:405–7. doi: 10.1126/science.8023163. [DOI] [PubMed] [Google Scholar]
- Rosenberg SM, Queitsch C. Combating evolution to fight disease. Science. 2014;343:1088–89. doi: 10.1126/science.1247472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rosenberg SM, Thullin C, Harris RS. Transient and heritable mutators in adaptive evolution in the lab and in nature. Genetics. 1998;148:1559–66. doi: 10.1093/genetics/148.4.1559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudner R, Murray A, Huda N. Is there a link between mutation rates and the stringent response in Bacillus subtilis? Ann N Y Acad Sci. 1999;870:418–22. doi: 10.1111/j.1749-6632.1999.tb08917.x. [DOI] [PubMed] [Google Scholar]
- Saint-Ruf C, Garfa-Taore M, Collin V, Cordier C, Francschi C, Matic I. Massive diversification in aging colonies of Escherichia coli. J Bacteriol. 2014;196:3059–73. doi: 10.1128/JB.01421-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakofsky CJ, Roberts SA, Malc E, Mieczkowski PA, Resnick MA, et al. Break-induced replication is a source of mutation clusters underlying kataegis. Cell Rep. 2014;7:1640–48. doi: 10.1016/j.celrep.2014.04.053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santos-Pereira JM, Aguilera A. R loops: new modulators of genome dynamics and function. Nat Rev Genet. 2015;16:583–97. doi: 10.1038/nrg3961. [DOI] [PubMed] [Google Scholar]
- Scanlon SE, Glazer PM. Hypoxic stress facilitates acute activation and chronic downregulation of Fanconi anemia proteins. Mol Cancer Res. 2014;12:1016–28. doi: 10.1158/1541-7786.MCR-13-0628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schenten D, Gerlach V, Guo C, Velasco-Miguel S, Hladik C, et al. DNA polymerase κ deficiency does not affect somatic hypermutation in mice. Eur J Immunol. 2002;32:3152–60. doi: 10.1002/1521-4141(200211)32:11<3152::AID-IMMU3152>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
- Shee C, Cox BD, Gu F, Luengas EM, Joshi MC, et al. Engineered proteins detect spontaneous DNA breakage in human and bacterial cells. eLife. 2013;2:e01222. doi: 10.7554/eLife.01222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shee C, Gibson JL, Darrow MC, Gonzalez C, Rosenberg SM. Impact of a stress-inducible switch to mutagenic repair of DNA breaks on mutation in Escherichia coli. PNAS. 2011;108:13659–64. doi: 10.1073/pnas.1104681108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shee C, Gibson JL, Rosenberg SM. Two mechanisms produce mutation hotspots at DNA breaks in Escherichia coli. Cell Rep. 2012;2:714–21. doi: 10.1016/j.celrep.2012.08.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shor E, Fox CA, Broach JR. The yeast environmental stress response regulates mutagenesis induced by proteotoxic stress. PLOS Genet. 2013;9:e1003680. doi: 10.1371/journal.pgen.1003680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slack A, Thornton PC, Magner DB, Rosenberg SM, Hastings PJ. On the mechanism of gene amplification induced under stress in Escherichia coli. PLOS Genet. 2006;2:e48. doi: 10.1371/journal.pgen.0020048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sottoriva A, Kang H, Ma Z, Graham TA, Salomon MP, et al. A Big Bang model of human colorectal tumor growth. Nat Genet. 2015;47:209–16. doi: 10.1038/ng.3214. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Specchia V, Piacentini L, Tritto P, Fanti L, D’Alessandro R, et al. Hsp90 prevents phenotypic variation by suppressing the mutagenic activity of transposons. Nature. 2010;463:662–65. doi: 10.1038/nature08739. [DOI] [PubMed] [Google Scholar]
- Stahl F. Bacterial genetics: a unicorn in the garden. Nature. 1988;335:112–13. doi: 10.1038/335112a0. [DOI] [PubMed] [Google Scholar]
- Strathern J, Shafer B, McGill C. DNA synthesis errors associated with double-strand-break repair. Genetics. 1995;140:965–72. doi: 10.1093/genetics/140.3.965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung HM, Yasbin RE. Adaptive, or stationary-phase, mutagenesis, a component of bacterial differentiation in Bacillus subtilis. J Bacteriol. 2002;184:5641–53. doi: 10.1128/JB.184.20.5641-5653.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung HM, Yeamans G, Ross CA, Yasbin RE. Roles of YqjH and YqjW, homologs of the Escherichia coli UmuC/DinB or Y superfamily of DNA polymerases, in stationary-phase mutagenesis and UV-induced mutagenesis of Bacillus subtilis. J Bacteriol. 2003;185:2153–60. doi: 10.1128/JB.185.7.2153-2160.