Abstract
Cellular barcodes are distinct DNA sequences that enable one to track specific cells across time or space. Recent advances in our ability to detect natural or synthetic cellular barcodes, paired with single-cell readouts of cell state, have markedly increased our knowledge of clonal dynamics and genealogies of the cells that compose a variety of tissues and organs. These advances hold promise to redefine our view of human disease. Here, we provide an overview of cellular barcoding approaches, discuss applications to gain new insights into disease mechanisms, and provide an outlook on future applications. We discuss unanticipated insights gained through barcoding in studies of cancer and blood cell production and describe how barcoding can be applied to a growing array of medical fields, particularly with the increasing recognition of clonal contributions in human diseases.
Tracing back the origins of tissues and organs has long been a fascination of developmental biologists and more recently has become an important component of medicine. Historically, lineage tracing has involved coating with or injection of dyes to follow cells as they contribute to embryos. In the modern era, techniques involving recombinases that drive fluorescent colors have elucidated precise populations of cells that give rise to their differentiated progeny. Recently, the next version of tracing cell origins has involved cellular barcoding, a technique that uses distinct DNA sequences to mark each cell and to follow the process of differentiation or self-renewal. Coupling this DNA marking with single-cell profiling technologies to measure gene expression or epigenetic marks has generated rich datasets that define the lineage tree and cell state of each differentiated cell in a tissue. Medicine has traditionally lacked the precision of lineage tracing to inform patient decisions. Even excellent animal models of disease could only examine populations of cells and tell which tissues are formed from a group of cells. Recently, cellular barcoding technologies have begun to provide the medical and biotechnology communities tools for enabling the reconstruction of high-resolution lineage trees and their application to clinically relevant questions.
Two medical fields have already begun lever-aging the insights of cellular barcoding studies: cancer and blood formation. For the oncologist, the concept of clones of cancer cells having distinct behaviors for tumor initiation, therapy resistance, and metastasis may help with treatment design (1). Currently, somatically acquired cancer driver mutations are used as a proxy for clonality (2, 3), the number of cells that participate in tissue growth. However, multiple clones and subclones can arise with the exact same oncogenic mutation, making it difficult to identify the “offending” clones. Now, cellular barcoding can examine a cancer, find the DNA barcode for many cells of the tumor, draw lineage trees to find the cell of origin, calculate the timing of that origin, examine growth rates of clones, and evaluate the transcriptional and chromatin state of each cell. Cellular barcoding has also been extensively applied to hematopoiesis, the process by which blood cells are made. The use of barcoding has begun to refine our understanding of the relative contributions of blood stem cells and progenitors to the maintenance of blood and immune cell production at steady state and under various stresses (4-8). Using barcoding to follow these stem cells and progenitors to examine the onset of blood cancers, such as myelodysplastic syndrome and leukemia, becomes possible (9-11). By combining lineage trees with gene expression or other functional readouts (12, 13), informed decisions can be made about when to treat patients and which drugs to give them. For example, therapy-resistant clones may harbor different expression signatures that separate them from the majority of cells in a tumor. Understanding such heterogeneity in cell states and behavior could lead to improved and more targeted therapeutic approaches.
In this Review, we provide a brief overview of cellular barcoding approaches that either take advantage of engineering in model systems or rely on the detection of naturally occurring somatic DNA changes, we discuss medical insights that have emerged from cellular barcoding studies, and we provide an outlook on further advances that will affect this field.
Cellular barcoding: An overview of technology and approaches
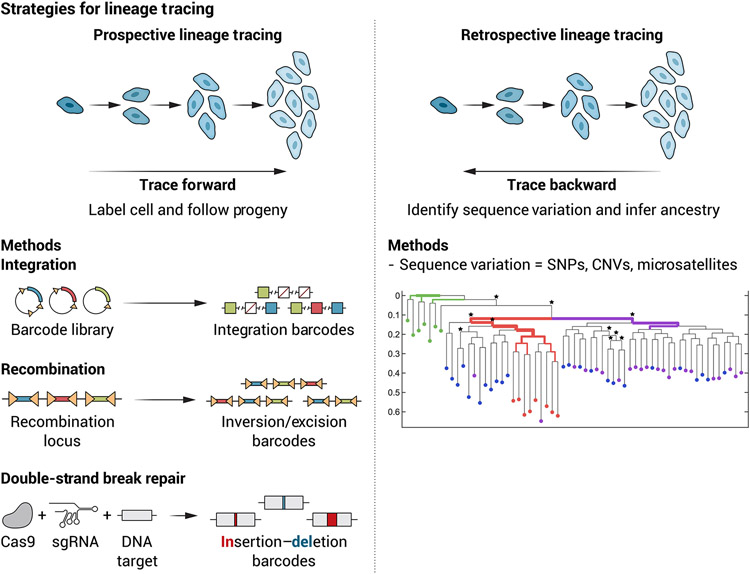
Barcoding techniques can determine the genealogy of related cells, providing a critical tool for exploring the phylogenetic principles of biological processes such as metazoan development, cancer progression and metastasis, and hematopoiesis (14-16). Historically, these efforts have often been categorized as “prospective” lineage tracing strategies, which involve labeling of individual cells (for example, with synthetic DNA sequence barcodes or fluorescent markers) and then following their descendants. Alternatively, “retrospective” lineage tracing strategies have been used and involve sequencing natural DNA variations that occur during the life of an organism (for example, single-nucleotide polymorphisms, copy-number variants, or short tandem repeats) (17, 18). Whereas prospective approaches can be used to study clonal dynamics in model systems (for example, the clonal progeny of a given cell type), retrospective approaches are particularly valuable for studying subclonal dynamics in patient-derived samples, such as elucidating which mutations contribute to tumor metastasis and when they occur (Fig. 1) (19-22). The resolution of such approaches has been limited by technical challenges, including sample heterogeneity (23, 24) and the slow rate and disperse nature of naturally occurring mutations. These technical limitations are now being mitigated by using single-cell resolution measurements or whole-genome sequencing (25-28).
Fig. 1. An overview of strategies for lineage tracing.
(Left) Some strategies for prospective lineage tracing. These approaches rely on engineered barcodes that are introduced through integration of the barcode into DNA, recombination of modified alleles that can evolve over time, or genome editing–mediated scar formation that can either occur at a single time point or evolve over time. This enables all the descendants of a particular marked cell to be tracked. (Right) The concept of retrospective lineage tracing, which uses the detection of natural barcodes in cells (such as single-nucleotide variants or other types of mutations) to track back and generate phylogenetic relationships between different cells. Such approaches can be made even more powerful by combining this barcoding with measurements of the cell state at single-cell resolution.
A key recent advance in retrospective lineage tracing has been the use of naturally occurring mutations of the mitochondrial genomes as an evolving barcode (11, 29). Critically, it is possible to exploit the high level of mitochondrial DNA, which is a natural byproduct of single-cell chromatin accessibility “ATAC-seq” (assay for transposase-accessible chromatin with high-throughput sequencing) approaches, to monitor the accumulation of mitochondrial mutations at single-cell resolution. This, together with the much more rapid (10- to 100-fold) rate of accumulation of mutations in the mitochondrial versus nuclear genomes, makes it possible to follow far more cells than is possible by using single-cell analysis of nuclear genome mutations. This approach allows for cellular barcoding of human cells without having to create a synthetic barcoding system and has been applied already to clonal tracking in human cancers. The current methods for mitochondrial DNA analyses are limited by the difficulty in accurately detecting mutations, particularly rarer variants, and by the challenges in linking the cell lineages with a detailed description of the cellular state (such as transcriptional or epigenomic states) (30, 31).
Some of the earliest contributions to the use of cellular barcodes encoded in DNA started with the classic analysis of hematopoietic stem cell clones after retroviral marking, which took advantage of each clone harboring a distinct retroviral insertion site in the genome (32). Such approaches have been advanced further more recently and provided an initial approach to assess the behavior of stem and progenitor cell populations (33, 34). These methods have even been applied to infer clonal dynamics after lentiviral gene therapy, providing key insights into clonal contributions to blood and immune reconstitution in these clinical settings (35). But these strategies also have limitations because they rely on a barcode being introduced at only a single time point upon retroviral insertion into the genome and cannot undergo further modification. These static barcodes can track clones but fail to provide further information as clones evolve.
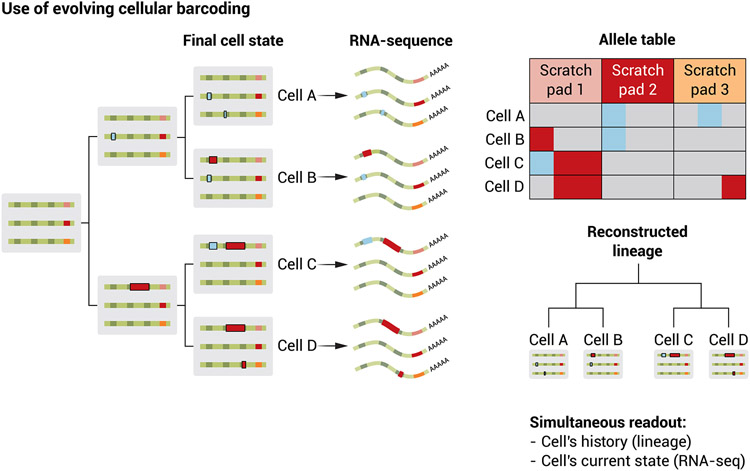
The recent development of Cas9-enabled lineage tracing techniques with single-cell RNA readouts (36-39) provides the potential to explore phylogenetic relationships in cancer progression and normal development at a vastly larger scale and finer resolution than was previously possible with methods involving static barcodes or fluorescent markers (33, 40-45). These newer methods most commonly rely on similar technical principles [reviewed in (15, 16)]. Briefly, Cas9 targets and cuts a defined, synthetic genomic locus introduced into a cell type of interest ( “scratch pad” or “target site”), resulting in a stable insertion-deletion (indel) allele that is inherited over subsequent generations. As cells divide, they accrue more Cas9-induced indels at additional sites that further distinguish successive clades of cells (Fig. 2) (46). At the end of the lineage tracing experiment, the indel alleles are collected from each individual cell by means of single-cell sequencing. Lineage information can be paired with single-cell gene expression profiles or other measures of the cell state to relate the phylogenetic history of a cell with its present state. Then, as in retrospective tracing approaches, various computational approaches (47, 48) can reconstruct a phylogenetic tree that best models subclonal cellular relationships (such as by maximum parsimony) from the observed alleles, although this remains a challenging problem for complex trees (49-52). By progressively introducing indel “scars” into the genomes of living cells by use of CRISPR-Cas9, it is possible to record in vivo and reconstruct high-depth lineage trees over months-long time scales that detail cellular parentage with single-cell resolution. Moreover, by pairing lineage information with the transcriptional state, it is possible to relate the phylogenetic history of a cell with its present state. Thus far, Cas9-enabled tracing has been successfully applied to study important aspects of metazoan biology, such as the cellular progenitor landscape in early mammalian embryogenesis (37, 53) and neural development in zebrafish (36, 54); the rates, routes, and drivers of tumor metastases (55); and the evolution of a tumor from initial acquisition of oncogenic mutations to formation of an aggressive and ultimately metastatic cancer (56).
Fig. 2. A simplified example of the use and value of evolving cellular barcoding.
Advances in cell barcoding now enable the use of evolving barcodes. In the simplified example shown, there are three independent integrated barcode regions that can each be modified at every cell division (for example, through genome editing–mediated scar formation). The combination of these different barcodes allows cellular hierarchies to be reconstructed. At the same time, other measurements of cell state can be obtained (such as the transcriptome or epigenome), providing rich information into the evolution of tissues in healthy and disease states.
Beyond simple barcoding information, the DNA scratch pad approach can in principle be used to develop multichannel molecular recorders that record critical information about the history of the cell [for example, the states a cell has passed through, whether a cell has been infected with a virus, or a cell’s three-dimensional (3D) environment]. Critically, such records can be made in an animal under normal physiological conditions or in disease states. Along this line, a number of strategies have been devised for recording in DNA scratch pads past transcriptional states of the cell (57, 58), including clever approaches that allow for recording the temporal order in which such cellular transcriptional programs occur (59, 60). As such approaches advance further, it will be interesting to assess to what extent past changes in cell state (such as the gene expression of a precursor) may contribute to alterations in lineage contributions in more differentiated cells (61). When such linkages are made, questions about how more recent cell states relate to earlier cell states can be addressed.
A key goal for lineage tracing efforts is to reconstruct the complete 3D cell fate map of a developing mammalian embryo analogous to the seminal studies of Caenorbhabditis elegans (62). In principle, the Cas9 DNA scratch pad approaches have the capacity to specifically mark every cell even in a complex animal such as a mouse. Moreover, current droplet-based (such as Chromium X) and split pool [such as Sci and SHARE-seq (simultaneous high-throughput ATAC and RNA expression with sequencing)] single-cell approaches now make it possible to routinely analyze millions of cells. However, the spatial information relating these cells is lost when using these approaches. Advances in spatial transcriptomic approaches may circumvent this problem, but achieving true single-cell resolution or preserving the spatial interactions between cells is currently not possible (63, 64). Parallel efforts by use of microscopy-based approaches hold great promise for simultaneous profiling of single-cell lineage and high-dimensional cellular information. For example, in the MEMOIR (memory by engineered mutagenesis with optical in situ readout) approach, information recorded in the DNA scratch pads is read out through microscopy by using fluorescent in situ hybridization (FISH), making it possible to visualize cell lineage relationships directly within their native tissue context (65, 66). However, the nature of DNA alterations that can be readily assayed by using current microscopy-based lineage tracing modalities limits the resolution of reconstructed trees and the time scales over which lineages can be recorded relative to what has been achieved with Cas9-based lineage tracing systems. In addition, FISH can currently only assess relatively small tissue sections, which limits applications.