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swanton C, McGranahan N, Starrett GJ, Harris RS. APOBEC enzymes: mutagenic fuel for cancer evolution and heterogeneity. Cancer Discov. 2015;5:704–12. doi: 10.1158/2159-8290.CD-15-0344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang HL, Tang HM, Mak KH, Hu S, Wang SS, et al. Cell survival, DNA damage, and oncogenic transformation after a transient and reversible apoptotic response. Mol Biol Cell. 2012;23:2240–52. doi: 10.1091/mbc.E11-11-0926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tenaillon O, Denamur E, Matic I. Evolutionary significance of stress-induced mutagenesis in bacteria. Trends Microbiol. 2004;12:264–70. doi: 10.1016/j.tim.2004.04.002. [DOI] [PubMed] [Google Scholar]
- Tlsty T, Albertini A, Miller J. Gene amplification in the lac region of E. coli. Cell. 1984;37:217–24. doi: 10.1016/0092-8674(84)90317-9. [DOI] [PubMed] [Google Scholar]
- Tomasetti C, Vogelstein B. Variation in cancer risk among tissues can be explained by the number of stem cell divisions. Science. 2015;347:78–81. doi: 10.1126/science.1260825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torkelson J, Harris RS, Lombardo MJ, Nagendran J, Thulin C, Rosenberg SM. Genome-wide hypermutation in a subpopulation of stationary-phase cells underlies recombination-dependent adaptive mutation. EMBO J. 1997;16:3303–11. doi: 10.1093/emboj/16.11.3303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsui H-CT, Feng G, Winkler M. Negative regulation of mutS and mutH repair gene expression by the Hfq and RpoS global regulators of Escherichia coli K-12. J Bacteriol. 1997;179:7476–87. doi: 10.1128/jb.179.23.7476-7487.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veening JW, Smits WK, Kuipers OP. Bistability, epigenetics, and bet-hedging in bacteria. Annu Rev Microbiol. 2008;62:193–210. doi: 10.1146/annurev.micro.62.081307.163002. [DOI] [PubMed] [Google Scholar]
- Velasco-Miguel S, Richardson J, Gerlach V, Lai W, Gao T, et al. Constitutive and regulated expression of the mouse Dinb (Polκ) gene encoding DNA polymerase κ. DNA Repair. 2003;2:91–106. doi: 10.1016/s1568-7864(02)00189-1. [DOI] [PubMed] [Google Scholar]
- Vieira VC, Leonard B, White EA, Starrett GJ, Temiz NA, et al. Human papillomavirus E6 triggers upregulation of the antiviral and cancer genomic DNA deaminase APOBEC3B. mBio. 2014;5:e02234. doi: 10.1128/mBio.02234-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Waters J, Leung ML, Unruh A, Roh W, et al. Clonal evolution in breast cancer revealed by single nucleus genome sequencing. Nature. 2014;512:155–60. doi: 10.1038/nature13600. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitesell L, Santagata S, Mendillo ML, Lin NU, Proia DA, Lindquist S. HSP90 empowers evolution of resistance to hormonal therapy in human breast cancer models. PNAS. 2014;111:18297–302. doi: 10.1073/pnas.1421323111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wimberly H, Shee C, Thornton PC, Sivaramakrishnan P, Rosenberg SM, Hastings PJ. R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli. Nat Commun. 2013;4:2115. doi: 10.1038/ncomms3115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Witkin E. The radiation sensitivity of Escherichia coli B: a hypothesis relating filament formation and prophage induction. PNAS. 1967;57:1275–79. doi: 10.1073/pnas.57.5.1275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Witkin E, George D. Ultraviolet mutagenesis in polA and UvrA polA derivatives of Escherichia coli B/R: evidence for an inducible error-prone repair system. Genetics. 1973;73:91–108. [PubMed] [Google Scholar]
- Wu S, Powers S, Zhu W, Hannun YA. Substantial contribution of extrinsic risk factors to cancer development. Nature. 2016;529:43–47. doi: 10.1038/nature16166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yates LR, Campbell PJ. Evolution of the cancer genome. Nat Rev Genet. 2012;13:795–806. doi: 10.1038/nrg3317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu AM, McVey M. Synthesis-dependent microhomology-mediated end joining accounts for multiple types of repair junctions. Nucleic Acids Res. 2010;38:5706–17. doi: 10.1093/nar/gkq379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang Q, Lambert G, Liao D, Kim H, Robin K, et al. Acceleration of emergence of bacterial antibiotic resistance in connected microenvironments. Science. 2011;333:1764–67. doi: 10.1126/science.1208747. [DOI] [PubMed] [Google Scholar]