Opportunities to gain insights into disease through barcoding
Cellular barcoding may have a major use in disease detection and prevention. Lineage has been brought to the forefront of medicine with the discovery that mutations occur in humans with clonal hematopoiesis during aging. Aging is also associated with TP53 mutant clones that lead to eyelid wrinkles (67), esophageal clones with NOTCH mutations (68), and colon clonal expansions (DNA mutations induced by a bacterial protein) (69, 70). Such changes are a natural process in aging, but the clonal expansions can predispose to disease. Following such expanding clones with cellular barcoding would allow for disease monitoring and could lead to earlier treatments with surgery or chemotherapy. Consider the current care of Barrett’s esophagus, which has a prevalence of 1.6% in the United States (71). Patients are followed with repeat biopsies to detect cancer, but currently, there are no lineage trees to predict the “offending” clones that could occur years before. Cellular barcoding would give the physician a precise detail of the activity of clones that lead to disease. Similarly, clonal hematopoiesis has been shown to be an inflammatory condition and associated with cardiovascular events (72). There is interest in diagnosing clones of cells that promote inflammation to be able to give individuals therapies that prevent the cardiovascular manifestations by identifying synthetic lethal vulnerabilities in the expanded clones. Other human disorders can be probed by using cellular barcoding. Somatic mosaicism accounts for substantial symptomatology. It is estimated that 2 to 3% of all Down syndrome cases are mosaic (73). Following patients by means of barcoding could establish genotypes associated with clone expansion that leads to pathophysiology.
Given that altered clonal architecture may underlie the etiology of a number of human diseases, there are emerging opportunities to study this process more deeply with both natural and synthetic barcoding to enable a higher-resolution view of disease pathology. Perhaps the clearest opportunity to apply such approaches is in the study of cancers, which are characterized by clonal evolutionary processes driven by both genetic and epigenetic alterations (74-76). Naturally occurring somatic mutations have been used for retrospective phylogenetic inference of the clonal dynamics in cancers (77,78). Although many of the somatic mutations that have been used in such studies may be drivers of the malignancy (79), some somatic changes—including passenger single-nucleotide variants, copy number alterations, and mitochondrial DNA mutations—have shown value as innocuous cell barcodes by which to reconstruct cancer clonal hierarchies (11, 80, 81). Such retrospective analyses have elucidated subclonal genetic and epigenetic alterations, factors contributing to metastatic spread of these diseases, and drivers of therapy resistance. Coupling such analyses with insights into transcriptional or epigenetic states has provided valuable insights into the drivers of clonal diversity, as exemplified by studies of human gliomas, leukemias, and cell lines that illuminate nongenetic drivers of clonal evolution and dominance (13, 82, 83).
Although the study of natural mutations has improved our knowledge of cancer evolution, many aspects of this process remain poorly understood and are challenging to validate experimentally beyond the observed associations. This has motivated the use of cell barcoding in tractable cancer model systems. Recent studies that used static barcoding of cancer cells have revealed how there can be substantial clonal diversity driven by nongenetic mechanisms that underlie properties observed in these cell populations, including subclonal therapy resistance and dominance (13, 41, 84). These findings highlight the potential insights that will likely emerge through studies of primary cancers with higher-resolution assessment of natural variants concomitant with cell state measurements, particularly if used over a span of time. In addition, the use of evolving engineered barcodes is providing a refined view of the clonal dynamics in disease, including in studies that have introduced these barcodes into cancer cell lines that are then transplanted into mice to track variable behavior in vivo (55, 85, 86) or through the use of engineered faithful cancer mouse models (56). For example, the use of evolving barcodes in a lung cancer mouse model has revealed how perturbation of additional tumor suppressors, including Lkb1 or Apc, can accelerate clonal evolution through distinct evolutionary mechanisms compared with what is seen in the absence of these perturbations (56).
Although the applications of cell barcoding in cancer have been in many ways a natural progression in a field that has been thinking about clonal evolution for decades (75), genome sequencing is providing clues on how clonal dynamics may have a broader scope of implications across a range of human tissues, even in conditions that are conventionally thought to be nonmalignant (87). For example, as individuals age, the hematopoietic system can be dominated by specific clones harboring somatic driver mutations that overlap with what is observed in blood cancers—a condition that is termed clonal hematopoiesis of indeterminate potential (CHIP) (72, 88). Although CHIP can predispose to acquisition of blood cancers, most individuals do not undergo such transformation. These observations have largely emerged from detection of somatic driver mutations in bulk populations of cells, but assessment of such mutations at the resolution of single cells will likely provide further critical insights into this process and how often such events can occur, as well as variation in properties between distinct clones (28). Cell barcoding in such states may also reveal cooperating pathways that are required to achieve clonal outgrowth, as illuminated through a recent study revealing how zebrafish asxl1 mutations that promote clonal hematopoiesis require resistance to inflammatory pathways to achieve clonal dominance (89). Interestingly, CHIP has been linked to a variety of age-related conditions beyond the risk for acquiring blood cancers, and higher-resolution studies of this process may provide an improved understanding of the mechanisms through which these risks emerge (90-94).
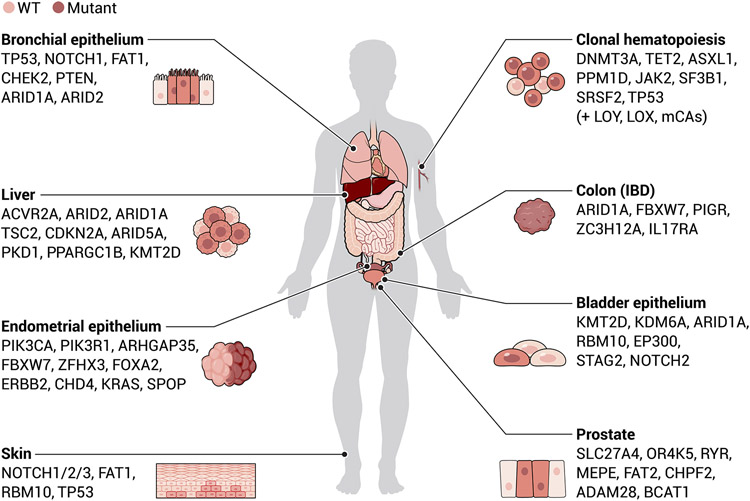
Clonal growths in nonmalignant states are not only a feature of the hematopoietic system, in which the measures of such states have been facilitated through the frequent sampling of blood, but also appear in a variety of other tissues (Fig. 3). Somatic driver mutations found in cancer have been identified in nonmalignant samples from the skin (95), bronchial epithelium (96), bladder (97), endometrium (98), prostate (99), colon (69), liver (100, 101), pancreas (102), and brain (103). In some cases, the expansion of such clones can be promoted in diseased states such as inflammatory bowel disease for the colon (104) or cirrhosis for the liver (100, 101). The precise risk of disease conferred by these clonal populations requires further study across this diverse group of tissues. Critically, such clonal populations that harbor somatic mutations may not always predispose to acquisition of a malignancy, and in some instances, these clonal expansions could prevent the outgrowth of malignant clones, as shown in the context of mouse models in the esophageal epithelium (105). Such findings may also be relevant in clinical scenarios, as suggested by individuals who have clonal hematopoiesis that involves DNMT3A mutations being less likely to have a disease relapse after blood stem cell transplantation (106). Further studies of these observations are needed to fully understand the mechanisms underlying the reduced risk of relapse.
Fig. 3. Nonmalignant clonal expansions can be found across a range of human tissues and organs.
A variety of human tissues and organs, including those highlighted here, have been shown to have age-related clonal expansions that are often driven by somatic mutations found in cancer. Examples of the affected tissues and some genes frequently mutated in these expansions are highlighted. The majority of these studies have relied on the use of bulk sequencing approaches, and with improved resolution of barcoding approaches, the implications of such clonal dynamics and its impact on disease will likely become clearer. IBD, inflammatory bowel disease.
Although somatic driver mutations can promote clonal outgrowths that are readily measured in bulk, single-cell monitoring of natural or synthetic barcodes provides an opportunity to gain a higher-resolution view of clonal dynamics that may contribute both to normal development and to disease predisposition. As our understanding of developmental processes continues to expand (25, 27, 107), we are learning more about how mosaic mutations acquired early during development could contribute to a range of human diseases, including cancer predisposition (108) and neuro-developmental disorders (109). Even in normal development in zebrafish, there appear to be examples of limited clonal contributions, as exemplified by studies of early heart development (110). However, in some ways this may only be the tip of the iceberg for understanding clonal contributions to development and disease. Although the somatic mutations detected may be drivers of or confer predisposition to disease, they could also serve as markers of clonality, and variation in this process itself may alter disease risk. For example, a range of estimates for clonal contributions to physiologic processes such as human hematopoiesis suggests that there may be tens to several hundred thousand distinct clones composing this process in healthy individuals (28, 111, 112). It is possible that a greater number of hematopoietic stem cell clones contribute to increased risks for blood cancers, as suggested through functional follow-up of genetic studies (113), or that a reduced amount of such stem cells could predispose to conditions such as aplastic anemia (114). Our ability to better detect natural barcodes or to engineer improved disease models with evolving barcodes will facilitate our knowledge of how disease arises and provide a higher-resolution view of this process. Moreover, although specific somatic changes such as T cell receptor rearrangements can facilitate assessment of clonal diversity in immune cell populations, the use of higher-resolution barcodes could enable the tracking of subclonal dynamics, particularly with findings emerging from therapeutic use of such cells (115).
Future innovations and application to diseases
There is clearly tremendous promise for cellular barcoding to enable a deeper understanding of human disease. Here, we highlight both technological innovations that will enable further insights as well as potential clinical applications that we envision will come to fruition in the future. There are some limitations of existing cellular barcoding approaches for both prospective and retrospective lineage tracing. Improvements in barcode detection, cell recovery, and avoidance of allelic dropout will enable higher-fidelity lineage reconstruction, which will be improved with advanced computational approaches that take advantage of the rich set of data generated (48, 50, 51). Increased throughput, which can be enabled through both droplet-based or split-pool approaches, will enable data to be generated on a greater diversity and larger number of cells and will enable more faithful lineage reconstruction in the context of disease. In particular, a major challenge relates to inferring lineage information from smaller clones that may be more poorly sampled by using existing methods, and with improved detection of barcodes in a greater number of cells, this issue can be overcome. The opportunity to integrate cellular barcodes with further data could illuminate the pathologic mechanisms that underlie disease. This includes the use of multichannel recorders, including those that enable temporal resolution, as well as multimodal analysis of cells (116).
We foresee there being two areas in which advances are likely to enable substantial progress. First, the development of spatial genomic analyses can be advanced by integrating barcode detection into these methods, including through adaptation of existing lineage tracing methods in single-cell spatial genomics or through the further development of microscopy-based approaches. This could not only enable insights into clonal hierarchies in disease contexts such as tumors but also enable the determination of how lineage relationships occur in 3D space. Such insights could refine therapeutic approaches, as exemplified by the use of traditional histology for determining tumor margins—an approach that could be improved and made more accurate through spatial genomic analyses. Second, we have focused almost entirely on cellular barcoding to elucidate cellular relationships in either primary disease samples or model systems. However, integration of perturbation approaches along with these cellular barcodes could enable insights into therapeutic targets or pathways that could alter the clonal composition in disease contexts (117). For example, if models of premalignant clonal states were analyzed with barcodes, this could enable newer regulators of clonal dominance to more accurately be identified and validated (89).
As these approaches advance further and as more data are generated, it is likely that limitations inherent to these approaches will become apparent. For example, the use of mitochondrial DNA mutations as natural cellular barcodes assumes that most mutations would not alter cellular metabolism or differentiation (11, 30). However, studies of specific mutations demonstrate how these variants are selected against in specific immune cell populations, including in T lymphocytes (118). As we obtain larger datasets, the impact of markers used as cellular barcodes may reveal that some of these do indeed affect the processes being studied, as has been clearly demonstrated through the study of somatic cancer driver mutations. Although we think that mutation rates may be constant in a cell and depend on cell division rates, recent studies have revealed how conditions such as inflammation can substantially alter the rate of mutation accrual in the nuclear genome (104) and have also revealed how nuclear mutations can arise in postmitotic tissues (119).
Beyond the technological advances, there are a number of highly promising clinical applications. At the moment, most cellular barcoding approaches are focused on helping to decipher the etiology and progression of disease. However, it may be possible to apply these approaches for prospective diagnostics. For example, if a patient was known to have a predisposition to acquiring cancer or had a premalignant condition, such diagnostics could be applied in a routine clinic visit to enable risk stratification or prediction that could tailor further clinical management on the patient’s likelihood of progressing to have a malignancy. Even in ongoing clinical studies, such as experimental gene therapy or genome editing trials, the malignancies observed in patients, such as those with sickle cell disease (120, 121), suggest a critical need to use these approaches both to study the basis for these cancers and to try to predict those patients who might develop such complications. From a therapeutic perspective, understanding subclonal properties of populations contributing to disease may enable the development of targeting approaches based on the concept of synthetic lethality that selectively target those clonal populations (for example, the drivers of clonal expansion may confer specific vulnerabilities to the dominant subclonal population).
The applications of cellular barcoding approaches in clinical medicine are likely to grow considerably, particularly with the increasing use of cellular therapies. For example, although clonal tracking of chimeric antigen receptor (CAR)–T cell clones has been valuable to detect durable persistence of this therapy for more than a decade (115), advances in molecular recording in the future could provide insights into the activities of these cellular therapies and enable the development of even more effective and targeted therapies in the future. In the setting of replacement of diseased epidermis by gene-modified cells, cellular barcodes from viral insertion demonstrated the limited contributions to regeneration by a subset of the stem cells introduced, suggesting opportunities for further clinical improvement as a result of insights from barcoding (122).
Conclusions
The advances in cellular barcoding in both primary human tissues and model systems offer an unprecedented opportunity to gain new insights into the mechanisms of disease. We have reviewed some advances in our understanding of cancer and hematopoiesis by dissecting the cellular hierarchies involved in these processes. Further applications of these approaches will undoubtedly advance a number of other medical fields, particularly with the growing recognition of clonal contributions in a diverse array of human diseases. Moreover, with further technological innovations, the opportunities to apply these approaches more broadly and prospectively in clinical medicine will continue to blossom. Barcoding provides us with a lens to define how clonal contributions can play a role in and help predict the outcome of disease.
ACKNOWLEDGMENTS
We thank members of the Sankaran, Weissman, and Zon laboratories, particularly C. Weng, J. D. Martin-Rufino, L. A. Liggett, L. Koblan, C. Baron, J. Henninger, and S. Avagyan, as well as A. Schier and C. Trapnell, for valuable comments and advice on this work.
Funding:
The work in V.G.S.’s laboratory is supported by the New York Stem Cell Foundation (NYSCF), a gift from the Lodish Family, the Edward P. Evans Foundation, the MPN Research Foundation, and National Institute of Health (NIH) grants R01 DK103794, R01 CA265726, and R01 HL146500. J.S.W.’s laboratory is supported by NIH grants 1RM1 HG009490-01 and 1U01 CA217882-01, the Milkyway Research Foundation, the Jameel Clinic at MIT, and the Ludwig Institute for Cancer Research. L.I.Z’s laboratory is supported by NIH grants P01HL131477, P01HL032262, U54DK110805, R24DK092760, R24OD017870, U01 HL134812, and R01HL144780-01; the Edward P. Evans Foundation; and the Alex’s Lemonade Stand Fund. V.G.S. is a NYSCF-Robertson Investigator. J.S.W. and L.I.Z. are investigators of the Howard Hughes Medical Institute.
Footnotes
Competing Interests: V.G.S. declares outside interest in Branch Biosciences, Ensoma, Novartis, Forma, Sana Biotechnology, and Cellarity. J.S.W. declares outside interest in 5 AM Ventures, Amgen, Chroma Medicine, KSQ Therapeutics, Maze Therapeutics, Tenaya Therapeutics, and Tessera Therapeutics. L.I.Z. declares outside interest in Branch Biosciences, Fate Therapeutics, CAMP4Therapeutics, and Scholar Rock.
REFERENCES AND NOTES
- 1.Kreso A et al. , Variable clonal repopulation dynamics influence chemotherapy response in colorectal cancer. Science 339, 543–548 (2013). doi: 10.1126/science.1227670 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Shlush LI et al. , Tracing the origins of relapse in acute myeloid leukaemia to stem cells. Nature 547, 104–108 (2017). doi: 10.1038/nature22993 [DOI] [PubMed] [Google Scholar]
- 3.Jan M et al. , Clonal evolution of preleukemic hematopoietic stem cells precedes human acute myeloid leukemia. Sci. Transl. Med 4, 149ra118 (2012). doi: 10.1126/scitranslmed.3004315 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Sun J et al. , Clonal dynamics of native haematopoiesis. Nature 514, 322–327 (2014). doi: 10.1038/nature13824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Pei W et al. , Resolving fates and single-cell transcriptomes of hematopoietic stem cell clones by PolyloxExpress barcoding. Cell Stem Cell 27, 383–395.e8 (2020). doi: 10.1016/j.stem.2020.07.018 [DOI] [PubMed] [Google Scholar]
- 6.Carrelha J et al. , Hierarchically related lineage-restricted fates of multipotent haematopoietic stem cells. Nature 554, 106–111 (2018). doi: 10.1038/nature25455 [DOI] [PubMed] [Google Scholar]
- 7.Rodriguez-Fraticelli AE et al. , Clonal analysis of lineage fate in native haematopoiesis. Nature 553, 212–216 (2018). doi: 10.1038/nature25168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Feng J et al. , Clonal lineage tracing reveals shared origin of conventional and plasmacytoid dendritic cells. Immunity 55, 405–422.e11 (2022). doi: 10.1016/j.immuni.2022.01.016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Van Egeren D et al. , Reconstructing the lineage histories and differentiation trajectories of individual cancer cells in myeloproliferative neoplasms. Cell Stem Cell 28, 514–523.e9 (2021). doi: 10.1016/j.stem.2021.02.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Williams N et al. , Life histories of myeloproliferative neoplasms inferred from phylogenies. Nature 602, 162–168 (2022). doi: 10.1038/s41586-021-04312-6 [DOI] [PubMed] [Google Scholar]
- 11.Ludwig LS et al. , Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell 176, 1325–1339.e22 (2019). doi: 10.1016/j.cell.2019.01.022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Umkehrer C et al. , Isolating live cell clones from barcoded populations using CRISPRa-inducible reporters. Nat. Biotechnol 39, 174–178 (2021). doi: 10.1038/s41587-020-0614-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Fennell KA et al. , Non-genetic determinants of malignant clonal fitness at single-cell resolution. Nature 601, 125–131 (2022). doi: 10.1038/s41586-021-04206-7 [DOI] [PubMed] [Google Scholar]
- 14.Stadler T, Pybus OG, Stumpf MPH, Phylodynamics for cell biologists. Science 371, eaah6266 (2021). doi: 10.1126/science.aah6266 [DOI] [PubMed] [Google Scholar]
- 15.Wagner DE, Klein AM, Lineage tracing meets single-cell omics: Opportunities and challenges. Nat. Rev. Genet 21, 410–427 (2020). doi: 10.1038/s41576-020-0223-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.VanHorn S, Morris SA, Next-generation lineage tracing and fate mapping to interrogate development. Dev. Cell 56, 7–21 (2021). doi: 10.1016/j.devcel.2020.10.021 [DOI] [PubMed] [Google Scholar]
- 17.Wu SS, Lee J-H, Koo B-K, Lineage tracing: Computational reconstruction goes beyond the limit of imaging. Mol. Cells 42, 104–112 (2019). [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tao L et al. , Retrospective cell lineage reconstruction in humans by using short tandem repeats. Cell Rep Methods 1, 100054 (2021). doi: 10.1016/j.crmeth.2021.100054 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Jamal-Hanjani M et al. , Tracking genomic cancer evolution for precision medicine: The lung TRACERx study. PLOS Biol. 12, e1001906 (2014). doi: 10.1371/journal.pbio.1001906 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hu Z et al. , Quantitative evidence for early metastatic seeding in colorectal cancer. Nat. Genet 51, 1113–1122 (2019). doi: 10.1038/s41588-019-0423-x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gerlinger M et al. , Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med 366, 883–892 (2012). doi: 10.1056/NEJMoa1113205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Shih DJH et al. , Genomic characterization of human brain metastases identifies drivers of metastatic lung adenocarcinoma. Nat. Genet 52, 371–377 (2020). doi: 10.1038/s41588-020-0592-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hong WS, Shpak M, Townsend JP, Inferring the origin of metastases from cancer phylogenies. Cancer Res. 75, 4021–4025 (2015). doi: 10.1158/0008-5472.CAN-15-1889 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Turajlic S, Swanton C, Metastasis as an evolutionary process. Science 352, 169–175 (2016). doi: 10.1126/science.aaf2784 [DOI] [PubMed] [Google Scholar]
- 25.Park S et al. , Clonal dynamics in early human embryogenesis inferred from somatic mutation. Nature 597, 393–397 (2021). doi: 10.1038/s41586-021-03786-8 [DOI] [PubMed] [Google Scholar]
- 26.Spencer Chapman M et al. , Lineage tracing of human development through somatic mutations. Nature 595, 85–90 (2021). doi: 10.1038/s41586-021-03548-6 [DOI] [PubMed] [Google Scholar]
- 27.Coorens THH et al. , Extensive phylogenies of human development inferred from somatic mutations. Nature 597, 387–392 (2021). doi: 10.1038/s41586-021-03790-y [DOI] [PubMed] [Google Scholar]
- 28.Mitchell E et al. , Clonal dynamics of haematopoiesis across the human lifespan. Nature 606, 343–350 (2022). doi: 10.1038/s41586-022-04786-y [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Xu J et al. , Single-cell lineage tracing by endogenous mutations enriched in transposase accessible mitochondrial DNA. eLife 8, e45105 (2019). doi: 10.7554/eLife.45105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lareau CA et al. , Massively parallel single-cell mitochondrial DNA genotyping and chromatin profiling. Nat. Biotechnol 39, 451–461 (2021). doi: 10.1038/s41587-020-0645-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Miller TE et al. , Mitochondrial variant enrichment from high-throughput single-cell RNA sequencing resolves clonal populations. Nat. Biotechnol 40, 1030–1034 (2022). doi: 10.1038/s41587-022-01210-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lemischka IR, Raulet DH, Mulligan RC, Developmental potential and dynamic behavior of hematopoietic stem cells. Cell 45, 917–927 (1986). doi: 10.1016/0092-8674(86)90566-0 [DOI] [PubMed] [Google Scholar]
- 33.Lu R, Neff NF, Quake SR, Weissman IL, Tracking single hematopoietic stem cells in vivo using high-throughput sequencing in conjunction with viral genetic barcoding. Nat. Biotechnol 29, 928–933 (2011). doi: 10.1038/nbt.1977 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lu R, Czechowicz A, Seita J, Jiang D, Weissman IL, Clonal-level lineage commitment pathways of hematopoietic stem cells in vivo. Proc. Natl. Acad. Sci. U.S.A 116, 1447–1456 (2019). doi: 10.1073/pnas.1801480116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Biasco L et al. , In vivo tracking of human hematopoiesis reveals patterns of clonal dynamics during early and steady-state reconstitution phases. Cell Stem Cell 19, 107–119 (2016). doi: 10.1016/j.stem.2016.04.016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Raj B et al. , Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nat. Biotechnol 36, 442–450 (2018). doi: 10.1038/nbt.4103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Chan MM et al. , Molecular recording of mammalian embryogenesis. Nature 570, 77–82 (2019). doi: 10.1038/s41586-019-1184-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Alemany A, Florescu M, Baron CS, Peterson-Maduro J, van Oudenaarden A, Whole-organism clone tracing using single-cell sequencing. Nature 556, 108–112 (2018). doi: 10.1038/nature25969 [DOI] [PubMed] [Google Scholar]
- 39.Spanjaard B et al. , Simultaneous lineage tracing and cell-type identification using CRISPR-Cas9-induced genetic scars. Nat. Biotechnol 36, 469–473 (2018). doi: 10.1038/nbt.4124 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Naik SH et al. , Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature 496, 229–232 (2013). doi: 10.1038/nature12013 [DOI] [PubMed] [Google Scholar]
- 41.Oren Y et al. , Cycling cancer persister cells arise from lineages with distinct programs. Nature 596, 576–582 (2021). doi: 10.1038/s41586-021-03796-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Biddy BA et al. , Single-cell mapping of lineage and identity in direct reprogramming. Nature 564, 219–224 (2018). ; doi: 10.1038/s41586-018-0744-4 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Weinreb C, Rodriguez-Fraticelli A, Camargo FD, Klein AM, Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science 367, eaaw3381 (2020). doi: 10.1126/science.aaw3381 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Livet J et al. , Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007). doi: 10.1038/nature06293 [DOI] [PubMed] [Google Scholar]
- 45.Snippert HJ et al. , Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010). doi: 10.1016/j.cell.2010.09.016 [DOI] [PubMed] [Google Scholar]
- 46.McKenna A et al. , Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science 353, aaf7907 (2016). doi: 10.1126/science.aaf7907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zafar H, Lin C, Bar-Joseph Z, Single-cell lineage tracing by integrating CRISPR-Cas9 mutations with transcriptomic data. Nat. Commun 11, 3055 (2020). doi: 10.1038/s41467-020-16821-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Jones MG et al. , Inference of single-cell phylogenies from lineage tracing data using Cassiopeia. Genome Biol. 21, 92 (2020). doi: 10.1186/s13059-020-02000-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Salvador-Martínez I, Grillo M, Averof M, Telford MJ, Is it possible to reconstruct an accurate cell lineage using CRISPR recorders? eLife 8, e40292 (2019). doi: 10.7554/eLife.40292 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Konno N et al. , Deep distributed computing to reconstruct extremely large lineage trees. Nat. Biotechnol 40, 566–575 (2022). doi: 10.1038/s41587-021-01111-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Wang S-W, Herriges MJ, Hurley K, Kotton DN, Klein AM, CoSpar identifies early cell fate biases from single-cell transcriptomic and lineage information. Nat. Biotechnol 40, 1066–1074 (2022). doi: 10.1038/s41587-022-01209-1 [DOI] [PubMed] [Google Scholar]
- 52.Kong W et al. , Capybara: A computational tool to measure cell identity and fate transitions. Cell Stem Cell 29, 635–649. e11 (2022). doi: 10.1016/j.stem.2022.03.001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kalhor R et al. , Developmental barcoding of whole mouse via homing CRISPR. Science 361, eaat9804 (2018). doi: 10.1126/science.aat9804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Raj B et al. , Emergence of neuronal diversity during vertebrate brain development. Neuron 108, 1058–1074.e6 (2020). doi: 10.1016/j.neuron.2020.09.023 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Quinn JJ et al. , Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts. Science 371, eabc1944 (2021). doi: 10.1126/science.abc1944 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Yang D et al. , Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell 185, 1905–1923. e25 (2022). doi: 10.1016/j.cell.2022.04.015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Tang W, Liu DR, Rewritable multi-event analog recording in bacterial and mammalian cells. Science 360, eaap8992 (2018). doi: 10.1126/science.aap8992 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Sheth RU, Wang HH, DNA-based memory devices for recording cellular events. Nat. Rev. Genet 19, 718–732 (2018). doi: 10.1038/s41576-018-0052-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Loveless TB et al. , Lineage tracing and analog recording in mammalian cells by single-site DNA writing. Nat. Chem. Biol 17, 739–747 (2021). doi: 10.1038/s41589-021-00769-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Choi J et al. , A time-resolved, multi-symbol molecular recorder via sequential genome editing. Nature 608, 98–107 (2022). doi: 10.1038/s41586-022-04922-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Packer JS et al. , A lineage-resolved molecular atlas of C. elegans embryogenesis at single-cell resolution. Science 365, eaax1971 (2019). doi: 10.1126/science.aax1971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Sulston JE, Schierenberg E, White JG, Thomson JN, The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev. Biol 100, 64–119 (1983). doi: 10.1016/0012-1606(83)90201-4 [DOI] [PubMed] [Google Scholar]
- 63.Rao A, Barkley D, França GS, Yanai I, Exploring tissue architecture using spatial transcriptomics. Nature 596, 211–220 (2021). doi: 10.1038/s41586-021-03634-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Srivatsan SR et al. , Embryo-scale, single-cell spatial transcriptomics. Science 373, 111–117 (2021). doi: 10.1126/science.abb9536 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Frieda KL et al. , Synthetic recording and in situ readout of lineage information in single cells. Nature 541, 107–111 (2017). doi: 10.1038/nature20777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Chow KK et al. , Imaging cell lineage with a synthetic digital recording system. Science 372, eabb3099 (2021). doi: 10.1126/science.abb3099 [DOI] [PubMed] [Google Scholar]
- 67.Jonason AS et al. , Frequent clones of p53-mutated keratinocytes in normal human skin. Proc. Natl. Acad. Sci. U.S.A 93, 14025–14029 (1996). doi: 10.1073/pnas.93.24.14025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Alcolea MP et al. , Differentiation imbalance in single oesophageal progenitor cells causes clonal immortalization and field change. Nat. Cell Biol 16, 615–622 (2014). doi: 10.1038/ncb2963 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Lee-Six H et al. , The landscape of somatic mutation in normal colorectal epithelial cells. Nature 574, 532–537 (2019). doi: 10.1038/s41586-019-1672-7 [DOI] [PubMed] [Google Scholar]
- 70.Pleguezuelos-Manzano C et al. , Mutational signature in colorectal cancer caused by genotoxic pks+ E. coli. Nature 580, 269–273 (2020). doi: 10.1038/s41586-020-2080-8 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Gilbert EW, Luna RA, Harrison VL, Hunter JG, Barrett’s esophagus: A review of the literature. J. Gastrointest. Surg 15, 708–718 (2011). doi: 10.1007/s11605-011-1485-y [DOI] [PubMed] [Google Scholar]
- 72.Bick AG et al. , Inherited causes of clonal haematopoiesis in 97,691 whole genomes. Nature 586, 763–768 (2020). doi: 10.1038/s41586-020-2819-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Shin M, Siffel C, Correa A, Survival of children with mosaic Down syndrome. Am. J. Med. Genet. A 152A, 800–801 (2010). doi: 10.1002/ajmg.a.33295 [DOI] [PubMed] [Google Scholar]
- 74.Marine J-C, Dawson S-J, Dawson MA, Non-genetic mechanisms of therapeutic resistance in cancer. Nat. Rev. Cancer 20, 743–756 (2020). doi: 10.1038/s41568-020-00302-4 [DOI] [PubMed] [Google Scholar]
- 75.Nowell PC, The clonal evolution of tumor cell populations. Science 194, 23–28 (1976). doi: 10.1126/science.959840 [DOI] [PubMed] [Google Scholar]
- 76.Prieto T, Landau DA, A heritable, non-genetic road to cancer evolution. Nature 601, 31–32 (2022). doi: 10.1038/d41586-021-03606-z [DOI] [PubMed] [Google Scholar]
- 77.Nangalia J, Campbell PJ, Genome sequencing during a patient’s journey through cancer. N. Engl. J. Med 381, 2145–2156 (2019). doi: 10.1056/NEJMra1910138 [DOI] [PubMed] [Google Scholar]
- 78.Cortés-Ciriano I, Gulhan DC, Lee JJ-K, Melloni GEM, Park PJ, Computational analysis of cancer genome sequencing data. Nat. Rev. Genet 23, 298–314 (2022). doi: 10.1038/s41576-021-00431-y [DOI] [PubMed] [Google Scholar]
- 79.Gerstung M et al. , The evolutionary history of 2,658 cancers. Nature 578, 122–128 (2020). doi: 10.1038/s41586-019-1907-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Gao R et al. , Delineating copy number and clonal substructure in human tumors from single-cell transcriptomes. Nat. Biotechnol 39, 599–608 (2021). doi: 10.1038/s41587-020-00795-2 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Schwartz R, Schäffer AA, The evolution of tumour phylogenetics: Principles and practice. Nat. Rev. Genet 18, 213–229 (2017). doi: 10.1038/nrg.2016.170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Chaligne R et al. , Epigenetic encoding, heritability and plasticity of glioma transcriptional cell states. Nat. Genet 53, 1469–1479 (2021). doi: 10.1038/s41588-021-00927-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Shaffer SM et al. , Memory Sequencing Reveals Heritable Single-Cell Gene Expression Programs Associated with Distinct Cellular Behaviors. Cell 182, 947–959.e17 (2020). doi: 10.1016/j.cell.2020.07.003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Goyal Y, Dardani IP, Busch GT, Emert B, Fingerman D, Kaur A, Jain N, Mellis IA, Li J, Kiani K, Fane ME, Weeraratna AT, Herlyn M, Raj A, Pre-determined diversity in resistant fates emerges from homogenous cells after anti-cancer drug treatment. bioRxiv 471833 [Preprint] (2021). doi: 10.1101/2021.12.08.471833 [DOI] [Google Scholar]
- 85.Zhang W et al. , The bone microenvironment invigorates metastatic seeds for further dissemination. Cell 184, 2471–2486.e20 (2021). doi: 10.1016/j.cell.2021.03.011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Simeonov KP et al. , Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states. Cancer Cell 39, 1150–1162.e9 (2021). doi: 10.1016/j.ccell.2021.05.005 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Mustjoki S, Young NS, Somatic mutations in “benign” disease. N. Engl. J. Med 384, 2039–2052 (2021). doi: 10.1056/NEJMra2101920 [DOI] [PubMed] [Google Scholar]
- 88.Jaiswal S, Ebert BL, Clonal hematopoiesis in human aging and disease. Science 366, eaan4673 (2019). doi: 10.1126/science.aan4673 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Avagyan S et al. , Resistance to inflammation underlies enhanced fitness in clonal hematopoiesis. Science 374, 768–772 (2021). doi: 10.1126/science.aba9304 [DOI] [PubMed] [Google Scholar]
- 90.Jaiswal S et al. , Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med 377, 111–121 (2017). doi: 10.1056/NEJMoa1701719 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Miller PG et al. , Association of clonal hematopoiesis with chronic obstructive pulmonary disease. Blood 139, 357–368 (2022). doi: 10.1182/blood.2021013531 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Kim PG et al. , Dnmt3a-mutated clonal hematopoiesis promotes osteoporosis. J. Exp. Med 218, e20211872 (2021). doi: 10.1084/jem.20211872 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Zekavat SM et al. , Hematopoietic mosaic chromosomal alterations increase the risk for diverse types of infection. Nat. Med 27, 1012–1024 (2021). doi: 10.1038/s41591-021-01371-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Bolton KL et al. , Clonal hematopoiesis is associated with risk of severe Covid-19. Nat. Commun 12, 5975 (2021). doi: 10.1038/s41467-021-26138-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Martincorena I et al. , Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015). doi: 10.1126/science.aaa6806 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Yoshida K et al. , Tobacco smoking and somatic mutations in human bronchial epithelium. Nature 578, 266–272 (2020). doi: 10.1038/s41586-020-1961-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 97.Lawson ARJ et al. , Extensive heterogeneity in somatic mutation and selection in the human bladder. Science; 370, 75–82 (2020). doi: 10.1126/science.aba8347 [DOI] [PubMed] [Google Scholar]
- 98.Moore L et al. , The mutational landscape of normal human endometrial epithelium. Nature 580, 640–646 (2020). doi: 10.1038/s41586-020-2214-z [DOI] [PubMed] [Google Scholar]
- 99.Cooper CS et al. , Analysis of the genetic phylogeny of multifocal prostate cancer identifies multiple independent clonal expansions in neoplastic and morphologically normal prostate tissue. Nat. Genet 47, 367–372 (2015). doi: 10.1038/ng.3221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Brunner SF et al. , Somatic mutations and clonal dynamics in healthy and cirrhotic human liver. Nature 574, 538–542 (2019). doi: 10.1038/s41586-019-1670-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Zhu M et al. , Somatic mutations increase hepatic clonal fitness and regeneration in chronic liver disease. Cell 177, 608–621.e12 (2019). doi: 10.1016/j.cell.2019.03.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Enge M et al. , Single-cell analysis of human pancreas reveals transcriptional signatures of aging and somatic mutation patterns. Cell 171, 321–330.e14 (2017). doi: 10.1016/j.cell.2017.09.004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Ganz J et al. , Rates and patterns of clonal oncogenic mutations in the normal human brain. Cancer Discov. 12, 172–185 (2022). doi: 10.1158/2159-8290.CD-21-0245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Olafsson S et al. , Somatic evolution in non-neoplastic IBD-affected colon. Cell 182, 672–684.e11 (2020). doi: 10.1016/j.cell.2020.06.036 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 105.Colom B et al. , Mutant clones in normal epithelium outcompete and eliminate emerging tumours. Nature 598, 510–514 (2021). doi: 10.1038/s41586-021-03965-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 106.Gibson CJ et al. , Donor clonal hematopoiesis and recipient outcomes after transplantation. J. Clin. Oncol 40, 189–201 (2022). doi: 10.1200/JCO.21.02286 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 107.Li R et al. , A body map of somatic mutagenesis in morphologically normal human tissues. Nature 597, 398–403 (2021). doi: 10.1038/s41586-021-03836-1 [DOI] [PubMed] [Google Scholar]
- 108.Pareja F et al. , Cancer-causative mutations occurring in early embryogenesis. Cancer Discov. 12, 949–957 (2022). doi: 10.1158/2159-8290.CD-21-1110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Bizzotto S, Walsh CA, Genetic mosaicism in the human brain: From lineage tracing to neuropsychiatric disorders. Nat. Rev. Neurosci 23, 275–286 (2022). doi: 10.1038/s41583-022-00572-x [DOI] [PubMed] [Google Scholar]
- 110.Gupta V, Poss KD, Clonally dominant cardiomyocytes direct heart morphogenesis. Nature 484, 479–484 (2012). doi: 10.1038/nature11045 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 111.Liggett LA, Sankaran VG, Unraveling hematopoiesis through the lens of genomics. Cell 182, 1384–1400 (2020). doi: 10.1016/j.cell.2020.08.030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 112.Lee-Six H et al. , Population dynamics of normal human blood inferred from somatic mutations. Nature 561, 473–478 (2018). doi: 10.1038/s41586-018-0497-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Bao EL et al. , Inherited myeloproliferative neoplasm risk affects haematopoietic stem cells. Nature 586, 769–775 (2020). doi: 10.1038/s41586-020-2786-7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 114.Young NS, Aplastic anemia. N. Engl. J. Med 379, 1643–1656 (2018). doi: 10.1056/NEJMra1413485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 115.Melenhorst JJ et al. , Decade-long leukaemia remissions with persistence of CD4+ CAR T cells. Nature 602, 503–509 (2022). doi: 10.1038/s41586-021-04390-6 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Nam AS, Chaligne R, Landau DA, Integrating genetic and non-genetic determinants of cancer evolution by single-cell multi-omics. Nat. Rev. Genet 22, 3–18 (2021). doi: 10.1038/s41576-020-0265-5 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 117.Tian L et al. , Clonal multi-omics reveals Bcor as a negative regulator of emergency dendritic cell development. Immunity 54, 1338–1351.e9 (2021). doi: 10.1016/j.immuni.2021.03.012 [DOI] [PubMed] [Google Scholar]
- 118.Walker MA et al. , Purifying selection against pathogenic mitochondrial DNA in human T cells. N. Engl. J. Med 383, 1556–1563 (2020). doi: 10.1056/NEJMoa2001265 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 119.Abascal F et al. , Somatic mutation landscapes at single-molecule resolution. Nature 593, 405–410 (2021). doi: 10.1038/s41586-021-03477-4 [DOI] [PubMed] [Google Scholar]
- 120.Liggett LA et al. , Clonal hematopoiesis in sickle cell disease. J. Clin. Invest 132, e156060 (2022). doi: 10.1172/JCI156060 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 121.Kaiser J, Gene therapy trials for sickle cell disease halted after two patients develop cancer. Science (2021). doi: 10.1126/science.abh1106 [DOI] [Google Scholar]
- 122.Hirsch T et al. , Regeneration of the entire human epidermis using transgenic stem cells. Nature 551, 327–332 (2017). doi: 10.1038/nature24487 [DOI] [PMC free article] [PubMed] [Google Scholar]